Bring More Data!—A Good Advice? Removing Separation in Logistic Regression by Increasing Sample Size
Abstract
1. Introduction
2. Materials and Methods
2.1. General
| x | |||
| 0 | 1 | ||
| y | 0 | ||
| 1 | |||
2.2. Simulation Study Setup
- ML after removing separation by ISS (ML+ISS);
- FC applied to the original data; and
- FC after removing separation by ISS (FC+ISS).
3. Results
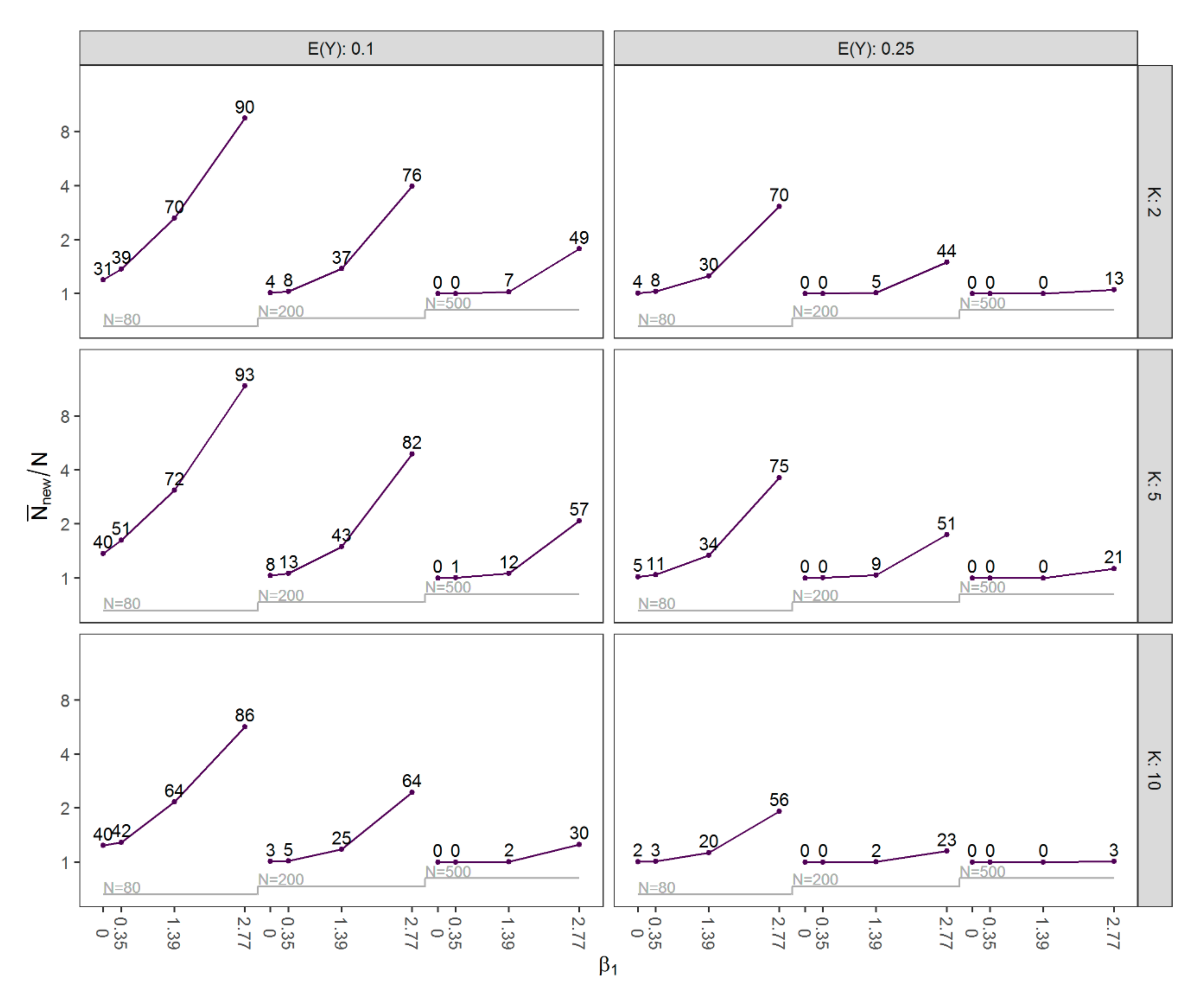
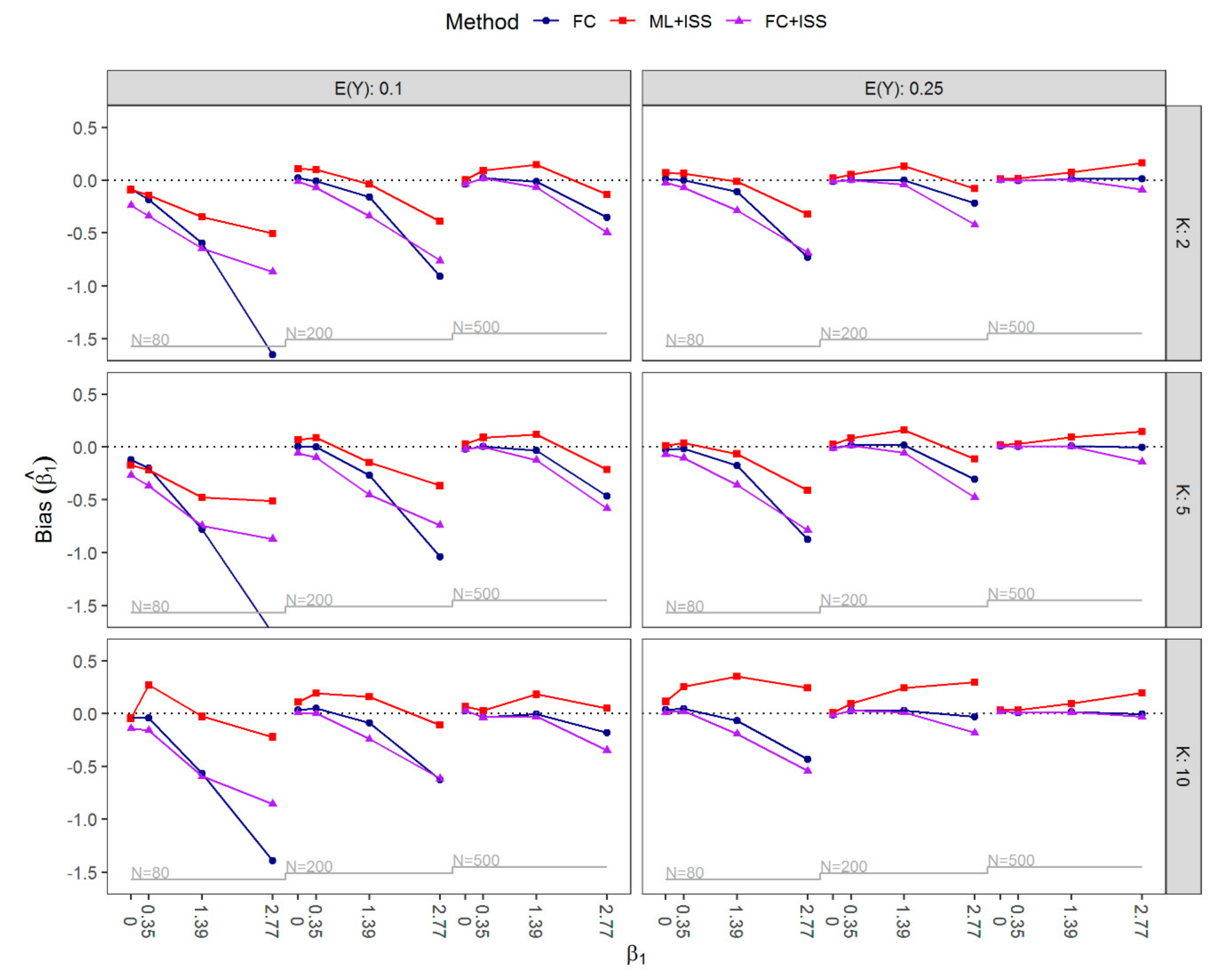
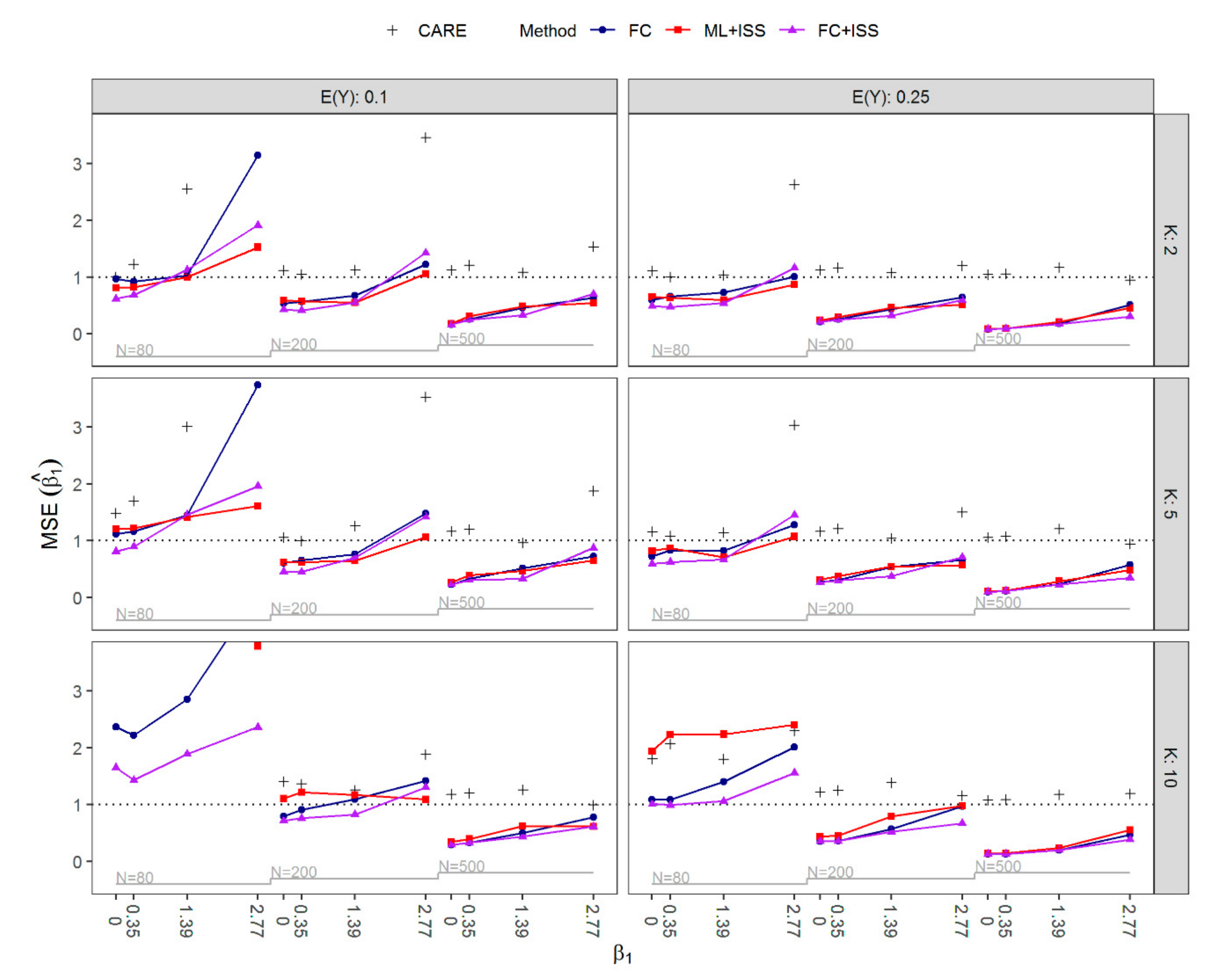
3.1. Results of Simulation Study
3.2. Examples
3.2.1. Bowel Preparation Study
3.2.2. European Passerine Birds Study
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Cox, D.R.; Snell, E.J. Analysis of binary data, 2nd ed.; Chapman and Hall/CRC: London, UK, 1989. [Google Scholar]
- Kosmidis, I. Bias in parametric estimation: Reduction and useful side-effects. WIREs Comput. Stat. 2014, 6, 185–196. [Google Scholar] [CrossRef]
- King, G.; Zeng, L. Logistic regression in rare events data. Polit. Anal. 2001, 9, 137–163. [Google Scholar] [CrossRef]
- Albert, A.; Anderson, J.A. On the existence of maximum likelihood estimates in logistic regression models. Biometrika 1984, 71, 1–10. [Google Scholar] [CrossRef]
- Heinze, G.; Schemper, M. A solution to the problem of separation in logistic regression. Stat. Med. 2002, 21, 2409–2419. [Google Scholar] [CrossRef] [PubMed]
- Mansournia, M.A.; Geroldinger, A.; Greenland, S.; Heinze, G. Separation in logistic regression: Causes, consequences, and control. Am. J. Epidemiol. 2018, 187, 864–870. [Google Scholar] [CrossRef] [PubMed]
- Courvoisier, D.S.; Combescure, C.; Agoritsas, T.; Gayet-Ageron, A.; Perneger, T.V. Performance of logistic regression modeling: Beyond the number of events per variable, the role of data structure. J. Clin. Epidemiol. 2011, 64, 993–1000. [Google Scholar] [CrossRef] [PubMed]
- van Smeden, M.; de Groot, J.A.H.; Moons, K.G.M.; Collins, G.S.; Altman, D.G.; Eijkemans, M.J.C.; Reitsma, J.B. No rationale for 1 variable per 10 events criterion for binary logistic regression analysis. BMC Med. Res. Methodol. 2016, 16, 163. [Google Scholar] [CrossRef] [PubMed]
- Firth, D. Bias reduction of maximum likelihood estimates. Biometrika 1993, 80, 27–38. [Google Scholar] [CrossRef]
- Hastie, T.; Tibshirani, R.; Friedman, J. The Elements of Statistical Learning, 2nd ed.; Springer New York Inc.: New York, NY, USA, 2001; pp. 119–128. [Google Scholar]
- Puhr, R.; Heinze, G.; Nold, M.; Lusa, L.; Geroldinger, A. Firth’s logistic regression with rare events: Accurate effect estimates and predictions? Stat. Med. 2017, 36, 2302–2317. [Google Scholar] [CrossRef] [PubMed]
- Greenland, S.; Mansournia, M.A. Penalization, bias reduction, and default priors in logistic and related categorical and survival regressions. Stat. Med. 2015, 34, 3133–3143. [Google Scholar] [CrossRef] [PubMed]
- Rousseeuw, P.J.; Christmann, A. Robustness against separation and outliers in logistic regression. Comput. Stat. Data Anal. 2003, 43, 315–332. [Google Scholar] [CrossRef]
- Morris, T.P.; White, I.R.; Crowther, M.J. Using simulation studies to evaluate statistical methods. Stat. Med. 2019, 38, 2074–2102. [Google Scholar] [CrossRef] [PubMed]
- Binder, H.; Sauerbrei, W.; Royston, P. Multivariable Model-Building with Continuous Covariates: 1. Performance Measures and Simulation Design; Technical Report FDM-Preprint 105; University of Freiburg Germany: Breisgau, Germany, 2011. [Google Scholar]
- Heinze, G.; Ploner, M.; Dunkler, D.; Southworth, H. logistf: Firth’s Bias Reduced Logistic Regression, R package version 1.22; CRAN: Vienna, Austria, 2014. [Google Scholar]
- Konis, K. SafeBinaryRegression: Safe Binary Regression, R package version 0.1-3; CRAN: Vienna, Austria, 2013. [Google Scholar]
- Kosmidis, I. brglm2: Bias Reduction in Generalized Linear Models, R package version 0.1.8; CRAN: Vienna, Austria, 2018. [Google Scholar]
- Armstrong, B.G. Optimizing Power in Allocating Resources to Exposure Assessment in an Epidemiologic Study. Am. J. Epidemiol. 1996, 144, 192–197. [Google Scholar] [CrossRef] [PubMed]
- Rücker, G.; Schwarzer, G. Presenting simulation results in a nested loop plot. BMC Med. Res. Methodol. 2014, 14, 129. [Google Scholar] [CrossRef] [PubMed]
- LoopR: A R Package for Creating Nested Loop Plots. Available online: https://github.com/matherealize/loopR (accessed on 1 October 2019).
- Waldmann, E.; Penz, D.; Majcher, B.; Zagata, J.; Šinkovec, H.; Heinze, G.; Dokladanska, A.; Szymanska, A.; Trauner, M.; Ferlitsch, A.; et al. Impact of high-volume, intermediate-volume and low-volume bowel preparation on colonoscopy quality and patient satisfaction: An observational study. United Eur. Gastroenterol. J. 2019, 7, 114–124. [Google Scholar] [CrossRef] [PubMed]
- Bandelj, P.; Blagus, R.; Trilar, T.; Vengust, M.; Vergles Rataj, A. Influence of phylogeny, migration and type of diet on the presence of intestinal parasites in the faeces of European passerine birds (Passeriformes). Wildl. Biol. 2015, 21, 227–233. [Google Scholar] [CrossRef]
- Gelman, A.; Jakulin, A.; Pittau, M.G.; Su, Y.S. A weakly informative default prior distribution for logistic and other regression models. Ann. Appl. Stat. 2008, 2, 1360–1383. [Google Scholar] [CrossRef]
- Greenland, S.; Mansournia, M.A.; Altman, D.G. Sparse data bias: A problem hiding in plain sight. BMJ 2016, 353, i1981. [Google Scholar] [CrossRef] [PubMed]
- Phillips, C.V. The economics of more research is needed. Int. J. Epidemiol. 2001, 30, 771–776. [Google Scholar] [CrossRef] [PubMed]
| Correlation of | Type | E( ) | ||
|---|---|---|---|---|
| binary | 0.8 | |||
| binary | 0.36 | |||
| binary | 0.5 | |||
| binary | 0.5 | |||
| ordinal | 1.11 | |||
| ordinal | 0.37 | |||
| continuous | 54.5 | |||
| continuous | 138.58 | |||
| continuous | 106.97 | |||
| - | continuous | 54.5 |
| Data Set | Bowel Purgative | [95% CI] | [95% CI] | |
|---|---|---|---|---|
| Preliminary version | A | 2149 | reference | |
| B | 239 | not available | 1.95 [−0.01, 6.8] | |
| C | 596 | 0.98 [−0.21, 2.18] | 0.85 [−0.13, 2.16] | |
| D | 1148 | −0.83 [−1.33, −0.34] | −0.83 [−1.32, −0.34] | |
| Final version | A | 2648 | reference | |
| B | 267 | 1.4 [−0.59, 3.39] | 1.01 [−0.62, 2.64] | |
| C | 799 | 0.83 [−0.11, 1.76] | 0.74 [−0.15, 1.64] | |
| D | 1286 | −0.83 [−1.28, −0.39] | −0.83 [−1.27, −0.39] | |
| Data Set | Diet | [95% CI] | [95% CI] | |
|---|---|---|---|---|
| Preliminary version | Granivorous | 17 | reference | |
| Insectivorous | 274 | not available | 1.53 [−0.7, 6.43] | |
| Omnivorous | 75 | not available | 2.17 [−0.02, 7.06] | |
| Final (ISS)version | Granivorous | 32 | reference | |
| Insectivorous | 276 | 0.75 [−0.82, 2.33] | 0.57 [−0.73, 2.26] | |
| Omnivorous | 77 | 1.42 [−0.15, 2.98] | 1.24 [−0.05, 2.91] | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Šinkovec, H.; Geroldinger, A.; Heinze, G. Bring More Data!—A Good Advice? Removing Separation in Logistic Regression by Increasing Sample Size. Int. J. Environ. Res. Public Health 2019, 16, 4658. https://doi.org/10.3390/ijerph16234658
Šinkovec H, Geroldinger A, Heinze G. Bring More Data!—A Good Advice? Removing Separation in Logistic Regression by Increasing Sample Size. International Journal of Environmental Research and Public Health. 2019; 16(23):4658. https://doi.org/10.3390/ijerph16234658
Chicago/Turabian StyleŠinkovec, Hana, Angelika Geroldinger, and Georg Heinze. 2019. "Bring More Data!—A Good Advice? Removing Separation in Logistic Regression by Increasing Sample Size" International Journal of Environmental Research and Public Health 16, no. 23: 4658. https://doi.org/10.3390/ijerph16234658
APA StyleŠinkovec, H., Geroldinger, A., & Heinze, G. (2019). Bring More Data!—A Good Advice? Removing Separation in Logistic Regression by Increasing Sample Size. International Journal of Environmental Research and Public Health, 16(23), 4658. https://doi.org/10.3390/ijerph16234658






