Bioinformatic Approach to Identify Positive Prognostic TGFB2-Dependent and Negative Prognostic TGFB2-Independent Biomarkers for Breast Cancers
Abstract
1. Introduction
2. Results
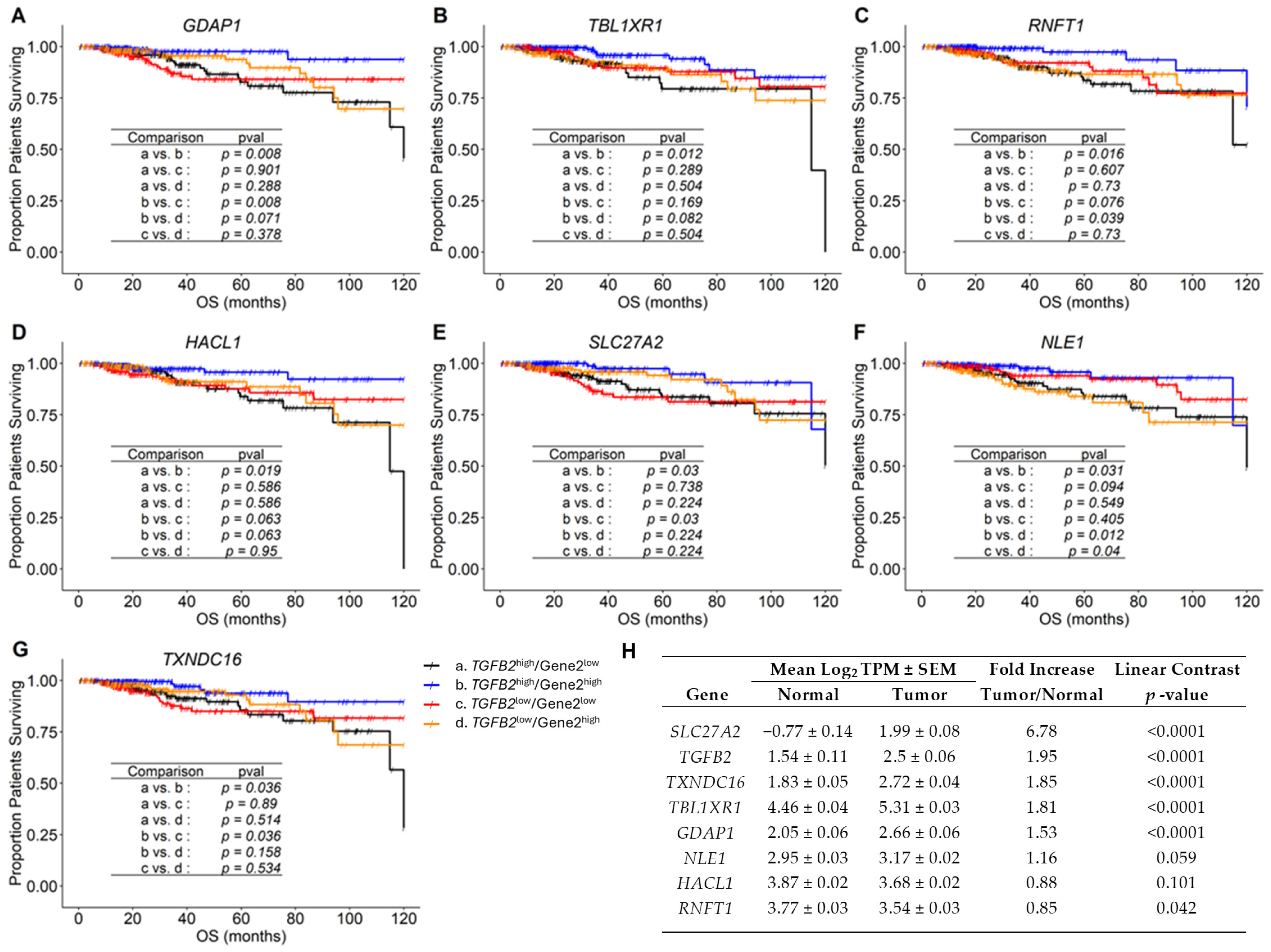
2.1. TGFB2-Dependent Positively Prognostic Gene2
2.2. TGFB2-Independent Gene2 Correlated with Increase in HR
3. Discussion
4. Materials and Methods
4.1. AI-Powered Chatbot Identified PubMed Abstracts for Manuscript Preparation
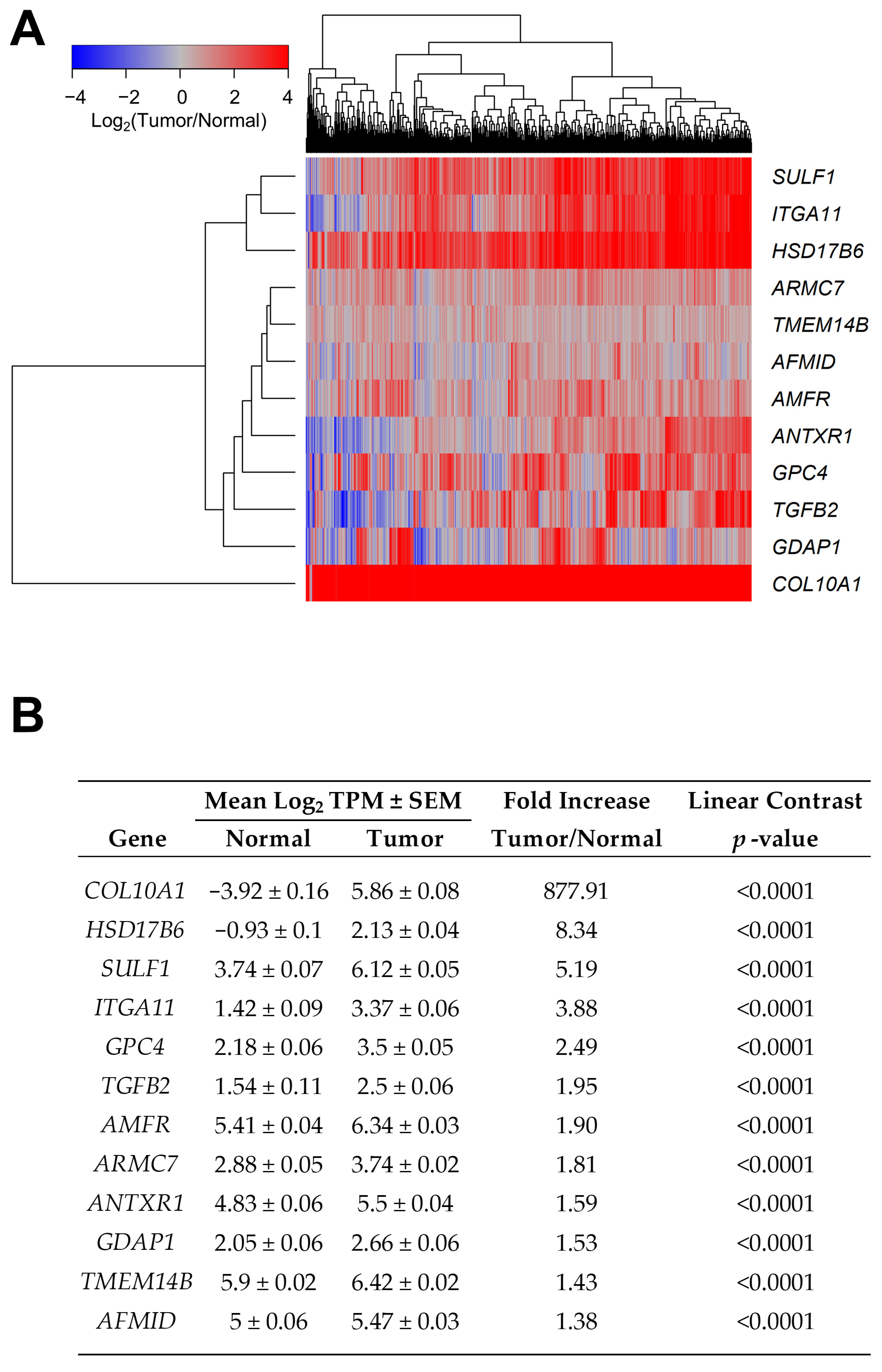
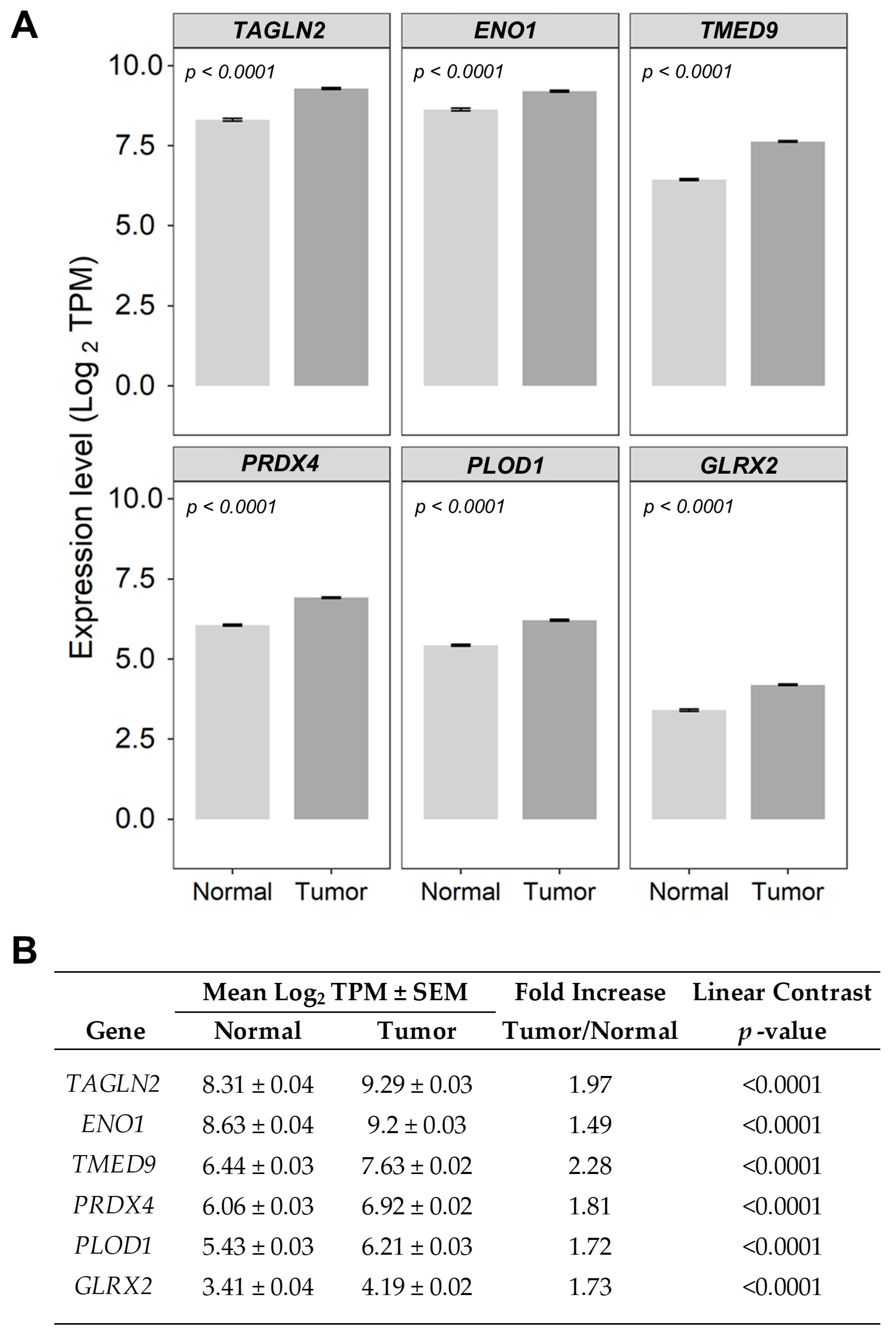
4.2. Differential Expression of mRNA Comparing Breast Cancer Tumors Versus Normal Samples
4.3. OS Outcomes for Breast Cancer
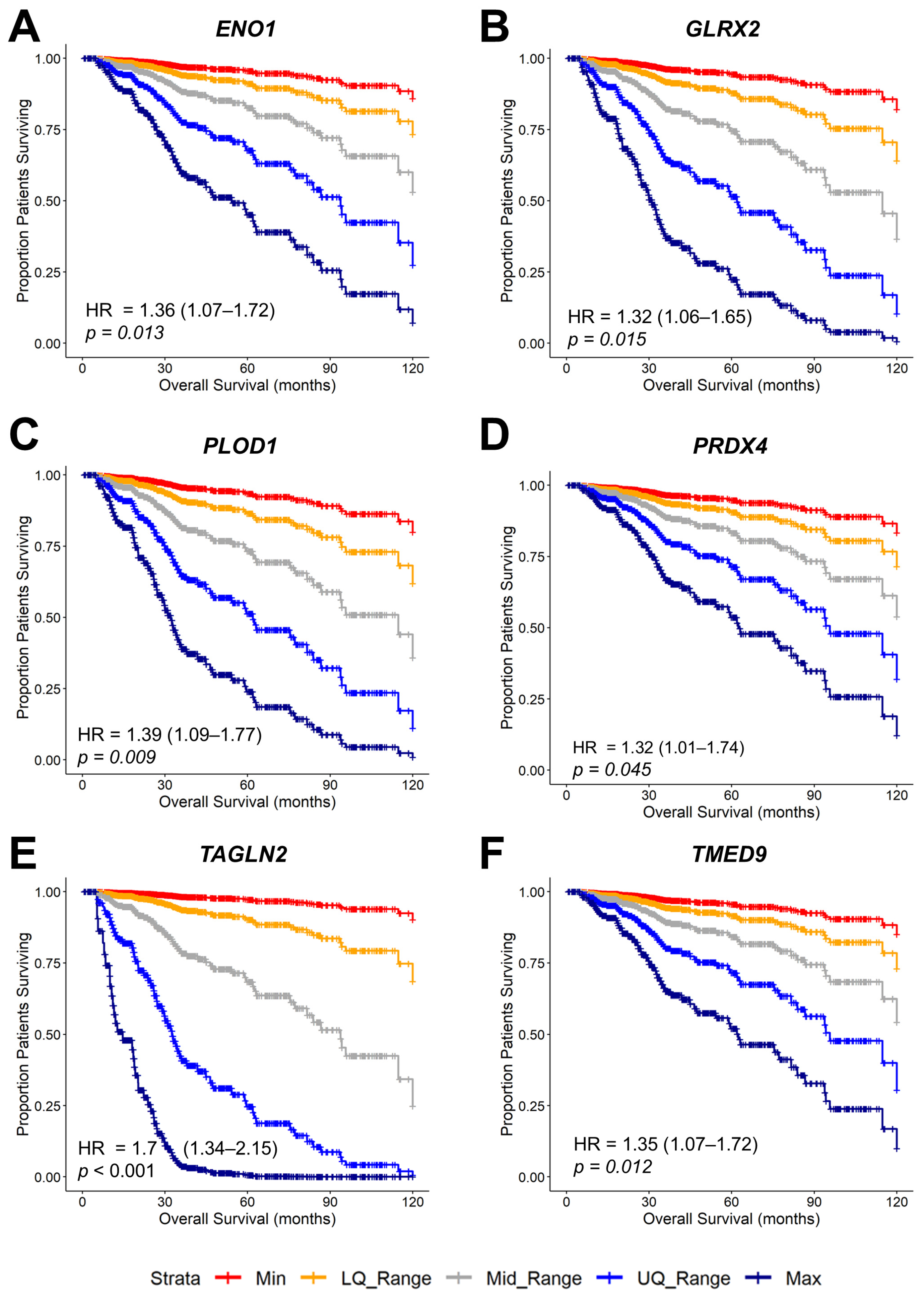
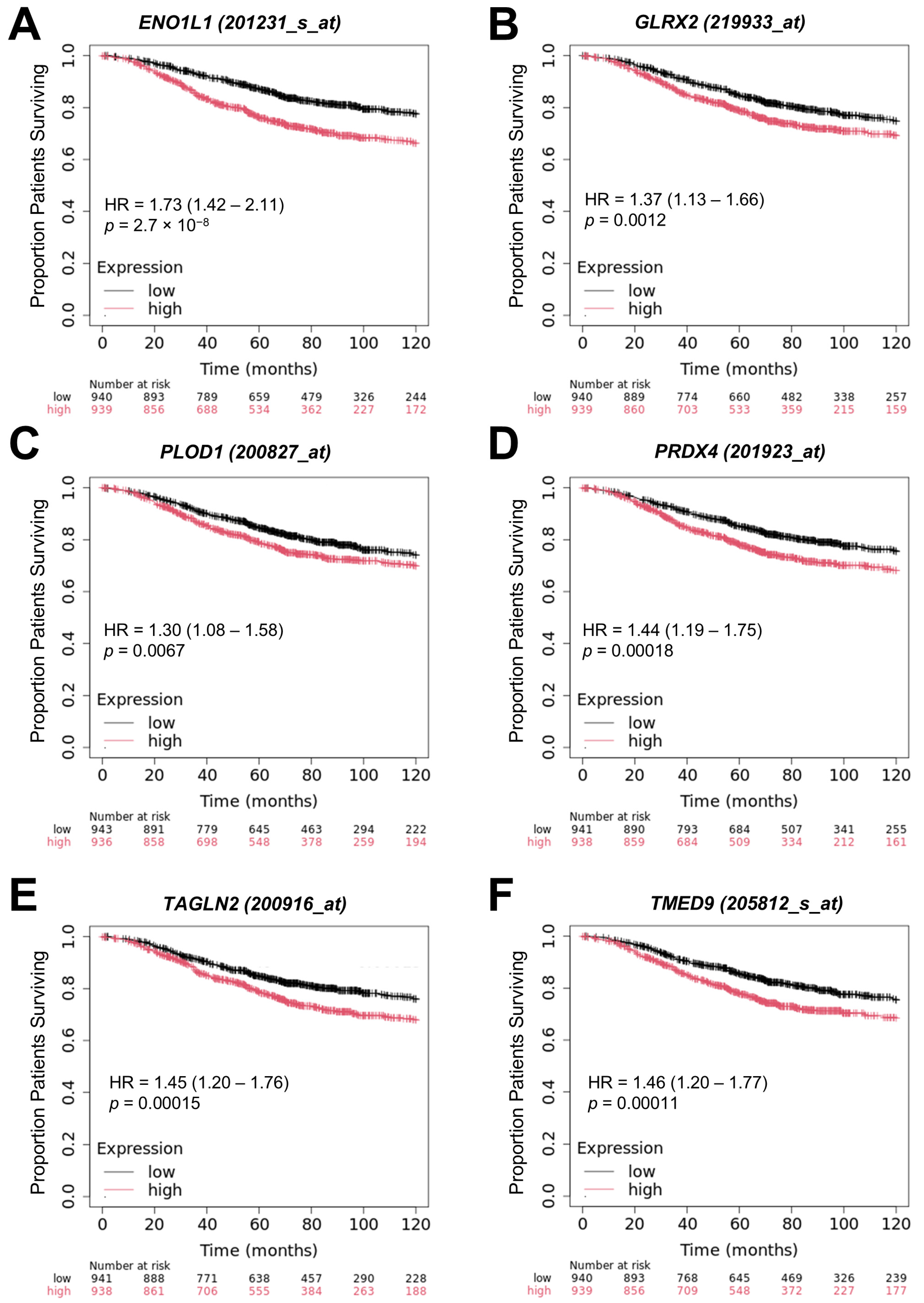
4.4. OS Analysis Using Kaplan–Meier Comparisons for Genes Identified in the TCGA Dataset
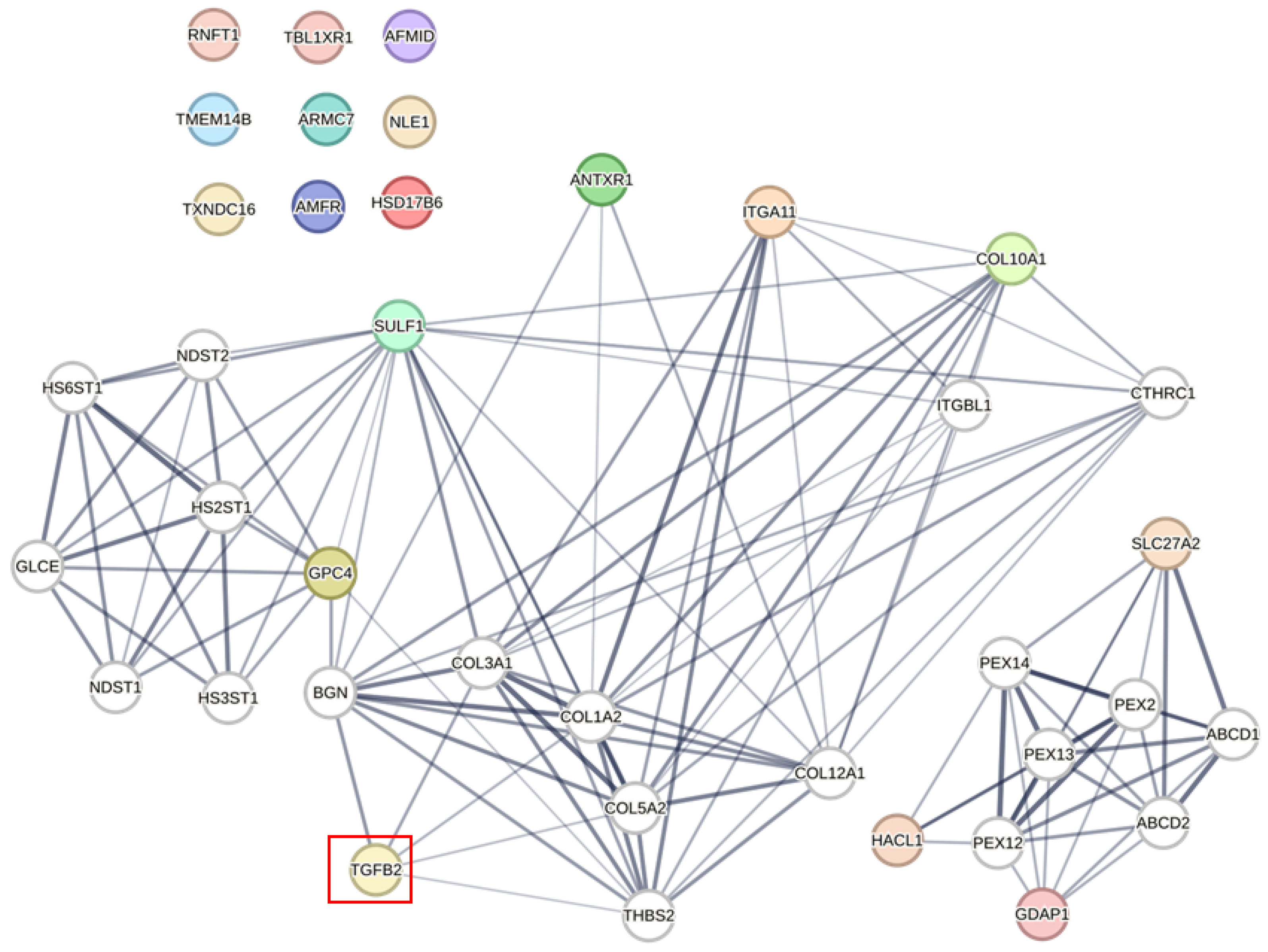
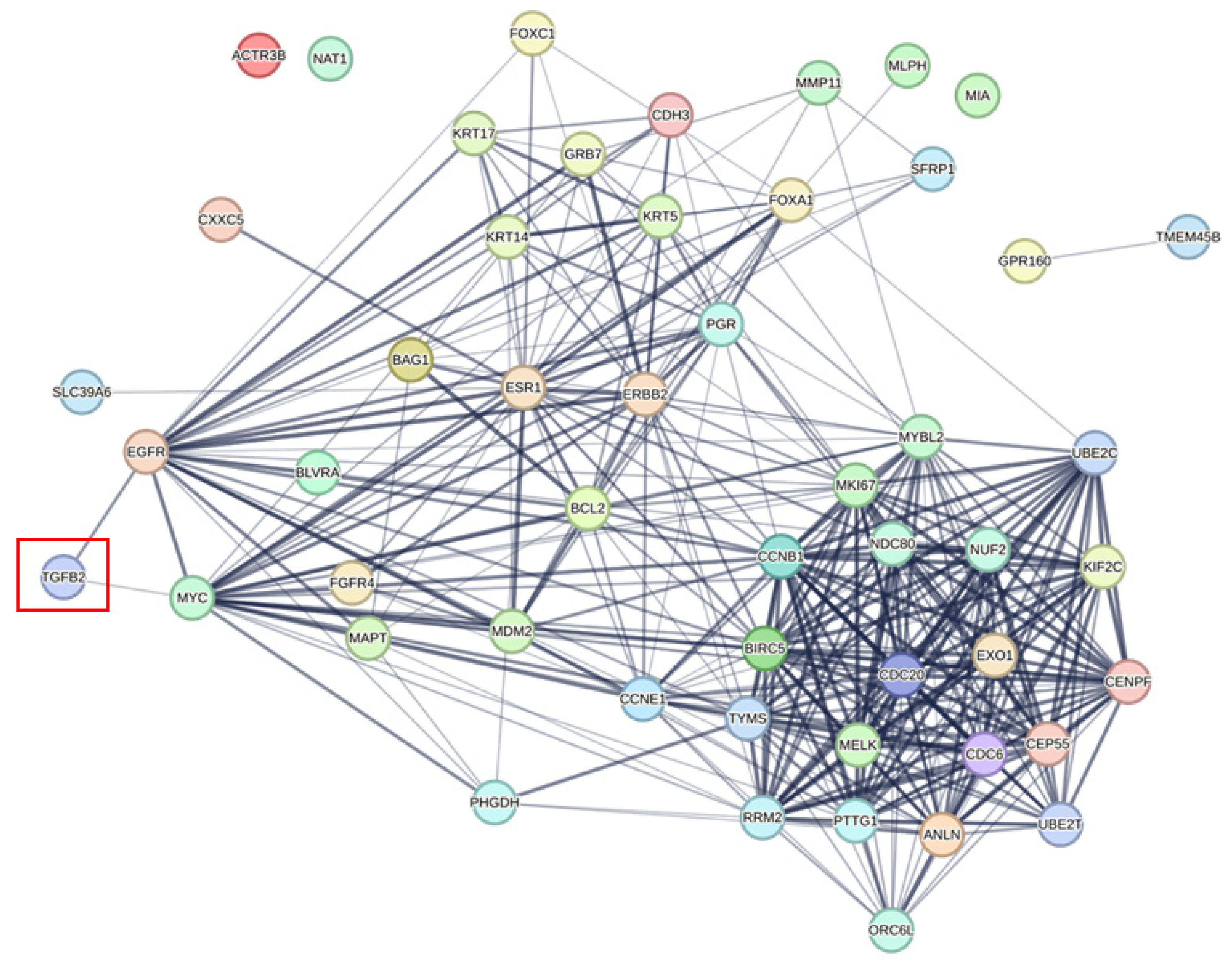
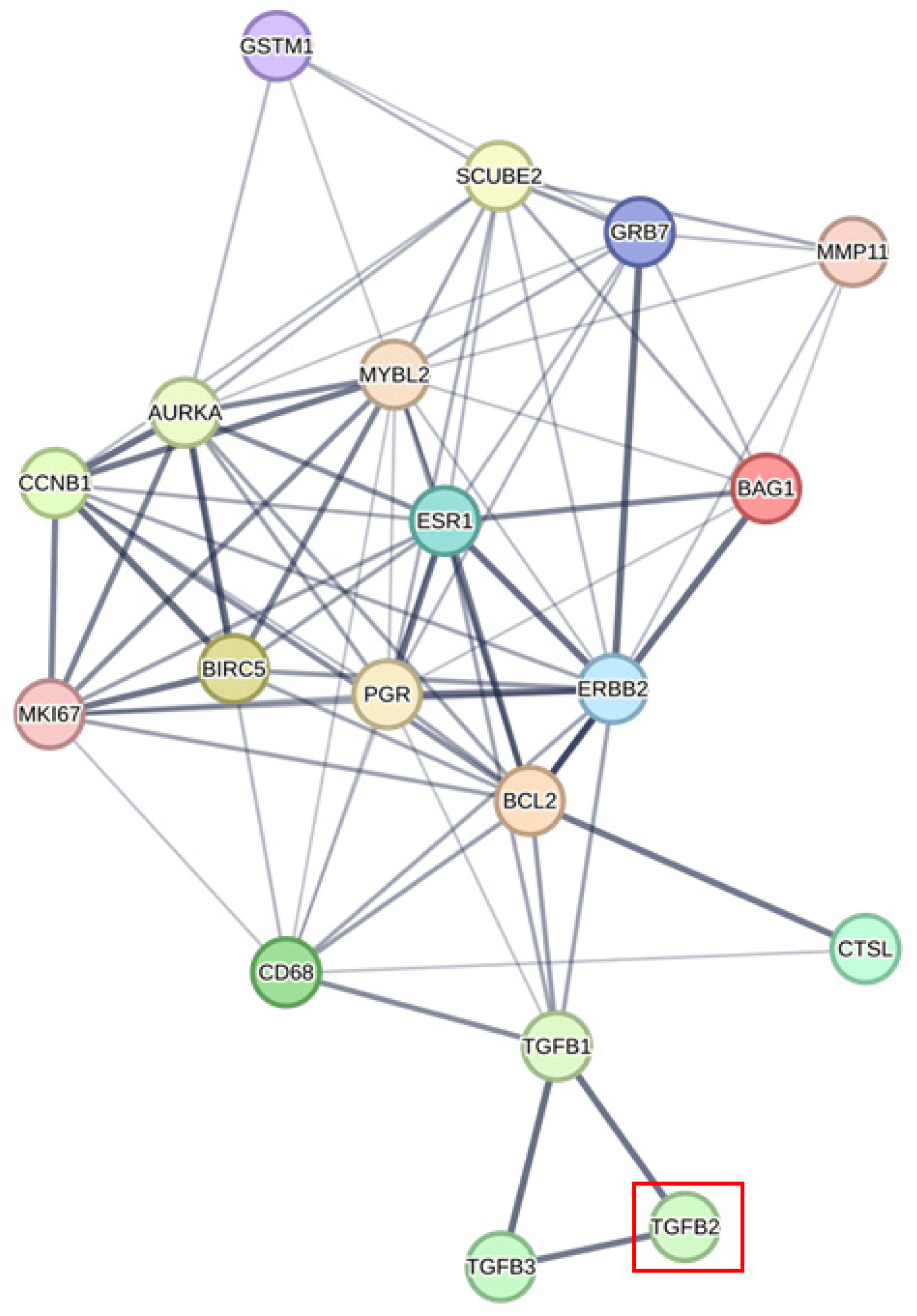
4.5. Identifying Prognostically Relevant Signaling Proteins Networked for TGFB2-Dependent Gene2, PAM50, and OncotypeDx Signature Genes Using the STRING Interaction Algorithm
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Allott, E.H.; Shan, Y.; Chen, M.; Sun, X.; Garcia-Recio, S.; Kirk, E.L.; Olshan, A.F.; Geradts, J.; Earp, H.S.; Carey, L.A.; et al. Bimodal Age Distribution at Diagnosis in Breast Cancer Persists Across Molecular and Genomic Classifications. Breast Cancer Res. Treat. 2020, 179, 185–195. [Google Scholar] [CrossRef]
- Guo, L.; Chen, G.; Zhang, W.; Zhou, L.; Xiao, T.; Di, X.; Wang, Y.; Feng, L.; Zhang, K. A High-Risk Luminal A Dominant Breast Cancer Subtype with Increased Mobility. Breast Cancer Res. Treat. 2019, 175, 459–472. [Google Scholar] [CrossRef]
- Horr, C.; Buechler, S.A. Breast Cancer Consensus Subtypes: A System for Subtyping Breast Cancer Tumors Based on Gene Expression. NPJ Breast Cancer 2021, 7, 136. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Qin, S.; Yi, Y.; Gao, H.; Liu, X.; Ma, F.; Guan, M. Delving into the Heterogeneity of Different Breast Cancer Subtypes and the Prognostic Models Utilizing ScRNA-Seq and Bulk RNA-Seq. Int. J. Mol. Sci. 2022, 23, 9936. [Google Scholar] [CrossRef]
- Ali, S.; Rehman, M.U.; Yatoo, A.M.; Arafah, A.; Khan, A.; Rashid, S.; Majid, S.; Ali, A.; Ali, N.M. TGF-β Signaling Pathway: Therapeutic Targeting and Potential for Anti-Cancer Immunity. Eur. J. Pharmacol. 2023, 947, 175678. [Google Scholar] [CrossRef]
- Cecerska-Heryć, E.; Jerzyk, A.; Goszka, M.; Polikowska, A.; Rachwalska, J.; Serwin, N.; Wojciuk, B.; Dołęgowska, B. TGF-β Signaling in Cancer: Mechanisms of Progression and Therapeutic Targets. Int. J. Mol. Sci. 2025, 26, 7326. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Mishra, L.; Deng, C.-X. The Role of TGF-β/SMAD4 Signaling in Cancer. Int. J. Biol. Sci. 2018, 14, 111–123. [Google Scholar] [CrossRef]
- Tzavlaki, K.; Moustakas, A. TGF-β Signaling. Biomolecules 2020, 10, 487. [Google Scholar] [CrossRef]
- Huang, C.-Y.; Chung, C.-L.; Hu, T.-H.; Chen, J.-J.; Liu, P.-F.; Chen, C.-L. Recent Progress in TGF-β Inhibitors for Cancer Therapy. Biomed. Pharmacother. 2021, 134, 111046. [Google Scholar] [CrossRef] [PubMed]
- Qazi, S.; Talebi, Z.; Trieu, V. Transforming Growth Factor Beta 2 (TGFB2) and Interferon Gamma Receptor 2 (IFNGR2) MRNA Levels in the Brainstem Tumor Microenvironment (TME) Significantly Impact Overall Survival in Pediatric DMG Patients. Biomedicines 2024, 12, 191. [Google Scholar] [CrossRef]
- Qazi, S.; Trieu, V. TGFB2 MRNA Levels Prognostically Interact with Interferon-Alpha Receptor Activation of IRF9 and IFI27, and an Immune Checkpoint LGALS9 to Impact Overall Survival in Pancreatic Ductal Adenocarcinoma. Int. J. Mol. Sci. 2024, 25, 11221. [Google Scholar] [CrossRef]
- Trieu, V.; Maida, A.E.; Qazi, S. Transforming Growth Factor Beta 2 (TGFB2) MRNA Levels, in Conjunction with Interferon-Gamma Receptor Activation of Interferon Regulatory Factor 5 (IRF5) and Expression of CD276/B7-H3, Are Therapeutically Targetable Negative Prognostic Markers in Low-Grade Gliomas. Cancers 2024, 16, 1202. [Google Scholar] [CrossRef]
- Figueroa, J.D.; Flanders, K.C.; Garcia-Closas, M.; Anderson, W.F.; Yang, X.R.; Matsuno, R.K.; Duggan, M.A.; Pfeiffer, R.M.; Ooshima, A.; Cornelison, R.; et al. Expression of TGF-β Signaling Factors in Invasive Breast Cancers: Relationships with Age at Diagnosis and Tumor Characteristics. Breast Cancer Res. Treat. 2010, 121, 727–735. [Google Scholar] [CrossRef]
- Soufla, G.; Porichis, F.; Sourvinos, G.; Vassilaros, S.; Spandidos, D.A. Transcriptional Deregulation of VEGF, FGF2, TGF-Β1, 2, 3 and Cognate Receptors in Breast Tumorigenesis. Cancer Lett. 2006, 235, 100–113. [Google Scholar] [CrossRef]
- Chakravarthy, D.; Green, A.R.; Green, V.L.; Kerin, M.J.; Speirs, V. Expression and Secretion of TGF-Beta Isoforms and Expression of TGF-Beta-Receptors I, II and III in Normal and Neoplastic Human Breast. Int. J. Oncol. 1999, 15, 187–281. [Google Scholar] [CrossRef] [PubMed]
- Koumoundourou, D.; Kassimatis, T.; Zolota, V.; Tzorakoeleftherakis, E.; Ravazoula, P.; Vassiliou, V.; Kardamakis, D.; Varakis, J. Prognostic Significance of TGFbeta-1 and PSmad2/3 in Breast Cancer Patients with T1-2,N0 Tumours. Anticancer Res. 2007, 27, 2613–2620. [Google Scholar] [PubMed]
- Ciftci, R.; Tas, F.; Yasasever, C.T.; Aksit, E.; Karabulut, S.; Sen, F.; Keskin, S.; Kilic, L.; Yildiz, İ.; Bozbey, H.U.; et al. High Serum Transforming Growth Factor Beta 1 (TGFB1) Level Predicts Better Survival in Breast Cancer. Tumor Biol. 2014, 35, 6941–6948. [Google Scholar] [CrossRef] [PubMed]
- Ghellal, A.; Li, C.; Hayes, M.; Byrne, G.; Bundred, N.; Kumar, S. Prognostic Significance of TGF Beta 1 and TGF Beta 3 in Human Breast Carcinoma. Anticancer Res. 2000, 20, 4413–4418. [Google Scholar]
- Auvinen, P.; Lipponen, P.; Johansson, R.; Syrjänen, K. Prognostic Significance of TGF-Beta 1 and TGF-Beta 2 Expressions in Female Breast Cancer. Anticancer Res. 1995, 15, 2627–2631. [Google Scholar]
- Huang, H.; Li, T.; Ye, G.; Zhao, L.; Zhang, Z.; Mo, D.; Wang, Y.; Zhang, C.; Deng, H.; Li, G.; et al. High Expression of COL10A1 Is Associated with Poor Prognosis in Colorectal Cancer. OncoTargets Ther. 2018, 11, 1571–1581. [Google Scholar] [CrossRef]
- Ke, T.-W.; Chang, S.-C.; Yeh, C.-M.; Lin, S.-H.; Yeh, K.-T. Comprehensive Bioinformatic Analysis Reveals Prognostic Significance and Functional Insights of Candidate Gene Expression in Colorectal Cancer. Sci. Rep. 2025, 15, 5659. [Google Scholar] [CrossRef]
- Liang, Y.; Xia, W.; Zhang, T.; Chen, B.; Wang, H.; Song, X.; Zhang, Z.; Xu, L.; Dong, G.; Jiang, F. Upregulated Collagen COL10A1 Remodels the Extracellular Matrix and Promotes Malignant Progression in Lung Adenocarcinoma. Front. Oncol. 2020, 10, 573534. [Google Scholar] [CrossRef]
- Zhang, M.; Jin, M.; Gao, Z.; Yu, W.; Zhang, W. High COL10A1 Expression Potentially Contributes to Poor Outcomes in Gastric Cancer with the Help of LEF1 and Wnt2. J. Clin. Lab. Anal. 2022, 36, e24612. [Google Scholar] [CrossRef]
- Xu, Q.; Zheng, J.; Su, Z.; Chen, B.; Gu, S. COL10A1 Promotes Tumorigenesis by Modulating CD276 in Pancreatic Adenocarcinoma. BMC Gastroenterol. 2023, 23, 397. [Google Scholar] [CrossRef]
- Bret, C.; Moreaux, J.; Schved, J.-F.; Hose, D.; Klein, B. SULFs in Human Neoplasia: Implication as Progression and Prognosis Factors. J. Transl. Med. 2011, 9, 72. [Google Scholar] [CrossRef]
- Zhang, B.; Luo, D.; Xiang, L.; Chen, J.; Fang, T. Investigating the Anti-Cancer Potential of Sulfatase 1 and Its Underlying Mechanism in Non-Small Cell Lung Cancer. Cytojournal 2024, 21, 52. [Google Scholar] [CrossRef] [PubMed]
- Dhanasekaran, R.; Nakamura, I.; Hu, C.; Chen, G.; Oseini, A.M.; Seven, E.S.; Miamen, A.G.; Moser, C.D.; Zhou, W.; van Kuppevelt, T.H.; et al. Activation of the Transforming Growth Factor-β/SMAD Transcriptional Pathway Underlies a Novel Tumor-promoting Role of Sulfatase 1 in Hepatocellular Carcinoma. Hepatology 2015, 61, 1269–1283. [Google Scholar] [CrossRef]
- Fang, X.; Chen, D.; Yang, X.; Cao, X.; Cheng, Q.; Liu, K.; Xu, P.; Wang, Y.; Xu, J.; Zhao, S.; et al. Cancer Associated Fibroblasts-Derived SULF1 Promotes Gastric Cancer Metastasis and CDDP Resistance through the TGFBR3-Mediated TGF-β Signaling Pathway. Cell Death Discov. 2024, 10, 111. [Google Scholar] [CrossRef]
- Yang, Y.; Ahn, J.; Edwards, N.J.; Benicky, J.; Rozeboom, A.M.; Davidson, B.; Karamboulas, C.; Nixon, K.C.J.; Ailles, L.; Goldman, R. Extracellular Heparan 6-O-Endosulfatases SULF1 and SULF2 in Head and Neck Squamous Cell Carcinoma and Other Malignancies. Cancers 2022, 14, 5553. [Google Scholar] [CrossRef] [PubMed]
- Lyu, Y.; Cheng, Y.; Wang, B.; Chen, L.; Zhao, S. Sulfatase 1 Expression in Pancreatic Cancer and Its Correlation with Clinicopathological Features and Postoperative Prognosis. Cancer Biomark. 2018, 22, 701–707. [Google Scholar] [CrossRef] [PubMed]
- Hammond, E.; Khurana, A.; Shridhar, V.; Dredge, K. The Role of Heparanase and Sulfatases in the Modification of Heparan Sulfate Proteoglycans within the Tumor Microenvironment and Opportunities for Novel Cancer Therapeutics. Front. Oncol. 2014, 4, 195. [Google Scholar] [CrossRef] [PubMed]
- Khurana, A.; Liu, P.; Mellone, P.; Lorenzon, L.; Vincenzi, B.; Datta, K.; Yang, B.; Linhardt, R.J.; Lingle, W.; Chien, J.; et al. HSulf-1 Modulates FGF2- and Hypoxia-Mediated Migration and Invasion of Breast Cancer Cells. Cancer Res. 2011, 71, 2152–2161. [Google Scholar] [CrossRef] [PubMed]
- Staub, J.; Chien, J.; Pan, Y.; Qian, X.; Narita, K.; Aletti, G.; Scheerer, M.; Roberts, L.R.; Molina, J.; Shridhar, V. Epigenetic Silencing of HSulf-1 in Ovarian Cancer:Implications in Chemoresistance. Oncogene 2007, 26, 4969–4978. [Google Scholar] [CrossRef]
- He, X.; Khurana, A.; Roy, D.; Kaufmann, S.; Shridhar, V. Loss of HSulf-1 Expression Enhances Tumorigenicity by Inhibiting Bim Expression in Ovarian Cancer. Int. J. Cancer 2014, 135, 1783–1789. [Google Scholar] [CrossRef]
- Yang, X.; Wei, M.; Huang, Y.; Yang, X.; Yuan, Z.; Huang, J.; Wei, J.; Tian, L. ITGA11, a Prognostic Factor Associated with Immunity in Gastric Adenocarcinoma. Int. J. Gen. Med. 2024, 17, 471–483. [Google Scholar] [CrossRef]
- Ando, T.; Kage, H.; Matsumoto, Y.; Zokumasu, K.; Yotsumoto, T.; Maemura, K.; Amano, Y.; Watanabe, K.; Nakajima, J.; Nagase, T.; et al. Integrin A11 in Non–Small Cell Lung Cancer Is Associated with Tumor Progression and Postoperative Recurrence. Cancer Sci. 2020, 111, 200–208. [Google Scholar] [CrossRef]
- Primac, I.; Maquoi, E.; Blacher, S.; Heljasvaara, R.; Van Deun, J.; Smeland, H.Y.H.; Canale, A.; Louis, T.; Stuhr, L.; Sounni, N.E.; et al. Stromal Integrin A11 Regulates PDGFRβ Signaling and Promotes Breast Cancer Progression. J. Clin. Investig. 2019, 129, 4609–4628. [Google Scholar] [CrossRef]
- Talia, M.; Cesario, E.; Cirillo, F.; Scordamaglia, D.; Di Dio, M.; Zicarelli, A.; Mondino, A.A.; Occhiuzzi, M.A.; De Francesco, E.M.; Belfiore, A.; et al. Cancer-Associated Fibroblasts (CAFs) Gene Signatures Predict Outcomes in Breast and Prostate Tumor Patients. J. Transl. Med. 2024, 22, 597. [Google Scholar] [CrossRef]
- Li, M.; Tian, X.; Guo, H.; Xu, X.; Liu, Y.; Hao, X.; Fei, H. A Novel LncRNA-MRNA-MiRNA Signature Predicts Recurrence and Disease-Free Survival in Cervical Cancer. Braz. J. Med. Biol. Res. 2021, 54, e11592. [Google Scholar] [CrossRef] [PubMed]
- Decruyenaere, P.; Giuili, E.; Verniers, K.; Anckaert, J.; De Grove, K.; Van der Linden, M.; Deeren, D.; Van Dorpe, J.; Offner, F.; Vandesompele, J. Exploring the Cell-Free Total RNA Transcriptome in Diffuse Large B-Cell Lymphoma and Primary Mediastinal B-Cell Lymphoma Patients as Biomarker Source in Blood Plasma Liquid Biopsies. Front. Oncol. 2023, 13, 1221471. [Google Scholar] [CrossRef]
- Takanami, I.; Takeuchi, K.; Watanabe, H.; Yanagawa, T.; Takagishi, K.; Raz, A. Significance Ofautocrine Motility Factor Receptor Gene Expression as a Prognostic Factor in Non-Small-Cell Lung Cancer. Int. J. Cancer 2001, 95, 384–387. [Google Scholar] [CrossRef]
- Huang, Z.; Zhang, N.; Zha, L.; Mao, H.-C.; Chen, X.; Xiang, J.-F.; Zhang, H.; Wang, Z.-W. Aberrant Expression of the Autocrine Motility Factor Receptor Correlates with Poor Prognosis and Promotes Metastasis in Gastric Carcinoma. Asian Pac. J. Cancer Prev. 2014, 15, 989–997. [Google Scholar] [CrossRef][Green Version]
- Jiang, W.G.; Raz, A.; Douglas-Jones, A.; Mansel, R.E. Expression of Autocrine Motility Factor (AMF) and Its Receptor, AMFR, in Human Breast Cancer. J. Histochem. Cytochem. 2006, 54, 231–241. [Google Scholar] [CrossRef]
- Halwani, Y.; Kojic, L.D.; Chan, S.K.; Phang, T.P.; Masoudi, H.; Jones, S.J.M.; Nabi, I.R.; Wiseman, S.M. Prognostic Significance of Autocrine Motility Factor Receptor Expression by Colorectal Cancer and Lymph Node Metastases. Am. J. Surg. 2015, 209, 884–889. [Google Scholar] [CrossRef]
- Marco, A. Evolutionary and Structural Analyses of GDAP1, Involved in Charcot-Marie-Tooth Disease, Characterize a Novel Class of Glutathione Transferase-Related Genes. Mol. Biol. Evol. 2003, 21, 176–187. [Google Scholar] [CrossRef] [PubMed]
- Magouliotis, D.E.; Sakellaridis, N.; Dimas, K.; Tasiopoulou, V.S.; Svokos, K.A.; Svokos, A.A.; Zacharoulis, D. In Silico Transcriptomic Analysis of the Chloride Intracellular Channels (CLIC) Interactome Identifies a Molecular Panel of Seven Prognostic Markers in Patients with Pancreatic Ductal Adenocarcinoma. Curr. Genom. 2020, 21, 119–127. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Chen, C.; Hu, Y.; Li, K.; Zhang, Y.; Chen, Z.; Nie, D.; Gao, R.; Huang, Y.; Zhong, M.; et al. High Expression of LOC541471, GDAP1, SOD1, and STK25 Is Associated with Poor Overall Survival of Patients with Acute Myeloid Leukemia. Cancer Med. 2023, 12, 9055–9067. [Google Scholar] [CrossRef]
- Shen, J.; Shu, M.; Xie, S.; Yan, J.; Pan, K.; Chen, S.; Li, X. A Six-Gene Prognostic Risk Prediction Model In Hepatitis B Virus-Associated Hepatocellular Carcinoma. Clin. Investig. Med. 2021, 44, E32–E44. [Google Scholar] [CrossRef]
- Wang, Y.; Chen, X.; Li, Y.; Zhang, Z.; Xia, L.; Jiang, J.; Chai, Y.; Wang, Z.; Wan, Y.; Li, T.; et al. SLC27A2 Is a Potential Immune Biomarker for Hematological Tumors and Significantly Regulates the Cell Cycle Progression of Diffuse Large B-Cell Lymphoma. BMC Med. Genom. 2024, 17, 105. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Zhang, Y.; Chang, K.; Hou, N.; Fan, P.; Ji, C.; Liu, L.; Wang, Z.; Li, R.; Wang, Y.; et al. Risk Assessment Model Based on Nucleotide Metabolism-Related Genes Highlights SLC27A2 as a Potential Therapeutic Target in Breast Cancer. J. Cancer Res. Clin. Oncol. 2024, 150, 258. [Google Scholar] [CrossRef]
- Feng, K.; Ma, R.; Li, H.; Yin, K.; Du, G.; Chen, X.; Liu, Z.; Yin, D. Upregulated SLC27A2/FATP2 in Differentiated Thyroid Carcinoma Promotes Tumor Proliferation and Migration. J. Clin. Lab. Anal. 2022, 36, e24148. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.; Li, J.; Zheng, Y.; Luo, L.; Huang, Y.; Hu, J.; Chen, Y. High Expression of SLC27A2 Predicts Unfavorable Prognosis and Promotes Inhibitory Immune Infiltration in Acute Lymphoblastic Leukemia. Transl. Oncol. 2024, 45, 101952. [Google Scholar] [CrossRef]
- Xu, N.; Xiao, W.; Meng, X.; Li, W.; Wang, X.; Zhang, X.; Yang, H. Up-Regulation of SLC27A2 Suppresses the Proliferation and Invasion of Renal Cancer by Down-Regulating CDK3-Mediated EMT. Cell Death Discov. 2022, 8, 351. [Google Scholar] [CrossRef]
- Kocatürk, B. Identification of Thioredoxin Domain Containing Family Members’ Expression Pattern and Prognostic Value in Diffuse Gliomas via in Silico Analysis. Cancer Med. 2023, 12, 3830–3844. [Google Scholar] [CrossRef]
- Li, X.; Liang, W.; Liu, J.; Lin, C.; Wu, S.; Song, L.; Yuan, Z. Transducin (β)-like 1 X-Linked Receptor 1 Promotes Proliferation and Tumorigenicity in Human Breast Cancer via Activation of Beta-Catenin Signaling. Breast Cancer Res. 2014, 16, 465. [Google Scholar] [CrossRef] [PubMed]
- Du, R.; Li, K.; Guo, K.; Chen, Z.; Zhao, X.; Han, L.; Bian, H. Two Decades of a Protooncogene TBL1XR1: From a Transcription Modulator to Cancer Therapeutic Target. Front. Oncol. 2024, 14, 1309687. [Google Scholar] [CrossRef] [PubMed]
- Ren, Z.; Ni, F.; Zhang, T.; Yuan, X.; Li, J.; Xiao, W. Knockdown of NLE1 Inhibits Development of Malignant Melanoma In Vitro and In Vivo NLE1 Promotes Development of Malignant Melanoma. Exp. Cell Res. 2021, 404, 112636, Erratum in Exp. Cell Res. 2021, 405, 112733. [Google Scholar] [CrossRef]
- Wang, Y.; Xu, Z. The Effect of Up-Regulation of NLE1 Gene Expression on the Invasion and Migration of Colon Cancer Cells and Its Mechanism. J. King Saud Univ. Sci. 2023, 35, 102669. [Google Scholar] [CrossRef]
- Xu, P.; Wang, L.; Mo, B.; Xie, X.; Hu, R.; Jiang, L.; Hu, F.; Ding, F.; Xiao, H. Identification of NLE1/CDK1 Axis as Key Regulator in the Development and Progression of Non-Small Cell Lung Cancer. Front. Oncol. 2023, 12, 985827. [Google Scholar] [CrossRef]
- Zhang, R.; Liu, D.; Feng, X.; Yang, Z.; Zhang, K. WD40 Protein NLE1 as a Novel Diagnostic Biomarker Promoting Hepatocellular Carcinoma Proliferation. Clin. Med. Insights Oncol. 2025, 19, 11795549251348902. [Google Scholar] [CrossRef]
- Ungefroren, H.; Lenschow, W.; Chen, W.-B.; Faendrich, F.; Kalthoff, H. Regulation of Biglycan Gene Expression by Transforming Growth Factor-β Requires MKK6-P38 Mitogen-Activated Protein Kinase Signaling Downstream of Smad Signaling. J. Biol. Chem. 2003, 278, 11041–11049. [Google Scholar] [CrossRef] [PubMed]
- Ungefroren, H.; Groth, S.; Ruhnke, M.; Kalthoff, H.; Fändrich, F. Transforming Growth Factor-β (TGF-β) Type I Receptor/ALK5-Dependent Activation of the GADD45β Gene Mediates the Induction of Biglycan Expression by TGF-β. J. Biol. Chem. 2005, 280, 2644–2652. [Google Scholar] [CrossRef]
- Verrecchia, F.; Chu, M.-L.; Mauviel, A. Identification of Novel TGF-β/Smad Gene Targets in Dermal Fibroblasts Using a Combined CDNA Microarray/Promoter Transactivation Approach. J. Biol. Chem. 2001, 276, 17058–17062. [Google Scholar] [CrossRef]
- Zhao, L.; Wang, P.; Sun, L.; Ma, W.; Yu, L. SP1/COL1A2/ZEB1 Axis Promotes TGF-Β2-Induced Lens Epithelial Cell Proliferation, Migration, Invasion and EMT Process. Exp. Eye Res. 2025, 251, 110220. [Google Scholar] [CrossRef]
- Han, Y.-L.; Luo, D.; Habaxi, K.; Tayierjiang, J.; Zhao, W.; Wang, W.; Aikebaier, W.; Wang, L. COL5A2 Inhibits the TGF-β and Wnt/β-Catenin Signaling Pathways to Inhibit the Invasion and Metastasis of Osteosarcoma. Front. Oncol. 2022, 12, 813809. [Google Scholar] [CrossRef]
- Nan, P.; Dong, X.; Bai, X.; Lu, H.; Liu, F.; Sun, Y.; Zhao, X. Tumor-Stroma TGF-Β1-THBS2 Feedback Circuit Drives Pancreatic Ductal Adenocarcinoma Progression via Integrin Avβ3/CD36-Mediated Activation of the MAPK Pathway. Cancer Lett. 2022, 528, 59–75. [Google Scholar] [CrossRef]
- Bastien, R.R.; Rodríguez-Lescure, Á.; Ebbert, M.T.; Prat, A.; Munárriz, B.; Rowe, L.; Miller, P.; Ruiz-Borrego, M.; Anderson, D.; Lyons, B.; et al. PAM50 Breast Cancer Subtyping by RT-QPCR and Concordance with Standard Clinical Molecular Markers. BMC Med. Genom. 2012, 5, 44. [Google Scholar] [CrossRef]
- Ohnstad, H.O.; Borgen, E.; Falk, R.S.; Lien, T.G.; Aaserud, M.; Sveli, M.A.T.; Kyte, J.A.; Kristensen, V.N.; Geitvik, G.A.; Schlichting, E.; et al. Prognostic Value of PAM50 and Risk of Recurrence Score in Patients with Early-Stage Breast Cancer with Long-Term Follow-Up. Breast Cancer Res. 2017, 19, 120. [Google Scholar] [CrossRef] [PubMed]
- Cichon, M.A.; Moruzzi, M.E.; Shqau, T.A.; Miller, E.; Mehner, C.; Ethier, S.P.; Copland, J.A.; Radisky, E.S.; Radisky, D.C. MYC Is a Crucial Mediator of TGFβ-Induced Invasion in Basal Breast Cancer. Cancer Res. 2016, 76, 3520–3530. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Ma, J.; Fan, Y.; Wang, Z.; Tian, R.; Ji, W.; Zhang, F.; Niu, R. TGF-β Transactivates EGFR and Facilitates Breast Cancer migration and Invasion Through Canonical Smad3 and ERK/Sp1 Signaling Pathways. Mol. Oncol. 2018, 12, 305–321. [Google Scholar] [CrossRef]
- Kalinsky, K.; Barlow, W.E.; Gralow, J.R.; Meric-Bernstam, F.; Albain, K.S.; Hayes, D.F.; Lin, N.U.; Perez, E.A.; Goldstein, L.J.; Chia, S.K.L.; et al. 21-Gene Assay to Inform Chemotherapy Benefit in Node-Positive Breast Cancer. N. Engl. J. Med. 2021, 385, 2336–2347. [Google Scholar] [CrossRef]
- Sun, X.; Bernhardt, S.M.; Glynn, D.J.; Hodson, L.J.; Woolford, L.; Evdokiou, A.; Yan, C.; Du, H.; Robertson, S.A.; Ingman, W.V. Attenuated TGFB Signalling in Macrophages Decreases Susceptibility to DMBA-Induced Mammary Cancer in Mice. Breast Cancer Res. 2021, 23, 39. [Google Scholar] [CrossRef]
- Chow, A.; Arteaga, C.L.; Wang, S.E. When Tumor Suppressor TGFβ Meets the HER2 (ERBB2) Oncogene. J. Mammary Gland. Biol. Neoplasia 2011, 16, 81–88. [Google Scholar] [CrossRef]
- Shi, Y.-Y.; Chen, X.-L.; Chen, Q.-X.; Yang, Y.-Z.; Zhou, M.; Ren, Y.-X.; Tang, L.-Y.; Ren, Z.-F. Association of Enolase-1 with Prognosis and Immune Infiltration in Breast Cancer by Clinical Stage. J. Inflamm. Res. 2023, 16, 493–503. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Liao, L.; An, C.; Wang, X.; Li, Z.; Xu, Z.; Liu, J.; Liu, S. α-Enolase Lies Downstream of MTOR/HIF1α and Promotes Thyroid Carcinoma Progression by Regulating CST1. Front. Cell Dev. Biol. 2021, 9, 670019. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; Yang, W.; Wu, C.; Ma, X.; Li, H.; Zheng, J. Enolase 1 Correlated with Cancer Progression and Immune-Infiltrating in Multiple Cancer Types: A Pan-Cancer Analysis. Front. Oncol. 2021, 10, 593706. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.K.; Sun, Y.; Lv, L.; Ping, Y. ENO1 and Cancer. Mol. Ther. Oncolytics 2022, 24, 288–298. [Google Scholar] [CrossRef]
- Almaguel, F.A.; Sanchez, T.W.; Ortiz-Hernandez, G.L.; Casiano, C.A. Alpha-Enolase: Emerging Tumor-Associated Antigen, Cancer Biomarker, and Oncotherapeutic Target. Front. Genet. 2021, 11, 614726. [Google Scholar] [CrossRef]
- Novelli, F. Alpha-Enolase i ENO1 i a Potential Target in Novel Immunotherapies. Front. Biosci. 2017, 22, 4526. [Google Scholar] [CrossRef]
- Chen, M.-L.; Yuan, T.-T.; Chuang, C.-F.; Huang, Y.-T.; Chung, I.-C.; Huang, W.-C. A Novel Enolase-1 Antibody Targets Multiple Interacting Players in the Tumor Microenvironment of Advanced Prostate Cancer. Mol. Cancer Ther. 2022, 21, 1337–1347. [Google Scholar] [CrossRef]
- Li, Y.; Liu, L.; Li, B. Role of ENO1 and Its Targeted Therapy in Tumors. J. Transl. Med. 2024, 22, 1025. [Google Scholar] [CrossRef]
- Lin, Y.; Zhang, W.; Liu, L.; Li, W.; Li, Y.; Li, B. ENO1 Promotes OSCC Migration and Invasion by Orchestrating IL-6 Secretion from Macrophages via a Positive Feedback Loop. Int. J. Mol. Sci. 2023, 24, 737. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Chen, B.; Zhu, S.; Zhang, J.; He, X.; Cao, G.; Chen, B. Hyperglycemia Promotes Snail-Induced Epithelial–Mesenchymal Transition of Gastric Cancer via Activating ENO1 Expression. Cancer Cell Int. 2019, 19, 344. [Google Scholar] [CrossRef]
- Zhao, M.; Fang, W.; Wang, Y.; Guo, S.; Shu, L.; Wang, L.; Chen, Y.; Fu, Q.; Liu, Y.; Hua, S.; et al. Enolase-1 Is a Therapeutic Target in Endometrial Carcinoma. Oncotarget 2015, 6, 15610–15627. [Google Scholar] [CrossRef]
- Ji, H.; Wang, J.; Guo, J.; Li, Y.; Lian, S.; Guo, W.; Yang, H.; Kong, F.; Zhen, L.; Guo, L.; et al. Progress in the Biological Function of Alpha-Enolase. Anim. Nutr. 2016, 2, 12–17. [Google Scholar] [CrossRef]
- Chung, I.-C.; Huang, W.-C.; Huang, Y.-T.; Chen, M.-L.; Tsai, A.-W.; Wu, P.-Y.; Yuan, T.-T. Unrevealed Roles of Extracellular Enolase-1 (ENO1) in Promoting Glycolysis and Pro-Cancer Activities in Multiple Myeloma via Hypoxia-Inducible Factor 1α. Oncol. Rep. 2023, 50, 205. [Google Scholar] [CrossRef]
- Song, Q.; Zhang, K.; Sun, T.; Xu, C.; Zhao, W.; Zhang, Z. Knockout of ENO1 Leads to Metabolism Reprogramming and Tumor Retardation in Pancreatic Cancer. Front. Oncol. 2023, 13, 1119886. [Google Scholar] [CrossRef]
- Su, Z.; You, L.; He, Y.; Chen, J.; Zhang, G.; Liu, Z. Multi-Omics Reveals the Role of ENO1 in Bladder Cancer and Constructs an Epithelial-Related Prognostic Model to Predict Prognosis and Efficacy. Sci. Rep. 2024, 14, 2189. [Google Scholar] [CrossRef] [PubMed]
- Jung, D.-W.; Kim, W.-H.; Park, S.-H.; Lee, J.; Kim, J.; Su, D.; Ha, H.-H.; Chang, Y.-T.; Williams, D.R. A Unique Small Molecule Inhibitor of Enolase Clarifies Its Role in Fundamental Biological Processes. ACS Chem. Biol. 2013, 8, 1271–1282. [Google Scholar] [CrossRef] [PubMed]
- Shen, C.; Liu, J.; Hu, D.; Liu, C.; Xie, F.; Wang, Y. Tumor-Intrinsic ENO1 Inhibition Promotes Antitumor Immune Response and Facilitates the Efficacy of Anti-PD-L1 Immunotherapy in Bladder Cancer. J. Exp. Clin. Cancer Res. 2025, 44, 207. [Google Scholar] [CrossRef]
- León-Letelier, R.A.; Sevillano-Mantas, A.M.; Chen, Y.; Park, S.; Vykoukal, J.; Fahrmann, J.F.; Ostrin, E.J.; Garrett, C.; Dou, R.; Cai, Y.; et al. Citrullinated ENO1 Vaccine Enhances PD-1 Blockade in Mice Implanted with Murine Triple-Negative Breast Cancer Cells. Vaccines 2025, 13, 629. [Google Scholar] [CrossRef]
- Li, J.; Tang, X.; Wen, X.; Ren, X.; Zhang, H.; Du, Y.; Lu, J. Mitochondrial Glrx2 Knockout Augments Acetaminophen-Induced Hepatotoxicity in Mice. Antioxidants 2022, 11, 1643. [Google Scholar] [CrossRef]
- Brzozowa-Zasada, M.; Piecuch, A.; Bajdak-Rusinek, K.; Gołąbek, K.; Michalski, M.; Janelt, K.; Matysiak, N. Glutaredoxin 2 Protein (Grx2) as an Independent Prognostic Factor Associated with the Survival of Colon Adenocarcinoma Patients. Int. J. Mol. Sci. 2024, 25, 1060. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Wang, Z.; Wang, T. Machine-Learning Prediction of a Novel Diagnostic Model Using Mitochondria-Related Genes for Patients with Bladder Cancer. Sci. Rep. 2024, 14, 9282. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.; Li, G.; Zhai, Y.; Huang, L.; Feng, Y.; Wang, D.; Zhang, W.; Hu, H. Redox Regulator GLRX Is Associated with Tumor Immunity in Glioma. Front. Immunol. 2020, 11, 580934. [Google Scholar] [CrossRef]
- Jiang, H.; Guo, W.; Yuan, S.; Song, L. PLOD1 Is a Prognostic Biomarker and Mediator of Proliferation and Invasion in Osteosarcoma. Biomed. Res. Int. 2020, 2020, 3418398. [Google Scholar] [CrossRef]
- Wang, Z.; Shi, Y.; Ying, C.; Jiang, Y.; Hu, J. Hypoxia-Induced PLOD1 Overexpression Contributes to the Malignant Phenotype of Glioblastoma via NF-ΚB Signaling. Oncogene 2021, 40, 1458–1475. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Wu, Y.; Su, X. PLOD1 Promotes Cell Growth and Aerobic Glycolysis by Regulating the SOX9/PI3K/Akt/MTOR Signaling Pathway in Gastric Cancer. Front. Biosci. 2021, 26, 322–334. [Google Scholar] [CrossRef]
- Yang, Z.; Zhou, Z.; Yang, M. Lysyl Hydroxylase PLOD1 Enhances Actin Network to Promote Confined Migration of Hepatocellular Carcinoma Cells via Binding with Septin2. J. Clin. Oncol. 2022, 40, e16111. [Google Scholar] [CrossRef]
- Wang, D.-D.; Li, L.; Fu, Y.-Q.; Yang, S.-J.; Chen, X.; Hou, J.-C.; Zhang, Q.; Tian, X.-X.; Tang, J.-H.; Zhang, J.; et al. Systematic Characterization of the Expression, Prognosis and Immune Characteristics of PLOD Family Genes in Breast Cancer. Aging 2024, 16, 11434–11445. [Google Scholar] [CrossRef]
- Zhang, J.; Tian, Y.; Mo, S.; Fu, X. Overexpressing PLOD Family Genes Predict Poor Prognosis in Pancreatic Cancer. Int. J. Gen. Med. 2022, 15, 3077–3096. [Google Scholar] [CrossRef] [PubMed]
- Li, S.-S.; Lian, Y.-F.; Huang, Y.-L.; Huang, Y.-H.; Xiao, J. Overexpressing PLOD Family Genes Predict Poor Prognosis in Gastric Cancer. J. Cancer 2020, 11, 121–131. [Google Scholar] [CrossRef]
- Yamada, Y.; Kato, M.; Arai, T.; Sanada, H.; Uchida, A.; Misono, S.; Sakamoto, S.; Komiya, A.; Ichikawa, T.; Seki, N. Aberrantly Expressed PLOD1 Promotes Cancer Aggressiveness in Bladder Cancer: A Potential Prognostic Marker and Therapeutic Target. Mol. Oncol. 2019, 13, 1898–1912. [Google Scholar] [CrossRef]
- Zhai, Z.; Wang, S.; Cao, Y.; Liu, J.; Zhao, Q.; Ji, Y.; Yang, X.; Tang, X.; Ma, J.; Du, P. Pan-Cancer Analysis Reveals the Potential of PLOD1 as a Prognostic and Immune Biomarker for Human Cancer. Biomedicines 2024, 12, 2653. [Google Scholar] [CrossRef]
- Yeh, T.-L.; Leissing, T.M.; Abboud, M.I.; Thinnes, C.C.; Atasoylu, O.; Holt-Martyn, J.P.; Zhang, D.; Tumber, A.; Lippl, K.; Lohans, C.T.; et al. Molecular and Cellular Mechanisms of HIF Prolyl Hydroxylase Inhibitors in Clinical Trials. Chem. Sci. 2017, 8, 7651–7668. [Google Scholar] [CrossRef]
- Jia, W.; Chen, P.; Cheng, Y. PRDX4 and Its Roles in Various Cancers. Technol. Cancer Res. Treat. 2019, 18, 1533033819864313. [Google Scholar] [CrossRef]
- Ding, N.; Jiang, H.; Thapa, P.; Hao, Y.; Alshahrani, A.; Allison, D.; Izumi, T.; Rangnekar, V.M.; Liu, X.; Wei, Q. Peroxiredoxin IV Plays a Critical Role in Cancer Cell Growth and Radioresistance Through the Activation of the Akt/GSK3 Signaling Pathways. J. Biol. Chem. 2022, 298, 102123. [Google Scholar] [CrossRef]
- Zhang, L.; Wu, K.; Hou, Y.; Li, X. Validation of the Interaction between PRDX4 and TXNDC5 in Gastric Cancer and the Significance of the PRDX4 Gene in Gastric Cancer Based on a Data Mining Analysis. Transl. Cancer Res. 2024, 13, 81–101. [Google Scholar] [CrossRef]
- Zhou, H.; Li, L.; Chen, J.; Hou, S.; Zhou, T.; Xiong, Y. Expression and Prognostic Value of PRDX Family in Colon Adenocarcinoma by Integrating Comprehensive Analysis and In Vitro and In Vivo Validation. Front. Oncol. 2023, 13, 1136738. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Guo, X.; Nakamura, Y.; Zhou, X.; Yamaguchi, R.; Zhang, J.; Ishigaki, Y.; Uramoto, H.; Yamada, S. Overexpression of PRDX4 Modulates Tumor Microenvironment and Promotes Urethane-Induced Lung Tumorigenesis. Oxid. Med. Cell Longev. 2020, 2020, 8262730. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.-W.; Bai, Y.; Guo, J.-Q.; Li, Y.-Y.; Liu, Y.-F.; Liang, C.; Xing, Y.-R.; Guo, H.-L.; Qi, T.-X.; Wu, J.; et al. Peroxiredoxin 4 as a Switch Regulating PTEN/AKT Axis in Alveolar Macrophages Activation. Signal Transduct. Target. Ther. 2025, 10, 352. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.-R.; Park, J.-S.; Karabulut, H.; Yasmin, F.; Jun, C.-D. Transgelin-2: A Double-Edged Sword in Immunity and Cancer Metastasis. Front. Cell Dev. Biol. 2021, 9, 606149. [Google Scholar] [CrossRef]
- Jeon, B.-N.; Kim, H.-R.; Chung, Y.S.; Na, B.-R.; Park, H.; Hong, C.; Fatima, Y.; Oh, H.; Kim, C.-H.; Jun, C.-D. Actin Stabilizer TAGLN2 Potentiates Adoptive T Cell Therapy by Boosting the Inside-Out Costimulation via Lymphocyte Function-Associated Antigen-1. Oncoimmunology 2018, 7, e1500674. [Google Scholar] [CrossRef]
- Yang, L.; Hong, Q.; Xu, S.; Kuang, X.; Di, G.; Liu, G.; Wu, J.; Shao, Z.; Yu, S. Downregulation of Transgelin 2 Promotes Breast Cancer Metastasis by Activating the Reactive Oxygen Species/Nuclear Factor-κB Signaling Pathway. Mol. Med. Rep. 2019, 20, 4045–4258. [Google Scholar] [CrossRef]
- Pan, T.; Wang, S.; Wang, Z. An Integrated Analysis Identified TAGLN2 As an Oncogene Indicator Related to Prognosis and Immunity in Pan-Cancer. J. Cancer 2023, 14, 1809–1836. [Google Scholar] [CrossRef]
- Zhuo, H.; Hou, J.; Hong, Z.; Yu, S.; Peng, H.; Zhang, L.; Xie, W.; Hong, X. TAGLN2 Induces Resistance Signature ISGs by Activating AKT-YBX1 Signal with Dual Pathways and Mediates the IFN-Related DNA Damage Resistance in Gastric Cancer. Cell Death Dis. 2024, 15, 608. [Google Scholar] [CrossRef]
- Wang, L.; Tan, H.; Huang, Y.; Guo, M.; Dong, Y.; Liu, C.; Zhao, H.; Liu, Z. TAGLN2 Promotes Papillary Thyroid Carcinoma Invasion via the Rap1/PI3K/AKT Axis. Endocr. Relat. Cancer 2023, 30, e210352. [Google Scholar] [CrossRef]
- Zhao, Z.; Lu, L.; Li, W. TAGLN2 Promotes the Proliferation, Invasion, Migration and Epithelial-mesenchymal Transition of Colorectal Cancer Cells by Activating STAT3 Signaling through ANXA2. Oncol. Lett. 2021, 22, 737. [Google Scholar] [CrossRef]
- Jin, H.; Zheng, W.; Hou, J.; Peng, H.; Zhuo, H. An Essential NRP1-Mediated Role for Tagln2 in Gastric Cancer Angiogenesis. Front. Oncol. 2021, 11, 653246. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Rajasekera, P.; Becker, V.; Biehn, S.; Beyer, S.; McElroy, J.; Becker, A.; Johnson, B.; Cui, T.; Sebastian, E.; et al. Small Molecule Inhibitors Targeting Transgelin-2-Actin Interaction for Therapeutic Intervention in Glioblastoma. Int. J. Radiat. Oncol. Biol. Phys. 2022, 114, e72–e73. [Google Scholar] [CrossRef]
- Zhou, L.; Li, H.; Yao, H.; Dai, X.; Gao, P.; Cheng, H. TMED Family Genes and Their Roles in Human Diseases. Int. J. Med. Sci. 2023, 20, 1732–1743. [Google Scholar] [CrossRef]
- Wang, H.; Wang, Y.; Tan, P.; Liu, Y.; Zhou, S.; Ma, W. Prognostic Value and Anti-Tumor Immunity Role of TMED9 in Pan-Cancer: A Bioinformatics Study. Transl. Cancer Res. 2024, 13, 5429–5445. [Google Scholar] [CrossRef]
- Yang, Y.-C.; Chien, M.-H.; Lai, T.-C.; Tung, M.-C.; Jan, Y.-H.; Chang, W.-M.; Jung, S.-M.; Chen, M.-H.; Yeh, C.-N.; Hsiao, M. Proteomics-Based Identification of TMED9 Is Linked to Vascular Invasion and Poor Prognoses in Patients with Hepatocellular Carcinoma. J. Biomed. Sci. 2021, 28, 29. [Google Scholar] [CrossRef]
- HAN, G.H.; YUN, H.; CHUNG, J.-Y.; KIM, J.-H.; CHO, H. TMED9 Expression Level as a Biomarker of Epithelial Ovarian Cancer Progression and Prognosis. Cancer Genom. Proteom. 2022, 19, 692–702. [Google Scholar] [CrossRef]
- Fang, Z.; Song, Y.-X.; Wo, G.-Q.; Zhou, H.-L.; Li, L.; Yang, S.-Y.; Chen, X.; Zhang, J.; Tang, J.-H. Screening of the Novel Immune-Suppressive Biomarkers of TMED Family and Whether Knockdown of TMED2/3/4/9 Inhibits Cell Migration and Invasion in Breast Cancer. Ann. Transl. Med. 2022, 10, 1280. [Google Scholar] [CrossRef]
- Mishra, S.; Bernal, C.; Silvano, M.; Anand, S.; Ruiz i Altaba, A. The Protein Secretion Modulator TMED9 Drives CNIH4/TGFα/GLI Signaling Opposing TMED3-WNT-TCF to Promote Colon Cancer Metastases. Oncogene 2019, 38, 5817–5837. [Google Scholar] [CrossRef] [PubMed]
- Ullah, M.A.; Tabassum, T.; Farzana, M.; Moin, A.T.; Zohora, U.S.; Rahman, M.S. Expression Analysis, Molecular Characterization and Prognostic Evaluation on TMED4 and TMED9 Gene Expression in Glioma. Biomed. Signal Process. Control. 2022, 78, 103922. [Google Scholar] [CrossRef]
- Dvela-Levitt, M.; Kost-Alimova, M.; Emani, M.; Kohnert, E.; Thompson, R.; Sidhom, E.-H.; Rivadeneira, A.; Sahakian, N.; Roignot, J.; Papagregoriou, G.; et al. Small Molecule Targets TMED9 and Promotes Lysosomal Degradation to Reverse Proteinopathy. Cell 2019, 178, 521–535.e23. [Google Scholar] [CrossRef]
- Goldman, M.J.; Craft, B.; Hastie, M.; Repečka, K.; McDade, F.; Kamath, A.; Banerjee, A.; Luo, Y.; Rogers, D.; Brooks, A.N.; et al. Visualizing and Interpreting Cancer Genomics Data via the Xena Platform. Nat. Biotechnol. 2020, 38, 675–678. [Google Scholar] [CrossRef]
- Vivian, J.; Rao, A.A.; Nothaft, F.A.; Ketchum, C.; Armstrong, J.; Novak, A.; Pfeil, J.; Narkizian, J.; Deran, A.D.; Musselman-Brown, A.; et al. Toil Enables Reproducible, Open Source, Big Biomedical Data Analyses. Nat. Biotechnol. 2017, 35, 314–316. [Google Scholar] [CrossRef]
- Denkert, C.; Budczies, J.; Darb-Esfahani, S.; Györffy, B.; Sehouli, J.; Könsgen, D.; Zeillinger, R.; Weichert, W.; Noske, A.; Buckendahl, A.; et al. A Prognostic Gene Expression Index in Ovarian Cancer—Validation across Different Independent Data Sets. J. Pathol. 2009, 218, 273–280. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Pena, J.; Tibor Fekete, J.; Páez, R.; Baliu-Piqué, M.; García-Saenz, J.Á.; García-Barberán, V.; Manzano, A.; Pérez-Segura, P.; Esparis-Ogando, A.; Pandiella, A.; et al. A Transcriptomic Immunologic Signature Predicts Favorable Outcome in Neoadjuvant Chemotherapy Treated Triple Negative Breast Tumors. Front. Immunol. 2019, 10, 2802. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Kirsch, R.; Koutrouli, M.; Nastou, K.; Mehryary, F.; Hachilif, R.; Gable, A.L.; Fang, T.; Doncheva, N.T.; Pyysalo, S.; et al. The STRING Database in 2023: Protein–Protein Association Networks and Functional Enrichment Analyses for Any Sequenced Genome of Interest. Nucleic Acids Res. 2023, 51, D638–D646. [Google Scholar] [CrossRef] [PubMed]
| mRNA | Zscore | mOS (months) |
|---|---|---|
| ENO1 | −1.08 | >120 |
| ENO1 | 1.27 | >120 |
| ENO1 | 3.62 | >120 |
| ENO1 | 6 | 93.8 |
| ENO1 | 8.32 | 54.2 |
| GLRX2 | −1.36 | >120 |
| GLRX2 | 1.56 | >120 |
| GLRX2 | 4.47 | 114.8 |
| GLRX2 | 7.39 | 61.9 |
| GLRX2 | 10.3 | 31 |
| PLOD1 | −0.92 | >120 |
| PLOD1 | 1.41 | >120 |
| PLOD1 | 3.73 | 114.8 |
| PLOD1 | 6.06 | 61.94 |
| PLOD1 | 8.39 | 32.09 |
| PRDX4 | −1.16 | >120 |
| PRDX4 | 1.03 | >120 |
| PRDX4 | 3.22 | >120 |
| PRDX4 | 5.42 | 95.7 |
| PRDX4 | 7.61 | 62.5 |
| TAGLN2 | −1.54 | >120 |
| TAGLN2 | 0.92 | >120 |
| TAGLN2 | 3.37 | 93.8 |
| TAGLN2 | 5.83 | 32.6 |
| TAGLN2 | 8.28 | 14.7 |
| TMED9 | −1.72 | >120 |
| TMED9 | 0.48 | >120 |
| TMED9 | 2.67 | >120 |
| TMED9 | 4.86 | 95.7 |
| TMED9 | 7.06 | 62.5 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qazi, S.; Richardson, S.; Potts, M.; Myers, S.; Trieu, V. Bioinformatic Approach to Identify Positive Prognostic TGFB2-Dependent and Negative Prognostic TGFB2-Independent Biomarkers for Breast Cancers. Int. J. Mol. Sci. 2025, 26, 11580. https://doi.org/10.3390/ijms262311580
Qazi S, Richardson S, Potts M, Myers S, Trieu V. Bioinformatic Approach to Identify Positive Prognostic TGFB2-Dependent and Negative Prognostic TGFB2-Independent Biomarkers for Breast Cancers. International Journal of Molecular Sciences. 2025; 26(23):11580. https://doi.org/10.3390/ijms262311580
Chicago/Turabian StyleQazi, Sanjive, Stephen Richardson, Mike Potts, Scott Myers, and Vuong Trieu. 2025. "Bioinformatic Approach to Identify Positive Prognostic TGFB2-Dependent and Negative Prognostic TGFB2-Independent Biomarkers for Breast Cancers" International Journal of Molecular Sciences 26, no. 23: 11580. https://doi.org/10.3390/ijms262311580
APA StyleQazi, S., Richardson, S., Potts, M., Myers, S., & Trieu, V. (2025). Bioinformatic Approach to Identify Positive Prognostic TGFB2-Dependent and Negative Prognostic TGFB2-Independent Biomarkers for Breast Cancers. International Journal of Molecular Sciences, 26(23), 11580. https://doi.org/10.3390/ijms262311580





