Abstract
Despite the significant advances in the field of protein engineering, general design principles to improve organic cosolvent resistance of enzymes still remain undiscovered. Previous studies drew conclusions to engineer enzymes for their use in water-miscible organic solvents based on few amino acid substitutions. In this study, we conduct a comparison of a Bacillus subtilis lipase A (BSLA) library—covering the full natural diversity of single amino acid substitutions at all 181 positions of BSLA—with three state of the art random mutagenesis methods: error-prone PCR (epPCR) with low and high mutagenesis frequency (epPCR-low and high) as well as a transversion-enriched Sequence Saturation Mutagenesis (SeSaM-Tv P/P) method. Libraries were searched for amino acid substitutions that increase the enzyme’s resistance to the water-miscible organic cosolvents 1,4-dioxane (DOX), 2,2,2-trifluoroethanol (TFE), and dimethyl sulfoxide (DMSO). Our analysis revealed that 5%–11% of all possible single substitutions (BSLA site-saturation mutagenesis (SSM) library) contribute to improved cosolvent resistance. However, only a fraction of these substitutions (7%–12%) could be detected in the three random mutagenesis libraries. To our knowledge, this is the first study that quantifies the capability of these diversity generation methods generally employed in directed evolution campaigns and compares them to the entire natural diversity with a single substitution. Additionally, the investigation of the BSLA SSM library revealed only few common beneficial substitutions for all three cosolvents as well as the importance of introducing surface charges for organic cosolvent resistance—most likely due to a stronger attraction of water molecules.
1. Introduction
Conventional directed evolution studies typically report between 6 and 12 beneficial amino acid substitutions after screening of a few thousand clones [1]. However, this resembles only a small fraction of all potentially beneficial positions, which makes it difficult to deduce general design principles for the rational engineering of currently not systematically studied, complex properties. One such property is an enzyme’s resistance to organic solvents. The capability of routinely performing biocatalysis in organic solvents would be a groundbreaking improvement in chemical production processes, e.g., by directly applying hydrophobic substrates as solvents, simplifying downstream processing or shifting the reaction equilibrium into the desired direction [2,3,4,5,6], and thereby would unify the two quite separated worlds of catalysis (chemo- and biocatalysis). Given that the protein sequence space is huge (e.g., an enzyme with 100 amino acids can theoretically exist as 1.27 × 10130 natural variants), the existence of enzymes that perform well in organic solvents seems likely. As organic solvents rarely occur in most ecosystems, there is, if at all, only a very limited evolutionary pressure on enzymes to adapt to an optimum efficiency in their presence.
In aqueous solutions, enzymes typically are structurally stabilized by the interplay of hydrophobic interactions, hydrogen bonds, electrostatic interactions, van der Waals forces, and disulfide bridges [7,8,9,10]. Additionally, a key feature of an enzyme is the hydration shell that maintains both its flexibility and catalytic activity [11,12,13,14,15]. As non-natural solvents, organic solvents can remove the essential water layer and/or disrupt the above-mentioned interactions and therefore have a negative influence on enzyme structure, function, and solubility [3,5]. While hydrophobic, water-immiscible organic solvents are less detrimental and have even been reported to be consistent with proper enzyme folding and activity [16,17], experimental results and molecular dynamic simulations suggested that polar, water-miscible organic solvents tend to penetrate into the enzyme’s essential water layer or strip it off [13,18,19,20,21]. In addition to disturbing the interaction between the enzyme surface and surface-bound water, polar organic solvents can interfere with enzymatic activity directly in the active site: For instance, dimethyl sulfoxide (DMSO) was reported to act as an inhibitor by interacting with the catalytic heme iron in the active site of cytochrome P450 BM-3 monooxygenase from Bacillus megaterium [22,23,24].
Up to now, a multitude of studies has been reported that utilize protein engineering approaches to overcome the limited applicability of enzymes in organic cosolvents. Examples mainly include hydrolases and oxidoreductases, but also transferases and lyases. A more detailed overview on this topic was recently published by Stepankova et al. [25]. Furthermore, the analysis of the three-dimensional structure of 29 organic solvent-stable lipases revealed that surface charge can contribute to organic solvent resistance by exclusion of the organic solvent molecules from interacting with the enzyme’s surface [26]. Despite these successes, our limited knowledge of structure function relationships in the presence of organic cosolvents may yet guide future protein engineering campaigns through directed enzyme evolution, which remains the preferred choice to identify beneficial amino acid substitutions. However, directed evolution studies rarely report more than a dozen beneficial amino acid substitutions, which would be essential to deduce general design principles for the rational engineering of cosolvent resistance.
In a previous study, we described the statistical analysis of 3000 mutations in the lipase gene bsla, which were generated using three state of the art random mutagenesis methods (error-prone PCR with low and high mutagenesis frequency (epPCR-low and epPCR-high) and Sequence Saturation Mutagenesis (SeSaM-Tv P/P); 1000 mutations each) [27]. Furthermore, a systematic investigation of the same enzyme to ionic liquid resistance [28] and detergent tolerance [29] using a site-saturation mutagenesis (SSM) library (termed BSLA SSM library) [28] was reported. This BSLA SSM library covers the natural single substitution diversity of BSLA (in total 181 positions; i.e., all possible BSLA variants that can be generated by single amino acid substitutions). It was generated in 181 SSM experiments followed by extensive sequencing and site-directed mutagenesis (SDM) to ensure full coverage of all single substituted BSLA variants. The current study advances beyond the statistical analysis of individual libraries and compares the three random mutagenesis libraries to the entire possible single substitution diversity provided by the BSLA SSM library for their ability to promote organic cosolvent resistance. We chose to investigate BSLA resistance in three water-miscible organic cosolvents with different relative permittivities [30,31,32]: 1,4-dioxane (DOX; apolar), 2,2,2-trifluoroethanol (TFE; polar protic), and DMSO (polar aprotic). Hence, this approach addresses the following fundamental protein engineering questions: (1) What is the potential that natural single substitution diversity—represented by the BSLA SSM library—provides to improve organic cosolvent resistance? (2) How capable are state of the art random mutagenesis methods in exploring this potential? (3) Can we derive general design principles to improve organic cosolvent resistance and thus guide future directed enzyme evolution experiments?
2. Results
The results section displayed below is organized in six parts that cover the comparison of both library generation methods, the quantity and identity of the single substitutions, and their respective repercussions for the cosolvent resistance of the resulting enzymes. Section 2.1 analyzes the residual activity of wild type BSLA (BSLA WT) as a reference for the generated enzyme variants. The following Section 2.2 visualizes the beneficial single substitutions of BSLA for each cosolvent and identifies those substitutions that positively affect resistance to multiple or all three cosolvents. Section 2.3 quantifies the beneficial substitutions for each of the diversity generating methods and all cosolvents, whereas Section 2.4 categorizes and quantifies the beneficial substitutions into buried and surface-exposed ones. In Section 2.5, a visualization of common and unique beneficial substitutions for each cosolvent is provided. The final Section 2.6 delivers a deeper analysis into the identity of amino acids that influence cosolvent resistance.
2.1. Selection of Organic Cosolvent Concentrations for BSLA SSM Library Screening
In order to determine suitable conditions for the screening of the BSLA SSM library, each of the three water-miscible organic cosolvents (DOX, TFE, DMSO) was tested at different concentrations using BSLA WT. Cosolvent concentrations resulting in a residual BSLA WT activity of ~30% were chosen for the screening assay (see Table 1).

Table 1.
Organic cosolvent concentrations and residual activity of wild type Bacillus subtilis lipase A (BSLA WT) under screening conditions.
After screening of the BSLA SSM library in both the presence and absence of the given cosolvent concentration, the residual activity of each BSLA variant (RV) and BSLA WT (RWT) was determined. Beneficial variants were defined based on the true standard deviation as described in the Material and Methods section.
2.2. Overview of Beneficial Substitutions
In this study, beneficial substitutions are defined as amino acid exchanges leading to significantly higher residual activity of the BSLA variant compared to BSLA WT (defined in more detail in the Materials and Methods section). An overview of all beneficial amino acid substitutions that improve the resistance of BSLA to the three water-miscible organic cosolvents DOX, TFE, and DMSO is given in Figure 1. This allows the comparison between the full natural single substitution diversity—represented by the BSLA SSM library—and three previously generated random mutagenesis libraries (epPCR-low, epPCR-high, and SeSaM) (Figure 1a–c). Figure 1d provides an overview of all common beneficial substitutions for two or three of the cosolvents.
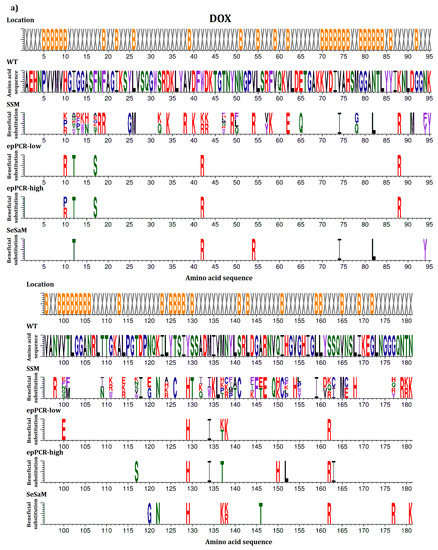
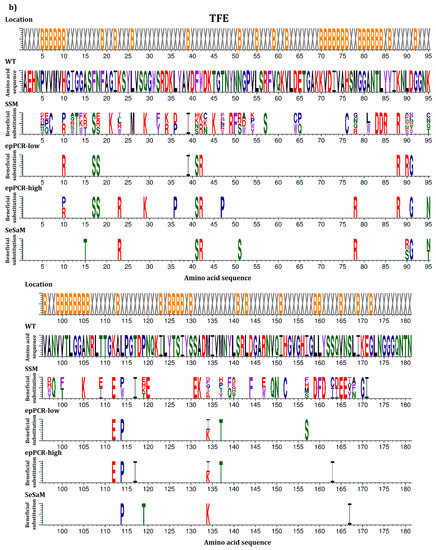
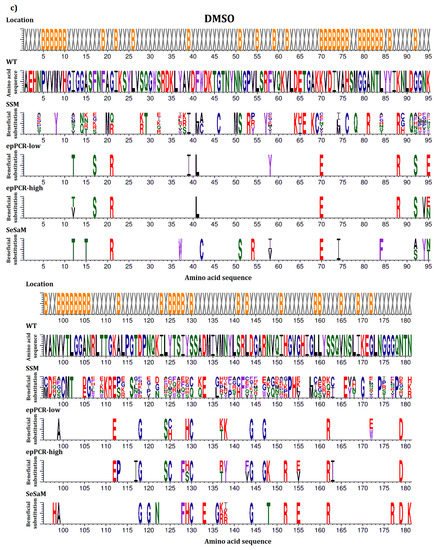
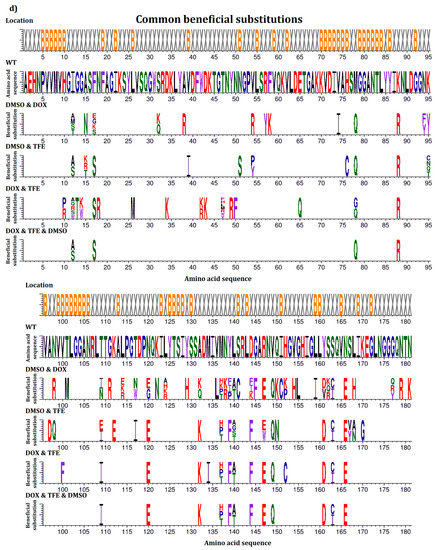
Figure 1.
Beneficial amino acid substitutions for improving the organic cosolvent resistance of BSLA to (a) DOX; (b) TFE and (c) DMSO. The BSLA WT sequence as well as beneficial substitutions found in the BSLA SSM library (SSM) and three random mutagenesis libraries (error-prone PCR with low and high mutagenesis frequency (epPCR-low and epPCR-high) and Sequence Saturation Mutagenesis (SeSaM)) are given. The figures are divided in two parts with BSLA amino acid positions 1–95 shown in the upper panel and 96–181 shown in the lower panel, respectively. Common beneficial substitutions (d) within the BSLA SSM library for improving resistance to more than one organic cosolvent are shown in different combinations (DMSO & DOX/DMSO & TFE/DOX & TFE/DOX, TFE, DMSO). Color-coding represents different amino acid categories as defined in the Material and Methods section: purple: aromatic amino acids (F, Y, W), black: hydrophobic amino acids (A, V, L, I, M), green: polar amino acids (S, T, N, Q), red: charged amino acids (D, E, H, K, R), blue: special amino acids (G, C, P). The location of the respective amino acids is indicated by X (surface exposed residue) and B (buried residue).
2.3. Number of Beneficial Amino Acid Substitutions
Table 2 summarizes the total number of beneficial amino acid substitutions that improved organic cosolvent resistance. Substitutions were identified after screening of the BSLA SSM library and three random mutagenesis libraries (epPCR-low; epPCR-high, SeSaM). The number of different beneficial substitutions after the screening of two and of all three random mutagenesis libraries is given to evaluate the complementarity and diversity of the three random mutagenesis methods.

Table 2.
Number of beneficial substitutions generated by BSLA SSM (site-saturation mutagenesis), epPCR-low, epPCR-high, SeSaM, and theoretical combinations.
The lowest number of beneficial substitutions was obtained for DOX, followed by TFE and DMSO. Five to eleven percent of all possible single substitutions (i.e., 159–371 of 3439 possible substitutions) contribute to improved cosolvent resistance. For DOX, the residual activity of BSLA was raised up to 76% by single amino acid substitution (Figure S1, Supplementary Materials), whereas for resistance to TFE and DMSO, the maximum residual activity was 85% and 76%, respectively. In the case of the three random mutagenesis methods, only 7%–12% of all beneficial substitutions identified by SSM were discovered. Screening of two random mutagenesis libraries increases the number of beneficial substitutions—especially when the SeSaM method is combined with any of the epPCR methods. The combination of two epPCR methods, however, only slightly increases the absolute number of beneficial substitutions. Combining the results obtained from all three random mutagenesis libraries results in the discovery of only 17%–18% of all beneficial substitutions. Similar trends were observed when looking only at the numbers of beneficial positions (see Table S1, Supplementary Materials). In contrast to beneficial substitutions, the term beneficial positions refers to positions with at least one beneficial amino acid substitution (i.e., a position with ≥2 different beneficial amino acid substitutions is counted as one beneficial position).
2.4. Location of Beneficial Amino Acid Substitutions in the BSLA 3D Structure
The beneficial amino acid substitutions were categorized into those located at the enzyme surface (termed exposed) or those buried by other amino acids as described in the Materials and Methods section (Table 3).

Table 3.
Location of beneficial amino acid substitutions within BSLA that improve organic cosolvent resistance to DOX, TFE, and DMSO.
In BSLA WT 71% of the amino acids are surface-exposed. In relation to this, 74%–87% beneficial substitutions detected in the BSLA SSM library were located in a surface-exposed position, with the side chain directed to the surrounding medium. When comparing the three random mutagenesis libraries to the full single substitution diversity, the location of beneficial substitutions obtained by epPCR-low, epPCR-high, and SeSaM differ only slightly for all three organic solvents. A very similar trend was observed when only the positions of beneficial substitutions were considered (i.e., a position with ≥2 different beneficial amino acid substitutions was counted as one beneficial position; see Table S2, Supplementary Materials).
2.5. Common Beneficial Substitutions for Different Organic Cosolvents
The entirety of beneficial BSLA substitutions provided by the BSLA SSM library causing resistance to each of the three tested cosolvents led us to ask which beneficial substitutions are common for several or even for all three cosolvents (Figure 2).
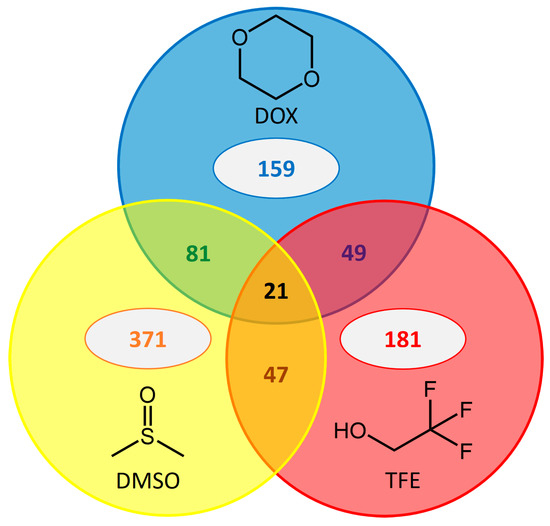
Figure 2.
Common beneficial substitutions in BSLA for three water-miscible organic solvents. The total number of beneficial substitutions for resistance to each of the three cosolvents is given in the center of the circles. The number of common beneficial substitutions for more than one cosolvent is given in the circles’ overlapping areas.
In total, 47–81 substitutions contribute to improved resistance against two cosolvents with the cosolvent pair DOX/DMSO sharing the highest number of beneficial substitutions. Twenty-one substitutions were found to be beneficial for resistance to all three cosolvents. A more detailed view considering the location of common beneficial substitutions (surface-exposed or buried; Table S3 in Supplementary Materials) revealed that in more than 92% of all cases, common beneficial substitutions were located at the enzyme’s surface. In particular, all of the 21 beneficial amino acid substitutions common for all three selected organic cosolvents were surface-exposed. A deeper investigation into the chemical nature of these common beneficial substitutions revealed that substitutions to charged and polar amino acids occurred most frequently at surface-exposed positions (Table S3 in Supplementary Materials). Due to the low number of common beneficial substitution at buried positions, no significant trend for any amino acid category was observable.
2.6. Occurrence of Beneficial Substitutions
A detailed analysis of the chemical characteristics of beneficial amino acids from each library (BSLA SSM library and the three random mutagenesis libraries) (Figure S2, Supplementary Materials) was conducted to gain a better picture of which beneficial amino acids were introduced most frequently by substitution. For DMSO, the five most frequent beneficial substitutions are R > K > E > Q > G, which corresponds to 37% of all generated beneficial substitutions. For TFE, 38% of beneficial substitutions are covered by R > K = W > F > D. Finally, for DOX, the five most beneficial substitutions are K > R > Q > F > I = H, which sum up to 52% of all beneficial substitutions. The primary type of beneficial amino acid substitutions are those to the positively charged amino acids (K and R; see Figure 3). Indeed, R is the predominant beneficial substitution for resistance to DOX in all three random mutagenesis libraries (see Figure S2; Supplementary Materials), which corresponds to the findings of the BSLA SSM library. Most of the aforementioned trends, however, cannot be observed in the three random mutagenesis libraries. Several of the most prominent beneficial substitutions of the BSLA SSM library were not obtainable by any of the three random mutagenesis methods (e.g., Q and F for DOX; W, F, and D for TFE; Q for DMSO; Figure S2, Supplementary Materials). Moreover, some substitutions that were frequent in the random mutagenesis libraries were not found among the five most prominent beneficial substitutions of the BSLA SSM library (e.g., T for DOX; S, I, and P for TFE; S, T for DMSO; Figure S2; Supplementary Materials). This illustrates that random mutagenesis methods fail in providing a representative picture of the full single substitution diversity to improve BSLA.
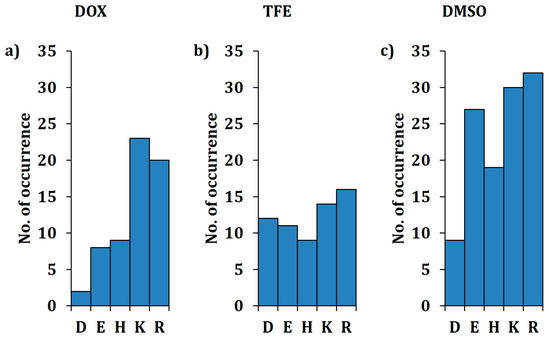
Figure 3.
Number of beneficial substitutions to charged amino acids for resistance to DOX (a), TFE (b) and DMSO (c). Numbers are based on the findings of the BSLA SSM library. Information about substitutions to all remaining amino acids is available in the Supplementary Materials (Figure S2).
In parallel to investigating the newly introduced amino acids, the amino acids of BSLA WT they were replacing were identified (Figure S3, Supplementary Materials). The five most frequently occurring amino acids within BSLA WT are G > V = N > L > S (48% of all amino acids). BSLA WT does not contain cysteine (Figure S4, Supplementary Materials). The BSLA SSM library results indicate that the amino acids G, N, V, and L are among the five most frequently replaced amino acids in BSLA mutants with improved resistance to DMSO as well as to DOX (Figure S3a, (i), Supplementary Materials). For TFE, the three amino acids L, V, and N were substituted most frequently (Figure S3e, Supplementary Materials).
BSLA shows a strong preference for specific amino acids to be replaced for improving organic cosolvent resistance. Among the variants with improved resistance to DMSO, 82% of all N in BSLA WT were replaced by a different amino acid. Interestingly, 80% of all R were also replaced, although R was also the primary beneficial amino acid substitution (Figure S2 (i), Supplementary Materials). Moreover, 75% of all F, A, and M were replaced as well. Among the improved variants to DOX, 75% of all F and M occurring in BSLA WT were replaced, while for resistance to TFE, 75% of all M were substituted. A comparably low frequency of replacement was observed in DMSO for P (25%), I (30%), and E (33%), in DOX for P (0%), Q (17%), and K (27%), and in TFE for G (17%), I (20%), and S (31%).
3. Discussion
Enzyme resistance to water-miscible organic solvents such as DOX, TFE, and DMSO clearly represents a desirable trait for many applications. Defining general design principles for the rational engineering of such properties, however, requires comprehensive knowledge about beneficial amino acid substitutions. Most directed evolution studies report the effects of few beneficial amino acid substitutions and omit the vast majority of possible but undetected beneficial amino acid exchanges. In this report, we compared a library containing all beneficial single amino acid exchanges of Bacillus subtilis lipase A (BSLA SSM library) to three state of the art random mutagenesis libraries (epPCR-low, epPCR-high, SeSaM) in order to answer the following three questions: (1) What is the potential that the total single substitution diversity represented by the BSLA SSM library provides to improve cosolvent resistance? (2) How capable are state of the art random mutagenesis methods in exploring this potential and (3) can we derive general design principles to improve cosolvent resistance to guide future directed enzyme evolution experiments?
3.1. Comparison of Multiple Mutagenesis Methods to Address Question (1) and (2)
Given that naturally occurring enzymes are adapted to environments occurring in nature (which usually do not contain organic cosolvents), it is not surprising that up to one in ten single amino acid substitutions in BSLA lead to improved organic cosolvent resistance. This ratio explains why beneficial substitutions were identified in previous directed evolution campaigns after the screening of only a few thousand clones [25,33]. However, our study clearly indicates that random mutagenesis methods fail to detect the vast majority (>88%) of the theoretically possible single substitution diversity that would be beneficial for resistance to the three selected organic cosolvents. The investigated random mutagenesis libraries (epPCR-low, -high, and SeSaM-Tv P/P) allow only the identification of a small fraction of all obtainable beneficial substitutions. Despite the fact that these shortcomings of random mutagenesis methods are commonly known [1,27] and accepted, this study is the first to quantify this phenomenon in the example of organic cosolvents. A more suitable approach to overcome the limitation of a single random mutagenesis method could be the combination of two different methods as shown in Table 2. The SeSaM-Tv P/P method was optimized to generate a complementary bias to the often transition-biased epPCR-based methods and to introduce subsequent mutations (>30%), which cannot be achieved by conventional epPCR approaches [27,34]. By performing different rounds of directed evolution utilizing epPCR and SeSaM-Tv P/P in alternating order, a larger pool of beneficial substitutions (up to 16% using the epPCR/SeSaM-Tv P/P combination vs. just up to 12% using epPCR only; Table 2) could be accessed. For instance, the combination of two different methods including epPCR has successfully proven to increase the stability of a lipase [35]. Nonetheless, even combining the diversity of all three random mutagenesis methods could increase the substitution rate to a maximum of 18% of all possible beneficial single substitutions (Table 2). This illustrates that state of the art random mutagenesis methods, as used in standard directed evolution campaigns, are quite limited in accessing the total single substitution diversity that nature offers.
3.2. General Design Principles Derived from the BSLA SSM Library to Address Question (3)
As a general trend, substitutions to charged amino acids comprise the highest fraction of beneficial amino acid substitutions in the BSLA SSM library. They are primarily located in surface-exposed positions, which is in accordance with previous findings stating that amino acid residues located at the protein surface mediate solvent resistance [25,35]. This is highlighted by the fact that all of the 21 common beneficial substitutions that improve resistance to DOX, TFE, and DMSO are located at the enzyme’s surface and mainly involve substitutions to charged and polar amino acids. This may imply that introducing charged residues could lead to the formation of new salt bridges that stabilize the enzyme in the presence of organic cosolvents. However, a detailed analysis of the enzyme’s three-dimensional structure indicates that this may not be the mechanism responsible for organic cosolvent resistance in BSLA. The loss of the salt bridge in BSLA WT between R107 and D144 in fact increased organic solvent resistance while simultaneously maintaining the enzyme’s stability in mutants of BSLA in this study. Beneficial substitutions at position D144 comprised mutations to F, W, I, G, T, and K for DMSO, to F, I, and K for DOX, and to F for TFE. None of these substitutions could sustain this salt bridge. In fact, the substitution D144K would even invert the charge at position 144. In silico introduction of the latter mutation using the FoldX method proposes that R107 compensates for the missing salt bridge by forming a hydrogen bond with the backbone of R142, which does not occur in BSLA WT. Therefore, both the charged side chains of R107 and the newly introduced K144 are available for other interactions. These might involve a stronger interaction of water with the enzyme’s surface. Water-miscible organic cosolvents tend to remove the enzyme’s essential water layer which renders it inactive [3,5,13]. For instance, it was reported that DMSO when applied as a cosolvent denatures a protein by solvation of non-polar amino acid side chains on the enzyme’s surface through methyl groups of the cosolvent [36]. Introduction of charged residues would assist in tightly binding water molecules to the enzyme. This would exclude cosolvent molecules from interacting with the surface and prevent the enzyme from denaturation and/or deactivation. In this respect, the observation made in this study is consistent with previous studies reporting the importance of introducing polar and charged amino acids [33,37,38] to increase resistance to water-miscible organic solvents. Similar trends were observed when the three-dimensional structures of 29 organic solvent-stable lipases were analyzed [26]. On the contrary, the replacement of surface charges by hydrophobic amino acids has been proposed previously to increase organic solvent resistance [2]. Indeed, a stability increase was reported for α-lytic protease in 84% dimethylformamide after surface charge substitution by hydrophobic amino acids [39]. However, Dror et al. illustrated that the introduction of a hydrophobic amino acid—while increasing an enzyme’s stability—simultaneously can reduce its activity in the presence of polar organic cosolvents [35]. A comprehensive structural analysis of the BSLA SSM library confirmed that in most cases a newly introduced, surface-exposed charge either (1) broke a preexisting salt bridge (see example above); (2) was introduced at a location distant from preexisting charges or (3) introduced a charge in proximity to an existing salt bridge. In all three cases, this leads to an increase of the local (positive or negative) charge on the enzyme’s surface. For all newly introduced charged residues capable of forming salt bridges, an additional analysis for other possible beneficial substitutions at the given position was conducted. In many of these cases, other beneficial substitutions incapable of forming salt bridges (e.g., substitutions to aromatic or hydrophobic amino acids) were detected (Figure 4).
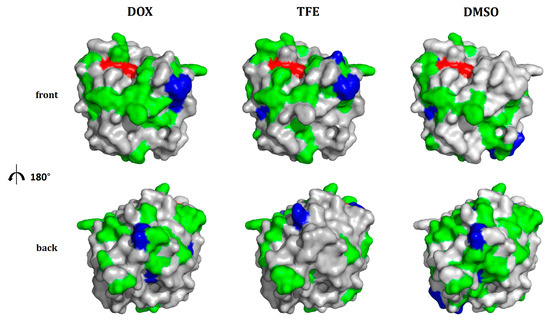
Figure 4.
Analysis of newly introduced charged residues for the formation of salt bridges. All beneficial substitutions to surface-exposed, charged amino acids found in the BSLA SSM library were analyzed for their ability to form salt bridges as described in the Materials and Methods section. Surface color represents the results of this analysis: red: catalytic triad residues (S77, D133, H156); blue: substitutions to charged amino acids that most likely lead to the formation of a salt bridge; green: remaining substitutions to charged amino acids (this includes charged amino acids that are too distant from oppositely charged amino acids to form salt bridges and positions at which substitutions to amino acids other than charged ones were possible as well; i.e., at these positions, the formation of a salt bridge is either impossible or one of several options to increase cosolvent resistance). All remaining residues of BSLA are colored in grey.
These findings lead to the conclusion that introducing charges does not necessarily stabilize the protein by forming salt bridges. Such salt bridges are important elements in stabilizing enzymes in aqueous solutions and they certainly participate in the stabilization of enzymes in water/organic cosolvent systems as well. However, in the present study we illustrate that the formation of salt bridges may be of minor importance for increased resistance to water-miscible organic solvents after the introduction of charged amino acids in BSLA. It is more likely that the majority of the introduced charges act in a water-attracting way to prevent cosolvent molecules from interacting with the enzyme. For example, the formation of new hydrogen bonds between residues on the enzyme’s surface was observed in organic cosolvent-stable enzyme variants in previous studies, [33,35,40]. Such hydrogen bonds could be established between surface residues (e.g., after the substitution to polar or charged amino acids) as well as with water molecules. A molecular dynamics simulation of Fusarium solani cutinase in five organic solvents of different polarity showed that—independent of the organic solvent—water clusters are preferentially formed in certain areas of the enzyme’s surface [18]. Future molecular dynamics simulations of improved BSLA variants in water/cosolvent systems will reveal if this is also the case for BSLA and if possible water clustering areas overlap with the location at which surface charges that do not form salt bridges were introduced (green areas in Figure 4). This may ultimately enable the identification of the underlying molecular principles for protein stabilization in organic cosolvents. In the absence of such molecular understanding and based on the results presented in this study, random mutagenesis might not be the ideal way to improve organic cosolvent resistance of enzymes. Designing smart libraries (e.g., enriched in enzyme variants with substitutions to charged amino acids at surface-exposed positions) could represent a more suitable approach until our molecular understanding allows us to deduce general design principles.
4. Materials and Methods
4.1. Generation of the BSLA SSM Library and Random Mutagenesis Libraries
A detailed description of the BSLA SSM library generation was reported in our previous studies [28,29]. Briefly, a library of 3620 unique variants (20 amino acids × 181 positions) was generated by site-saturation mutagenesis (SSM) and site-directed mutagenesis (SDM). In this report, the term BSLA SSM library refers to the combined mutant libraries (SDM and SSM) which encode the full natural diversity of BSLA with single amino acid exchanges (termed natural single substitution diversity). A coverage of 100% was confirmed by extensive sequencing.
The generation of the three random mutagenesis libraries was described before [27]. Error-prone PCR (epPCR) was performed using two different concentrations of MnCl2 (0.1 mM and 0.3 mM) to obtain a library with high mutagenesis frequency (epPCR-high, ~4 aa per sequence) and low mutagenesis frequency (epPCR-low, ~1 aa per sequence). Additionally, a transversion-enriched Sequence Saturation Mutagenesis (SeSaM-Tv P/P) experiment was performed in accordance with a protocol described before [27,34] in order to obtain a third random mutagenesis library. Single clones of each library were randomly picked and sent for sequencing. For each random mutagenesis library, 1000 mutations were analyzed and compared to the BSLA SSM library.
4.2. Amino Acid Categorization
For graphical presentation, amino acids were grouped into different categories according to their chemical properties: Aromatic amino acids (F, Y, W), hydrophobic amino acids (A, V, L, I, M), polar amino acids (S, T, N, Q), and charged amino acids (D, E, H, K, R). The remaining amino acids (G, C, P) were combined in a category termed special.
4.3. BSLA SSM Library Expression in 96-Well Plates
Cultivation of BSLA SSM library variants was performed as previously described [28]. Briefly, three wild type (WT) clones and one empty vector clone (controls) alongside 92 clones of the BSLA SSM libraries were incubated in 150 mL of a modified auto-induction medium [41] (see Supplementary Materials) supplemented with ampicillin in a 96-well plate (V-bottomed, polystyrene MTP, Greiner Bio-One, Frickenhausen, Germany) for 16 h (37 °C, 70% humidity, 900 rpm). After centrifugation of the expression culture (4000 g, 20 min, 4 °C, Eppendorf 5804R/5810R, Eppendorf, Wesseling, Germany) the supernatant containing the secreted BSLA was used for activity measurements.
4.4. MTP-Based p-Nitrophenyl Butyrate (pNPB) Assay for BSLA Activity Measurement
BSLA activity measurements were performed as described previously [28]. BSLA converts pNPB to p-nitrophenolate upon hydrolysis, which was detected photometrically at 410 nm. In each well, 90 µL of triethanolamine (TEA) buffer (50 mM, pH 7.4) was added to 10 µL of culture supernatant and freshly prepared substrate solution (100 µL) containing pNPB (0.5 mM) and acetonitrile (10% (v/v)). Acetonitrile was present in all reactions to provide pNPB solubility and thus to ensure reliable activity measurements. Absorption measurement at 410 nm was performed at room temperature for 8 min in a microtiter plate reader (Tecan infinite M200Pro, Tecan Group Ltd., Männedorf, Switzerland). A 96-well plate only containing BSLA WT was used to determine the assay’s apparent standard deviation. Additionally, a 96-well plate exclusively containing empty vector (EV) controls was measured. These blanks were subtracted for the determination of the true standard deviation.
4.5. Optimization of Screening Conditions in Organic Solvents
In this study, 1,4-dioxane (DOX), 2,2,2-trifluoroethanol (TFE), and dimethyl sulfoxide (DMSO) were used as water-miscible organic cosolvents. The three cosolvents are characterized by a wide range of polarity (see Table S4, Supplementary Materials). Different concentrations of the cosolvent in TEA buffer (50 mM, pH 7.4) were prepared. Organic cosolvent concentrations for BSLA SSM library screening were chosen to ensure a residual BSLA WT activity of ~30%. Similar values have proven to be a good compromise between the enzyme’s resistance on the one hand and reliably determining the enzyme’s activity compared to BSLA WT in a previous study [28]. The pH of the incubation mixture (BSLA lipase, TEA buffer, organic cosolvent) and the pH of the reaction mixture were measured and pH shifts of less than 0.22 pH units were observed.
4.6. Screening of the BSLA SSM Library in the Presence of the Three Organic Cosolvents
In each well of a DOX-resistant polypropylene MTP (flat-bottomed, Greiner Bio-One, Frickenhausen, Germany), 10 µL of culture supernatant was incubated with 90 µL of the organic cosolvent (concentration in Table 1) for 2 h at room temperature on an MTP shaker (1000 rpm, Microtiter shaker TiMix 5, Edmund Bühler GmbH, Hechingen, Germany). For each well, the supernatant was incubated in the presence and absence of organic cosolvent (DOX, TFE or DMSO). Freshly prepared pNPB substrate solution (100 µL) was added and the absorption at 410 nm was measured at room temperature for 8 min (Tecan infinite M200Pro, Tecan Group Ltd., Männedorf, Switzerland). The BSLA activity (WT or variant) in the presence of organic cosolvent was divided by the activity without DOX, TFE, or DMSO to yield the residual BSLA activity (equation in Supporting Materials).
Beneficial positions were defined as positions harboring at least one substitution leading to residual activity (Rv) that was significantly higher (beyond 3σ) than the wild type activity (RWT) (Rv ≥ RWT + 3σ; σ: true standard deviation of the screening system). Visual representations of all generated beneficial variants were created by means of the WebLogo 3 server: http://weblogo.threeplusone.com/ [42].
4.7. Computational Methods
The classification into surface-exposed and buried amino acid residues was done based on solvent accessible surface calculations of amino acids residues. The solvent-accessible surface was calculated for each residue with the BIOVIA Discovery Studio software package [43] using a water probe with a radius of 1.5 Å and a cutoff of <5% maximum solvent accessibility for buried residues based on the chain A of the BSLA X-ray crystal structure (PDB ID: 1I6W) [44].
For the analysis of charged amino acid residues, every histidine was considered as potentially positively charged. Structural models of BSLA variants were constructed in YASARA Structure version 13.9.8 [45] using the YASARA-FoldX plugin [46,47]. A FoldX mutation run including rotamer search exploring alternative conformations (3 independent runs) of mutation and the surrounding side chains below 6 Å distance from the mutated residue was performed during the FoldX energy minimization employing a probability-based rotamer library. Stabilization energy calculation were computed with FoldX version 3.0 Beta [47] using standard settings. The starting coordinates for the FoldX in silico mutagenesis experiment were taken from the X-ray crystal structure of the chain A of BSLA WT (PDB-code: 1I6W) [44]. Oppositely charged residues with their Cβ distance <8–10 Å were considered as “potentially salt-bridge-forming” to ensure satisfaction of geometric distance and direction criteria [48] for potential salt-bridges. For all beneficial substitutions to charged amino acids potentially forming salt bridges, non-charged beneficial substitutions at the same position were also analyzed. The surface representation of this salt bridge analysis was generated using PyMOL [49].
5. Conclusions
In this work, we presented a quantitative comparison of organic cosolvent (DOX, TFE, DMSO) resistant BSLA mutants detected in a BSLA SSM library (representing the entire single amino acid substitution diversity) and three state of the art random mutagenesis libraries. In summary, the BSLA SSM library demonstrates that 5%–11% of all possible single substitutions contribute to improving cosolvent resistance. The three random mutagenesis methods, however, generated only 7%–12% of this fraction. Furthermore, the comprehensive analysis of the BSLA SSM library revealed that there is no easy way of protein engineering to simultaneously enhance enzyme resistance to different types of water-miscible organic cosolvents. Although a considerable amount of beneficial mutations was found to improve BSLA resistance to DOX (apolar), TFE (polar protic), or DMSO (polar aprotic) individually, only a fraction of them was common for all three cosolvents. This illustrates that even when the entire single substitution diversity of a protein is available, our understanding of its organic cosolvent resistance still remains incomplete. Hence, it seems counterintuitive to deduce general design principles from only a few amino acid exchanges as it is commonly done in random mutagenesis-based directed evolution campaigns. Yet, the analysis of the BSLA SSM library provides valuable insights into the role of surface-exposed charged amino acid residues for the resistance to organic cosolvents. A structural investigation revealed that this is due to the attraction of water rather than to the formation of new salt bridges. Thus, the introduction of charges at the enzyme’s surface could be a reasonable strategy to promote resistance to water-miscible organic solvents and may assist future protein engineering studies to improve biocatalysis in organic cosolvents.
Supplementary Materials
The following are available online at www.mdpi.com/2073-4344/7/5/142/s1. Auto-induction medium, definition of residual activity (equation). Table S1: Number of beneficial amino acid positions in BSLA generated by SSM, epPCR-low, epPCR-high, SeSaM and theoretical combinations, Table S2: Location of amino acid positions within BSLA with at least one beneficial substitution improving organic cosolvent resistance to DOX, TFE, and DMSO, Table S3: Number, category and location of common beneficial amino acid substitutions and positions for resistance to more than one organic solvent, Table S4: Relative permittivity of DOX, TFE, and DMSO, Figure S1: Distribution of residual activity of beneficial BSLA variants for three organic cosolvents, Figure S2: Number of beneficial amino acid substitutions, Figure S3: Numbers of replaced amino acids, Figure S4: Amino acid distribution of BSLA WT.
Acknowledgments
The authors wish to thank the Deutsche Forschungsgemeinschaft (DFG) for financial support through the Research Training Group “BioNoCo–Biocatalysis using non-conventional media” (GRK1166). Assistance by Alexander Fulton in generating the BSLA SSM library and editorial advice by Christoph Gertler are gratefully acknowledged.
Author Contributions
V.J.F.-M., K.E.J., and U.S. conceived and designed the research; V.J.F.-M. and J.Z. performed the experiments; U.M., L.Z., M.B., and M.D.D. analyzed the data; U.M., L.Z., M.D.D., and U.S. wrote the paper.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Cheng, F.; Zhu, L.; Schwaneberg, U. Directed evolution 2.0: Improving and deciphering enzyme properties. Chem. Commun. 2015, 51, 9760–9772. [Google Scholar] [CrossRef] [PubMed]
- Arnold, F.H. Engineering enzymes for non-aqueous solvents. Trends Biotechnol. 1990, 8, 244–249. [Google Scholar] [CrossRef]
- Dordick, J.S. Enzymatic catalysis in monophasic organic solvents. Enzyme Microb. Technol. 1989, 11, 194–211. [Google Scholar] [CrossRef]
- Halling, P.J. Biocatalysis in multi-phase reaction mixtures containing organic liquids. Biotechnol. Adv. 1987, 5, 47–84. [Google Scholar] [CrossRef]
- Serdakowski, A.L.; Dordick, J.S. Enzyme activation for organic solvents made easy. Trends Biotechnol. 2008, 26, 48–54. [Google Scholar] [CrossRef] [PubMed]
- Zaks, A.; Klibanov, A.M. Enzyme-catalyzed processes in organic solvents. Proc. Natl. Acad. Sci. USA 1985, 82, 3192–3196. [Google Scholar] [CrossRef] [PubMed]
- Dill, K.A. Dominant forces in protein folding. Biochemistry 1990, 29, 7133–7155. [Google Scholar] [CrossRef] [PubMed]
- Pace, C.N.; Shirley, B.A.; McNutt, M.; Gajiwala, K. Forces contributing to the conformational stability of proteins. FASEB J. 1996, 10, 75–83. [Google Scholar] [PubMed]
- Prabhu, N.; Sharp, K. Protein-solvent interactions. Chem. Rev. 2006, 106, 1616–1623. [Google Scholar] [CrossRef] [PubMed]
- Wang, W. Instability, stabilization, and formulation of liquid protein pharmaceuticals. Int. J. Pharm. 1999, 185, 129–188. [Google Scholar] [CrossRef]
- Bone, S.; Pethig, R. Dielectric studies of protein hydration and hydration-induced flexibility. J. Mol. Biol. 1985, 181, 323–326. [Google Scholar] [CrossRef]
- Daniel, R.M.; Dunn, R.V.; Finney, J.L.; Smith, J.C. The role of dynamics in enzyme activity. Annu. Rev. Biophys. Biomol. Struct. 2003, 32, 69–92. [Google Scholar] [CrossRef] [PubMed]
- Gorman, L.A.S.; Dordick, J.S. Organic solvents strip water off enzymes. Biotechnol. Bioeng. 1992, 39, 392–397. [Google Scholar] [CrossRef] [PubMed]
- Poole, P.L.; Finney, J.L. Hydration-induced conformational and flexibility changes in lysozyme at low water content. Int. J. Biol. Macromol. 1983, 5, 308–310. [Google Scholar] [CrossRef]
- Rupley, J.A.; Careri, G. Protein hydration and function. Adv. Protein Chem. 1991, 41, 37–172. [Google Scholar] [PubMed]
- Gao, X.G.; Maldonado, E.; Pérez-Montfort, R.; Garza-Ramos, G.; de Gómez-Puyou, M.T.; Gómez-Puyou, A.; Rodríguez-Romero, A. Crystal structure of triosephosphate isomerase from Trypanosoma cruzi in hexane. Proc. Natl. Acad. Sci. USA 1999, 96, 10062–10067. [Google Scholar] [CrossRef] [PubMed]
- Zaks, A.; Klibanov, A.M. The effect of water on enzyme action in organic media. J. Biol. Chem. 1988, 263, 8017–8021. [Google Scholar] [PubMed]
- Micaêlo, N.M.; Soares, C.M. Modeling hydration mechanisms of enzymes in nonpolar and polar organic solvents. FEBS J. 2007, 274, 2424–2436. [Google Scholar] [CrossRef] [PubMed]
- Wedberg, R.; Abildskov, J.; Peters, G.H. Protein dynamics in organic media at varying water activity studied by molecular dynamics simulation. J. Phys. Chem. 2012, 116, 2575–2585. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Russell, A.J. The role of hydration in enzyme activity and stability: 1. Water adsorption by alcohol dehydrogenase in a continuous gas phase reactor. Biotechnol. Bioeng. 1996, 49, 700–708. [Google Scholar] [CrossRef]
- Yang, L.; Dordick, J.S.; Garde, S. Hydration of enzyme in nonaqueous media is consistent with solvent dependence of its activity. Biophys. J. 2004, 87, 812–821. [Google Scholar] [CrossRef] [PubMed]
- Kuper, J.; Tee, K.L.; Wilmanns, M.; Roccatano, D.; Schwaneberg, U.; Wong, T.S. The role of active-site Phe87 in modulating the organic co-solvent tolerance of cytochrome P450 BM3 monooxygenase. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 2012, 68, 1013–1017. [Google Scholar] [CrossRef] [PubMed]
- Kuper, J.; Wong, T.S.; Roccatano, D.; Wilmanns, M.; Schwaneberg, U. Understanding a mechanism of organic cosolvent inactivation in heme monooxygenase P450 BM-3. J. Am. Chem. Soc. 2007, 129, 5786–5787. [Google Scholar] [CrossRef] [PubMed]
- Roccatano, D.; Wong, T.S.; Schwaneberg, U.; Zacharias, M. Toward understanding the inactivation mechanism of monooxygenase P450 BM-3 by organic cosolvents: A molecular dynamics simulation study. Biopolymers 2006, 83, 467–476. [Google Scholar] [CrossRef] [PubMed]
- Stepankova, V.; Bidmanova, S.; Koudelakova, T.; Prokop, Z.; Chaloupkova, R.; Damborsky, J. Strategies for stabilization of enzymes in organic solvents. ACS Catal. 2013, 3, 2823–2836. [Google Scholar] [CrossRef]
- Chakravorty, D.; Parameswaran, S.; Dubey, V.K.; Patra, S. Unraveling the rationale behind organic solvent stability of lipases. Appl. Biochem. Biotechnol. 2012, 167, 439–461. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Kardashliev, T.; Ruff, A.J.; Bocola, M.; Schwaneberg, U. Lessons from diversity of directed evolution experiments by an analysis of 3000 mutations. Biotechnol. Bioeng. 2014, 111, 2380–2389. [Google Scholar] [CrossRef] [PubMed]
- Frauenkron-Machedjou, V.J.; Fulton, A.; Zhu, L.; Anker, C.; Bocola, M.; Jaeger, K.E.; Schwaneberg, U. Towards understanding directed evolution: More than half of all amino acid positions contribute to ionic liquid resistance of Bacillus subtilis lipase A. ChemBioChem 2015, 16, 937–945. [Google Scholar] [CrossRef] [PubMed]
- Fulton, A.; Frauenkron-Machedjou, V.J.; Skoczinski, P.; Wilhelm, S.; Zhu, L.; Schwaneberg, U.; Jaeger, K.E. Exploring the protein stability landscape: Bacillus subtilis lipase A as a model for detergent tolerance. ChemBioChem 2015, 16, 930–936. [Google Scholar] [CrossRef] [PubMed]
- Secundo, F.; Fialà, S.; Fraaije, M.W.; de Gonzalo, G.; Meli, M.; Zambianchi, F.; Ottolina, G. Effects of water miscible organic solvents on the activity and conformation of the baeyer–villiger monooxygenases from Thermobifida fusca and Acinetobacter calcoaceticus: A comparative study. Biotechnol. Bioeng. 2011, 108, 491–499. [Google Scholar] [CrossRef] [PubMed]
- Uematsu, M.; Frank, E. Static dielectric constant of water and steam. J. Phys. Chem. Ref. Data 1980, 9, 1291–1306. [Google Scholar] [CrossRef]
- MacGregor, W.S. The chemical and physical properties of DMSO. Ann. N. Y. Acad. Sci. 1967, 141, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Korman, T.P.; Sahachartsiri, B.; Charbonneau, D.M.; Huang, G.L.; Beauregard, M.; Bowie, J.U. Dieselzymes: Development of a stable and methanol tolerant lipase for biodiesel production by directed evolution. Biotechnol. Biofuels 2013, 6, 70. [Google Scholar] [CrossRef] [PubMed]
- Mundhada, H.; Marienhagen, J.; Scacioc, A.; Schenk, A.; Roccatano, D.; Schwaneberg, U. SeSaM-Tv-II generates a protein sequence space that is unobtainable by epPCR. ChemBioChem 2011, 12, 1595–1601. [Google Scholar] [CrossRef] [PubMed]
- Dror, A.; Shemesh, E.; Dayan, N.; Fishman, A. Protein engineering by random mutagenesis and structure-guided consensus of Geobacillus stearothermophilus lipase T6 for enhanced stability in methanol. Appl. Environ. Microbiol. 2014, 80, 1515–1527. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.; Bagchi, B. Comparative study of protein unfolding in aqueous urea and dimethyl sulfoxide solutions: Surface polarity, solvent specificity, and sequence of secondary structure melting. J. Phys. Chem. 2014, 118, 5691–5697. [Google Scholar] [CrossRef] [PubMed]
- Park, H.J.; Joo, J.C.; Park, K.; Kim, Y.H.; Yoo, Y.J. Prediction of the solvent affecting site and the computational design of stable Candida antarctica lipase B in a hydrophilic organic solvent. J. Biotechnol. 2013, 163, 346–352. [Google Scholar] [CrossRef] [PubMed]
- Sears, P.; Schuster, M.; Wang, P.; Witte, K.; Wong, C.H. Engineering subtilisin for peptide coupling: Studies on the effects of counterions and site-specific modifications on the stability and specificity of the enzyme. J. Am. Chem. Soc. 1994, 116, 6521–6530. [Google Scholar] [CrossRef]
- Martinez, P.; Arnold, F.H. Surface charge substitutions increase the stability of. alpha.-lytic protease in organic solvents. J. Am. Chem. Soc. 1991, 113, 6336–6337. [Google Scholar] [CrossRef]
- Reetz, M.T.; Soni, P.; Fernández, L.; Gumulya, Y.; Carballeira, J.D. Increasing the stability of an enzyme toward hostile organic solvents by directed evolution based on iterative saturation mutagenesis using the B-FIT method. Chem. Commun. 2010, 46, 8657–8658. [Google Scholar] [CrossRef] [PubMed]
- Studier, F.W. Protein production by auto-induction in high-density shaking cultures. Protein Expr. Purif. 2005, 41, 207–234. [Google Scholar] [CrossRef] [PubMed]
- Crooks, G.E.; Hon, G.; Chandonia, J.M.; Brenner, S.E. WebLogo: A sequence logo generator. Genome Res. 2004, 14, 1188–1190. [Google Scholar] [CrossRef] [PubMed]
- Dassault Systèmes BIOVIA. Discovery Studio Modeling Environment, 4th ed.; Dessault Systèmes BIOVIA: San Diego, CA, USA, 2015. [Google Scholar]
- van Pouderoyen, G.; Eggert, T.; Jaeger, K.E.; Dijkstra, B.W. The crystal structure of Bacillus subtilis lipase: A minimal α/β hydrolase fold enzyme. J. Mol. Biol. 2001, 309, 215–226. [Google Scholar] [CrossRef] [PubMed]
- Krieger, E.; Koraimann, G.; Vriend, G. Increasing the precision of comparative models with YASARA NOVA – a self-parameterizing force field. Proteins Struct. Funct. Bioinform. 2002, 47, 393–402. [Google Scholar] [CrossRef] [PubMed]
- Van Durme, J.; Delgado, J.; Stricher, F.; Serrano, L.; Schymkowitz, J.; Rousseau, F. A graphical interface for the FoldX forcefield. Bioinformatics 2011, 27, 1711–1712. [Google Scholar] [CrossRef] [PubMed]
- Guerois, R.; Nielsen, J.E.; Serrano, L. Predicting changes in the stability of proteins and protein complexes: A study of more than 1000 mutations. J. Mol. Biol. 2002, 320, 369–387. [Google Scholar] [CrossRef]
- Donald, J.E.; Kulp, D.W.; DeGrado, W.F. Salt bridges: Geometrically specific, designable interactions. Proteins Struct. Funct. Bioinform. 2011, 79, 898–915. [Google Scholar] [CrossRef] [PubMed]
- The PyMOL Molecular Graphics System, version 1.3r1; Llc Schrödinger: New York, NY, USA, 2010.
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).