Non-Histone Lysine Modifications in Tumor Microenvironment: Mechanisms and Therapeutic Opportunities
Abstract
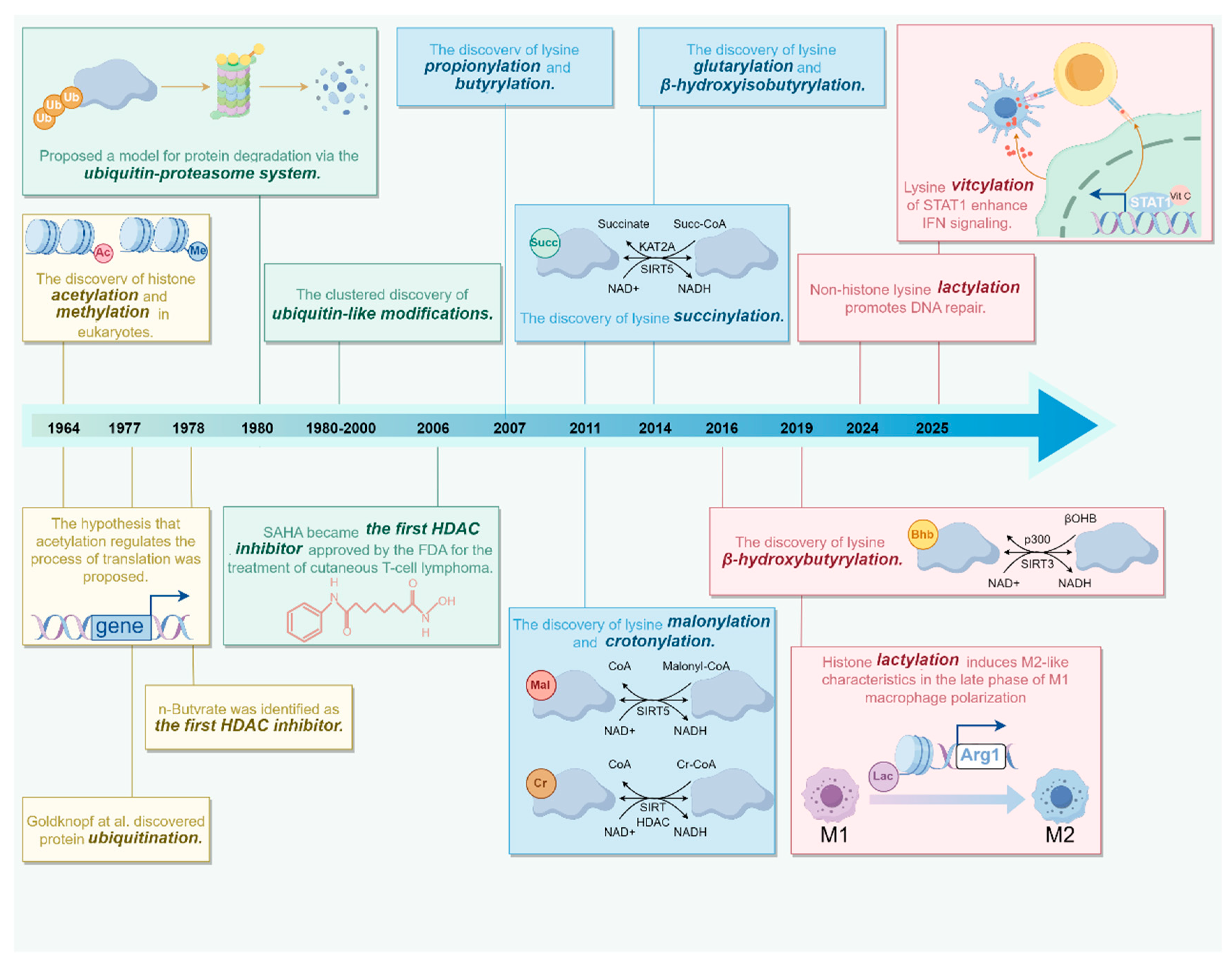
1. Introduction
2. Non-Histone Lysine Modifications in Tumors
2.1. The Overview of Non-Histone Lysine Modifications in Tumors
2.2. Regulation of Tumor-Associated Proteins Through Non-Histone Lysine PTMs
3. Crosstalk of Non-Histone Lysine PTMs in Tumors
3.1. Competitive Occupancy of Different PTMs at Identical Lysine Sites
3.2. Interdependence of Different PTMs at Neighboring Modification Sites
3.3. Regulating the Activity of Modifying Enzymes Through PTMs
3.4. Hierarchical Modification of Ubiquitination
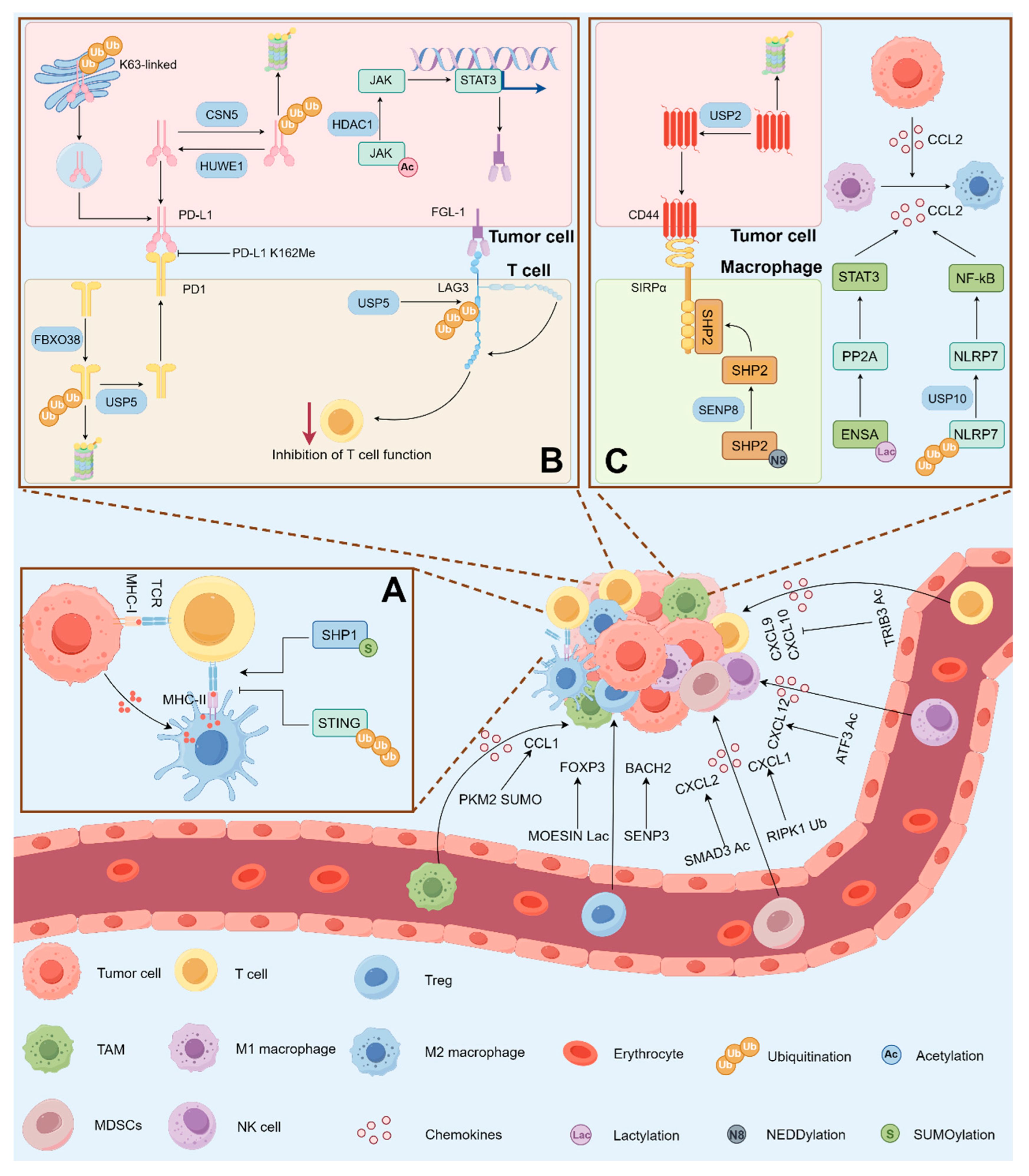
4. Non-Histone Lysine PTMs in TIME
4.1. Non-Histone Lysine PTMs in T Cells
4.2. Non-Histone Lysine PTMs in Macrophages
4.3. Non-Histone Lysine PTMs in Other Immune Cells
4.4. Lactate and Non-Histone Lactylation in TIME
5. Therapeutic Strategies Targeting Lysine PTMs
5.1. Targeting Substrates and Their Upstream/Downstream Pathways
5.2. Targeting Modifying Enzymes
5.3. Dietary Interventions
5.4. Challenges and Future Perspectives
| Targets | Cancer | Treatment | Phases | Study Status | NCT Number |
|---|---|---|---|---|---|
| Targeting methylation for cancer therapy | |||||
| EZH2 | Non-Hodgkin Lymphoma and Solid Tumor | AXT-1003 | Phase1 | Recruiting | NCT06484985 |
| Non-Hodgkin Lymphoma | AXT-1003 | Phase1 | Terminated | NCT05965505 | |
| Ovarian Cancer | CPI-0209 | Phase1 | Recruiting | NCT05942300 | |
| Peripheral T Cell Lymphoma | SHR2554 | Phase1 | Not Yet Recruiting | NCT06712173 | |
| Solid Tumor and Lymphoma | SHR2554 | Phase1/Phase2 | Recruiting | NCT04407741 | |
| Lymphoma | Tazemetostat | Phase2 | Recruiting | NCT06692452 | |
| Lymphoma | Tazemetostat | Phase2 | Not Yet Recruiting | NCT06068881 | |
| Follicular Lymphoma | Tazemetostat | Phase1/Phase2 | Recruiting | NCT05551936 | |
| Peripheral Nerve Sheath Tumor | Tazemetostat | Phase2 | Active Not Recruiting | NCT04917042 | |
| Melanoma | Tazemetostat | Phase1/Phase2 | Active Not Recruiting | NCT04557956 | |
| Lymphoma | Tazemetostat | Phase3 | Recruiting | NCT04224493 | |
| Non-Small Cell Lung Cancer | Tulmimetostat | Phase1/Phase2 | Not Yet Recruiting | NCT05467748 | |
| Advanced Tumor and Lymphoma | XNW5004 | Phase1/Phase2 | Recruiting | NCT06558513 | |
| Solid Tumors | XNW5004 | Phase1/Phase2 | Recruiting | NCT06022757 | |
| DOT1L | Acute Myeloid Leukemia | Pinometostat | Phase1/Phase2 | Completed | NCT03701295 |
| Targeting acetylation for cancer therapy | |||||
| HDAC | Colon Adenocarcinoma | Chidamide | Phase2 | Recruiting | NCT06709885 |
| Peripheral T-cell Lymphoma | Chidamide | Phase1/Phase2 | Recruiting | NCT06421948 | |
| Acute Lymphoblastic Leukemia | Chidamide | Phase2 | Recruiting | NCT06220487 | |
| Metastatic Colorectal Cancer | Sodium Valproate | Phase2 | Recruiting | NCT05694936 | |
| Breast Cancer | chidamide | Phase2 | Unknown | NCT05438706 | |
| Non-Hodgkin Lymphoma and Solid Tumor | Chidamide | Phase1/Phase2 | Unknown | NCT05320640 | |
| Neuroendocrine Tumor | Chidamide | Phase2 | Unknown | NCT05113355 | |
| Neuroendocrine Tumor | Chidamide | Phase2 | Unknown | NCT05076786 | |
| Solid Tumors and Lymphoma | Entinostat | Phase1/Phase2 | Recruiting | NCT05053971 | |
| Cervical Cancer | Chidamide | Phase1/Phase2 | Unknown | NCT04651127 | |
| T-cell Leukemia | Romidepsin | Phase1 | Withdrawn | NCT04639843 | |
| Non-Small Cell Lung Cancer | Entinostat | Phase1 | Completed | NCT04631029 | |
| Non-Hodgkin Lymphoma | Chidamide | Phase1/Phase2 | Unknown | NCT04553393 | |
| Hodgkin Lymphoma | Chidamide | Phase2 | Unknown | NCT04514081 | |
| Peripheral T-cell Lymphoma | Chidamide | Phase2 | Recruiting | NCT04512534 | |
| Non-Hodgkin Lymphoma | Chidamide | Phase1/Phase2 | Recruiting | NCT04337606 | |
| Breast Cancer | Entinostat | Phase1 | Terminated | NCT04296942 | |
| Hodgkin Lymphoma | Chidamide | Phase2 | Recruiting | NCT04233294 | |
| Lymphoma and Follicular | Abexinostat | Phase2 | Active Not Recruiting | NCT03934567 | |
| Solid Tumor and Lymphoma | Entinostat | Phase1 | Withdrawn | NCT03925428 | |
| Melanoma | Vorinostat | EARLYPhase1 | Withdrawn | NCT03022565 | |
| KAT6A/B | Solid Tumors | OP-3136 | Phase1 | Recruiting | NCT06784193 |
| Targeting ubiquitination for cancer therapy | |||||
| MDM2 | Acute Myeloid Leukemia | Navtemadlin | Phase1 | Active Not Recruiting | NCT04190550 |
| Acute Myeloid Leukemia | Siremadlin | Phase1/Phase2 | Terminated | NCT05447663 | |
| Acute Myeloid Leukemia | Siremadlin | Phase1 | Terminated | NCT05155709 | |
| Acute Myeloid Leukemia | Siremadlin | Phase1 | Terminated | NCT04496999 | |
| Soft-tissue Sarcoma | Siremadlin | Phase1/Phase2 | Recruiting | NCT05180695 | |
| Solid Tumor | Siremadlin | Phase2 | Recruiting | NCT04116541 | |
| Solid Tumor | Milademetan | Phase1/Phase2 | Withdrawn | NCT06090318 | |
| Solid Tumor | Milademetan | Phase2 | Terminated | NCT05012397 | |
| Dedifferentiated Liposarcoma | Milademetan | Phase3 | Terminated | NCT04979442 | |
| Breast Cancer | Milademetan | Phase2 | Terminated | NCT05932667 | |
| Neuroblastoma | APG-2575 | Phase1 | Recruiting | NCT05701306 | |
| Liposarcoma | APG-2575 | Phase1/Phase2 | Recruiting | NCT04785196 | |
| Leukemia | APG-2575 | Phase1/Phase2 | Recruiting | NCT04358393 | |
| Leukemia | APG-2575 | Phase1 | Recruiting | NCT04275518 | |
| CUL4-DDB1-CRBM-RBX1 E3 complex | Acute Myeloid Leukemia | CC-90009 | Phase1 | Terminated | NCT04336982 |
| CBL-B | Advanced Malignancies | NX-1607 | Phase1 | Recruiting | NCT05107674 |
| CRL4 | Hematological Malignancies | KPG-818 | Phase1 | Completed | NCT04283097 |
| USP1 | Solid Tumor | KSQ-4279 | Phase1 | Active Not Recruiting | NCT05240898 |
| Targeting SUMOylation for cancer therapy | |||||
| SUMO-activating enzyme | Head and Neck Cancer | TAK-981 | EARLYPhase1 | Completed | NCT04065555 |
| Solid Tumors | TAK-981 | Phase1/Phase2 | Completed | NCT04381650 | |
| Solid Tumors | TAK-981 | Phase1 | Terminated | NCT05976334 | |
| Targeting neddylation for cancer therapy | |||||
| NEDD8-activating enzyme | Acute Myelogenous Leukemia | Pevonedistat | Phase1 | Terminated | NCT04172844 |
| Acute Myeloid Leukemia | Pevonedistat | Phase2 | Completed | NCT04266795 | |
| Acute Myeloid Leukemia and Myelodysplastic Syndromes | Pevonedistat | Phase2 | Completed | NCT04712942 | |
| Cholangiocarcinoma and Hepatocellular Carcinoma | Pevonedistat | Phase2 | Active Not Recruiting | NCT04175912 | |
| Multiple Myeloma | Pevonedistat | Phase1 | Completed | NCT03770260 | |
| Non-Small Cell Lung Cancer | Pevonedistat | Phase2 | Active Not Recruiting | NCT03965689 | |
| Solid Tumor | Pevonedistat | Phase1/Phase2 | Terminated | NCT04800627 | |
6. Conclusions and Future Perspectives
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| PTMs | Post-translational modifications |
| TIME | tumor immune microenvironment |
| KMTs | lysine methyltransferases |
| KDMs | lysine demethylases |
| SAM | S-adenosylmethionine |
| KATs | lysine acetyltransferases |
| KDACs | lysine deacetylases |
| ASCC | activating signal cointegrator complex |
| DSB | double-strand break |
| HR | homologous recombination |
| UBLs | ubiquitin-like proteins |
| HDACi | HDAC inhibitor |
| ICIs | immune checkpoint inhibitors |
| TAMs | tumor-associated macrophages |
| Tregs | intratumoral regulatory T cells |
| MDSCs | myeloid-derived suppressor cells |
| NK cells | natural killer cells |
| FFA | free fatty acid |
| −SG diet | serine/glycine-free diet |
| PROTACs | proteolysis-targeting chimeras |
References
- Walsh, C.T.; Garneau-Tsodikova, S.; Gatto, G.J., Jr. Posttranslational Modifications: The Chemistry of Proteome Diversifications. Angew. Chem. Int. Ed. Engl. 2005, 44, 7342–7372. [Google Scholar] [CrossRef]
- Lee, J.M.; Hammarén, H.M.; Savitski, M.M.; Baek, S.H. Control of Protein Stability by Post-Translational Modifications. Nat. Commun. 2023, 14, 201. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, R.; Hei, H. Advances in Post-Translational Modifications of Proteins and Cancer Immunotherapy. Front. Immunol. 2023, 14, 1229397. [Google Scholar] [CrossRef]
- Pan, S.; Chen, R. Pathological Implication of Protein Post-Translational Modifications in Cancer. Mol. Asp. Med. 2022, 86, 101097. [Google Scholar] [CrossRef]
- Zhang, Z.; Tan, M.; Xie, Z.; Dai, L.; Chen, Y.; Zhao, Y. Identification of Lysine Succinylation as a New Post-Translational Modification. Nat. Chem. Biol. 2011, 7, 58–63. [Google Scholar] [CrossRef]
- Hao, B.; Chen, K.; Zhai, L.; Liu, M.; Liu, B.; Tan, M. Substrate and Functional Diversity of Protein Lysine Post-Translational Modifications. Genom. Proteom. Bioinform. 2024, 22, qzae019. [Google Scholar] [CrossRef]
- Hamamoto, R.; Saloura, V.; Nakamura, Y. Critical Roles of Non-Histone Protein Lysine Methylation in Human Tumorigenesis. Nat. Rev. Cancer 2015, 15, 110–124. [Google Scholar] [CrossRef] [PubMed]
- Allfrey, V.G.; Faulkner, R.; Mirsky, A.E. Acetylation and Methylation of Histones and Their Possible Role in the Regulation of Rna Synthesis. Proc. Natl. Acad. Sci. USA 1964, 51, 786–794. [Google Scholar] [CrossRef] [PubMed]
- Murray, K. The Occurrence of Epsilon-N-Methyl Lysine in Histones. Biochemistry 1964, 3, 10–15. [Google Scholar] [CrossRef] [PubMed]
- Bao, Y.; Ma, Y.; Huang, W.; Bai, Y.; Gao, S.; Xiu, L.; Xie, Y.; Wan, X.; Shan, S.; Chen, C.; et al. Regulation of Autophagy and Cellular Signaling through Non-Histone Protein Methylation. Int. J. Biol. Macromol. 2025, 291, 139057. [Google Scholar] [CrossRef]
- Luo, Y.; Lu, J.; Lei, Z.; Zhu, H.; Rao, D.; Wang, T.; Fu, C.; Zhang, Z.; Xia, L.; Huang, W. Lysine Methylation Modifications in Tumor Immunomodulation and Immunotherapy: Regulatory Mechanisms and Perspectives. Biomark. Res. 2024, 12, 74. [Google Scholar] [CrossRef]
- Mazur, P.K.; Reynoird, N.; Khatri, P.; Jansen, P.W.; Wilkinson, A.W.; Liu, S.; Barbash, O.; Van Aller, G.S.; Huddleston, M.; Dhanak, D.; et al. Smyd3 Links Lysine Methylation of Map3k2 to Ras-Driven Cancer. Nature 2014, 510, 283–287. [Google Scholar] [CrossRef]
- Lukinović, V.; Hausmann, S.; Roth, G.S.; Oyeniran, C.; Ahmad, T.; Tsao, N.; Brickner, J.R.; Casanova, A.G.; Chuffart, F.; Benitez, A.M.; et al. Smyd3 Impedes Small Cell Lung Cancer Sensitivity to Alkylation Damage through Rnf113a Methylation-Phosphorylation Cross-Talk. Cancer Discov. 2022, 12, 2158–2179. [Google Scholar] [CrossRef]
- Narita, T.; Weinert, B.T.; Choudhary, C. Functions and Mechanisms of Non-Histone Protein Acetylation. Nat. Rev. Mol. Cell Biol. 2019, 20, 156–174, Erratum in Nat. Rev. Mol. Cell Biol. 2019, 20, 508. [Google Scholar] [CrossRef]
- Choudhary, C.; Weinert, B.T.; Nishida, Y.; Verdin, E.; Mann, M. The Growing Landscape of Lysine Acetylation Links Metabolism and Cell Signalling. Nat. Rev. Mol. Cell Biol. 2014, 15, 536–550. [Google Scholar] [CrossRef]
- Wang, J.; Qian, J.; Hu, Y.; Kong, X.; Chen, H.; Shi, Q.; Jiang, L.; Wu, C.; Zou, W.; Chen, Y.; et al. Arhgap30 Promotes P53 Acetylation and Function in Colorectal Cancer. Nat. Commun. 2014, 5, 4735. [Google Scholar] [CrossRef]
- Goldknopf, I.L.; French, M.F.; Musso, R.; Busch, H. Presence of Protein A24 in Rat Liver Nucleosomes. Proc. Natl. Acad. Sci. USA 1977, 74, 5492–5495. [Google Scholar] [CrossRef] [PubMed]
- Ciechanover, A. The Unravelling of the Ubiquitin System. Nat. Rev. Mol. Cell Biol. 2015, 16, 322–324. [Google Scholar] [CrossRef] [PubMed]
- Deng, L.; Meng, T.; Chen, L.; Wei, W.; Wang, P. The Role of Ubiquitination in Tumorigenesis and Targeted Drug Discovery. Signal Transduct. Target. Ther. 2020, 5, 11. [Google Scholar] [CrossRef] [PubMed]
- Swatek, K.N.; Komander, D. Ubiquitin Modifications. Cell Res. 2016, 26, 399–422. [Google Scholar] [CrossRef]
- Ye, Q.; Ma, J.; Wang, Z.; Li, L.; Liu, T.; Wang, B.; Zhu, L.; Lei, Y.; Xu, S.; Wang, K.; et al. Dtx3l-Mediated Tirr Nuclear Export and Degradation Regulates DNA Repair Pathway Choice and Parp Inhibitor Sensitivity. Nat. Commun. 2024, 15, 10596. [Google Scholar] [CrossRef]
- Ma, J.; Shi, Q.; Cui, G.; Sheng, H.; Botuyan, M.V.; Zhou, Y.; Yan, Y.; He, Y.; Wang, L.; Wang, Y.; et al. Spop Mutation Induces Replication over-Firing by Impairing Geminin Ubiquitination and Triggers Replication Catastrophe Upon Atr Inhibition. Nat. Commun. 2021, 12, 5779. [Google Scholar] [CrossRef]
- Liu, F.; Chen, J.; Li, K.; Li, H.; Zhu, Y.; Zhai, Y.; Lu, B.; Fan, Y.; Liu, Z.; Chen, X.; et al. Ubiquitination and Deubiquitination in Cancer: From Mechanisms to Novel Therapeutic Approaches. Mol. Cancer 2024, 23, 148. [Google Scholar] [CrossRef] [PubMed]
- Cajee, U.F.; Hull, R.; Ntwasa, M. Modification by Ubiquitin-Like Proteins: Significance in Apoptosis and Autophagy Pathways. Int. J. Mol. Sci. 2012, 13, 11804–11831. [Google Scholar] [CrossRef]
- Kerscher, O.; Felberbaum, R.; Hochstrasser, M. Modification of Proteins by Ubiquitin and Ubiquitin-Like Proteins. Annu. Rev. Cell Dev. Biol. 2006, 22, 159–180. [Google Scholar] [CrossRef]
- Subramanian, S.; Bates, S.E.; Wright, J.J.; Espinoza-Delgado, I.; Piekarz, R.L. Clinical Toxicities of Histone Deacetylase Inhibitors. Pharmaceuticals 2010, 3, 2751–2767. [Google Scholar] [CrossRef]
- Zhang, S.; Yu, Q.; Li, Z.; Zhao, Y.; Sun, Y. Protein Neddylation and Its Role in Health and Diseases. Signal Transduct. Target. Ther. 2024, 9, 85. [Google Scholar] [CrossRef]
- Zhou, H.; Deng, N.; Li, Y.; Hu, X.; Yu, X.; Jia, S.; Zheng, C.; Gao, S.; Wu, H.; Li, K. Distinctive Tumorigenic Significance and Innovative Oncology Targets of Sumoylation. Theranostics 2024, 14, 3127–3149. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Sprung, R.; Tang, Y.; Ball, H.; Sangras, B.; Kim, S.C.; Falck, J.R.; Peng, J.; Gu, W.; Zhao, Y. Lysine Propionylation and Butyrylation Are Novel Post-Translational Modifications in Histones. Mol. Cell Proteom. 2007, 6, 812–819. [Google Scholar] [CrossRef] [PubMed]
- Peng, C.; Lu, Z.; Xie, Z.; Cheng, Z.; Chen, Y.; Tan, M.; Luo, H.; Zhang, Y.; He, W.; Yang, K.; et al. The First Identification of Lysine Malonylation Substrates and Its Regulatory Enzyme. Mol. Cell. Proteom. 2011, 10, M111.012658. [Google Scholar] [CrossRef]
- Tan, M.; Luo, H.; Lee, S.; Jin, F.; Yang, J.S.; Montellier, E.; Buchou, T.; Cheng, Z.; Rousseaux, S.; Rajagopal, N.; et al. Identification of 67 Histone Marks and Histone Lysine Crotonylation as a New Type of Histone Modification. Cell 2011, 146, 1016–1028. [Google Scholar] [CrossRef]
- Tan, M.; Peng, C.; Anderson, K.A.; Chhoy, P.; Xie, Z.; Dai, L.; Park, J.; Chen, Y.; Huang, H.; Zhang, Y.; et al. Lysine Glutarylation Is a Protein Posttranslational Modification Regulated by Sirt5. Cell Metab. 2014, 19, 605–617. [Google Scholar] [CrossRef]
- Dai, L.; Peng, C.; Montellier, E.; Lu, Z.; Chen, Y.; Ishii, H.; Debernardi, A.; Buchou, T.; Rousseaux, S.; Jin, F.; et al. Lysine 2-Hydroxyisobutyrylation Is a Widely Distributed Active Histone Mark. Nat. Chem. Biol. 2014, 10, 365–370. [Google Scholar] [CrossRef]
- Xie, Z.; Zhang, D.; Chung, D.; Tang, Z.; Huang, H.; Dai, L.; Qi, S.; Li, J.; Colak, G.; Chen, Y.; et al. Metabolic Regulation of Gene Expression by Histone Lysine Β-Hydroxybutyrylation. Mol. Cell 2016, 62, 194–206. [Google Scholar] [CrossRef]
- Zhang, D.; Tang, Z.; Huang, H.; Zhou, G.; Cui, C.; Weng, Y.; Liu, W.; Kim, S.; Lee, S.; Perez-Neut, M.; et al. Metabolic Regulation of Gene Expression by Histone Lactylation. Nature 2019, 574, 575–580. [Google Scholar] [CrossRef]
- Yu, X.; Yang, J.; Xu, J.; Pan, H.; Wang, W.; Yu, X.; Shi, S. Histone Lactylation: From Tumor Lactate Metabolism to Epigenetic Regulation. Int. J. Biol. Sci. 2024, 20, 1833–1854. [Google Scholar] [CrossRef]
- Zong, Z.; Ren, J.; Yang, B.; Zhang, L.; Zhou, F. Emerging Roles of Lysine Lactyltransferases and Lactylation. Nat. Cell Biol. 2025, 27, 563–574. [Google Scholar] [CrossRef] [PubMed]
- Zhu, R.; Ye, X.; Lu, X.; Xiao, L.; Yuan, M.; Zhao, H.; Guo, D.; Meng, Y.; Han, H.; Luo, S.; et al. Acss2 Acts as a Lactyl-Coa Synthetase and Couples Kat2a to Function as a Lactyltransferase for Histone Lactylation and Tumor Immune Evasion. Cell Metab. 2025, 37, 361–376.e7. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Liu, C.; Li, R.; Zhou, L.; Ran, Y.; Yang, Q.; Huang, H.; Lu, H.; Song, H.; Yang, B.; et al. Aars1 and Aars2 Sense L-Lactate to Regulate Cgas as Global Lysine Lactyltransferases. Nature 2024, 634, 1229–1237. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Wu, J.; Zhai, L.; Zhang, T.; Yin, H.; Gao, H.; Zhao, F.; Wang, Z.; Yang, X.; Jin, M.; et al. Metabolic Regulation of Homologous Recombination Repair by Mre11 Lactylation. Cell 2024, 187, 294–311.e21. [Google Scholar] [CrossRef]
- Chen, H.; Li, Y.; Li, H.; Chen, X.; Fu, H.; Mao, D.; Chen, W.; Lan, L.; Wang, C.; Hu, K.; et al. Nbs1 Lactylation Is Required for Efficient DNA Repair and Chemotherapy Resistance. Nature 2024, 631, 663–669. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Sun, L.; Gao, P.; Hu, H. Lactylation in Cancer: Current Understanding and Challenges. Cancer Cell 2024, 42, 1803–1807. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Yu, J.; Li, F.; Ge, S. Oncometabolites Drive Tumorigenesis by Enhancing Protein Acylation: From Chromosomal Remodelling to Nonhistone Modification. J. Exp. Clin. Cancer Res. 2022, 41, 144. [Google Scholar] [CrossRef] [PubMed]
- Hassin, O.; Oren, M. Drugging P53 in Cancer: One Protein, Many Targets. Nat. Rev. Drug Discov. 2023, 22, 127–144. [Google Scholar] [CrossRef]
- Chuikov, S.; Kurash, J.K.; Wilson, J.R.; Xiao, B.; Justin, N.; Ivanov, G.S.; McKinney, K.; Tempst, P.; Prives, C.; Gamblin, S.J.; et al. Regulation of P53 Activity through Lysine Methylation. Nature 2004, 432, 353–360. [Google Scholar] [CrossRef]
- Huang, J.; Perez-Burgos, L.; Placek, B.J.; Sengupta, R.; Richter, M.; Dorsey, J.A.; Kubicek, S.; Opravil, S.; Jenuwein, T.; Berger, S.L. Repression of P53 Activity by Smyd2-Mediated Methylation. Nature 2006, 444, 629–632. [Google Scholar] [CrossRef]
- Gu, W.; Roeder, R.G. Activation of P53 Sequence-Specific DNA Binding by Acetylation of the P53 C-Terminal Domain. Cell 1997, 90, 595–606. [Google Scholar] [CrossRef]
- Gu, W.; Shi, X.L.; Roeder, R.G. Synergistic Activation of Transcription by Cbp and P53. Nature 1997, 387, 819–823. [Google Scholar] [CrossRef]
- Luo, J.; Su, F.; Chen, D.; Shiloh, A.; Gu, W. Deacetylation of P53 Modulates Its Effect on Cell Growth and Apoptosis. Nature 2000, 408, 377–381. [Google Scholar] [CrossRef]
- Luo, J.; Nikolaev, A.Y.; Imai, S.; Chen, D.; Su, F.; Shiloh, A.; Guarente, L.; Gu, W. Negative Control of P53 by Sir2alpha Promotes Cell Survival under Stress. Cell 2001, 107, 137–148. [Google Scholar] [CrossRef]
- Vaziri, H.; Dessain, S.K.; Ng Eaton, E.; Imai, S.I.; Frye, R.A.; Pandita, T.K.; Guarente, L.; Weinberg, R.A. Hsir2(Sirt1) Functions as an Nad-Dependent P53 Deacetylase. Cell 2001, 107, 149–159. [Google Scholar] [CrossRef]
- Jakobsen, S.T.; Jensen, R.A.M.; Madsen, M.S.; Ravnsborg, T.; Vaagenso, C.S.; Siersbæk, M.S.; Einarsson, H.; Andersson, R.; Jensen, O.N.; Siersbæk, R. Myc Activity at Enhancers Drives Prognostic Transcriptional Programs through an Epigenetic Switch. Nat. Genet. 2024, 56, 663–674, Erratum in Nat. Genet. 2024, 56, 1764. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Li, Y.; Yang, J.; Sun, Y.; He, Y.; Wang, Y.; Liang, Y.; Chen, X.; Chen, T.; Han, D.; et al. Circcfl1 Promotes Tnbc Stemness and Immunoescape Via Deacetylation-Mediated C-Myc Deubiquitylation to Facilitate Mutant Tp53 Transcription. Adv. Sci. 2024, 11, e2404628. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Gao, Y.; Xue, S.; Zhao, L.; Jiang, H.; Zhang, T.; Li, Y.; Zhao, C.; Wu, F.; Siqin, T.; et al. Scarb2 Drives Hepatocellular Carcinoma Tumor Initiating Cells Via Enhanced Myc Transcriptional Activity. Nat. Commun. 2023, 14, 5917. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Chen, Z.; He, X.; Wang, J.; Zhong, J.; Zou, Y.; Zheng, X.; Lin, Y.; Zhang, R.; Kang, T.; et al. Sumoylation of Setd8 Promotes Tumor Growth by Methylating and Stabilizing Myc in Bladder Cancer. Adv. Sci. 2025, 12, e2501734. [Google Scholar] [CrossRef]
- Semenza, G.L. Hypoxia-Inducible Factors: Mediators of Cancer Progression and Targets for Cancer Therapy. Trends Pharmacol. Sci. 2012, 33, 207–214. [Google Scholar] [CrossRef]
- Luo, Y.; Yang, Z.; Yu, Y.; Zhang, P. Hif1α Lactylation Enhances Kiaa1199 Transcription to Promote Angiogenesis and Vasculogenic Mimicry in Prostate Cancer. Int. J. Biol. Macromol. 2022, 222 Pt B, 2225–2243. [Google Scholar] [CrossRef]
- Kim, Y.; Nam, H.J.; Lee, J.; Park, D.Y.; Kim, C.; Yu, Y.S.; Kim, D.; Park, S.W.; Bhin, J.; Hwang, D.; et al. Methylation-Dependent Regulation of Hif-1α Stability Restricts Retinal and Tumour Angiogenesis. Nat. Commun. 2016, 7, 10347. [Google Scholar] [CrossRef]
- Liu, X.; Chen, Z.; Xu, C.; Leng, X.; Cao, H.; Ouyang, G.; Xiao, W. Repression of Hypoxia-Inducible Factor A Signaling by Set7-Mediated Methylation. Nucleic Acids Res. 2015, 43, 5081–5098. [Google Scholar] [CrossRef]
- Zhang, X.; Di, Y.; Wang, Y.; Qin, J.; Ye, L.; Wen, X.; Ke, Z.; Wang, Z.; He, W. Sirt5-Mediated Desuccinylation of Ppa2 Enhances Hif-1alpha-Dependent Adaptation to Hypoxic Stress and Colorectal Cancer Metastasis. EMBO J. 2025, 44, 2514–2540. [Google Scholar] [CrossRef]
- Kim, S.C.; Sprung, R.; Chen, Y.; Xu, Y.; Ball, H.; Pei, J.; Cheng, T.; Kho, Y.; Xiao, H.; Xiao, L.; et al. Substrate and Functional Diversity of Lysine Acetylation Revealed by a Proteomics Survey. Mol. Cell 2006, 23, 607–618. [Google Scholar] [CrossRef]
- Choudhary, C.; Kumar, C.; Gnad, F.; Nielsen, M.L.; Rehman, M.; Walther, T.C.; Olsen, J.V.; Mann, M. Lysine Acetylation Targets Protein Complexes and Co-Regulates Major Cellular Functions. Science 2009, 325, 834–840. [Google Scholar] [CrossRef]
- Leutert, M.; Entwisle, S.W.; Villén, J. Decoding Post-Translational Modification Crosstalk with Proteomics. Mol. Cell Proteom. 2021, 20, 100129. [Google Scholar] [CrossRef]
- Fu, B.; Xiong, Y.; Sha, Z.; Xue, W.; Xu, B.; Tan, S.; Guo, D.; Lin, F.; Wang, L.; Ji, J.; et al. Septin2 Suppresses an Ifn-Γ-Independent, Proinflammatory Macrophage Activation Pathway. Nat. Commun. 2023, 14, 7441. [Google Scholar] [CrossRef]
- Ju, J.; Zhang, H.; Lin, M.; Yan, Z.; An, L.; Cao, Z.; Geng, D.; Yue, J.; Tang, Y.; Tian, L.; et al. The Alanyl-Trna Synthetase Aars1 Moonlights as a Lactyltransferase to Promote Yap Signaling in Gastric Cancer. J. Clin. Investig. 2024, 134, e174587. [Google Scholar] [CrossRef]
- Zhao, H.; Gao, S.; Han, Y.; Xie, D.; Xuan, L.; Huang, X.; Luo, J.; Ran, Q.; Li, G.; Guo, H.; et al. Conversion of Ku80 K568 Crotonylation to Sumoylation Facilitates DNA Non-Homologous End Joining and Cancer Radioresistance. Signal Transduct. Target. Ther. 2025, 10, 127. [Google Scholar] [CrossRef]
- Wang, T.; Lu, Z.; Han, T.; Wang, Y.; Gan, M.; Wang, J.B. Deacetylation of Glutaminase by Hdac4 Contributes to Lung Cancer Tumorigenesis. Int. J. Biol. Sci. 2022, 18, 4452–4465. [Google Scholar] [CrossRef]
- Wu, L.; Yan, X.; Sun, R.; Ma, Y.; Yao, W.; Gao, B.; Zhang, Q.; You, J.; Wang, H.; Han, Q.; et al. Sirt3 Restricts Tumor Initiation Via Promoting Lonp1 Deacetylation and K63 Ubiquitination. J. Transl. Med. 2023, 21, 81. [Google Scholar] [CrossRef]
- Arango, D.; Sturgill, D.; Alhusaini, N.; Dillman, A.A.; Sweet, T.J.; Hanson, G.; Hosogane, M.; Sinclair, W.R.; Nanan, K.K.; Mandler, M.D.; et al. Acetylation of Cytidine in Mrna Promotes Translation Efficiency. Cell 2018, 175, 1872–1886.e24. [Google Scholar] [CrossRef]
- Liao, L.; He, Y.; Li, S.J.; Yu, X.M.; Liu, Z.C.; Liang, Y.Y.; Yang, H.; Yang, J.; Zhang, G.G.; Deng, C.M.; et al. Lysine 2-Hydroxyisobutyrylation of Nat10 Promotes Cancer Metastasis in an Ac4c-Dependent Manner. Cell Res. 2023, 33, 355–371. [Google Scholar] [CrossRef]
- Liu, J.; Zhai, M.; Chen, Y.; Wei, Y.; Li, F.; Chen, Y.; Duan, B.; Xing, L.; Du, H.; Jiang, M.; et al. Acetylation-Dependent Deubiquitinase Usp26 Stabilizes Bag3 to Promote Breast Cancer Progression. Cancer Lett. 2024, 597, 217005. [Google Scholar] [CrossRef]
- Liu, W.; Zhan, Z.; Zhang, M.; Sun, B.; Shi, Q.; Luo, F.; Zhang, M.; Zhang, W.; Hou, Y.; Xiao, X.; et al. Kat6a, a Novel Regulator of Β-Catenin, Promotes Tumorigenicity and Chemoresistance in Ovarian Cancer by Acetylating Cop1. Theranostics 2021, 11, 6278–6292. [Google Scholar] [CrossRef]
- Ohtake, F.; Saeki, Y.; Sakamoto, K.; Ohtake, K.; Nishikawa, H.; Tsuchiya, H.; Ohta, T.; Tanaka, K.; Kanno, J. Ubiquitin Acetylation Inhibits Polyubiquitin Chain Elongation. EMBO Rep. 2015, 16, 192–201. [Google Scholar] [CrossRef]
- Wu, Q.; Cheng, Z.; Zhu, J.; Xu, W.; Peng, X.; Chen, C.; Li, W.; Wang, F.; Cao, L.; Yi, X.; et al. Suberoylanilide Hydroxamic Acid Treatment Reveals Crosstalks among Proteome, Ubiquitylome and Acetylome in Non-Small Cell Lung Cancer A549 Cell Line. Sci. Rep. 2015, 5, 9520. [Google Scholar] [CrossRef]
- Elia, A.E.; Boardman, A.P.; Wang, D.C.; Huttlin, E.L.; Everley, R.A.; Dephoure, N.; Zhou, C.; Koren, I.; Gygi, S.P.; Elledge, S.J. Quantitative Proteomic Atlas of Ubiquitination and Acetylation in the DNA Damage Response. Mol. Cell 2015, 59, 867–881. [Google Scholar] [CrossRef]
- Singh, R.K.; Zerath, S.; Kleifeld, O.; Scheffner, M.; Glickman, M.H.; Fushman, D. Recognition and Cleavage of Related to Ubiquitin 1 (Rub1) and Rub1-Ubiquitin Chains by Components of the Ubiquitin-Proteasome System. Mol. Cell Proteom. 2012, 11, 1595–1611. [Google Scholar] [CrossRef][Green Version]
- Rausch, L.; Kallies, A. Molecular Mechanisms Governing Cd8 T Cell Differentiation and Checkpoint Inhibitor Response in Cancer. Annu. Rev. Immunol. 2025, 43, 515–543. [Google Scholar] [CrossRef]
- Nagarsheth, N.; Wicha, M.S.; Zou, W. Chemokines in the Cancer Microenvironment and Their Relevance in Cancer Immunotherapy. Nat. Rev. Immunol. 2017, 17, 559–572. [Google Scholar] [CrossRef]
- Shang, S.; Yang, Y.W.; Chen, F.; Yu, L.; Shen, S.H.; Li, K.; Cui, B.; Lv, X.X.; Zhang, C.; Yang, C.; et al. Trib3 Reduces Cd8(+) T Cell Infiltration and Induces Immune Evasion by Repressing the Stat1-Cxcl10 Axis in Colorectal Cancer. Sci. Transl. Med. 2022, 14, eabf0992. [Google Scholar] [CrossRef]
- Zhang, R.; Yu, C.; Zeh, H.J.; Kroemer, G.; Klionsky, D.J.; Tang, D.; Kang, R. Tax1bp1-Dependent Autophagic Degradation of Sting1 Impairs Anti-Tumor Immunity. Autophagy 2025, 21, 1802–1823, Erratum in Autophagy 2025, 10, 1–2. [Google Scholar] [CrossRef]
- Liu, C.; Wang, X.; Qin, W.; Tu, J.; Li, C.; Zhao, W.; Ma, L.; Liu, B.; Qiu, H.; Yuan, X. Combining Radiation and the Atr Inhibitor Berzosertib Activates Sting Signaling and Enhances Immunotherapy Via Inhibiting Shp1 Function in Colorectal Cancer. Cancer Commun. 2023, 43, 435–454. [Google Scholar] [CrossRef]
- Baumeister, S.H.; Freeman, G.J.; Dranoff, G.; Sharpe, A.H. Coinhibitory Pathways in Immunotherapy for Cancer. Annu. Rev. Immunol. 2016, 34, 539–573. [Google Scholar] [CrossRef]
- Lim, S.O.; Li, C.W.; Xia, W.; Cha, J.H.; Chan, L.C.; Wu, Y.; Chang, S.S.; Lin, W.C.; Hsu, J.M.; Hsu, Y.H.; et al. Deubiquitination and Stabilization of Pd-L1 by Csn5. Cancer Cell 2016, 30, 925–939. [Google Scholar] [CrossRef]
- Meng, X.; Liu, X.; Guo, X.; Jiang, S.; Chen, T.; Hu, Z.; Liu, H.; Bai, Y.; Xue, M.; Hu, R.; et al. Fbxo38 Mediates Pd-1 Ubiquitination and Regulates Anti-Tumour Immunity of T Cells. Nature 2018, 564, 130–135. [Google Scholar] [CrossRef]
- Zhang, J.; Bu, X.; Wang, H.; Zhu, Y.; Geng, Y.; Nihira, N.T.; Tan, Y.; Ci, Y.; Wu, F.; Dai, X.; et al. Cyclin D-Cdk4 Kinase Destabilizes Pd-L1 Via Cullin 3-Spop to Control Cancer Immune Surveillance. Nature 2018, 553, 91–95, Erratum in Nature 2019, 571, E10. [Google Scholar] [CrossRef]
- Ma, X.; Jia, S.; Wang, G.; Liang, M.; Guo, T.; Du, H.; Li, S.; Li, X.; Huangfu, L.; Guo, J.; et al. Trim28 Promotes the Escape of Gastric Cancer Cells from Immune Surveillance by Increasing Pd-L1 Abundance. Signal Transduct. Target. Ther. 2023, 8, 246. [Google Scholar] [CrossRef]
- Yu, X.; Li, W.; Liu, H.; Wang, X.; Coarfa, C.; Cheng, C.; Yu, X.; Zeng, Z.; Cao, Y.; Young, K.H.; et al. Pd-L1 Translocation to the Plasma Membrane Enables Tumor Immune Evasion through Mib2 Ubiquitination. J. Clin. Investig. 2023, 133, e160456. [Google Scholar] [CrossRef]
- Gao, Y.; Nihira, N.T.; Bu, X.; Chu, C.; Zhang, J.; Kolodziejczyk, A.; Fan, Y.; Chan, N.T.; Ma, L.; Liu, J.; et al. Acetylation-Dependent Regulation of Pd-L1 Nuclear Translocation Dictates the Efficacy of Anti-Pd-1 Immunotherapy. Nat. Cell Biol. 2020, 22, 1064–1075. [Google Scholar] [CrossRef]
- Huang, C.; Ren, S.; Chen, Y.; Liu, A.; Wu, Q.; Jiang, T.; Lv, P.; Song, D.; Hu, F.; Lan, J.; et al. Pd-L1 Methylation Restricts Pd-L1/Pd-1 Interactions to Control Cancer Immune Surveillance. Sci. Adv. 2023, 9, eade4186. [Google Scholar] [CrossRef]
- Xiao, X.; Shi, J.; He, C.; Bu, X.; Sun, Y.; Gao, M.; Xiang, B.; Xiong, W.; Dai, P.; Mao, Q.; et al. Erk and Usp5 Govern Pd-1 Homeostasis Via Deubiquitination to Modulate Tumor Immunotherapy. Nat. Commun. 2023, 14, 2859, Erratum in Nat. Commun. 2025, 16, 5357. [Google Scholar] [CrossRef]
- Liu, D.; Li, M.; Zhao, Z.; Zhou, L.; Zhi, F.; Guo, Z.; Cui, J. Targeting the Trim14/Usp14 Axis Enhances Immunotherapy Efficacy by Inducing Autophagic Degradation of Pd-L1. Cancer Res. 2024, 84, 2806–2819. [Google Scholar] [CrossRef]
- Hsieh, H.C.; Young, M.J.; Chen, K.Y.; Su, W.C.; Lin, C.C.; Yen, Y.T.; Hung, J.J.; Wang, Y.C. Deubiquitinase Usp24 Activated by Il-6/Stat3 Enhances Pd-1 Protein Stability and Suppresses T Cell Antitumor Response. Sci. Adv. 2025, 11, eadt4258. [Google Scholar] [CrossRef]
- Liang, L.; Kuang, X.; He, Y.; Zhu, L.; Lau, P.; Li, X.; Luo, D.; Gong, L.; Zhou, W.; Zhang, F.; et al. Alterations in Pd-L1 Succinylation Shape Anti-Tumor Immune Responses in Melanoma. Nat. Genet. 2025, 57, 680–693. [Google Scholar] [CrossRef]
- Jiang, Y.; Dai, A.; Huang, Y.; Li, H.; Cui, J.; Yang, H.; Si, L.; Jiao, T.; Ren, Z.; Zhang, Z.; et al. Ligand-Induced Ubiquitination Unleashes Lag3 Immune Checkpoint Function by Hindering Membrane Sequestration of Signaling Motifs. Cell 2025, 188, 2354–2371. [Google Scholar] [CrossRef]
- Xu, T.; Fang, Y.; Gu, Y.; Xu, D.; Hu, T.; Yu, T.; Xu, Y.Y.; Shen, H.Y.; Ma, P.; Shu, Y. Hdac Inhibitor Saha Enhances Antitumor Immunity Via the Hdac1/Jak1/Fgl1 Axis in Lung Adenocarcinoma. J. Immunother. Cancer 2024, 12, e010077. [Google Scholar] [CrossRef]
- Yao, C.C.; Sun, R.M.; Yang, Y.; Zhou, H.Y.; Meng, Z.W.; Chi, R.; Xia, L.L.; Ji, P.; Chen, Y.Y.; Zhang, G.Q.; et al. Accumulation of Branched-Chain Amino Acids Reprograms Glucose Metabolism in Cd8(+) T Cells with Enhanced Effector Function and Anti-Tumor Response. Cell Rep. 2023, 42, 112186. [Google Scholar] [CrossRef]
- Hu, Z.; Chen, Y.; Lei, J.; Wang, K.; Pan, Z.; Zhang, L.; Xu, X.; Li, W.; Zhang, L.; Qin, X.; et al. Sirt7 Regulates T-Cell Antitumor Immunity through Modulation Bcaa and Fatty Acid Metabolism. Cell Death Differ. 2025, 32, 1777–1790. [Google Scholar] [CrossRef]
- Wang, W.; Green, M.; Choi, J.E.; Gijón, M.; Kennedy, P.D.; Johnson, J.K.; Liao, P.; Lang, X.; Kryczek, I.; Sell, A.; et al. Cd8(+) T Cells Regulate Tumour Ferroptosis during Cancer Immunotherapy. Nature 2019, 569, 270–274. [Google Scholar] [CrossRef]
- Han, Y.; Zhang, Y.Y.; Pan, Y.Q.; Zheng, X.J.; Liao, K.; Mo, H.Y.; Sheng, H.; Wu, Q.N.; Liu, Z.X.; Zeng, Z.L.; et al. Il-1β-Associated Nnt Acetylation Orchestrates Iron-Sulfur Cluster Maintenance and Cancer Immunotherapy Resistance. Mol. Cell 2023, 83, 1887–1902.e8. [Google Scholar] [CrossRef] [PubMed]
- Xiang, X.; Wang, J.; Lu, D.; Xu, X. Targeting Tumor-Associated Macrophages to Synergize Tumor Immunotherapy. Signal Transduct. Target. Ther. 2021, 6, 75. [Google Scholar] [CrossRef]
- Mantovani, A.; Marchesi, F.; Malesci, A.; Laghi, L.; Allavena, P. Tumour-Associated Macrophages as Treatment Targets in Oncology. Nat. Rev. Clin. Oncol. 2017, 14, 399–416. [Google Scholar] [CrossRef]
- Pollard, J.W. Tumour-Educated Macrophages Promote Tumour Progression and Metastasis. Nat. Rev. Cancer 2004, 4, 71–78. [Google Scholar] [CrossRef]
- Hou, P.P.; Luo, L.J.; Chen, H.Z.; Chen, Q.T.; Bian, X.L.; Wu, S.F.; Zhou, J.X.; Zhao, W.X.; Liu, J.M.; Wang, X.M.; et al. Ectosomal Pkm2 Promotes Hcc by Inducing Macrophage Differentiation and Remodeling the Tumor Microenvironment. Mol. Cell 2020, 78, 1192–1206.e10. [Google Scholar] [CrossRef]
- DeNardo, D.G.; Ruffell, B. Macrophages as Regulators of Tumour Immunity and Immunotherapy. Nat. Rev. Immunol. 2019, 19, 369–382. [Google Scholar] [CrossRef]
- Dai, P.; Sun, Y.; Huang, Z.; Liu, Y.T.; Gao, M.; Liu, H.M.; Shi, J.; He, C.; Xiang, B.; Yao, Y.; et al. Usp2 Inhibition Unleashes Cd47-Restrained Phagocytosis and Enhances Anti-Tumor Immunity. Nat. Commun. 2025, 16, 4564. [Google Scholar] [CrossRef]
- Kharitonenkov, A.; Chen, Z.; Sures, I.; Wang, H.; Schilling, J.; Ullrich, A. A Family of Proteins That Inhibit Signalling through Tyrosine Kinase Receptors. Nature 1997, 386, 181–186. [Google Scholar] [CrossRef]
- Li, Y.; Zhou, H.; Liu, P.; Lv, D.; Shi, Y.; Tang, B.; Xu, J.; Zhong, T.; Xu, W.; Zhang, J.; et al. Shp2 Deneddylation Mediates Tumor Immunosuppression in Colon Cancer Via the Cd47/Sirpα Axis. J. Clin. Investig. 2023, 133, e162870. [Google Scholar] [CrossRef]
- Muntjewerff, E.M.; Meesters, L.D.; van den Bogaart, G. Antigen Cross-Presentation by Macrophages. Front. Immunol. 2020, 11, 1276. [Google Scholar] [CrossRef]
- Zhang, N.; Sun, L.; Zhou, S.; Ji, C.; Cui, T.; Chu, Q.; Ye, J.; Liang, S.; Ma, K.; Liu, Y.; et al. Cholangiocarcinoma Pdha1 Succinylation Suppresses Macrophage Antigen Presentation Via Alpha-Ketoglutaric Acid Accumulation. Nat. Commun. 2025, 16, 3177. [Google Scholar] [CrossRef]
- Lin, Y.; Xu, J.; Lan, H. Tumor-Associated Macrophages in Tumor Metastasis: Biological Roles and Clinical Therapeutic Applications. J. Hematol. Oncol. 2019, 12, 76. [Google Scholar] [CrossRef]
- Zhang, B.; Zhang, Z.; Li, L.; Qin, Y.R.; Liu, H.; Jiang, C.; Zeng, T.T.; Li, M.Q.; Xie, D.; Li, Y.; et al. Tspan15 Interacts with Btrc to Promote Oesophageal Squamous Cell Carcinoma Metastasis Via Activating Nf-Κb Signaling. Nat. Commun. 2018, 9, 1423. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Wang, J.; Zhao, J.; Wang, H.; Chen, J.; Wu, J. Hmga2 Facilitates Colorectal Cancer Progression Via Stat3-Mediated Tumor-Associated Macrophage Recruitment. Theranostics 2022, 12, 963–975. [Google Scholar] [CrossRef]
- Li, B.; Qi, Z.P.; He, D.L.; Chen, Z.H.; Liu, J.Y.; Wong, M.W.; Zhang, J.W.; Xu, E.P.; Shi, Q.; Cai, S.L.; et al. Nlrp7 Deubiquitination by Usp10 Promotes Tumor Progression and Tumor-Associated Macrophage Polarization in Colorectal Cancer. J. Exp. Clin. Cancer Res. 2021, 40, 126. [Google Scholar] [CrossRef]
- Bauer, C.A.; Kim, E.Y.; Marangoni, F.; Carrizosa, E.; Claudio, N.M.; Mempel, T.R. Dynamic Treg Interactions with Intratumoral Apcs Promote Local Ctl Dysfunction. J. Clin. Investig. 2014, 124, 2425–2440. [Google Scholar] [CrossRef]
- Yu, X.; Lao, Y.; Teng, X.L.; Li, S.; Zhou, Y.; Wang, F.; Guo, X.; Deng, S.; Chang, Y.; Wu, X.; et al. Senp3 Maintains the Stability and Function of Regulatory T Cells Via Bach2 Desumoylation. Nat. Commun. 2018, 9, 3157. [Google Scholar] [CrossRef]
- Zhao, Y.; Du, J.; Shen, X. Targeting Myeloid-Derived Suppressor Cells in Tumor Immunotherapy: Current, Future and Beyond. Front. Immunol. 2023, 14, 1157537. [Google Scholar] [CrossRef] [PubMed]
- Liang, M.; Sun, Z.; Chen, X.; Wang, L.; Wang, H.; Qin, L.; Zhao, W.; Geng, B. E3 Ligase Trim28 Promotes Anti-Pd-1 Resistance in Non-Small Cell Lung Cancer by Enhancing the Recruitment of Myeloid-Derived Suppressor Cells. J. Exp. Clin. Cancer Res. 2023, 42, 275. [Google Scholar] [CrossRef] [PubMed]
- Yu, B.; Luo, F.; Sun, B.; Liu, W.; Shi, Q.; Cheng, S.Y.; Chen, C.; Chen, G.; Li, Y.; Feng, H. Kat6a Acetylation of Smad3 Regulates Myeloid-Derived Suppressor Cell Recruitment, Metastasis, and Immunotherapy in Triple-Negative Breast Cancer. Adv. Sci. 2021, 8, e2100014. [Google Scholar] [CrossRef]
- Wu, S.Y.; Fu, T.; Jiang, Y.Z.; Shao, Z.M. Natural Killer Cells in Cancer Biology and Therapy. Mol. Cancer 2020, 19, 120. [Google Scholar] [CrossRef]
- Zhu, J.; Wang, F.; Wang, L.; Dai, B.; Xu, G.; Zhao, L.; Jiang, H.; Gao, W.; Zhang, T.; Zhao, C.; et al. Hdac Inhibition Increases Cxcl12 Secretion to Recruit Natural Killer Cells in Peripheral T-Cell Lymphoma. Cancer Res. 2024, 84, 2450–2467. [Google Scholar] [CrossRef]
- Caronni, N.; Simoncello, F.; Stafetta, F.; Guarnaccia, C.; Ruiz-Moreno, J.S.; Opitz, B.; Galli, T.; Proux-Gillardeaux, V.; Benvenuti, F. Downregulation of Membrane Trafficking Proteins and Lactate Conditioning Determine Loss of Dendritic Cell Function in Lung Cancer. Cancer Res. 2018, 78, 1685–1699. [Google Scholar] [CrossRef]
- Mendler, A.N.; Hu, B.; Prinz, P.U.; Kreutz, M.; Gottfried, E.; Noessner, E. Tumor Lactic Acidosis Suppresses Ctl Function by Inhibition of P38 and Jnk/C-Jun Activation. Int. J. Cancer 2012, 131, 633–640. [Google Scholar] [CrossRef] [PubMed]
- Kumagai, S.; Koyama, S.; Itahashi, K.; Tanegashima, T.; Lin, Y.T.; Togashi, Y.; Kamada, T.; Irie, T.; Okumura, G.; Kono, H.; et al. Lactic Acid Promotes Pd-1 Expression in Regulatory T cells in Highly Glycolytic Tumor Microenvironments. Cancer Cell 2022, 40, 201–218.e9. [Google Scholar] [CrossRef]
- Sun, K.; Zhang, X.; Shi, J.; Huang, J.; Wang, S.; Li, X.; Lin, H.; Zhao, D.; Ye, M.; Zhang, S.; et al. Elevated Protein Lactylation Promotes Immunosuppressive Microenvironment and Therapeutic Resistance in Pancreatic Ductal Adenocarcinoma. J. Clin. Investig. 2025, 135, e187024. [Google Scholar] [CrossRef]
- Gu, J.; Zhou, J.; Chen, Q.; Xu, X.; Gao, J.; Li, X.; Shao, Q.; Zhou, B.; Zhou, H.; Wei, S.; et al. Tumor Metabolite Lactate Promotes Tumorigenesis by Modulating Moesin Lactylation and Enhancing Tgf-Β Signaling in Regulatory T cells. Cell Rep. 2022, 39, 110986. [Google Scholar] [CrossRef]
- Chen, J.; Zhao, D.; Wang, Y.; Liu, M.; Zhang, Y.; Feng, T.; Xiao, C.; Song, H.; Miao, R.; Xu, L.; et al. Lactylated Apolipoprotein C-Ii Induces Immunotherapy Resistance by Promoting Extracellular Lipolysis. Adv. Sci. 2024, 11, e2406333. [Google Scholar] [CrossRef]
- Jia, H.; Jiang, L.; Shen, X.; Ye, H.; Li, X.; Zhang, L.; Hu, Y.; Song, D.; Jia, H.; Wang, Z. Post-Translational Modifications of Cancer Immune Checkpoints: Mechanisms and Therapeutic Strategies. Mol. Cancer 2025, 24, 193. [Google Scholar] [CrossRef]
- Xiong, H.J.; Yu, H.Q.; Zhang, J.; Fang, L.; Wu, D.; Lin, X.T.; Xie, C.M. Elevated Fbxl6 Activates Both Wild-Type Kras and Mutant Kras(G12d) and Drives Hcc Tumorigenesis Via the Erk/Mtor/Prelid2/Ros Axis in Mice. Mil. Med. Res. 2023, 10, 68. [Google Scholar]
- Feng, X.; Zhang, H.; Meng, L.; Song, H.; Zhou, Q.; Qu, C.; Zhao, P.; Li, Q.; Zou, C.; Liu, X.; et al. Hypoxia-Induced Acetylation of Pak1 Enhances Autophagy and Promotes Brain Tumorigenesis Via Phosphorylating Atg5. Autophagy 2021, 17, 723–742. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Yang, J.; Liao, W.; Liu, X.; Zhang, H.; Wang, S.; Wang, D.; Feng, J.; Yu, L.; Zhu, W.G. Cytosolic Foxo1 Is Essential for the Induction of Autophagy and Tumour Suppressor Activity. Nat. Cell Biol. 2010, 12, 665–675. [Google Scholar] [CrossRef] [PubMed]
- Riggs, M.G.; Whittaker, R.G.; Neumann, J.R.; Ingram, V.M. N-Butyrate Causes Histone Modification in Hela and Friend Erythroleukaemia Cells. Nature 1977, 268, 462–464. [Google Scholar] [CrossRef]
- Candido, E.P.; Reeves, R.; Davie, J.R. Sodium Butyrate Inhibits Histone Deacetylation in Cultured Cells. Cell 1978, 14, 105–113. [Google Scholar] [CrossRef]
- Marks, P.A.; Breslow, R. Dimethyl Sulfoxide to Vorinostat: Development of This Histone Deacetylase Inhibitor as an Anticancer Drug. Nat. Biotechnol. 2007, 25, 84–90. [Google Scholar] [CrossRef]
- Shen, H.; Qi, X.; Hu, Y.; Wang, Y.; Zhang, J.; Liu, Z.; Qin, Z. Targeting Sirtuins for Cancer Therapy: Epigenetics Modifications and Beyond. Theranostics 2024, 14, 6726–6767. [Google Scholar] [CrossRef]
- Zhou, B.; Yang, Y.; Pang, X.; Shi, J.; Jiang, T.; Zheng, X. Quercetin Inhibits DNA Damage Responses to Induce Apoptosis Via Sirt5/Pi3k/Akt Pathway in Non-Small Cell Lung Cancer. Biomed. Pharmacother. 2023, 165, 115071. [Google Scholar] [CrossRef]
- Zhou, M.; Niu, H.; Cui, D.; Huang, G.; Li, J.; Tian, H.; Xu, X.; Liang, F.; Chen, R. Resveratrol Triggers Autophagy-Related Apoptosis to Inhibit the Progression of Colorectal Cancer Via Inhibition of Foxq1. Phytother. Res. 2024, 38, 3218–3239. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Shi, L.; Xie, N.; Liu, Z.; Qian, M.; Meng, F.; Xu, Q.; Zhou, M.; Cao, X.; Zhu, W.G.; et al. Sirt7 Antagonizes Tgf-Β Signaling and Inhibits Breast Cancer Metastasis. Nat. Commun. 2017, 8, 318. [Google Scholar] [CrossRef] [PubMed]
- Simon, R.P.; Robaa, D.; Alhalabi, Z.; Sippl, W.; Jung, M. Katching-up on Small Molecule Modulators of Lysine Acetyltransferases. J. Med. Chem. 2016, 59, 1249–1270. [Google Scholar] [CrossRef]
- Bhat, K.P.; Ümit Kaniskan, H.; Jin, J.; Gozani, O. Epigenetics and Beyond: Targeting Writers of Protein Lysine Methylation to Treat Disease. Nat. Rev. Drug Discov. 2021, 20, 265–286. [Google Scholar] [CrossRef] [PubMed]
- Biggar, K.K.; Wang, Z.; Li, S.S. Snapshot: Lysine Methylation Beyond Histones. Mol. Cell 2017, 68, 1016–1016.E1. [Google Scholar] [CrossRef]
- Klein, A.M.; de Queiroz, R.M.; Venkatesh, D.; Prives, C. The Roles and Regulation of Mdm2 and Mdmx: It Is Not Just About P53. Genes. Dev. 2021, 35, 575–601. [Google Scholar] [CrossRef]
- Nencioni, A.; Caffa, I.; Cortellino, S.; Longo, V.D. Fasting and Cancer: Molecular Mechanisms and Clinical Application. Nat. Rev. Cancer 2018, 18, 707–719. [Google Scholar] [CrossRef]
- Hwang, C.Y.; Choe, W.; Yoon, K.S.; Ha, J.; Kim, S.S.; Yeo, E.J.; Kang, I. Molecular Mechanisms for Ketone Body Metabolism, Signaling Functions, and Therapeutic Potential in Cancer. Nutrients 2022, 14, 4932. [Google Scholar] [CrossRef]
- Qin, J.; Huang, X.; Gou, S.; Zhang, S.; Gou, Y.; Zhang, Q.; Chen, H.; Sun, L.; Chen, M.; Liu, D.; et al. Ketogenic Diet Reshapes Cancer Metabolism through Lysine Β-Hydroxybutyrylation. Nat. Metab. 2024, 6, 1505–1528. [Google Scholar] [CrossRef] [PubMed]
- Shimazu, T.; Hirschey, M.D.; Newman, J.; He, W.; Shirakawa, K.; Le Moan, N.; Grueter, C.A.; Lim, H.; Saunders, L.R.; Stevens, R.D.; et al. Suppression of Oxidative Stress by Β-Hydroxybutyrate, an Endogenous Histone Deacetylase Inhibitor. Science 2013, 339, 211–214. [Google Scholar] [CrossRef] [PubMed]
- Geeraerts, S.L.; Heylen, E.; De Keersmaecker, K.; Kampen, K.R. The Ins and Outs of Serine and Glycine Metabolism in Cancer. Nat. Metab. 2021, 3, 131–141. [Google Scholar] [CrossRef]
- Maddocks, O.D.K.; Athineos, D.; Cheung, E.C.; Lee, P.; Zhang, T.; van den Broek, N.J.F.; Mackay, G.M.; Labuschagne, C.F.; Gay, D.; Kruiswijk, F.; et al. Modulating the Therapeutic Response of Tumours to Dietary Serine and Glycine Starvation. Nature 2017, 544, 372–376, Erratum in Nature 2017, 548, 122. [Google Scholar] [CrossRef]
- Muthusamy, T.; Cordes, T.; Handzlik, M.K.; You, L.; Lim, E.W.; Gengatharan, J.; Pinto, A.F.M.; Badur, M.G.; Kolar, M.J.; Wallace, M.; et al. Serine Restriction Alters Sphingolipid Diversity to Constrain Tumour Growth. Nature 2020, 586, 790–795. [Google Scholar] [CrossRef] [PubMed]
- Tong, H.; Jiang, Z.; Song, L.; Tan, K.; Yin, X.; He, C.; Huang, J.; Li, X.; Jing, X.; Yun, H.; et al. Dual Impacts of Serine/Glycine-Free Diet in Enhancing Antitumor Immunity and Promoting Evasion Via Pd-L1 Lactylation. Cell Metab. 2024, 36, 2493–2510.e9. [Google Scholar] [CrossRef]
- Zong, Z.; Xie, F.; Wang, S.; Wu, X.; Zhang, Z.; Yang, B.; Zhou, F. Alanyl-Trna Synthetase, Aars1, Is a Lactate Sensor and Lactyltransferase That Lactylates P53 and Contributes to Tumorigenesis. Cell 2024, 187, 2375–2392.e33. [Google Scholar]
- He, X.; Wang, Q.; Cheng, X.; Wang, W.; Li, Y.; Nan, Y.; Wu, J.; Xiu, B.; Jiang, T.; Bergholz, J.S.; et al. Lysine Vitcylation Is a Vitamin C-Derived Protein Modification That Enhances Stat1-Mediated Immune Response. Cell 2025, 188, 1858–1877.e21. [Google Scholar] [CrossRef]
- Bornes, K.E.; Moody, M.A.; Huckaba, T.M.; Benz, M.C.; McConnell, E.C.; Foroozesh, M.; Barnes, V.H.; Collins-Burow, B.M.; Burow, M.E.; Watt, T.J.; et al. Lysine Deacetylase Inhibitors Have Low Selectivity in Cells and Exhibit Predominantly Off-Target Effects. FEBS Open Bio 2025, 15, 94–107. [Google Scholar] [CrossRef]
- Wang, X.; Jiang, W.; Du, Y.; Zhu, D.; Zhang, J.; Fang, C.; Yan, F.; Chen, Z.S. Targeting Feedback Activation of Signaling Transduction Pathways to Overcome Drug Resistance in Cancer. Drug Resist. Updates 2022, 65, 100884. [Google Scholar] [CrossRef]
- Xu, X.; Wang, Q.; Guo, K.; Xu, J.; Lu, Y.; Chen, H.; Hu, W.; Fu, Y.; Sun, L.; He, Y.; et al. Cd47 Blockade Reverses Resistance to Hdac Inhibitor by Liberating Anti-Tumor Capacity of Macrophages. J. Exp. Clin. Cancer Res. 2025, 44, 67. [Google Scholar] [CrossRef]
- Wang, M.; Wang, J.; Wang, R.; Jiao, S.; Wang, S.; Zhang, J.; Zhang, M. Identification of a Monoclonal Antibody That Targets Pd-1 in a Manner Requiring Pd-1 Asn58 Glycosylation. Commun. Biol. 2019, 2, 392. [Google Scholar] [CrossRef] [PubMed]
- Nalawansha, D.A.; Crews, C.M. Protacs: An Emerging Therapeutic Modality in Precision Medicine. Cell Chem. Biol. 2020, 27, 998–1014. [Google Scholar] [CrossRef]
- Vander Heiden, M.G.; Cantley, L.C.; Thompson, C.B. Understanding the Warburg Effect: The Metabolic Requirements of Cell Proliferation. Science 2009, 324, 1029–1033. [Google Scholar] [CrossRef]
- Chen, J.; Huang, Z.; Chen, Y.; Tian, H.; Chai, P.; Shen, Y.; Yao, Y.; Xu, S.; Ge, S.; Jia, R. Lactate and Lactylation in Cancer. Signal Transduct. Target. Ther. 2025, 10, 38. [Google Scholar] [CrossRef] [PubMed]
- Venne, A.S.; Kollipara, L.; Zahedi, R.P. The Next Level of Complexity: Crosstalk of Posttranslational Modifications. Proteomics 2014, 14, 513–524. [Google Scholar] [CrossRef]
- Dang, F.; Wei, W. Targeting the Acetylation Signaling Pathway in Cancer Therapy. Semin. Cancer Biol. 2022, 85, 209–218. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Zhang, H.; Gao, P. Metabolic Reprogramming and Epigenetic Modifications on the Path to Cancer. Protein Cell 2022, 13, 877–919. [Google Scholar] [CrossRef] [PubMed]



| Immune Cells | PTMs | Target Protein | Sites | Writer | Eraser | Involved Pathways | Main Effects | References |
|---|---|---|---|---|---|---|---|---|
| T cell | Acetylation | TRIB3 | K240 | p300 | - | P300/TRIB3/EGFR/STAT1/CXCL10 | Enhance T cell infiltration | [79] |
| PD-L1 | K263 | p300 | HDAC2 | P300-HDAC2/PD-L1 | Inhibit the immune function of T cell | [88] | ||
| JAK | K1109 | HDAC1 | HDAC1/JAK1/STAT3/FGL1 | Inhibit the immune function of T cell | [95] | |||
| NNT | K1042 | PCAF | - | IL-1β/PCAF/NNT/NADPH | Inhibit the immune function of T cell | [99] | ||
| Ubiquitination | STING1 | K244 | - | - | STING1/TAX1BP1 | Enhance T cell infiltration | [80] | |
| PD-L1 | - | TRIM28 | - | TRIM28/TBK1/PD-L1 | Inhibit the immune function of T cell | [86] | ||
| LAG3 | K498 | c-Cbl, Cbl-b | - | FGL1/LAG3/c-Cbl, Cbl-b | Inhibit the immune function of T cell | [94] | ||
| SUMOylation | SHP1 | K127 | - | - | SHP1/TRAF6-STING/p65 | Enhance T cell infiltration | [81] | |
| Methylation | PD-L1 | K162 | SETD7 | LSD2 | SETD7-LSD2/PD-L1 | Inhibit the immune function of T cell | [89] | |
| Macrophage | SUMOylation | PKM2 | - | UBC9 | - | UBC9/PKM2/ARRDC1 | Promote monocyte-to-TAM differentiation | [103] |
| Ubiquitination | CD47 | - | - | USP2 | USP2/CD47/SIRPα | Enhance macrophage phagocytosis | [105] | |
| NLRP7 | K379 | - | USP10 | USP10/NLRP7/NF-κB/CCL2 | Promote M2 polarization of macrophages | [113] | ||
| Neddylation | SHP2 | K358, K364 | - | SENP8 | SENP8/SHP2/SIPRα | Enhance macrophage phagocytosis | [107] | |
| Succinylation | PDHA1 | K83 | DLST | - | PDHA1/α-KG/OXGR1/MAPK | Suppress macrophage antigen presentation | [109] | |
| Lactylation | ENSA | K63 | P300 | - | ENSA/PP2A/STAT3/CCL2 | Promote M2 polarization of macrophages | [124] | |
| Treg | Lactylation | MOESIN | K72 | - | - | MOESIN/TGF-β/SMAD3 | Promote Tregs accumulation | [125] |
| APOC2 | K70 | P300 | SIRT1, HDAC3 | P300/APOC2/FFA | Promote Tregs accumulation | [126] | ||
| Succinylation | BACH2 | K172 | - | SENP3 | SENP3/BACH2 | Promote Tregs accumulation | [115] | |
| MDSC | Ubiquitination | RIPK1 | - | TRIM28 | - | TRIM28/RIPK1/NF-Κb/CXCL1 | Enhance MDSCs infiltration | [117] |
| Acetylation | SMAD3 | K20, K117 | KAT6A | - | KAT6A/SMAD3/TRIM24 | Enhance MDSCs infiltration | [118] | |
| NK cell | Acetylation | ATF3 | K136, K139 | - | HDAC3 | HDAC3/ATF3/CXCL12 | Enhance NK cells infiltration | [120] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sun, K.; Xiao, S.; Huang, Q.; Zhang, S.; Li, Q.; Hu, C. Non-Histone Lysine Modifications in Tumor Microenvironment: Mechanisms and Therapeutic Opportunities. Int. J. Mol. Sci. 2025, 26, 11229. https://doi.org/10.3390/ijms262211229
Sun K, Xiao S, Huang Q, Zhang S, Li Q, Hu C. Non-Histone Lysine Modifications in Tumor Microenvironment: Mechanisms and Therapeutic Opportunities. International Journal of Molecular Sciences. 2025; 26(22):11229. https://doi.org/10.3390/ijms262211229
Chicago/Turabian StyleSun, Kai, Shuying Xiao, Qibo Huang, Suhang Zhang, Qilin Li, and Chuanyu Hu. 2025. "Non-Histone Lysine Modifications in Tumor Microenvironment: Mechanisms and Therapeutic Opportunities" International Journal of Molecular Sciences 26, no. 22: 11229. https://doi.org/10.3390/ijms262211229
APA StyleSun, K., Xiao, S., Huang, Q., Zhang, S., Li, Q., & Hu, C. (2025). Non-Histone Lysine Modifications in Tumor Microenvironment: Mechanisms and Therapeutic Opportunities. International Journal of Molecular Sciences, 26(22), 11229. https://doi.org/10.3390/ijms262211229




