Theoretical Explanation for the Rarity of Antibody-Dependent Enhancement of Infection (ADE) in COVID-19
Abstract
1. Introduction
2. Results
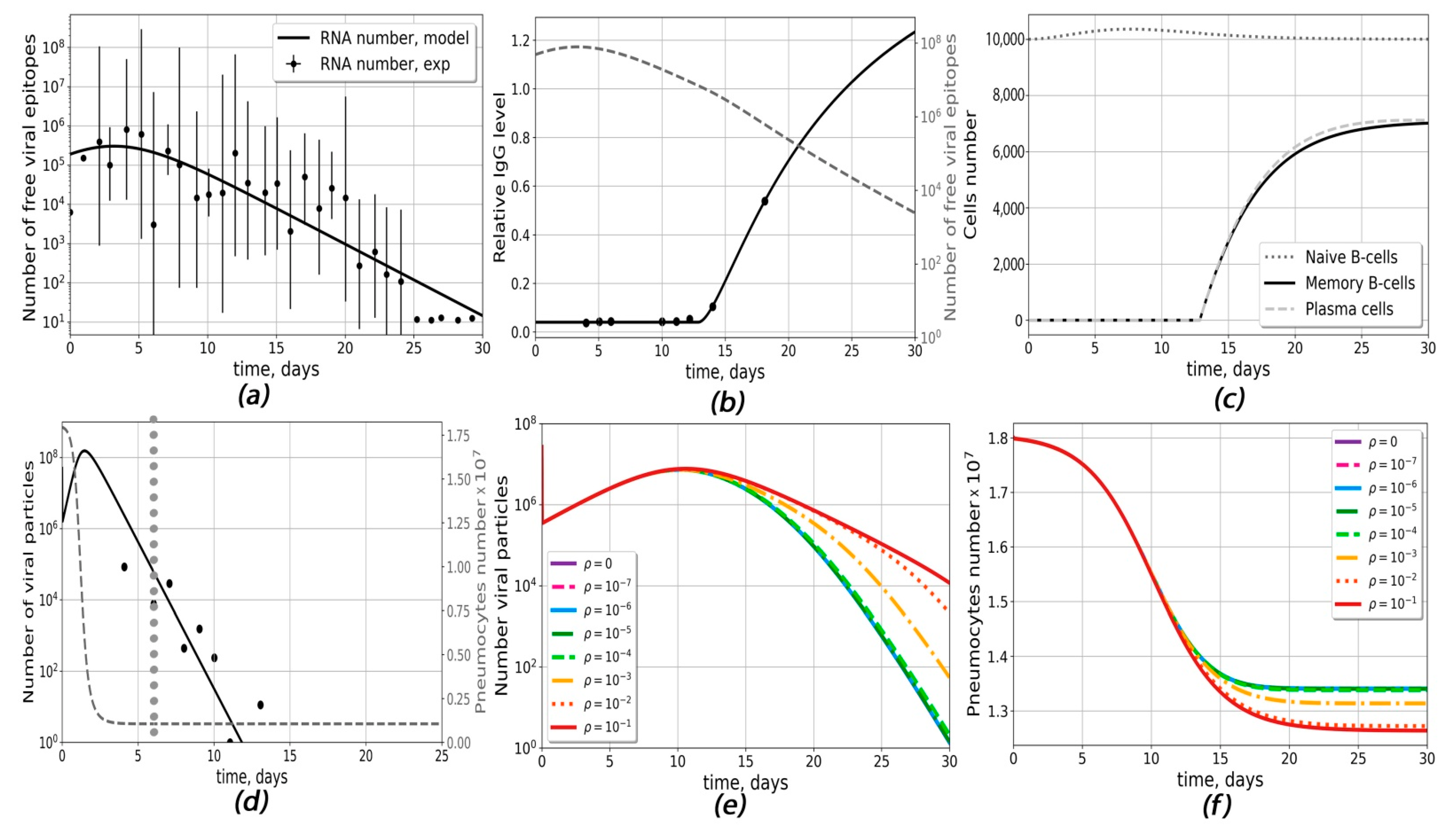
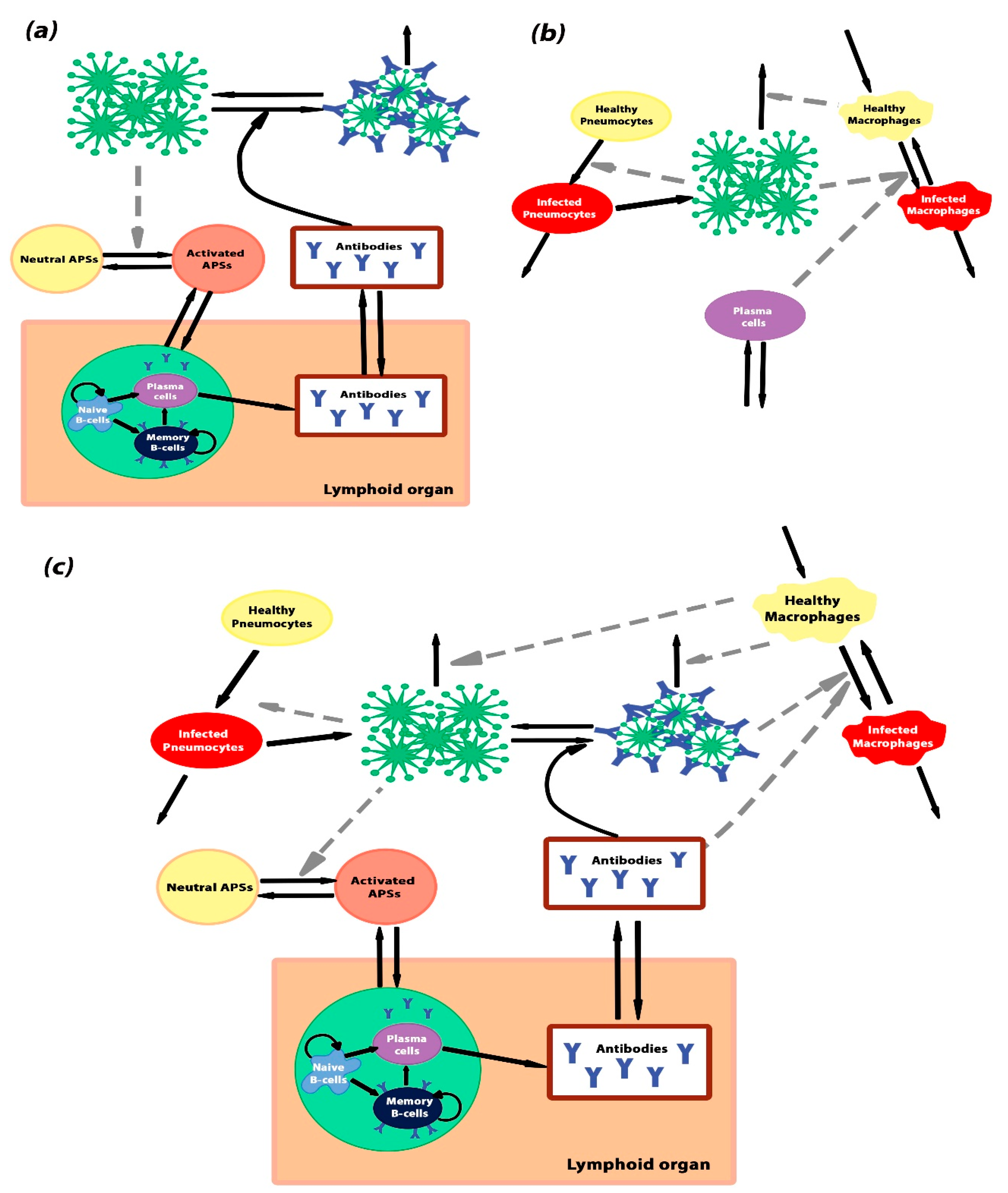
2.1. Construction of Antibody Response Generation Module (Model Ab)
2.2. The Infection Propagation Model (Model Cv) Suggests Possibilities of ADE Occurrence in COVID-19
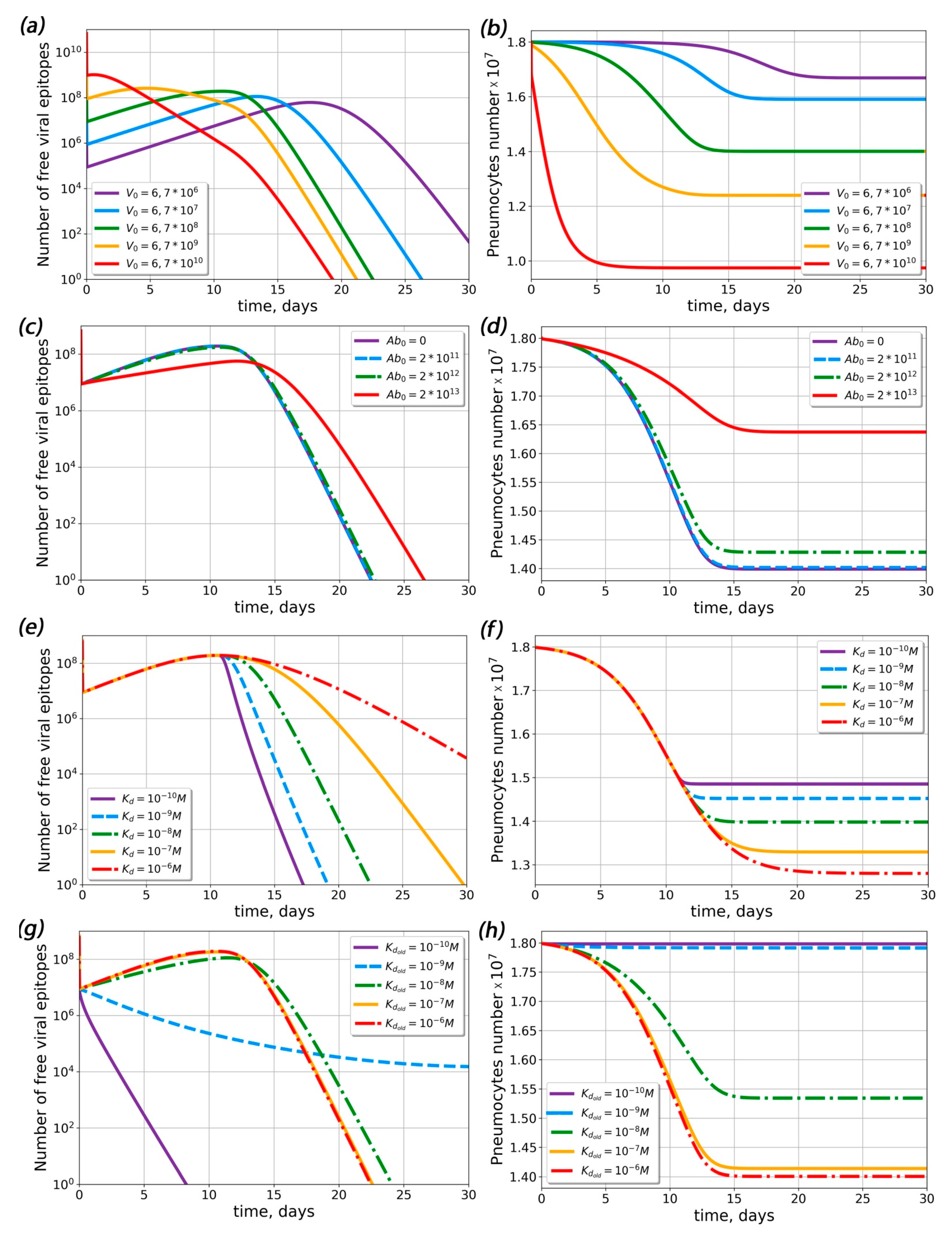
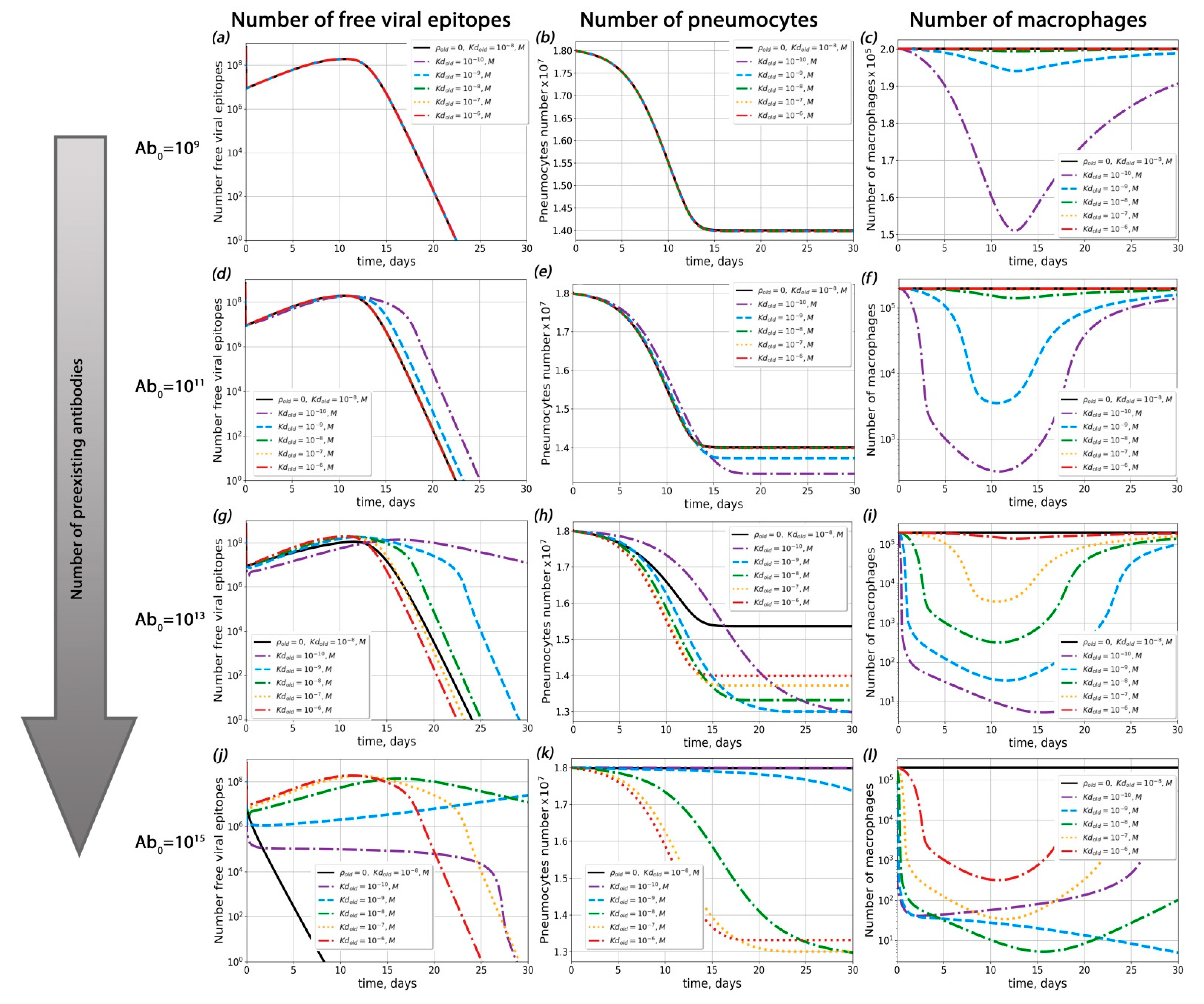
2.3. The Complete Model Is Capable of Describing Viral Load and Assessing the Influence of Pre-Existing Antibody Concentration on the Course of Disease
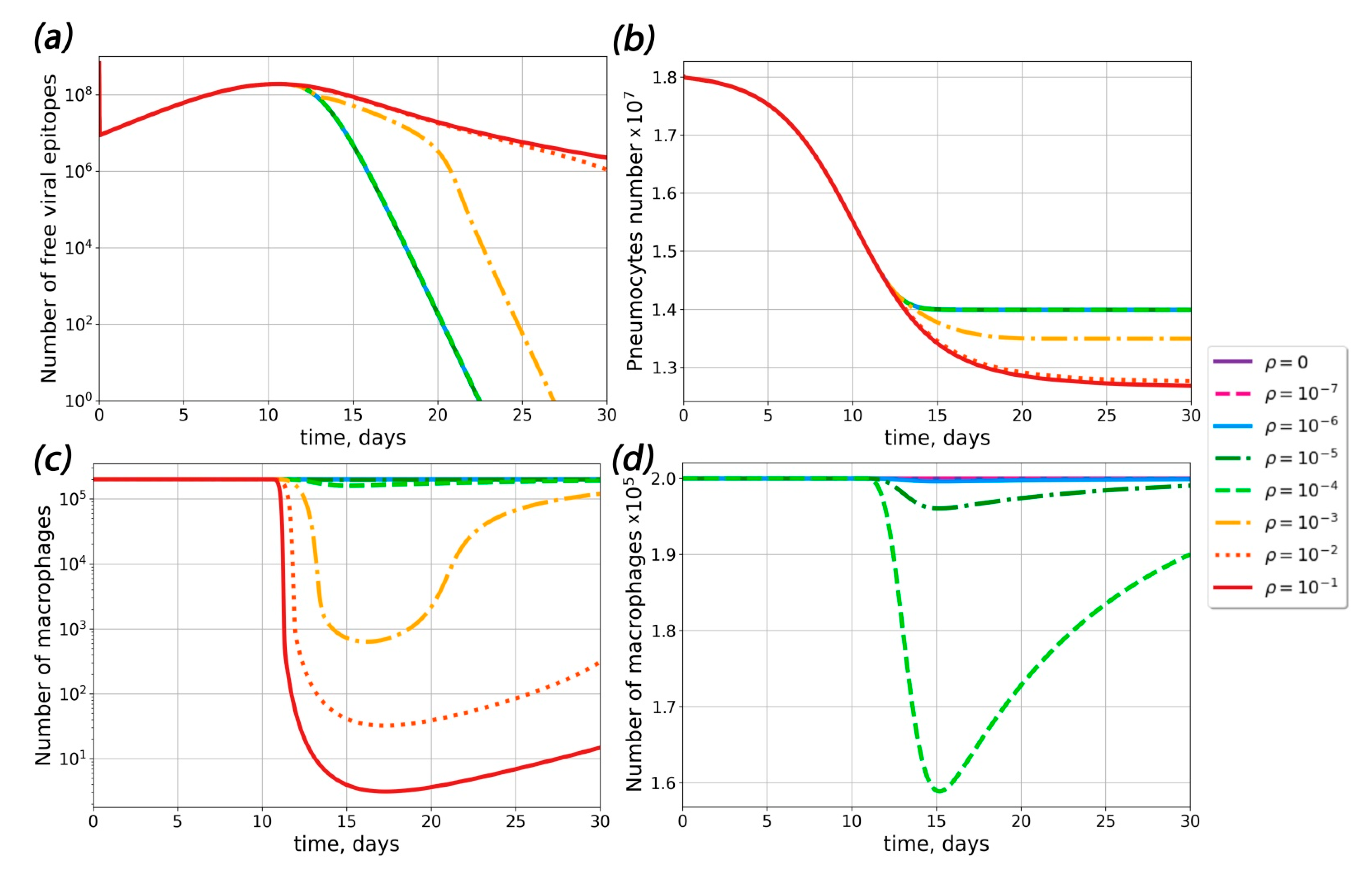
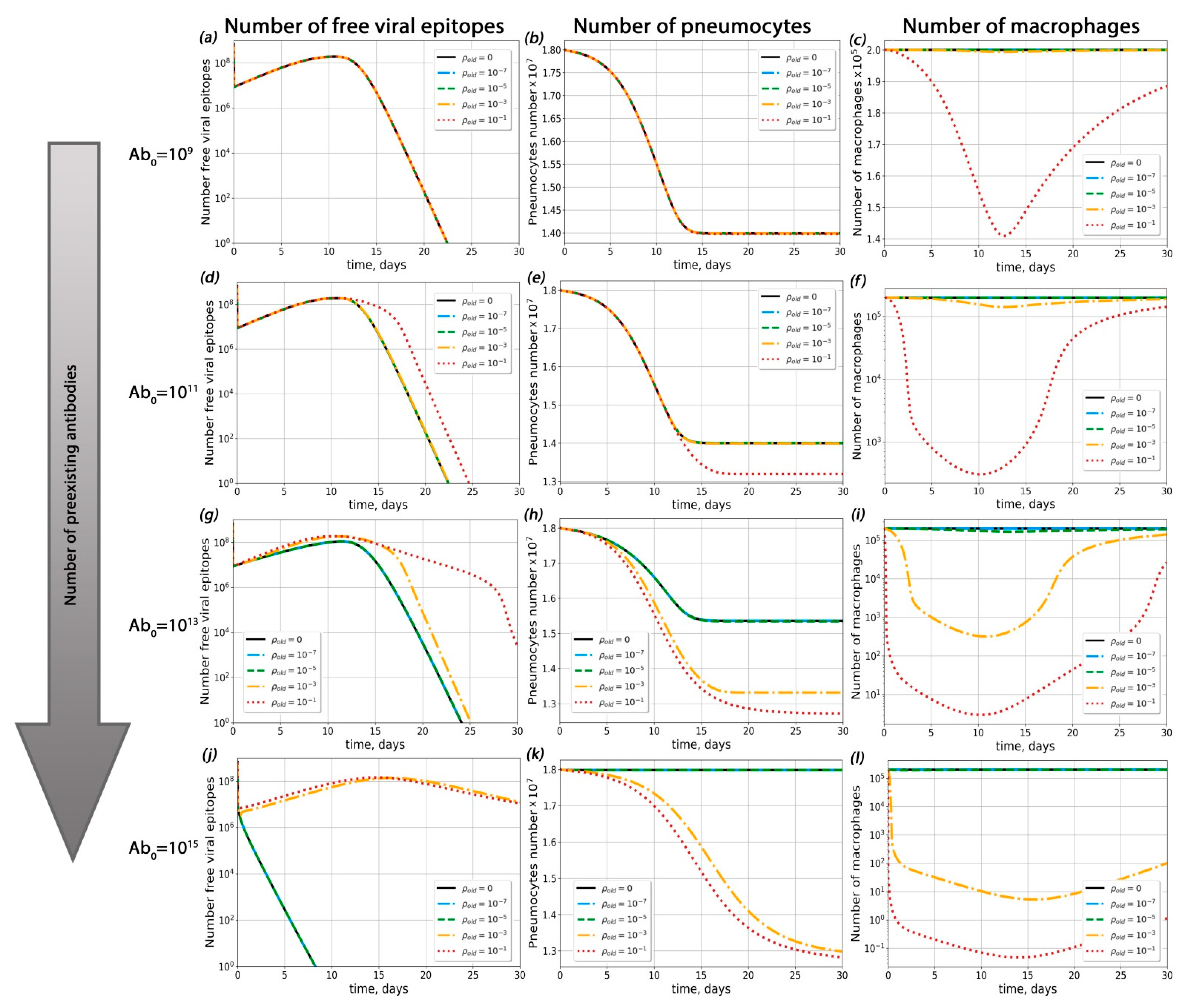
2.4. Theoretical Conditions for ADE Occurrence in COVID-19
3. Discussion
4. Materials and Methods
4.1. General Principles for the Construction of the Mathematical Models
4.2. Model Ab—The Antibody Response
4.3. Model Cv—Propagation of Infection
4.4. Complete Model—A Combination of Model Ab and Model Cv
4.5. Model Integration, Analysis and Parameter Estimation
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ACE 2 | Angiotensin-converting enzyme 2 |
| ADE | Antibody-dependent enhancement |
| APS | Antigen-presenting site |
| Bm | Memory B-cell |
| COVID-19 | Coronavirus disease (COVID-19) |
| HIV | Human immunodeficiency viruses |
| PCR | Polymerase chain reaction |
| Pl | Plasma cell |
References
- Wölfel, R.; Corman, V.M.; Guggemos, W.; Seilmaier, M.; Zange, S.; Müller, M.A.; Niemeyer, D.; Jones, T.C.; Vollmar, P.; Rothe, C.; et al. Virological Assessment of Hospitalized Patients with COVID-2019. Nature 2020, 581, 465–469. [Google Scholar] [CrossRef] [PubMed]
- Ita, K. Coronavirus Disease (COVID-19): Current Status and Prospects for Drug and Vaccine Development. Arch. Med. Res. 2021, 52, 15–24. [Google Scholar] [CrossRef] [PubMed]
- Grigorova, I. Overview of the Neutralizing Antibody and Memory B Cell Response Kinetics in SARS-CoV-2 Convalescent and/or MRNA Vaccinated Individuals. Syst. Biol. Physiol. Rep. 2021, 1, 1. [Google Scholar] [CrossRef]
- Arvin, A.M.; Fink, K.; Schmid, M.A.; Cathcart, A.; Spreafico, R.; Havenar-Daughton, C.; Lanzavecchia, A.; Corti, D.; Virgin, H.W. A Perspective on Potential Antibody-Dependent Enhancement of SARS-CoV-2. Nature 2020, 584, 353–363. [Google Scholar] [CrossRef]
- Tirado, S.M.C.; Yoon, K.-J. Antibody-Dependent Enhancement of Virus Infection and Disease. Viral. Immunol. 2003, 16, 69–86. [Google Scholar] [CrossRef]
- Iwasaki, A.; Yang, Y. The Potential Danger of Suboptimal Antibody Responses in COVID-19. Nat. Rev. Immunol. 2020, 20, 339–341. [Google Scholar] [CrossRef]
- Winarski, K.L.; Tang, J.; Klenow, L.; Lee, J.; Coyle, E.M.; Manischewitz, J.; Turner, H.L.; Takeda, K.; Ward, A.B.; Golding, H.; et al. Antibody-Dependent Enhancement of Influenza Disease Promoted by Increase in Hemagglutinin Stem Flexibility and Virus Fusion Kinetics. Proc. Natl. Acad. Sci. USA 2019, 116, 15194–15199. [Google Scholar] [CrossRef]
- Takada, A.; Feldmann, H.; Ksiazek, T.G.; Kawaoka, Y. Antibody-Dependent Enhancement of Ebola Virus Infection. J. Virol. 2003, 77, 7539–7544. [Google Scholar] [CrossRef]
- Dejnirattisai, W.; Jumnainsong, A.; Onsirisakul, N.; Fitton, P.; Vasanawathana, S.; Limpitikul, W.; Puttikhunt, C.; Edwards, C.; Duangchinda, T.; Supasa, S.; et al. Cross-Reacting Antibodies Enhance Dengue Virus Infection in Humans. Science 2010, 328, 745–748. [Google Scholar] [CrossRef]
- Gorlani, A.; Forthal, D.N. Antibody-Dependent Enhancement and the Risk of HIV Infection. Curr. HIV Res. 2013, 11, 421–426. [Google Scholar] [CrossRef]
- Kulkarni, R. Antibody-Dependent Enhancement of Viral Infections. In Dynamics of Immune Activation in Viral Diseases; Bramhachari, P., Ed.; Springer: Singapore, 2020; pp. 9–41. [Google Scholar] [CrossRef]
- Tay, M.Z.; Wiehe, K.; Pollara, J. Antibody-Dependent Cellular Phagocytosis in Antiviral Immune Responses. Front. Immunol. 2019, 10, 332. [Google Scholar] [CrossRef]
- Taylor, A.; Foo, S.-S.; Bruzzone, R.; Dinh, L.V.; King, N.J.C.; Mahalingam, S. Fc Receptors in Antibody-Dependent Enhancement of Viral Infections. Immunol. Rev. 2015, 268, 340–364. [Google Scholar] [CrossRef]
- Coperchini, F.; Chiovato, L.; Croce, L.; Magri, F.; Rotondi, M. The Cytokine Storm in COVID-19: An Overview of the Involvement of the Chemokine/Chemokine-Receptor System. Cytokine Growth Factor Rev. 2020, 53, 25–32. [Google Scholar] [CrossRef]
- Wang, S.; Wang, J.; Yu, X.; Jiang, W.; Chen, S.; Wang, R.; Wang, M.; Jiao, S.; Yang, Y.; Wang, W.; et al. Antibody-Dependent Enhancement (ADE) of SARS-CoV-2 Pseudoviral Infection Requires FcγRIIB and Virus-Antibody Complex with Bivalent Interaction. Commun. Biol. 2022, 5, 1–9. [Google Scholar] [CrossRef]
- Bournazos, S.; Gupta, A.; Ravetch, J.V. The Role of IgG Fc Receptors in Antibody-Dependent Enhancement. Nat. Rev. Immunol. 2020, 20, 633–643. [Google Scholar] [CrossRef]
- Jaume, M.; Yip, M.S.; Cheung, C.Y.; Leung, H.L.; Li, P.H.; Kien, F.; Dutry, I.; Callendret, B.; Escriou, N.; Altmeyer, R.; et al. Anti-Severe Acute Respiratory Syndrome Coronavirus Spike Antibodies Trigger Infection of Human Immune Cells via a PH- and Cysteine Protease-Independent FcγR Pathway. J. Virol. 2011, 85, 10582–10597. [Google Scholar] [CrossRef]
- Kam, Y.W.; Kien, F.; Roberts, A.; Cheung, Y.C.; Lamirande, E.W.; Vogel, L.; Chu, S.L.; Tse, J.; Guarner, J.; Zaki, S.R.; et al. Antibodies against Trimeric S Glycoprotein Protect Hamsters against SARS-CoV Challenge despite Their Capacity to Mediate FcgammaRII-Dependent Entry into B Cells in Vitro. Vaccine 2007, 25, 729–740. [Google Scholar] [CrossRef]
- Wan, Y.; Shang, J.; Sun, S.; Tai, W.; Chen, J.; Geng, Q.; He, L.; Chen, Y.; Wu, J.; Shi, Z.; et al. Molecular Mechanism for Antibody-Dependent Enhancement of Coronavirus Entry. J. Virol. 2019, 94, e02015-19. [Google Scholar] [CrossRef]
- Eroshenko, N.; Gill, T.; Keaveney, M.K.; Church, G.M.; Trevejo, J.M.; Rajaniemi, H. Implications of Antibody-Dependent Enhancement of Infection for SARS-CoV-2 Countermeasures. Nat. Biotechnol. 2020, 38, 789–791. [Google Scholar] [CrossRef]
- Lee, W.S.; Wheatley, A.K.; Kent, S.J.; DeKosky, B.J. Antibody-Dependent Enhancement and SARS-CoV-2 Vaccines and Therapies. Nat. Microbiol. 2020, 5, 1185–1191. [Google Scholar] [CrossRef]
- Ricke, D.O. Two Different Antibody-Dependent Enhancement (ADE) Risks for SARS-CoV-2 Antibodies. Front. Immunol. 2021, 12, 443. [Google Scholar] [CrossRef]
- Banerjee, A.; Nasir, J.A.; Budylowski, P.; Yip, L.; Aftanas, P.; Christie, N.; Ghalami, A.; Baid, K.; Raphenya, A.R.; Hirota, J.A.; et al. Isolation, Sequence, Infectivity, and Replication Kinetics of Severe Acute Respiratory Syndrome Coronavirus 2. Emerg. Infect. Dis. 2020, 26, 2054–2063. [Google Scholar] [CrossRef]
- Abassi, Z.; Knaney, Y.; Karram, T.; Heyman, S.N. The Lung Macrophage in SARS-CoV-2 Infection: A Friend or a Foe? Front. Immunol. 2020, 11, 1312. [Google Scholar] [CrossRef]
- Junqueira, C.; Crespo, Â.; Ranjbar, S.; de Lacerda, L.B.; Lewandrowski, M.; Ingber, J.; Parry, B.; Ravid, S.; Clark, S.; Schrimpf, M.R.; et al. FcγR-Mediated SARS-CoV-2 Infection of Monocytes Activates Inflammation. Nature 2022, 606, 576–584. [Google Scholar] [CrossRef]
- Hui, K.P.Y.; Cheung, M.-C.; Perera, R.A.P.M.; Ng, K.-C.; Bui, C.H.T.; Ho, J.C.W.; Ng, M.M.T.; Kuok, D.I.T.; Shih, K.C.; Tsao, S.-W.; et al. Tropism, Replication Competence, and Innate Immune Responses of the Coronavirus SARS-CoV-2 in Human Respiratory Tract and Conjunctiva: An Analysis in Ex-Vivo and In-Vitro Cultures. Lancet Respir. Med. 2020, 8, 687–695. [Google Scholar] [CrossRef]
- Zheng, J.; Wang, Y.; Li, K.; Meyerholz, D.K.; Allamargot, C.; Perlman, S. Severe Acute Respiratory Syndrome Coronavirus 2–Induced Immune Activation and Death of Monocyte-Derived Human Macrophages and Dendritic Cells. J. Infect. Dis. 2020, 223, 785–795. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, T.S.; de Sá, K.S.G.; Ishimoto, A.Y.; Becerra, A.; Oliveira, S.; Almeida, L.; Gonçalves, A.V.; Perucello, D.B.; Andrade, W.A.; Castro, R.; et al. Inflammasomes Are Activated in Response to SARS-CoV-2 Infection and Are Associated with COVID-19 Severity in Patients. J. Exp. Med. 2021, 218, e20201707. [Google Scholar] [CrossRef]
- Szekely, L.; Bozoky, B.; Bendek, M.; Ostad, M.; Lavignasse, P.; Haag, L.; Wu, J.; Jing, X.; Gupta, S.; Saccon, E.; et al. Pulmonary Stromal Expansion and Intra-Alveolar Coagulation Are Primary Causes of COVID-19 Death. Heliyon 2021, 7, e07134. [Google Scholar] [CrossRef]
- Desforges, M.; Miletti, T.C.; Gagnon, M.; Talbot, P.J. Activation of Human Monocytes after Infection by Human Coronavirus 229E. Virus Res. 2007, 130, 228–240. [Google Scholar] [CrossRef]
- Gómez-Rial, J.; Rivero-Calle, I.; Salas, A.; Martinón-Torres, F. Role of Monocytes/Macrophages in Covid-19 Pathogenesis: Implications for Therapy. Infect. Drug Resist. 2020, 13, 2485–2493. [Google Scholar] [CrossRef]
- Beeraka, N.M.; Tulimilli, S.V.; Karnik, M.; Sadhu, S.P.; Pragada, R.R.; Aliev, G.; Madhunapantula, S.V. The Current Status and Challenges in the Development of Vaccines and Drugs against Severe Acute Respiratory Syndrome-Corona Virus-2 (SARS-CoV-2). Biomed. Res. Int. 2021, 2021, 8160860. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.; Yan, R.; Liu, M.; Liu, Z.; Wang, Y.; Luan, D.; Wu, K.; Song, Z.; Sun, T.; Ma, Y.; et al. Antibody-Dependent Enhancement (ADE) of SARS-CoV-2 Infection in Recovered COVID-19 Patients: Studies Based on Cellular and Structural Biology Analysis. medRxiv 2020. [Google Scholar] [CrossRef]
- To, K.K.-W.; Tsang, O.T.-Y.; Leung, W.-S.; Tam, A.R.; Wu, T.-C.; Lung, D.C.; Yip, C.C.-Y.; Cai, J.-P.; Chan, J.M.-C.; Chik, T.S.-H.; et al. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: An observational cohort study. Lancet Infect. Dis. 2020, 20, 565–574. [Google Scholar] [CrossRef]
- Zhu, N.; Wang, W.; Liu, Z.; Liang, C.; Wang, W.; Ye, F.; Huang, B.; Zhao, L.; Wang, H.; Zhou, W.; et al. Morphogenesis and Cytopathic Effect of SARS-CoV-2 Infection in Human Airway Epithelial Cells. Nat. Commun. 2020, 11, 3910. [Google Scholar] [CrossRef]
- Cerón Gómez, M.; Yang, H.M. A Simple Mathematical Model to Describe Antibody-Dependent Enhancement in Heterologous Secondary Infection in Dengue. Math. Med. Biol. 2019, 36, 411–438. [Google Scholar] [CrossRef]
- Fain, B.; Dobrovolny, H.M. Initial Inoculum and the Severity of COVID-19: A Mathematical Modeling Study of the Dose-Response of SARS-CoV-2 Infections. Epidemiologia 2020, 1, 5–15. [Google Scholar] [CrossRef]
- Lee, H.Y.; Topham, D.J.; Park, S.Y.; Hollenbaugh, J.; Treanor, J.; Mosmann, T.R.; Jin, X.; Ward, B.M.; Miao, H.; Holden-Wiltse, J.; et al. Simulation and Prediction of the Adaptive Immune Response to Influenza a Virus Infection. J. Virol. 2009, 83, 7151–7165. [Google Scholar] [CrossRef]
- Danchin, A.; Pagani-Azizi, O.; Turinici, G.; Yahiaoui, G. COVID-19 Adaptive Humoral Immunity Models: Weakly Neutralizing Versus Antibody-Disease Enhancement Scenarios. Acta Biotheor. 2022, 70, 23. [Google Scholar] [CrossRef]
- Schieffelin, J.S.; Costin, J.M.; Nicholson, C.O.; Orgeron, N.M.; Fontaine, K.A.; Isern, S.; Michael, S.F.; Robinson, J.E. Neutralizing and Non-Neutralizing Monoclonal Antibodies against Dengue Virus E Protein Derived from a Naturally Infected Patient. Virol. J. 2010, 7, 28. [Google Scholar] [CrossRef]
- Hegazy, A.N.; Krönke, J.; Angermair, S.; Schwartz, S.; Weidinger, C.; Keller, U.; Treskatsch, S.; Siegmund, B.; Schneider, T. Anti-SARS-CoV2 Antibody-Mediated Cytokine Release Syndrome in a Patient with Acute Promyelocytic Leukemia. BMC Infect. Dis. 2022, 22, 537. [Google Scholar] [CrossRef]
- Nechipurenko, Y.D.; Anashkina, A.A.; Matveeva, O.V. Change of Antigenic Determinants of SARS-CoV-2 Virus S-Protein as a Possible Cause of Antibody-Dependent Enhancement of Virus Infection and Cytokine Storm. Biophysics 2020, 65, 703–709. [Google Scholar] [CrossRef]
- Zaichuk, T.A.; Nechipurenko, Y.D.; Adzhubey, A.A.; Onikienko, S.B.; Chereshnev, V.A.; Zainutdinov, S.S.; Kochneva, G.V.; Netesov, S.V.; Matveeva, O.V. The Challenges of Vaccine Development against Betacoronaviruses: Antibody Dependent Enhancement and Sendai Virus as a Possible Vaccine Vector. Mol. Biol. 2020, 54, 812–826. [Google Scholar] [CrossRef]
- Yang, X.; Zhang, X.; Zhao, X.; Yuan, M.; Zhang, K.; Dai, J.; Guan, X.; Qiu, H.-J.; Li, Y. Antibody-Dependent Enhancement: ″Evil″ Antibodies Favorable for Viral Infections. Viruses 2022, 14, 1739. [Google Scholar] [CrossRef]
- Okuya, K.; Hattori, T.; Saito, T.; Takadate, Y.; Sasaki, M.; Furuyama, W.; Marzi, A.; Ohiro, Y.; Konno, S.; Hattori, T.; et al. Multiple Routes of Antibody-Dependent Enhancement of SARS-CoV-2 Infection. Microbiol. Spectr. 2022, 10, e01553-21. [Google Scholar] [CrossRef]
- Clark, N.M.; Janaka, S.K.; Hartman, W.; Stramer, S.; Goodhue, E.; Weiss, J.; Evans, D.T.; Connor, J.P. Anti-SARS-CoV-2 IgG and IgA Antibodies in COVID-19 Convalescent Plasma Do Not Enhance Viral Infection. PLoS ONE 2022, 17, e0257930. [Google Scholar] [CrossRef]
- Dassarma, B.; Tripathy, S.; Matsabisa, M. Emergence of Ancient Convalescent Plasma (CP) Therapy: To Manage COVID-19 Pandemic. Transfus. Clin. Et Biol. 2021, 28, 123–127. [Google Scholar] [CrossRef]
- Turner, J.S.; Benet, Z.L.; Grigorova, I.L. Signals 1, 2 and B Cell Fate or: Where, When and for How Long? Immunol. Rev. 2020, 296, 9–23. [Google Scholar] [CrossRef]
- Reis, R.F.; Pigozzo, A.B.; Bonin, C.R.B.; de Melo Quintela, B.; Pompei, L.T.; Vieira, A.C.; de Lima e Silva, L.; Xavier, M.P.; Weber dos Santos, R.; Lobosco, M. A Validated Mathematical Model of the Cytokine Release Syndrome in Severe COVID-19. Front. Mol. Biosci. 2021, 8, 680. [Google Scholar] [CrossRef]
- Goutelle, S.; Maurin, M.; Rougier, F.; Barbaut, X.; Bourguignon, L.; Ducher, M.; Maire, P. The Hill Equation: A Review of Its Capabilities in Pharmacological Modelling. Fundam. Clin. Pharmacol. 2008, 22, 633–648. [Google Scholar] [CrossRef]
- Bonin, C.R.B.; Fernandes, G.C.; dos Santos, R.W.; Lobosco, M. A Qualitatively Validated Mathematical-Computational Model of the Immune Response to the Yellow Fever Vaccine. BMC Immunol. 2018, 19, 15. [Google Scholar] [CrossRef]
- Nguyen, D.C.; Joyner, C.J.; Sanz, I.; Lee, F.E.-H. Factors Affecting Early Antibody Secreting Cell Maturation into Long-Lived Plasma Cells. Front. Immunol. 2019, 10, 2138. [Google Scholar] [CrossRef]
- Wang, S.; Pan, Y.; Wang, Q.; Miao, H.; Brown, A.N.; Rong, L. Modeling the Viral Dynamics of SARS-CoV-2 Infection. Math. Biosci. 2020, 328, 108438. [Google Scholar] [CrossRef] [PubMed]
- Ajmeriya, S.; Kumar, A.; Karmakar, S.; Rana, S.; Singh, H. Neutralizing Antibodies and Antibody-Dependent Enhancement in COVID-19: A Perspective. J. Indian Inst. Sci. 2022, 102, 1–17. [Google Scholar] [CrossRef]
- Bergmann, F.T.; Hoops, S.; Klahn, B.; Kummer, U.; Mendes, P.; Pahle, J.; Sahle, S. COPASI and Its Applications in Biotechnology. J. Biotechnol. 2017, 261, 215–220. [Google Scholar] [CrossRef]
- Petzold, L. Automatic Selection of Methods for Solving Stiff and Nonstiff Systems of Ordinary Differential Equations. SIAM J. Sci. Stat. Comput. 1983, 4, 136–148. [Google Scholar] [CrossRef]
- Kapellos, T.S.; Taylor, L.; Lee, H.; Cowley, S.A.; James, W.S.; Iqbal, A.J.; Greaves, D.R. A Novel Real Time Imaging Platform to Quantify Macrophage Phagocytosis. Biochem. Pharmacol. 2016, 116, 107–119. [Google Scholar] [CrossRef]
- Bromage, E.; Stephens, R.; Hassoun, L. The Third Dimension of ELISPOTs: Quantifying Antibody Secretion from Individual Plasma Cells. J. Immunol. Methods 2009, 346, 75–79. [Google Scholar] [CrossRef]
- Seydoux, E.; Homad, L.J.; MacCamy, A.J.; Parks, K.R.; Hurlburt, N.K.; Jennewein, M.F.; Akins, N.R.; Stuart, A.B.; Wan, Y.-H.; Feng, J.; et al. Characterization of Neutralizing Antibodies from a SARS-CoV-2 Infected Individual. bioRxiv, 2020; preprint. [Google Scholar] [CrossRef]
- Itoua Maïga, R.; Bonnaure, G.; Tremblay Rochette, J.; Néron, S. Human CD38hiCD138+ Plasma Cells Can Be Generated In Vitro from CD40-Activated Switched-Memory B Lymphocytes. J. Immunol. Res. 2014, 2014, 635108. [Google Scholar] [CrossRef] [PubMed]
- Amanna, I.J.; Slifka, M.K. Mechanisms That Determine Plasma Cell Lifespan and the Duration of Humoral Immunity. Immunol. Rev. 2010, 236, 125–138. [Google Scholar] [CrossRef]
- Marino, S.; Hogue, I.B.; Ray, C.J.; Kirschner, D.E. A Methodology for Performing Global Uncertainty and Sensitivity Analysis in Systems Biology. J. Theor. Biol. 2008, 254, 178–196. [Google Scholar] [CrossRef] [PubMed]
- Saltelli, A.; Ratto, M.; Tarantola, S.; Campolongo, F. Sensitivity Analysis for Chemical Models. Chem. Rev. 2005, 105, 2811–2828. [Google Scholar] [CrossRef] [PubMed]
- Sender, R.; Bar-On, Y.M.; Gleizer, S.; Bernshtein, B.; Flamholz, A.; Phillips, R.; Milo, R. The total number and mass of SARS-CoV-2 virions. Proc. Natl. Acad. Sci. USA 2021, 118, e2024815118. [Google Scholar] [CrossRef] [PubMed]
- Chi, X.; Liu, X.; Wang, C.; Zhang, X.; Li, X.; Hou, J.; Ren, L.; Jin, Q.; Wang, J.; Yang, W. Humanized single domain antibodies neutralize SARS-CoV-2 by targeting the spike receptor binding domain. Nat. Commun. 2020, 11, 4528. [Google Scholar] [CrossRef]
- Kamath, A.T.; Henri, S.; Battye, F.; Tough, D.F.; Shortman, K. Developmental kinetics and lifespan of dendritic cells in mouse lymphoid organs. Blood 2002, 100, 1734–1741. [Google Scholar] [CrossRef]
- Sathe, P.; Shortman, K. The steady-state development of splenic dendritic cells. Mucosal Immunol. 2008, 1, 425–431. [Google Scholar] [CrossRef]
- Liu, K.; Waskow, C.; Liu, X.; Yao, K.; Hoh, J.; Nussenzweig, M. Origin of dendritic cells in peripheral lymphoid organs of mice. Nat. Immunol. 2007, 8, 578–583. [Google Scholar] [CrossRef]
- Hartley, G.E.; Edwards, E.S.; Aui, P.M.; Varese, N.; Stojanovic, S.; McMahon, J.; Peleg, A.Y.; Boo, I.; Drummer, H.E.; Hogarth, P.M.; et al. Rapid generation of durable B cell memory to SARS-CoV-2 spike and nucleocapsid proteins in COVID-19 and convalescence. Sci. Immunol. 2020, 5, eabf8891. [Google Scholar] [CrossRef]
- Mamani-Matsuda, M.; Cosma, A.; Weller, S.; Faili, A.; Staib, C.; Garçon, L.; Hermine, O.; Beyne-Rauzy, O.; Fieschi, C.; Pers, J.O.; et al. The human spleen is a major reservoir for long-lived vaccinia virus–specific memory B cells. Blood 2008, 111, 4653–4659. [Google Scholar] [CrossRef]
- Bujoreanu, I.; Gupta, V. Anatomy, Lymph Nodes. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2022. Available online: http://www.ncbi.nlm.nih.gov/books/NBK557717/ (accessed on 18 July 2022).
- Alberts, B.; Johnson, A.; Lewis, J.; Raff, M.; Roberts, K.; Walter, P. B Cells and Antibodies. In Molecular Biology of the Cell, 4th ed.; Garland Science: New Yoek, NY, USA, 2002. Available online: https://www.ncbi.nlm.nih.gov/books/NBK26884/ (accessed on 19 July 2022).
- Hibi, T.; Dosch, H.M. Limiting dilution analysis of the B cell compartment in human bone marrow. Eur. J. Immunol. 1986, 16, 139–145. [Google Scholar] [CrossRef]
- Arvola, M.; Gustafsson, E.; Svensson, L.; Jansson, L.; Holmdahl, R.; Heyman, B.; Okabe, M.; Mattsson, R. Immunoglobulin-Secreting Cells of Maternal Origin Can Be Detected in B Cell-Deficient Mice1. Biol. Reprod. 2000, 63, 1817–1824. [Google Scholar] [CrossRef]
- Pawelek, K.A.; Huynh, G.T.; Quinlivan, M.; Cullinane, A.; Rong, L.; Perelson, A.S. Modeling Within-Host Dynamics of Influenza Virus Infection Including Immune Responses. PLoS Comput. Biol. 2012, 8, e1002588. [Google Scholar] [CrossRef]
- Yap, T.F.; Liu, Z.; Shveda, R.A.; Preston, D.J. A predictive model of the temperature-dependent inactivation of coronaviruses. Appl. Phys. Lett. 2020, 117, 060601. [Google Scholar] [CrossRef]
- Laidlaw, B.J.; Cyster, J.G. Transcriptional regulation of memory B cell differentiation. Nat. Rev. Immunol. 2021, 21, 209–220. [Google Scholar] [CrossRef]
- Ke, Z.; Oton, J.; Qu, K.; Cortese, M.; Zila, V.; McKeane, M.; Nakane, T.; Zivanov, J.; Neufeldt, C.J.; Cerikan, B.; et al. Structures and distributions of SARS-CoV-2 spike proteins on intact virions. Nature 2020, 588, 498–502. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Boldova, A.E.; Korobkin, J.D.; Nechipurenko, Y.D.; Sveshnikova, A.N. Theoretical Explanation for the Rarity of Antibody-Dependent Enhancement of Infection (ADE) in COVID-19. Int. J. Mol. Sci. 2022, 23, 11364. https://doi.org/10.3390/ijms231911364
Boldova AE, Korobkin JD, Nechipurenko YD, Sveshnikova AN. Theoretical Explanation for the Rarity of Antibody-Dependent Enhancement of Infection (ADE) in COVID-19. International Journal of Molecular Sciences. 2022; 23(19):11364. https://doi.org/10.3390/ijms231911364
Chicago/Turabian StyleBoldova, Anna E., Julia D. Korobkin, Yury D. Nechipurenko, and Anastasia N. Sveshnikova. 2022. "Theoretical Explanation for the Rarity of Antibody-Dependent Enhancement of Infection (ADE) in COVID-19" International Journal of Molecular Sciences 23, no. 19: 11364. https://doi.org/10.3390/ijms231911364
APA StyleBoldova, A. E., Korobkin, J. D., Nechipurenko, Y. D., & Sveshnikova, A. N. (2022). Theoretical Explanation for the Rarity of Antibody-Dependent Enhancement of Infection (ADE) in COVID-19. International Journal of Molecular Sciences, 23(19), 11364. https://doi.org/10.3390/ijms231911364







