3.1. Collection of Real BCG Dataset
To non-intrusively and conveniently collect BCG signal, an unobtrusive BCG signal acquisition system (named RS-611) is employed in this study [
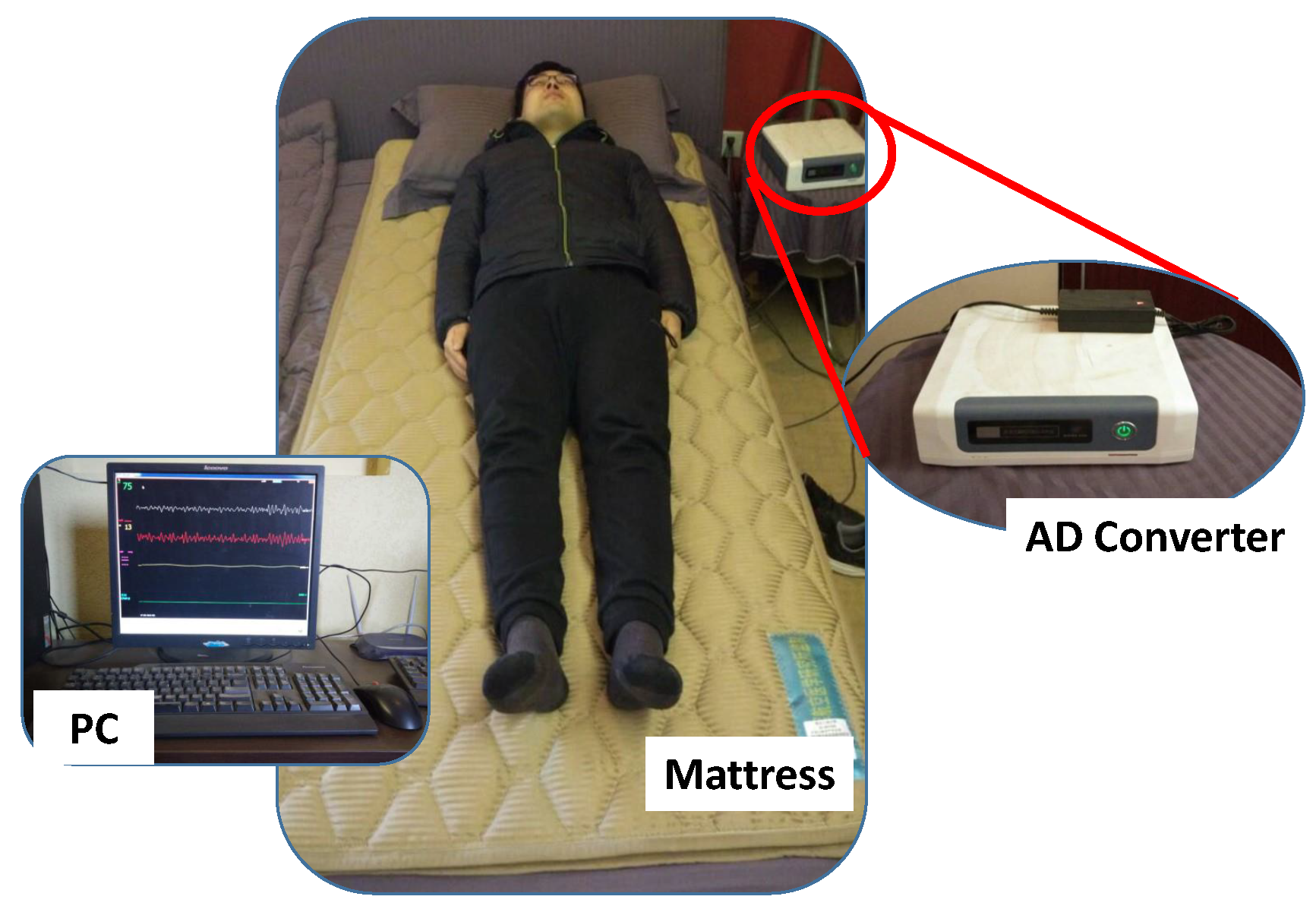
22]. The RS-611 is a certificated medical device developed by Institute of Air-force Aviation Medicine (Beijing, China). As shown in
Figure 1, the RS-611 is comprised of a micro-movement sensitive mattress (MSM), an analog-digital (AD) converter and a terminal PC. The MSM is the main function part of this system, in which two hydraulic pressure sensors (oil tubes) in the shape of strips are embedded. Particularly, one is located at the upper part of the mattress (i.e., chest area) and the other one is placed at the leg region. Both of them are long enough and placed in parallel, which guarantees that proper BCG signal can be captured regardless of the size of the subject. When a subject lies on the MSM, the changes of original pressure caused by cardiac activities (i.e., heartbeats) are recorded, amplified, and converted to digital signal by utilizing the 16-bit-resolution AD converter with a sample rate of 100 Hz, forming a time series of composite pressure data. Afterwards, BCG signal is separated from the original composite pressure signal by performing drift compensation and digital filtering based on the built-in software, and eventually displayed on the PC. In [
23], a real-time comparison between BCG signal collected by RS-611 and ECG signal collected by a commercial ECG collector (Prince 180D,
http://www.healforce.com/en/) was made, whose experimental results indicated that each wave trough of the collected BCG signal corresponded to a local minimum point of the collected ECG signal, which demonstrated the suitability of the collected BCG signal for analyzing and diagnosing cardiovascular diseases. In comparison with other wearable signal acquisition devices like ECG Holter [
24], smart watch [
25], smart chair [
26], PPG based non-invasive blood pressure estimator [
27] and cuff-based mercury sphygmomanometer, the MSM-based RS-611 is completely unobtrusive and the users do not need to attach any electrodes or devices on their body. Furthermore, the RS-611 does not require complex configurations, which is especially suitable for the elderly.
In this study, 175 staff from our university were recruited as subjects. All of them gave their informed consent for inclusion before they participated in the study. The study was conducted in accordance with the Declaration of Helsinki, and the protocol was approved by the Medical Experimental Ethical Inspection Institute of Northwestern Polytechnical University (No.20170078). First, we conducted routine examinations to obtain basic physiological index. In particular, the blood pressure was measured by a trained nurse by using a standard mercury sphygmomanometer. At each examination, blood pressure was measured in the left arm twice after five minutes of rest. According to the guidelines of the American Society of Hypertension [
28], the subjects were asked to be in seated position when measured. For each subject, blood pressure was recorded in four separate sessions within 3 weeks, and the averaged values were then used to represent the final systolic and diastolic blood pressure values. Subjects with blood pressure values greater than 140/90 mmHg were viewed as hypertensive patients, while the rest were regarded as normotensive subjects. In particular, subjects were excluded if they met any of the following criteria [
29]: (1) use of any anti-hypertensive drugs when measuring blood pressures, (2) Body Mass Index (BMI) >30, (3) history or clinical evidence of congestive heart failure or myocardial infarction, (4) atrial fibrillation, (5) diabetes mellitus, (6) alcoholics or smokers. After applying these exclusions, 68 hypertensive patients and 76 normotensive subjects were eligible for our experiments.
Before collecting BCG signal, we instructed each subject about how to use the RS-611 properly. In addition, the day before the experiment they were asked to sleep on the bed for about one hour for the sake of gathering feedbacks. We adjusted the indoor temperature, indoor light intensity, noise level and other factors that may affect their normal sleep according to their feedbacks, aiming to make sure that can sleep well through the whole night. Finally, for each subject one night of BCG signal was collected when they were sleeping, since the physiological indexes are relatively stable and not affected by other factors during sleep. The next morning, subjects were asked some questions, for example, when did they fall asleep, when did they get up, and how did they sleep. If the subjects did not sleep well or the sleep time was less than seven hours, the corresponding BCG recordings were considered inappropriate for research and were therefore discarded. Furthermore, a manual visual inspection was conducted on each BCG recording. BCG recordings that contained too many drastic fluctuations were also excluded from this research. After the inspection process, the final BCG dataset left 128 BCG recordings in total, including 61 hypertensive patients and 67 normotensive subjects. The statistical results of the BCG dataset are summarized in
Table 1.
3.3. RR intervals Sequence Extraction
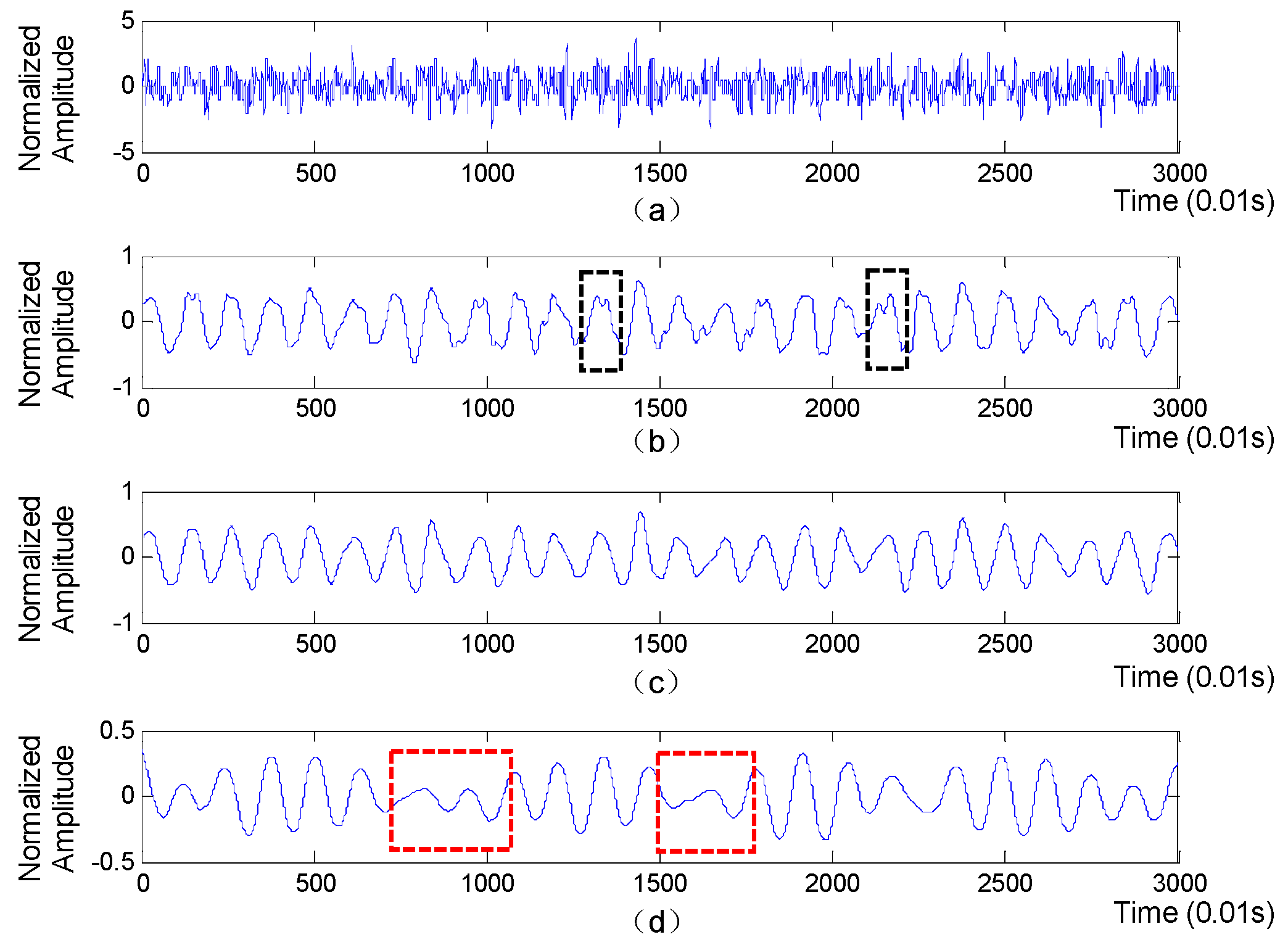
A segment of collected BCG signal is shown in
Figure 3 (top subplot), from which we will extract accurate RR intervals sequence for further extraction of HRV-related features.
The extraction of RR intervals sequence mainly consists of three steps as follows:
Signal normalization: Since the amplitude value of BCG signal is influenced by body weight to a certain extent, we first eliminate BCG signal amplitude difference caused by different body weight by normalizing the BCG signal using Z-score method [
13], which is written as:
where
is the
value of BCG signal,
and
represent the mean value and the standard deviation of the BCG signal respectively, and
is the normalization form of
. Afterwards, we design an elliptic bandpass filter to filter the noise component that is irrelative to the heart beats. Based on extensive experiments, the passband corner frequency and the stopband corner frequency are set to 5/6 Hz and 13/6 Hz respectively and the passband ripple and the stopband attenuation are set to 0.2 and 8 respectively.
Wavelet decomposition: Discrete wavelet transform (DWT) is a signal processing technique that is often employed to extract useful information from non-stationary time series, including physiological signals [
14,
15]. The utilization of DWT usually has a wonderful advantage that it can decompose the time series into multiple time-frequency resolutions, by which the details of the signal in time and frequency domains can be clearly displayed. Therefore, in this paper we utilize DWT to decompose BCG signal and further select the most appropriate approximation layer that can clearly reflect the heartbeats. Concretely, the BCG signal is first decomposed into detail component and approximation component at the first level, which is implemented by passing it through a pair of high-pass and low-pass filters as follows:
where
represent the
kth sample of BCG signal, and
and
are high-pass filter and low-pass filter respectively. The detail and approximation components are comprised of detailed coefficients and approximant coefficients, respectively. Afterwards, the above process is iteratively applied to the approximation component at each level until a specified level is reached. Finally, the approximation coefficients at each level are separately utilized to reconstruct an approximation layer of the original BCG signal. The approximation layers derived from BCG signal is shown in
Figure 3, with the ‘db6’ function used as the wavelet basis. Obviously, compared with the 5th approximation layer, the parts in red boxes mean that the 6th approximation layer may miss some heartbeats, while the parts in black boxes indicate that the 4th approximation layer is prone to generate some fake wave peaks, which will lead to inaccurate RR intervals. Moreover, the waveform of the 5th approximation layer is more regular and smoother than that of the 4th and 5th approximation layers. Therefore, the 5th approximation layer is finally selected to extract RR intervals.
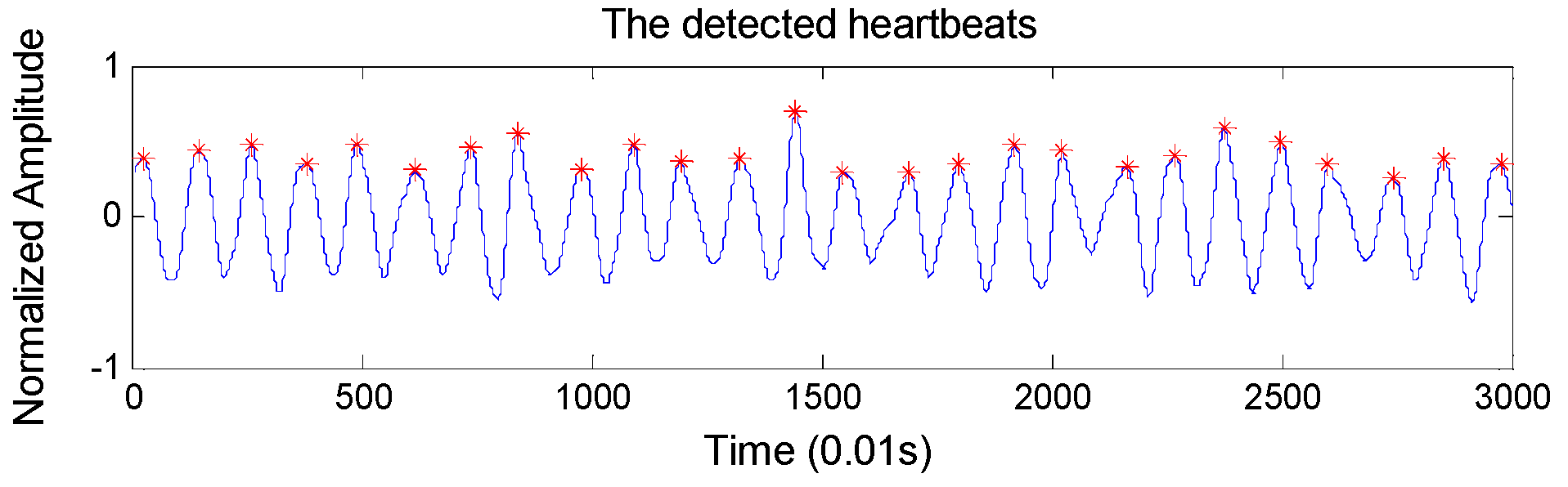
RR intervals extraction: To locate the heartbeats accurately, we design an overlapping sliding window to detect BCG wave peaks automatically. Concretely, the window size and the overlapping size are set to 100 samples and 60 samples respectively (the sample frequency of the BCG signal is 100 Hz), considering that the normal heart rate range is 60–100 per minute. The detected heartbeats are shown in
Figure 4, which shows very high heartbeat detection performance. In particular, it achieves an accuracy of 98.4% in detecting the heartbeats compared with ECG monitor.
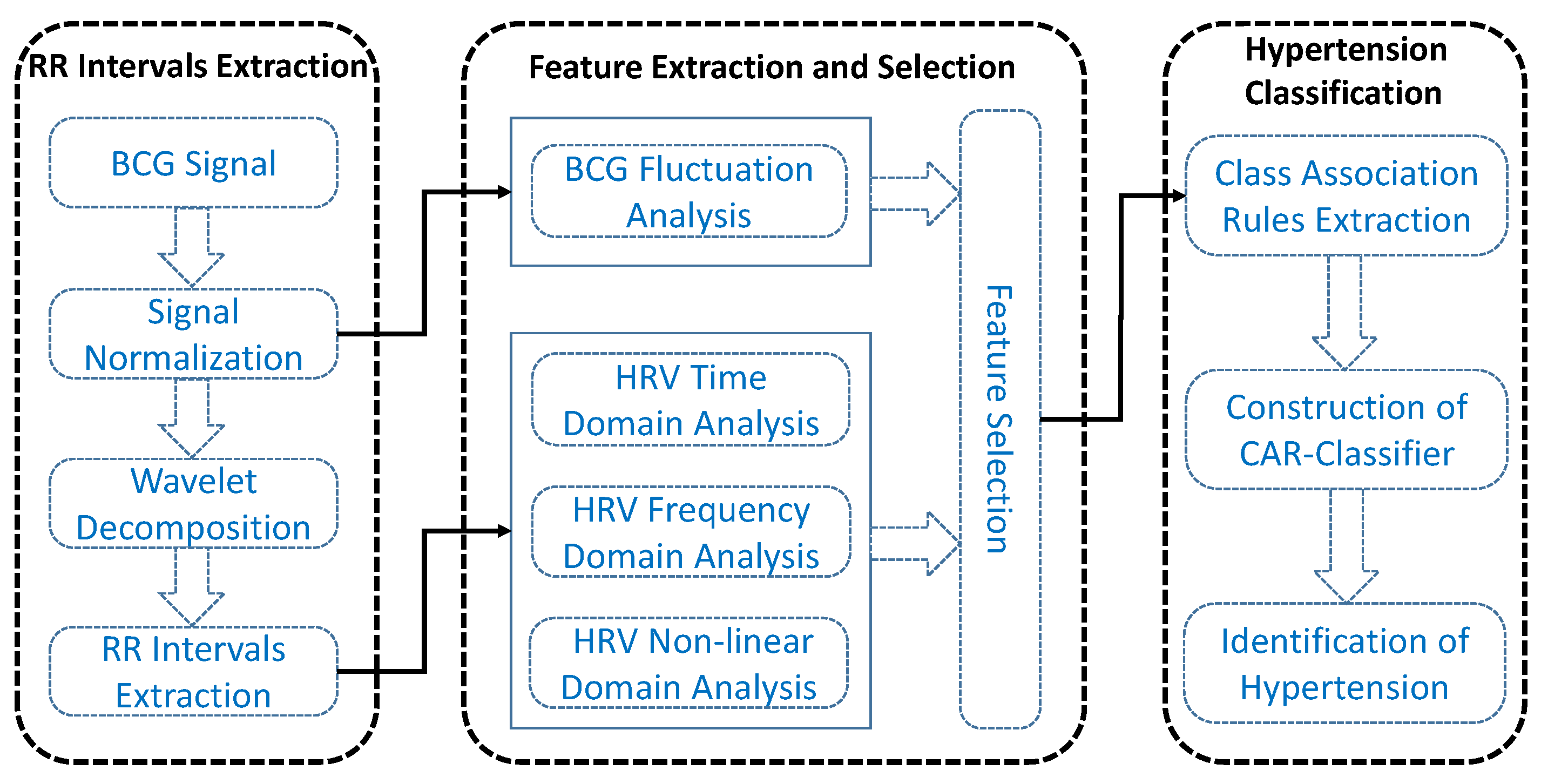
3.5. Hypertension Classification
In this study, we discriminate hypertensive patients and normotensive subjects by designing a CAR-Classifier, which integrates association rules mining and classification together. By this method, the association relationship among multi-dimensional features can be fully investigated. Furthermore, it also generates a set of CARs, which are helpful for doctors to analyze patients’ condition in-depth [
41,
42,
43,
44,
45,
46].
Before elaborating the proposed CAR-Classifier [
47,
48], we formulize some definitions that are relevant to the CAR-Classifier. In CAR-based classification problems, the dataset is usually viewed as a relational table. Let
D = {
d1,
d2,…,
dN} represent the dataset, which contains
N instances described by
k − 1 distinct attributes. Let
Y = {
y1,
y2,…,
yM} denote a finite set of class labels that has
M known classes. Each instance belongs to one of the
M classes. To simplify the narrative and make it easier to understand, we treat the class label as a special attribute, thus, each instance has
k distinct attributes. Attributes in
D can be discrete or continuous, while all of them should be preprocessed uniformly before mining CARs. Concretely, for any discrete attribute, all the possible values are mapped to a set of consecutive positive integers. Similarly, for any continuous attribute, its value range is discretized into several intervals which are further mapped to consecutive positive integers. For example, assume that an instance is described by three attributes (i.e., attribute
A, attribute
B and class label
C) which are discretized into three intervals, four intervals and five categories, respectively. In this case, the intervals of A, B and C are mapped with numbers 1 through 3, numbers 1 through 4, and numbers 1 through 5, respectively. Based on this mappings, an instance can be represented as a set of
<attribute, value> pairs, and each pair is referred as an
item. Let
I = {
item1,
item2,…,
itemL} denote the set of items in
D, which contains
L items. Afterwards, we define
itemSet as a subset of
I. Accordingly, an
itemSet consisting of
k items is called
k-itemSet. Particularly, an
itemSet that contains only one item is called 1-
itemSet. Moreover, if a
k-itemSet (
k ≥ 2) contains just one
item derived from the class attribute, we refer this
k-itemSet as a candidate class association rule (CCAR), which can be represented as a form of follows:
where
y denotes the
item derived from the class attribute, while
ruleItems represents the rest
items of the
k-itemSet. A CCAR is with
support (sup) s signifies that
s% of the instances in
D contain the
ruleItems and are labeled with
y. Meanwhile, a CCAR has
confidence (
conf)
c means that
c% of instances in
D that contain the
ruleItems are labeled with
y. Let
RuleitemsCount and
CandidateCount represent the number of instances in
D that contain the
ruleItems, and the number of instances in
D that contain the
ruleItems and are labeled with
y, respectively. Thus, the
sup and
conf of a given CCAR can be computed as follows:
where
is the number of the instances in
D. Particularly, a CCAR whose
sup is greater than a user-specified
minimum support (minSup) is called frequent candidate class association rule (FCCAR), otherwise called infrequent candidate class association rule (ICCAR). Furthermore, if the
conf of a FCCAR is greater than a user-specified
minimum confidence (minConf), we call this FCCAR a class association rule (CAR).
Next, we give a simple example to illustrate the relationship among the CCAR, FCCAR and CAR. Given a CCAR with form of:
where
A and
B are attributes, and
C is the class label. Actually, the
and
are the
ruleItems and
y in Equation 7, respectively. Assume that the
RuleitemsCount and the
CandidateCount of the {<
A, 1>, <
B, 3>} are 3 and 2 respectively, and the number of instances in
D is 10. Thus, the
sup of the CCAR is 2/10 = 20%, and the confidence is 2/3 = 66.7%. If the
minSup is specified as 10%, we say that this CCAR is a FCCAR since its
sup is greater than the
minSup. Furthermore, if the
minConf is set less than 66.7%, this CCAR is actually a CAR, and hence can be utilized when constructing the CAR-Classifier.
Based on the concepts defined above, the construction of the CAR-Classifier is elaborated as follows. Concretely, building a CAR-Classifier mainly consists of two stages [
34,
39,
49], i.e., the rule generation stage (RG-Stage) and the classifier build stage (CB-Stage). The former aims to mine all CARs, i.e., CCARs that satisfy specific
minSup and
minConf requirements, while the latter is to build the CAR-Classifier based on the extracted CARs.
RG-Stage: Since the features extracted in
Section 3.4 are continuous values, and cannot be directly used to mine CARs [
50], we adopt the equal-width strategy to discretize the feature values into some equal-width intervals. Based on the advice given by doctors, each feature is divided into five equal intervals which represent very low, low, medium, high, very high, respectively. Accordingly, we use numbers 1 through 5 to represent these five intervals. In addition, the class attribute consists of two categories, i.e., hypertensive and normotensive, which corresponds to numbers 1 and 2 respectively.
In order to mine CARs, we design a rule generation algorithm based on the well-known Apriori algorithm [
46]. It extracts all the CARs by making multiple traversals over the dataset. Concretely, in the 1
th traversal, it picks out all the frequent
1-itemSets. In the
k + 1
th traversal, it first generate frequent (
k + 1)-
itemSets based on the set of frequent
k-itemSets obtained in the
kth pass. Then it singles out all CCARs from the obtained frequent (
k + 1)-
itemSets. Afterwards, the
sup and
conf of these CCARs are computed. At the end of the current traversal, CARs are picked out from the obtained CCARs based on the computed
sup and
conf, and the user-specified
minSup and
minConf. Eventually, the CARs extracted from each traversals conjointly make up the final CAR set. The details of the proposed rule generation algorithm is described in Algorithm 1, where the function
candidateGen() represents the linkage of two (
k − 1)-
itemSets into one
k-itemSets.
| Algorithm 1: The Rule Generation Algorithm |
Definition:Let k-itemSet denote an itemSet that contains k items. Let Pk denote the set of k-itemSet that may be frequent. Let Ck denote the set of k-itemSet that has a class label-derived item, whose sup and conf are denoted as supCount and confValue respectively. Let Fk denote the set of frequent candidate rules. Let CARk denote the set of CARs that have k items.
|
| Input: |
| Output: |
| 1, F1={frequent 1-items} |
| 2, for(k=2; Fk-1≠Ø; k++) do |
| 3, Pk=link(Fk-1); |
| 4, Ck={p∈ Pk∣p has a class label-derived item}; |
| 5, Fk={c∈Ck∣c.supCount ≥ minConf} |
| 6, CARk={f∈Fk ∣ f.confValue ≥ minConf } |
| 7, end |
| 8, RuleSet=∪kCARk |
CB-Stage: For the sake of narrative, the set of CARs is thereafter referred to as RuleSet. To improve the classification performance, the most powerful CARs should be employed to construct the CAR-Classifier. To this end, we first sort the RuleSet in descending order, where the CARs at the front are of higher classification ability than the CARs at the back. When building the CAR-Classifier, the CARs at the front have the priority to be included into the CAR-classifier. Particularly, the sorting method employed in this paper is as follows:
Given two CARs named ri and rj, ri >rj (also called ri precedes rj or ri has a higher precedence relative to rj) if:
- (1)
the conf of ri is larger than that of rj, or
- (2)
their conf are the same, but the sup of ri is larger than that of rj, or
- (3)
both the conf and sup of ri and rj are the same, but ri has more items than rj.
Afterwards, we build the CAR-Classifier based on the sorted RuleSet. Concretely, for each CAR r in sorted RuleSet, we first traverse the dataset D to figure out whether r can match at least one instance (i.e., the discretized items derived from the instance can completely cover the antecedent of rule r). If so, we mark the rule r, and remove the instances that can be classified by r from D. Until no more instances can be correctly classified, the majority class of the rest instances in D is chosen as a default class. Eventually, all the marked CARs and the default class conjointly make up the CAR-Classifier. The construction of CAR-Classifier is described in Algorithm 2. Particularly, when using the CAR-Classifier to classify an unknown instance, the class label of the CAR that firstly matches the instance is viewed as the classification result. If there is no matched CAR, the instance will be labeled with the default class.
| Algorithm 2: The Classifier Build Algorithm |
Definition:Let RuleSet denote the set of extracted CARs, and the ith CAR in RuleSet is denoted as ri. Let D denote the dataset, and the ith case in D is denoted as di.
|
Input:The RuleSet The dataset D.
|
Output:The CAR-Classifier, which is denoted as C. The default class, which is denoted as DClass.
|
| 1, RuleSet=sort(RuleSet) |
| 2, for(i=1; i<RuleSet.size(); i++) do |
| 3, temp = Ø; |
| 4, for(j=1; j<D.size(), j++) do |
| 5, if dj is correctly classified by ri then |
| 6, store dj in temp and mark ri; |
| 7, end |
| 8, if ri is marked then |
| 9, insert ri at the end of C; |
| 10, delete all the cases in temp form D. |
| 11, end |
| 12, if D ≠ ∅ then |
| 13, DClass = the majority class label of the rest instances in D; |
| 14, C=C∪DClass; |