Geometric Understanding of Local Fluctuation Distribution of Conduction Time in Lined-Up Cardiomyocyte Network in Agarose-Microfabrication Multi-Electrode Measurement Assay
Abstract
1. Introduction
2. Materials and Methods
2.1. Embryonic Mouse Primary Cardiomyocytes
2.2. Human Embryonic Stem Cell-Derived Cardiomyocytes
2.3. Agarose Microfabrication
2.4. Cell Culture
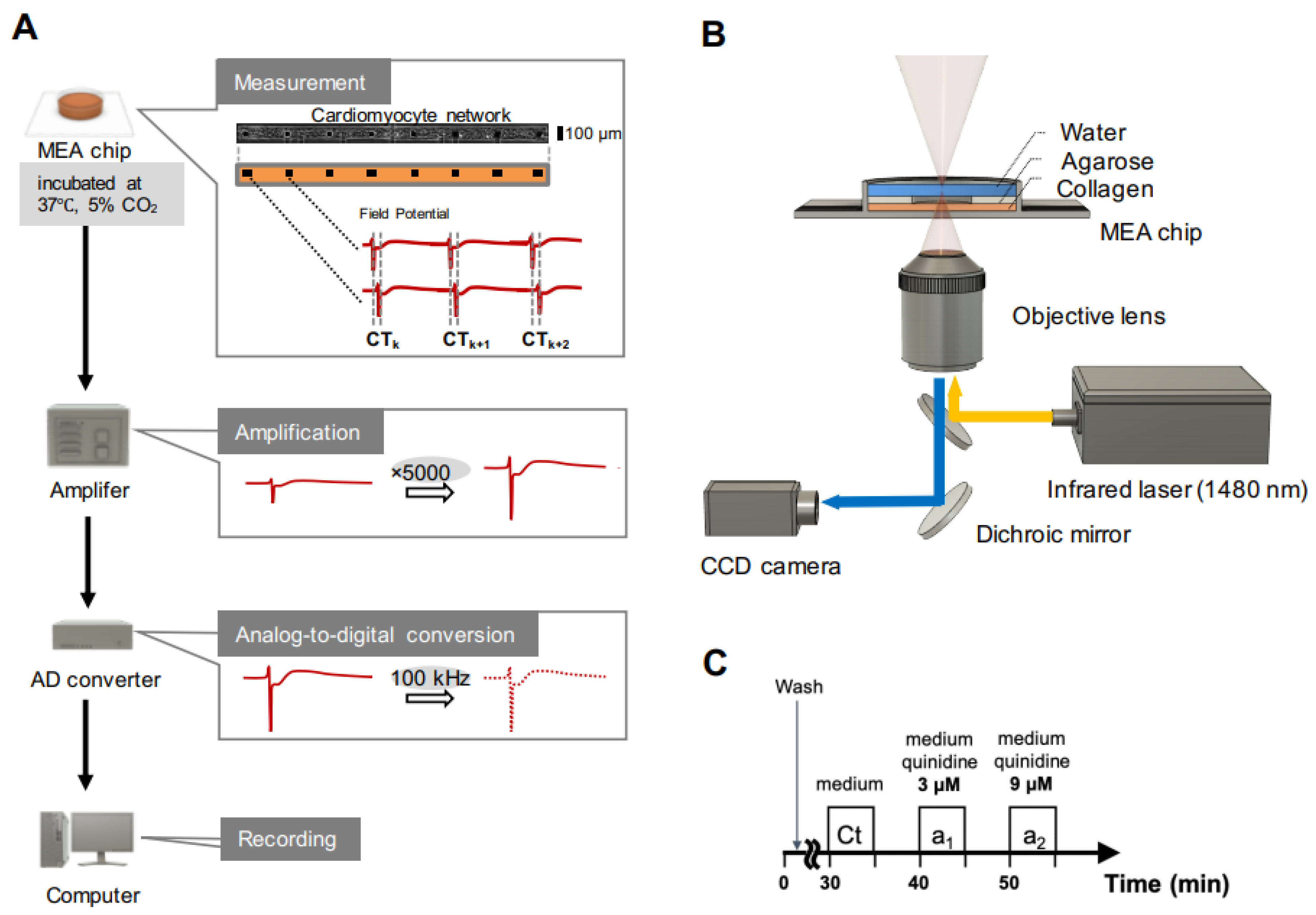
2.5. Measurement System
2.6. Data Analysis
2.7. Drug Administration
2.8. Cell Staining
2.9. Statistical Analysis
2.10. Simulation
3. Results and Discussion
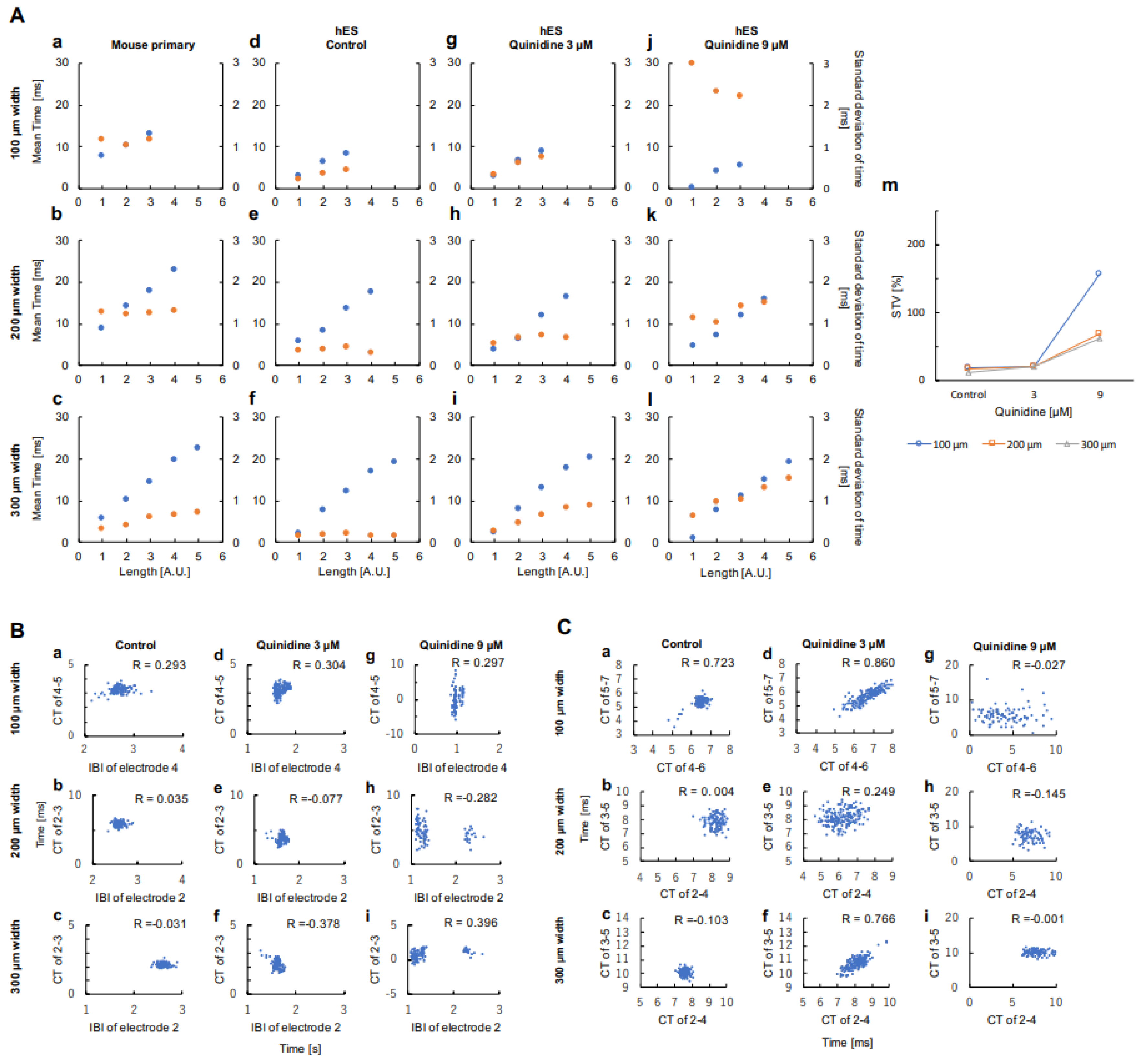
3.1. Distribution of Conduction Time in Width Controlled Cardiomyocyte Networks
3.2. Effect of Fast Inward-Sodium-Current Blocking on the Distribution of Conduction Time
3.3. Correlation of Beating Intervals and Conduction Time
3.4. Correlation of Conduction Time and Its Fluctuation between Neighboring Units
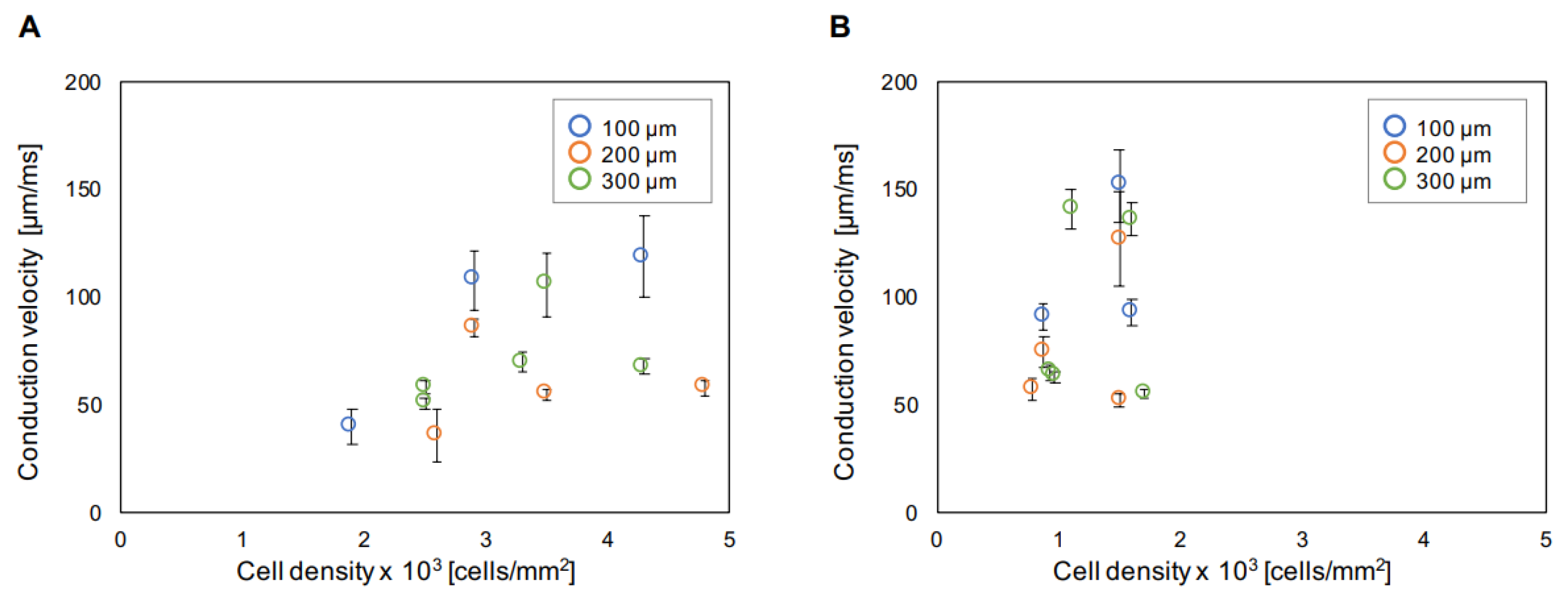
3.5. Influence of Cell Density for Conduction Velocity in Unit Length
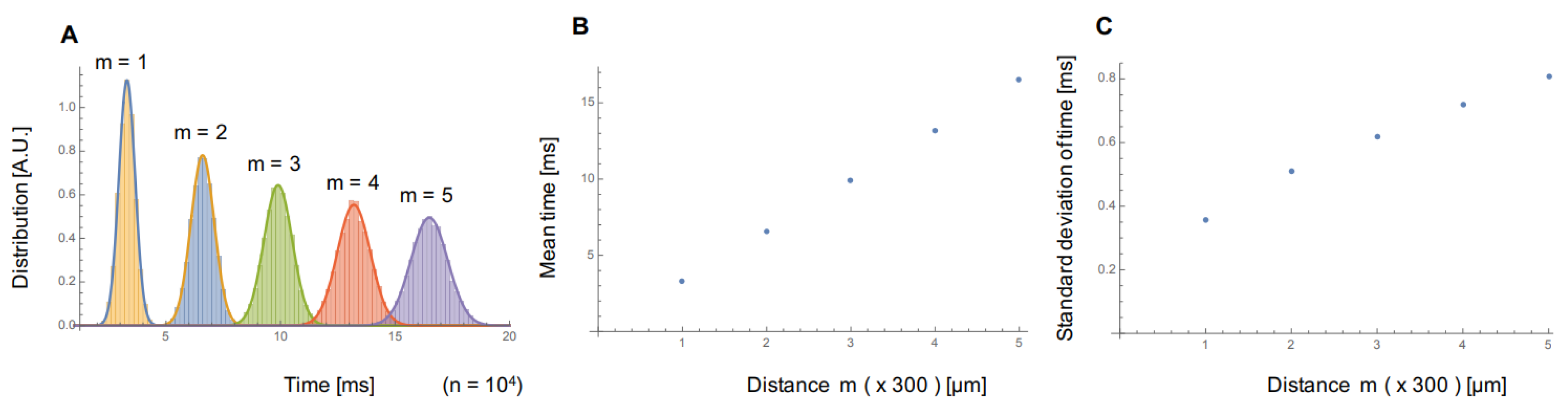
3.6. Can the Faster Firing Regulation Explain These Conduction Characteristics?
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Brill, M.H.; Waxman, S.G.; Moore, J.W.; Joyner, R.W. Conduction velocity and spike configuration in myelinated fibres: Computed dependence on internode distance. J. Neurol. Neurosurg. Psychiatry 1977, 40, 769–774. [Google Scholar] [CrossRef] [PubMed]
- Lang, E.J.; Rosenbluth, J. Role of myelination in the development of a uniform olivocerebellar conduction time. J. Neurophysiol. 2003, 89, 2259–2270. [Google Scholar] [CrossRef] [PubMed]
- Salami, M.; Itami, C.; Tsumoto, T.; Kimura, F. Change of conduction velocity by regional myelination yields constant latency irrespective of distance between thalamus and cortex. Proc. Natl. Acad. Sci. USA 2003, 100, 6174–6179. [Google Scholar] [CrossRef] [PubMed]
- Goshima, K.; Tonomura, Y. Synchronized beating of embryonic mouse myocardial cells mediated by FL cells in monolayer culture. Exp. Cell Res. 1969, 56, 387–392. [Google Scholar] [CrossRef]
- Vassalle, M. The relationship among cardiac pacemakers. Overdrive suppression. Circ. Res. 1977, 41, 269–277. [Google Scholar] [CrossRef]
- Hatano, A.; Okada, J.; Washio, T.; Hisada, T.; Sugiura, S. A three-dimensional simulation model of cardiomyocyte integrating excitation-contraction coupling and metabolism. Biophys. J. 2011, 101, 2601–2610. [Google Scholar] [CrossRef]
- Keener, J.; Sneyd, J. Mathematical Physiology; Springer: New York, NY, USA, 1998. [Google Scholar]
- Murray, J. Mathematical Biology; Springer: Berlin/Heiderberg, Germany, 2002. [Google Scholar]
- Kuramoto, Y. Chemical Oscillations, Waves, and Turbulence; Springer: New York, NY, USA, 1984. [Google Scholar]
- Winfree, A. The Geometry of Biological Time; Springer: New York, NY, USA, 2001. [Google Scholar]
- Kori, K.; Kawamura, Y.; Masuda, N. Structure of cell networks critically determines oscillation regularity. J. Theor. Biol. 2012, 297, 61–72. [Google Scholar] [CrossRef][Green Version]
- Keener, J.P.; Hoppensteadt, F.C.; Rinzel, J. Integrate-and-fire models of nerve membrane response to oscillatory input. SIAM J. Appl. Math. 1981, 41, 503–517. [Google Scholar] [CrossRef]
- Burkitt, A.N. A review of the integrate-and-fire neuron model: I. Homogeneous synaptic input. Biol. Cybern. 2006, 95, 1–12. [Google Scholar] [CrossRef]
- Sacerdote, L.; Giraudo, M.T. Stochastic integrable and fire models: A review on mathematical methods and their applications. In Stochastic Biomathematical Models with Applications to Neuronal Modeling; Bachar, M., Batzel, J., Ditlevsen, S., Eds.; Springer: Berlin/Heidelberg, Germany, 2013. [Google Scholar]
- Kaneko, T.; Nomura, F.; Hamada, T.; Abe, Y.; Takamori, H.; Sakakura, T.; Takasuna, K.; Sanbuissho, A.; Hyllner, J.; Sartipy, P.; et al. On-chip in vitro cell-network pre-clinical cardiac toxicity using spatiotemporal human cardiomyocyte measurement on a chip. Sci. Rep. 2014, 4, 4670. [Google Scholar] [CrossRef]
- Nozaki, Y.; Honda, Y.; Watanabe, H.; Saiki, S.; Koyabu, K.; Itoh, T.; Nagasawa, C.; Nakamori, C.; Nakayama, C.; Iwasaki, H.; et al. CSAHi study: Validation of multi-electrode array systems (MEA60/2100) for prediction of drug-induced proarrhythmia using human iPS cell-derived cardiomyocytes-assessment of inter-facility and cells lot-to-lot-variability. Regul. Toxicol. Pharmacol. 2016, 77, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Asahi, Y.; Hamada, T.; Hattori, A.; Matsuura, K.; Odaka, M.; Nomura, F.; Kaneko, T.; Abe, Y.; Takasuna, K.; Sanbuissho, A.; et al. On-chip spatiotemporal electrophysiological analysis of human stem cell derived cardiomyocytes enables quantitative assessment of proarrhythmia in drug development. Sci. Rep. 2018, 8. [Google Scholar] [CrossRef] [PubMed]
- Asahi, Y.; Nomura, F.; Abe, Y.; Doi, M.; Sakakura, T.; Takasuna, K.; Yasuda, K. Electrophysiological evaluation of pentamidine and 17-AAG in human stem cell-derived cardiomyocytes for safety assessment. Eur. J. Pharmacol. 2019, 842, 221–230. [Google Scholar] [CrossRef] [PubMed]
- Kojima, K.; Moriguchi, H.; Hattori, A.; Kaneko, T.; Yasuda, K. Two-dimensional network formation of cardiac myocytes in agar microculture chip with 1480 nm infrared laser photo-thermal etching. Lab. Chip. 2003, 3, 292–296. [Google Scholar] [CrossRef] [PubMed]
- Kojima, K.; Kaneko, T.; Yasuda, K. Role of the community effect of cardiomyocyte in the entrainment and reestablishment of stable beating rhythms. Biochem. Biophys. Res. Commun. 2006, 351, 209–215. [Google Scholar] [CrossRef]
- Sokolow, M. Some quantitative aspects of treatment with quinidine. Ann. Intern. Med. 1956, 45, 582–588. [Google Scholar] [CrossRef]
- Wu, L.; Guo, D.; Li, H.; Hackett, J.; Yan, G.X.; Jiao, Z.; Antzelevitch, C.; Shryock, J.C.; Belardinelli, L. Role of late sodium current in modulating the proarrhythmic and antiarrhythmic effects of quinidine. Heart Rhythm 2008, 5, 1726–1734. [Google Scholar] [CrossRef]
- Paul, A.A.; Witchel, H.J.; Hancox, J.C. Inhibition of the current of heterologously expressed HERG potassium channels by flecainide and comparison with quinidine, propafenone and lignocaine. Br. J. Pharmacol. 2002, 136, 717–729. [Google Scholar] [CrossRef]
- Bazett, H.C. An analysis of the time-relations of electrocardiograms. Heart Vessel. 1920, 7, 353–370. [Google Scholar] [CrossRef]
- Fridericia, L.S. Die Sytolendauer im Elektrokardiogramm bei normalen Menschen und bei Herzkranken. Acta Medica Scand. 1920, 53, 460–469. [Google Scholar]
- Hayashi, T.; Tokihiro, T.; Kurihara, H.; Yasuda, K. Community effect of cardiomyocytes in beating rhythms is determined by stable cells. Sci. Rep. 2017, 7, 15450. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, T.; Tokihiro, T.; Kurihara, H.; Nomura, F.; Yasuda, K. Integrate and fire model with refractory period for synchronization of two cardiomyocytes. J. Theor. Biol. 2018, 437, 141–148. [Google Scholar] [CrossRef] [PubMed]
- Bub, G.; Shrier, A. Propagation through heterogeneous substrates in simple excitable media models. Chaos 2002, 12, 747. [Google Scholar] [CrossRef] [PubMed]
- Bub, G.; Shrier, A.; Glass, L. Spiral wave generation in heterogeneous excitable media. Phys. Rev. Lett. 2002, 88, 4. [Google Scholar] [CrossRef] [PubMed]
- Bub, G.; Shrier, A.; Glass, L. Global organization of dynamics in oscillatory heterogeneous excitable media. Phys. Rev. Lett. 2005, 94. [Google Scholar] [CrossRef] [PubMed]
- Greenberg, J.M.; Hastings, S.P. Spatial patterns for discrete models of diffusion in excitable media. SIAM J. Appl. Math. 1978, 34, 515–523. [Google Scholar] [CrossRef]

| Conduction Time (Mean ± S.D.) [ms] | |||||||
|---|---|---|---|---|---|---|---|
| Propagation Distance [μm] | |||||||
| Cell Type | Width [μm] | Sample | 300 | 600 | 900 | 1200 | 1500 |
| Mouse primary | 100 | 1 | 8.11 ± 0.747 | 10.8 ± 0.576 | 13.6 ± 0.692 | ||
| 2 | 7.91 ± 0.599 | 16.7 ± 0.703 | 28.5 ± 0.780 | 39.2 ± 1.38 | |||
| 200 | 1 | 8.19 ± 1.77 | 13.8 ± 1.68 | 17.3 ± 1.75 | 22.6 ± 1.80 | ||
| 2 | 5.51 ± 0.409 | 9.00 ± 0.469 | 16.9 ± 0.603 | 29.4 ± 1.06 | |||
| 300 | 1 | 5.94 ± 0.256 | 10.4 ± 0.333 | 14.7 ± 0.457 | 19.9 ± 0.447 | 22.7 ± 0.473 | |
| 2 | 6.63 ± 0.296 | 12.6 ± 0.445 | 23.2 ± 0.915 | ||||
| hES | 100 | 3.23 ± 0.210 | 6.55 ± 0.193 | 8.54 ± 0.212 | |||
| 200 | 5.80 ± 0.327 | 8.23 ± 0.361 | 13.6 ± 0.422 | 17.6 ± 0.298 | |||
| 300 | 2.14 ± 0.140 | 7.62 ± 0.189 | 12.2 ± 0.212 | 17.0 ± 0.155 | 19.2 ± 0.148 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sakamoto, K.; Aoki, S.; Tanaka, Y.; Shimoda, K.; Hondo, Y.; Yasuda, K. Geometric Understanding of Local Fluctuation Distribution of Conduction Time in Lined-Up Cardiomyocyte Network in Agarose-Microfabrication Multi-Electrode Measurement Assay. Micromachines 2020, 11, 1105. https://doi.org/10.3390/mi11121105
Sakamoto K, Aoki S, Tanaka Y, Shimoda K, Hondo Y, Yasuda K. Geometric Understanding of Local Fluctuation Distribution of Conduction Time in Lined-Up Cardiomyocyte Network in Agarose-Microfabrication Multi-Electrode Measurement Assay. Micromachines. 2020; 11(12):1105. https://doi.org/10.3390/mi11121105
Chicago/Turabian StyleSakamoto, Kazufumi, Shota Aoki, Yuhei Tanaka, Kenji Shimoda, Yoshitsune Hondo, and Kenji Yasuda. 2020. "Geometric Understanding of Local Fluctuation Distribution of Conduction Time in Lined-Up Cardiomyocyte Network in Agarose-Microfabrication Multi-Electrode Measurement Assay" Micromachines 11, no. 12: 1105. https://doi.org/10.3390/mi11121105
APA StyleSakamoto, K., Aoki, S., Tanaka, Y., Shimoda, K., Hondo, Y., & Yasuda, K. (2020). Geometric Understanding of Local Fluctuation Distribution of Conduction Time in Lined-Up Cardiomyocyte Network in Agarose-Microfabrication Multi-Electrode Measurement Assay. Micromachines, 11(12), 1105. https://doi.org/10.3390/mi11121105





