DDX21, a Host Restriction Factor of FMDV IRES-Dependent Translation and Replication
Abstract
:1. Introduction
2. Materials and Methods
2.1. Cell Lines, Viruses, and Plasmid Constructs
2.2. Antibodies and Reagents
2.3. Quantitative Real-Time PCR
2.4. Knockdown and Overexpression
2.5. TCID50
2.6. Western Blot
2.7. Dual-Luciferase Assay
2.8. In Vitro Transcription
2.9. RNA Pulldown Assay
2.10. Nuclear Cytosol Fractionation Assay
2.11. Confocal Microscopy
2.12. Virus Infection
2.13. Proteasome, Lysosome, and Caspase Inhibitor Assays
2.14. RNA Immunoprecipitation and RTPCR
2.15. Statistical Analysis
3. Results
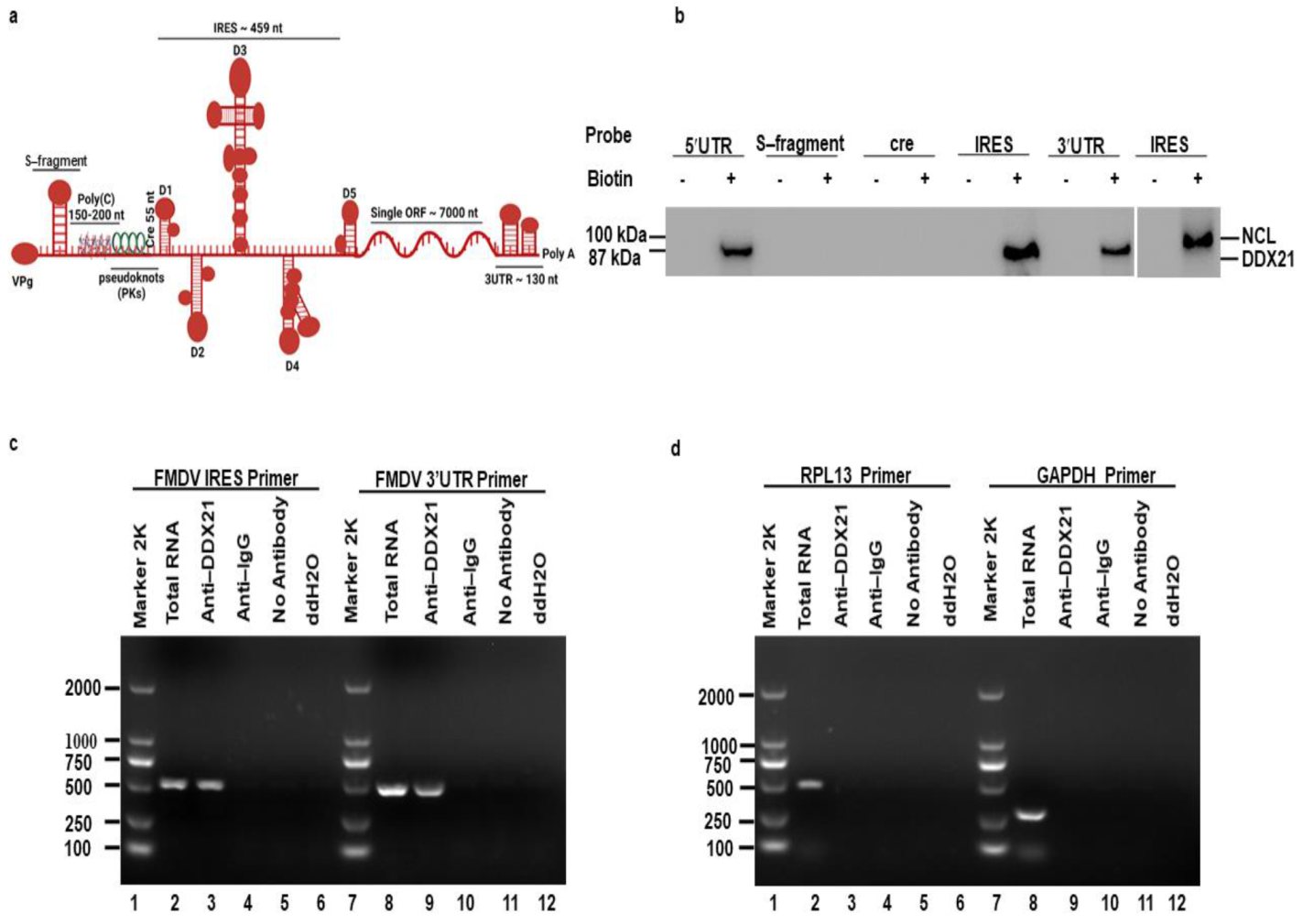
3.1. DDX21 Co-Precipitates with the FMDV IRES
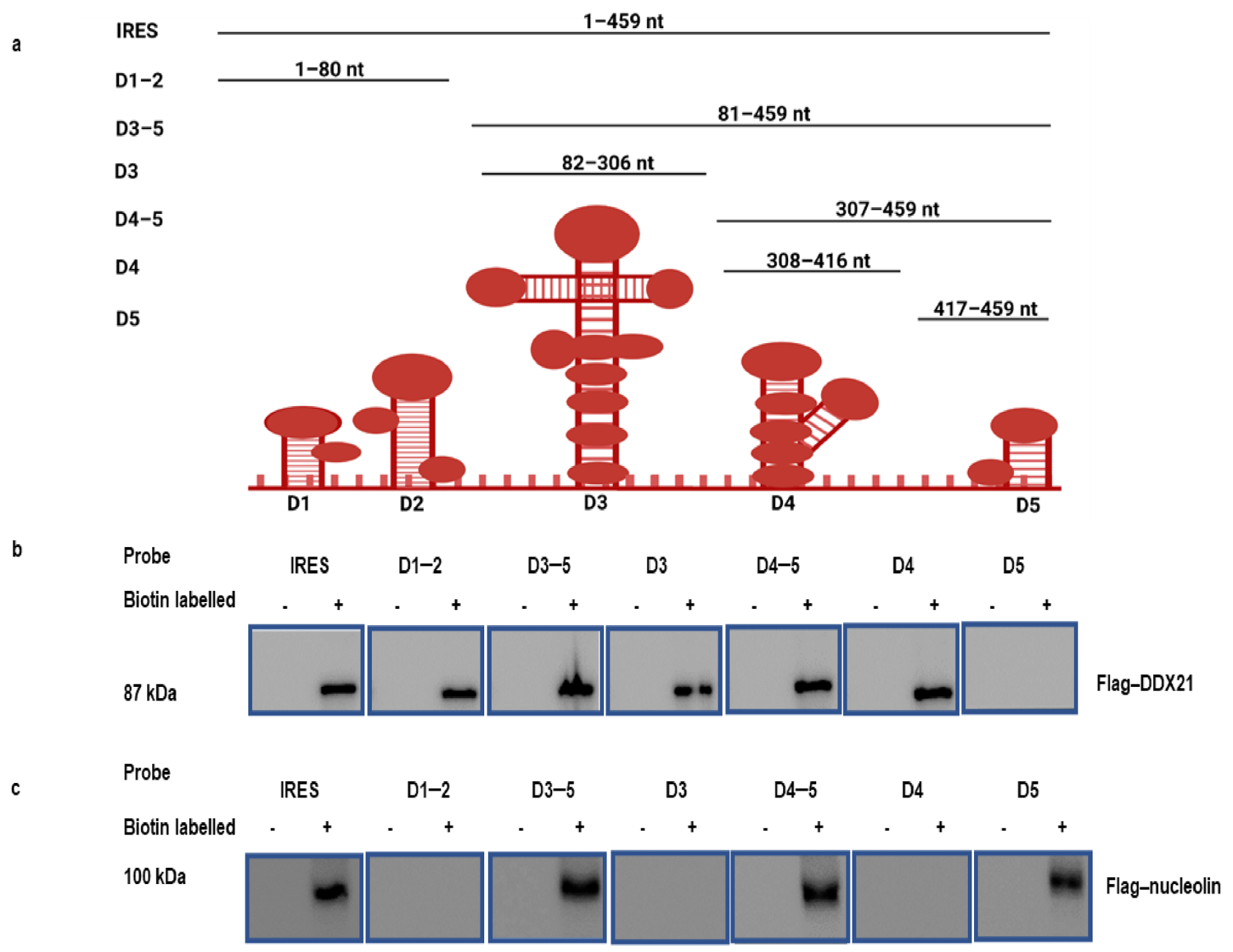
3.2. DDX21 Pulldown with the FMDV IRES Regions
3.3. DDX21 Negatively Regulates FMDV Replication
3.4. DDX21 Negatively Regulates FMDV, CSFV, and SVV IRES-Dependent Translation
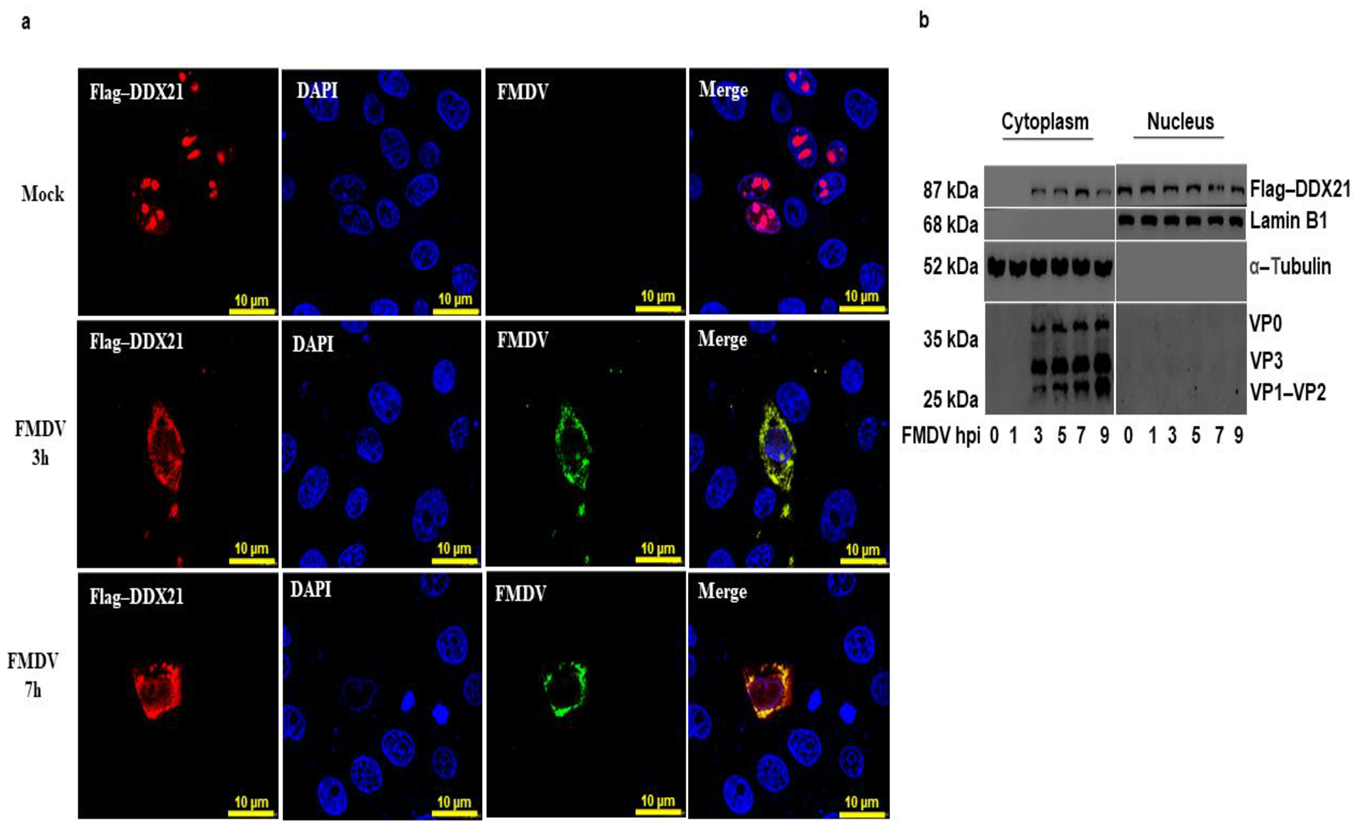
3.5. DDX21 Translocates to the Cytoplasm during FMDV Infection
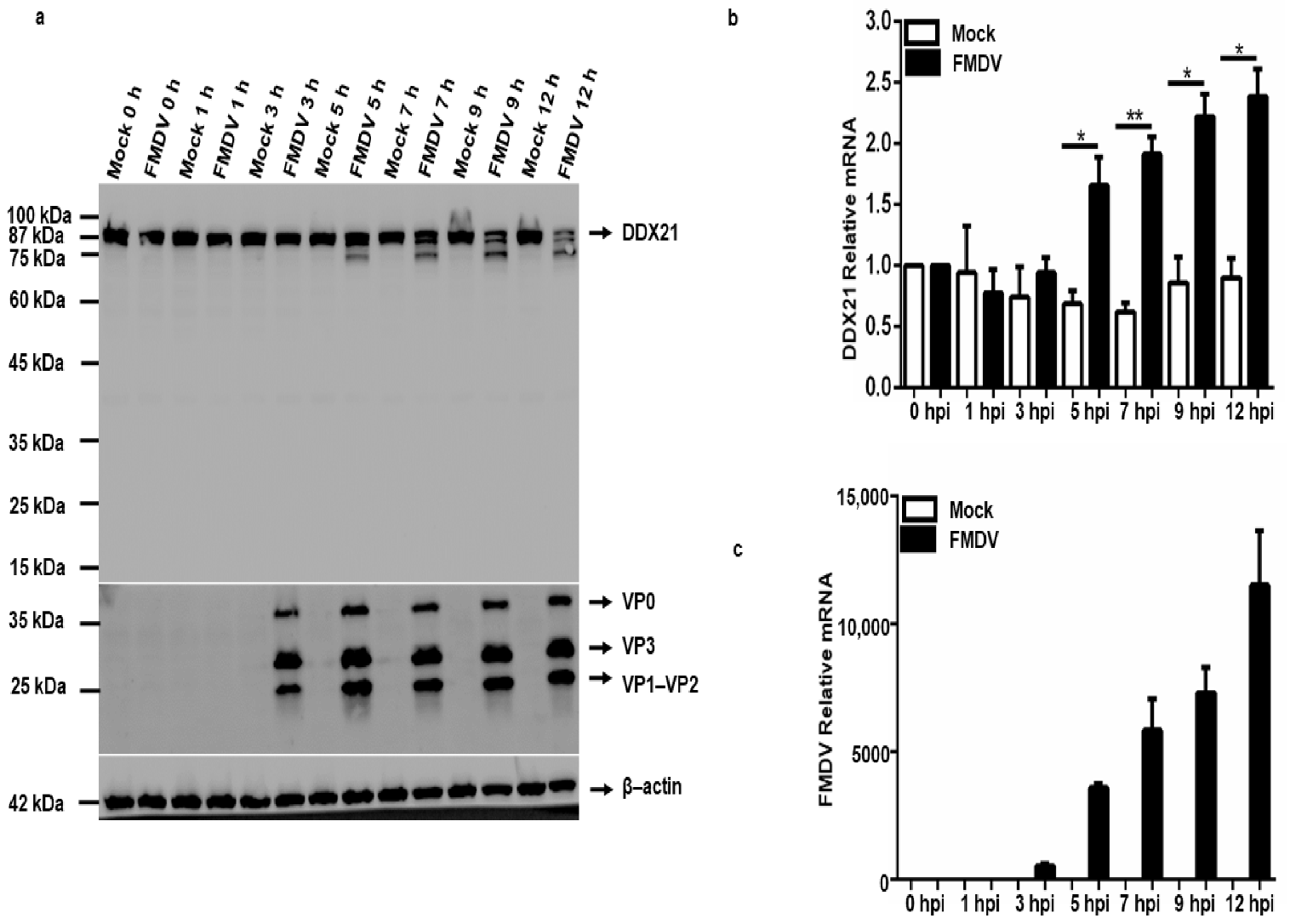
3.6. FMDV Infection Causes Degradation of the DDX21 Protein
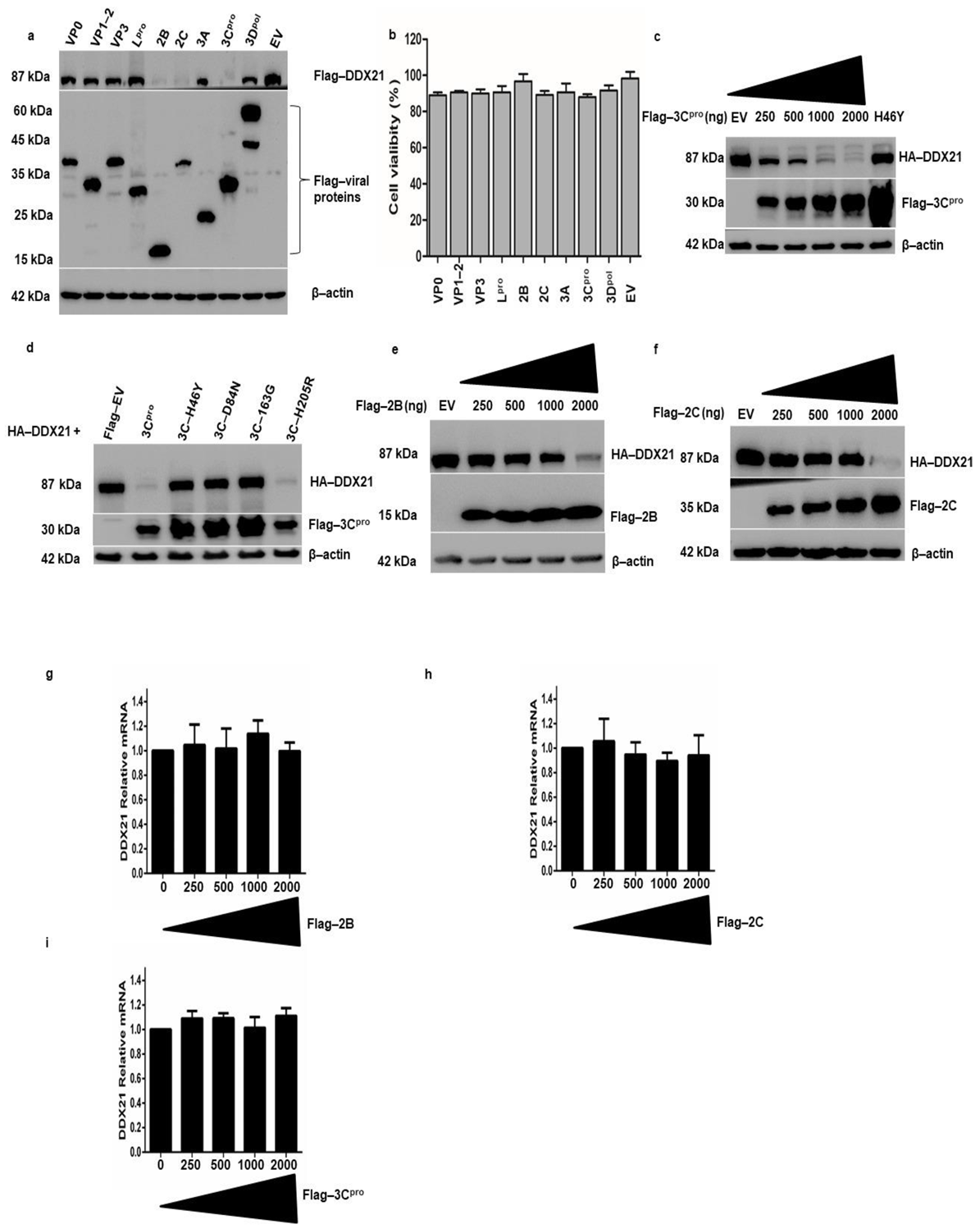
3.7. FMDV 2B, 2C, and 3Cpro Decrease DDX21 Protein Levels, and 3Cpro Catalytic Triad Active Site Residues Are Required for DDX21 Degradation
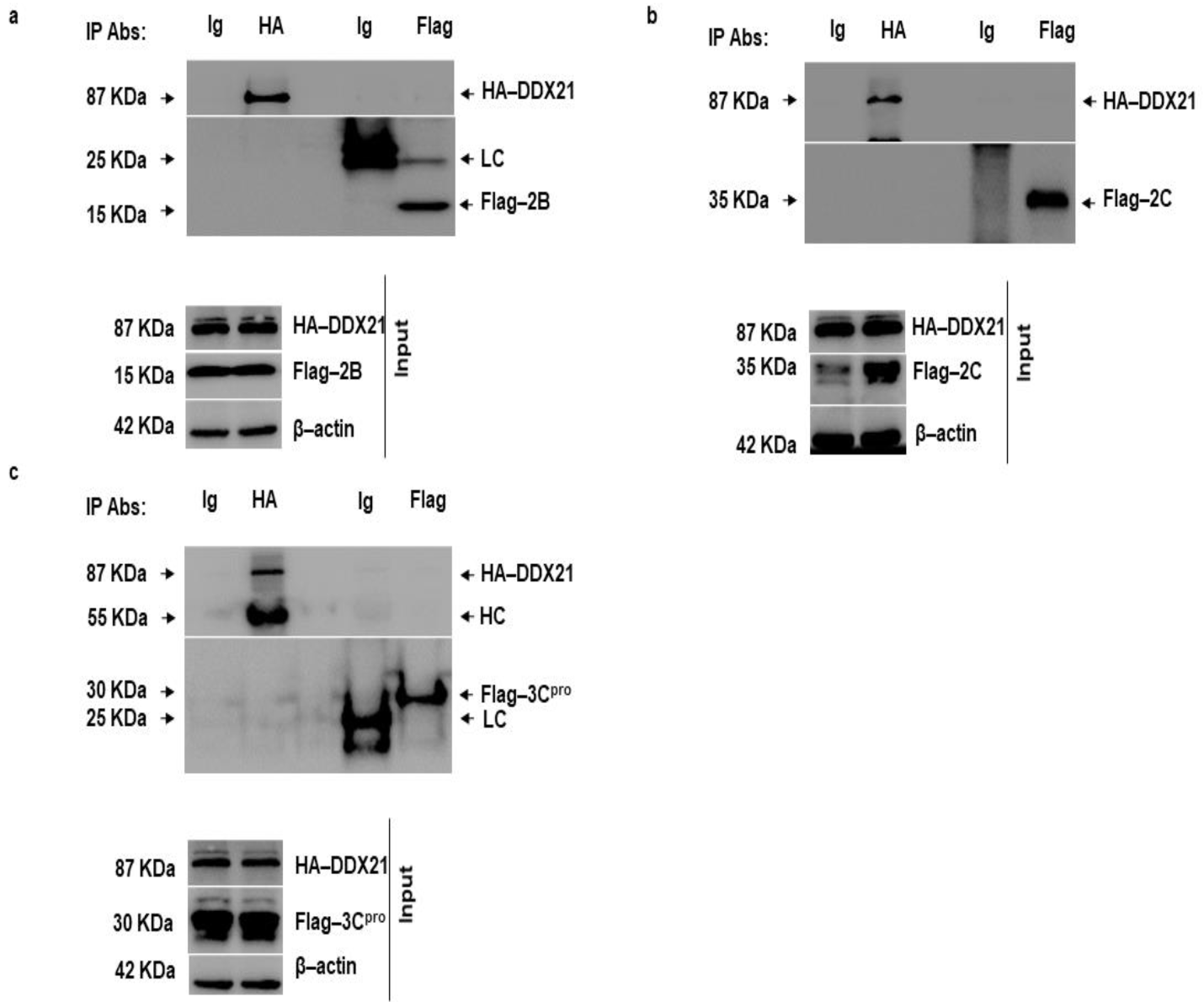
3.8. DDX21 Does Not Interact with FMDV 2B, 2C, and 3Cpro
3.9. Lysosomal and Caspase Pathway-Dependent Degradation of DDX21
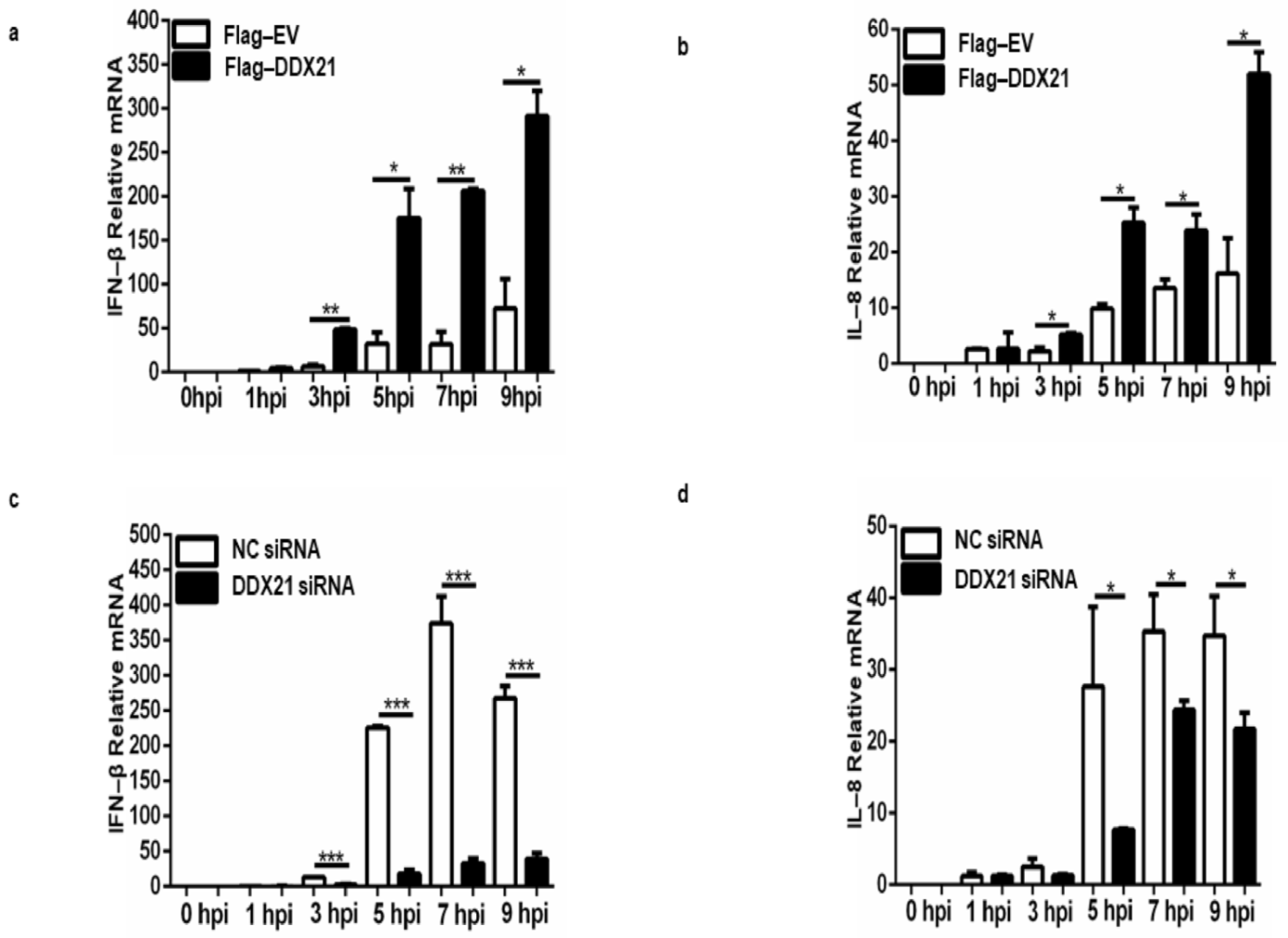
3.10. DDX21 Positively Regulates IFN-β and IL-8 Production
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Pelletier, J.; Sonenberg, N. The Organizing Principles of Eukaryotic Ribosome Recruitment. Annu. Rev. Biochem. 2019, 88, 307–335. [Google Scholar] [CrossRef] [PubMed]
- Jackson, R.J.; Hellen, C.U.; Pestova, T.V. The mechanism of eukaryotic translation initiation and principles of its regulation. Nat. Rev. Mol. Cell Biol. 2010, 11, 113–127. [Google Scholar] [CrossRef] [Green Version]
- Ramanathan, A.; Robb, G.B.; Chan, S.H. mRNA capping: Biological functions and applications. Nucleic Acids Res. 2016, 44, 7511–7526. [Google Scholar] [CrossRef] [PubMed]
- Jackson, R.J. The current status of vertebrate cellular mRNA IRESs. Cold Spring Harb. Perspect. Biol. 2013, 5, a011569. [Google Scholar] [CrossRef]
- Lee, K.M.; Chen, C.J.; Shih, S.R. Regulation Mechanisms of Viral IRES-Driven Translation. Trends Microbiol. 2017, 25, 546–561. [Google Scholar] [CrossRef]
- Kozak, M. A second look at cellular mRNA sequences said to function as internal ribosome entry sites. Nucleic Acids Res. 2005, 33, 6593–6602. [Google Scholar] [CrossRef] [PubMed]
- Leppek, K.; Das, R.; Barna, M. Functional 5′ UTR mRNA structures in eukaryotic translation regulation and how to find them. Nat. Rev. Mol. Cell Biol. 2018, 19, 158–174. [Google Scholar] [CrossRef]
- Yang, Y.; Wang, Z. IRES-mediated cap-independent translation, a path leading to hidden proteome. J. Mol. Cell Biol. 2019, 11, 911–919. [Google Scholar] [CrossRef] [Green Version]
- Godet, A.C.; David, F.; Hantelys, F.; Tatin, F.; Lacazette, E.; Garmy-Susini, B.; Prats, A.C. IRES Trans-Acting Factors, Key Actors of the Stress Response. Int. J. Mol. Sci. 2019, 20, 924. [Google Scholar] [CrossRef] [Green Version]
- Lozano, G.; Martinez-Salas, E. Structural insights into viral IRES-dependent translation mechanisms. Curr. Opin. Virol. 2015, 12, 113–120. [Google Scholar] [CrossRef] [Green Version]
- Mailliot, J.; Martin, F. Viral internal ribosomal entry sites: Four classes for one goal. Wiley Interdiscip. Rev. RNA 2018, 9, 924. [Google Scholar] [CrossRef] [PubMed]
- Anderson, E.C.; Hunt, S.L.; Jackson, R.J. Internal initiation of translation from the human rhinovirus-2 internal ribosome entry site requires the binding of Unr to two distinct sites on the 5′ untranslated region. J. Gen. Virol. 2007, 88, 3043–3052. [Google Scholar] [CrossRef] [PubMed]
- Pelletier, J.; Sonenberg, N. Internal initiation of translation of eukaryotic mRNA directed by a sequence derived from poliovirus RNA. Nature 1988, 334, 320–325. [Google Scholar] [CrossRef]
- Chase, A.J.; Daijogo, S.; Semler, B.L. Inhibition of poliovirus-induced cleavage of cellular protein PCBP2 reduces the levels of viral RNA replication. J. Virol. 2014, 88, 3192–3201. [Google Scholar] [CrossRef] [Green Version]
- Hung, C.T.; Kung, Y.A.; Li, M.L.; Brewer, G.; Lee, K.M.; Liu, S.T.; Shih, S.R. Additive Promotion of Viral Internal Ribosome Entry Site-Mediated Translation by Far Upstream Element-Binding Protein 1 and an Enterovirus 71-Induced Cleavage Product. PLoS Pathog. 2016, 12, e1005959. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, J.Y.; Li, M.L.; Huang, P.N.; Chien, K.Y.; Horng, J.T.; Shih, S.R. Heterogeneous nuclear ribonuclear protein K interacts with the enterovirus 71 5′ untranslated region and participates in virus replication. J. Gen. Virol. 2008, 89, 2540–2549. [Google Scholar] [CrossRef]
- Rai, D.K.; Lawrence, P.; Kloc, A.; Schafer, E.; Rieder, E. Analysis of the interaction between host factor Sam68 and viral elements during foot-and-mouth disease virus infections. Virol. J. 2015, 12, 224. [Google Scholar] [CrossRef] [Green Version]
- Fernandez-Chamorro, J.; Francisco-Velilla, R.; Ramajo, J.; Martinez-Salas, E. Rab1b and ARF5 are novel RNA-binding proteins involved in FMDV IRES-driven RNA localization. Life Sci. Alliance 2019, 2. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Khawaja, A.; Vopalensky, V.; Pospisek, M. Understanding the potential of hepatitis C virus internal ribosome entry site domains to modulate translation initiation via their structure and function. Wiley Interdiscip. Rev. RNA 2015, 6, 211–224. [Google Scholar] [CrossRef] [PubMed]
- Rijnbrand, R.; van der Straaten, T.; van Rijn, P.A.; Spaan, W.J.; Bredenbeek, P.J. Internal entry of ribosomes is directed by the 5′ noncoding region of classical swine fever virus and is dependent on the presence of an RNA pseudoknot upstream of the initiation codon. J. Virol. 1997, 71, 451–457. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hellen, C.U.; de Breyne, S. A distinct group of hepacivirus/pestivirus-like internal ribosomal entry sites in members of diverse picornavirus genera: Evidence for modular exchange of functional noncoding RNA elements by recombination. J. Virol. 2007, 81, 5850–5863. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wilson, J.E.; Powell, M.J.; Hoover, S.E.; Sarnow, P. Naturally occurring dicistronic cricket paralysis virus RNA is regulated by two internal ribosome entry sites. Mol. Cell. Biol. 2000, 20, 4990–4999. [Google Scholar] [CrossRef] [Green Version]
- Hertz, M.I.; Thompson, S.R. In vivo functional analysis of the Dicistroviridae intergenic region internal ribosome entry sites. Nucleic Acids Res. 2011, 39, 7276–7288. [Google Scholar] [CrossRef]
- Cevallos, R.C.; Sarnow, P. Factor-independent assembly of elongation-competent ribosomes by an internal ribosome entry site located in an RNA virus that infects penaeid shrimp. J. Virol. 2005, 79, 677–683. [Google Scholar] [CrossRef] [Green Version]
- Burks, J.M.; Zwieb, C.; Muller, F.; Wower, I.K.; Wower, J. In silico analysis of IRES RNAs of foot-and-mouth disease virus and related picornaviruses. Arch. Virol. 2011, 156, 1737–1747. [Google Scholar] [CrossRef]
- Liu, W.; Yang, D.; Sun, C.; Wang, H.; Zhao, B.; Zhou, G.; Yu, L. hnRNP K Is a Novel Internal Ribosomal Entry Site-Transacting Factor That Negatively Regulates Foot-and-Mouth Disease Virus Translation and Replication and Is Antagonized by Viral 3C Protease. J. Virol. 2020, 94, e00803-20. [Google Scholar] [CrossRef]
- Fajardo, T., Jr.; Rosas, M.F.; Sobrino, F.; Martinez-Salas, E. Exploring IRES region accessibility by interference of foot-and-mouth disease virus infectivity. PLoS ONE 2012, 7, e41382. [Google Scholar] [CrossRef] [Green Version]
- Fernandez, N.; Fernandez-Miragall, O.; Ramajo, J.; Garcia-Sacristan, A.; Bellora, N.; Eyras, E.; Briones, C.; Martinez-Salas, E. Structural basis for the biological relevance of the invariant apical stem in IRES-mediated translation. Nucleic Acids Res. 2011, 39, 8572–8585. [Google Scholar] [CrossRef]
- de Breyne, S.; Yu, Y.; Unbehaun, A.; Pestova, T.V.; Hellen, C.U. Direct functional interaction of initiation factor eIF4G with type 1 internal ribosomal entry sites. Proc. Natl. Acad. Sci. USA 2009, 106, 9197–9202. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Niepmann, M.; Petersen, A.; Meyer, K.; Beck, E. Functional involvement of polypyrimidine tract-binding protein in translation initiation complexes with the internal ribosome entry site of foot-and-mouth disease virus. J. Virol. 1997, 71, 8330–8339. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pilipenko, E.V.; Pestova, T.V.; Kolupaeva, V.G.; Khitrina, E.V.; Poperechnaya, A.N.; Agol, V.I.; Hellen, C.U. A cell cycle-dependent protein serves as a template-specific translation initiation factor. Genes Dev. 2000, 14, 2028–2045. [Google Scholar]
- Pacheco, A.; Lopez de Quinto, S.; Ramajo, J.; Fernandez, N.; Martinez-Salas, E. A novel role for Gemin5 in mRNA translation. Nucleic Acids Res. 2009, 37, 582–590. [Google Scholar] [CrossRef]
- Galan, A.; Lozano, G.; Pineiro, D.; Martinez-Salas, E. G3BP1 interacts directly with the FMDV IRES and negatively regulates translation. FEBS J. 2017, 284, 3202–3217. [Google Scholar] [CrossRef] [Green Version]
- Sun, C.; Liu, M.; Chang, J.; Yang, D.; Zhao, B.; Wang, H.; Zhou, G.; Weng, C.; Yu, L. Heterogeneous Nuclear Ribonucleoprotein L Negatively Regulates Foot-and-Mouth Disease Virus Replication through Inhibition of Viral RNA Synthesis by Interacting with the Internal Ribosome Entry Site in the 5′ Untranslated Region. J. Virol. 2020, 94, e00282-20. [Google Scholar] [CrossRef]
- Abdullah, S.W.; Han, S.; Wu, J.; Zhang, Y.; Bai, M.; Jin, Y.; Zhi, X.; Guan, J.; Sun, S.; Guo, H. The DDX23 Negatively Regulates Translation and Replication of Foot-and-Mouth Disease Virus and Is Degraded by 3C Proteinase. Viruses 2020, 12, 1348. [Google Scholar] [CrossRef]
- Zhou, Y.; Wu, W.; Xie, L.; Wang, D.; Ke, Q.; Hou, Z.; Wu, X.; Fang, Y.; Chen, H.; Xiao, S.; et al. Cellular RNA Helicase DDX1 Is Involved in Transmissible Gastroenteritis Virus nsp14-Induced Interferon-Beta Production. Front. Immunol. 2017, 8, 940. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jarmoskaite, I.; Russell, R. RNA helicase proteins as chaperones and remodelers. Annu. Rev. Biochem. 2014, 83, 697–725. [Google Scholar] [CrossRef] [Green Version]
- Meier-Stephenson, V.; Mrozowich, T.; Pham, M.; Patel, T.R. DEAD-box helicases: The Yin and Yang roles in viral infections. Biotechnol. Genet. Eng. Rev. 2018, 34, 3–32. [Google Scholar] [CrossRef] [PubMed]
- Taschuk, F.; Cherry, S. DEAD-Box Helicases: Sensors, Regulators, and Effectors for Antiviral Defense. Viruses 2020, 12, 181. [Google Scholar] [CrossRef] [Green Version]
- Dehghani, M.; Lasko, P. Multiple Functions of the DEAD-Box Helicase Vasa in Drosophila Oogenesis. Results Probl. Cell Differ. 2017, 63, 127–147. [Google Scholar] [CrossRef] [PubMed]
- Ali, M.A.M. The DEAD-box protein family of RNA helicases: Sentinels for a myriad of cellular functions with emerging roles in tumorigenesis. Int. J. Clin. Oncol. 2021, 26, 795–825. [Google Scholar] [CrossRef] [PubMed]
- Xing, Y.H.; Yao, R.W.; Zhang, Y.; Guo, C.J.; Jiang, S.; Xu, G.; Dong, R.; Yang, L.; Chen, L.L. SLERT Regulates DDX21 Rings Associated with Pol I Transcription. Cell 2017, 169, 664–678.e616. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Liu, C.H.; Zhou, L.; Krug, R.M. Cellular DDX21 RNA helicase inhibits influenza A virus replication but is counteracted by the viral NS1 protein. Cell Host Microbe 2014, 15, 484–493. [Google Scholar] [CrossRef] [Green Version]
- Chen, Z.; Li, Z.; Hu, X.; Xie, F.; Kuang, S.; Zhan, B.; Gao, W.; Chen, X.; Gao, S.; Li, Y.; et al. Structural Basis of Human Helicase DDX21 in RNA Binding, Unwinding, and Antiviral Signal Activation. Adv. Sci. 2020, 7, 2000532. [Google Scholar] [CrossRef]
- Dong, Y.; Ye, W.; Yang, J.; Han, P.; Wang, Y.; Ye, C.; Weng, D.; Zhang, F.; Xu, Z.; Lei, Y. DDX21 translocates from nucleus to cytoplasm and stimulates the innate immune response due to dengue virus infection. Biochem. Biophys. Res. Commun. 2016, 473, 648–653. [Google Scholar] [CrossRef] [PubMed]
- Hammond, J.A.; Zhou, L.; Lamichhane, R.; Chu, H.Y.; Millar, D.P.; Gerace, L.; Williamson, J.R. A Survey of DDX21 Activity During Rev/RRE Complex Formation. J. Mol. Biol. 2018, 430, 537–553. [Google Scholar] [CrossRef] [PubMed]
- Hao, H.; Han, T.; Xuan, B.; Sun, Y.; Tang, S.; Yue, N.; Qian, Z. Dissecting the Role of DDX21 in Regulating Human Cytomegalovirus Replication. J. Virol. 2019, 93, e01222-19. [Google Scholar] [CrossRef]
- Watanabe, Y.; Ohtaki, N.; Hayashi, Y.; Ikuta, K.; Tomonaga, K. Autogenous translational regulation of the Borna disease virus negative control factor X from polycistronic mRNA using host RNA helicases. PLoS Pathog. 2009, 5, e1000654. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Z.; Kim, T.; Bao, M.; Facchinetti, V.; Jung, S.Y.; Ghaffari, A.A.; Qin, J.; Cheng, G.; Liu, Y.J. DDX1, DDX21, and DHX36 helicases form a complex with the adaptor molecule TRIF to sense dsRNA in dendritic cells. Immunity 2011, 34, 866–878. [Google Scholar] [CrossRef] [Green Version]
- Tsai, S.Y.; Segovia, J.A.; Chang, T.H.; Morris, I.R.; Berton, M.T.; Tessier, P.A.; Tardif, M.R.; Cesaro, A.; Bose, S. DAMP molecule S100A9 acts as a molecular pattern to enhance inflammation during influenza A virus infection: Role of DDX21-TRIF-TLR4-MyD88 pathway. PLoS Pathog. 2014, 10, e1003848. [Google Scholar] [CrossRef]
- Fu, S.Z.; Yang, W.P.; Ru, Y.; Zhang, K.S.; Wang, Y.; Liu, X.T.; Li, D.; Zheng, H.X. DDX56 cooperates with FMDV 3A to enhance FMDV replication by inhibiting the phosphorylation of IRF3. Cell. Signal. 2019, 64, 109393. [Google Scholar] [CrossRef] [PubMed]
- Han, S.; Sun, S.; Li, P.; Liu, Q.; Zhang, Z.; Dong, H.; Sun, M.; Wu, W.; Wang, X.; Guo, H. Ribosomal Protein L13 Promotes IRES-Driven Translation of Foot-and-Mouth Disease Virus in a Helicase DDX3-Dependent Manner. J. Virol. 2020, 94, e01679-19. [Google Scholar] [CrossRef]
- Klein, J. Understanding the molecular epidemiology of foot-and-mouth-disease virus. Infect. Genet. Evol. 2009, 9, 153–161. [Google Scholar] [CrossRef]
- Ao, D.; Guo, H.C.; Sun, S.Q.; Sun, D.H.; Fung, T.S.; Wei, Y.Q.; Han, S.C.; Yao, X.P.; Cao, S.Z.; Liu, D.X.; et al. Viroporin Activity of the Foot-and-Mouth Disease Virus Non-Structural 2B Protein. PLoS ONE 2015, 10, e0125828. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, H.; Xue, Q.; Cao, W.; Yang, F.; Ma, L.; Liu, W.; Zhang, K.; Liu, X.; Zhu, Z.; Zheng, H. Foot-and-mouth disease virus nonstructural protein 2B interacts with cyclophilin A, modulating virus replication. FASEB J. 2018, 32, 6706–6723. [Google Scholar] [CrossRef]
- Gladue, D.P.; O’Donnell, V.; Baker-Branstetter, R.; Holinka, L.G.; Pacheco, J.M.; Fernandez-Sainz, I.; Lu, Z.; Brocchi, E.; Baxt, B.; Piccone, M.E.; et al. Foot-and-mouth disease virus nonstructural protein 2C interacts with Beclin1, modulating virus replication. J. Virol. 2012, 86, 12080–12090. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, H.; Zhu, Z.; Xue, Q.; Yang, F.; Cao, W.; Zhang, K.; Liu, X.; Zheng, H. Foot-and-Mouth Disease Virus Antagonizes NOD2-Mediated Antiviral Effects by Inhibiting NOD2 Protein Expression. J. Virol. 2019, 93, e00124-19. [Google Scholar] [CrossRef] [Green Version]
- Du, Y.; Bi, J.; Liu, J.; Liu, X.; Wu, X.; Jiang, P.; Yoo, D.; Zhang, Y.; Wu, J.; Wan, R.; et al. 3Cpro of foot-and-mouth disease virus antagonizes the interferon signaling pathway by blocking STAT1/STAT2 nuclear translocation. J. Virol. 2014, 88, 4908–4920. [Google Scholar] [CrossRef] [Green Version]
- Dong, H.; Liu, P.; Bai, M.; Wang, K.; Feng, R.; Zhu, D.; Sun, Y.; Mu, S.; Li, H.; Harmsen, M.; et al. Structural and molecular basis for foot-and-mouth disease virus neutralization by two potent protective antibodies. Protein Cell 2021, in press. [Google Scholar] [CrossRef]
- Han, S.; Wang, X.; Guan, J.; Wu, J.; Zhang, Y.; Li, P.; Liu, Z.; Abdullah, S.W.; Zhang, Z.; Jin, Y.; et al. Nucleolin promotes IRES-driven translation of foot-and-mouth disease virus by supporting the assembly of translation initiation complexes. J. Virol. 2021, in press. [Google Scholar] [CrossRef]
- Zhi, X.; Zhang, Y.; Sun, S.; Zhang, Z.; Dong, H.; Luo, X.; Wei, Y.; Lu, Z.; Dou, Y.; Wu, R.; et al. NLRP3 inflammasome activation by Foot-and-mouth disease virus infection mainly induced by viral RNA and non-structural protein 2B. RNA Biol. 2020, 17, 335–349. [Google Scholar] [CrossRef]
- Carrillo, C.; Tulman, E.R.; Delhon, G.; Lu, Z.; Carreno, A.; Vagnozzi, A.; Kutish, G.F.; Rock, D.L. Comparative genomics of foot-and-mouth disease virus. J. Virol. 2005, 79, 6487–6504. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bai, X.; Li, P.; Sun, P.; Li, Y.; Bao, H.; Lu, Z.; Cao, Y.; Guo, J.; Liu, Z. Virulence comparison of genetic engineering virus containing different length poly(C) tract of foot-and-mouth disease virus. Wei Sheng Wu Xue Bao 2008, 48, 1654–1658. [Google Scholar] [PubMed]
- Zhu, Z.; Yang, F.; Cao, W.; Liu, H.; Zhang, K.; Tian, H.; Dang, W.; He, J.; Guo, J.; Liu, X.; et al. The Pseudoknot Region of the 5′ Untranslated Region Is a Determinant of Viral Tropism and Virulence of Foot-and-Mouth Disease Virus. J. Virol. 2019, 93, e02039-18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mason, P.W.; Bezborodova, S.V.; Henry, T.M. Identification and characterization of a cis-acting replication element (cre) adjacent to the internal ribosome entry site of foot-and-mouth disease virus. J. Virol. 2002, 76, 9686–9694. [Google Scholar] [CrossRef] [Green Version]
- Diaz-Toledano, R.; Lozano, G.; Martinez-Salas, E. In-cell SHAPE uncovers dynamic interactions between the untranslated regions of the foot-and-mouth disease virus RNA. Nucleic Acids Res. 2017, 45, 1416–1432. [Google Scholar] [CrossRef]
- Kolupaeva, V.G.; Hellen, C.U.; Shatsky, I.N. Structural analysis of the interaction of the pyrimidine tract-binding protein with the internal ribosomal entry site of encephalomyocarditis virus and foot-and-mouth disease virus RNAs. RNA 1996, 2, 1199–1212. [Google Scholar]
- Andreev, D.E.; Fernandez-Miragall, O.; Ramajo, J.; Dmitriev, S.E.; Terenin, I.M.; Martinez-Salas, E.; Shatsky, I.N. Differential factor requirement to assemble translation initiation complexes at the alternative start codons of foot-and-mouth disease virus RNA. RNA 2007, 13, 1366–1374. [Google Scholar] [CrossRef] [Green Version]
- Zhu, Z.; Wang, G.; Yang, F.; Cao, W.; Mao, R.; Du, X.; Zhang, X.; Li, C.; Li, D.; Zhang, K.; et al. Foot-and-Mouth Disease Virus Viroporin 2B Antagonizes RIG-I-Mediated Antiviral Effects by Inhibition of Its Protein Expression. J. Virol. 2016, 90, 11106–11121. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- DiMaio, D. Viruses, masters at downsizing. Cell Host Microbe 2012, 11, 560–561. [Google Scholar] [CrossRef] [Green Version]
- Kim, D.S.; Camacho, C.V.; Nagari, A.; Malladi, V.S.; Challa, S.; Kraus, W.L. Activation of PARP-1 by snoRNAs Controls Ribosome Biogenesis and Cell Growth via the RNA Helicase DDX21. Mol. Cell 2019, 75, 1270–1285. [Google Scholar] [CrossRef] [PubMed]
- McRae, E.K.S.; Dupas, S.J.; Booy, E.P.; Piragasam, R.S.; Fahlman, R.P.; McKenna, S.A. An RNA guanine quadruplex regulated pathway to TRAIL-sensitization by DDX21. RNA 2020, 26, 44–57. [Google Scholar] [CrossRef]
- Marcaida, M.J.; Kauzlaric, A.; Duperrex, A.; Sulzle, J.; Moncrieffe, M.C.; Adebajo, D.; Manley, S.; Trono, D.; Dal Peraro, M. The Human RNA Helicase DDX21 Presents a Dimerization Interface Necessary for Helicase Activity. iScience 2020, 23, 101811. [Google Scholar] [CrossRef] [PubMed]
- McRae, E.K.S.; Booy, E.P.; Moya-Torres, A.; Ezzati, P.; Stetefeld, J.; McKenna, S.A. Human DDX21 binds and unwinds RNA guanine quadruplexes. Nucleic Acids Res. 2017, 45, 6656–6668. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Santoriello, C.; Sporrij, A.; Yang, S.; Flynn, R.A.; Henriques, T.; Dorjsuren, B.; Custo Greig, E.; McCall, W.; Stanhope, M.E.; Fazio, M.; et al. RNA helicase DDX21 mediates nucleotide stress responses in neural crest and melanoma cells. Nat. Cell Biol. 2020, 22, 372–379. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Moreno, M.; Noerenberg, M.; Ni, S.; Järvelin, A.I.; González-Almela, E.; Lenz, C.E.; Bach-Pages, M.; Cox, V.; Avolio, R.; Davis, T. System-wide profiling of RNA-binding proteins uncovers key regulators of virus infection. Mol. Cell 2019, 74, 196–211.e111. [Google Scholar] [CrossRef] [Green Version]
- Lin, J.Y.; Li, M.L.; Shih, S.R. Far upstream element binding protein 2 interacts with enterovirus 71 internal ribosomal entry site and negatively regulates viral translation. Nucleic Acids Res. 2009, 37, 47–59. [Google Scholar] [CrossRef]
- Sun, X.; Belser, J.A.; Maines, T.R. Adaptation of H9N2 Influenza Viruses to Mammalian Hosts: A Review of Molecular Markers. Viruses 2020, 12, 541. [Google Scholar] [CrossRef]
- Chan, Y.K.; Gack, M.U. Viral evasion of intracellular DNA and RNA sensing. Nat. Rev. Microbiol. 2016, 14, 360–373. [Google Scholar] [CrossRef]
- Clementz, M.A.; Chen, Z.; Banach, B.S.; Wang, Y.; Sun, L.; Ratia, K.; Baez-Santos, Y.M.; Wang, J.; Takayama, J.; Ghosh, A.K.; et al. Deubiquitinating and interferon antagonism activities of coronavirus papain-like proteases. J. Virol. 2010, 84, 4619–4629. [Google Scholar] [CrossRef] [Green Version]
- Gao, Y.; Sun, S.Q.; Guo, H.C. Biological function of Foot-and-mouth disease virus non-structural proteins and non-coding elements. Virol. J. 2016, 13, 107. [Google Scholar] [CrossRef] [Green Version]
- Visser, L.J.; Medina, G.N.; Rabouw, H.H.; de Groot, R.J.; Langereis, M.A.; de Los Santos, T.; van Kuppeveld, F.J.M. Foot-and-Mouth Disease Virus Leader Protease Cleaves G3BP1 and G3BP2 and Inhibits Stress Granule Formation. J. Virol. 2019, 93, e00922-18. [Google Scholar] [CrossRef] [Green Version]
- Xue, Q.; Liu, H.; Zeng, Q.; Zheng, H.; Xue, Q.; Cai, X. The DEAD-Box RNA Helicase DDX1 Interacts with the Viral Protein 3D and Inhibits Foot-and-Mouth Disease Virus Replication. Virol. Sin. 2019, 34, 610–617. [Google Scholar] [CrossRef] [Green Version]
- Wang, D.; Fang, L.; Li, K.; Zhong, H.; Fan, J.; Ouyang, C.; Zhang, H.; Duan, E.; Luo, R.; Zhang, Z.; et al. Foot-and-mouth disease virus 3C protease cleaves NEMO to impair innate immune signaling. J. Virol. 2012, 86, 9311–9322. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Belsham, G.J.; McInerney, G.M.; Ross-Smith, N. Foot-and-mouth disease virus 3C protease induces cleavage of translation initiation factors eIF4A and eIF4G within infected cells. J. Virol. 2000, 74, 272–280. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, J.; Wang, Y.; Liu, J.; Ding, L.; Zhang, Q.; Li, X.; Cao, H.; Tang, J.; Zheng, S.J. A critical role of N-myc and STAT interactor (Nmi) in foot-and-mouth disease virus (FMDV) 2C-induced apoptosis. Virus Res. 2012, 170, 59–65. [Google Scholar] [CrossRef]
- Moffat, K.; Howell, G.; Knox, C.; Belsham, G.J.; Monaghan, P.; Ryan, M.D.; Wileman, T. Effects of foot-and-mouth disease virus nonstructural proteins on the structure and function of the early secretory pathway: 2BC but not 3A blocks endoplasmic reticulum-to-Golgi transport. J. Virol. 2005, 79, 4382–4395. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, H.; Kim, A.Y.; Choi, J.; Park, S.Y.; Park, S.H.; Kim, J.S.; Lee, S.I.; Park, J.H.; Park, C.K.; Ko, Y.J. Foot-and-Mouth Disease Virus Evades Innate Immune Response by 3C-Targeting of MDA5. Cells 2021, 10, 271. [Google Scholar] [CrossRef] [PubMed]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abdullah, S.W.; Wu, J.; Zhang, Y.; Bai, M.; Guan, J.; Liu, X.; Sun, S.; Guo, H. DDX21, a Host Restriction Factor of FMDV IRES-Dependent Translation and Replication. Viruses 2021, 13, 1765. https://doi.org/10.3390/v13091765
Abdullah SW, Wu J, Zhang Y, Bai M, Guan J, Liu X, Sun S, Guo H. DDX21, a Host Restriction Factor of FMDV IRES-Dependent Translation and Replication. Viruses. 2021; 13(9):1765. https://doi.org/10.3390/v13091765
Chicago/Turabian StyleAbdullah, Sahibzada Waheed, Jin’en Wu, Yun Zhang, Manyuan Bai, Junyong Guan, Xiangtao Liu, Shiqi Sun, and Huichen Guo. 2021. "DDX21, a Host Restriction Factor of FMDV IRES-Dependent Translation and Replication" Viruses 13, no. 9: 1765. https://doi.org/10.3390/v13091765
APA StyleAbdullah, S. W., Wu, J., Zhang, Y., Bai, M., Guan, J., Liu, X., Sun, S., & Guo, H. (2021). DDX21, a Host Restriction Factor of FMDV IRES-Dependent Translation and Replication. Viruses, 13(9), 1765. https://doi.org/10.3390/v13091765







