Bioinformatic Extraction of Functional Genetic Diversity from Heterogeneous Germplasm Collections for Crop Improvement
Abstract
1. Introduction
2. Material and Methods
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Vasudevan, K.; Vera Cruz, C.M.; Gruissem, W.; Bhullar, N.K. Large scale germplasm screening for identification of novel rice blast resistance sources. Front. Plant Sci. 2014, 5, 505. [Google Scholar] [CrossRef] [PubMed]
- Chen, A.; Sun, J.; Matthews, A.; Armas-Egas, L.; Chen, N.; Hamill, S.; Mintoff, S.; Tran-Nguyen, L.T.T.; Batley, J.; Aitken, E.A.B. Assessing variations in host resistance to Fusarium oxysporum f sp. cubense race 4 in Musa species, with a focus on the subtropical race 4. Front. Microbiol. 2019, 10, A1062. [Google Scholar] [CrossRef]
- Mansfield, B.D.; Mumm, R.H. Survey of plant density tolerance in U.S. maize germplasm. Crop. Sci. 2014, 54, 157–173. [Google Scholar] [CrossRef]
- Kurtz, B.; Gardner, C.A.C.; Millard, M.J.; Nickson, T.; Smith, J.S.C. Global access to maize germplasm provided by the US National Plant Germplasm System and by US plant breeders. Crop. Sci. 2016, 56, 931–941. [Google Scholar] [CrossRef]
- Mastrodomenico, A.T.; Hendrix, C.C.; Below, F.E. Nitrogen use efficiency and the genetic variation of maize expired Plant Variety Protection germplasm. Agriculture 2018, 8, 3. [Google Scholar] [CrossRef]
- Williams, J.W.; Jackson, S.T. Novel climates, no analog communities, and ecological surprises. Front. Ecol. Environ. 2007, 5, 475–482. [Google Scholar] [CrossRef]
- Murdock, T.Q.; Taylor, S.W.; Flower, A.; Mehlenbacher, A.; Montenegro, A.; Zwiers, F.W.; Alfaro, R.; Spittlehouse, D.L. Pest outbreak distribution and forest management impacts in a changing climate in British Columbia. Environ. Sci. Policy 2012, 26, 75–89. [Google Scholar] [CrossRef]
- Brown, A.H.D. Core collections: A practical approach to genetic resources management. Genome 1989, 31, 818–824. [Google Scholar] [CrossRef]
- Upadhyaya, H.D.; Pundir, R.P.S.; Dwivedi, S.L.; Gowda, C.L.L.; Reddy, V.G.; Singh, S. Developing a mini core collection of sorghum for diversified utilization of germplasm. Crop. Sci. 2009, 49, 1769–1780. [Google Scholar] [CrossRef]
- Tanksley, S.D.; McCouch, S.R. Seed banks and molecular maps: Unlocking genetic potential from the wild. Science 1997, 22, 1063–1066. [Google Scholar] [CrossRef]
- Lauter, N.; Doebley, J. Genetic variation for phenotypically invariant traits detected in teosinte: Implications for the evolution of novel forms. Genetics 2002, 160, 333–342. [Google Scholar] [PubMed]
- Le Rouzic, A.; Carlborg, Ö. Evolutionary potential of hidden genetic variation. Trends Ecol. Evolut. 2008, 23, 33–37. [Google Scholar] [CrossRef]
- Paaby, A.B.; Rockman, M.V. Cryptic genetic variation: Evolution’s hidden substrate. Nat. Rev. Genet. 2014, 15, 247–258. [Google Scholar] [CrossRef]
- McKhann, H.I.; Camilleri, C.; Bérard, A.; Bataillon, T.; David, J.L.; Reboud, X.; Le Corre, V.; Caloustian, C.; Gut, I.G.; Brunel, D. Nested core collections maximizing genetic diversity in Arabidopsis thaliana. Plant J. 2004, 38, 193–202. [Google Scholar] [CrossRef] [PubMed]
- Belaj, A.; Dominguez-Garcia, M.C.; Atienza, S.G.; Urdíroz, N.M.; De la Rosa, R.; Satovic, Z.; Martín, A.; Kilian, A.; Trujillo, I.; Valpuesta, V.; et al. Developing a core collection of olive (Olea europaea L.) based on molecular markers (DArTs, SSRs, SNPs) and agronomic traits. Tree Genet. Genomes 2012, 8, 365–378. [Google Scholar] [CrossRef]
- Gross, B.L.; Volk, G.M.; Richards, C.M.; Reeves, P.A.; Henk, A.D.; Forsline, P.L.; Szewc-McFadden, A.; Fazio, G.; Chao, C.T. Diversity captured in the USDA-ARS national plant germplasm system apple core collection. J. Amer. Soc. Hort. Sci. 2013, 138, 375–381. [Google Scholar] [CrossRef]
- Bataillon, T.M.; David, J.L.; Schoen, D.J. Neutral genetic markers and conservation genetics: Simulated germplasm collections. Genetics 1996, 144, 409–417. [Google Scholar]
- Reeves, P.A.; Panella, L.W.; Richards, C.M. Retention of agronomically important variation in germplasm core collections: Implications for allele mining. Theor. Appl. Genet. 2012, 124, 1155–1171. [Google Scholar] [CrossRef]
- Upadhyaya, H.D.; Bramel, P.J.; Singh, S. Development of a chickpea core subset using geographic distribution and quantitative traits. Crop. Sci. 2001, 41, 206–210. [Google Scholar] [CrossRef]
- Upadhyaya, H.D.; Gowda, C.L.L.; Pundir, R.P.S.; Reddy, V.G.; Singh, S. Development of core subset of finger millet germplasm using geographical origin and data on 14 quantitative traits. Genet. Resour. Crop. Evol. 2006, 53, 679–685. [Google Scholar] [CrossRef]
- Yan, W.; Rutger, J.N.; Bryant, R.J.; Bockelman, H.E.; Fjellstrom, R.G.; Chen, M.H.; Tai, T.H.; McClung, A.M. Development and evaluation of core subset of the USDA rice germplasm collection. Crop. Sci. 2007, 47, 869–878. [Google Scholar] [CrossRef]
- Parra-Quijano, M.; Iriondo, J.M.; Torres, E. Improving representativeness of genebank collections through species distribution models, gap analysis and ecogeographical maps. Biodivers Conserv. 2012, 21, 79–96. [Google Scholar] [CrossRef]
- Reeves, P.A.; Richards, C.M. Capturing haplotypes in germplasm core collections using bioinformatics. Genet. Resour. Crop. Evol. 2017, 64, 1821–1828. [Google Scholar] [CrossRef]
- Reeves, P.A.; Richards, C.M. Biases induced by using geography and environment to guide ex situ conservation. Conserv. Genet. 2018, 19, 1281–1293. [Google Scholar] [CrossRef]
- Mascher, M.; Schreiber, M.; Scholz, U.; Graner, A.; Reif, J.C.; Stein, N. Genebank genomics bridges the gap between the conservation of crop diversity and plant breeding. Nat. Genet. 2019, 51, 1076–1081. [Google Scholar] [CrossRef]
- Lasky, J.R.; Des Marais, D.L.; McKay, J.K.; Richards, J.H.; Juenger, T.E.; Keitt, T.H. Characterizing genomic variation of Arabidopsis thaliana: The roles of geography and climate. Mol. Ecol. 2012, 21, 5512–5529. [Google Scholar] [CrossRef]
- Geraldes, A.; Farzaneh, N.; Grassa, C.J.; McKown, A.D.; Guy, R.D.; Mansfield, S.D.; Douglas, C.J.; Cronk, Q.C.B. Landscape genomics of Populus trichocarpa: The role of hybridization, limited gene flow, and natural selection in shaping patterns of population structure. Evolution 2014, 68, 3260–3280. [Google Scholar] [CrossRef]
- Lasky, J.R.; Upadhyaya, H.D.; Ramu, P.; Deshpande, S.; Hash, C.T.; Bonnette, J.; Juenger, T.E.; Hyma, K.; Acharya, C.; Mitchell, S.E.; et al. Genome environment associations in sorghum landraces predict adaptive traits. Sci. Adv. 2015, 1, e1400218. [Google Scholar] [CrossRef]
- Scheet, P.; Stephens, M. A fast and flexible statistical model for large scale population genotype data: Applications to inferring missing genotypes and haplotypic phase. Am. J. Hum. Genet. 2006, 78, 629–644. [Google Scholar] [CrossRef]
- Xu, Z.; Zhang, D.; Hu, J.; Zhou, X.; Ye, X.; Reichel, K.L.; Stewart, N.R.; Syrenne, R.D.; Yang, X.; Gao, P.; et al. Comparative genome analysis of lignin biosynthesis gene families across the plant kingdom. BMC Bioinformatics 2009, 10 (Suppl. S11), S3. [Google Scholar] [CrossRef]
- Schlenker, W.; Roberts, M.J. Nonlinear temperature effects indicate severe damages to U.S. crop yields under climate change. Proc. Nat. Acad. Sci. USA 2009, 106, 15594–15598. [Google Scholar] [CrossRef] [PubMed]
- Lobell, D.B.; Roberts, M.J.; Schlenker, W.; Braun, N.; Little, B.B.; Rejesus, R.M.; Hammer, G.L. Greater sensitivity to drought accompanies maize yield increase in the U.S. Midwest. Sci. 2014, 344, 516–519. [Google Scholar] [CrossRef] [PubMed]
- Walthall, C.L.; Hatfield, J.; Backlund, P.; Lengnick, L.; Marshall, E.; Walsh, M.; Adkins, S.; Aillery, M.; Ainsworth, E.A.; Ammann, C.; et al. Climate Change and Agriculture in the United States: Effects and Adaptation. USDA Technical Bulletin 1935; USDA: Washington, DC, USA, 2012.
- Deerwester, S.; Dumais, S.T.; Furnas, G.W.; Landauer, T.K.; Harshman, R. Indexing by latent semantic analysis. J. Am. Soc. Inf. Sci. 1990, 41, 391–407. [Google Scholar] [CrossRef]
- Landauer, T.K.; Foltz, P.W.; Laham, D. Introduction to latent semantic analysis. Discourse Process. 1998, 25, 259–284. [Google Scholar] [CrossRef]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine learning in Python. J. Mach. Learn Res. 2011, 12, 2825–2830. [Google Scholar]
- Hamblin, M.T.; Salas Fernandez, M.G.; Casa, A.M.; Mitchell, S.E.; Paterson, A.H.; Kresovich, S. Equilibrium processes cannot explain high levels of short- and medium-range linkage disequilibrium in the domesticated grass Sorghum bicolor. Genetics 2005, 171, 1247–1256. [Google Scholar] [CrossRef]
- Van Treuren, R.; van Hintum, T.J.L. Marker assisted reduction of redundancy in germplasm collections: Genetic and economic aspects. Acta. Hort. 2003, 623, 139–149. [Google Scholar] [CrossRef]
- Singh, N.; Wu, S.; Raupp, W.J.; Sehgal, S.; Arora, S.; Tiwari, V.; Vikram, P.; Singh, S.; Chhuneja, P.; Gill, B.S.; et al. Efficient curation of genebanks using next generation sequencing reveals substantial duplication of germplasm accessions. Sci. Rep. 2019, 9, 650. [Google Scholar] [CrossRef]
- Milner, S.G.; Jost, M.; Taketa, S.; Mazón, E.R.; Himmelbach, A.; Oppermann, M.; Weise, S.; Knüpffer, H.; Basterrechea, M.; König, P.; et al. Genebank genomics highlights the diversity of a global barley collection. Nat. Genet. 2019, 51, 319–326. [Google Scholar] [CrossRef]
- Hu, Z.; Olatoye, M.O.; Marla, S.; Morris, G.P. An integrated genotyping by sequencing polymorphism map for over 10,000 sorghum genotypes. Plant Genome 2019, 12, 180044. [Google Scholar] [CrossRef]
- Wang, W.; Mauleon, R.; Hu, Z.; Chebotarov, D.; Tai, S.; Wu, Z.; Li, M.; Zheng, T.; Fuentes, R.R.; Zhang, F.; et al. Genomic variation in 3,010 diverse accessions of Asian cultivated rice. Nature 2018, 557, 43–49. [Google Scholar] [CrossRef] [PubMed]
- Oksman-Caldentey, K.M.; Saito, K. Integrating genomics and metabolomics for engineering plant metabolic pathways. Curr. Opin. Biotechnol. 2005, 16, 174–179. [Google Scholar] [CrossRef]
- Jupe, F.; Witek, K.; Verweij, W.; Sliwka, J.; Pritchard, L.; Etherington, G.J.; Maclean, D.; Cock, P.J.; Leggett, R.M.; Bryan, G.J.; et al. Resistance gene enrichment sequencing (RenSeq) enables reannotation of the NB-LRR gene family from sequenced plant genomes and rapid mapping of resistance loci in segregating populations. Plant J. 2013, 76, 530–544. [Google Scholar] [CrossRef] [PubMed]
- Valliyodan, B.; Nguyen, H.T. Understanding regulatory networks and engineering for enhanced drought tolerance in plants. Curr. Opin. Plant Biol. 2006, 9, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Loqué, D.; Scheller, H.V.; Pauly, M. Engineering of plant cell walls for enhanced biofuel production. Curr. Opin. Plant Biol. 2015, 25, 151–161. [Google Scholar] [CrossRef] [PubMed]
- Ort, D.R.; Merchant, S.S.; Alric, J.; Barkan, A.; Blankenship, R.E.; Bock, R.; Croce, R.; Hanson, M.R.; Hibberd, J.M.; Long, S.P.; et al. Redesigning photosynthesis to sustainably meet global food and bioenergy demand. Proc. Natl. Acad. Sci. USA 2015, 28, 8529–8536. [Google Scholar] [CrossRef] [PubMed]
- Wallace, D.H.; Ozbun, J.L.; Munger, H.M. Physiological genetics of crop yield. Adv. Agron. 1972, 24, 97–146. [Google Scholar]
- Gur, A.; Zamir, D. Unused natural variation can lift yield barriers in plant breeding. PLoS Biol. 2004, 2, e245. [Google Scholar] [CrossRef]
- Ashraf, M.; Harris, P.J.C. Abiotic Stresses. Plant Resistance Through Breeding and Molecular Approaches; Haworth Press: New York, NY, USA, 2005. [Google Scholar]
- Das, G.; Rao, G.J.N. Molecular marker assisted gene stacking for biotic and abiotic stress resistance genes in an elite rice cultivar. Front. Plant Sci. 2015, 6, 698. [Google Scholar] [CrossRef]
- Ceccarelli, S.; Grando, S.; Maatougui, M.; Michael, M.; Slash, M.; Haghparast, R.; Rahmanian, M.; Taheri, A.; Al-Yassin, A.; Benbelkacem, A.; et al. Plant breeding and climate changes. J. Agric. Sci. 2010, 148, 627–637. [Google Scholar] [CrossRef]
- Hanson, J.O.; Rhodes, J.R.; Riginos, C.; Fuller, R.A. Environmental and geographic variables are effective surrogates for genetic variation in conservation planning. Proc. Nat. Acad. Sci. USA 2017, 114, 12755–12760. [Google Scholar] [CrossRef] [PubMed]
- Friedman, C.; Borlawsky, T.; Shagina, L.; Xing, H.R.; Lussier, Y.A. Bio Ontology and text: Bridging the modeling gap. Bioinformatics 2006, 22, 2421–2429. [Google Scholar] [CrossRef]
- Sam, L.T.; Mendonça, E.A.; Li, J.; Blake, J.; Friedman, C.; Lussier, Y.A. PhenoGO: An integrated resource for the multiscale mining of clinical and biological data. BMC Bioinform. 2009, 10, S8. [Google Scholar] [CrossRef] [PubMed]
- Doebley, J.; Lukens, L. Transcriptional regulators and the evolution of plant form. Plant Cell 1998, 10, 1075–1082. [Google Scholar] [CrossRef] [PubMed]
- Wray, G.A. The evolutionary significance of cis-regulatory mutations. Nat. Rev. Genet. 2007, 8, 206–216. [Google Scholar] [CrossRef] [PubMed]
- Carroll, S.B. Evo-devo and an expanding evolutionary synthesis: A genetic theory of morphological evolution. Cell 2008, 134, 25–36. [Google Scholar] [CrossRef]
- Cambria, E.; White, B. Jumping NLP curves: A review of natural language processing research. IEEE Comput. Intell. Mag. 2014, 9, 48–57. [Google Scholar] [CrossRef]
- Rodríguez-Leal, D.; Lemmon, Z.H.; Man, J.; Bartlett, M.E.; Lippman, Z.B. Engineering quantitative trait variation for crop improvement by genome editing. Cell 2017, 171, 470–480. [Google Scholar] [CrossRef]
- Lemmon, Z.H.; Reem, N.T.; Dalrymple, J.; Soyk, S.; Swartwood, K.E.; Rodriguez-Leal, D.; Van Eck, J.; Lippman, Z.B. Rapid improvement of domestication traits in an orphan crop by genome editing. Nat. Plants 2018, 4, 766–770. [Google Scholar] [CrossRef] [PubMed]
- Atwell, S.; Huang, Y.S.; Vilhjálmsson, B.J.; Willems, G.; Horton, M.; Li, Y.; Meng, D.; Platt, A.; Tarone, A.M.; Hu, T.T.; et al. Genome-wide association study of 107 phenotypes in Arabidopsis thaliana inbred lines. Nature 2010, 465, 627–631. [Google Scholar] [CrossRef]
- Golicz, A.A.; Batley, J.; Edwards, D. Towards plant pangenomics. Plant Biotechnol. J. 2016, 14, 1099–1105. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.; Gonda, I.; Sun, H.; Ma, Q.; Bao, K.; Tieman, D.M.; Burzynski-Chang, E.A.; Fish, T.L.; Stromberg, K.A.; Sacks, G.L.; et al. The tomato pan-genome uncovers new genes and a rare allele regulating fruit flavor. Nat. Genet. 2019, 51, 1044–1051. [Google Scholar] [CrossRef]
- Hübner, S.; Bercovich, N.; Todesco, M.; Mandel, J.R.; Odenheimer, J.; Ziegler, E.; Lee, J.S.; Baute, G.J.; Owens, G.L.; Grassa, C.J.; et al. Sunflower pan-genome analysis shows that hybridization altered gene content and disease resistance. Nat. Plants 2019, 5, 54–62. [Google Scholar] [CrossRef] [PubMed]
- Tao, Y.; Zhao, X.; Mace, E.; Henry, R.; Jordan, D. Exploring and exploiting pan-genomics for crop improvement. Mol. Plant 2019, 12, 156–169. [Google Scholar] [CrossRef] [PubMed]
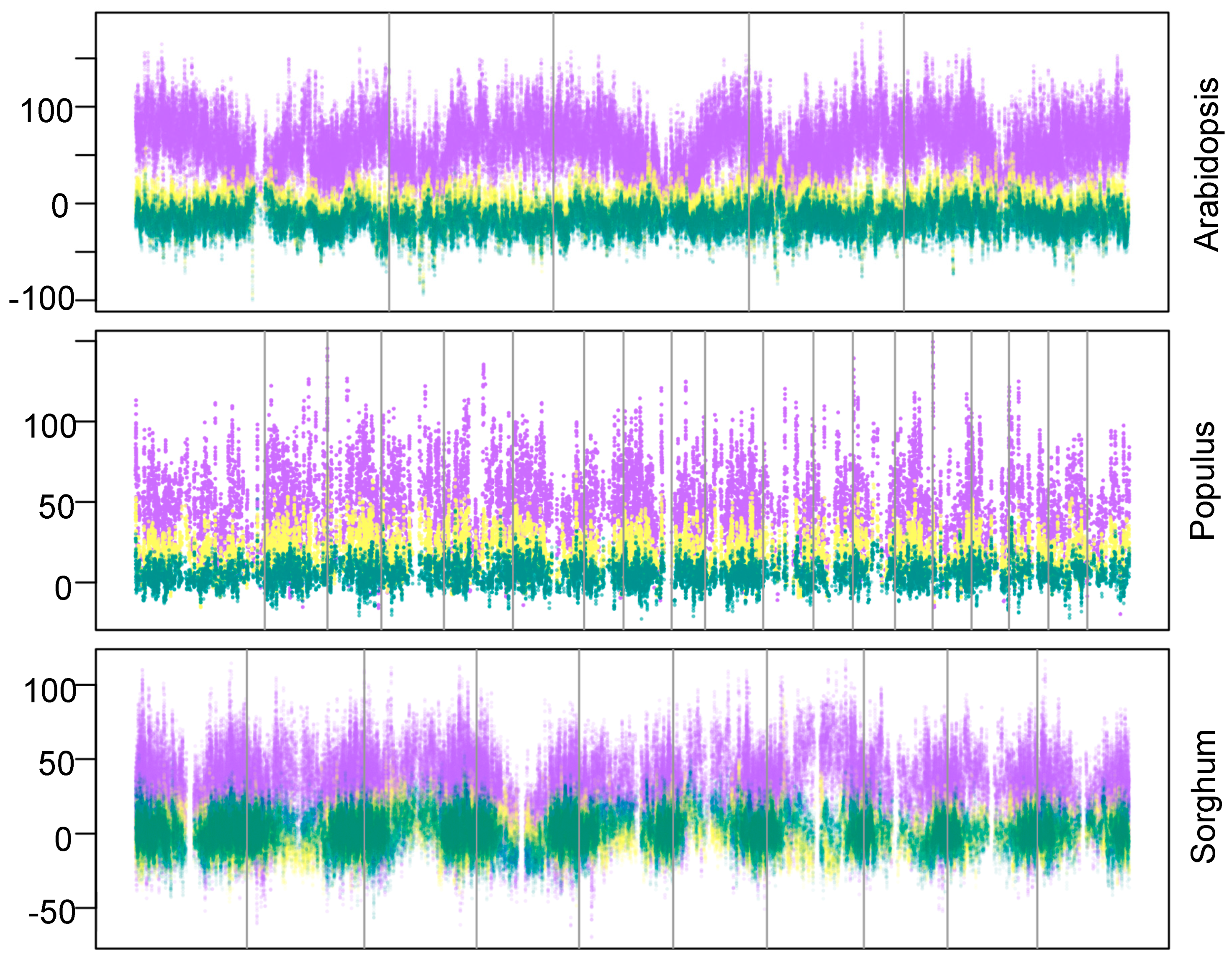
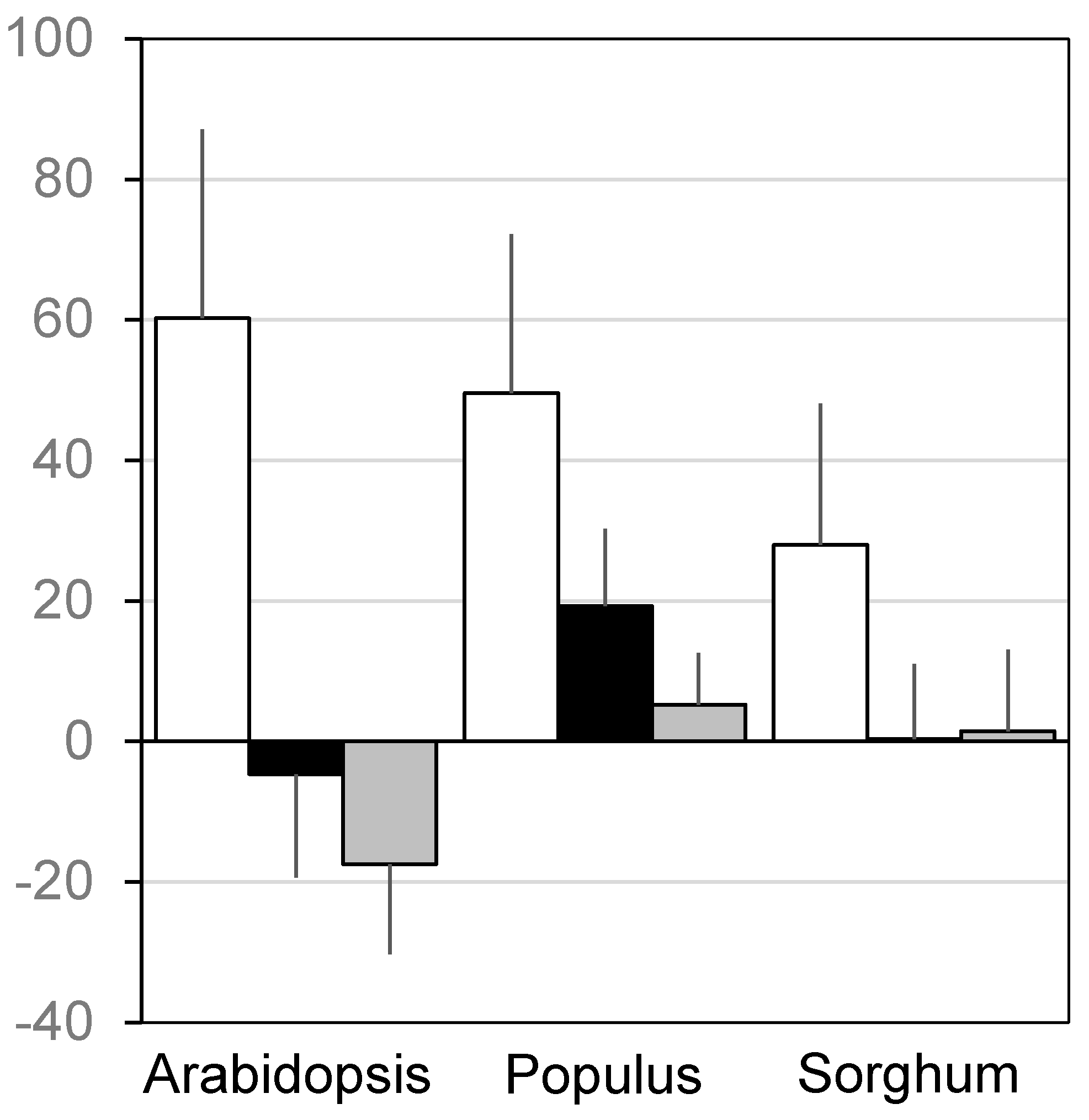
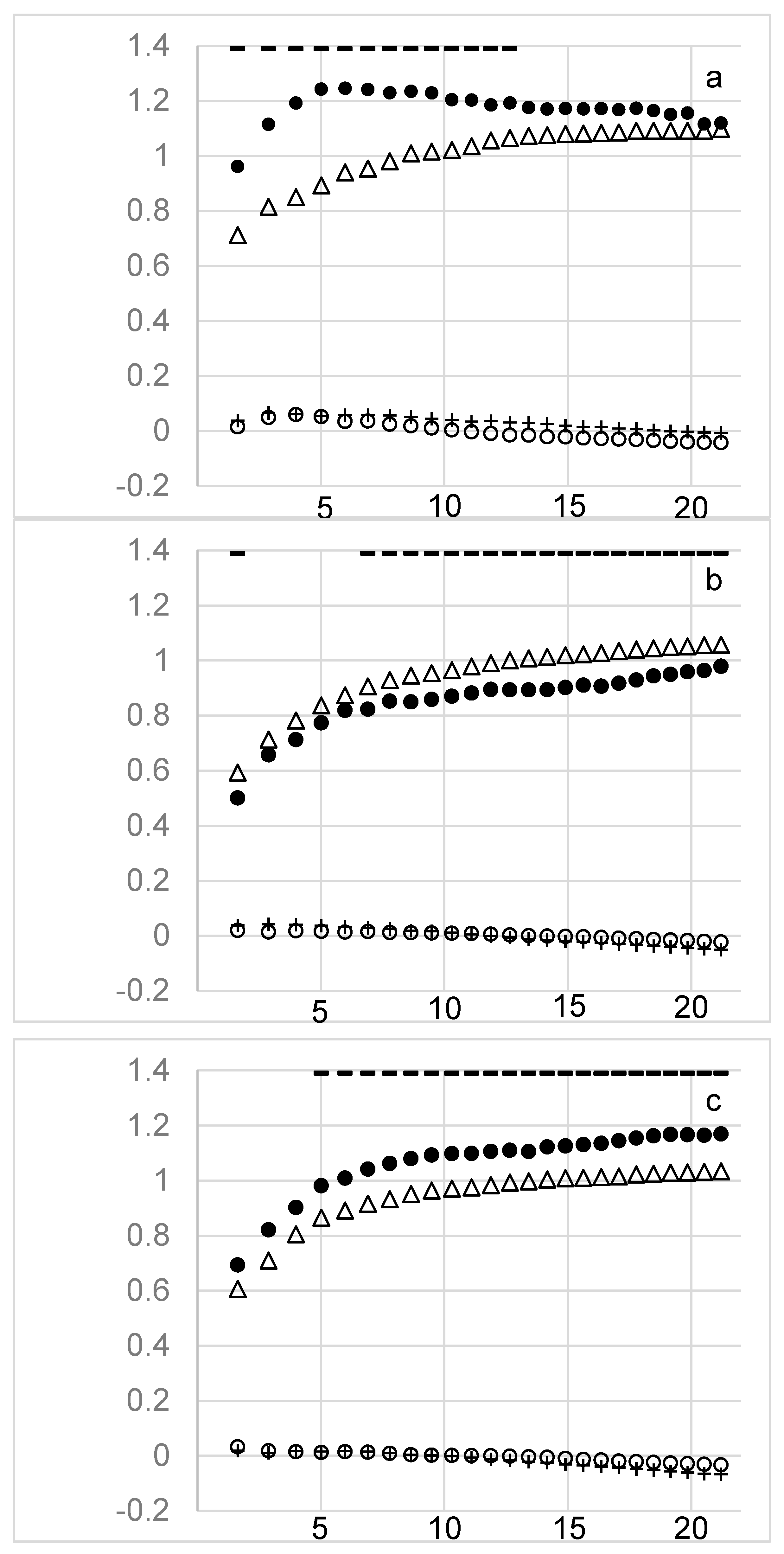
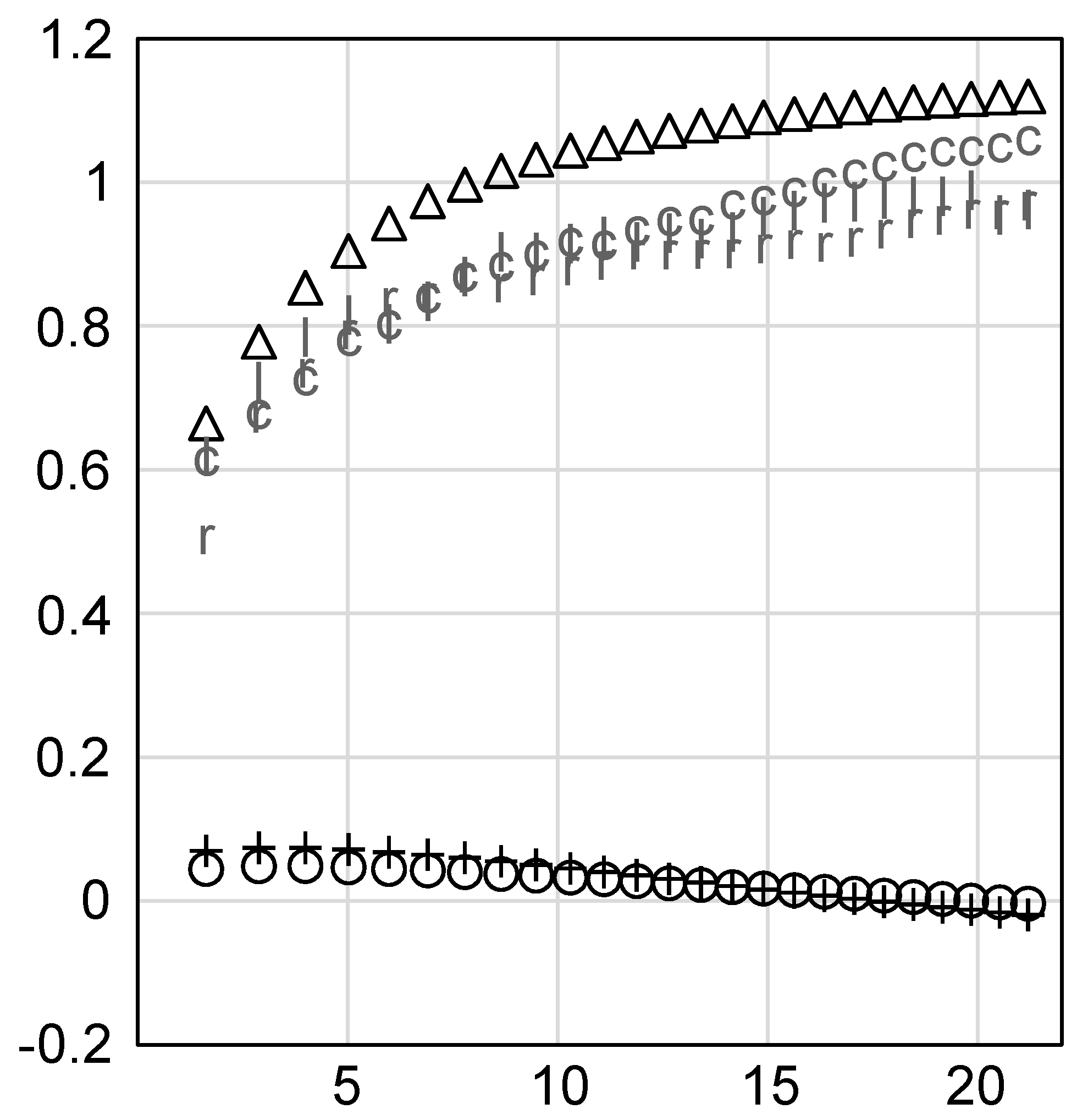

| ID | Term | ID | Term |
|---|---|---|---|
| GO:0010114 | response to red light | GO:0019740 | nitrogen utilization |
| GO:0010018 | far-red light signaling pathway | GO:0071481 | cellular response to X-ray |
| GO:0009408 | response to heat | GO:0016209 | antioxidant activity |
| GO:0015979 | photosynthesis | GO:0045087 | innate immune response |
| GO:0009409 | response to cold | GO:0080167 | response to karrikin |
| GO:0031990 | mRNA export from nucleus in response to heat stress | GO:0071480 | cellular response to gamma radiation |
| GO:0009640 | photomorphogenesis | GO:0019915 | lipid storage |
| GO:0009651 | response to salt stress | GO:0048316 | seed development |
| GO:0009555 | pollen development | GO:0006281 | DNA repair |
| GO:0006979 | response to oxidative stress | GO:0009411 | response to UV |
| GO:0034599 | cellular response to oxidative stress | GO:0032502 | developmental process |
| GO:0009266 | response to temperature stimulus | GO:0009646 | response to absence of light |
| GO:0042538 | hyperosmotic salinity response | GO:0043207 | response to external biotic stimulus |
| GO:0009631 | cold acclimation | GO:0006974 | cellular response to DNA damage stimulus |
| GO:0071470 | cellular response to osmotic stress | GO:0030332 | cyclin binding |
| GO:0006338 | chromatin remodeling | GO:0019760 | glucosinolate metabolic process |
| GO:0070483 | detection of hypoxia | GO:0009414 | response to water deprivation |
| GO:0050826 | response to freezing | GO:0019684 | photosynthesis, light reaction |
| GO:0048577 | negative regulation of short-day photoperiodism, flowering | GO:0016132 | brassinosteroid biosynthetic process |
| GO:0048364 | root development | GO:0016131 | brassinosteroid metabolic process |
| GO:0019748 | secondary metabolic process | GO:0015250 | water channel activity |
| GO:0043044 | ATP-dependent chromatin remodeling | GO:0006995 | cellular response to nitrogen starvation |
| GO:0042276 | error-prone translesion synthesis | GO:0009611 | response to wounding |
| GO:0030154 | cell differentiation | GO:0009793 | embryo development ending in seed dormancy |
| GO:0009908 | flower development | GO:0010014 | meristem initiation |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Reeves, P.A.; Tetreault, H.M.; Richards, C.M. Bioinformatic Extraction of Functional Genetic Diversity from Heterogeneous Germplasm Collections for Crop Improvement. Agronomy 2020, 10, 593. https://doi.org/10.3390/agronomy10040593
Reeves PA, Tetreault HM, Richards CM. Bioinformatic Extraction of Functional Genetic Diversity from Heterogeneous Germplasm Collections for Crop Improvement. Agronomy. 2020; 10(4):593. https://doi.org/10.3390/agronomy10040593
Chicago/Turabian StyleReeves, Patrick A., Hannah M. Tetreault, and Christopher M. Richards. 2020. "Bioinformatic Extraction of Functional Genetic Diversity from Heterogeneous Germplasm Collections for Crop Improvement" Agronomy 10, no. 4: 593. https://doi.org/10.3390/agronomy10040593
APA StyleReeves, P. A., Tetreault, H. M., & Richards, C. M. (2020). Bioinformatic Extraction of Functional Genetic Diversity from Heterogeneous Germplasm Collections for Crop Improvement. Agronomy, 10(4), 593. https://doi.org/10.3390/agronomy10040593





