A New Method of Identifying Characteristic Points in the Impedance Cardiography Signal Based on Empirical Mode Decomposition
Abstract
1. Introduction
2. Methods
2.1. Impedance Cardiography
2.2. Empirical Mode Decomposition (EMD)
2.2.1. Standard EMD
- 1.
- First, the maxima and minima of the sequence are identified.
- 2.
- Next, the upper and lower envelopes over found extremes are constructed by cubic spline line interpolation.
- 3.
- The mean value of the upper and lower envelopes is estimated. This average value is subtracted from the original series, , and the resulting signal is treated as the potential candidate to be the first IMF. Each IMF must fulfill two conditions:
- The number of extrema (maxima and minima) and the number of zero crossings must be equal to or differ at most by one;
- The average value of the upper and lower envelopes defined by local maxima and minima must be zero.
- 4.
- If the conditions are not fulfilled, the procedure is repeated starting from step 1, this time with as an input signal.
- 5.
- After the identification of the first IMF, this component is subtracted from the input series. Then, the obtained residuum must meet the following stopping criterion:
- The residuum signal has only one extremum (minimum or maximum) or is represented by a constant/monotonic function. That kind of residuum signal characterizes the trend of the time series.
- 6.
- The whole procedure of this sifting process ends when a residual signal is found. If the criterion of being residual sequences is not fulfilled yet, the procedure is repeated from step 1 with a residuum as an input series. A diagram of the main standard EMD stages is presented in Figure 3.
2.2.2. Ensemble Empirical Mode Decomposition
- 1.
- A white noise sequence is added to the time series under consideration .
- 2.
- The noisy signal is decomposed into IMFs through the standard procedure of empirical mode decomposition described in the previous subsection.
- 3.
- The first two steps (1 and 2) are repeated for different realizations of white noise.
- 4.
- EEMD-based IMFs are estimated by averaging the ensemble of IMFs:where is the -order IMF and i stands for the number of realizations.
3. Results
3.1. New Method of ICG Characteristic Point Identification
3.2. Test of the Algorithm
3.2.1. The Database
3.2.2. Algorithm vs. Expert—Statistical Analysis of the Emerging Differences
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ECG | Electrocardiogram |
| ICG | Impedance Cardiogram |
| EMD | Empirical Mode Decomposition |
| EEMD | Ensemble Empirical Mode Decomposition |
| CEEMD | Complete Mode Empirical Decomposition |
| IMF | Intrinsic Mode Function |
| SV | Stroke Volume |
| LVET | Left Ventricular Ejection Time |
| SNR | Signal-to-Noise Ratio |
| HR | Heart Rate |
| PEP | Pre-Ejection Period |
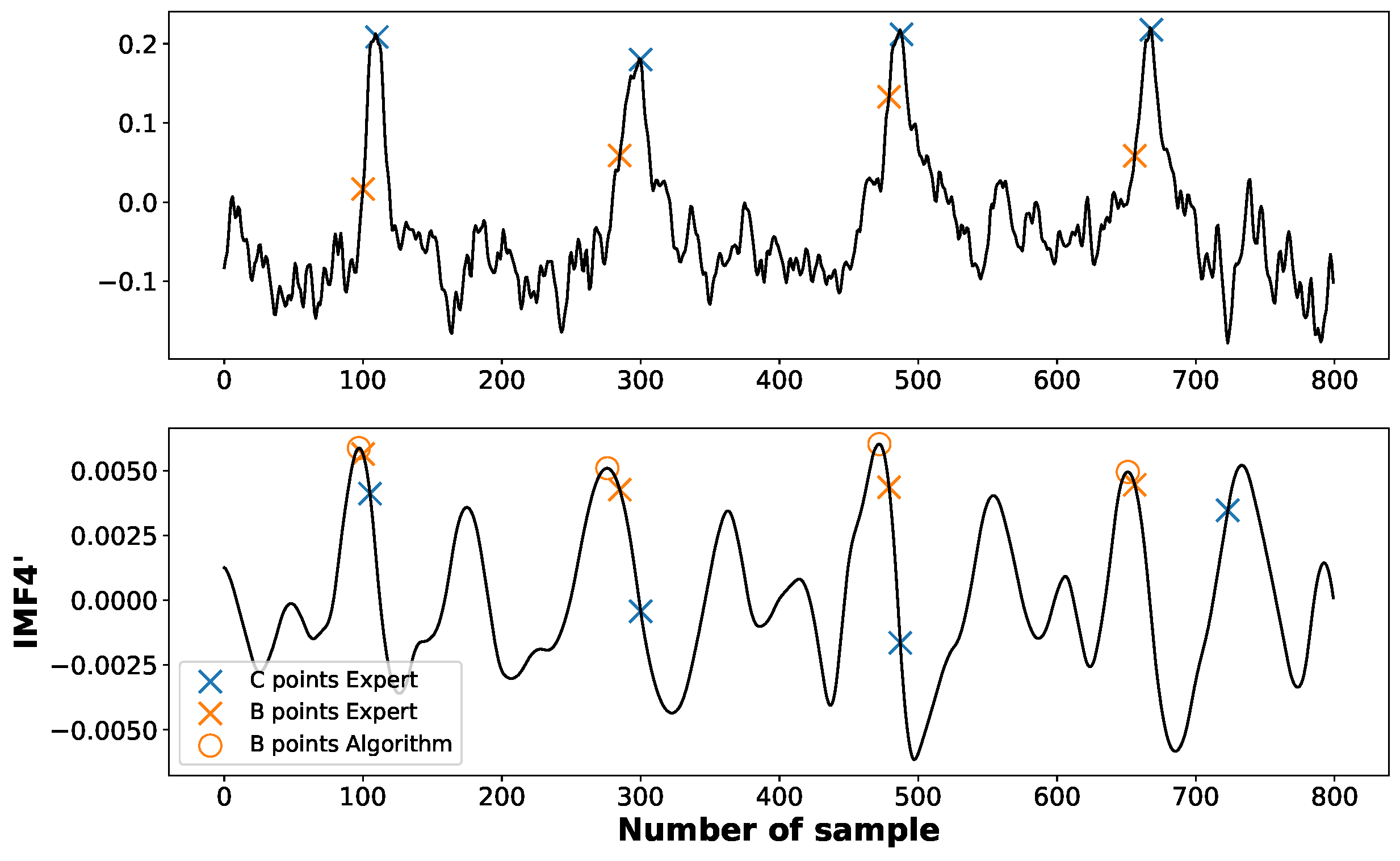
Appendix A. Discrepancies between an Expert and the Algorithm




References
- Miller, J.; Horvath, S. Impedance cardiography. Psychophysiology 1978, 15, 80–91. [Google Scholar] [CrossRef]
- Siedlecka, J.; Siedlecki, P.; Bortkiewicz, A. Impedance cardiography-Old method, new opportunities. Part I. Clinical applications. Int. J. Occup. Med. Environ. Health 2015, 28, 27–33. [Google Scholar] [CrossRef]
- Summers, R.L.; Shoemaker, W.C.; Peacock, W.F.; Ander, D.S.; Coleman, T.G. Bench to bedside: Electrophysiologic and clinical principles of noninvasive hemodynamic monitoring using impedance cardiography. Acad. Emerg. Med. 2003, 10, 669–680. [Google Scholar] [CrossRef]
- Woltjer, H.; Bogaard, H.; De Vries, P. The technique of impedance cardiography: A review. Eur. Heart J. 1997, 18, 1396–1403. [Google Scholar] [CrossRef]
- Staelens, A.; Tomsin, K.; Grieten, L.; Oben, J.; Mesens, T.; Spaanderman, M.; Jacquemyn, Y.; Gyselaers, W. Non-invasive assessment of gestational hemodynamics: Benefits and limitations of impedance cardiography versus other techniques. Expert Rev. Med. Devices 2013, 10, 765–779. [Google Scholar] [CrossRef]
- Nabian, M.; Yin, Y.; Wormwood, J.; Quigley, K.S.; Barrett, L.F.; Ostadabbas, S. An open-source feature extraction tool for the analysis of peripheral physiological data. IEEE J. Transl. Eng. Health Med. 2018, 6, 1–11. [Google Scholar] [CrossRef]
- Benouar, S.; Hafid, A.; Attari, M.; Kedir-Talha, M.; Seoane, F. Systematic variability in ICG recordings results in ICG complex subtypes–steps towards the enhancement of ICG characterization. J. Electr. Bioimpedance 2018, 9, 72–82. [Google Scholar] [CrossRef]
- Chabchoub, S.; Mansouri, S.; Ben Salah, R. Signal processing techniques applied to impedance cardiography ICG signals—A review. J. Med. Eng. Technol. 2022, 46, 243–260. [Google Scholar] [CrossRef]
- Naidu, S.; Pandey, P.C.; Pandey, V.K. Automatic detection of characteristic points in impedance cardiogram. In Proceedings of the 2011 Computing in Cardiology, Hangzhou, China, 18–21 September 2011; IEEE: New York, NY, USA, 2011; pp. 497–500. [Google Scholar]
- Naidu, S.; Bagal, U.R.; Pandey, P.C.; Hardas, S.; Khambete, N.D. Detection of characteristic points of impedance cardiogram and validation using Doppler echocardiography. In Proceedings of the 2014 Annual IEEE India Conference (INDICON), Pune, India, 11–13 December 2014; IEEE: New York, NY, USA, 2014; pp. 1–6. [Google Scholar]
- Bagal, U.R.; Pandey, P.C.; Naidu, S.; Hardas, S. Detection of opening and closing of the aortic valve using impedance cardiography and its validation by echocardiography. Biomed. Phys. Eng. Express 2017, 4, 015012. [Google Scholar] [CrossRef]
- Ono, T.; Miyamura, M.; Yasuda, Y.; Ito, T.; Saito, T.; Ishiguro, T.; Yoshizawa, M.; Yambe, T. Beat-to-beat evaluation of systolic time intervals during bicycle exercise using impedance cardiography. Tohoku J. Exp. Med. 2004, 203, 17–29. [Google Scholar] [CrossRef]
- Carvalho, P.; Paiva, R.P.; Henriques, J.; Antunes, M.; Quintal, I.; Muehlsteff, J. Robust Characteristic Points for ICG-Definition and Comparative Analysis. In Proceedings of the BIOSIGNALS, Rome, Italy, 26–29 January 2011; pp. 161–168. [Google Scholar]
- Árbol, J.R.; Perakakis, P.; Garrido, A.; Mata, J.L.; Fernández-Santaella, M.C.; Vila, J. Mathematical detection of aortic valve opening (B point) in impedance cardiography: A comparison of three popular algorithms. Psychophysiology 2017, 54, 350–357. [Google Scholar] [CrossRef] [PubMed]
- Forouzanfar, M.; Baker, F.C.; de Zambotti, M.; McCall, C.; Giovangrandi, L.; Kovacs, G.T. Toward a better noninvasive assessment of preejection period: A novel automatic algorithm for B-point detection and correction on thoracic impedance cardiogram. Psychophysiology 2018, 55, e13072. [Google Scholar] [CrossRef] [PubMed]
- Forouzanfar, M.; Baker, F.C.; Colrain, I.M.; Goldstone, A.; de Zambotti, M. Automatic analysis of pre-ejection period during sleep using impedance cardiogram. Psychophysiology 2019, 56, e13355. [Google Scholar] [CrossRef] [PubMed]
- Lozano, D.L.; Norman, G.; Knox, D.; Wood, B.L.; Miller, B.D.; Emery, C.F.; Berntson, G.G. Where to B in dZ/dt. Psychophysiology 2007, 44, 113–119. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Sun, H.H.; Van De Water, J.M. An advanced signal processing technique for impedance cardiography. IEEE Trans. Biomed. Eng. 1995, 42, 224–230. [Google Scholar] [CrossRef]
- Shuguang, Z.; Yanhong, F.; Hailong, Z.; Min, T. Detection of impedance cardioaraphy’s characteristic points based on wavelet transform. In Proceedings of the 2005 IEEE Engineering in Medicine and Biology 27th Annual Conference, Shanghai, China, 1–4 September 2005; IEEE: New York, NY, USA, 2006; pp. 2730–2732. [Google Scholar]
- Rizzi, M.; D’Aloia, M.; Castagnolo, B. High sensitivity and noise immune method to detect impedance cardiography characteristic points using wavelet transform. J. Appl. Sci. 2009, 9, 1412–1421. [Google Scholar] [CrossRef]
- Liu, S.; Yue, K.; Yang, H.; Liu, L.; Duan, X.; Guo, T. Study on cardiac impedance signal feature point extraction. In Proceedings of the 2017 IEEE 3rd Information Technology and Mechatronics Engineering Conference (ITOEC), Chongqing, China, 3–5 October 2017; IEEE: New York, NY, USA, 2017; pp. 790–793. [Google Scholar]
- Shyu, L.Y.; Lin, Y.S.; Liu, C.P.; Hu, W.C. The detection of impedance cardiogram characteristic points using wavelet transform. Comput. Biol. Med. 2004, 34, 165–175. [Google Scholar] [CrossRef]
- Podtaev, S.; Stepanov, R.; Dumler, A.; Chugainov, S.; Tziberkin, K. Wavelet analysis of the impedance cardiogram waveforms. In Journal of Physics: Conference Series; IOP Publishing: Bristol, UK, 2012; Volume 407, p. 012003. [Google Scholar]
- Stepanov, R.; Podtaev, S.; Dumler, A.; Chugainov, S. Assessment of cardiac time intervals by wavelet transform of the impedance cardiogram. Technol. Health Care 2016, 24, S803–S809. [Google Scholar] [CrossRef]
- Kabir, M.A.; Shahnaz, C. Denoising of ECG signals based on noise reduction algorithms in EMD and wavelet domains. Biomed. Signal Process. Control 2012, 7, 481–489. [Google Scholar] [CrossRef]
- Zhang, D.; Wang, S.; Li, F.; Tian, S.; Wang, J.; Ding, X.; Gong, R. An efficient ECG denoising method based on empirical mode decomposition, sample entropy, and improved threshold function. Wirel. Commun. Mob. Comput. 2020, 2020, 8811962. [Google Scholar] [CrossRef]
- El Bouny, L.; Khalil, M.; Adib, A. ECG signal filtering based on CEEMDAN with hybrid interval thresholding and higher order statistics to select relevant modes. Multimed. Tools Appl. 2019, 78, 13067–13089. [Google Scholar] [CrossRef]
- Moghtaderi, A.; Flandrin, P.; Borgnat, P. Trend filtering via empirical mode decompositions. Comput. Stat. Data Anal. 2013, 58, 114–126. [Google Scholar] [CrossRef]
- Trybek, P.; Nowakowski, M.; Machura, L. Multifractal characteristics of external anal sphincter based on sEMG signals. Med. Eng. Phys. 2018, 55, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Maji, C.; Sengupta, P.; Batabyal, A.; Chaudhuri, H. Nonlinear and statistical analysis of ECG signals from arrhythmia affected cardiac system through the EMD process. arXiv 2020, arXiv:2002.03840. [Google Scholar]
- Riaz, F.; Hassan, A.; Rehman, S.; Niazi, I.K.; Dremstrup, K. EMD-based temporal and spectral features for the classification of EEG signals using supervised learning. IEEE Trans. Neural Syst. Rehabil. Eng. 2015, 24, 28–35. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Zhou, W.; Yuan, Q.; Geng, S.; Cai, D. Feature extraction and recognition of ictal EEG using EMD and SVM. Comput. Biol. Med. 2013, 43, 807–816. [Google Scholar] [CrossRef]
- Anas, E.M.A.; Lee, S.Y.; Hasan, M.K. Exploiting correlation of ECG with certain EMD functions for discrimination of ventricular fibrillation. Comput. Biol. Med. 2011, 41, 110–114. [Google Scholar] [CrossRef]
- Hotradat, M.; Balasundaram, K.; Masse, S.; Nair, K.; Nanthakumar, K.; Umapathy, K. Empirical mode decomposition based ECG features in classifying and tracking ventricular arrhythmias. Comput. Biol. Med. 2019, 112, 103379. [Google Scholar] [CrossRef]
- Pal, S.; Mitra, M. Empirical mode decomposition based ECG enhancement and QRS detection. Comput. Biol. Med. 2012, 42, 83–92. [Google Scholar] [CrossRef]
- Slimane, Z.E.H.; Naït-Ali, A. QRS complex detection using empirical mode decomposition. Digit. Signal Process. 2010, 20, 1221–1228. [Google Scholar] [CrossRef]
- Hossain, M.B.; Bashar, S.K.; Walkey, A.J.; McManus, D.D.; Chon, K.H. An accurate QRS complex and P wave detection in ECG signals using complete ensemble empirical mode decomposition with adaptive noise approach. IEEE Access 2019, 7, 128869–128880. [Google Scholar] [CrossRef]
- Rezgui, D.; Lachiri, Z. EMD method for automatic ECG fiducial points detection. In Proceedings of the 2016 International Image Processing, Applications and Systems (IPAS), Hammamet, Tunisia, 5–7 November 2016; IEEE: New York, NY, USA, 2016; pp. 1–5. [Google Scholar]
- Nimunkar, A.J.; Tompkins, W.J. R-peak detection and signal averaging for simulated stress ECG using EMD. In Proceedings of the 2007 29th Annual International Conference of the IEEE Engineering in Medicine and Biology Society, Lyon, France, 22–26 August 2007; IEEE: New York, NY, USA, 2007; pp. 1261–1264. [Google Scholar]
- Li, H.; Wang, X.; Chen, L.; Li, E. Denoising and R-peak detection of electrocardiogram signal based on EMD and improved approximate envelope. Circuits, Syst. Signal Process. 2014, 33, 1261–1276. [Google Scholar] [CrossRef]
- Tan, X.; Chen, X.; Hu, X.; Ren, R.; Zhou, B.; Fang, Z.; Xia, S. EMD-based electrocardiogram delineation for a wearable low-power ECG monitoring device. Can. J. Electr. Comput. Eng. 2014, 37, 212–221. [Google Scholar]
- Hasan, M.A.; Chauhan, V.S.; Krishnan, S. Beat-to-beat T-wave alternans detection using the Ensemble Empirical Mode Decomposition method. Comput. Biol. Med. 2016, 77, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Panigrahy, D.; Sahu, P. Denoising of Electrocardiogram (ECG) signal by using empirical mode decomposition (EMD) with non-local mean (NLM) technique. Biocybern. Biomed. Eng. 2018, 38, 297–312. [Google Scholar] [CrossRef]
- Zhang, M.; Wei, G. An integrated EMD adaptive threshold denoising method for reduction of noise in ECG. PLoS ONE 2020, 15, e0235330. [Google Scholar] [CrossRef] [PubMed]
- Han, G.; Lin, B.; Xu, Z. Electrocardiogram signal denoising based on empirical mode decomposition technique: An overview. J. Instrum. 2017, 12, P03010. [Google Scholar] [CrossRef]
- Jenitta, J.; Rajeswari, A. Denoising of ECG signal based on improved adaptive filter with EMD and EEMD. In Proceedings of the 2013 IEEE Conference on Information & Communication Technologies, Tamil Nadu, India, 11–12 April 2013; IEEE: New York, NY, USA, 2013; pp. 957–962. [Google Scholar]
- Singh, G.; Kaur, G.; Kumar, V. ECG denoising using adaptive selection of IMFs through EMD and EEMD. In Proceedings of the 2014 International Conference on Data Science & Engineering (ICDSE), Kochi, India, 26–28 August 2014; IEEE: New York, NY, USA, 2014; pp. 228–231. [Google Scholar]
- Tang, J.; Zou, Q.; Tang, Y.; Liu, B.; Zhang, X.k. Hilbert-Huang transform for ECG de-noising. In Proceedings of the 2007 1st International Conference on Bioinformatics and Biomedical Engineering, Wuhan, China, 6–8 July 2007; IEEE: New York, NY, USA, 2007; pp. 664–667. [Google Scholar]
- Weng, B.; Blanco-Velasco, M.; Barner, K.E. ECG denoising based on the empirical mode decomposition. In Proceedings of the 2006 International Conference of the IEEE Engineering in Medicine and Biology Society, New York, NY, USA, 30 August–3 September 2006; IEEE: New York, NY, USA, 2006; pp. 1–4. [Google Scholar]
- Chabchoub, S.; Mansouri, S.; Salah, R.B. Detection of valvular heart diseases using impedance cardiography ICG. Biocybern. Biomed. Eng. 2018, 38, 251–261. [Google Scholar] [CrossRef]
- Chen, X.; Zhou, T.; Li, D.; Zhang, C.; Jia, P.; Ma, J.; Zhang, J.; Wang, G.; Fang, J. Evaluating the clinical value of oscillatory cardiopulmonary coupling in patients with obstructive sleep apnea hypopnea syndrome by impedance cardiogram. Sleep Med. 2016, 19, 75–84. [Google Scholar] [CrossRef]
- Salah, I.B.; De la Rosa, R.; Ouni, K.; Salah, R.B. Automatic diagnosis of valvular heart diseases by impedance cardiography signal processing. Biomed. Signal Process. Control 2020, 57, 101758. [Google Scholar] [CrossRef]
- Janušauskas, A.; Marozas, V.; Lukoševičius, A. Ensemble empirical mode decomposition based feature enhancement of cardio signals. Med. Eng. Phys. 2013, 35, 1059–1069. [Google Scholar] [CrossRef] [PubMed]
- De Ridder, S.; Neyt, X.; Pattyn, N.; Migeotte, P.F. Comparison between EEMD, wavelet and FIR denoising: Influence on event detection in impedance cardiography. In Proceedings of the 2011 Annual International Conference of the IEEE Engineering in Medicine and Biology Society, Boston, MA, USA, 30 August–3 September 2011; IEEE: New York, NY, USA, 2011; pp. 806–809. [Google Scholar]
- Xie, Y.; Song, R.; Yang, D.; Yu, H.; Sun, C.; Xie, Q.; Xu, R.X. Motion robust ICG measurements using a two-step spectrum denoising method. Physiol. Meas. 2021, 42, 095004. [Google Scholar] [CrossRef]
- Cybulski, G.; Strasz, A.; Niewiadomski, W.; Gąsiorowska, A. Impedance cardiography: Recent advancements. Cardiol. J. 2012, 19, 550–556. [Google Scholar] [CrossRef] [PubMed]
- Sobotnicki, A.; Pałko, T.; Mocha, J.; Czerw, M. Evaluation of volumetric parameters of the ventricular assist device using bioimpedance method. J. Med. Informatics Technol. 2012, 19, 117–123. [Google Scholar]
- Sobotnicki, A.; Gacek, A.; Pałko, T.; Mocha, J.; Badura, G.; Czerw, M. Determination of stroke volume of the ventricular assist device using bioimpedance method. J. Med. Informatics Technol. 2013, 22, 235–241. [Google Scholar]
- Krzesinski, P.; Sobotnicki, A.; Gacek, A.; Siebert, J.; Walczak, A.; Murawski, P.; Gielerak, G. Noninvasive Bioimpedance Methods From the Viewpoint of Remote Monitoring in Heart Failure. JMIR MHealth UHealth 2021, 9, e25937. [Google Scholar] [CrossRef]
- Wtorek, J.; Polinski, A. The contribution of blood-flow-induced conductivity changes to measured impedance. IEEE Trans. Biomed. Eng. 2004, 52, 41–49. [Google Scholar] [CrossRef]
- Freeborn, T.J. A survey of fractional-order circuit models for biology and biomedicine. IEEE J. Emerg. Sel. Top. Circuits Syst. 2013, 3, 416–424. [Google Scholar] [CrossRef]
- Cybulski, G. Ambulatory impedance cardiography. In Ambulatory Impedance Cardiography; Springer: Berlin/Heidelberg, Germany, 2011; pp. 39–56. [Google Scholar]
- Kubicek, W. Development and evaluation of an impedance cardiac output system. Aerosp. Med. 1966, 37, 1208–1212. [Google Scholar]
- Van Wyk, L.; Gupta, S.; Lawrenson, J.; de Boode, W.P. Accuracy and Trending Ability of Electrical Biosensing Technology for Non-invasive Cardiac Output Monitoring in Neonates: A Systematic Qualitative Review. Front. Pediatr. 2022, 10, 851850. [Google Scholar] [CrossRef]
- Huang, N.E.; Shen, Z.; Long, S.R.; Wu, M.C.; Shih, H.H.; Zheng, Q.; Yen, N.C.; Tung, C.C.; Liu, H.H. The empirical mode decomposition and the Hilbert spectrum for nonlinear and non-stationary time series analysis. Proc. R. Soc. London Ser. A Math. Phys. Eng. Sci. 1998, 454, 903–995. [Google Scholar] [CrossRef]
- Wu, Z.; Huang, N.E. Ensemble empirical mode decomposition: A noise-assisted data analysis method. Adv. Adapt. Data Anal. 2009, 1, 1–41. [Google Scholar] [CrossRef]
- Torres, M.E.; Colominas, M.A.; Schlotthauer, G.; Flandrin, P. A complete ensemble empirical mode decomposition with adaptive noise. In Proceedings of the 2011 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), Prague, Czech Republic, 22–27 May 2011; IEEE: New York, NY, USA, 2011; pp. 4144–4147. [Google Scholar]
- Rodríguez, R.; Mexicano, A.; Bila, J.; Cervantes, S.; Ponce, R. Feature extraction of electrocardiogram signals by applying adaptive threshold and principal component analysis. J. Appl. Res. Technol. 2015, 13, 261–269. [Google Scholar] [CrossRef]
- Sahoo, S.; Biswal, P.; Das, T.; Sabut, S. Denoising of ECG signal and QRS detection using Hilbert transform and adaptive thresholding. Procedia Technol. 2016, 25, 68–75. [Google Scholar] [CrossRef]
- Pale, U.; Meier, D.; Müller, O.; Valdes, A.A.; Alonso, D.A. ReBeatICG database. Zenodo 2021. [Google Scholar] [CrossRef]
- Dell’Agnola, F.; Momeni, N.; Arza, A.; Atienza, D. Cognitive Workload Monitoring in Virtual Reality Based Rescue Missions with Drones. In Virtual, Augmented and Mixed Reality. Design and Interaction; Chen, J.Y.C., Fragomeni, G., Eds.; Springer International Publishing: Cham, Switzerland, 2020; pp. 397–409. [Google Scholar]
- Li, J.; Deng, H.; Li, P.; Yu, B. Real-time infrared gas detection based on an adaptive Savitzky–Golay algorithm. Appl. Phys. B 2015, 120, 207–216. [Google Scholar] [CrossRef]
- Acharya, D.; Rani, A.; Agarwal, S.; Singh, V. Application of adaptive Savitzky—Golay filter for EEG signal processing. Perspect. Sci. 2016, 8, 677–679. [Google Scholar] [CrossRef]
- Pale, U.; Müller, N.; Arza, A.; Atienza, D. ReBeatICG: Real-time Low-Complexity Beat-to-beat Impedance Cardiogram Delineation Algorithm. In Proceedings of the 2021 43rd Annual International Conference of the IEEE Engineering in Medicine & Biology Society (EMBC), Scotland, UK, 1–5 November 2021; pp. 5618–5624. [Google Scholar] [CrossRef]
- Association for the Advancement of Medical Instrumentation; American National Standards Institute. Testing and Reporting Performance Results of Cardiac Rhythm and ST-Segment Measurement Algorithms; ANSI/AAMI, The Association: Washington, DC, USA, 1999. [Google Scholar]
- Ben Salah, I.; Ouni, K. Denoising of the impedance cardiographie signal (ICG) for a best detection of the characteristic points. In Proceedings of the 2017 2nd International Conference on Bio-Engineering for Smart Technologies (BioSMART), Paris, France, 30 August –1 September 2017; pp. 1–4. [Google Scholar] [CrossRef]










| The Point | Description | Hemodynamic Parameters |
|---|---|---|
| B | The onset of rapid upstroke towards the C point. It represents the moment of aortic valve opening. | PEP, LVET, SV, CO |
| C | Point with the greatest amplitude in one cardiac cycle. It represents the maximum aortic flow. | HR, SV, CO, Heather Index |
| X | The minimum ICG signal in one cardiac cycle. It represents the moment of aortic valve closing. | LVET, SV, CO |
| The Hemodynamic Parameter | Definition |
|---|---|
| PEP (Pre-ejection period) | The time between electrical systole (Q point in ECG) and opening of the aortic valve (B point in ICG). |
| LVET (Left Ventricular Ejection Time) | The period of blood flow across the aortic valve. The time between B and X points in the ICG signal. |
| HR (Heart Rate) | The frequency of the heartbeat. The mean number of C points occurrences in one minute. |
| Heather Index | Cardiac contractility index defined as . |
| SV (Stroke Volume) | Amount of blood ejected from the left ventricle during one cycle |
| CO (Cardiac Output) | Amount of blood ejected from the left ventricle in one minute |
| The Characteristic Point | EMD/EEMD Components | Method of Identification |
|---|---|---|
| C point | EMD 1, 2, 3, 4 | Point of the largest amplitude occurring in the combinations of EMD lower–order components ( function). |
| B point | EEMD 4 | First maximum preceeding C point in the first derivative of the fourth component obtained from EEMD |
| X point | EEMD 3, 4, 5 | First minimum after C point found in the combinations of EEMD higher–order components |
| Parameter | Algorithm | Expert | |
|---|---|---|---|
| LVET | 0.272 | 0.252 | 0.020 |
| 1.727 | 1.500 | 0.226 | |
| LVET × | 0.465 | 0.382 | 0.083 |
| LVET (CW) | 0.272 | 0.252 | 0.020 |
| (CW) | 2.053 | 1.920 | 0.151 |
| LVET × (CW) | 0.558 | 0.468 | 0.091 |
| Point | Number of Well Predicted Points | Number of All Points | Percentage Accuracy |
|---|---|---|---|
| B | 775 | 779 | 96.92 |
| B_CW | 584 | 623 | 93.74 |
| C | 764 | 799 | 98.07 |
| C_CW | 610 | 623 | 97.91 |
| X | 690 | 799 | 88.58 |
| X_CW | 519 | 623 | 83.31 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Trybek, P.; Sobotnicka, E.; Wawrzkiewicz-Jałowiecka, A.; Machura, Ł.; Feige, D.; Sobotnicki, A.; Richter-Laskowska, M. A New Method of Identifying Characteristic Points in the Impedance Cardiography Signal Based on Empirical Mode Decomposition. Sensors 2023, 23, 675. https://doi.org/10.3390/s23020675
Trybek P, Sobotnicka E, Wawrzkiewicz-Jałowiecka A, Machura Ł, Feige D, Sobotnicki A, Richter-Laskowska M. A New Method of Identifying Characteristic Points in the Impedance Cardiography Signal Based on Empirical Mode Decomposition. Sensors. 2023; 23(2):675. https://doi.org/10.3390/s23020675
Chicago/Turabian StyleTrybek, Paulina, Ewelina Sobotnicka, Agata Wawrzkiewicz-Jałowiecka, Łukasz Machura, Daniel Feige, Aleksander Sobotnicki, and Monika Richter-Laskowska. 2023. "A New Method of Identifying Characteristic Points in the Impedance Cardiography Signal Based on Empirical Mode Decomposition" Sensors 23, no. 2: 675. https://doi.org/10.3390/s23020675
APA StyleTrybek, P., Sobotnicka, E., Wawrzkiewicz-Jałowiecka, A., Machura, Ł., Feige, D., Sobotnicki, A., & Richter-Laskowska, M. (2023). A New Method of Identifying Characteristic Points in the Impedance Cardiography Signal Based on Empirical Mode Decomposition. Sensors, 23(2), 675. https://doi.org/10.3390/s23020675








