First Detection and Identification of Southern Tomato Virus Infecting Tomatoes in Oklahoma with Complete Genome Characterization and Insights into Global Genetic Diversity
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection and Total RNA Extraction
2.2. High Throughput Sequencing and RT-PCR Confirmation
2.3. Multiple Sequence Alignment and Phylogenetic Analysis
2.4. Analysis of Genetic Diversity and Population Structure of STV
2.5. Recombination Detection and Pairwise Identity Analysis
2.6. Bayesian Evolutionary Analysis Sampling Trees (BEAST) Analysis
2.7. Protein Disorder Analysis
2.8. Analysis of Selective Pressure
3. Results
3.1. Genome Assembly of STV Isolates and RT-PCR Confirmation
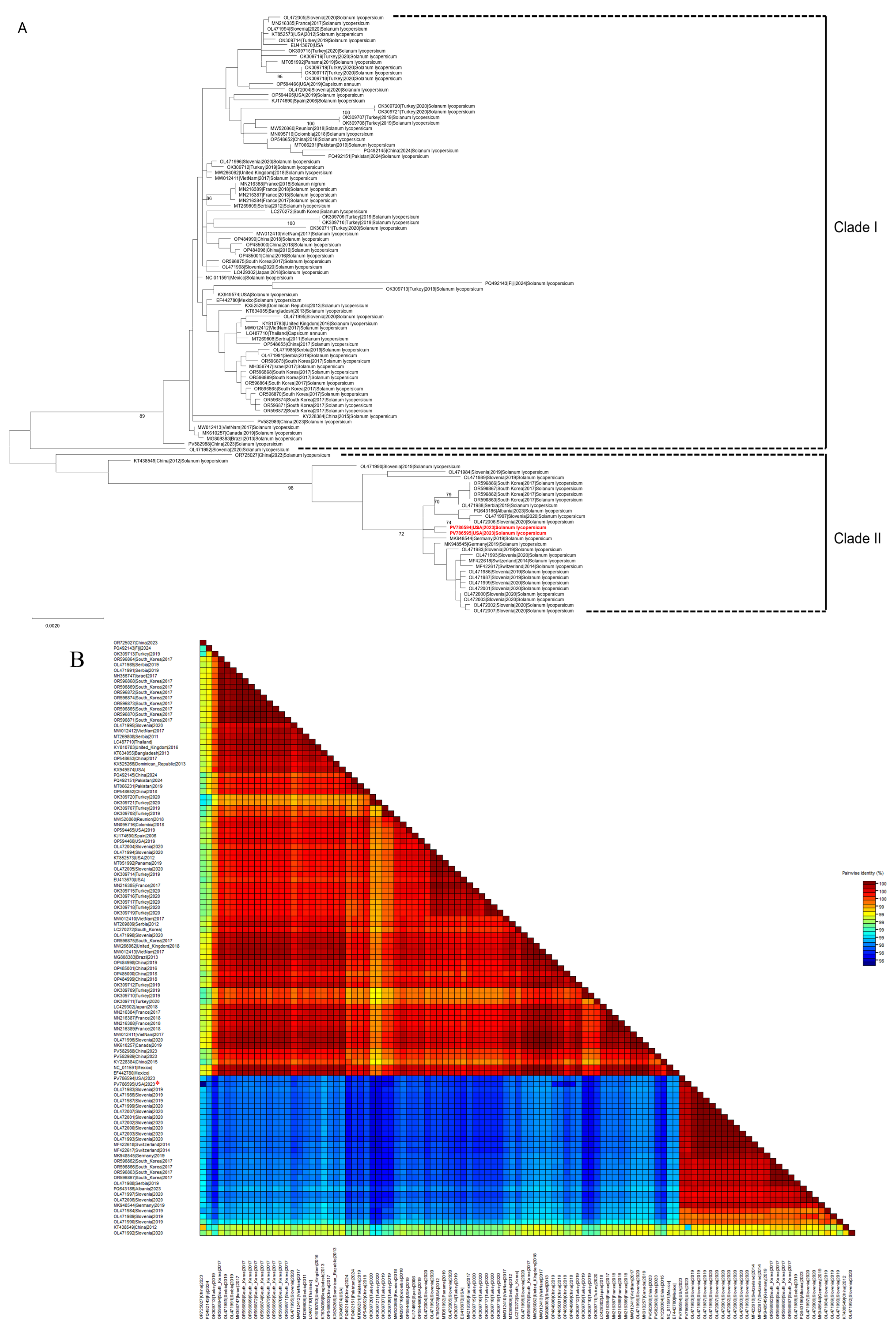
3.2. Phylogenetic Analysis
3.3. Genetic Diversity, Genetic Differentiation, and Migration in STV Populations
3.4. Recombination in the STV Population
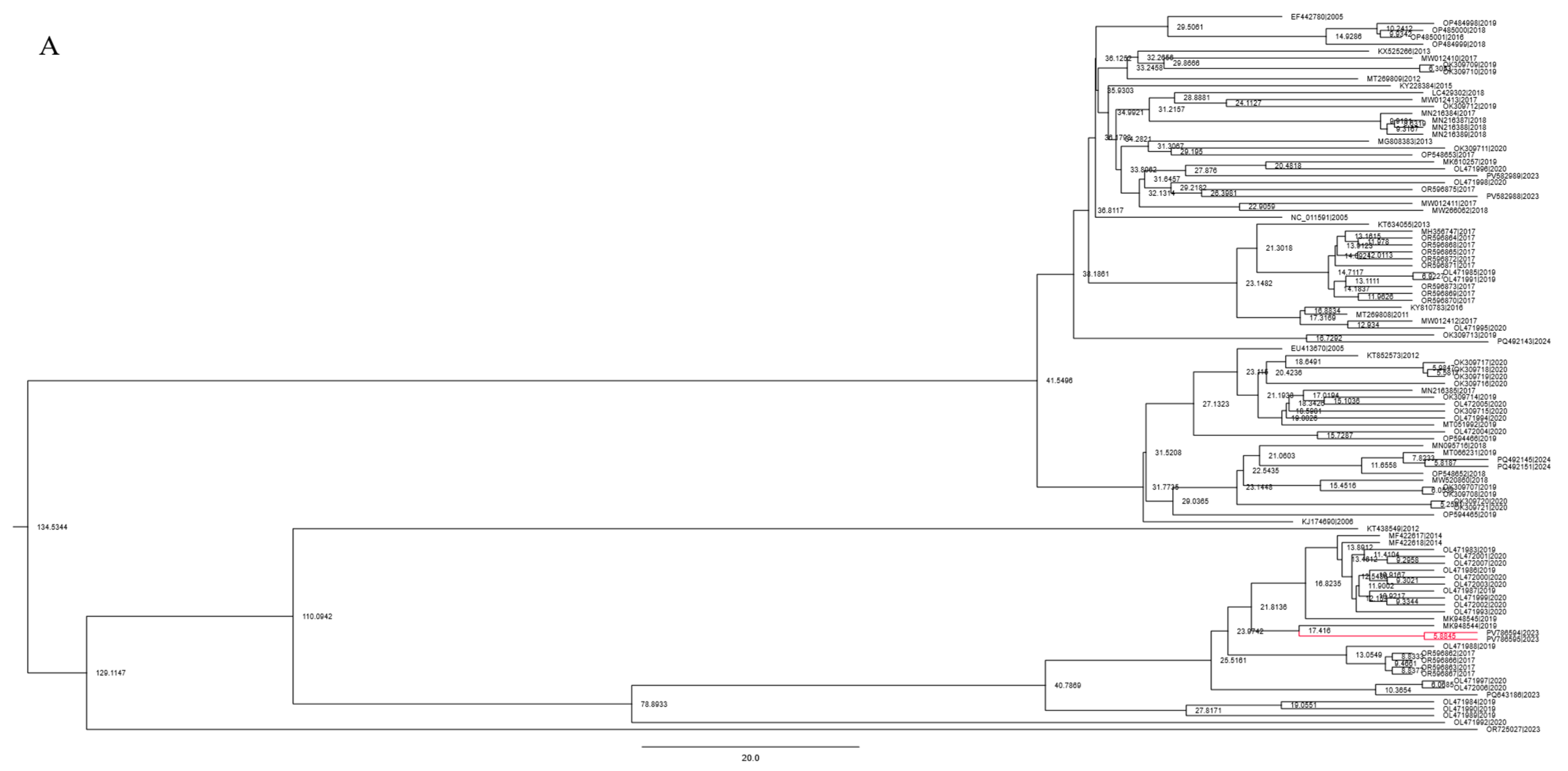
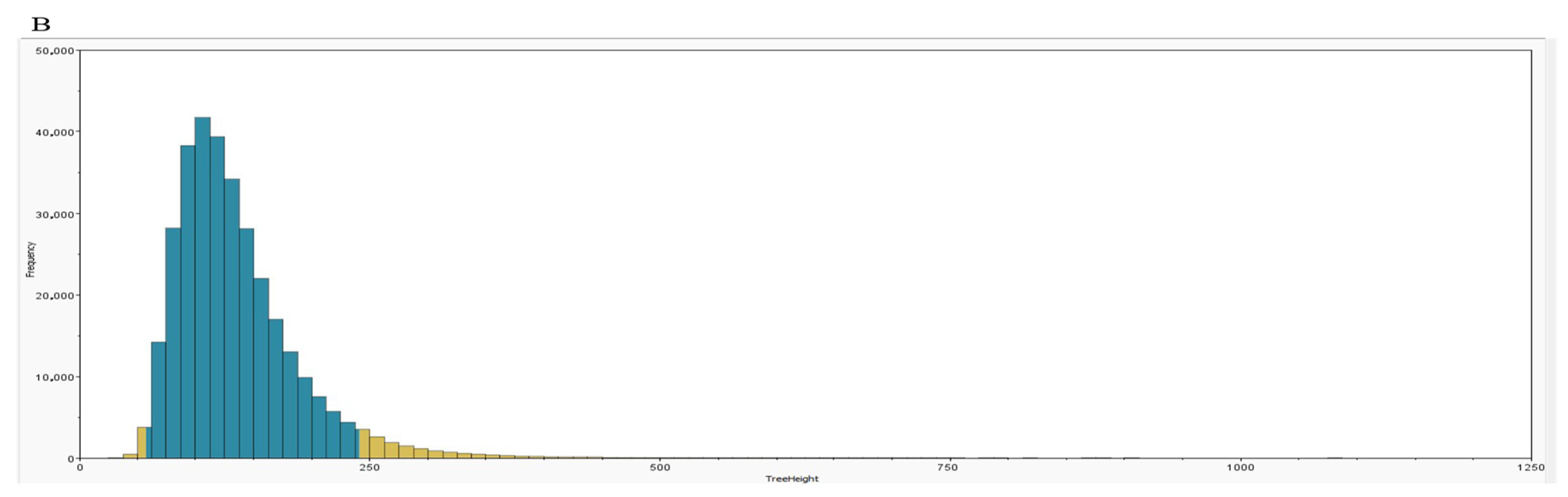
3.5. Bayesian Phylogenetic Analysis and Substitution Rate
3.6. Comparison of Intrinsically Disordered Regions in STV Proteins
3.7. Selective Pressure Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Panno, S.; Davino, S.; Caruso, A.G.; Bertacca, S.; Crnogorac, A.; Mandić, A.; Noris, E.; Matić, S. A Review of the Most Common and Economically Important Diseases That Undermine the Cultivation of Tomato Crop in the Mediterranean Basin. Agronomy 2021, 11, 2188. [Google Scholar] [CrossRef]
- Harvey, M.; Quilley, S.; Beynon, H. Exploring the Tomato: Transformations in Nature, Economy and Society; Edward Elgar: Cheltenham, UK, 2002. [Google Scholar]
- Raiola, A.; Rigano, M.M.; Calafiore, R.; Frusciante, L.; Barone, A. Enhancing the Health-Promoting Effects of Tomato Fruit for Biofortified Food. Mediat. Inflamm. 2014, 2014, 139873. [Google Scholar] [CrossRef]
- Food and Agriculture Organization of the United Nations (FAO). FAOSTAT Statistical Database. 2024. Available online: https://www.fao.org/faostat/en/#home (accessed on 21 March 2025).
- Rashid, T.S.; Sijam, K.; Awla, H.K.; Saud, H.M.; Kadir, J. Pathogenicity Assay and Molecular Identification of Fungi and Bacteria Associated with Diseases of Tomato in Malaysia. Am. J. Plant Sci. 2016, 7, 949–957. [Google Scholar] [CrossRef]
- Sabanadzovic, S.; Valverde, R.A.; Brown, J.K.; Martin, R.R.; Tzanetakis, I.E. Southern Tomato Virus: The Link Between the Families Totiviridae and Partitiviridae. Virus Res. 2009, 140, 130–137. [Google Scholar] [CrossRef]
- Magdalena, C.; Slimen, A.B.; Mitri, E.; Orges, C.; Frasheri, D.; Merkuri, J.; Parrella, G.; Elbeaino, T. Molecular Detection and Characterization of Viruses Infecting Greenhouse-Grown Tomatoes in Albania. Phytopathol. Mediterr. 2025, 64, 77–86. [Google Scholar] [CrossRef]
- Padmanabhan, C.; Zheng, Y.; Li, R.; Fei, Z.; Ling, K.-S. Complete Genome Sequence of Southern Tomato Virus Naturally Infecting Tomatoes in Bangladesh. Genome Announc. 2015, 3, e01522-15. [Google Scholar] [CrossRef] [PubMed]
- Padmanabhan, C.; Zheng, Y.; Li, R.; Sun, S.-E.; Zhang, D.; Liu, Y.; Fei, Z.; Ling, K.-S. Complete Genome Sequence of Southern Tomato Virus Identified in China Using Next-Generation Sequencing. Genome Announc. 2015, 3, e01226-15. [Google Scholar] [CrossRef]
- Xu, C.; Sun, X.; Taylor, A.; Jiao, C.; Xu, Y.; Cai, X.; Wang, X.; Ge, C.; Pan, G.; Wang, Q.; et al. Diversity, Distribution, and Evolution of Tomato Viruses in China Uncovered by Small RNA Sequencing. J. Virol. 2017, 91, 10–1128. [Google Scholar] [CrossRef]
- Hussain, M.D.; Farooq, T.; Chen, X.; Jiang, T.; Zang, L.; Shakeel, M.T.; Zhou, T. Molecular Detection of Southern Tomato Amalgavirus Prevalent in Tomatoes and Its Genomic Characterization with Global Evolutionary Dynamics. Viruses 2022, 14, 2481. [Google Scholar] [CrossRef]
- Bados, J.P.; Gómez, J.; Pérez, A.; Salazar, J.; Marín, M. Molecular Detection of Three RNA Viruses in Tomato (Solanum Lycopersicum L.) Leaf Tissue and Seeds in Antioquia (Colombia). Rev. Colomb. De Cienc. Hortícolas 2025, 19, e18657. [Google Scholar]
- Candresse, T.; Marais, A.; Faure, C. First Report of Southern Tomato Virus on Tomatoes in Southwest France. Plant Dis. 2013, 97, 1124. [Google Scholar] [CrossRef] [PubMed]
- Gaafar, Y.; Lüddecke, P.; Heidler, C.; Hartrick, J.; Sieg-Müller, A.; Hübert, C.; Wichura, A.; Ziebell, H. First Report of Southern Tomato Virus in German Tomatoes. New Dis. Rep 2019, 40, 1. [Google Scholar] [CrossRef]
- James, A.; Andronis, C.; Kryovrysanaki, N.; Goumenaki, E.; Kalantidis, K.; Katsarou, K. First Report of SOUTHERN Tomato Virus from Tomato (Solanum Lycopersicum) in Greece. Plant Dis. 2023, 107, 237. [Google Scholar] [CrossRef]
- Iacono, G.; Hernandez-Llopis, D.; Alfaro-Fernandez, A.; Davino, M.; Font, M.; Panno, S.; Galipenso, L.; Rubio, L.; Davino, S. First Report of Southern Tomato Virus in Tomato Crops in Italy. New Dis. Rep. 2015, 32, 27. [Google Scholar] [CrossRef]
- Oh, J.; Lee, H.-K.; Park, C.-Y.; Yeom, Y.-A.; Min, H.-G.; Yang, H.-J.; Jeong, R.-D.; Kim, H.; Moon, J.-S.; Lee, S.-H. First Report of Southern Tomato Virus in Tomato (Solanum Lycopersicum) in Korea. Plant Dis. 2018, 102, 1467. [Google Scholar] [CrossRef]
- Galipienso, L.; Elvira-González, L.; Velasco, L.; Herrera-Vásquez, J.Á.; Rubio, L. Detection of Persistent Viruses by High-Throughput Sequencing in Tomato and Pepper from Panama: Phylogenetic and Evolutionary Studies. Plants 2021, 10, 2295. [Google Scholar] [CrossRef]
- Vučurović, A.; Kutnjak, D.; Mehle, N.; Stanković, I.; Pecman, A.; Bulajić, A.; Krstić, B.; Ravnikar, M. Detection of Four New Tomato Viruses in Serbia Using Post Hoc High-Throughput Sequencing Analysis of Samples from a Large-Scale Field Survey. Plant Dis. 2021, 105, 2325–2332. [Google Scholar] [CrossRef]
- Verbeek, M.; Dullemans, A.; Espino, A.; Botella, M.; Alfaro-Fernández, A.; Font, M. First Report of Southern Tomato Virus in Tomato in the Canary Islands, Spain. J. Plant Pathol. 2015, 97, 392. [Google Scholar]
- Randa-Zelyüt, F.; Fox, A.; Karanfil, A. Population Genetic Dynamics of Southern Tomato Virus from Turkey. J. Plant Pathol. 2023, 105, 211–224. [Google Scholar] [CrossRef]
- Harju, V.; Skelton, A.; Lazenby, M.; Rimmer, T.; Buxton-Kirk, A.; Fowkes, A.; Forde, S.; Ward, R.; Frew, L.; Barker, R.; et al. First Detection, Symptoms and Management of Southern Tomato Virus in the United Kingdom. New Dis. Rep. 2021, 43, e12014. [Google Scholar] [CrossRef]
- Grenfell, B.T.; Pybus, O.G.; Gog, J.R.; Wood, J.L.; Daly, J.M.; Mumford, J.A.; Holmes, E.C. Unifying the Epidemiological and Evolutionary Dynamics of Pathogens. Science 2004, 303, 327–332. [Google Scholar] [CrossRef]
- Acosta-Leal, R.; Duffy, S.; Xiong, Z.; Hammond, R.; Elena, S.F. Advances in Plant Virus Evolution: Translating Evolutionary Insights into Better Disease Management. Phytopathology 2011, 101, 1136–1148. [Google Scholar] [CrossRef]
- Elvira-González, L.; Rubio, L.; Galipienso, L. Geographically Distant Isolates of the Persistent Southern Tomato Virus (STV) Show Very Low Genetic Diversity in the Putative Coat Protein Gene. Virus Genes 2020, 56, 668–672. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Suleski, M.; Sanderford, M.; Sharma, S.; Tamura, K. MEGA12: Molecular Evolutionary Genetic Analysis Version 12 for Adaptive and Green Computing. Mol. Biol. Evol. 2024, 41, msae263. [Google Scholar] [CrossRef] [PubMed]
- Rozas, J.; Ferrer-Mata, A.; Sánchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sánchez-Gracia, A. DnaSP 6: DNA Sequence Polymorphism Analysis of Large Data Sets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef] [PubMed]
- Hudson, R.R.; Boos, D.D.; Kaplan, N.L. A Statistical Test for Detecting Geographic Subdivision. Mol. Biol. Evol. 1992, 9, 138–151. [Google Scholar] [CrossRef] [PubMed]
- Hudson, R.R. A New Statistic for Detecting Genetic Differentiation. Genetics 2000, 155, 2011–2014. [Google Scholar] [CrossRef]
- Rousset, F. Genetic Differentiation and Estimation of Gene Flow from f-Statistics Under Isolation by Distance. Genetics 1997, 145, 1219–1228. [Google Scholar] [CrossRef]
- Jin, W.; Zhang, Y.; Su, X.; Xie, Z.; Wang, R.; Du, Z.; Wang, Y.; Qiu, Y. Genetic Diversity Analysis of Lychnis Mottle Virus and First Identification of Angelica Sinensis Infection. Heliyon 2023, 9, e17006. [Google Scholar] [CrossRef]
- Martin, D.P.; Murrell, B.; Golden, M.; Khoosal, A.; Muhire, B. RDP4: Detection and Analysis of Recombination Patterns in Virus Genomes. Virus Evol. 2015, 1, vev003. [Google Scholar] [CrossRef]
- Weaver, S.; Shank, S.D.; Spielman, S.J.; Li, M.; Muse, S.V.; Kosakovsky Pond, S.L. Datamonkey 2.0: A Modern Web Application for Characterizing Selective and Other Evolutionary Processes. Mol. Biol. Evol. 2018, 35, 773–777. [Google Scholar] [CrossRef] [PubMed]
- Muhire, B.M.; Varsani, A.; Martin, D.P. SDT: A Virus Classification Tool Based on Pairwise Sequence Alignment and Identity Calculation. PLoS ONE 2014, 9, e108277. [Google Scholar] [CrossRef] [PubMed]
- Bouckaert, R.; Vaughan, T.G.; Barido-Sottani, J.; Duchêne, S.; Fourment, M.; Gavryushkina, A.; Heled, J.; Jones, G.; Kühnert, D.; De Maio, N.; et al. BEAST 2.5: An Advanced Software Platform for Bayesian Evolutionary Analysis. PLoS Comput. Biol. 2019, 15, e1006650. [Google Scholar] [CrossRef] [PubMed]
- Rambaut, A.; Drummond, A.J.; Xie, D.; Baele, G.; Suchard, M.A. Posterior Summarization in Bayesian Phylogenetics Using Tracer 1.7. Syst. Biol. 2018, 67, 901–904. [Google Scholar] [CrossRef]
- Charon, J.; Theil, S.; Nicaise, V.; Michon, T. Protein Intrinsic Disorder Within the Potyvirus Genus: From Proteome-Wide Analysis to Functional Annotation. Mol. Biosyst. 2016, 12, 634–652. [Google Scholar] [CrossRef]
- Charon, J.; Barra, A.; Walter, J.; Millot, P.; Hébrard, E.; Moury, B.; Michon, T. First Experimental Assessment of Protein Intrinsic Disorder Involvement in an RNA Virus Natural Adaptive Process. Mol. Biol. Evol. 2018, 35, 38–49. [Google Scholar] [CrossRef]
- Walter, J.; Charon, J.; Hu, Y.; Lachat, J.; Leger, T.; Lafforgue, G.; Barra, A.; Michon, T. Comparative Analysis of Mutational Robustness of the Intrinsically Disordered Viral Protein VPg and of Its Interactor eIF4E. PLoS ONE 2019, 14, e0211725. [Google Scholar] [CrossRef]
- Byrne, M.; Kashyap, A.; Esquirol, L.; Ranson, N.; Sainsbury, F. The Structure of a Plant-Specific Partitivirus Capsid Reveals a Unique Coat Protein Domain Architecture with an Intrinsically Disordered Protrusion. Commun. Biol. 2021, 4, 1155. [Google Scholar] [CrossRef]
- LaTourrette, K.; Holste, N.M.; Garcia-Ruiz, H. Polerovirus Genomic Variation. Virus Evol. 2021, 7, veab102. [Google Scholar] [CrossRef]
- Mészáros, B.; Erdős, G.; Dosztányi, Z. IUPred2A: Context-Dependent Prediction of Protein Disorder as a Function of Redox State and Protein Binding. Nucleic Acids Res. 2018, 46, W329–W337. [Google Scholar] [CrossRef]
- Murrell, B.; Moola, S.; Mabona, A.; Weighill, T.; Sheward, D.; Kosakovsky Pond, S.L.; Scheffler, K. FUBAR: A Fast, Unconstrained Bayesian Approximation for Inferring Selection. Mol. Biol. Evol. 2013, 30, 1196–1205. [Google Scholar] [CrossRef] [PubMed]
- Murrell, B.; Wertheim, J.O.; Moola, S.; Weighill, T.; Scheffler, K.; Kosakovsky Pond, S.L. Detecting Individual Sites Subject to Episodic Diversifying Selection. PLoS Genet. 2012, 8, e1002764. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N. What Does It Mean to Be Natively Unfolded? Eur. J. Biochem. 2002, 269, 2–12. [Google Scholar] [CrossRef] [PubMed]
- Szilagyi, A.; Györffy, D.; Zavodszky, P. The Twilight Zone Between Protein Order and Disorder. Biophys. J. 2008, 95, 1612–1626. [Google Scholar] [CrossRef]
- Rivarez, M.P.S.; Vučurović, A.; Mehle, N.; Ravnikar, M.; Kutnjak, D. Global Advances in Tomato Virome Research: Current Status and the Impact of High-Throughput Sequencing. Front. Microbiol. 2021, 12, 671925. [Google Scholar] [CrossRef]
- Fukuhara, T.; Tabara, M.; Koiwa, H.; Takahashi, H. Effect of Asymptomatic Infection with Southern Tomato Virus on Tomato Plants. Arch. Virol. 2020, 165, 11–20. [Google Scholar] [CrossRef]
- Elvira Gonzalez, L.; Peiró, R.; Rubio, L.; Galipienso, L. Persistent Southern Tomato Virus (STV) Interacts with Cucumber Mosaic and/or Pepino Mosaic Virus in Mixed-Infections Modifying Plant Symptoms, Viral Titer and Small RNA Accumulation. Microorganisms 2021, 9, 689. [Google Scholar] [CrossRef]
- Van De Vossenberg, B.T.; Visser, M.; Bruinsma, M.; Koenraadt, H.M.; Westenberg, M.; Botermans, M. Real-Time Tracking of Tomato Brown Rugose Fruit Virus (ToBRFV) Outbreaks in the Netherlands Using Nextstrain. PLoS ONE 2020, 15, e0234671. [Google Scholar] [CrossRef]
- Steinhauer, D.A.; Domingo, E.; Holland, J.J. Lack of Evidence for Proofreading Mechanisms Associated with an RNA Virus Polymerase. Gene 1992, 122, 281–288. [Google Scholar] [CrossRef]
- Laemmlen, F.; Van Maren, A.; Endo, R.; Valverde, R. A Tomato Decline of Unknown Etiology in Imperial Valley. CA Phytopathol. 1985, 75, 1287. [Google Scholar]
- Alcalá-Briseño, R.I.; Coşkan, S.; Londoño, M.A.; Polston, J.E. Genome Sequence of Southern Tomato Virus in Asymptomatic Tomato “Sweet Hearts”. Genome Announc. 2017, 5, 10–1128. [Google Scholar] [CrossRef]
- Dias, N.P.; Hu, R.; Hale, F.A.; Hansen, Z.R.; Wszelaki, A.; Domier, L.L.; Hajimorad, M. Viromes of Field-Grown Tomatoes and Peppers in Tennessee Revealed by RNA Sequencing Followed by Bioinformatic Analysis. Plant Health Prog. 2023, 24, 207–213. [Google Scholar] [CrossRef]
- Ma, Y.; Marais, A.; Lefebvre, M.; Faure, C.; Candresse, T. Metagenomic Analysis of Virome Cross-Talk Between Cultivated Solanum Lycopersicum and Wild Solanum Nigrum. Virology 2020, 540, 38–44. [Google Scholar] [CrossRef] [PubMed]
- Murphy, J.F.; Bowen, K.L. Synergistic Disease in Pepper Caused by the Mixed Infection of Cucumber Mosaic Virus and Pepper Mottle Virus. Phytopathology 2006, 96, 240–247. [Google Scholar] [CrossRef]
- Syller, J. Facilitative and Antagonistic Interactions Between Plant Viruses in Mixed Infections. Mol. Plant Pathol. 2012, 13, 204–216. [Google Scholar] [CrossRef]
- Krupovic, M.; Dolja, V.V.; Koonin, E.V. Plant Viruses of the Amalgaviridae Family Evolved via Recombination Between Viruses with Double-Stranded and Negative-Strand RNA Genomes. Biol. Direct 2015, 10, 12. [Google Scholar] [CrossRef]
- Duffy, S.; Shackelton, L.A.; Holmes, E.C. Rates of Evolutionary Change in Viruses: Patterns and Determinants. Nat. Rev. Genet. 2008, 9, 267–276. [Google Scholar] [CrossRef]
- Chao, L.; Rang, C.U.; Wong, L.E. Distribution of Spontaneous Mutants and Inferences about the Replication Mode of the RNA Bacteriophage φ6. J. Virol. 2002, 76, 3276–3281. [Google Scholar] [CrossRef]
- Choi, H.; Jo, Y.; Chung, H.; Choi, S.Y.; Kim, S.-M.; Hong, J.-S.; Lee, B.C.; Cho, W.K. Phylogenetic and Phylodynamic Analyses of Soybean Mosaic Virus Using 305 Coat Protein Gene Sequences. Plants 2022, 11, 3256. [Google Scholar] [CrossRef]
- Mao, Y.; Sun, X.; Shen, J.; Gao, F.; Qiu, G.; Wang, T.; Nie, X.; Zhang, W.; Gao, Y.; Bai, Y. Molecular Evolutionary Analysis of Potato Virus y Infecting Potato Based on the VPg Gene. Front. Microbiol. 2019, 10, 1708. [Google Scholar] [CrossRef]
- Zhai, W.; Nielsen, R.; Slatkin, M. An Investigation of the Statistical Power of Neutrality Tests Based on Comparative and Population Genetic Data. Mol. Biol. Evol. 2009, 26, 273–283. [Google Scholar] [CrossRef]
- Sztuba-Solińska, J.; Urbanowicz, A.; Figlerowicz, M.; Bujarski, J.J. RNA-RNA Recombination in Plant Virus Replication and Evolution. Annu. Rev. Phytopathol. 2011, 49, 415–443. [Google Scholar] [CrossRef]
- Kozlowski, L.P.; Bujnicki, J.M. MetaDisorder: A Meta-Server for the Prediction of Intrinsic Disorder in Proteins. BMC Bioinform. 2012, 13, 111. [Google Scholar] [CrossRef]
- Katuwawala, A.; Oldfield, C.J.; Kurgan, L. Accuracy of Protein-Level Disorder Predictions. Brief. Bioinform. 2020, 21, 1509–1522. [Google Scholar] [CrossRef]
- Wright, P.E.; Dyson, H.J. Intrinsically Unstructured Proteins: Re-Assessing the Protein Structure-Function Paradigm. J. Mol. Biol. 1999, 293, 321–331. [Google Scholar] [CrossRef]
- Uversky, V.N. Intrinsically Disordered Proteins and Their “Mysterious”(meta) Physics. Front. Phys. 2019, 7, 10. [Google Scholar] [CrossRef]
- Gitlin, L.; Hagai, T.; LaBarbera, A.; Solovey, M.; Andino, R. Rapid Evolution of Virus Sequences in Intrinsically Disordered Protein Regions. PLoS Pathog. 2014, 10, e1004529. [Google Scholar] [CrossRef]
- Xue, B.; Blocquel, D.; Habchi, J.; Uversky, A.V.; Kurgan, L.; Uversky, V.N.; Longhi, S. Structural Disorder in Viral Proteins. Chem. Rev. 2014, 114, 6880–6911. [Google Scholar] [CrossRef] [PubMed]
- Mishra, P.M.; Verma, N.C.; Rao, C.; Uversky, V.N.; Nandi, C.K. Intrinsically Disordered Proteins of Viruses: Involvement in the Mechanism of Cell Regulation and Pathogenesis. Prog. Mol. Biol. Transl. Sci. 2020, 174, 1–78. [Google Scholar] [PubMed]
- Lafforgue, G.; Michon, T.; Charon, J. Analysis of the Contribution of Intrinsic Disorder in Shaping Potyvirus Genetic Diversity. Viruses 2022, 14, 1959. [Google Scholar] [CrossRef] [PubMed]
| Primer Name | Sequence (5′-3′) | Annealing Temperature (°C) | Amplicon Size (bp) |
|---|---|---|---|
| STV 1F | GCGAGAGCGATAAATTTAGTAAGCTAC | 53 | 673 |
| STV 1R | TTGACAATCTTACGCTGCAGATCAG | ||
| STV 2F | GAGAAGAGGACACTGCAGTACAA | 54 | 503 |
| STV 2R | GTAGATATCCTCCATCAGACTCT |
| Gene | No. of Sequences | Total no. of Nucleotide Sites | No. of Nucleotide Sites a | S | H | Hd | k | π | Eta | Theta (Per Sequence) | Theta (Per Site) | Tajima’s Test | Fu and Li’s D Test | Fu and Li’s F Test |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Complete genome | 108 | 3475 | 3190 | 184 | 70 | 0.979 | 23.64123 | 0.00741 | 193 | 36.72895 | 0.01151 | −1.18568 NS | −3.22915 * | −2.79709 * |
| p42 | 130 | 1134 | 1134 | 67 | 47 | 0.898 | 6.23280 | 0.00550 | 69 | 12.68173 | 0.01118 | −1.60530 NS | −3.21261 * | −3.02698 * |
| RdRp | 108 | 2289 | 2289 | 129 | 60 | 0.960 | 17.48165 | 0.00764 | 138 | 26.26215 | 0.01147 | −1.10373 NS | −2.62916 * | −2.35996 NS |
| Fusion protein | 108 | 3190 | 3190 | 184 | 70 | 0.979 | 23.64123 | 0.00741 | 193 | 37.72895 | 0.01151 | −1.18568 NS | −3.22915 * | −2.79709 * |
| Gene | Date Range | Mean Substitution Rate (Subs/Site/Year) | HPD a | TMRCA b | HPD a |
|---|---|---|---|---|---|
| Fusion protein | 2005–2024 | 6.514 × 10−5 | 3.4575 × 10−5–9.6661 × 10−5 | 135.137 | 56.8965–241.3992 |
| p42 | 2005–2024 | 1.184 × 10−4 | 5.7164 × 10−5–1.8674 × 10−4 | 58.546 | 23.4486–102.1011 |
| RdRp | 2005–2024 | 4.866 × 10−5 | 2.0639 × 10−5–7.7596 × 10−5 | 188.263 | 64.1967–368.4334 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jindal, S.; Ali, A. First Detection and Identification of Southern Tomato Virus Infecting Tomatoes in Oklahoma with Complete Genome Characterization and Insights into Global Genetic Diversity. Viruses 2025, 17, 1193. https://doi.org/10.3390/v17091193
Jindal S, Ali A. First Detection and Identification of Southern Tomato Virus Infecting Tomatoes in Oklahoma with Complete Genome Characterization and Insights into Global Genetic Diversity. Viruses. 2025; 17(9):1193. https://doi.org/10.3390/v17091193
Chicago/Turabian StyleJindal, Salil, and Akhtar Ali. 2025. "First Detection and Identification of Southern Tomato Virus Infecting Tomatoes in Oklahoma with Complete Genome Characterization and Insights into Global Genetic Diversity" Viruses 17, no. 9: 1193. https://doi.org/10.3390/v17091193
APA StyleJindal, S., & Ali, A. (2025). First Detection and Identification of Southern Tomato Virus Infecting Tomatoes in Oklahoma with Complete Genome Characterization and Insights into Global Genetic Diversity. Viruses, 17(9), 1193. https://doi.org/10.3390/v17091193







