GENDIS: Genetic Discovery of Shapelets
Abstract
1. Introduction
1.1. Background
1.2. Related Work
1.3. Our Contribution
2. Materials and Methods
2.1. Time Series Matrix and Label Vector
2.2. Shapelets and Shapelet Sets
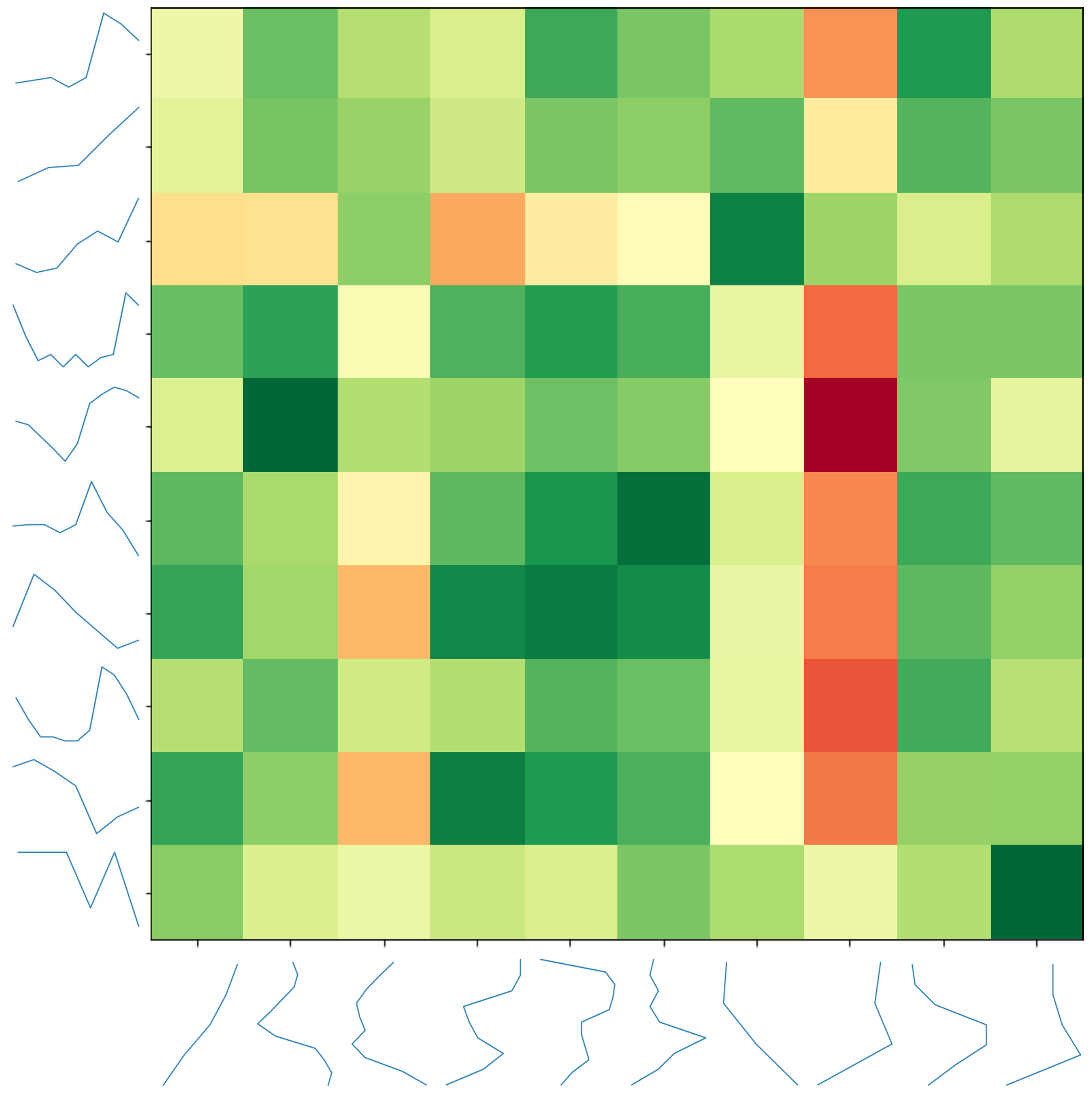
2.3. Distance Matrix Calculation
2.4. Shapelet Set Discovery
2.5. Genetic Discovery of Interpretable Shapelets
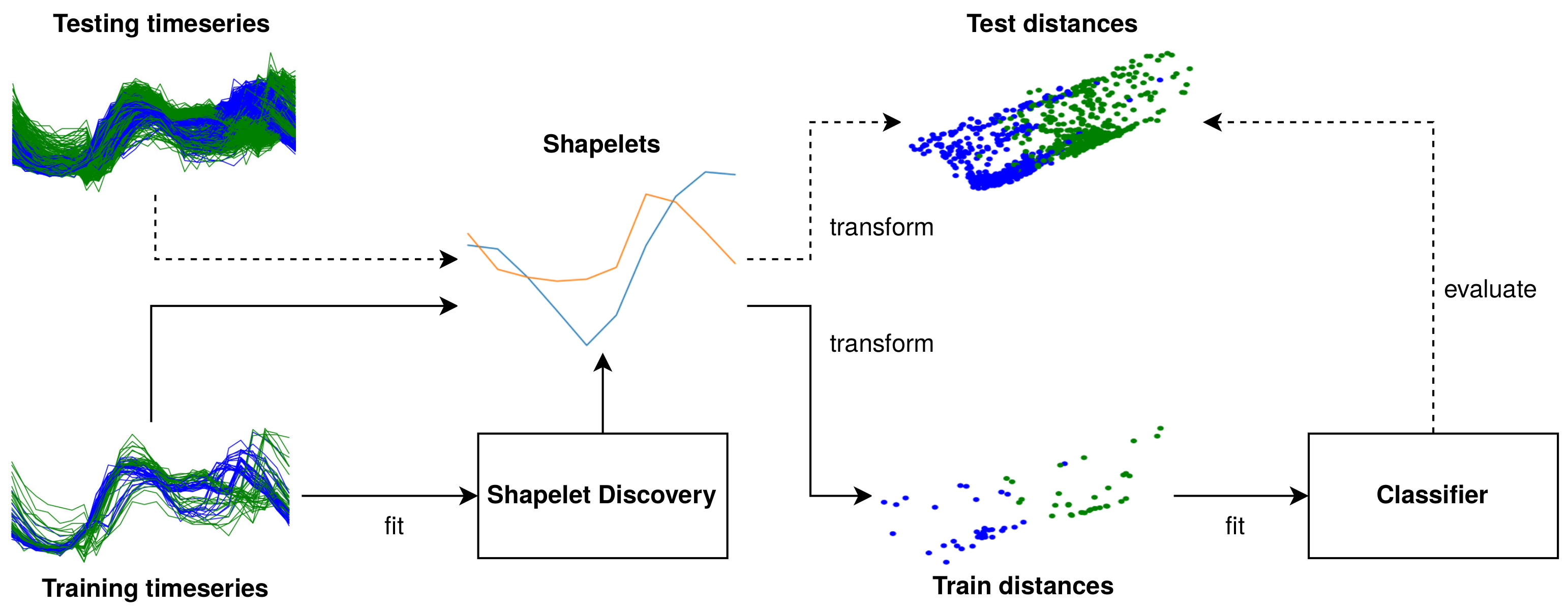
2.5.1. Conceptual Overview
2.5.2. Initialization
2.5.3. Fitness
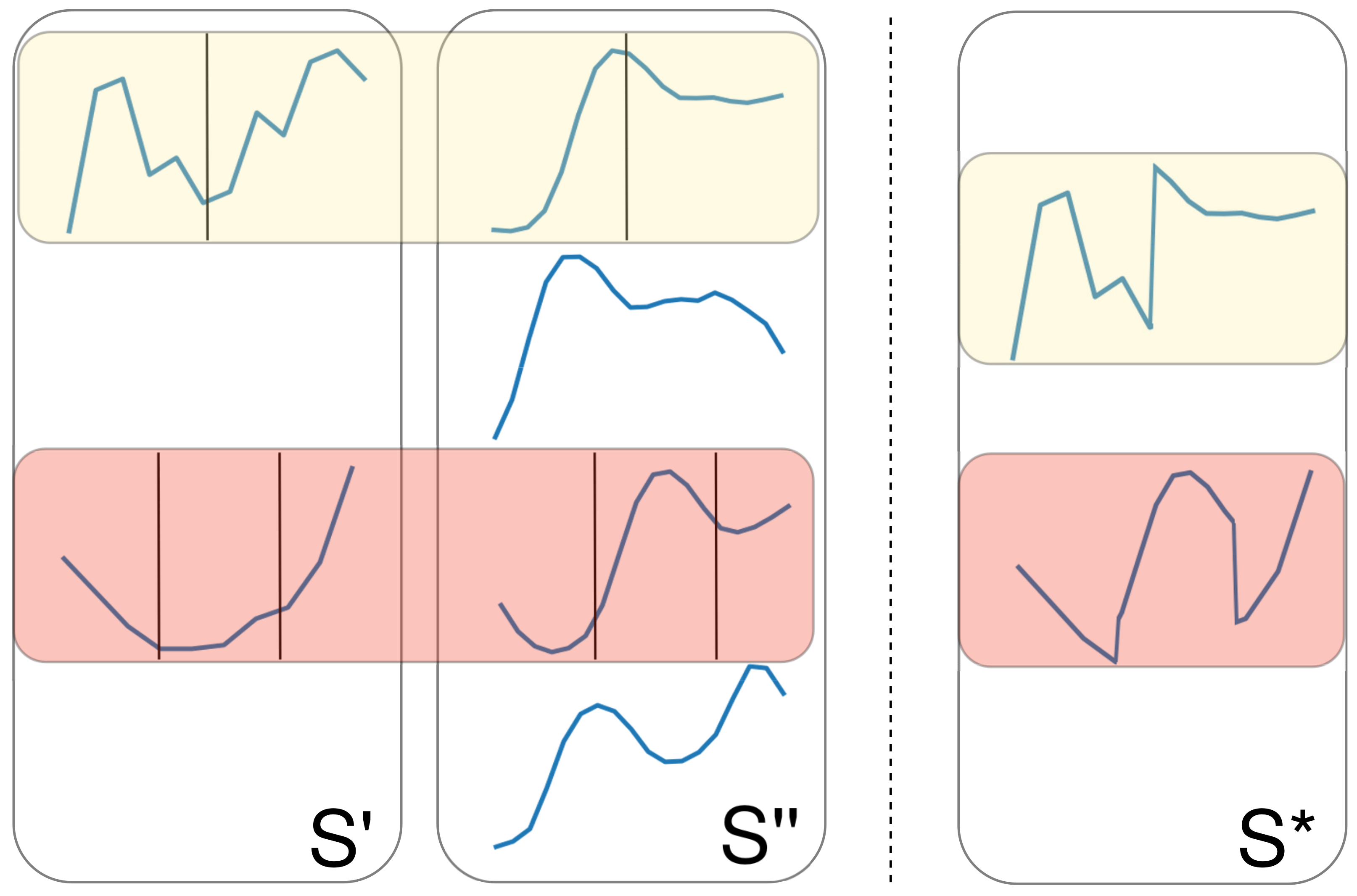
2.5.4. Crossover
2.5.5. Mutations
2.5.6. Selection, Elitism, and Early Stopping
2.5.7. List of All Hyper-Parameters
- Maximum shapelets per candidate (W): the maximum number of shapelets in a newly generated individual during initialization (default: ).
- Population size (P): the total number of candidates that are evaluated and evolved in every iteration (default: 100).
- Maximum number of generations (G): the maximum number of iterations the algorithm runs (default: 100).
- Early stopping patience (): the algorithm preemptively stops evolving when no better individual has been found for iterations (default: 10).
- Mutation probability (): the probability that a mutation operator gets applied to an individual in each iteration (default: ).
- Crossover probability (): the probability that a crossover operator is applied on a pair of individuals in each iteration (default: ).
- Maximum shapelet length (): the maximum length of the shapelets in each shapelet set (individual) (default: M).
- The operations used during the initialization, crossover, and mutation phases are configurable as well (default: all mentioned operations).
3. Results and Discussion
3.1. Efficiency of Genetic Operators
3.1.1. Datasets
3.1.2. Initialization Operators
- Initializing the individuals with K-means (Initialization 1)
- Randomly initializing the shapelet sets (Initialization 2)
- Using both initialization operations
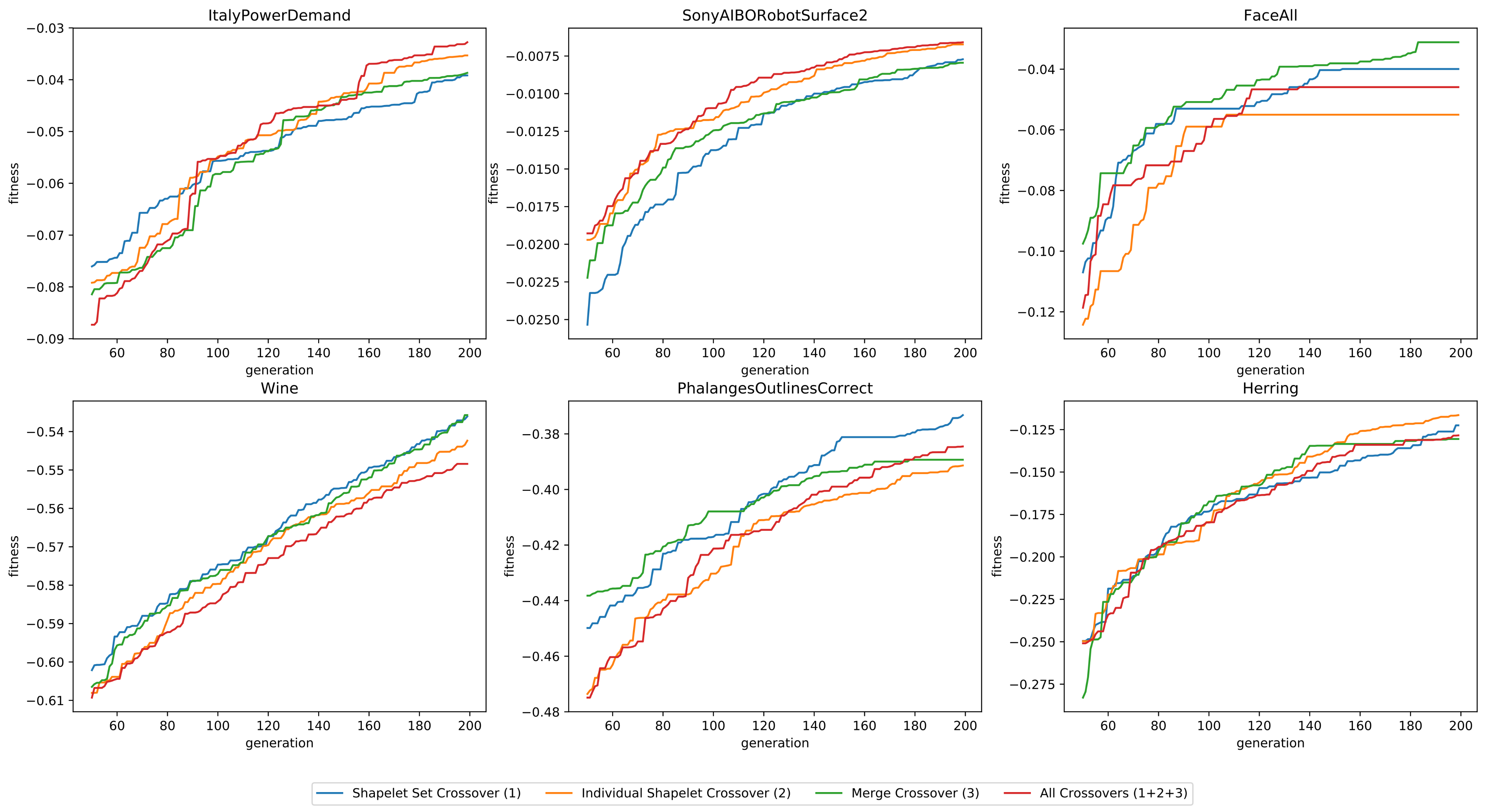
3.1.3. Crossover Operators
- Using solely point crossovers on the shapelet sets (Crossover 1)
- Using solely point crossovers on individual shapelets (Crossover 2)
- Using solely merge crossovers (Crossover 3)
- Using all three crossover operations
3.1.4. Mutation Operators
- Masking a random part of a shapelet (Mutation 1)
- Removing a random shapelet from the set (Mutation 2)
- Adding a shapelet, randomly sampled from the data, to the set (Mutation 3)
- Using all three mutation operations
3.2. Evaluating Sets of Candidates Versus Single Candidates
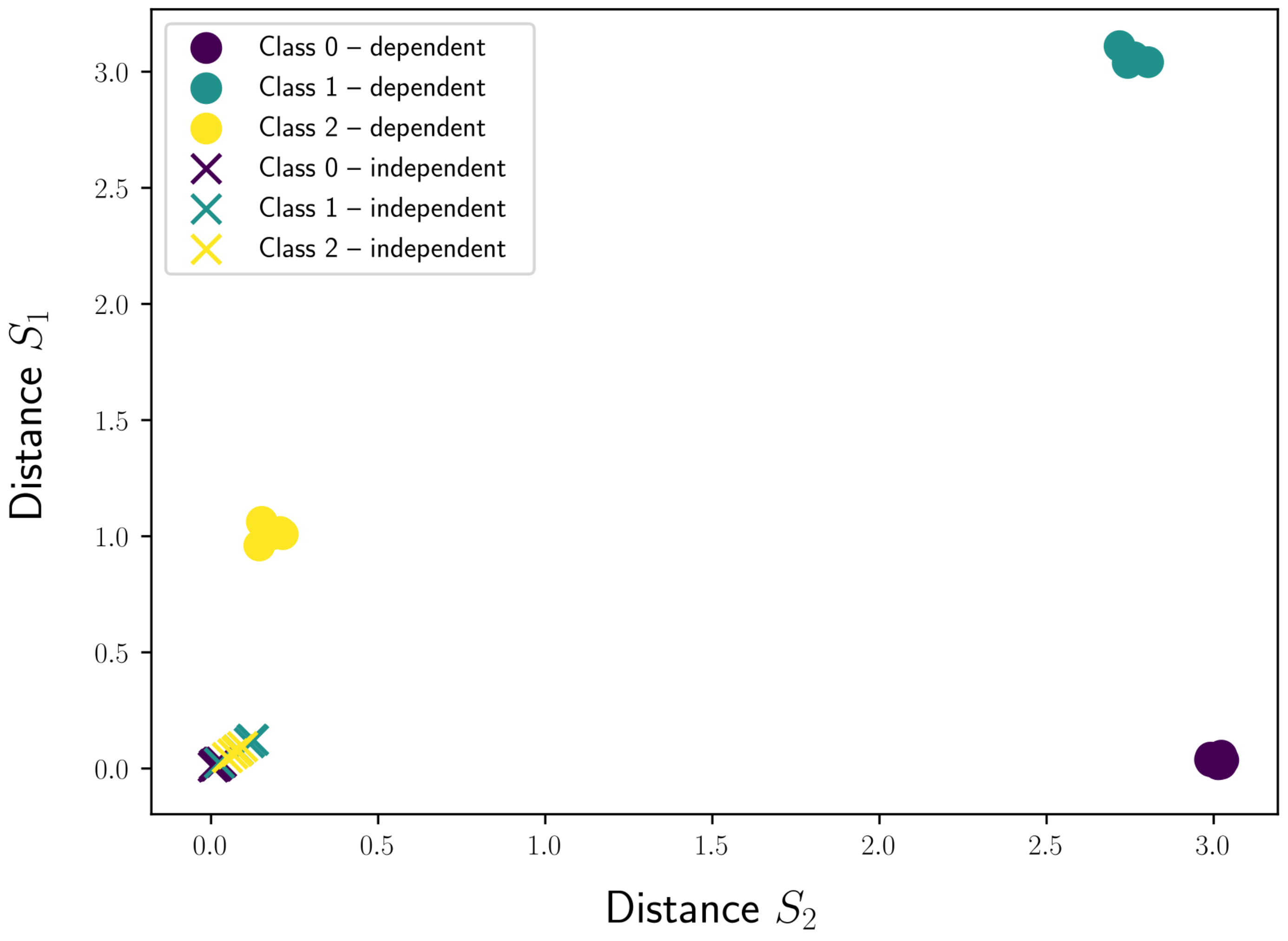
3.3. Discovering Shapelets Outside the Data
3.4. Stability
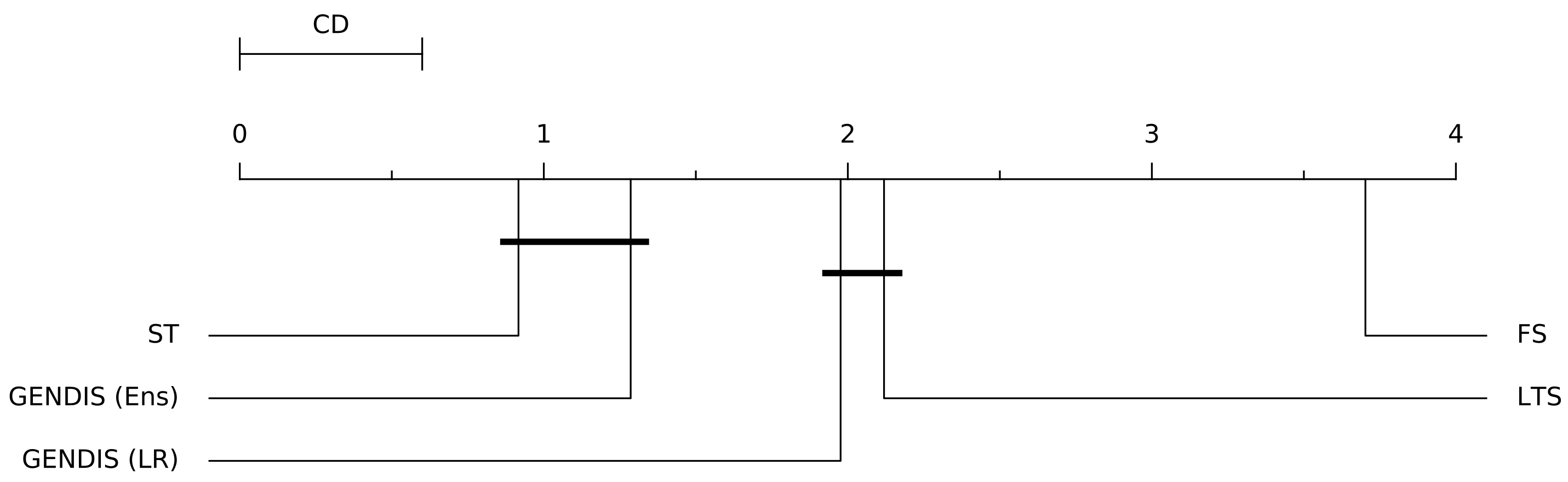
3.5. Comparing gendis to fs, st, and lts
4. Conclusions and Future Work
- evolutionary algorithms are gradient-free, allowing for an easy configuration of the optimization objective, which does not need to be differentiable
- only the maximum length of all shapelets has to be tuned, as opposed to the number of shapelets and the length of each shapelet, due to the fact that gendis evaluates entire sets of shapelets
- easy control over the runtime of the algorithm
- the possibility of discovering shapelets that do not need to be a subsequence of the input time series
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Chaovalitwongse, W.A.; Prokopyev, O.A.; Pardalos, P.M. Electroencephalogram (EEG) time series classification: Applications in epilepsy. Ann. Oper. Res. 2006, 148, 227–250. [Google Scholar] [CrossRef]
- Liu, L.; Peng, Y.; Liu, M.; Huang, Z. Sensor-based human activity recognition system with a multilayered model using time series shapelets. Knowl. Based Syst. 2015, 90, 138–152. [Google Scholar] [CrossRef]
- Li, D.; Bissyandé, T.F.; Kubler, S.; Klein, J.; Le Traon, Y. Profiling household appliance electricity usage with n-gram language modeling. In Proceedings of the 17th IEEE International Conference on Industrial Technology (ICIT), Taipei, Taiwan, 14–17 March 2016; pp. 604–609. [Google Scholar] [CrossRef]
- Grabocka, J.; Schilling, N.; Wistuba, M.; Schmidt-Thieme, L. Learning time-series shapelets. In Proceedings of the 20th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, New York, NY, USA, 24–27 August 2014; pp. 392–401. [Google Scholar] [CrossRef]
- Abanda, A.; Mori, U.; Lozano, J.A. A review on distance based time series classification. Data Min. Knowl. Discov. 2019, 33, 378–412. [Google Scholar] [CrossRef]
- Ye, L.; Keogh, E. Time series shapelets: A new primitive for data mining. In Proceedings of the 15th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, Paris, France, 28 June–1 July 2009; pp. 947–956. [Google Scholar] [CrossRef]
- Mueen, A.; Keogh, E.; Young, N. Logical-shapelets: An expressive primitive for time series classification. In Proceedings of the 17th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Diego, CA, USA, 21–24 August 2011; pp. 1154–1162. [Google Scholar] [CrossRef]
- Rakthanmanon, T.; Keogh, E. Fast shapelets: A scalable algorithm for discovering time series shapelets. In Proceedings of the 2013 SIAM International Conference on Data Mining, Austin, TX, USA, 2–4 May 2013; pp. 668–676. [Google Scholar] [CrossRef]
- Lin, J.; Keogh, E.; Lonardi, S.; Chiu, B. A symbolic representation of time series, with implications for streaming algorithms. In Proceedings of the 8th ACM SIGMOD Workshop on Research Issues in Data Mining and Knowledge Discovery, San Diego, CA, USA, 13 June 2003; pp. 2–11. [Google Scholar] [CrossRef]
- Lines, J.; Davis, L.M.; Hills, J.; Bagnall, A. A shapelet transform for time series classification. In Proceedings of the 18th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, Beijing, China, 12–16 August 2012; pp. 289–297. [Google Scholar] [CrossRef]
- Hills, J.; Lines, J.; Baranauskas, E.; Mapp, J.; Bagnall, A. Classification of time series by shapelet transformation. Data Min. Knowl. Discov. 2014, 28, 851–881. [Google Scholar] [CrossRef]
- Bostrom, A.; Bagnall, A. Binary shapelet transform for multiclass time series classification. In Transactions on Large-Scale Data-and Knowledge-Centered Systems XXXII: Special Issue on Big Data Analytics and Knowledge Discovery; Springer: Heidelberg, Germany, 2017; pp. 24–46. [Google Scholar] [CrossRef]
- Bagnall, A.; Lines, J.; Bostrom, A.; Large, J.; Keogh, E. The great time series classification bake off: A review and experimental evaluation of recent algorithmic advances. Data Min. Knowl. Discov. 2017, 31, 606–660. [Google Scholar] [CrossRef] [PubMed]
- Grabocka, J.; Wistuba, M.; Schmidt-Thieme, L. Scalable discovery of time-series shapelets. arXiv 2015, arXiv:1503.03238. [Google Scholar]
- Wistuba, M.; Grabocka, J.; Schmidt-Thieme, L. Ultra-fast shapelets for time series classification. arXiv 2015, arXiv:1503.05018. [Google Scholar]
- Hou, L.; Kwok, J.T.; Zurada, J.M. Efficient learning of timeseries shapelets. In Proceedings of the 30th AAAI Conference on Artificial Intelligence, Phoenix, AZ, USA, 12–17 February 2016; pp. 1209–1215. [Google Scholar] [CrossRef]
- Wang, Y.; Emonet, R.; Fromont, E.; Malinowski, S.; Menager, E.; Mosser, L.; Tavenard, R. Learning interpretable shapelets for time series classification through adversarial regularization. arXiv 2019, arXiv:1906.00917. [Google Scholar]
- Guilleme, M.; Malinowski, S.; Tavenard, R.; Renard, X. Localized Random Shapelets. In Proceedings of the 2nd International Workshop on Advanced Analysis and Learning on Temporal Data, Würzburg, Germany, 16–20 September 2019; pp. 85–97. [Google Scholar] [CrossRef]
- Raychaudhuri, D.S.; Grabocka, J.; Schmidt-Thieme, L. Channel masking for multivariate time series shapelets. arXiv 2017, arXiv:1711.00812. [Google Scholar]
- Medico, R.; Ruyssinck, J.; Deschrijver, D.; Dhaene, T. Learning Multivariate Shapelets with Multi-Layer Neural Networks. Advanced Course on Data Science & Machine Learning. 2018. Available online: https://biblio.ugent.be/publication/8569397/file/8569684 (accessed on 1 February 2021).
- Bostrom, A.; Bagnall, A. A shapelet transform for multivariate time series classification. arXiv 2017, arXiv:1712.06428. [Google Scholar]
- Mitchell, M. An Introduction to Genetic Algorithms; MIT Press: Cambridge, UK, 1998. [Google Scholar]
- Julstrom, B.A. Seeding the population: Improved performance in a genetic algorithm for the rectilinear steiner problem. In Proceedings of the 1994 ACM Symposium on Applied Computing, Phoenix, AZ, USA, 6–8 March 1994; pp. 222–226. [Google Scholar] [CrossRef]
- Sheblé, G.B.; Brittig, K. Refined genetic algorithm-economic dispatch example. IEEE Trans. Power Syst. 1995, 10, 117–124. [Google Scholar] [CrossRef]
- Berndt, D.J.; Clifford, J. Using dynamic time warping to find patterns in time series. In Proceedings of the 3rd International Conference on Knowledge Discovery and Data Mining, Newport Beach, CA, USA, 14–17 August 1997; Volume 10, pp. 359–370. [Google Scholar] [CrossRef]
- Large, J.; Lines, J.; Bagnall, A. The heterogeneous ensembles of standard classification algorithms (HESCA): The whole is greater than the sum of its parts. arXiv 2017, arXiv:1710.09220. [Google Scholar]
- Dietterich, T.G. Ensemble methods in machine learning. In Proceedings of the International Workshop on Multiple Classifier Systems, Cagliari, Italy, 21–23 June 2000; pp. 1–15. [Google Scholar] [CrossRef]
- Friedman, M. The use of ranks to avoid the assumption of normality implicit in the analysis of variance. J. Am. Stat. Assoc. 1937, 32, 675–701. [Google Scholar] [CrossRef]
- Holm, S. A simple sequentially rejective multiple test procedure. Scand. J. Stat. 1979, 6, 65–70. [Google Scholar] [CrossRef]
- Benavoli, A.; Corani, G.; Mangili, F. Should we really use post-hoc tests based on mean-ranks? J. Mach. Learn. Res. 2016, 17, 1–10. [Google Scholar] [CrossRef]









| Dataset | #Cls | TS_len | #Train | #Test |
|---|---|---|---|---|
| ItalyPowerDemand | 2 | 24 | 67 | 1029 |
| SonyAIBORobotSurface2 | 2 | 65 | 27 | 953 |
| FaceAll | 14 | 131 | 560 | 1690 |
| Wine | 2 | 234 | 57 | 54 |
| PhalangesOutlinesCorrect | 2 | 80 | 1800 | 858 |
| Herring | 2 | 512 | 64 | 64 |
| Dataset | #Cls | TS_len | #Train | #Test | GENDIS | ST | LTS | FS | #Shapes | |
|---|---|---|---|---|---|---|---|---|---|---|
| Ens | LR | |||||||||
| Adiac | 37 | 176 | 390 | 391 | 66.2 | 69.8 | 76.8 | 42.9 | 55.5 | 39 |
| ArrowHead | 3 | 251 | 36 | 175 | 79.4 | 82.0 | 85.1 | 84.1 | 67.5 | 39 |
| Beef | 5 | 470 | 30 | 30 | 51.5 | 58.8 | 73.6 | 69.8 | 50.2 | 41 |
| BeetleFly | 2 | 512 | 20 | 20 | 90.6 | 87.5 | 87.5 | 86.2 | 79.6 | 42 |
| BirdChicken | 2 | 512 | 20 | 20 | 90.0 | 90.5 | 92.7 | 86.4 | 86.2 | 45 |
| CBF | 3 | 128 | 30 | 900 | 99.1 | 97.6 | 98.6 | 97.7 | 92.4 | 43 |
| Car | 4 | 577 | 60 | 60 | 82.5 | 83.0 | 90.2 | 85.6 | 73.6 | 48 |
| ChlorineConcentration | 3 | 166 | 467 | 3840 | 60.9 | 57.5 | 68.2 | 58.6 | 56.6 | 30 |
| CinCECGTorso | 4 | 1639 | 40 | 1380 | 92.1 | 91.5 | 91.8 | 85.5 | 74.1 | 57 |
| Coffee | 2 | 286 | 28 | 28 | 98.9 | 98.6 | 99.5 | 99.5 | 91.7 | 44 |
| Computers | 2 | 720 | 250 | 250 | 75.4 | 72.7 | 78.5 | 65.4 | 50.0 | 38 |
| CricketX | 12 | 300 | 390 | 390 | 73.1 | 66.6 | 77.7 | 74.4 | 47.9 | 41 |
| CricketY | 12 | 300 | 390 | 390 | 69.8 | 64.3 | 76.2 | 72.6 | 50.9 | 41 |
| CricketZ | 12 | 300 | 390 | 390 | 72.6 | 65.9 | 79.8 | 75.4 | 46.6 | 40 |
| DiatomSizeReduction | 4 | 345 | 16 | 306 | 97.4 | 96.5 | 91.1 | 92.7 | 87.3 | 45 |
| DistalPhalanxOutlineAgeGroup | 3 | 80 | 400 | 139 | 84.4 | 83.2 | 81.9 | 81.0 | 74.5 | 32 |
| DistalPhalanxOutlineCorrect | 2 | 80 | 600 | 276 | 82.4 | 81.5 | 82.9 | 82.2 | 78.0 | 32 |
| DistalPhalanxTW | 6 | 80 | 400 | 139 | 76.7 | 76.0 | 69.0 | 65.9 | 62.3 | 33 |
| ECG200 | 2 | 96 | 100 | 100 | 86.3 | 86.5 | 84.0 | 87.1 | 80.6 | 30 |
| ECG5000 | 5 | 140 | 500 | 4500 | 94.0 | 93.8 | 94.3 | 94.0 | 92.2 | 34 |
| ECGFiveDays | 2 | 136 | 23 | 861 | 99.9 | 100.0 | 95.5 | 98.5 | 98.6 | 33 |
| Earthquakes | 2 | 512 | 322 | 139 | 78.4 | 73.7 | 73.7 | 74.2 | 74.7 | 44 |
| ElectricDevices | 7 | 96 | 8926 | 7711 | 83.7 | 77.6 | 89.5 | 70.9 | 26.2 | 31 |
| FaceAll | 14 | 131 | 560 | 1690 | 94.5 | 92.6 | 96.8 | 92.6 | 77.2 | 38 |
| FaceFour | 4 | 350 | 24 | 88 | 93.2 | 94.1 | 79.4 | 95.7 | 86.9 | 41 |
| FacesUCR | 14 | 131 | 200 | 2050 | 90.1 | 89.0 | 90.9 | 93.9 | 70.1 | 39 |
| FiftyWords | 50 | 270 | 450 | 455 | 72.9 | 71.8 | 71.3 | 69.4 | 51.2 | 39 |
| Fish | 7 | 463 | 175 | 175 | 87.0 | 90.5 | 97.4 | 94.0 | 74.2 | 50 |
| FordA | 2 | 500 | 3601 | 1320 | 90.8 | 90.7 | 96.5 | 89.5 | 78.5 | 37 |
| FordB | 2 | 500 | 3636 | 810 | 89.5 | 89.8 | 91.5 | 89.0 | 78.3 | 38 |
| GunPoint | 2 | 150 | 50 | 150 | 96.9 | 95.7 | 99.9 | 98.3 | 93.0 | 39 |
| Ham | 2 | 431 | 109 | 105 | 72.9 | 77.2 | 80.8 | 83.2 | 67.7 | 37 |
| HandOutlines | 2 | 2709 | 1000 | 370 | 89.7 | 91.0 | 92.4 | 83.7 | 84.1 | 41 |
| Haptics | 5 | 1092 | 155 | 308 | 45.2 | 43.9 | 51.2 | 47.8 | 35.6 | 55 |
| Herring | 2 | 512 | 64 | 64 | 59.6 | 61.8 | 65.3 | 62.8 | 55.8 | 42 |
| InlineSkate | 7 | 1882 | 100 | 550 | 43.8 | 39.3 | 39.3 | 29.9 | 25.7 | 58 |
| InsectWingbeatSound | 11 | 256 | 220 | 1980 | 57.3 | 57.5 | 61.7 | 55.0 | 48.8 | 36 |
| ItalyPowerDemand | 2 | 24 | 67 | 1029 | 95.6 | 96.0 | 95.3 | 95.2 | 90.9 | 31 |
| LargeKitchenAppliances | 3 | 720 | 375 | 375 | 91.0 | 90.4 | 93.3 | 76.5 | 41.9 | 33 |
| Lightning2 | 2 | 637 | 60 | 61 | 80.9 | 79.1 | 65.9 | 75.9 | 48.0 | 39 |
| Lightning7 | 7 | 319 | 70 | 73 | 78.2 | 76.3 | 72.4 | 76.5 | 10.1 | 39 |
| Mallat | 8 | 1024 | 55 | 2345 | 98.2 | 97.3 | 97.2 | 95.1 | 89.3 | 58 |
| Dataset | #Cls | TS_len | #Train | #Test | GENDIS | ST | LTS | FS | #Shapes | |
|---|---|---|---|---|---|---|---|---|---|---|
| Ens | LR | |||||||||
| Meat | 3 | 448 | 60 | 60 | 98.7 | 98.8 | 96.6 | 81.4 | 92.4 | 48 |
| MedicalImages | 10 | 99 | 381 | 760 | 72.4 | 68.6 | 69.1 | 70.4 | 60.9 | 37 |
| MiddlePhalanxOutlineAgeGroup | 3 | 80 | 400 | 154 | 74.4 | 73.2 | 69.4 | 67.9 | 61.3 | 30 |
| MiddlePhalanxOutlineCorrect | 2 | 80 | 600 | 291 | 80.7 | 79.6 | 81.5 | 82.2 | 71.6 | 30 |
| MiddlePhalanxTW | 6 | 80 | 399 | 154 | 62.2 | 63.1 | 57.9 | 54.0 | 51.9 | 37 |
| MoteStrain | 2 | 84 | 20 | 1252 | 86.3 | 86.6 | 88.2 | 87.6 | 79.3 | 36 |
| NonInvasiveFatalECGThorax1 | 42 | 750 | 1800 | 1965 | 84.7 | 89.4 | 94.7 | 60.0 | 71.0 | 41 |
| NonInvasiveFatalECGThorax2 | 42 | 750 | 1800 | 1965 | 87.1 | 92.3 | 95.4 | 73.9 | 75.8 | 37 |
| OSULeaf | 6 | 427 | 200 | 242 | 76.2 | 75.8 | 93.4 | 77.1 | 67.9 | 45 |
| OliveOil | 4 | 570 | 30 | 30 | 86.1 | 88.8 | 88.1 | 17.2 | 76.5 | 53 |
| PhalangesOutlinesCorrect | 2 | 80 | 1800 | 858 | 80.8 | 78.7 | 79.4 | 78.3 | 73.0 | 30 |
| Plane | 7 | 144 | 105 | 105 | 99.2 | 99.3 | 100.0 | 99.5 | 97.0 | 34 |
| ProximalPhalanxOutlineAgeGroup | 3 | 80 | 400 | 205 | 84.1 | 83.9 | 84.1 | 83.2 | 79.7 | 32 |
| ProximalPhalanxOutlineCorrect | 2 | 80 | 600 | 291 | 86.2 | 85.7 | 88.1 | 79.3 | 79.7 | 30 |
| ProximalPhalanxTW | 6 | 80 | 400 | 205 | 81.7 | 80.2 | 80.3 | 79.4 | 71.6 | 31 |
| RefrigerationDevices | 3 | 720 | 375 | 375 | 68.6 | 62.4 | 76.1 | 64.2 | 57.4 | 34 |
| ScreenType | 3 | 720 | 375 | 375 | 52.8 | 52.8 | 67.6 | 44.5 | 36.5 | 37 |
| ShapeletSim | 2 | 500 | 20 | 180 | 100.0 | 100.0 | 93.4 | 93.3 | 100.0 | 34 |
| ShapesAll | 60 | 512 | 600 | 600 | 79.6 | 79.3 | 85.4 | 76.0 | 59.8 | 44 |
| SmallKitchenAppliances | 3 | 720 | 375 | 375 | 74.3 | 74.2 | 80.2 | 66.3 | 33.3 | 37 |
| SonyAIBORobotSurface1 | 2 | 70 | 20 | 601 | 96.0 | 95.6 | 88.8 | 90.6 | 91.8 | 34 |
| SonyAIBORobotSurface2 | 2 | 65 | 27 | 953 | 88.6 | 87.0 | 92.4 | 90.0 | 84.9 | 34 |
| StarLightCurves | 3 | 1024 | 1000 | 8236 | 95.9 | 95.3 | 97.7 | 88.8 | 90.8 | 30 |
| Strawberry | 2 | 235 | 613 | 370 | 95.2 | 95.1 | 96.8 | 92.5 | 91.7 | 34 |
| SwedishLeaf | 15 | 128 | 500 | 625 | 88.7 | 87.7 | 93.9 | 89.9 | 75.8 | 37 |
| Symbols | 6 | 398 | 25 | 995 | 93.4 | 92.5 | 86.2 | 91.9 | 90.8 | 43 |
| SyntheticControl | 6 | 60 | 300 | 300 | 98.8 | 98.7 | 98.7 | 99.5 | 92.0 | 39 |
| ToeSegmentation1 | 2 | 277 | 40 | 228 | 92.0 | 90.7 | 95.4 | 93.4 | 90.4 | 40 |
| ToeSegmentation2 | 2 | 343 | 36 | 130 | 93.1 | 90.6 | 94.7 | 94.3 | 87.3 | 36 |
| Trace | 4 | 275 | 100 | 100 | 100.0 | 99.9 | 100.0 | 99.6 | 99.8 | 25 |
| TwoLeadECG | 2 | 82 | 23 | 1139 | 95.8 | 96.6 | 98.4 | 99.4 | 92.0 | 36 |
| TwoPatterns | 4 | 128 | 1000 | 4000 | 95.8 | 93.1 | 95.2 | 99.4 | 69.6 | 33 |
| UWaveGestureLibraryAll | 8 | 945 | 896 | 3582 | 94.9 | 94.8 | 94.2 | 68.0 | 76.6 | 44 |
| UWaveGestureLibraryX | 8 | 315 | 896 | 3582 | 80.2 | 77.9 | 80.6 | 80.4 | 69.4 | 42 |
| UWaveGestureLibraryY | 8 | 315 | 896 | 3582 | 71.5 | 69.5 | 73.7 | 71.8 | 59.1 | 40 |
| UWaveGestureLibraryZ | 8 | 315 | 896 | 3582 | 74.4 | 72.2 | 74.7 | 73.7 | 63.8 | 42 |
| Wafer | 2 | 152 | 1000 | 6164 | 99.4 | 99.3 | 100.0 | 99.6 | 98.1 | 24 |
| Wine | 2 | 234 | 57 | 54 | 86.5 | 86.4 | 92.6 | 52.4 | 79.4 | 39 |
| WordSynonyms | 25 | 270 | 267 | 638 | 68.4 | 62.8 | 58.2 | 58.1 | 46.1 | 42 |
| Worms | 5 | 900 | 181 | 77 | 65.6 | 59.9 | 71.9 | 64.2 | 62.2 | 47 |
| WormsTwoClass | 2 | 900 | 181 | 77 | 74.1 | 69.1 | 77.9 | 73.6 | 70.6 | 50 |
| Yoga | 2 | 426 | 300 | 3000 | 83.3 | 80.2 | 82.3 | 83.3 | 72.1 | 40 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vandewiele, G.; Ongenae, F.; De Turck, F. GENDIS: Genetic Discovery of Shapelets. Sensors 2021, 21, 1059. https://doi.org/10.3390/s21041059
Vandewiele G, Ongenae F, De Turck F. GENDIS: Genetic Discovery of Shapelets. Sensors. 2021; 21(4):1059. https://doi.org/10.3390/s21041059
Chicago/Turabian StyleVandewiele, Gilles, Femke Ongenae, and Filip De Turck. 2021. "GENDIS: Genetic Discovery of Shapelets" Sensors 21, no. 4: 1059. https://doi.org/10.3390/s21041059
APA StyleVandewiele, G., Ongenae, F., & De Turck, F. (2021). GENDIS: Genetic Discovery of Shapelets. Sensors, 21(4), 1059. https://doi.org/10.3390/s21041059








