Abstract
Disease outbreaks caused by introduced Phytophthora species have been increasing in British forests and woodlands in recent years. A better knowledge of the Phytophthora communities already present in the UK is of great importance when developing management and mitigation strategies for these diseases. To do this, soils were sampled in “disturbed” sites, meaning sites frequently visited by the public, with recent and new plantings or soil disturbances versus more “natural” forest and woodland sites with little disturbance or management. Phytophthora diversity was assessed using high-throughput Illumina sequencing targeting the widely accepted barcoding Internal Transcribed Spacer 1 (ITS1) region of rRNA and comparing it with the mitochondrial cytochrome c oxidase I (COI) gene. Isolation of Phytophthora was run in parallel. Nothophytophthora spp. and Phytophthora spp. were detected in 79 and 41 of the 132 locations of the 14 studied sites when using ITS or COI, respectively. A total of 20 Phytophthora amplicon sequence variants (ASVs) were assigned to known Phytophthora species from eight clades (1a, 2, 2b, 3a, 5, 6b, 7a, 8b, 8c, 8d, 10a, and 10b) and 12 ASVs from six clades (1a, 2c, 3a, 3b, 6b, 7a, 8b, 8c, and 8d) when using ITS or COI, respectively. Only at two locations were the results in agreement for ITS, COI, and isolation. Additionally, 21 and 17 unknown Phytophthora phylotypes were detected using the ITS and COI, respectively. Several Phytophthora spp. within clades 7 and 8, including very important forest pathogens such as P. austrocedri and P. ramorum, were identified and found more frequently at “disturbed” sites. Additionally, eight ASVs identified as Nothophytophthora spp. were detected representing the first report of species within this new genus in Britain. Only three species not known to be present in Britain (P. castaneae, P. capsici, and P. fallax) were detected with the ITS primers and not with COI. To confirm the presence of these or any potential new Phytophthora species, sites should be re-sampled for confirmation. Additionally, there is a need to confirm if these species are a threat to British trees and try to establish any eradication measures required to mitigate Phytophthora spread in Britain.
1. Introduction
In recent years, several outbreaks caused by Phytophthora (P. ramorum, P. kernoviae, P. lateralis, P. pseudosyringae and P. austrocedri) have emerged in Britain and also worldwide to cause significant mortality on a range of woody hosts [1,2,3,4,5,6,7]. As we witness the devastating impact of Phytophthora on the natural ecosystem, an increasing number of studies have looked at species composition in diverse environments such as nurseries [8,9,10,11], agricultural and urban environments [12,13,14], and semi- and natural environments [15,16,17,18,19,20]. As a result, many new species have been recovered with a lot of these not formally described yet. In some cases, these newly discovered species are endemic to limited geographic areas, and their potential impact on the environment remains to be elucidated [16,20,21,22]. Approximately 180 Phytophthora species have been described with an estimate of 326 species in total covering 12 phylogenetic clades [12].
The spread of Phytophthora has been linked to human activities such as plant movement/trade and other pathways such as soil on shoes, machinery, and water [23,24,25,26,27]. The soil environment plays an integral role in the spread and establishment of Phytophthora pathogens, which can persist long term in soil in the form of resilient thick-walled spores. Waterlogged soils may also harbor free-swimming zoospores, the main mechanism by which Phytophthora infects plants. A better knowledge of the diversity of Phytophthora and the mechanisms of spread from site to site is of great importance in developing management and mitigation strategies for these diseases.
The identification of Phytophthora species is mainly based on the sequencing of the Internal Transcribed Spacer (ITS) region [28,29,30]. The ITS region is considered as the best available genetic marker to identify Phytophthora to clade level, since almost all known Phytophthora taxa have been sequenced for the ITS region [31]. New technologies such as next-generation sequencing (NGS) using Illumina or 454 technologies base their identification on the ITS region [21,32,33,34,35]. These new techniques have demonstrated greater depth in detecting Phytophthora communities than traditional cultural methods followed by Sanger sequencing. However, those studies also highlighted the ongoing difficulty of differentiating some closely related Phytophthora species which cannot be discriminated based on their ITS sequence. In such cases, species identification has been limited to species complexes or clusters. To improve the accuracy of DNA sequence-based identification, many markers have been potentially identified [36,37,38]. Several studies have indicated that DNA barcoding with the cytochrome c oxidase subunit I (COI or cox1) is an approach that can be used as a practical option to identify Phytophthora due to its highest resolution within the genus [31,39,40]. It is also complementary to ITS analyses because it uses the mitochondrial genome instead of nuclear DNA and therefore cannot identify hybrids, which are increasingly found in the Phytophthora community [17,41,42,43].
As part of the research effort to identify Phytophthora species that are a potential risk for the wider environment, this study is looking at the presence of Phytophthora in UK sites that we catalogue as “disturbed” sites (sites frequently visited by the public, with recent and new plantings and a link to nurseries) and from semi-natural forest and woodland sites (with little disturbance or management, i.e., “undisturbed”). Since 1600, most woods in Britain have had some human interference such as cultural activity or felling; therefore, the term “semi-natural” forest is more accurately used for those settings classified as “undisturbed”. Based on the information already published, we hypothesize that Phytophthora diversity will be greater in “disturbed” sites than “undisturbed” sites. In addition, for the first time, we test the applicability of the COI genetic marker in a metabarcoding study to identify Phytophthora communities and compare its discriminative power with the ITS.
2. Materials and Methods
2.1. Soil Sampling and Phytophthora Isolation
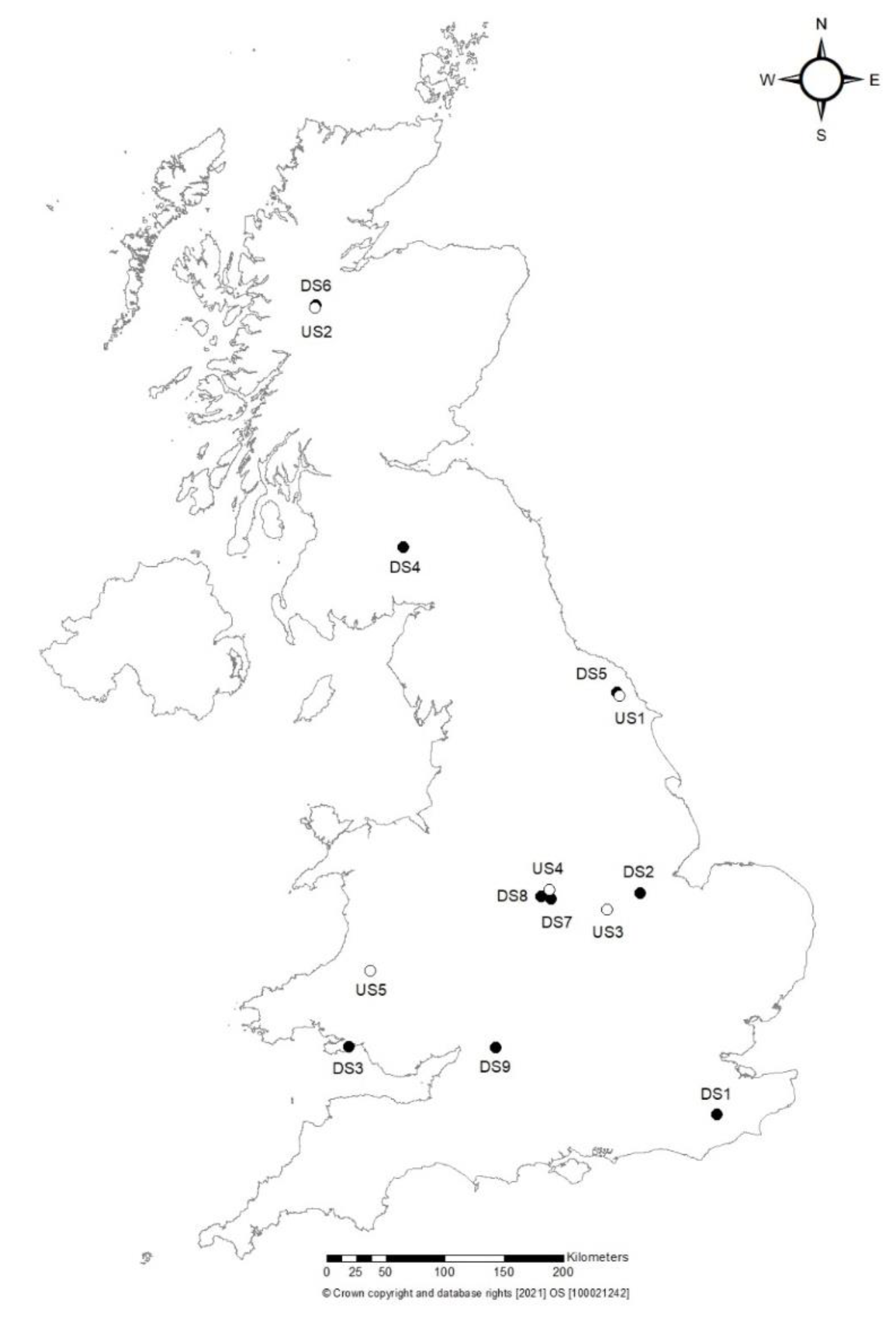
From May to November of 2016 soil samples were collected from England, Scotland and Wales from nine “disturbed” sites (called DS1 to DS9) including three gardens/arboreta sites open to the public (DS1, DS3, DS9), four woodlands or recreational parks sites frequently visited by hikers, bikers, and children parties (DS2, DS5, DS7, DS8), and two sites with recent and new plantings and a link to nurseries (DS4, DS6). We also sampled five more “natural” forest and woodland sites (with little disturbance or management, i.e., “undisturbed” called US1 to US5) (Figure 1; Table 1).
Figure 1.
Map of Britain showing the sites where samples were collected for this study.

Table 1.
Description of site and locations sampled in the study with nearest hosts and health status, and the Phytophthora/Nothophytophthora species identified by using Internal Transcribed Spacer (ITS) and cytochrome c oxidase I (COI) regions or by isolation from soil.
Soil samples (500–1000 g) were collected from around symptomatic trees or asymptomatic trees/shrubs. The sampling was done to cover as many different areas/hosts as possible in order to try and identify whether there are any patterns to Phytophthora diversity. At each site, the distribution of sampling depended on the presence/distribution of known current or previous Phytophthora outbreaks, or the location of known asymptomatic hosts. Each sample was recorded with a global positioning system (GPS) reading, and the genus/species of host in the proximity and their ill/health status was also recorded (Table 1).
The soil was collected at four points around each individual tree, taking samples within 1 m horizontal distance from the root collar of each tree. The soil samples per tree were pooled and mixed in a single grip-seal polythene bag. In the laboratory, soil samples were mixed well together removing any stones or organic material and then were divided in two parts: one to perform Phytophthora baiting and the other to perform DNA extraction.
Baiting from soil was performed by placing the soil in containers and flooding them with distilled water about 3–4 cm above the soil level. Debris and organic matter were removed from the surface of the water and young leaves of Castanea, Quercus and Rhododendron or Lupinus seedlings were placed carefully on the surface of the water as described by Jung et al. [44,45]. The leaf/seedlings-baits were removed four or five days after incubation at room temperature, and isolations were made onto synthetic mucor agar (SMA) medium (amended as per Brasier et al. [1]) and incubated at 20 °C for 48 h. Then, resulting Phytophthora colonies were transferred to potato-dextrose agar (PDA) medium.
For DNA extraction, soil samples were air dried in metal trays at 80 °C for 72 h in an oven. Once dried, samples were homogenized, sieved through a 2 mm sieve, and stored at room temperature until DNA extraction.
In total, 14 sites, nine “disturbed” sites (DS1 to DS9) and five “undisturbed” sites (US1 to US5) were sampled. At each site, different locations were selected for sampling, making a total of 132 samples. The samples collected within the “disturbed” sites were ten different locations at sites DS1, DS2, DS3, DS4, DS5, and DS6. Five locations were selected at sites DS7 and DS8. At site DS9, 11 locations were sampled; the additional sampled area corresponded to an area where an outbreak of P. ramorum occurred and was eradicated in 2009. Within the “undisturbed” sites, ten different locations were selected within sites (US2 to US5) and 11 locations were selected at US1. A total of 81 samples were collected from the “disturbed” sites (DS1 to DS9) and 51 samples were collected from the “undisturbed” sites (US1 to US5) (Table 1).
2.2. Soil DNA Extraction and PCR Amplification
From each soil sample, DNA was extracted from three 0.25-g subsamples using the PowerSoil® DNA isolation kit (MoBio Laboratories Inc., Carlsbad, CA, USA) according to the manufacturer’s instructions. After DNA extraction, a clean-up was carried out using either the Jet-QuickTM DNA purification kit (Genomed GmbH, Löhne, Germany) or DNA Clean & ConcentratorTM kit (Zymo Research, Irvine, CA, USA) according to the manufacturer’s instructions. Extracted DNA was used to perform NGS analysis of the ITS rRNA and cytochrome c oxidase subunit I (COI) regions. A < 250 bp region of the ribosomal RNA (rRNA) Internal Transcribed Spacer (ITS1) was amplified using a nested PCR approach with primer pairs 18Ph2F and 5.8S-1R in the first round and ITS6 and 5.8S-1R in the second round according to the protocol of Scibetta et al. [46]. Amplification of the COI region was performed using a semi-nested PCR approach with primers cox1levup (TCAWCWMGATGGCTTTTTTCAAC)/cox1levlo (CYTCHGGRTGWCCRAAAAACCAAA) (Choi et al., 2015) in the first round and primers HVshort (GNATGAAYAAYATHAGYTTYTGG)/cox1levlo in the second round. Primer HVshort was designed in this study to amplify an internal portion of the COI region of appropriate size for Illumina sequencing. For the COI region, the amplification conditions were the same as those reported in Choi et al. [40].
Cultures obtained from soil baiting were identified by amplification and sequencing of their ITS region using the universal primers ITS-6 [28] and ITS-4 [47].
2.3. Positive Controls for Illumina Sequencing
Initially, a development of positive controls for Illumina sequencing runs was performed. Control DNA mixes were set up containing the following ten Phytophthora species: P. boehmeriae, P. capsici, P. castaneae, P. fallax, P. foliorum, P. idaei, P. obscura, P. plurivora, P. rubi and P. siskiyouensis. Except for P. capsici, P. castaneae and P. fallax, all species are known to be present in the UK. Some of those species such as P. rubi and P. idaei are common in the UK but not known to be present in forests. Genomic DNA of Phytophthora spp. isolates was extracted from mycelium harvested from cultures on V8 agar using the Plant DNAeasyTM mini kit (Qiagen, Germany) according to the manufacturer’s instructions. A SYBR green quantitative real-time PCR assay was carried out in a Roche LightCycler® 480 Real Time PCR System with the primers 18Ph2F and 5.8S-1R, using Roche SYBR Green I master mix with the same PCR conditions as described in Scibetta et al. [46]. The DNA of each Phytophthora species was diluted 2.5, 5, 10, and 25 times and cycle threshold (Ct) values at each concentration were determined. DNA dilutions corresponding to approximately similar Ct values of 25 ± 3 (amplification curves) were pooled at equal volumes for the DNA control mix to account for differences in ITS copy number [32]. Thus, in theory, the Illumina sequencing potential for each species should be equal within the pooled control mix.
2.4. Illumina Sequencing Library Preparation and Sequencing
The triplicated PCR amplicons from the same forest soil sample that were positive for Phytophthora spp. using the nested PCR approaches described above were pooled and prepared for NGS following the protocols described for 16S Metagenomic Sequencing Library Preparation according to the manufacturer’s instructions [48]. In brief, 1 μL of the pooled DNA preparations was amplified using the nested PCR as indicated above with the exception that in the second round, the primers ITS6 and 5.8S-1R (ITS region) or HVshort and cox1levlo (COI region) both with overhang adapters were used to ensure compatibility with the Illumina indices and sequencing adapters. PCR products were cleaned up using Agencourt® AMpure® XP beads followed by index PCR in KAPA HiFi HotStart ReadyMix to attach dual indices to each sample using the Nextera XT Index Kit to ensure that each sample could be uniquely identified during the sequencing run. Then, a second PCR clean-up (as above) was carried out, and the DNA of each sample visualized on an Agilent 2200 TapeStation (Agilent Technologies, Edinburgh, UK) and quantified using a Qubit® fluorimeter (Life Technologies, Paisley, UK). At least three independent quantification measurements were performed, and the libraries consisting of a maximum of 96 samples were diluted using 10 mM Tris ph 8.5 to 4 nM before pooling. The ITS libraries were sequenced using two runs of a 500-cycle MiSeq Reagent Nano Kit v2 (Illumina, San Diego, CA, USA), which has a maximum output of 1 M reads for the ITS region, and using 20% of a 600-cycle MiSeq v3 run, which has a maximum output of 25 M reads (Illumina, San Diego, CA, USA) for the COI region. The sequencing was done at the James Hutton Institute, Dundee, UK and at the Fundación Parque Científico de Madrid, Madrid, Spain for the COI samples.
2.5. Bioinformatic and Statistical Analyses
ITS rRNA and COI mtDNA libraries were trimmed using Trim-Galore software [49], removing short-length, low quality, and PCR adapter sequences. The quality of the DNA libraries before and after the trimming step were checked with FASTQC [50], and all output files were merged into a single report using MultiQC [51].
Then, high-quality and trimmed sequences were processed within Insights Into Microbial Ecology 2 (QIIME 2 v2020.2) [52]. An amplicon sequence variants (ASVs) table was generated by means of the DADA2 pipeline [53], and potential chimeras were detected and removed using subsequently both uchime-denovo and uchime-ref approaches from the V-SEARCH algorithm [54]. For the uchime-ref, we used the complete “BARCODE OF LIFE DATA SYSTEM v4 (http://www.barcodinglife.org)” database updated on 11 June 2020 with a record of 11,234,685 specimens from Animals, Plants, Fungi and Protists [55]. Then, ASV sequences were clustered at 99% of identity using the V-SEARCH algorithm [54] to collapse very close sequences and create a non-redundant ASVs table. Then, an initial taxonomic assignment was made with GenBank nt database (updated on 13 June 2020) using BLASTN implemented in the “qiime brocc classify-brocc” plugin (https://github.com/kylebittinger/q2-brocc (accessed on 24 September 2018)), with a p-min-genus-id (minimum percent identity for genus-level consensus) parameter fixed to 90% of identity. Then, those ASV sequences identified as Phytophthora spp. were checked against two curated databases: (i) CPSM, which is a database provided by Dr. Treena Burgess (Centre of Phytophthora Science and Management; Murdoch University, WA, Australia) and (ii) BOLD v4, The Barcode of Life Database V4. Finally, those ASV sequences showing <99% BLASTN identity with any of the sequences in the reference databases (CPMS and BOLD), were submitted to a phylogenetic analysis to show their positions in the different Phytophthora taxonomic clades. For that, COI and ITS ASV sequences and curated DNA sequences from the CPSM database were aligned using MAFFT [56], using the “auto” mode to select an appropriate alignment strategy. Alignments were visualized, and the consensus region of the alignment between the ASVs and the reference sequences of the database was extracted. Subsequently, the consensus regions of the alignment were processed by trimAl [57] using the flag automated1 for the automated removal of spurious sequences or poorly aligned regions from multiple sequence alignment. To select the substitution model that best fits our data, we ran MODELTEST [58] with the default parameters, which found that GTR+I+G4 was the best model of nucleotide substitution according to BIC (Bayesian information criterion) and AIC (Akaike information criterion) model selection techniques. Finally, maximum likelihood trees (ML) were calculated using IQtree v2.0.3 [59] for DNA sequences with a 1000 bootstrap and option bnni to reduce the risk of overestimating branch supports with UFBoot [60]. ML trees were visualized with FigTree v1.4.4 (https://github.com/rambaut/figtree/ (accessed on 1 January 2021)) software.
To determine the potential anthropogenic impact on the woodlands of UK among the different sites catalogued as “disturbed” and “undisturbed”, we performed a permutational multivariate analysis of variance (PERMANOVA) [61] for each matrix of Phytophthora spp. identified with ITS and COI regions using the Bray–Curtis (abundance) and Jaccard (present/absence) distance-based dissimilarity matrices. Counts data were transformed using the Hellinger method [62] before analysis. Additionally, to evaluate differences in the variability of species composition among groups, we used a multivariate analogue of Levene’s test for homogeneity of variances (Betadisp) with 999 permutations for each case to compare the mean distance-to-centroid of study group [63]. In order to visualize the spatial ordination of these results, principal coordinates analysis (PCoA) plots were calculated for each comparison. All multivariate analyses were performed in the statistical software R with the VEGAN package [64].
3. Results
3.1. Sequencing Output and Performance of Control Reactions
After the quality control step of the NGS libraries, we obtained a total of 1,192,022 and 1,931,588 high-quality sequences, with an average length of 290 and 242 bp, for ITS and COI regions, respectively. In both trimmed libraries, the Q score of quality was higher than 36. The DADA2 workflow generated 147 and 1537 ASVs, which after the elimination of artefacts and unidentified sequences, a total of 97 ASVs (with a mean length of 554 bp) and 710 ASVs (with a mean length of 444 bp) were kept for further analysis, for the COI and ITS regions, respectively. No reads were detected in any of the negative control samples included in each plate and run.
In the control reactions, a total of 11,131 and 6506 assembled sequences were generated from the ITS and COI sequences, respectively, comprising 10 Phytophthora species. For ITS, all Phytophthora spp. included in the control samples, except P. boehmeriae, were detected. However, P. foliorum, P. idaei and P. siskiyouensis were detected in 66.7% of control samples, and the rest of Phytophthora spp. were detected in all control samples (Table 2). For COI, all Phytophthora spp., except P. foliorum, were detected in the control samples. P. rubi was detected in 33.3% of control samples, P. idaei and P. plurivora were detected in 66.7% of control samples, and the rest of Phytophthora spp. were detected in all control samples (Table 2). No false positive Phytophthora species for any of the control reactions were detected. For ITS, and concerning the percentage of reads assigned to each Phytophthora spp., five Phytophthora spp. represented more than 85.8% of reads (from 12.0 to 24%), including P. capsici, P. castaneae, P. obscura, P. plurivora and P. rubi, whereas the remaining Phytophthora spp. represented 14.2% of reads only (from 1.6 to 5.4% of reads). For COI, five Phytophthora spp. represented more than 79.0% of reads, including P. capsici, P. fallax, P. idaei, P. obscura and P. siskiyouensis (from 9.1 to 30.8%), whereas the remaining Phytophthora spp. represented the remaining 21% of reads (from 1.5 to 7.9%) (Table 2).

Table 2.
Frequency of reads assigned to each Phytophthora sp. in the control samples and incidence of detection.
3.2. Analysis of Sequences from British Soil Samples
Amplification with COI primers yielded more total PCR positive samples than when using ITS primers, with the exception of “disturbed“ samples from DS2 and DS4 locations, when considering all PCRs amplifications for each of the three DNA extractions (N = 396 each) performed for each location (Supplementary Materials Figure S1A). For all “disturbed” and “undisturbed“ sites sampled, a positive PCR amplification was obtained in more than 80% of the locations sampled both when using primers targeting the ITS and COI region, with the exception of the DS5 “disturbed“ site and the US5 “undisturbed“ site in which the proportion of locations showing at least one successful PCR amplification when using ITS primers was much lower (Supplementary Materials Figure S1B).
However, NGS analyses of both the ITS and COI regions revealed that some of the amplified products did not belonged to sequences of Phytophthora, especially those reads obtained with primers targeting the COI region showed less specificity than ITS primers. Thus, ITS primers were very specific and most of the reads belonged to Phytophthora spp. (93% of reads); only a few sequences belonged to other oomycetes such as Peronospora spp. (3%), Bremia spp. (1%), Pythium spp. (0.9%), Nothophytophthora spp. (0.1%), and other uncultured ASV sequences of the Peronosporaceae family (2%) (Supplementary Materials Figure S2).
On the other hand, NGS of the COI region revealed that the largest group of sequences belonged to Oomycota (71%). However, the design of the primers was much less specific than those used for ITS, amplifying sequences from a wide range of organisms such as Bacteria (2, all of them Rhizobiales), Fungi (5%), Metazoa (0.9, mainly arthropods), Rhodophytes (Florideophyceae), or red algae (0.3%) from the Plantae kingdom, Phaeophyceae or brown algae (9%), Bacillariophyta or diatoms (3%), and other kingdoms much less abundant <0.1% (Supplementary Materials Figure S3). From all oomycetes, sequences assigned to the genus level of Phytophthora spp. represented 16% of them, and other genera from the Peronosporaceae family such as Peronospora (3%), Nothophytophthora (1%), and others uncultured Peronosporaceae (12%) were identified. In addition, different species from families such as Pythiaceae (41% of oomycete sequences) (Globisporangium spp., Pythium spp., Phytopythium spp., and others uncultured Pythiaceae), Saprolegniaceae (3%, Achlya spp., Saprolegnia spp., and Aphanomyces spp.) and other unidentified oomycetes (23%) were also identified (Supplementary Materials Figure S3).
3.3. Identification of Phytophthora Phylotypes in Britain by NGS
From all the reads identified as Phytophthora using the ITS region, 79 ASVs were identified. Of those, 20 ASVs were assigned to known species/phylotypes clusters of Phytophthora spp., and 21 ASVs were assigned to unknown/uncultured species of Phytophthora. Phytophthora spp. were detected in 79 locations of the 132 on 14 studied sites (Table 1, Supplementary Materials Table S1 and Figure S4). In this case, the databases of CPSM and BOLD were used for taxonomy assignation and comparison, and when a discrepancy was found, the procedure described by Idphy using the Genbank database was also performed. Most of the ASV taxonomic assignations were in agreement between the two databases used and corresponded to the following species: P. austrocedri, P. cactorum, P. capsici, P. castaneae, P. cinnamomi, P. fallax, P. foliorum, P. idaei, P. kernoviae, P. lacustris, P. obscura, P. primulae, P. pseudosyringae, P. siskiyouensis, P. ramorum, P. syringae and P. uniformis. The following few species did not agree with the databases or could not be reliably discriminated: P. megasperma/crassamura, P. plurivora/citricola and P. rubi/fragariae (Table 1, Supplementary Materials Table S1 and Figure S4). The other 21 ASVs were named as Phytophthora sp. uncultured 1a to 21a. Their sequences showed high similarity with sequences of already known/undescribed Phytophthora species such as P. alni/uniformis, P. cambivora, P. capsici/glovera, P. europaea/megasperma, P. iranica/clandestina, P. melonis/sinensis, P. quercina/P. sp. ohioensis, P. sojae and P. uliginosa, but their ITS sequence homology was below 99%, and the phylogenetic reconstruction also showed them to be very close but as independent branches (Supplementary Materials Figure S4). Most of these unidentified ASVs belonged to Clade 7, and the remaining belonged to Clades 1, 2, 3, 6, and 11. In addition, a new species of Nothophytophthora (ASV-56), placed in the same branch of the phylogenetic tree and showing more than 99% identity with N. vietnamensis and N. intricata, was only detected in one location from US4 (Table 1, Supplementary Materials Table S1 and Figure S4).
Results from NGS analysis of COI sequences from British soil samples revealed a total of 54 ASVs clustered within the genus Phytophthora that were assigned to 12 known Phytophthora species and 17 unknown Phytophthora phylotypes (named Phytophthora sp. uncultured from 1b to 17b), which were identified in 12 out of the 14 sites of the study (Table 1; Supplementary Materials Table S2 and Figure S5). The taxonomy assignation was obtained after comparing the two curated Phytophthora databases with results from GenBank using the procedure described by Idphy (https://idtools.org/id/phytophthora/molecular.php, accessed on September 2019). The main species detected using COI sequences that were in agreement with the three databases were P. cactorum, P. cinnamomi, P. europaea, P. gonapodyides, P. megasperma, P. plurivora, P. pseudosyringae, P. primulae, P. quercina, P. ramorum, P. syringae and P. uliginosa. The 17 uncultured phylotypes did not show a high percentage of similarity to any of the species present in the database, since homology in the COI sequences was below 99% or could not be reliably discriminated with any reference sequence using the ML phylogenetic reconstruction (Table 1 and Supplementary Materials Figure S5). It is important to highlight that 14 ASVs classified as Phytophthora spp. (Phytophthora uncultured 12b to 17b) showed a homology of sequences between 93 and 95% with known reference sequences such as P. agathidicida, P. castaneae, P. heveae, P. litchii/himalsilva and P. nierderhauserii/quercetorum. Moreover, these were clustered, although with low bootstrap support, in a new independent group according to phylogenetic analysis, which may indicate that they belong to a new unidentified clade. In addition, eight ASVs were assigned to the genus Nothophytophthora (Nothophytophthora spp. from 1 to 8) and were detected at the following sites DS7, DS8, DS9, US3, and US4 (Table 1 and Supplementary Materials Figure S5).
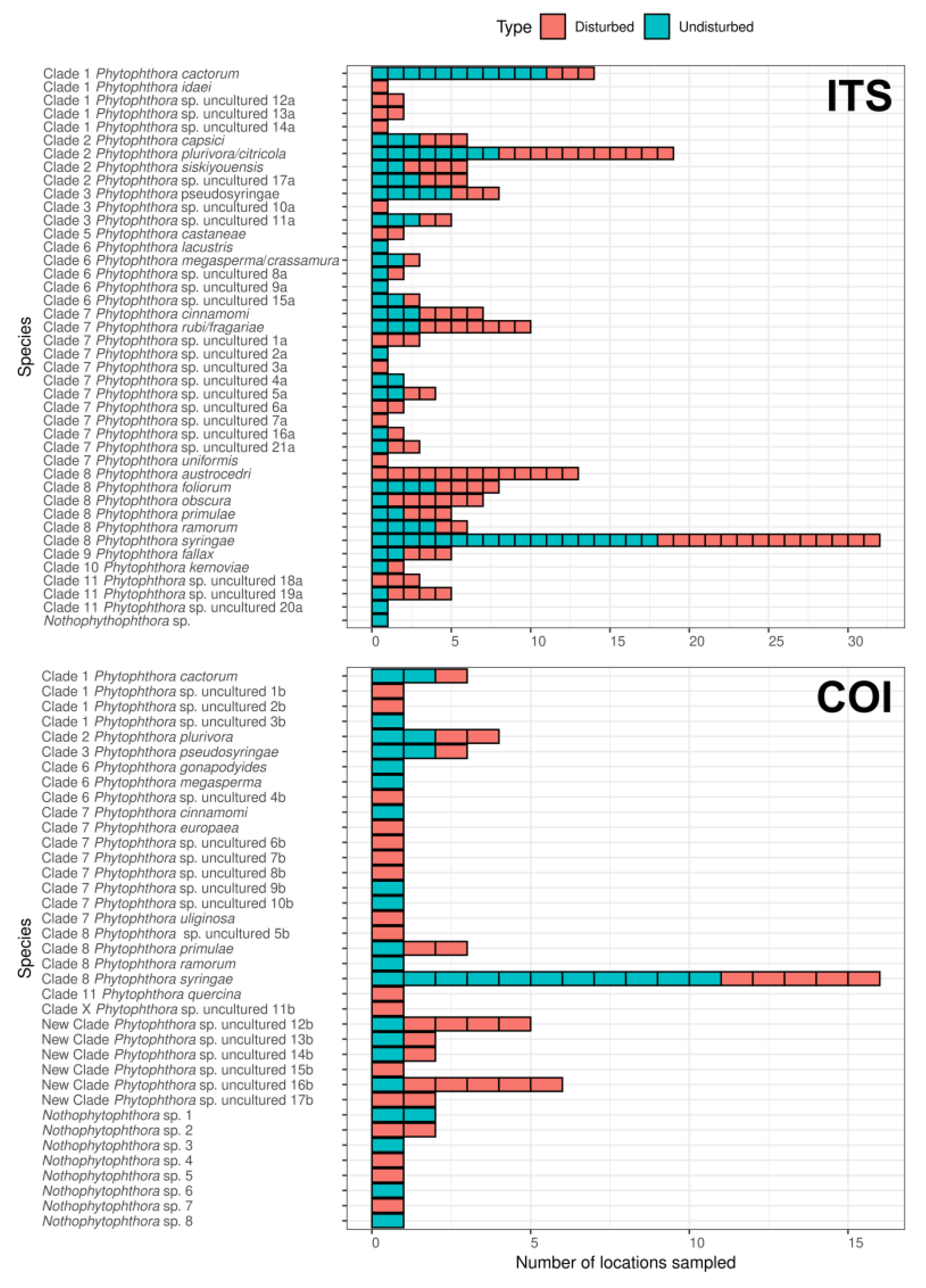
Eight Phytophthora species were detected in the soil samples when using both ITS and COI regions including P. cactorum, P. cinnamomi, P. megasperma, P. plurivora/citricola, P. primulae, P. pseudosyringae, P. ramorum and P. syringae. On the other hand, 12 species were only detected when using the ITS region including P. austrocedri, P. capsici, P. castaneae, P. fallax, P. foliorum, P. idaei, P. kernoviae, P. lacustris, P. obscura, P. rubi/fragariae, P. siskiyouensis and P. uniformis, whereas four species were detected only when using the COI region including P. europaea, P. gonapodyides, P. quercina, and P. uliginosa (Table 1, Figure 2 and Supplementary Materials Tables S1 and S2).
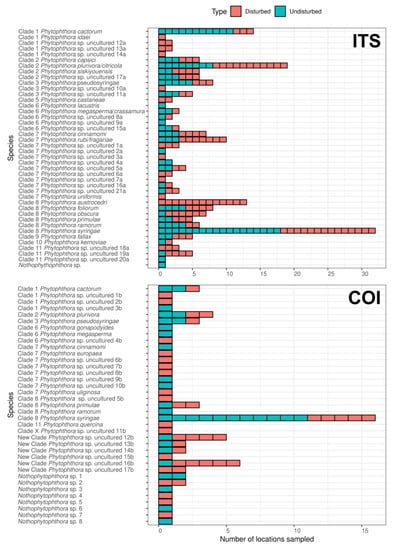
Figure 2.
Occurrence of Phytophthora spp. and Nothophytophthora spp. in British soils in the different location sampled in “disturbed” or “undisturbed” sites determined based on next-generation sequencing (NGS) analyses of the ITS and COI regions.
Phytophthora syringae was the most abundant and most prevalent species among all samples for both ITS and COI. This species was detected in six or 10 of the 14 studied sites and in 32 or 16 locations sampled when using the ITS or COI region, respectively, and both in “disturbed” and “undisturbed” sites (Table 1, Figure 2; Supplementary Materials Tables S1 and S2). It also represents, with eight or four associated ASVs, 35% (4.363) or 26% (83.642) of the total reads detected when using ITS (Supplementary Materials Table S1) or COI (Supplementary Materials Table S2), respectively. Other Phytophthora spp. that were also found as most prevalent when using the ITS region included P. plurivora, which was detected in 10 sites at 19 locations, and P. primulae, which was detected in three sites and five locations (Table 1, Figure 2); when using the COI region, P. plurivora and P. primulae were detected at four and three sites, respectively with a similar number of locations (Table 1, Figure 2). When using the ITS region, several Phytophthora spp. appeared also to be widespread, including P. cactorum, P. obscura, P. pseudosyringae, P. rubi/fragariae and P. siskiyouensis. These were detected in at least five sites and a number of locations ranging from six to 19 locations (Figure 2; Supplementary Materials Table S1). Interestingly, when using the COI region, two unknown Phytophthora spp. (New Clade Phytophthora uncultured 12b and 16b) appeared as the most abundant species being detected in five and four sites and in five and six locations, respectively (Figure 2; Supplementary Materials Table S2).
3.4. Identification of Phytophthora by Isolation
Eight Phytophthora species (P. cactorum, P. chlamydospora, P. cinnamomi, P. cryptogea, P. gonapodyides, P. megasperma, P. plurivora and P. ramorum) were isolated from the sampled soils at five of the “disturbed” sites (DS1, DS2, DS3, DS5, and DS9) and two of the “undisturbed” sites (US3 and US4) (Table 1). Only on two of the locations were the following species confirmed by ITS, COI, and isolation: P. plurivora on DS5 (DS5-4) and US3 (US3-2). At one of the sites, P. plurivora was isolated at site DS2 (DS2-10) and only confirmed by ITS, and P. cactorum was isolated at US3 (US3-9) and confirmed only by ITS. However, there were sites where Phytophthora species were isolated, and they were not detected by NGS. This was the case at the sites DS1 (P. cryptogea, P. cinnamomi and P. megasperma), DS2 (P. plurivora), DS3 (P. cinnamomi, P. chlamydospora and P. plurivora), DS9 (P. chlamydospora, P. gonapodyides, P. plurivora and P. ramorum), US3 (P. chlamydospora and P. plurivora) and US4 (P. chlamydospora, P. gonapodyides and P. plurivora).
3.5. Phytophthora spp. Community Composition Differs between Disturbed and Undisturbed Sites
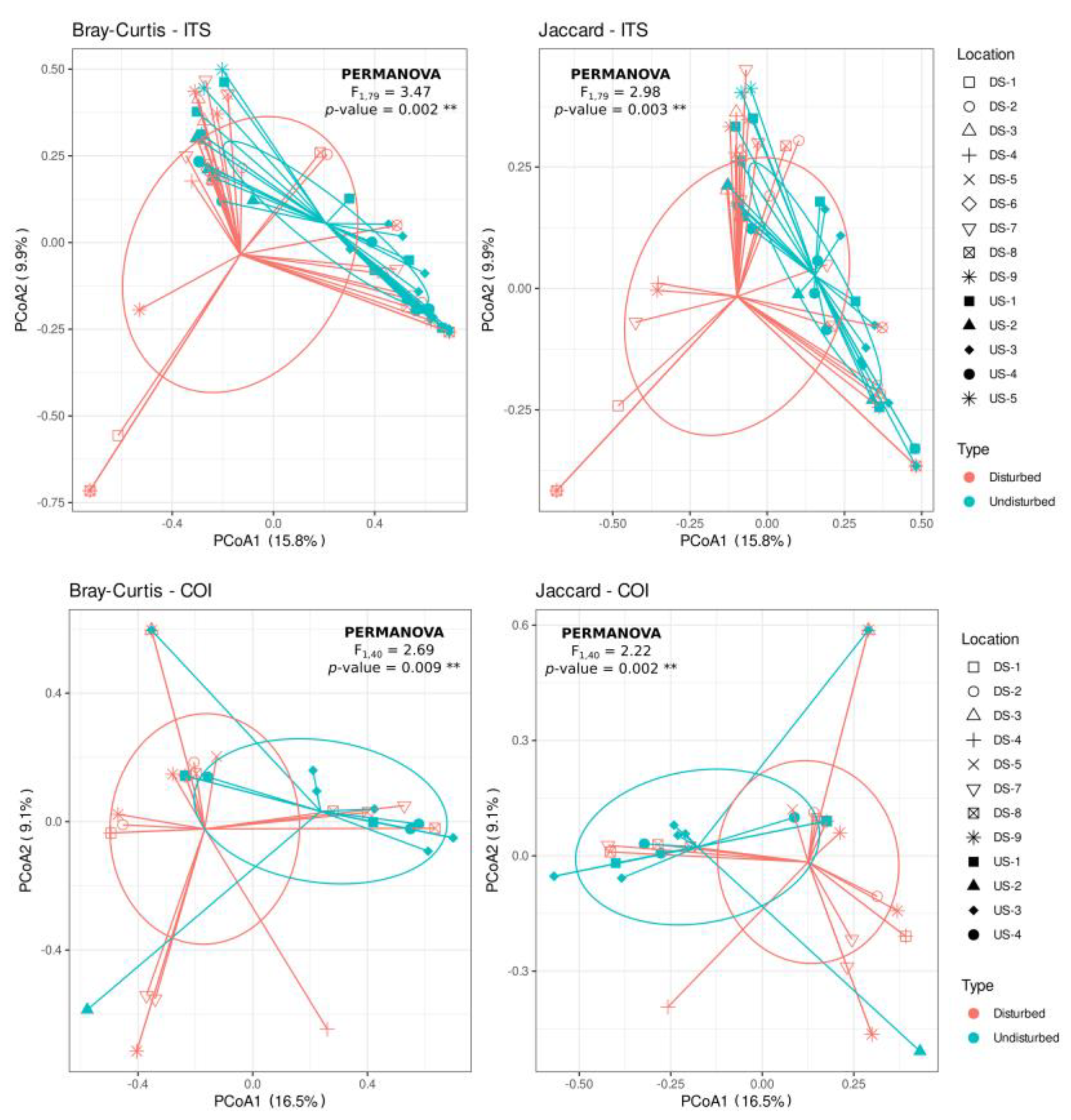
PCoA of Bray–Curtis and Jaccard distances showed a trend to differentiate Phytophthora communities according to the disturbance of the sites (i.e., “disturbed” versus “undisturbed”) (Figure 3). Thus, when using both the ITS or COI regions, there was a significant effect of soil disturbance on the composition of Phytophthora species found on those soils, both for Bray–Curtis (PERMANOVA, F > 2.687, p < 0.009) and Jaccard (PERMANOVA, F < 2.982, p < 0.003) distances (Supplementary Materials Table S3). However, the dispersion of the species communities across samples differed significantly for Bray–Curtis distance-based matrices (F > 4.497, p < 0.045) but not for the Jaccard distances (F < 3.427, p > 0.080), which indicated that for Bray–Curtis ordination (abundance), there is a heterogeneous dispersion, whereas that does not occur for Jaccard ordination (presence/absence) (Supplementary Materials Table S3).
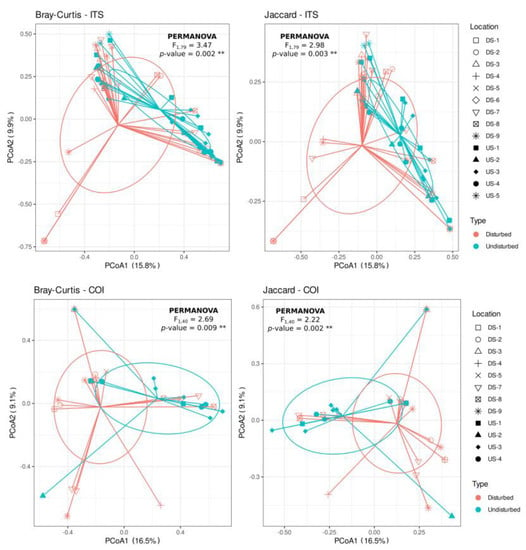
Figure 3.
Principal coordinates analysis (PCoA) of Bray–Curtis and Jaccard distance matrices from Phytophthora spp. and Nothophytophthora spp. community composition data obtained using COI and ITS markers. PCoA were plotted in the form of a spider diagram with “legs” joining samples that belong to the same cluster (centroid). Shapes on the figure legend correspond to the locations and colors to the sites (“disturbed” and “undisturbed”) sampled.
Concerning differences between “disturbed“ and “undisturbed” sites, a higher number of Phytophthora spp. were detected at “disturbed“ sites, both when using ITS (36 species) or COI (26 species) as compared to “undisturbed“ sites, where 29 and 20 species were detected when using the ITS and COI region, respectively (Figure 2; Supplementary Materials Tables S1 and S2).
When using the ITS region, of the total of 41 Phytophthora species identified, 13 were only detected in “disturbed” soils (P. austrocedri, P. castaneae, P. idaei, P. uniformis, and the uncultured Phytophthora 1a, 3a, 6a, 7a, 10a, 12a, 13a, 14a, and 18a) and five were detected in “undisturbed” soils (P. lacustris, and the uncultured Phytophthora 2a, 9a, 4a, and 20a) (Figure 2; Supplementary Materials Table S1). The unique Nothophytophthora sp. identified was detected in one location at the “undisturbed” site US4. The remaining 23 were detected in both types of soils. Phytophthora austrocedri was only detected at “disturbed” sites and showed a remarkably high prevalence at one of the sampled sites (DS1) (Figure 2; Supplementary Materials Table S1).
From the total of 29 Phytophthora species identified when using the COI gene, 13 Phytophthora spp. were detected only in “disturbed” soils (P. europaea, P. quercina, P. uliginosa, and several uncultured Phytophthora spp. from Clade 1 (Phytophthora sp. uncultured 1b, 2b), Clade 6 (Phytophthora sp. uncultured 4b), Clade 7 (Phytophthora sp. uncultured 6b, 7b, 8b), Clade 1 (Phytophthora sp. uncultured 5b), Clade X (Phytophthora sp. uncultured 11b), and from the new candidate Clade (Phytophthora sp. uncultured 15b and 17b)) (Figure 2; Supplementary Materials Table S2). On the other hand, seven were located exclusively in “undisturbed” soils (P. cinnamomi, P. gonapodyides, P. megasperma, P. ramorum, and two uncultured Phytophthora spp. from Clade 1 and one from Clade 7). From the eight Nothophytophthora spp. detected, four were detected at “disturbed” sites (DS7, DS8, and DS9) and a similar number were detected at “undisturbed” sites (US2 and US3) (Figure 2; Supplementary Materials Table S2).
4. Discussion
Although it is not known how many Phytophthora species are already present in British soils, at least 57 species have been detected in the UK [65,66]. Techniques such as high-throughput sequencing (HTS) technologies or NGS have been used recently to detect alien species in soil samples. While some of these techniques have been used to detect communities (all organisms) in environmental samples [33,67,68], others have been targeting a selected group of them. In the case of Phytophthora, there are several studies that have been targeting the ITS region, and they also have shown the lack of discrimination of Phytophthora species within some clades such as clades 2 and 3 [32,35,69]. Some authors have indicated the higher resolution of COI as compared to ITS within the genus Phytophthora [31,39,40]. However, there are no previous NGS studies comparing the Phytophthora community in soils using both ITS and COI. This is the first study where additionally to the ITS region, the COI region was targeted in order to investigate if the detection of oomycetes in soils could be improved and the discrimination of Phytophthora within clades could be better resolved.
To test the consistency of PCR amplification of Phytophthora present in the samples and compare the efficiency of ITS versus COI primers, we included control DNA mixes containing 10 Phytophthora spp. We found considerable variability in the number of Illumina reads generated per species that also differed between both markers. With ITS, we were able to amplify all the species included in the control samples except P. boehmeriae. A similar result was recently found by Riddell et al. [35] with the same species of Phytophthora. On the contrary, when using the COI region, we were not able to detect P. foliorum. One potential explanation to account for the lack of amplification of those Phytophthora sequences or a different amplification efficiency among the remaining ones may be due to the presence of several base mismatches in the primers used in the second round of PCR, which can induce some PCR bias, since the Ct values of the DNA dilutions of each Phytophthora spp. included in the control reactions were in general consistent across each species used in the DNA control mixture [35].
Although the ITS region has proven valuable in Phytophthora diagnostics, the intraspecific and intra-genomic sequence variation of this region is an important consideration for the accurate identification of several species, emphasizing the need for the further validation and refinement of reference databases. In our study, we found that, as expected, the COI region was able to better differentiate several Phytophthora spp. from clades that are difficult to discriminate when using the ITS region. However, as a drawback, several oomycete sequences and sequences from other Phyla were detected using the COI region, which generated a lower number of Phytophthora reads compared to the ITS. Nevertheless, we were able to confirm the presence of several Phytophthora spp. in British soils using both markers. It would be interesting to test whether other genes that have shown good for discrimination of Phytophthora spp. isolates when using pure cultures [31,37] may be used in NGS analysis and provide complementary results to that obtained for COI and ITS.
Thus, a total of 20 ASVs from eight clades (1a, 2, 2b, 3a, 5, 6b, 7a, 8b, 8c, 8d, 10a, and 10b) and 12 ASVs from six clades (1a, 2c, 3a, 3b, 6b, 7a, 8b, 8c, and 8d) were assigned to known Phytophthora species using the ITS and COI, respectively. Of those, only eight Phytophthora species were detected by both ITS and COI: P. cactorum, P. cinnamomi, P. megasperma, P. plurivora, P. primulae, P. pseudosyringae, P. ramorum and P. syringae. The results on these species were in agreement in 21 of the sampled locations at the different sites. No shared Phytophthora species were detected at four of the studied sites, and there were 51 locations where no Phytophthora was amplified by ITS or COI. Only in one site, US3, ITS and COI were in agreement for all the locations, and at the site US4, they were in agreement in seven of the 10 locations. NGS analysis provides evidence of the presence of Phytophthora DNA, but re-sampling the sites to obtain living cultures from soil or from host material should be attempted to confirm their presence. It is especially important for the new records of Britain. Special care should be taken for processing samples, since soil storage could decrease the viability of propagules and might diminish the success of pathogen isolation.
The results from this study also detected one and eight ASVs belonging to Nothophytophthora spp., when using the ITS or COI markers, respectively. This is the first report of species of Nothophytophthora detected through NGS, and to date, species within this new genus have not been detected in Britain. In this study, Nothophytophthora species were detected at five sites. Only one Nothophytophthora sp. was detected with ITS and COI at site US4 (US4-10), whereas with COI, eight Nothophytophthora spp. (1–8) were detected in five sites: DS7 (Nothophytophthora sp. 4), DS8 (Nothophytophthora sp. 2 and sp. 5), DS9 (Nothophytophthora sp. 7), US3 (Nothophytophthora sp. 1 and sp. 6), and US4 (Nothophytophthora sp. 3 and sp. 8). Nothophytophthora was described recently as a new sister genus to Phytophthora from natural ecosystems [70]. Pathogenicity of Nothophytophthora species still needs to be determined, but their association with symptomatic plants suggest they might be pathogenic. Resampling at these sites would be recommended to obtain isolates of these new species.
This study also compares which Phytophthora species are present at “disturbed” sites where diseases caused by these pathogens are common and at “undisturbed” sites where neither Phytophthora nor diseased plants associated with Phytophthora have been recorded previously.
This study shows that Phytophthora species within clade 8 were the most commonly identified by either ITS and COI (P. austrocedri, P. foliorum, P. obscura, P. primulae, P. ramorum and P. syringae) followed by Phytophthora species within clade 7 (P. cinnamomi, P. europaea, P. uniformis, P. uliginosa). The abundance of Phytophthora species within clade 8 in “disturbed” soils in Britain was also reported by Riddell et al. [35]. In their study, they also detected P. lateralis, which is a very significant pathogen of Chamaecyparis lawsoniana in the US and has been recently detected in the UK [4]. However, this species was not detected in this study. On the other hand, in this study, P. foliorum, also from clade 8, was detected in three “disturbed” sites (DS7, DS8, DS9) and in two “undisturbed” sites (US3, US5), and this species was not detected in their study. Phytophthora foliorum was only detected in the UK in 2016 on Rhododendron ponticum in Scotland [71]. This species was described causing leaf blight on azaleas in the US in 2006 [72] and has been found in nurseries in Europe [8].
Clades 7 and 8 contain very important pathogens threatening forest and woodlands tree species worldwide. Of the six species detected in clade 8, four of them (P. austrocedri, P. foliorum, P. obscura and P. ramorum) were only described between 2002 and 2012 and are considered recently introduced in UK. At present, P. austrocedri and P. ramorum are causing great environmental and economic losses in the UK. Phytophthora austrocedri was only discovered in Britain in 2011 causing dieback and mortality in northern England and Scotland on Juniperus communis, which is one of the three native conifers in the UK [73,74,75]. This species was described from southern Argentina where it is responsible for the mortality of their native cypress Austrocedrus chilensis [76]. The lack of genetic diversity of P. austrocedri in the UK and their aggressiveness on a native host support the hypothesis that P. austrocedri has been recently introduced into Britain [74,75]. In this study, this species was detected at four of the “disturbed” sites (DS1, DS4, DS7, and DS9) but not at any of the “undisturbed” sites, which would support the above hypothesis.
Phytophthora ramorum was described in Germany [77] and is responsible for sudden oak death in US. In the UK, this is a regulated organism and was initially discovered on Viburnum tinus in the south of England [78], and then, it was recorded mainly on Rhododendron or other ornamentals in nurseries, plant retail outlets, gardens, parks, and woodlands [79,80,81,82,83]. However, it was in 2009 that P. ramorum was observed causing mortality in mature and juvenile plantations of commercially grown Japanese larch (Larix kaempferi), and since then, it has been causing great economical losses. Other tree species have also been affected by P. ramorum, Fagus sylvatica, Nothofagus obliqua, Castanea sativa, Betula pendula, Tsuga heterophylla and Pseudotsuga menziesii amongst others [84]. In this study P. ramorum was detected at two of the “disturbed” sites (DS2 and DS4), and at three of the “undisturbed” sites (US1, US3, and US5). This is surprising as this species is to be expected mainly at “disturbed sites” where new plantings have occurred recently or at sites in close proximity to affected larch. Further investigations and isolations from soil samples need to be attempted at these ”undisturbed” sites. However, it is interesting that P. ramorum was isolated from DS9 from a site where the disease was eradicated in 2009. The affected plant back in 2009 was a Rhododendron, which was removed and burned. That area was grassed over, and there are not woody hosts growing there. However, P. ramorum was isolated from the soil underneath the grass, confirming that the pathogen can survive at least nine years in the soil after eradication. These results indicate that re-sampling the sites where some Phytophthora outbreaks have occurred would be advisable to test whether viable propagules of the pathogen are still present in those soils.
Phytophthora obscura was described in the USA in 2012 [85] on foliage of Kalmia latifolia, in substrate of Pieris, and in soil around Aesculus hippocastanum in Germany. This species was detected at three sites in British soils [35]. In this study, it was detected at five “disturbed” sites (DS2, DS3, DS4, DS7, and DS8) and at one “undisturbed” site (US1). More information is needed regarding this species and the possible threat to British trees.
In this study, Phytophthora phylotypes that could not be assigned to known Phytophthora species were identified by both ITS (22) and COI (17). Some of these species could be identified as known Phytophthora species if the cutting point was lowered, but the approach taken in this study was conservative, and only sequences with ≥99% similarity were identified to the species level. When the similarity was below 99%, phylogenetic trees were constructed to place them in their clades and show their genetic distances within the clades. As a result of this approach, some of these species might be known species, and some of them could be new to science. For example, Clade 1 Phytophthora sp. uncultured 12 has a high similarity with species detected in at least two previous studies [32,86], or Clade 6 Phytophthora sp. uncultured 9 has a high similarity to species detected by Reeser et al. [87]. Therefore, it is important to try to isolate these species to confirm the results obtained here and to study their biological features and their pathogenicity. In Britain, since the mid-1990s, new or invasive Phytophthora species have been described affecting trees in different environments. Two of these species were novel and were detected in the UK before being discovered in other countries. This was the case for example of Phytophthora alni, which was described in 1993 from riparian alder (Alnus glutinosa) in Britain [88] and P. kernoviae, which was discovered in 2003 in the southwest of England on Fagus sylvatica and Rhododendron ponticum (Brasier et al., 2005). Phytophthora kernoviae is also a regulated organism in Britain and was detected in this study at site DS6 (young plantation) and US2, which is in close proximity to DS6. These two sites are next to a nursery, and inspection of the nursery would be recommended. The closest host plant for a soil sample from DS6 was a healthy young Pinus sylvestris, and for US2, it was a mature dead Juniperus communis. In the latter, P. austrocedri was expected, but none were detected at this site. However, the detection of P. kernoviae close to a healthy Scots pine needs to be followed up, as recently, P. pluvialis and P. kernoviae have been associated with needle diseases of Pinus radiata in New Zealand [89], and this could be an emerging problem on pine.
The detection of species previously unknown in Britain and detected in this study were P. castaneae, P. capsici, and P. fallax. These species were only detected with the ITS primers and not with COI. They are host specific and have been described mainly on Castanea, Solanaceae, and Eucalyptus, respectively [90,91]. Confirmation of their presence at the sites is recommended.
5. Conclusions
This study shows that Phytophthora species are present at “disturbed” and “undisturbed” sites in British soils independent of the presence of declined trees. As we hypothesized, more Phytophthora species were detected at “disturbed“ than at “undisturbed“ sites, and this was confirmed by NGS of both ITS and COI regions.
This is the first study using COI to study the Phytophthora community in soils, and the results presented showed that COI primers were less specific than the ITS when targeting Phytophthora species. However, COI amplified more Oomycota and a wider range of organisms than when using ITS primers.
This is the first detection of Nothophytophthora species in British soils, where it was detected at both “disturbed” and “undisturbed” sites, and the first detection of Nothophytophthora species using NGS analysis with both ITS and COI regions.
Of the known Phytophthora species detected in this study, only three are not known to be present in the UK. Therefore, re-sampling of these sites to confirm their presence by isolation would be recommended not only for these three species but for all the potential new Phytophthora species detected by both ITS and COI regions.
Supplementary Materials
The following are available online at https://www.mdpi.com/1999-4907/12/2/229/s1. Figure S1: Frequency of total samples (A) and locations (B) sampled in “disturbed” and “undisturbed” soils showing positive PCR amplification using primers targeting ITS or COI region. Figure S2: Krona chart showing the relative abundance and diversity from phylum to genus level of all ASVs obtained by NGS using the ITS marker. Figure S3: Krona chart showing the relative abundance and diversity from the phylum to genus level of all ASVs obtained by NGS using the COI marker. Figure S4: Maximum-likelihood tree of ITS sequences obtained in this study and reference sequences inferred in IQ-TREE with the GTR + I + G4 model. Values at branches correspond to bootstraps support. Figure S5: Maximum-likelihood tree of COI sequences obtained in this study and reference sequences inferred in IQ-TREE with the GTR + I + G4 model. Values at branches correspond to bootstraps support. Table S1: Distribution of the different Amplicon Sequence Variants of Phytophthora/Nothophytophthora spp. and number of reads identified using the ITS marker across the 14 sites “disturbed” and “undisturbed” sites sampled in this study; Table S2: Distribution of the different Amplicon Sequence Variants of Phytophthora/Nothophytophthora spp. and number of reads identified using the COI marker across the 14 sites “disturbed” and “undisturbed” sites sampled in this study; Table S3: Permutational multivariate analysis of variance (PERMANOVA) and permutational analysis of multivariate dispersion (Betadisper) results based on Bray–Curtis and Jaccard distances using Hellinger data transformation, for Phytophthora/Nothophytophthora spp. community composition data using ITS and COI marker genes.
Author Contributions
Conception and design of the research, B.B.L., B.H. and A.P.-S.; soil sampling, B.H. and A.P.-S.; isolation and soil processing, B.H., L.A.S. and A.P.-S.; acquisition of data, B.B.L., M.M.-B., B.H. and A.P.-S.; analysis and interpretation of data, L.F.A.-G., B.B.L., B.H., M.M.-B. and A.P.-S.; statistical and bioinformatics analysis, L.F.A.-G. and B.B.L.; writing the manuscript, L.F.A.-G., B.B.L., B.H. and A.P.-S. All authors have read and agreed to the published version of the manuscript.
Funding
This work has received funding from the European Union’s Horizon 2020 research and innovation programme under grant agreement No 635646, POnTE (Pest Organisms Threatening Europe).
Data Availability Statement
The raw sequence data have been deposited in the Sequence Read Archive (SRA) database at the NCBI under BioProject accession number PRJNA690943 and PRJNA691575.
Acknowledgments
From Forest Research (FR), we would like to thank Alex Lewis for producing the map for Figure 1 and Caroline Gorton for her help processing some of the soil samples. We would like to thank Liz Richardson for helping on site selection, and we thank Forestry England and other site owners for allowing FR to collect the soil samples and for being part of this study. We would also like to thank Treena Burgess (Centre of Phytophthora Science and Management; Murdoch University, WA, Australia) for providing two curated Phytophthora databases for ITS and COI sequences.
Conflicts of Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Brasier, C.M.; Beales, P.A.; Kirk, S.A.; Denman, S.; Rose, J. Phytophthora kernoviae sp. nov., an invasive pathogen causing bleeding stem lesions on forest trees and foliar necrosis of ornamentals in the UK. Mycol. Res. 2005, 109, 853–859. [Google Scholar] [CrossRef] [PubMed]
- Brasier, C.; Webber, J. Sudden larch death. Nat. Cell Biol. 2010, 466, 824–825. [Google Scholar] [CrossRef] [PubMed]
- Robin, C.; Piou, D.; Feau, N.; Douzon, G.; Schenck, N.; Hansen, E.M. Root and aerial infections of Chamaecyparis lawsoniana by Phytophthora lateralis: A new threat for European countries. For. Pathol. 2010, 41, 417–424. [Google Scholar] [CrossRef]
- Green, S.; Brasier, C.M.; Schlenzig, A.; McCracken, A.; Macaskill, G.A.; Wilson, M.; Webber, J.F. The destructive invasive pathogen Phytophthora lateralis found on Chamaecyparis lawsoniana across the UK. For. Pathol. 2012, 43, 19–28. [Google Scholar] [CrossRef]
- Hansen, E.M. Phytophthora Species Emerging as Pathogens of Forest Trees. Curr. For. Rep. 2015, 1, 16–24. [Google Scholar] [CrossRef]
- Scanu, B.; Webber, J.F. Dieback and mortality of Nothofagus in Britain: Ecology, pathogenicity and sporulation potential of the causal agent Phytophthora pseudosyringae. Plant Pathol. 2016, 65, 26–36. [Google Scholar] [CrossRef]
- Greslebin, A.G.; Vélez, M.L.; Green, S. Phytophthora Diseases: Mal del Ciprés; American Phytopathological Society: St. Paul, MN, USA, 2018. [Google Scholar]
- Jung, T.; Orlikowski, L.; Henricot, B.; Abad-Campos, P.; Aday, A.G.; Casal, O.A.; Bakonyi, J.; Cacciola, S.O.; Cech, T.; Chavarriaga, D.; et al. Widespread Phytophthora infestations in European nurseries put forest, semi-natural and horticultural ecosystems at high risk of Phytophthora diseases. For. Pathol. 2015, 46, 134–163. [Google Scholar] [CrossRef]
- Redekar, N.R.; Eberhart, J.L.; Parke, J.L. Diversity of Phytophthora, Pythium, and Phytopythium Species in Recycled Irrigation Water in a Container Nursery. Phytobiomes J. 2019, 3, 31–45. [Google Scholar] [CrossRef]
- Rooney-Latham, S.; Blomquist, C.L.; Kosta, K.L.; Gou, Y.Y.; Woods, P.W. Phytophthora Species are Common on Nursery Stock Grown for Restoration and Revegetation Purposes in California. Plant Dis. 2019, 103, 448–455. [Google Scholar] [CrossRef]
- Molnar, C.; Nikolaeva, E.; Kim, S.; Olson, T.; Bily, D.; Kim, J.-E.; Kang, S. Phytophthora Diversity in Pennsylvania Nurseries and Greenhouses Inferred from Clinical Samples Collected over Four Decades. Microorganism 2020, 8, 1056. [Google Scholar] [CrossRef]
- Scott, P.; Bader, M.K.-F.; Burgess, T.; Hardy, G.; Williams, N. Global biogeography and invasion risk of the plant pathogen genus Phytophthora. Environ. Sci. Policy 2019, 101, 175–182. [Google Scholar] [CrossRef]
- Hulbert, J.M.; Agne, M.C.; Burgess, T.I.; Roets, F.; Wingfield, M.J. Urban environments provide opportunities for early detections of Phytophthora invasions. Biol. Invasions 2017, 19, 3629–3644. [Google Scholar] [CrossRef]
- Paap, T.; Burgess, T.I.; Wingfield, M.J. Urban trees: Bridge-heads for forest pest invasions and sentinels for early detection. Biol. Invasions 2017, 19, 3515–3526. [Google Scholar] [CrossRef]
- Hüberli, D.; Hardy, G.E.S.J.; White, D.; Williams, N.; Burgess, T.I. Fishing for Phytophthora from Western Australia’s waterways: A distribution and diversity survey. Australas. Plant Pathol. 2013, 42, 251–260. [Google Scholar] [CrossRef]
- Jung, T.; Jung, M.H.; Cacciola, S.O.; Cech, T.; Bakonyi, J.; Seress, D.; Mosca, S.; Schena, L.; Seddaiu, S.; Pane, A.; et al. Multiple new cryptic pathogenic Phytophthora species from Fagaceae forests in Austria, Italy and Portugal. IMA Fungus 2017, 8, 219–244. [Google Scholar] [CrossRef] [PubMed]
- Jung, T.; Jung, M.H.; Scanu, B.; Seress, D.; Kovács, M.G.; Maia, C.; Pérez-Sierra, A.; Chang, T.-T.; Chandelier, A.; Heungens, K.; et al. Six new Phytophthora species from ITS Clade 7a including two sexually functional heterothallic hybrid species detected in natural ecosystems in Taiwan. Pers. Mol. Phylogeny Evol. Fungi 2017, 38, 100–135. [Google Scholar] [CrossRef]
- Jung, T.; Pérez-Sierra, A.; Durán, A.; Jung, M.H.; Balci, Y.; Scanu, B. Canker and decline diseases caused by soil- and airborne Phytophthora species in forests and woodlands. Pers. Mol. Phylogeny Evol. Fungi 2018, 40, 182–220. [Google Scholar] [CrossRef] [PubMed]
- Jung, T.; La Spada, F.; Pane, A.; Aloi, F.; Evoli, M.; Jung, M.H.; Scanu, B.; Faedda, R.; Rizza, C.; Puglisi, I.; et al. Diversity and Distribution of Phytophthora Species in Protected Natural Areas in Sicily. Forests 2019, 10, 259. [Google Scholar] [CrossRef]
- Jung, T.; Scanu, B.; Brasier, C.M.; Webber, J.; Milenković, I.; Corcobado, T.; Tomšovský, M.; Pánek, M.; Bakonyi, J.; Maia, C.; et al. A Survey in Natural Forest Ecosystems of Vietnam Reveals High Diversity of both New and Described Phytophthora Taxa including P. ramorum. Forests 2020, 11, 93. [Google Scholar] [CrossRef]
- Burgess, T.I.; White, D.; McDougall, K.M.; Garnas, J.; Dunstan, W.A.; Català, S.; Carnegie, A.J.; Worboys, S.; Cahill, D.; Vettraino, A.-M.; et al. Distribution and diversity of Phytophthora across Australia. Pac. Conserv. Biol. 2017, 23, 150–162. [Google Scholar] [CrossRef]
- Bourret, B.T.; Aram, K.; Mehl, K.H.; Rizzo, M.D.; Rooney-Latham, S.; Swiecki, J.T.; Frankel, J.S. Ten new provisional species of Phytophthora and Nothophytophthora from California. In Proceedings of the Seventh Sudden Oak Death Science and Management Symposium: Healthy Plants in a World with Phytophthora, San Francisco, CA, USA, 25–27 June 2019; General Technical Report PSW-GTR-268. US Forest Service: Albany, CA, USA, 2020; Volume 268, pp. 46–47. [Google Scholar]
- Brasier, C.M. The biosecurity threat to the UK and global environment from international trade in plants. Plant Pathol. 2008, 57, 792–808. [Google Scholar] [CrossRef]
- Moralejo, E.; Pérez-Sierra, A.M.; Álvarez, L.A.; Belbahri, L.; Lefort, F.; Descals, E. Multiple alien Phytophthora taxa discovered on diseased ornamental plants in Spain. Plant Pathol. 2009, 58, 100–110. [Google Scholar] [CrossRef]
- O’Hanlon, R.; Choiseul, J.; Corrigan, M.; Catarame, T.; Destefanis, M. Diversity and detections of Phytophthora species from trade and non-trade environments in Ireland. EPPO Bull. 2016, 46, 594–602. [Google Scholar] [CrossRef]
- Frankel, S.J.; Conforti, C.; Hillman, J.; Ingolia, M.; Shor, A.; Benner, D.; Alexander, J.M.; Bernhardt, E.; Swiecki, T.J. Phytophthora Introductions in Restoration Areas: Responding to Protect California Native Flora from Human-Assisted Pathogen Spread. Forests 2020, 11, 1291. [Google Scholar] [CrossRef]
- Giordana, G.; Kitzberger, T.; La Manna, L. Anthropogenic Factors Control the Distribution of a Southern Conifer Phytophthora Disease in a Peri-Urban Area of Northern Patagonia, Argentina. Forests 2020, 11, 1183. [Google Scholar] [CrossRef]
- Cooke, D.; Drenth, A.; Duncan, J.; Wagels, G.; Brasier, C. A Molecular Phylogeny of Phytophthora and Related Oomycetes. Fungal Genet. Biol. 2000, 30, 17–32. [Google Scholar] [CrossRef]
- Seifert, K.A. Progress towards DNA barcoding of fungi. Mol. Ecol. Resour. 2009, 9, 83–89. [Google Scholar] [CrossRef] [PubMed]
- Grünwald, N.J.; Martin, F.N.; Larsen, M.M.; Sullivan, C.M.; Press, C.M.; Coffey, M.D.; Hansen, E.M.; Parke, J.L. Phytophthora-ID.org: A Sequence-Based Phytophthora Identification Tool. Plant Dis. 2011, 95, 337–342. [Google Scholar] [CrossRef]
- Yang, X.; Hong, C. Differential Usefulness of Nine Commonly Used Genetic Markers for Identifying Phytophthora Species. Front. Microbiol. 2018, 9, 2334. [Google Scholar] [CrossRef]
- Català, S.; Pérez-Sierra, A.; Abad-Campos, P. The Use of Genus-Specific Amplicon Pyrosequencing to Assess Phytophthora Species Diversity Using eDNA from Soil and Water in Northern Spain. PLoS ONE 2015, 10, e0119311. [Google Scholar] [CrossRef]
- Vannini, A.; Bruni, N.; Tomassini, A.; Franceschini, S.; Vettraino, A.M. Pyrosequencing of environmental soil samples reveals biodiversity of the Phytophthora resident community in chestnut forests. FEMS Microbiol. Ecol. 2013, 85, 433–442. [Google Scholar] [CrossRef] [PubMed]
- Prigigallo, M.I.; Abdelfattah, A.; Cacciola, S.O.; Faedda, R.; Sanzani, S.M.; Cooke, D.E.L.; Schena, L. Metabarcoding Analysis of Phytophthora Diversity Using Genus-Specific Primers and 454 Pyrosequencing. Phytopathology 2016, 106, 305–313. [Google Scholar] [CrossRef] [PubMed]
- Riddell, C.E.; Frederickson-Matika, D.; Armstrong, A.C.; Elliot, M.; Forster, J.; Hedley, P.E.; Morris, J.; Thorpe, P.; El Cooke, D.; Pritchard, L.; et al. Metabarcoding reveals a high diversity of woody host-associated Phytophthora spp. in soils at public gardens and amenity woodlands in Britain. PeerJ 2019, 7, e6931. [Google Scholar] [CrossRef]
- Kroon, L.; Bakker, F.; Bosch, G.V.D.; Bonants, P.; Flier, W. Phylogenetic analysis of Phytophthora species based on mitochondrial and nuclear DNA sequences. Fungal Genet. Biol. 2004, 41, 766–782. [Google Scholar] [CrossRef] [PubMed]
- Blair, J.E.; Coffey, M.D.; Park, S.-Y.; Geiser, D.M.; Kang, S. A multi-locus phylogeny for Phytophthora utilizing markers derived from complete genome sequences. Fungal Genet. Biol. 2008, 45, 266–277. [Google Scholar] [CrossRef]
- Martin, F.N.; Blair, J.E.; Coffey, M.D. A combined mitochondrial and nuclear multilocus phylogeny of the genus Phytophthora. Fungal Genet. Biol. 2014, 66, 19–32. [Google Scholar] [CrossRef] [PubMed]
- Robideau, G.P.; De Cock, A.W.A.M.; Coffey, M.D.; Voglmayr, H.; Brouwer, H.; Bala, K.; Chitty, D.W.; Désaulniers, N.; Eggertson, Q.A.; Gachon, C.M.M.; et al. DNA barcoding of oomycetes with cytochrome c oxidase subunit I and internal transcribed spacer. Mol. Ecol. Resour. 2011, 11, 1002–1011. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.-J.; Beakes, G.; Glockling, S.; Kruse, J.; Nam, B.; Nigrelli, L.; Ploch, S.; Shin, H.-D.; Shivas, R.G.; Telle, S.; et al. Towards a universal barcode of oomycetes—A comparison of the cox1 and cox2 loci. Mol. Ecol. Resour. 2015, 15, 1275–1288. [Google Scholar] [CrossRef]
- Brasier, C.M.; Kirk, S.A. Comparative aggressiveness of standard and variant hybrid alder phytophthoras, Phytophthora cambivora and other Phytophthora species on bark of Alnus, Quercus and other woody hosts. Plant Pathol. 2001, 50, 218–229. [Google Scholar] [CrossRef]
- Man, W.A.; de Cock, A.W.; Summerbell, R.C. Natural hybrids of resident and introduced Phytophthora species proliferating on multiple new hosts. Eur. J. Plant Pathol. 2006, 117, 25–33. [Google Scholar] [CrossRef]
- Nagel, J.H.; Gryzenhout, M.; Slippers, B.; Wingfield, M.J.; Hardy, G.E.S.J.; Stukely, M.J.; Burgess, T.I. Characterization of Phytophthora hybrids from ITS clade 6 associated with riparian ecosystems in South Africa and Australia. Fungal Biol. 2013, 117, 329–347. [Google Scholar] [CrossRef]
- Jung, T.; Blaschke, H.; Neumann, P. Isolation, identification and pathogenicity of Phytophthora species from declining oak stands. For. Pathol. 1996, 26, 253–272. [Google Scholar] [CrossRef]
- Jung, T.; Blaschke, H.; Osswald, W. Involvement of soilborne Phytophthora species in Central European oak decline and the effect of site factors on the disease. Plant Pathol. 2000, 49, 706–718. [Google Scholar] [CrossRef]
- Scibetta, S.; Schena, L.; Chimento, A.; Cacciola, S.O.; Cooke, D.E. A molecular method to assess Phytophthora diversity in environmental samples. J. Microbiol. Methods 2012, 88, 356–368. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M., Gelfand, D., Shinsky, J., White, T., Eds.; Academic Press: New York, NY, USA, 1990; pp. 315–322. [Google Scholar]
- Illumina. 16S Metagenomic Sequencing Library Preparation; Illumina: San Diego, CA, USA, 2013; p. 28. [Google Scholar]
- Krueger, F. Trim Galore!: A Wrapper Tool Around Cutadapt and FastQC to Consistently Apply Quality and Adapter Trimming to FastQ Files; Babraham Institute: Babraham, UK, 2015. [Google Scholar]
- Babraham Bioinformatics. FastQC v. 0.11.2. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 10 January 2020).
- Ewels, P.; Magnusson, M.; Lundin, S.; Käller, M. MultiQC: Summarize analysis results for multiple tools and samples in a single report. Bioinformatics 2016, 32, 3047–3048. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef]
- Callahan, B.J.; Mcmurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef]
- Rognes, T.; Flouri, T.; Nichols, B.; Quince, C.; Mahé, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ 2016, 4, e2584. [Google Scholar] [CrossRef] [PubMed]
- Ratnasingham, S.; Hebert, P.D.N. BARCODING: Bold: The Barcode of Life Data System. Mol. Ecol. Notes 2007, 7, 355–364. Available online: http://www.barcodinglife.org (accessed on 14 February 2021). [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Capella-Gutiérrez, S.; Silla-Martínez, J.M.; Gabaldón, T. trimAl: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 2009, 25, 1972–1973. [Google Scholar] [CrossRef]
- Darriba, D.; Posada, D.; Kozlov, A.M.; Stamatakis, A.; Morel, B.; Flouri, T. ModelTest-NG: A New and Scalable Tool for the Selection of DNA and Protein Evolutionary Models. Mol. Biol. Evol. 2020, 37, 291–294. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.-T.; Schmidt, H.A.; Von Haeseler, A.; Minh, B.Q. IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef] [PubMed]
- Hoang, D.T.; Chernomor, O.; Von Haeseler, A.; Minh, B.Q.; Vinh, L.S. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol. Biol. Evol. 2018, 35, 518–522. [Google Scholar] [CrossRef] [PubMed]
- Anderson, M.J. A new method for non-parametric multivariate analysis of variance. Austral Ecol. 2001, 26, 32–46. [Google Scholar] [CrossRef]
- Legendre, P.; Gallagher, E.D. Ecologically meaningful transformations for ordination of species data. Oecologia 2001, 129, 271–280. [Google Scholar] [CrossRef]
- Anderson, M.J. Distance-Based Tests for Homogeneity of Multivariate Dispersions. Biometrics 2005, 62, 245–253. [Google Scholar] [CrossRef]
- Oksanen, J.; Blanchet, F.G.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, R.B.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Vegan: Community Ecology Package. R Package Version 2.5-2. 2018. Available online: https://cran.r-project.org/web/packages/vegan/index.html (accessed on 14 February 2021).
- Cooke, D.E.L. Threats Posed by Phytophthora to Scottish Plant Health; A Review of Previous Findings, Pathways of Entry and Further Spread and the Status of Diagnostic Techniques. RESAS Phytophthora Risk Review. Scottish Government Report. Available online: https://cran.r-project.org/web/packages/vegan/index.html (accessed on 28 November 2020).
- Farr, D.F.; Rossman, A.Y. Fungal Databases, US National Fungus Collections, ARS, USDA. Available online: https://nt.ars-grin.gov/fungaldatabases/ (accessed on 14 February 2021).
- Roesch, L.F.W.; Fulthorpe, R.R.; Riva, A.; Casella, G.; Hadwin, A.K.M.; Kent, A.D.; Daroub, S.H.; Camargo, F.A.O.; Farmerie, W.G.; Triplett, E.W. Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J. 2007, 1, 283–290. [Google Scholar] [CrossRef] [PubMed]
- Jumpponen, A.; Jones, K.L. Massively parallel 454 sequencing indicates hyperdiverse fungal communities in temperate Quercus macrocarpa phyllosphere. New Phytol. 2009, 184, 438–448. [Google Scholar] [CrossRef] [PubMed]
- Vettraino, A.; Bonants, P.; Tomassini, A.; Bruni, N.; Vannini, A. Pyrosequencing as a tool for the detection of Phytophthora species: Error rate and risk of false Molecular Operational Taxonomic Units. Lett. Appl. Microbiol. 2012, 55, 390–396. [Google Scholar] [CrossRef]
- Jung, T.; Scanu, B.; Bakonyi, J.; Seress, D.; Kovács, G.; Durán, A.; Von Stowasser, E.S.; Schena, L.; Mosca, S.; Thu, P.; et al. Nothophytophthora gen. nov., a new sister genus of Phytophthora from natural and semi-natural ecosystems. Pers. Mol. Phylogeny Evol. Fungi 2017, 39, 143–174. [Google Scholar] [CrossRef] [PubMed]
- Schlenzig, A.; Purser, E.; Perez-Sierra, A. First finding of Phytophthora foliorum in the United Kingdom. New Dis. Rep. 2016, 34, 2. [Google Scholar] [CrossRef]
- Donahoo, R.; Blomquist, C.L.; Thomas, S.L.; Moulton, J.K.; Cooke, D.E.; Lamour, K.H. Phytophthora foliorum sp. nov., a new species causing leaf blight of azalea. Mycol. Res. 2006, 110, 1309–1322. [Google Scholar] [CrossRef]
- Green, S.; Hendry, S.; Macaskill, G.; Laue, B.; Steele, H. Dieback and mortality of Juniperus communis in Britain associated with Phytophthora austrocedrae. New Dis. Rep. 2012, 26, 2. [Google Scholar] [CrossRef]
- Green, S.; Elliot, M.; Armstrong, A.; Hendry, S.J. Phytophthora austrocedrae emerges as a serious threat to juniper (Juniperus communis) in Britain. Plant Pathol. 2014, 64, 456–466. [Google Scholar] [CrossRef]
- Henricot, B.; Pérez-Sierra, A.; Armstrong, A.C.; Sharp, P.M.; Green, S. Morphological and Genetic Analyses of the Invasive Forest Pathogen Phytophthora austrocedri Reveal that Two Clonal Lineages Colonized Britain and Argentina from a Common Ancestral Population. Phytopathology 2017, 107, 1532–1540. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Greslebin, A.G.; Hansen, E.M.; Sutton, W. Phytophthora austrocedrae sp. nov., a new species associated with Austrocedrus chilensis mortality in Patagonia (Argentina). Mycol. Res. 2007, 111, 308–316. [Google Scholar] [CrossRef] [PubMed]
- Werres, S.; Marwitz, R.; In’T Veld, W.A.M.; De Cock, A.W.A.M.; Bonants, P.J.M.; De Weerdt, M.; Themann, K.; Ilieva, E.; Baayen, R.P. Phytophthora ramorum sp. nov., a new pathogen on Rhododendron and Viburnum. Mycol. Res. 2001, 105, 1155–1165. [Google Scholar] [CrossRef]
- Lane, C.R.; Beales, P.A.; Hughes, K.J.D.; Griffin, R.L.; Munro, D.; Brasier, C.M.; Webber, J.F. First outbreak of Phytophthora ramorum in England, on Viburnum tinus. Plant Pathol. 2003, 52, 414. [Google Scholar] [CrossRef]
- Brasier, C.M.; Denman, S.; Rose, J.; Kirk, S.A.; Hughes, K.J.D.; Griffin, R.L.; Lane, C.R.; Inman, A.J.; Webber, J.F. First report of ramorum bleeding canker on Quercus falcata, caused by Phytophthora ramorum. Plant Pathol. 2004, 53, 804. [Google Scholar] [CrossRef]
- Denman, S.; Kirk, S.A.; Brasier, C.M.; Hughes, K.J.D.; Griffin, R.; Hobdon, E.; Webber, J.F. Foliar infection of sweet chestnut (Castanea sativa) by Phytophthora ramorum in the UK. Plant Pathol. 2005, 54, 581. [Google Scholar] [CrossRef]
- Giltrap, P.M.; Inman, A.J.; Barton, V.C.; Barnes, A.V.; Lane, C.R.; Hughes, K.J.D.; Tomlinson, J.; Dean, M.L.; Izzard, K. First report of ramorum dieback (Phytophthora ramorum) on Hamamelis virginiana in the UK. Plant Pathol. 2004, 53, 526. [Google Scholar] [CrossRef]
- Hughes, K.J.D.; Giltrap, P.M.; Barton, V.C.; Hobden, E.; Tomlinson, J.A.; Barber, P. On-site real-time PCR detection of Phytophthora ramorum causing dieback of Parrotia persica in the UK. Plant Pathol. 2006, 55, 813. [Google Scholar] [CrossRef]
- Inman, A.J.; Townend, V.C.; Barnes, A.V.; Lane, C.R.; Hughes, K.J.D.; Griffin, R.L.; Eales, S.J. First report of ramorum dieback (Phytophthora ramorum) on Pieris in England. Plant Pathol. 2003, 52, 785. [Google Scholar] [CrossRef]
- Webber, J.; Mullett, M.; Brasier, C. Dieback and mortality of plantation Japanese larch (Larix kaempferi) associated with infection by Phytophthora ramorum. New Dis. Rep. 2010, 22, 19. [Google Scholar] [CrossRef]
- Grünwald, N.; Werres, S.; Goss, E.M.; Taylor, C.R.; Fieland, V.J. Phytophthora obscura sp. nov., a new species of the novel Phytophthora subclade 8d. Plant Pathol. 2011, 61, 610–622. [Google Scholar] [CrossRef]
- Català, S.; Berbegal, M.; Pérez-Sierra, A.; Abad-Campos, P. Metabarcoding and development of new real-time specific assays reveal Phytophthora species diversity in holm oak forests in eastern Spain. Plant Pathol. 2016, 66, 115–123. [Google Scholar] [CrossRef]
- Reeser, P.W.; Sutton, W.; Hansen, E.M.; Remigi, P.; Adams, G.C. Phytophthora species in forest streams in Oregon and Alaska. Mycologia 2011, 103, 22–35. [Google Scholar] [CrossRef]
- Brasier, C.M.; Rose, J.; Gibbs, J.N. An unusual Phytophthora associated with widespread alder mortality in Britain. Plant Pathol. 1995, 44, 999–1007. [Google Scholar] [CrossRef]
- Fraser, S.; Gomez-Gallego, M.; Gardner, J.; Bulman, L.S.; Denman, S.; Williams, N.M. Impact of weather variables and season on sporulation of Phytophthora pluvialis and Phytophthora kernoviae. For. Pathol. 2020, 50, e12588. [Google Scholar] [CrossRef]
- Dick, M.A.; Dobbie, K.; Cooke, D.E.; Brasier, C.M. Phytophthora captiosa sp. nov. and P. fallax sp. nov. causing crown dieback of Eucalyptus in New Zealand. Mycol. Res. 2006, 110, 393–404. [Google Scholar] [CrossRef] [PubMed]
- Wheller, T.; Erwin, D.C.; Ribeiro, O.K. Phytophthora Diseases Worldwide. Mycologia 1998, 90, 1092. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).