Mapping Foliar C, N, and P Concentrations in An Ecological Restoration Area with Mixed Plant Communities Based on LiDAR and Hyperspectral Data
Abstract
1. Introduction
2. Materials and Methods
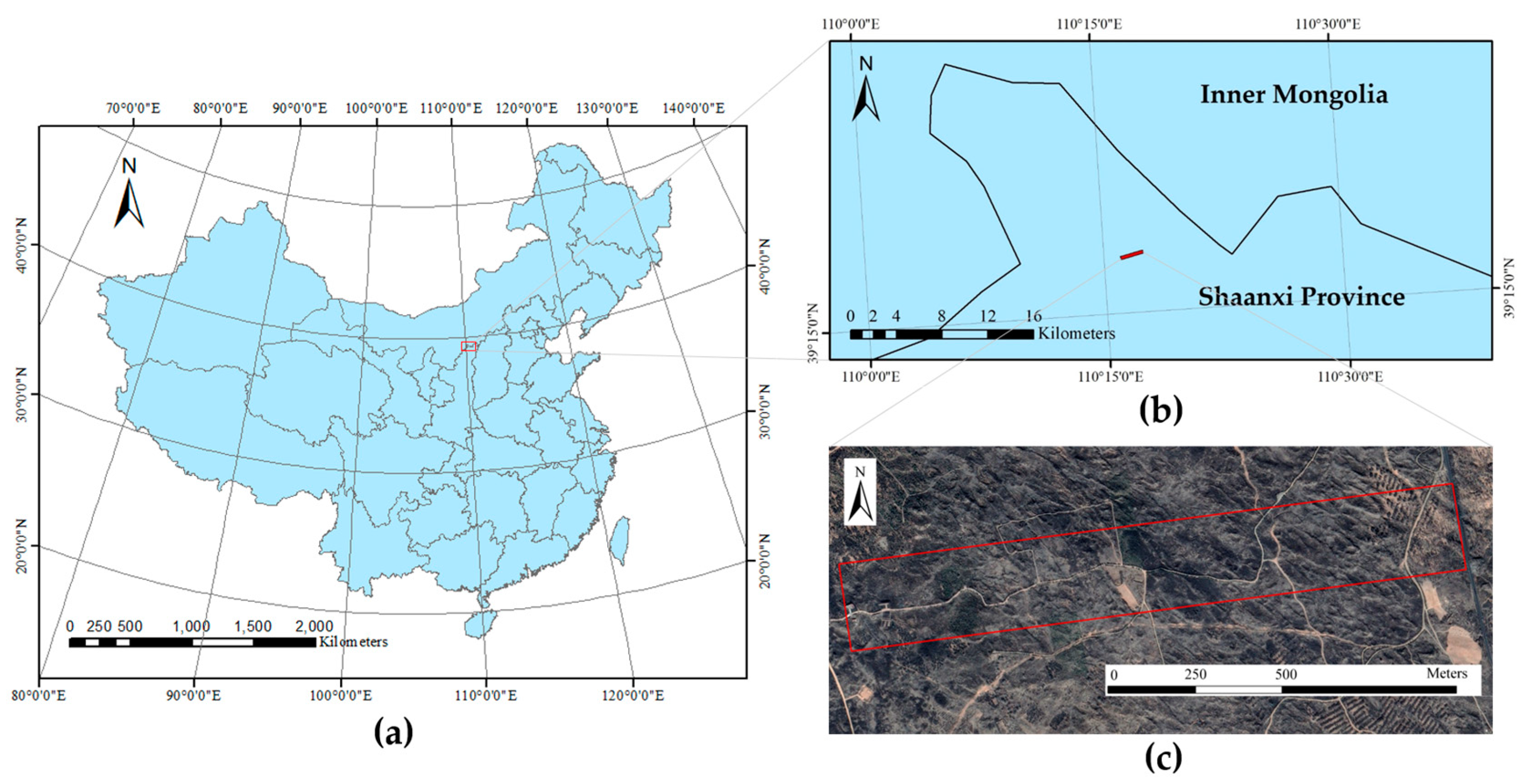
2.1. Study Area
2.2. Data
2.2.1. Remote Sensing Data
2.2.2. Field Data
2.3. Methods
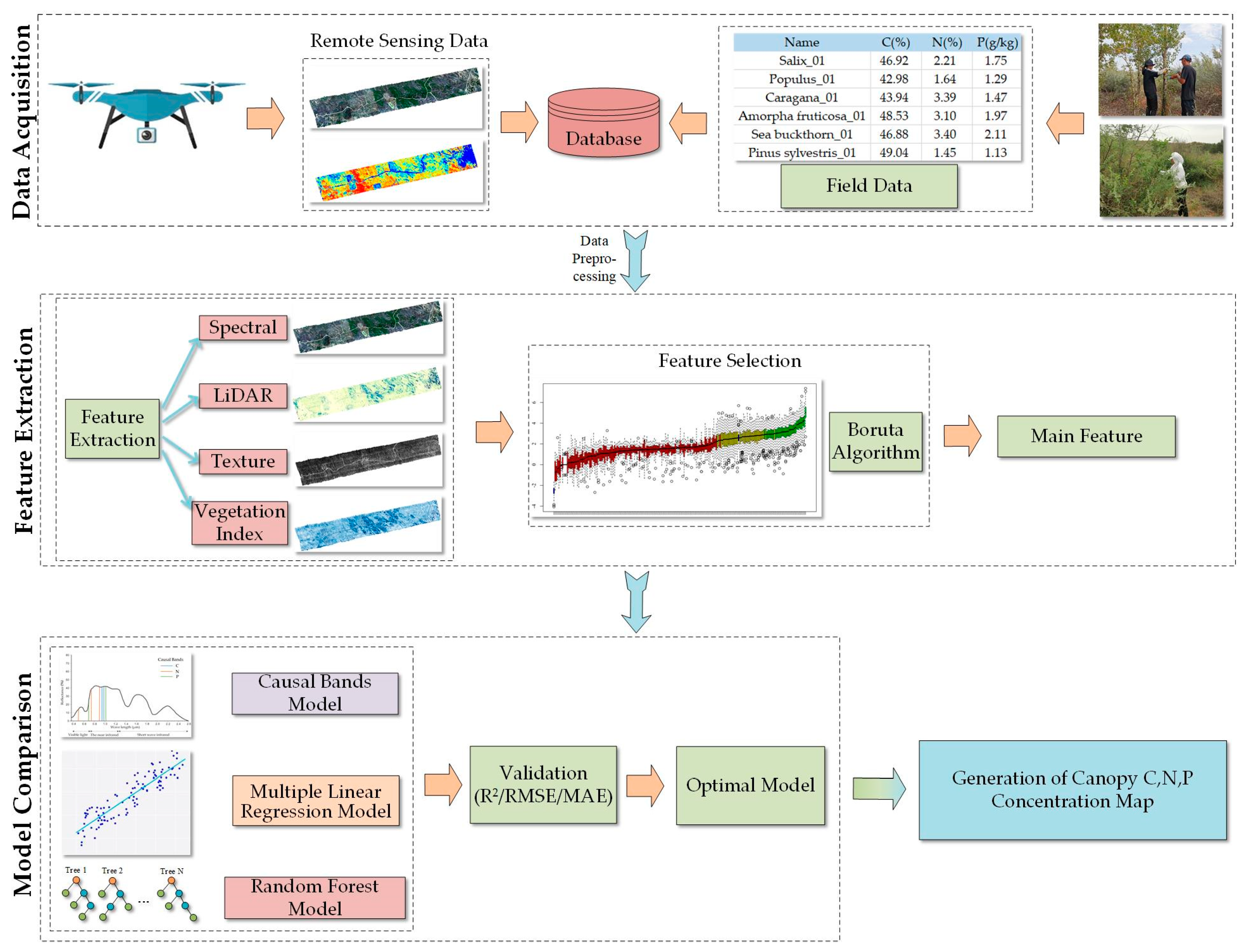
2.3.1. Technical Process
2.3.2. Extraction of Features
- Spectrum
- 2.
- Texture
- 3.
- Vegetation indices
- 4.
- Height and vegetation structure parameters
2.3.3. Selection of Features
- Build random forest: construct a random forest model using the original dataset. This typically involves multiple decision trees, each trained on a random subset.
- Calculate the importance of the original features: for each feature, compute its relative importance using the random forest model. This is achieved by measuring the contribution of the feature to the model’s predictive accuracy.
- Create shadow features: for each original feature, generate a corresponding “shadow” feature. A shadow feature is created by randomly permuting the values of the original feature.
- Build an extended random forest: build another random forest model using a dataset that includes both the original and shadow features.
- Compute the importance of the shadow features: calculate the relative importance of each shadow feature in the extended random forest.
- Compare the importance of the original and shadow features: for each original feature, compare its actual importance with the average importance of its shadow features. If the original feature’s importance is higher than the average importance of its shadow features, retain the feature; otherwise, label it as unimportant.
- Repeat steps 4–6: iterate through steps 4–6 until the stopping criteria are met, such as reaching a specified number of features or marking all features as important.
- Final feature selection: retain the features labeled as important for modeling or further analysis.
2.3.4. Estimation Model
- Causal band model
- 2.
- Multiple linear regression algorithm
- 3.
- Random forest algorithm
- n samples randomly drawn from the training set are put back.
- Create a decision tree from a dataset consisting of these n samples.
- At each node: randomly select d features without putting them back.
- Use maximized learning gain or other methods to split nodes based on these features.
- Repeat steps 1–2 several (k) times.
- Finally, take the average value according to the estimation results of these decision trees as the final estimation results.
2.3.5. Validation
3. Results
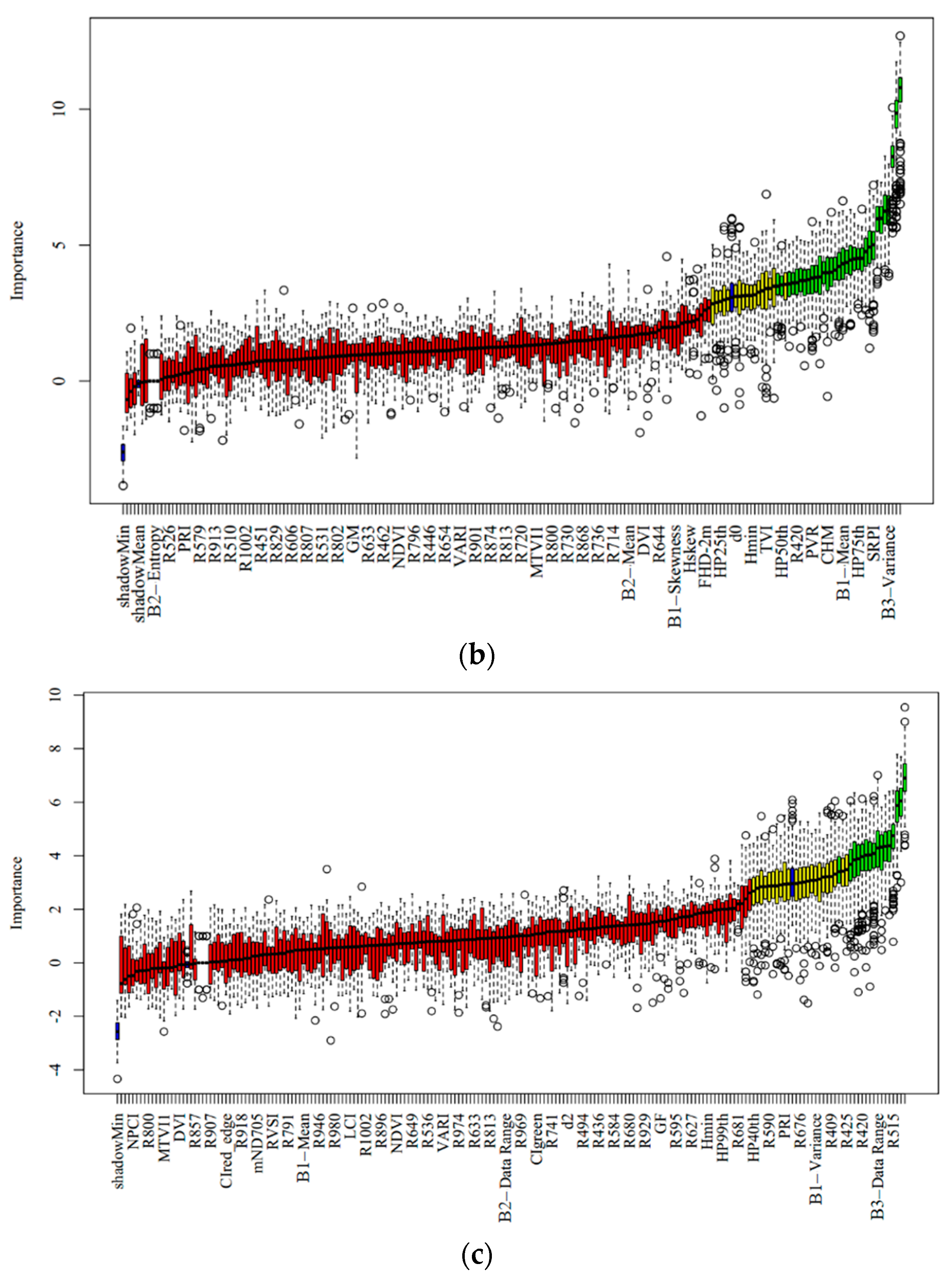
3.1. Selected Feature
3.2. Accuracy of the Estimation Model
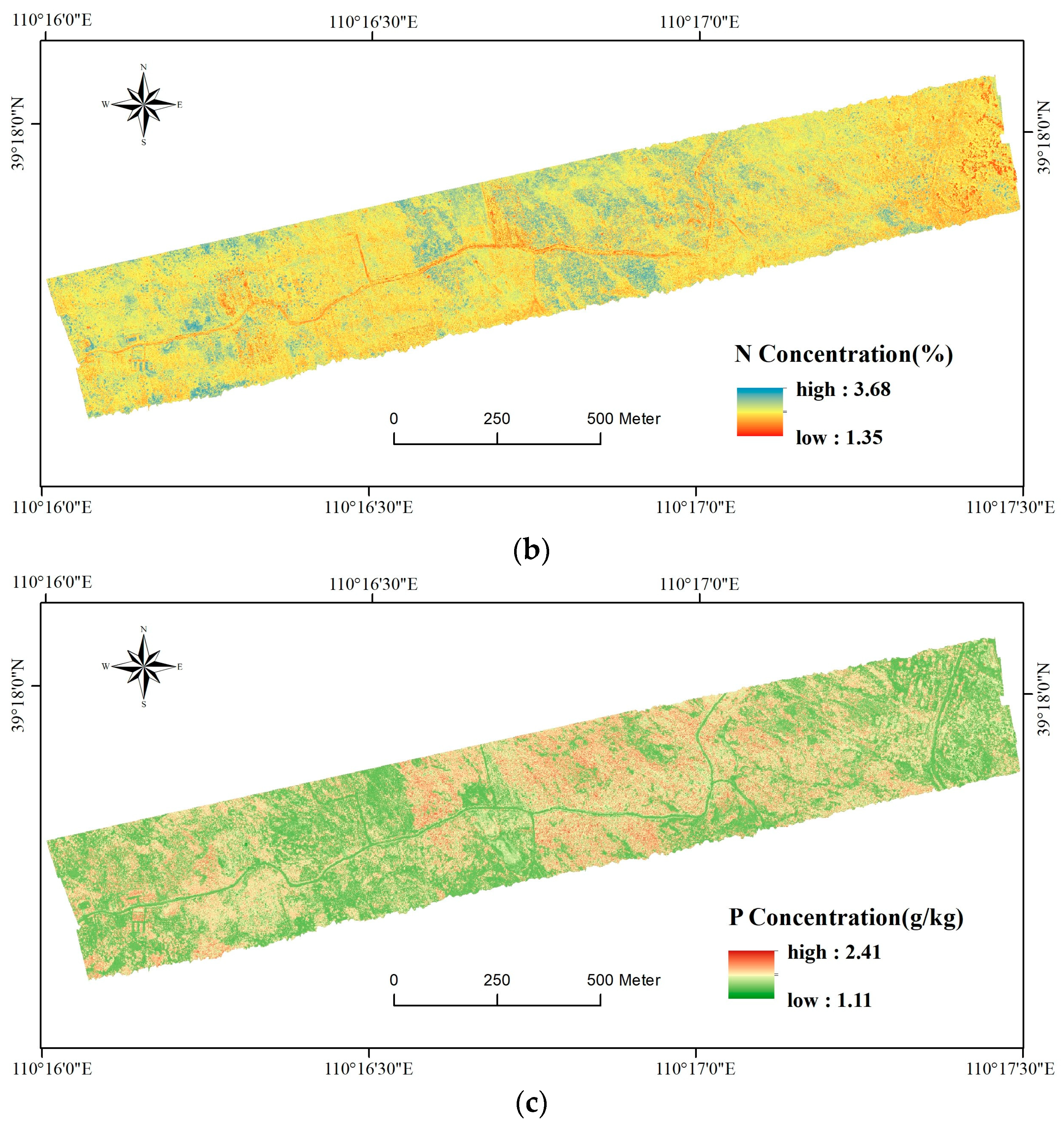
3.3. Map of Foliar C, N, and P Concentrations
4. Discussion
4.1. The Role of Features in the Model
4.2. Implications of the Foliar C, N, and P Map
4.3. Limitations and Future Work
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sun, X.; Yuan, L.; Liu, M.; Liang, S.; Li, D.; Liu, L. Quantitative estimation for the impact of mining activities on vegetation phenology and identifying its controlling factors from Sentinel-2 time series. Int. J. Appl. Earth Obs. Geoinf. 2022, 111, 102814. [Google Scholar] [CrossRef]
- Chugh, Y.P.; Schladweiler, B.K.; Skilbred, C. Sustainable and responsible mining through sound mine closure. Int. J. Coal Sci. Technol. 2023, 10, 14. [Google Scholar] [CrossRef]
- Fischer, J.; Riechers, M.; Loos, J.; Martin-Lopez, B.; Temperton, V.M. Making the UN Decade on ecosystem restoration a social-ecological endeavour. Trends Ecol. Evol. 2020, 36, 20–28. [Google Scholar] [CrossRef] [PubMed]
- Krzyszowska Waitkus, A. Sustainable reclamation practices for a large surface coal mine in shortgrass prairie, semiarid environment (Wyoming, USA): Case study. Int. J. Coal Sci. Technol. 2022, 9, 32. [Google Scholar] [CrossRef]
- Banerjee, B.P.; Raval, S. Mapping sensitive vegetation communities in mining eco-space using UAV-LiDAR. Int. J. Coal Sci. Technol. 2022, 9, 40. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhou, W. Remote sensing of vegetation fraction for monitoring reclamation dynamics: A case study in Pingshuo Mining area. In Proceedings of the 2016 IEEE International Geoscience and Remote Sensing Symposium (IGARSS), Beijing, China, 10–15 July 2016; pp. 5197–5200. [Google Scholar] [CrossRef]
- Ustin, S. Remote sensing of canopy chemistry. Proc. Natl. Acad. Sci. USA 2013, 110, 804–805. [Google Scholar] [CrossRef] [PubMed]
- Gastauer, M.; Silva, R.J.; Junior, C.F.C.; Ramos, S.J.; Filho, P.; Neto, A.; Siqueira, J.O. Mine land rehabilitation: Modern ecological approaches for more sustainable mining. J. Clean. Prod. 2018, 172, 1409–1422. [Google Scholar] [CrossRef]
- McGroddy, M.E.; Daufresne, T.; Hedin, L.O. Scaling of C: N: P stoichiometry in forests worldwide: Implications of terrestrial redfield-type ratios. Ecology 2004, 85, 2390–2401. [Google Scholar] [CrossRef]
- Han, W.; Fang, J.; Reich, P.B.; Ian Woodward, F.; Wang, Z. Biogeography and variability of eleven mineral elements in plant leaves across gradients of climate, soil and plant functional type in China. Ecol. Lett. 2011, 14, 788–796. [Google Scholar] [CrossRef]
- Xing, K.; Zhao, M.; Niinemets, U.; Niu, S.; Tian, J.; Jiang, Y.; Chen, H.; White, P.J.; Guo, D.; Ma, Z. Relationships between leaf carbon and macronutrients across woody species and forest ecosystems highlight how carbon is allocated to leaf structural function. Front. Plant Sci. 2021, 12, 674932. [Google Scholar] [CrossRef]
- Högberg, P.; Näsholm, T.; Franklin, O.; Högberg, M.N. Tamm review: On the nature of the nitrogen limitation to plant growth in Fennoscandian boreal forests. For. Ecol. Manag. 2017, 403, 161–185. [Google Scholar] [CrossRef]
- Urbina, I.; Sardans, J.; Grau, O.; Beierkuhnlein, C.; Jentsch, A.; Kreyling, J.; Peñuelas, J. Plant community composition affects the species biogeochemical niche. Ecosphere 2017, 8, e01801. [Google Scholar] [CrossRef]
- Güsewell, S. N:P ratios in terrestrial plants: Variation and functional significance. New Phytol. 2004, 164, 243–266. [Google Scholar] [CrossRef] [PubMed]
- Reich, P.B.; Oleksyn, J. Global patterns of plant leaf N and P in relation to temperature and latitude. Proc. Natl. Acad. Sci. USA 2004, 101, 11001–11006. [Google Scholar] [CrossRef] [PubMed]
- Xiong, J.; Yu, M.; Cheng, X.; Wang, C.; Zou, H. Effects of light and N-P supply ratios on growth and stoichiometric of Schimasuperba. Acta Ecol. Sin. 2021, 41, 2140–2150. [Google Scholar] [CrossRef]
- Avolio, M.L.; Koerner, S.E.; La Pierre, K.J.; Wilcox, K.R.; Wilson, G.W.T.; Smith, M.D.; Collins, S.L. Changes in plant community composition, not diversity, during a decade of nitrogen and phosphorus additions drive above-ground productivity in a tallgrass prairie. J. Ecol. 2014, 102, 1649–1660. [Google Scholar] [CrossRef]
- Stiles, W.A.V.; Rowe, E.C.; Dennis, P. Long-term nitrogen and phosphorus enrichment alters vegetation species composition and reduces carbon storage in upland soil. Sci. Total Environ. 2017, 593–594, 688–694. [Google Scholar] [CrossRef]
- McKenna, P.B.; Lechner, A.M.; Phinn, S.; Erskine, P.D. Remote sensing of mine site rehabilitation for ecological outcomes: A global systematic review. Remote Sens. 2020, 12, 3535. [Google Scholar] [CrossRef]
- Yang, Y.; Tang, J.; Zhang, Y.; Zhang, S.; Zhou, Y.; Hou, H.; Liu, R. Reforestation improves vegetation coverage and biomass, but not spatial structure, on semi-arid mine dumps. Ecol. Eng. 2022, 175, 106508. [Google Scholar] [CrossRef]
- Wang, J.; Wang, T.; Shi, T.; Wu, G.; Skidmore, A.K. A wavelet-based area parameter for indirectly estimating copper concentration in Carex Leaves from canopy reflectance. Remote Sens. 2015, 7, 15340–15360. [Google Scholar] [CrossRef]
- Dou, Z.; Cui, L.; Li, J.; Zhu, Y.; Gao, C.; Pan, X.; Lei, Y.; Zhang, M.; Zhao, X.; Li, W. Hyperspectral estimation of the chlorophyll content in short-term and long-term restorations of mangrove in Quanzhou Bay Estuary, China. Sustainability 2018, 10, 1127. [Google Scholar] [CrossRef]
- Liu, W.; Li, M.; Zhang, M.; Wang, D.; Guo, Z.; Long, S.; Yang, S.; Wang, H.; Li, W.; Hu, Y.; et al. Estimating leaf mercury content in Phragmites australis based on leaf hyperspectral reflectance. Ecosyst. Health Sustain. 2020, 6, 1726211. [Google Scholar] [CrossRef]
- Ren, H.; Zhao, Y.; Xiao, W.; Hu, H. A review of UAV monitoring in mining areas: Current status and future perspectives. Int. J. Coal Sci. Technol. 2019, 6, 320–333. [Google Scholar] [CrossRef]
- Nie, S.; Wang, C.; Zeng, H.; Xi, X.; Li, G. Above-ground biomass estimation using airborne discrete-return and full-waveform LiDAR data in a coniferous forest. Ecol. Indic. 2017, 78, 221–228. [Google Scholar] [CrossRef]
- Tang, J.; Liang, J.; Yang, Y.; Zhang, S.; Hou, H.; Zhu, X. Revealing the structure and composition of the restored vegetation cover in semi-arid mine dumps based on LiDAR and Hyperspectral Images. Remote Sens. 2022, 14, 978. [Google Scholar] [CrossRef]
- Ewald, M.; Aerts, R.; Lenoir, J.; Fassnacht, F.; Nicolas, M.; Skowronek, S.; Piat, J.; Honnay, O.; Garzón-López, C.; Feilhauer, H.; et al. LiDAR derived forest structure data improves predictions of canopy N and P concentrations from imaging spectroscopy. Remote Sens. Environ. 2018, 211, 13–25. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, T.; Guo, Y.; Skidmore, A.; Zhang, Z.; Tang, R.; Song, S.; Tang, Z. Estimating community-level plant functional traits in a species-rich alpine meadow using UAV image spectroscopy. Remote Sens. 2022, 14, 3399. [Google Scholar] [CrossRef]
- Bi, K.; Gao, S.; Niu, Z.; Zhang, C.; Huang, N. Estimating leaf chlorophyll and nitrogen contents using active hyperspectral LiDAR and partial least square regression method. J. Appl. Remote Sens. 2019, 13, 034513. [Google Scholar] [CrossRef]
- Yu, F.; Feng, S.; Yao, W.; Wang, D.; Xing, S.; Xu, T. BAS-ELM based UAV hyperspectral remote sensing inversion modeling of rice canopy nitrogen content. Int. J. Precis. Agric. Aviat. 2018, 1, 59–64. [Google Scholar] [CrossRef]
- Ye, X.; Abe, S.; Zhang, S. Estimation and mapping of nitrogen content in apple trees at leaf and canopy levels using hyperspectral imaging. Precis. Agric. 2020, 21, 198–225. [Google Scholar] [CrossRef]
- Cui, L.; Dou, Z.; Liu, Z.; Zuo, X.; Lei, Y.; Li, J.; Zhao, X.; Zhai, X.; Pan, X.; Li, W. Hyperspectral inversion of phragmites communis carbon, nitrogen, and phosphorus stoichiometry using three models. Remote Sens. 2020, 12, 1998. [Google Scholar] [CrossRef]
- Wang, Z.; Chen, J.; Zhang, J.; Tan, X.; Raza, M.A.; Ma, J.; Zhu, Y.; Yang, F.; Yang, W. Assessing canopy nitrogen and carbon content in maize by canopy spectral reflectance and uninformative variable elimination. Crop J. 2022, 10, 15. [Google Scholar] [CrossRef]
- Li, K.; Chen, Y.; Xu, Z.; Huang, X.; Hu, X.; Wang, X. Hyperspectral estimation method of chlorophyll content in Phyllostachys pubescens under pest stress. Spectrosc. Spectr. Anal. 2020, 40, 2578–2583. [Google Scholar] [CrossRef]
- Lohmann, P.; Koch, A.; Schaeffer, M. Approaches to the filtering of laser scanner data. Int. Arch. Photogramm. Remote Sens. 2000, 33, 540–547. [Google Scholar]
- Gitelson, A.A.; Keydan, G.P.; Merzlyak, M.N. Three-band model for noninvasive estimation of chlorophyll, carotenoids, and anthocyanin contents in higher plant leaves. Geophys. Res. Lett. 2006, 33, 431–433. [Google Scholar] [CrossRef]
- Maire, G.L.; Fran, O.C.; Dufrêne, E. Towards universal broad leaf chlorophyll indices using prospect simulated database and hyperspectral reflectance measurements. Remote Sens. Environ. 2004, 89, 1–28. [Google Scholar] [CrossRef]
- Richardson, A.J.; Wiegand, C.L. Distinguishing vegetation from soil background information. Photogramm. Eng. Remote Sens. 1977, 43, 100–120. [Google Scholar]
- Huete, A.; Justice, C.; Liu, H. Development of vegetation and soil indices for MODIS-EOS. Remote Sens. Environ. 1994, 49, 224–234. [Google Scholar] [CrossRef]
- Datt, B. A new reflectance index for remote sensing of chlorophyll content in higher plants: Tests using eucalyptus leaves. J. Plant Physiol. 1999, 154, 30–36. [Google Scholar] [CrossRef]
- Daughtry, C.S.T.; Walthall, C.L.; Kim, M.S.; De Colstoun, E.B.; McMurtrey Iii, J.E. Estimating corn leaf chlorophyll concentration from leaf and canopy re-flectance. Remote Sens. Environ. 2000, 74, 229–239. [Google Scholar] [CrossRef]
- Sims, D.A.; Gamon, J.A. Relationships between leaf pigment content and spectral reflectance across a wide range of species, leaf structures and developmental stages. Remote Sens. Environ. 2002, 81, 337–354. [Google Scholar] [CrossRef]
- Qi, J.; Chehbouni, A.; Huete, A.R.; Kerr, Y.H.; Sorooshian, S. A modified soil adjusted vegetation index. Remote Sens. Environ. 1994, 48, 119–126. [Google Scholar] [CrossRef]
- Driss, H.; John, R.M.; Elizabeth, P.; Pablo, J.Z.T.; Ian, B.S. Hyperspectral vegetation indices and novel algorithms for predicting green lai of crop canopies: Modeling and validation in the context of precision agriculture. Remote Sens. Environ. 2004, 10, 100–120. [Google Scholar] [CrossRef]
- Miller, J.R.; Hare, E.W.; Wu, J. Quantitative characterization of the vegetation red edge reflectance 1. An inverted-gaussian reflectance model. Int. J. Remote Sens. 1990, 11, 1755–1773. [Google Scholar] [CrossRef]
- Peuelas, J.; Gamon, J.A.; Fredeen, A.L.; Merino, J.; Field, C.B. Reflectance indices associated with physiological changes in nitrogen- and water-limited sunflower leaves. Remote Sens. Environ. 1994, 48, 135–146. [Google Scholar] [CrossRef]
- Rao, N.R.; Garg, P.K.; Ghosh, S.K.; Dadhwal, V.K. Estimation of leaf total chlorophyll and nitrogen concentrations using hyperspectral satellite imagery. J. Agric. Sci. 2008, 146, 65–75. [Google Scholar] [CrossRef]
- Thenot, F.; Méthy, M.; Winkel, T. The Photochemical Reflectance Index (PRI) as a water-stress index. Int. J. Remote Sens. 2002, 23, 5135–5139. [Google Scholar] [CrossRef]
- Blackburn, G.A. Spectral Indices for estimating photosynthetic pigment concentrations: A test using senescent tree leaves. Int. J. Remote Sens. 1998, 19, 657–675. [Google Scholar] [CrossRef]
- Metternicht, G. Vegetation indices derived from high-resolution airborne videography for precision crop management. Int. J. Remote Sens. 2003, 24, 2855–2877. [Google Scholar] [CrossRef]
- Schlerf, M.; Atzberger, C.; Hill, J. Remote sensing of forest biophysical variables using HyMap imaging spectrometer data. Remote Sens. Environ. 2005, 95, 177–194. [Google Scholar] [CrossRef]
- Merton, R.; Huntington, J. Early simulation of the ARIES-1 satellite sensor for multi-temporal vegetation research derived from AVIRIS. In Proceedings of the Summaries of the Eight JPL Airborne Earth Science Workshop, Pasadena, CA, USA, 8–11 February 1999; JPL Publication: Pasadena, CA, USA, 1999; Volume 99, pp. 299–307. [Google Scholar]
- Maccioni, A.; Agati, G.; Mazzinghi, P. New vegetation indices for remote measurement of chlorophylls based on leaf directional reflectance spectra. J. Photochem. Photobiol. B Biol. 2001, 61, 52–61. [Google Scholar] [CrossRef] [PubMed]
- Buschman, C.; Nagel, E. In vivo spectroscopy and internal optics of leaves as a basis for remote sensing of vegetation. Int. J. Remote Sens. 1993, 14, 711–722. [Google Scholar] [CrossRef]
- Huete, R.A. A Soil-Adjusted Vegetation Index (SAVI). Remote Sens. Environ. 1988, 20, 100–120. [Google Scholar] [CrossRef]
- McKee, T.B.; Doesken, N.J.; Kliest, J. The relationship of drought frequency and duration to time scales. In Proceedings of the Eighth Conference on Applied Climatology, American Meteorological Society, Anaheim, CA, USA, 17–22 January 1993; pp. 179–184. [Google Scholar]
- Broge, N.H.; Leblanc, E. Comparing prediction power and stability of broadband and hyperspectral vegetation indices for estimation of green leaf area index and canopy Chlorophyll density. Remote Sens. Environ. 2001, 76, 156–172. [Google Scholar] [CrossRef]
- Gitelson, A.A.; Kaufman, Y.J.; Stark, R.; Rundquist, D. Novel algorithms for remote estimation of vegetation fraction. Remote Sens. Environ. 2002, 80, 76–87. [Google Scholar] [CrossRef]
- Zarco-Tejada, P.J.; Miller, J.R.; Noland, T.L.; Mohammed, G.H.; Sampson, P.H. Scaling-up and model inversion methods with narrowband optical indices for chlorophyll content estimation in closed forest canopies with hyperspectral data. IEEE Trans. Geosci. Remote Sens. 2001, 39, 1491–1507. [Google Scholar] [CrossRef]
- Penuelas, J.; Pinol, J.; Ogaya, R.; Filella, I. Estimation of plant water concentration by the reflectance Water Index WI (R900/R970). Int. J. Remote Sens. 1997, 18, 2869–2875. [Google Scholar] [CrossRef]
- Kursa, M.B.; Rudnicki, W.R. Feature selection with the Boruta package. J. Stat. Softw. 2010, 36, 11. [Google Scholar] [CrossRef]
- Porder, S.; Asner, G.P.; Vitousek, P.M. Ground-based and remotely sensed nutrient availability across a tropical landscape. Proc. Natl. Acad. USA 2005, 102, 10909–10912. [Google Scholar] [CrossRef]
- Kokaly, R.F.; Asner, G.P.; Ollinger, S.V.; Martin, M.E.; Wessman, C.A. Characterizing canopy biochemistry from imaging spectroscopy and its application to ecosystem studies. Remote Sens. Environ. 2009, 113 (Suppl. 1), S78–S91. [Google Scholar] [CrossRef]
- Curran, P.J. Remote sensing of foliar chemistry. Remote Sens. Environ. 1989, 30, 271–278. [Google Scholar] [CrossRef]
- Wang, J.; Shi, T.; Liu, H.; Wu, G. Successive projections algorithm-based three-band vegetation index for foliar phosphorus estimation. Ecol. Indic. 2016, 67, 12–20. [Google Scholar] [CrossRef]
- Berger, K.; Verrelst, J.; Féret, J.B.; Hank, T.; Wocher, M.; Mauser, W.; Camps-Valls, G. Retrieval of aboveground crop nitrogen content with a hybrid machine learning method. Int. J. Appl. Earth Obs. Geoinf. 2020, 92, 174. [Google Scholar] [CrossRef]
- Homolová, L.; Malenovský, Z.; Clevers, J.G.P.W.; García-Santos, G.; Schaepman, M.E. Review of optical-based remote sensing for plant trait mapping. Ecol. Complex. 2013, 15, 1–16. [Google Scholar] [CrossRef]
- Huber, S.; Kneubühler, M.; Psomas, A.; Itten, K.; Zimmermann, N.E. Estimating foliar biochemistry from hyperspectral data in mixed forest canopy. For. Ecol. Manag. 2008, 256, 491–501. [Google Scholar] [CrossRef]
- van Deventer, H.; Cho, M.A.; Mutanga, O.; Ramoelo, A. Capability of models to predict leaf N and P across four seasons for six sub-tropical forest evergreen trees. ISPRS J. Photogramm. Remote Sens. 2015, 101, 209–220. [Google Scholar] [CrossRef]
- Peng, Y.; Zhang, M.; Xu, Z.; Yang, T.; Su, Y.; Zhou, T.; Wang, H.; Wang, Y.; Lin, Y. Estimation of leaf nutrition status in degraded vegetation based on field survey and hyperspectral data. Sci. Rep. 2020, 10, 4361. [Google Scholar] [CrossRef] [PubMed]
- Loozen, Y.; Rebel, K.T.; de Jong, S.M.; Lu, M.; Ollinger, S.V.; Wassen, M.J.; Karssenberg, D. Mapping canopy nitrogen in European forests using remote sensing and environmental variables with the random forests method. Remote Sens. Environ. 2020, 247, 111933. [Google Scholar] [CrossRef]
- Zhang, J.; He, N.; Liu, C.; Xu, L.; Chen, Z.; Li, Y.; Wang, R.; Yu, G.; Sun, W.; Xiao, C.; et al. Variation and evolution of C:N ratio among different organs enable plants to adapt to N-limited environments. Glob. Change Biol. 2020, 26, 2534–2543. [Google Scholar] [CrossRef]
- Song, S.; Xiong, K.; Chi, Y. Ecological stoichiometric characteristics of plant–soil–microorganism of grassland ecosystems under different restoration modes in the karst desertification area. Agronomy 2023, 13, 2016. [Google Scholar] [CrossRef]
- Koerselman, W.; Meuleman, A. The vegetation N:P ratio: A new tool to detect the nature of nutrient limitation. J. Appl. Ecol. 1996, 33, 1441–1450. [Google Scholar] [CrossRef]
- Yang, X.; Yuan, Z.; Ye, Y.; Wang, D.; Hua, K.; Guo, Z. Winter wheat total nitrogen content estimation based on UAV hyperspectral remote sensing. Spectrosc. Spectr. Anal. 2022, 42, 3269–3274. [Google Scholar] [CrossRef]
- Lapaz Olveira, A.; Saínz Rozas, H.; Castro-Franco, M.; Carciochi, W.; Nieto, L.; Balzarini, M.; Ciampitti, I.; Reussi Calvo, N. Monitoring corn nitrogen concentration from radar (C-SAR), optical, and sensor satellite data fusion. Remote Sens. 2023, 15, 824. [Google Scholar] [CrossRef]
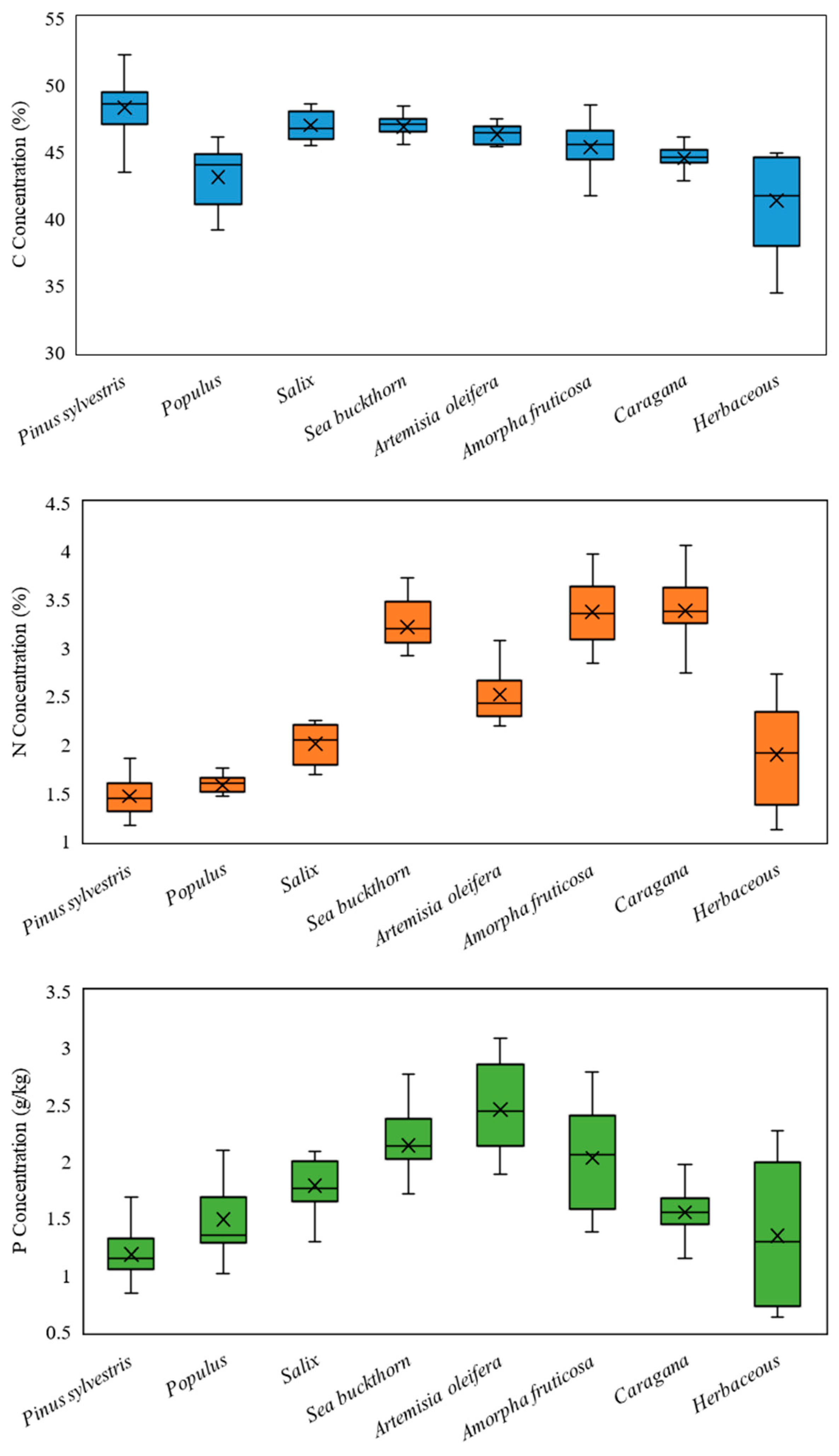







| Min | Max | Mean | Standard Deviation | |
|---|---|---|---|---|
| C (%) | 34.46 | 52.28 | 45.80 | 2.36 |
| N (%) | 1.09 | 4.06 | 2.62 | 0.87 |
| P (g/kg) | 0.63 | 3.08 | 1.73 | 0.49 |
| No. | Texture Feature | Formula |
|---|---|---|
| 1 | Mean | |
| 2 | Variance | |
| 3 | Entropy | |
| 4 | Data range | |
| 5 | Skewness |
| No. | Indices | Formula |
|---|---|---|
| 1 | CIgreen [36] | |
| 2 | CIred_edge [36] | |
| 3 | DD [37] | |
| 4 | DVI [38] | |
| 5 | EVI [39] | |
| 6 | GM [36] | |
| 7 | GNDVI [36] | |
| 8 | LCI [40] | |
| 9 | MCARI [41] | |
| 10 | mND705 [42] | |
| 11 | MSAVI [43] | |
| 12 | mSR705 [37] | |
| 13 | MTVI1 [44] | |
| 14 | NDI [36] | |
| 15 | NDVI [45] | |
| 16 | NPCI [46] | |
| 17 | PBI [47] | |
| 18 | PRI [48] | |
| 19 | PSNDa [49] | |
| 20 | PSNDb [49] | |
| 21 | PVR [50] | |
| 22 | RVI [51] | |
| 23 | RVSI [52] | |
| 24 | R680 [53] | Reflectance at 680 nm |
| 25 | R800 [54] | Reflectance at 800 nm |
| 26 | SAVI [55] | |
| 27 | SPI [56] | |
| 28 | SRPI [46] | |
| 29 | TVI [57] | |
| 30 | VARI [58] | |
| 31 | VOGa [59] | |
| 32 | VOG2 [59] | |
| 33 | WI [60] |
| No. | LiDAR Features | Variable Symbols |
|---|---|---|
| 1 | Maximum height | Hmax |
| 2 | Minimum height | Hmin |
| 3 | Average height | Hmean |
| 4 | Height kurtosis | Hkurt |
| 5 | Median | Hmedian_z |
| 6 | Height skewness | Hskew |
| 7 | Height standard deviation | Hstd |
| 8 | Height variance | Hvar |
| 9 | Canopy relief ratio | Hcrr |
| 10 | Canopy density metrics | d0, d1, d2, d3, d4, d5, d6, d7, d8, d9 |
| 11 | Height percentile | HP1st, HP5th, HP10th, HP20th, HP25th, HP30th, HP40th, HP50th, HP60th, HP70th, HP75th, HP80th, HP90th, HP95th, HP99th |
| 12 | Canopy height | CHM |
| 13 | Canopy cover | CC |
| 14 | Gap fraction | GF |
| 15 | Leaf area index | LAI |
| 16 | Foliage height diversity | FHD-1m, FHD-2m, FHD-3m |
| Model | Feature Combination | R2 | RMSE | MAE | |
|---|---|---|---|---|---|
| C | Causal bands | (970, 990 nm) | 0.07 | 5.16 | 3.64 |
| Multiple linear regression model | Hyperspectral features | 0.34 | 4.69 | 3.11 | |
| Hyperspectral + LiDAR | 0.42 | 2.24 | 2.13 | ||
| Random forest model | Hyperspectral features | 0.38 | 3.47 | 2.74 | |
| Hyperspectral + LiDAR | 0.56 | 1.19 | 1.00 | ||
| N | Causal bands | (510, 700–750, 910 nm) | 0.20 | 0.92 | 0.60 |
| Multiple linear regression model | Hyperspectral features | 0.46 | 0.87 | 0.52 | |
| Hyperspectral + LiDAR | 0.48 | 0.85 | 0.48 | ||
| Random forest model | Hyperspectral features | 0.53 | 0.57 | 0.46 | |
| Hyperspectral + LiDAR | 0.53 | 0.57 | 0.46 | ||
| P | Causal bands | (400–900 nm) | 0.32 | 0.52 | 0.45 |
| Multiple linear regression model | Hyperspectral features | 0.37 | 0.51 | 0.36 | |
| Hyperspectral + LiDAR | 0.39 | 0.50 | 0.36 | ||
| Random forest model | Hyperspectral features | 0.43 | 0.40 | 0.31 | |
| Hyperspectral + LiDAR | 0.44 | 0.40 | 0.31 |
| Hyperspectral Features | C | N | P | LiDAR Features | C | N | P |
|---|---|---|---|---|---|---|---|
| R399 | √ | d6 | √ | ||||
| R404 | √ | √ | d8 | √ | |||
| R409 | √ | d9 | √ | ||||
| R415 | √ | √ | Hmax | √ | |||
| R420 | √ | √ | Hmean | √ | √ | ||
| R451 | √ | Hmedian_z | √ | √ | |||
| R462 | √ | HP1st | √ | ||||
| R473 | √ | HP10th | √ | ||||
| R483 | √ | HP20th | √ | ||||
| R504 | √ | HP25th | √ | ||||
| R510 | √ | HP30th | √ | ||||
| R515 | √ | HP40th | √ | √ | |||
| R758 | √ | HP50th | √ | ||||
| R763 | √ | HP60th | √ | √ | |||
| EVI | √ | √ | HP70th | √ | √ | ||
| GNDVI | √ | HP75th | √ | √ | |||
| MTVI1 | √ | HP80th | √ | √ | √ | ||
| NPCI | √ | HP90th | √ | √ | |||
| PVR | √ | HP95th | √ | √ | |||
| R680 | √ | HP99th | √ | ||||
| SRPI | √ | Hstd | √ | √ | |||
| WI | √ | √ | Hvar | √ | √ | ||
| B1-Data Range | √ | CC | √ | ||||
| B1-Mean | √ | √ | CHM | √ | √ | ||
| B1-Variance | √ | FHD-1m | √ | √ | √ | ||
| B3-Data Range | √ | √ | FHD-2m | √ | |||
| B3-Variance | √ | √ | FHD-3m | √ | |||
| / | / | / | / | GF | √ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, Y.; Dong, J.; Tang, J.; Zhao, J.; Lei, S.; Zhang, S.; Chen, F. Mapping Foliar C, N, and P Concentrations in An Ecological Restoration Area with Mixed Plant Communities Based on LiDAR and Hyperspectral Data. Remote Sens. 2024, 16, 1624. https://doi.org/10.3390/rs16091624
Yang Y, Dong J, Tang J, Zhao J, Lei S, Zhang S, Chen F. Mapping Foliar C, N, and P Concentrations in An Ecological Restoration Area with Mixed Plant Communities Based on LiDAR and Hyperspectral Data. Remote Sensing. 2024; 16(9):1624. https://doi.org/10.3390/rs16091624
Chicago/Turabian StyleYang, Yongjun, Jing Dong, Jiajia Tang, Jiao Zhao, Shaogang Lei, Shaoliang Zhang, and Fu Chen. 2024. "Mapping Foliar C, N, and P Concentrations in An Ecological Restoration Area with Mixed Plant Communities Based on LiDAR and Hyperspectral Data" Remote Sensing 16, no. 9: 1624. https://doi.org/10.3390/rs16091624
APA StyleYang, Y., Dong, J., Tang, J., Zhao, J., Lei, S., Zhang, S., & Chen, F. (2024). Mapping Foliar C, N, and P Concentrations in An Ecological Restoration Area with Mixed Plant Communities Based on LiDAR and Hyperspectral Data. Remote Sensing, 16(9), 1624. https://doi.org/10.3390/rs16091624






