Hybrid Genome Assembly and Annotation of the Basidiomycete Fungus Candolleomyces candolleanus Strain CMU-8613 Using a Cost-Effective Iterative Pipeline
Abstract
1. Introduction
2. Results
2.1. Primary Assembly
2.2. Genome Polishing
- Racon: Uses the original long reads to build a consensus based on partial-order graphs (POA) and correct errors;
- Pilon: Uses aligned Illumina short reads for the assembly to correct specific errors (SNPs and small indels) with high accuracy;
- Racon + Medaka (R+M): Combines Racon with Medaka, another long-read polishing tool that uses neural networks and is optimized for ONT data, aiming to improve consensus accuracy;
- Racon + Medaka + Pilon (R+M+P): A hybrid approach where Racon and Medaka are first used to improve the long-read consensus, followed by Pilon with short reads for fine base-level correction.
2.3. Framework Using the Genome of Coprinellus Micaceus as an External Reference
2.4. Iterative Scaffolding Using the NECAT Assembly as a Reference
2.5. Comparison of Genome Annotation and Functional Analysis
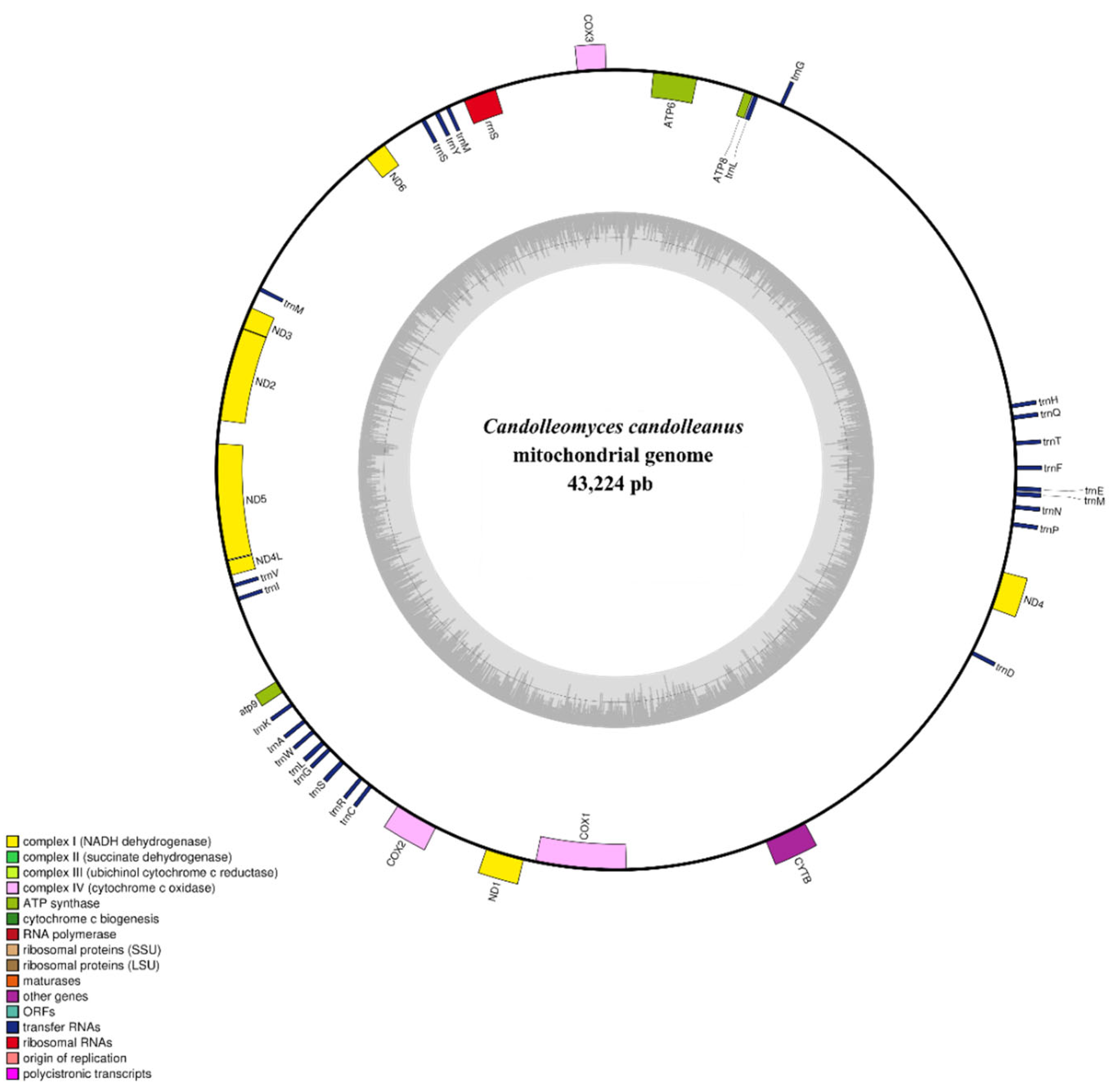
2.6. Features of the Mitochondrial Genome
2.7. Secondary Metabolite Gene Clusters (BGCs)
2.8. Number of CAZyme-Encoding Genes
2.9. Type of Mating in the CMU-8613 Strain
2.10. Phylogenetic Analysis
3. Discussion
4. Materials and Methods
4.1. Study Strain
4.2. DNA Extraction
4.3. Illumina Sequencing
4.4. Oxford Nanopore Sequencing
4.5. Genome Assembly and Polishing
4.6. Genome Arrangement
4.7. Genome Annotation
4.8. Mitochondrial Genome Assembly
4.9. Identification of CAZyme-Encoding Genes
4.10. Identification of the MAT-A/MAT-B Genes
4.11. BLASTn Search and Phylogenetic Analysis
4.12. Operating Environment
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AA | Auxiliary Activities |
| CAZymes | Carbohydrate-Active enzymes |
| CBM | Carbohydrate-Binding Modules |
| CE | Carbohydrate Esterases |
| GH | Glycoside Hydrolases |
| GT | Glycosyl-Transferases |
| PL | Polysaccharide Lyases |
| NCBI | National Center for Biotechnology Information |
| CDS | Protein-coding genes |
| BGC | Biosynthetic Gene Cluster |
| COG | Clusters of Orthologous Genes |
| ITS | Internal transcribed spacer ribosomal region |
| LSU | Gene for the Large Ribosomal Subunit |
References
- Wächter, D.; Melzer, A. Proposal for a subdivision of the family Psathyrellaceae based on a taxon-rich phylogenetic analysis with iterative multigene guide tree. Mycol. Prog. 2020, 19, 1151–1265. [Google Scholar] [CrossRef]
- Li, Q.; Haqnawaz, M.; Niazi, A.R.; Khalid, A.N. Phylogenetic studies on the genus Candolleomyces (Psathyrellaceae, Basidiomycota) occurring in the bed of the Indus River, Punjab, Pakistan, reveal three new species. MycoKeys 2025, 112, 165–182. [Google Scholar] [CrossRef]
- Padamsee, M.; Matheny, P.B.; Dentinger, B.T.M.; McLaughlin, D.J. The mushroom family Psathyrellaceae: Evidence for large-scale polyphyly of the genus Psathyrella. Mol. Phylogenet. Evol. 2008, 46, 415–429. [Google Scholar] [CrossRef] [PubMed]
- Janusz, G.; Pawlik, A.; Sulej, J.; Świderska-Burek, U.; Jarosz-Wilkołazka, A.; Paszczyński, A. Lignin degradation: Microorganisms, enzymes involved, genomes analysis and evolution. FEMS Microbiol. Rev. 2017, 41, 941–962. [Google Scholar] [CrossRef] [PubMed]
- Cicero, I.; Mirabile, G.; Venturella, G. Potential medicinal fungi from freshwater environments as resources of bioactive compounds. J. Fungi 2025, 11, 54. [Google Scholar] [CrossRef]
- Liktor-Busa, E.; Kovács, B.; Urbán, E.; Hohmann, J.; Ványolós, A. Investigation of Hungarian mushrooms for antibacterial activity and synergistic effects with standard antibiotics against resistant bacterial strains. Lett. Appl. Microbiol. 2016, 62, 437–443. [Google Scholar] [CrossRef]
- Pan, Y.; Zheng, W.; Yang, S. Chemical and activity investigation on metabolites produced by an endophytic fungi Psathyrella candolleana from the seed of Ginkgo biloba. Nat. Prod. Res. 2020, 34, 3130–3133. [Google Scholar] [CrossRef]
- Liu, Y.P.; Dai, Q.; Pu, C.J.; Wang, M.; Li, Z.H.; Liu, J.K.; Feng, T. Psathyrellanic acid, a monocyclic diterpenoid from the Basidiomycete Psathyrella candolleana. Nat. Prod. Commun. 2019, 14, 1934578X19850958. [Google Scholar] [CrossRef]
- Liu, Y.P.; Dai, Q.; Wang, W.X.; He, J.; Li, Z.H.; Feng, T.; Liu, J.K. Psathyrins: Antibacterial diterpenoids from Psathyrella candolleana. J. Nat. Prod. 2020, 83, 1725–1729. [Google Scholar] [CrossRef]
- Wu, H.; Yang, H.X.; Li, Z.H.; Feng, T.; Liu, J.K. Psathyrellins A–E, antibacterial guanacastane diterpenoids from mushroom Psathyrella candolleana. Nat. Prod. Bioprospect. 2021, 11, 447–452. [Google Scholar] [CrossRef]
- He, J.; Du, J.X.; Zhao, Q.R.; Liu, Y.P.; Yang, Y.L.; Liu, J.K.; Feng, T. Biochemical and genetic basis of guanacastane diterpene biosynthesis in basidiomycete fungi. Org. Lett. 2023, 25, 5345–5349. [Google Scholar] [CrossRef] [PubMed]
- Lohse, M.; Drechsel, O.; Kahlau, S.; Bock, R. OrganellarGenomeDRAW: A suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucleic Acids Res. 2013, 41, W575–W581. [Google Scholar] [CrossRef]
- Narh-Mensah, D.L.; Wingfield, B.D.; Coetzee, M.P. A practical approach to genome assembly and annotation of Basidiomycota using the example of Armillaria. BioTechniques 2023, 75, 115–128. [Google Scholar] [CrossRef] [PubMed]
- Schafhauser, T.; Wibberg, D.; Binder, A.; Rückert, C.; Busche, T.; Wohlleben, W.; Kalinowski, J. Genome assembly and genetic traits of the pleuromutilin-producer Clitopilus passeckerianus DSM1602. J. Fungi 2022, 8, 862. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro-Tomé, L.M.; Quintanilha-Peixoto, G.; Costa-Rezende, D.H.; Salvador-Montoya, C.A.; Cardoso, D.; Araújo, D.S.; Góes-Neto, A. Comparative genomics and stable isotope analysis reveal the saprotrophic-pathogenic lifestyle of a neotropical fungus. mBio 2024, 15, e01423-24. [Google Scholar] [CrossRef]
- Utomo, C.; Tanjung, Z.A.; Aditama, R.; Pratomo, A.D.M.; Buana, R.F.N.; Putra, H.S.G.; Liwang, T. Whole-genome sequencing of Ganoderma boninense, the causal agent of basal stem rot disease in oil palm, via combined short- and long-read sequencing. Sci. Rep. 2024, 14, 10520. [Google Scholar] [CrossRef]
- Xie, Y.; Zhong, Y.; Chang, J.; Kwan, H.S. Chromosome-level de novo assembly of Coprinopsis cinerea A43mut B43mut pab1-1# 326 and genetic variant identification of mutants using Nanopore MinION sequencing. Fungal Genet. Biol. 2021, 146, 103485. [Google Scholar]
- Yoshitake, K.; Shirasawa, K.; Kojima, K.K.; Asakawa, S.; Tanaka, N.; Kurokochi, H. Nearly telomere-to-telomere genome assembly of the L. edodes diploid genome. Fungal Genet. Biol. 2025, 179, 104005. [Google Scholar] [CrossRef]
- Castanera, R.; Borgognone, A.; Pisabarro, A.G.; Ramírez, L. Biology, dynamics, and applications of transposable elements in basidiomycete fungi. Appl. Microbiol. Biotechnol. 2017, 101, 1337–1350. [Google Scholar] [CrossRef]
- Chen, Y.; Nie, F.; Xie, S.Q.; Zheng, Y.F.; Dai, Q.; Bray, T.; Xiao, C.L. Efficient assembly of nanopore reads via highly accurate and intact error correction. Nat. Commun. 2021, 12, 60. [Google Scholar] [CrossRef]
- Kolmogorov, M.; Yuan, J.; Lin, Y.; Pevzner, P.A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 2019, 37, 540–546. [Google Scholar] [CrossRef]
- Wright, R.; Woof, K. The genome sequence of the glistening inkcap, Coprinellus micaceus Coprinellus; Coprinellus micaceus ((Bull.) Vilgalys, Hopple & Jacq. Johnson, 2001). Wellcome Open Res. 2024, 9, 677. [Google Scholar] [CrossRef]
- Stajich, J.E. Fungal genomes and insights into the evolution of the kingdom. Microbiol. Spectr. 2017, 5, FUNK-0055-2016. [Google Scholar] [CrossRef]
- Ruiz-Dueñas, F.J.; Barrasa, J.M.; Sánchez-García, M.; Camarero, S.; Miyauchi, S.; Serrano, A.; Martínez, A.T. Genomic analysis enlightens Agaricales lifestyle evolution and increasing peroxidase diversity. Mol. Biol. Evol. 2021, 38, 1428–1446. [Google Scholar] [CrossRef]
- Hess, J.; Skrede, I.; Wolfe, B.E.; LaButti, K.; Ohm, R.A.; Grigoriev, I.V.; Pringle, A. Transposable element dynamics among asymbiotic and ectomycorrhizal Amanita fungi. Genome Biol. Evol. 2014, 6, 1564–1578. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Lu, D.; Wang, S.; Zhao, Y.; Gao, S.; Han, R.; Hu, S. Genome assembly and pathway analysis of edible mushroom Agrocybe cylindracea. Genom. Proteom. Bioinform. 2020, 18, 341–351. [Google Scholar] [CrossRef]
- Wang, S.R.; Zhang, J.P.; He, Y.R.; Chang, M.C.; Meng, J.L. Characterization of the complete mitochondrial genome of Coprinellus micaceus, a wild saprobic mushroom in China. Mitochondrial DNA Part B 2021, 6, 1979–1981. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.C.; Xie, T.C.; Feng, X.L.; Wang, Z.X.; Lin, C.; Li, G.M.; Qi, J. The first five mitochondrial genomes for the family Nidulariaceae reveal novel gene rearrangements, intron dynamics, and phylogeny of Agaricales. Int. J. Mol. Sci. 2023, 24, 12599. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Wang, Y.; Yao, W.; Shen, J.; Chen, M.; Gao, M.; Liu, N. The 256 kb mitochondrial genome of Clavaria fumosa is the largest among phylum Basidiomycota and is rich in introns and intronic ORFs. IMA Fungus 2020, 11, 26. [Google Scholar] [CrossRef]
- Stajich, J.E.; Wilke, S.K.; Ahrén, D.; Au, C.H.; Birren, B.; Borodovsky, M.; Pukkila, P.J. Insights into evolution of multicellular fungi from the assembled chromosomes of the mushroom Coprinopsis cinerea (Coprinus cinereus). Proc. Natl. Acad. Sci. USA 2010, 107, 11889–11894. [Google Scholar] [CrossRef]
- Schueller, A.; Studt-Reinhold, L.; Strauss, J. How to completely squeeze a fungus—advanced genome mining tools for novel bioactive substances. Pharmaceutics 2022, 14, 1837. [Google Scholar] [CrossRef] [PubMed]
- Chiang, C.Y.; Ohashi, M.; Tang, Y. Deciphering chemical logic of fungal natural product biosynthesis through heterologous expression and genome mining. Nat. Prod. Rep. 2023, 40, 89–127. [Google Scholar] [CrossRef]
- Duan, Y.; Han, H.; Qi, J.; Gao, J.M.; Xu, Z.; Wang, P.; Liu, C. Genome sequencing of Inonotus obliquus reveals insights into candidate genes involved in secondary metabolite biosynthesis. BMC Genom. 2022, 23, 314. [Google Scholar] [CrossRef]
- Xie, X.; Zhao, L.; Song, Y.; Qiao, Y.; Wang, Z.X.; Qi, J. Genome-wide characterization and metabolite profiling of Cyathus olla: Insights into the biosynthesis of medicinal compounds. BMC Genom. 2024, 25, 618. [Google Scholar] [CrossRef] [PubMed]
- Saini, S.; Sharma, K.K. Fungal lignocellulolytic enzymes and lignocellulose: A critical review on their contribution to multiproduct biorefinery and global biofuel research. Int. J. Biol. Macromol. 2021, 193, 2304–2319. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Li, Q.; Liu, C.; Meng, E.; Zhang, B. Microbial degradation of lignocellulose for sustainable biomass utilization and future research perspectives. Sustainability 2025, 17, 4223. [Google Scholar] [CrossRef]
- Coelho, M.A.; Bakkeren, G.; Sun, S.; Hood, M.E.; Giraud, T. Fungal sex: The basidiomycota. Microbiol. Spectr. 2017, 5, FUNK-0046-2016. [Google Scholar] [CrossRef]
- Chu, C.; Li, D.; Gu, L.; Yang, S.; Liu, C. Evidence for the existence of mating subtypes within the Schizophyllum commune: Mating behavior and genetic divergence. J. Fungi 2025, 11, 277. [Google Scholar] [CrossRef]
- Mukhtar, I.; Arredondo-Santoyo, M.; Vázquez-Garciduenas, S.; Vázquez-Marrufo, G. Isolation and molecular identification of laccase-producing saprophytic/phytopathogenic mushroom-forming fungi from various ecosystems in Michoacan State, Mexico. Acta Mycol. 2019, 54, 1119. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Koren, S.; Walenz, B.P.; Berlin, K.; Miller, J.R.; Bergman, N.H.; Phillippy, A.M. Canu: Scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017, 27, 722–736. [Google Scholar] [CrossRef] [PubMed]
- Li, H. Minimap2: Pairwise alignment for nucleotide sequences. Bioinformatics 2018, 34, 3094–3100. [Google Scholar] [CrossRef]
- Vaser, R.; Sović, I.; Nagarajan, N.; Šikić, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 2017, 27, 737–746. [Google Scholar] [CrossRef]
- Medaka. Available online: https://github.com/nanoporetech/medaka (accessed on 11 August 2025).
- Walker, B.J.; Abeel, T.; Shea, T.; Priest, M.; Abouelliel, A.; Sakthikumar, S.; Earl, A.M. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 2014, 9, e112963. [Google Scholar] [CrossRef] [PubMed]
- Guan, D.; McCarthy, S.A.; Wood, J.; Howe, K.; Wang, Y.; Durbin, R. Identifying and removing haplotypic duplication in primary genome assemblies. Bioinformatics 2020, 36, 2896–2898. [Google Scholar] [CrossRef]
- Alonge, M.; Lebeigle, L.; Kirsche, M.; Jenike, K.; Ou, S.; Aganezov, S.; Soyk, S. Automated assembly scaffolding using RagTag elevates a new tomato system for high-throughput genome editing. Genome Biol. 2022, 23, 258. [Google Scholar] [CrossRef] [PubMed]
- Darling, A.E.; Mau, B.; Perna, N.T. progressiveMauve: Multiple genome alignment with gene gain, loss and rearrangement. PLoS ONE 2010, 5, e11147. [Google Scholar] [CrossRef]
- Funannotate. Available online: https://github.com/nextgenusfs/funannotate (accessed on 30 August 2025).
- Blin, K.; Shaw, S.; Vader, L.; Szenei, J.; Reitz, Z.L.; Augustijn, H.E.; Weber, T. antiSMASH 8.0: Extended gene cluster detection capabilities and analyses of chemistry, enzymology, and regulation. Nucleic Acids Res. 2025, 53, W32–W38. [Google Scholar] [CrossRef]
- Matsuda, Y. Global genome mining in fungi using FunBGCeX. Methods Enzymol. 2025, 717, 99–316. [Google Scholar]
- Uliano-Silva, M.; Ferreira, J.G.R.; Krasheninnikova, K.; Formenti, G.; Abueg, L.; Torrance, J.; McCarthy, S.A. MitoHiFi: A python pipeline for mitochondrial genome assembly from PacBio high fidelity reads. BMC Bioinform. 2023, 24, 288. [Google Scholar] [CrossRef]
- Allio, R.; Schomaker-Bastos, A.; Romiguier, J.; Prosdocimi, F.; Nabholz, B.; Delsuc, F. MitoFinder: Efficient automated large-scale extraction of mitogenomic data in target enrichment phylogenomics. Mol. Ecol. Resour. 2020, 20, 892–905. [Google Scholar] [CrossRef]
- Zheng, J.; Ge, Q.; Yan, Y.; Zhang, X.; Huang, L.; Yin, Y. dbCAN3: Automated carbohydrate-active enzyme and substrate annotation. Nucleic Acids Res. 2023, 51, W115–W121. [Google Scholar] [CrossRef]
- Asif, M.; Izhar, A.; Niazi, A.R.; Khalid, A.N. Candolleomyces asiaticus sp. nov. (Psathyrellaceae, Agaricales), a novel species from Punjab, Pakistan. Eur. J. Taxon. 2022, 826, 176–187. [Google Scholar] [CrossRef]
- Nayana, P.K.; Pradeep, C.K. New species and new record of Candolleomyces (Psathyrellaceae) from India. Botany 2023, 101, 472–484. [Google Scholar] [CrossRef]
- Yang, K.L.; Lin, J.Y.; Li, G.M.; Yang, Z.L. Mushrooms adapted to seawater: Two new species of Candolleomyces (Basidiomycota, Agaricales) from China. J. Fungi 2023, 9, 1204. [Google Scholar] [CrossRef]
- Izhar, A.; Asif, M.; Khan, Z.; Khalid, A.N. Introducing two new members of the genus Candolleomyces (Agaricales) from Punjab, Pakistan. Plant Syst. Evol. 2023, 309, 40. [Google Scholar] [CrossRef]
- Örstadius, L.; Ryberg, M.; Larsson, E. Molecular phylogenetics and taxonomy in Psathyrellaceae (Agaricales) with focus on psathyrelloid species: Introduction of three new genera and 18 new species. Mycol. Prog. 2015, 14, 25. [Google Scholar] [CrossRef]
- Büttner, E.; Karich, A.; Nghi, D.H.; Lange, M.; Liers, C.; Kellner, H.; Hofrichter, M.; Ullrich, R. Candolleomyces eurysporus, a new Psathyrellaceae (Agaricales) species from the tropical Cúc Phương National Park, Vietnam. Austrian J. Mycol. 2020, 28, 79–92. [Google Scholar]
- Moreno, G.; Heykoop, M.; Esqueda, M.; Olariaga, I. Another lineage of secotioid fungi is discovered: Psathyrella secotioides sp. nov. from Mexico. Mycol. Prog. 2015, 14, 34. [Google Scholar] [CrossRef]
- Haqnawaz, M.; Niazi, A.R.; Khalid, A.N. A study on the genus Candolleomyces (Agaricales: Psathyrellaceae) from Punjab, Pakistan. BMC Microbiol. 2023, 23, 181. [Google Scholar] [CrossRef] [PubMed]
- Han, X.X.; Phurbu, D.; Ma, G.F.; Li, Y.Z.; Mei, Y.J.; Liu, D.M.; Cao, B. A taxonomic study of Candolleomyces specimens from China revealed seven new species. J. Fungi 2024, 10, 499. [Google Scholar] [CrossRef] [PubMed]
- Bau, T.; Yan, J.Q. Two new rare species of Candolleomyces with pale spores from China. MycoKeys 2021, 80, 149. [Google Scholar] [CrossRef]
- Yan, J.Q.; Bau, T. The Northeast Chinese species of Psathyrella (Agaricales, Psathyrellaceae). MycoKeys 2018, 33, 85. [Google Scholar] [CrossRef]
- Battistin, E.; Chiarello, O.; Vizzini, A.; Örstadius, L.; Larsson, E. Morphological characterisation and phylogenetic placement of the very rare species Psathyrella sulcatotuberculosa. Sydowia 2014, 66, 171–181. [Google Scholar]
- Larsson, E.; Örstadius, L. Fourteen coprophilous species of Psathyrella identified in the Nordic countries using morphology and nuclear rDNA sequence data. Mycol. Res. 2008, 112, 1165–1185. [Google Scholar] [CrossRef]
- Nie, C.; Wang, S.N.; Tkalčec, Z.; Yan, J.Q.; Hu, Y.; Ge, Y.; Mešić, A. Coprinus leucostictus rediscovered after a century, epitypified, and its generic position in Hausknechtia resolved by multigene phylogenetic analysis of Psathyrellaceae. Diversity 2022, 14, 699. [Google Scholar] [CrossRef]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 2019, 20, 1160–1166. [Google Scholar] [CrossRef]
- Nguyen, L.T.; Schmidt, H.A.; Von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Trifinopoulos, J.; Nguyen, L.T.; von Haeseler, A.; Minh, B.Q. W-IQ-TREE: A fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 2016, 44, W232–W235. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Teslenko, M.; Van Der Mark, P.; Ayres, D.L.; Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef] [PubMed]








| Criterion | Canu | Flye | NECAT |
|---|---|---|---|
| Complete BUSCOs | 96.5% (S: 92.5%, D: 4.1%) | 96.7% (S: 93.9%, D: 2.8%) | 92.4% (S: 90.4%, D: 2.0%) |
| Fragmented BUSCOs | 0.4% | 0.3% | 0.4% |
| Missing BUSCOs | 3.0% | 3.0% | 7.2% (worse) |
| Contig/N50 | 95 Kb (worst) | 183 Kb | 291 KB (best) |
| Number of contigs | 1785 (more fragmented) | 771 | 369 (less fragmented) |
| Total assembly size | 58.6 MB (major) | 55.9 Mb | 47.1 MB (smaller) |
| Advantages | Good recovery of the genome | Better recovery of complete genes | More continuous assembly |
| Disadvantages | Highly fragmented assembly | Intermediate continuity | More gene loss, shorter assembly |
| NECAT Assembly | ||||||||
| Polished | C (%) | S (%) | D (%) | F (%) | M (%) | Contigs | Total length (Mb) | N50 Contig |
| Racon | 92.5 | 90.6 | 1.8 | 0.5 | 7.0 | 296 | 46.8 | 1 Mb |
| Pilon | 92.4 | 90.4 | 2.0 | 0.5 | 7.1 | 1352 | 47.9 | 91 Kb |
| Racon + Medaka | 92.6 | 90.6 | 1.9 | 0.5 | 7.0 | 296 | 47.3 | 845 Kb |
| Racon + Medaka + Pilon | 92.6 | 90.6 | 1.9 | 0.5 | 6.9 | 296 | 47.1 | 842 Kb |
| Flye Assembly | ||||||||
| Racon | 96.5 | 93.9 | 2.7 | 0.4 | 3.1 | 392 | 55.6 | 584 Kb |
| Pilon | 96.6 | 93.7 | 2.9 | 0.3 | 3.1 | 1508 | 56.6 | 81 Kb |
| Racon + Medaka | 96.7 | 94.0 | 2.7 | 0.4 | 2.9 | 386 | 56.0 | 623 Kb |
| Racon + Medaka + Pilon | 96.3 | 93.7 | 2.6 | 1.3 | 2.4 | 384 | 55.8 | 620 Kb |
| Criterion | Canu (vs. Primary Assembly) | Flye (vs. Primary Assembly) | NECAT (vs. Primary Assembly) |
|---|---|---|---|
| Complete BUSCOs (%) | 96.5 (S: 92.4, D: 4.1) (=) | 96.7 (S: 94.0, D: 2.8) (=) | 92.6 (S: 90.6, D: 2.0) (+0.2) |
| Fragmented BUSCOs (%) | 0.4 (=) | 0.3 (=) | 0.4 (=) |
| BUSCOs absents (%) | 3.1 (+0.1) | 2.9 (−0.1) | 7.2 (=) |
| Contig N50 (Kb) | 95 (=) | 183 (=) | 291 (=) |
| Scaffold N50 (Kb) | 114 (+19) | 251 (+68) | 1000 (+709) |
| Number of contigs | 1785 (=) | 774 (+3) | 368 (−1) |
| Number of scaffolds | 1686 (−99) | 702 (−69) | 288 (−81) |
| Total assembled size (Mb) | 59.3 (+0.7) | 56.4 (+0.5) | 47.5 (+0.4) |
| Criterion | Canu | Flye |
|---|---|---|
| Complete BUSCOs (%) | 96.5 (S: 92.3, D: 4.2) (=) | 96.7 (S: 93.9, D: 2.8) (=) |
| Fragmented BUSCOs (%) | 0.4 (=) | 0.3 (=) |
| BUSCOs absents (%) | 3.1 (+0.1) | 3.0 (=) |
| Contig N50 (Kb) | 95 (=) | 183 (=) |
| Scaffold N50 (Kb) | 2000 (+1905) | 2000 (+1817) |
| Number of contigs | 1782 (−3) | 770 (−1) |
| Number of scaffolds | 882 (−903) | 370 (−401) |
| Total assembled size (Mb) | 60.6 (+2.0) | 56.4 (+0.5) |
| Criterion | NECAT Assembly | Flye Assembly | Observation |
|---|---|---|---|
| Complete BUSCOs (%) | 92.6 (S: 90.6, D: 1.9) | 96.3 (S: 94.0, D: 2.7) | Flye more complete |
| Fragmented BUSCOs (%) | 0.5 | 1.3 | Similar |
| BUSCOs absents (%) | 6.9 | 2.4 | Flye with less loss |
| Contig N50 (Kb) | 842 | 620 | NECAT more continuous (contig) |
| Scaffold N50 (Kb) | 1000 | 2000 | Flye more continuous (scaffold) |
| Number of contigs | 296 | 384 | NECAT less fragmented (contig) |
| Number of scaffolds | 286 | 365 | NECAT less fragmented (scaffold) |
| Total assembled size (Mb) | 47.2 | 55.8 | Flye larger |
| Metrics | Flye Assembly | NECAT Assembly | Observations |
| Number of Genes | 15, 550 | 13, 481 | Flye predicts +2069 genes |
| Number of mRNA | 15, 259 | 13, 206 | Correlates with more genes in Flye |
| Number of tRNA | 291 | 281 | Similar in both |
| Complete CDS | 15, 088 | 13, 063 | Mayor gene integrity in Flye |
| Total exons | 86, 010 | 75, 300 | Flye has more exons |
| Multi-section transcripts | 13, 664 | 11, 918 | Most apparent genetic complexity in Flye |
| Single exon | 1, 595 | 1, 288 | Low proportion in both |
| Average gene length (bp) | 1813.4 | 1816.4 | Very similar in both |
| Average exon length (bp) | 251.9 | 250.9 | Very similar in both |
| Average protein length (amino acids) | 499.1 | 501.6 | Very similar in both |
| Parameter | FunBGCex | antiSMASH |
|---|---|---|
| BGCs (Flye) | 25 | 16 |
| BGCs (NECAT) | 23 | 15 |
| Main types | Terpenes, NRPS, PPPS, DMATS, UbiA | Terpenes, NRPS, PKS, NRPS-PKS hybrids |
| Sensitivity | High (detects partial fragments or variants) | Moderate (focused on entire regions) |
| Structural precision | Minor (possible redundancy) | Mayor (clear definition of boundaries) |
| Match with known BGCs | High | High |
| FunBGCex | antiSMASH | |||
|---|---|---|---|---|
| Type of BGC | Flye | NECAT | Flye | NECAT |
| Terpene/TC (Class 1, SHC/OSC, AstC) | 14 | 12 | 10 | 9 |
| PPPS (Prenyl pyrophosphate synthase) | 3 | 3 | – | – |
| PT (Prenyltransferase type DMATS or UbiA) | 3 | 3 | – | – |
| NRPS-like | 2 | 2 | 2 | 2 |
| NRPS | 1 | 1 | 1 | 1 |
| NRPS-PKS | 1 | 1 | 1 | 1 |
| PKS/NR-PKS | 1 | 1 | 1 | 1 |
| Siderophore | – | – | 1 | 1 |
| Total | 25 | 23 | 16 | 15 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Villa-Villa, E.M.; Vázquez-Garcidueñas, M.S.; Vázquez-Marrufo, G. Hybrid Genome Assembly and Annotation of the Basidiomycete Fungus Candolleomyces candolleanus Strain CMU-8613 Using a Cost-Effective Iterative Pipeline. Int. J. Mol. Sci. 2026, 27, 509. https://doi.org/10.3390/ijms27010509
Villa-Villa EM, Vázquez-Garcidueñas MS, Vázquez-Marrufo G. Hybrid Genome Assembly and Annotation of the Basidiomycete Fungus Candolleomyces candolleanus Strain CMU-8613 Using a Cost-Effective Iterative Pipeline. International Journal of Molecular Sciences. 2026; 27(1):509. https://doi.org/10.3390/ijms27010509
Chicago/Turabian StyleVilla-Villa, Edgar Manuel, Ma. Soledad Vázquez-Garcidueñas, and Gerardo Vázquez-Marrufo. 2026. "Hybrid Genome Assembly and Annotation of the Basidiomycete Fungus Candolleomyces candolleanus Strain CMU-8613 Using a Cost-Effective Iterative Pipeline" International Journal of Molecular Sciences 27, no. 1: 509. https://doi.org/10.3390/ijms27010509
APA StyleVilla-Villa, E. M., Vázquez-Garcidueñas, M. S., & Vázquez-Marrufo, G. (2026). Hybrid Genome Assembly and Annotation of the Basidiomycete Fungus Candolleomyces candolleanus Strain CMU-8613 Using a Cost-Effective Iterative Pipeline. International Journal of Molecular Sciences, 27(1), 509. https://doi.org/10.3390/ijms27010509








