Bioinformatic Exploration of the Targets of Xylem Sap miRNAs in Maize under Cadmium Stress
Abstract
1. Introduction
2. Results
2.1. High-Throughput Sequencing of sRNAs in Maize Xylem Sap
2.2. Identification of Xylem Sap Cd-Responsive miRNAs
2.3. Target Predictions of Xylem Sap miRNAs
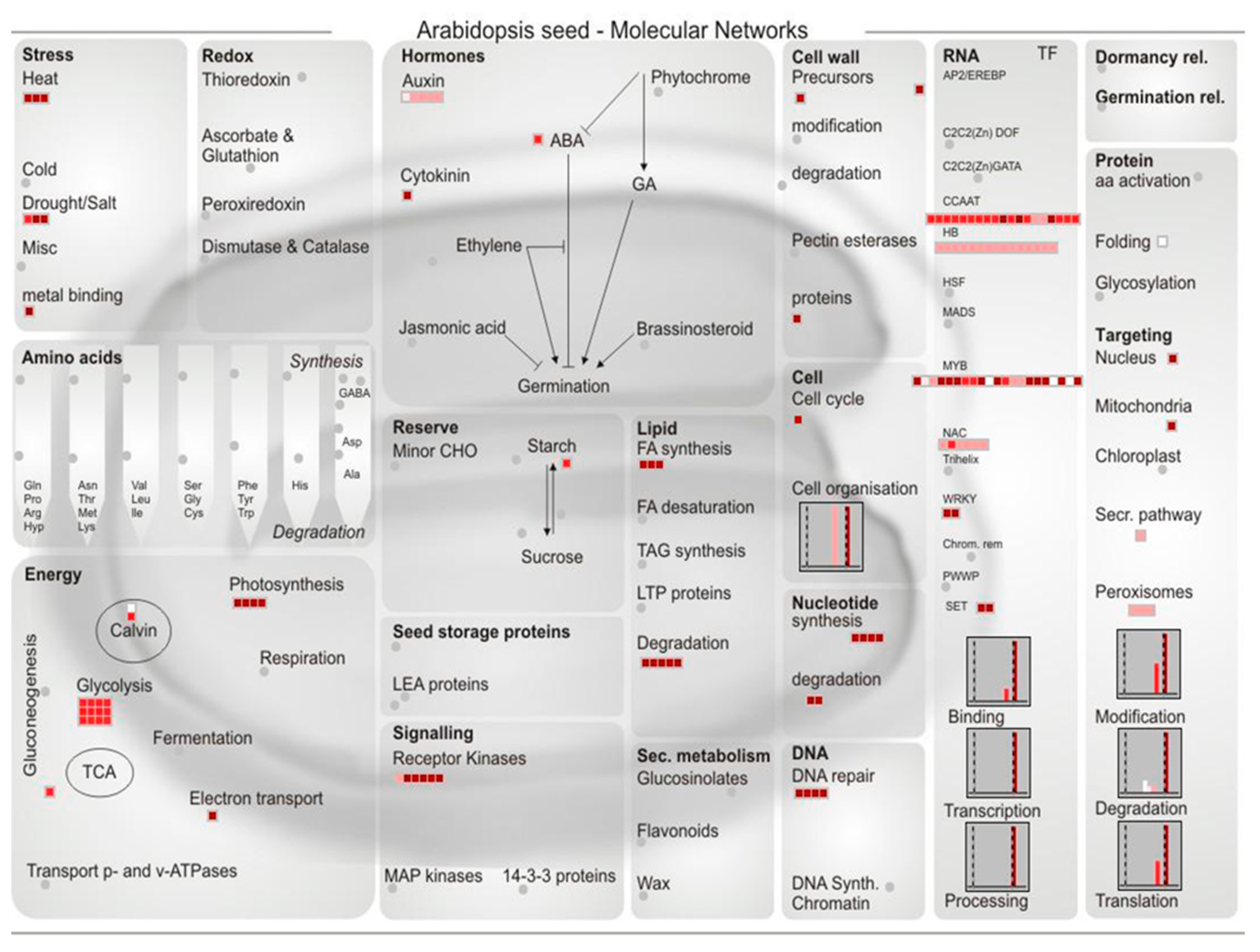
2.4. The Function Classification of the Predicted miRNAs Targets
2.5. The miRNAs Cleavable Targets
3. Discussion
3.1. The Long-Distance Transport of miRNAs
3.2. The Potential Cleavable Targets of miRNAs in Xylem Sap
4. Materials and Methods
4.1. Plant Materials and Cd Treatment
4.2. Sampling of Xylem Sap
4.3. Small RNA Library Preparation and Sequencing
4.4. Small RNA Analysis
4.5. Identification of Known and Novel miRNAs
4.6. Differential Expression Analysis of sRNAs Under Cd Stress
4.7. The Prediction of miRNA Targets
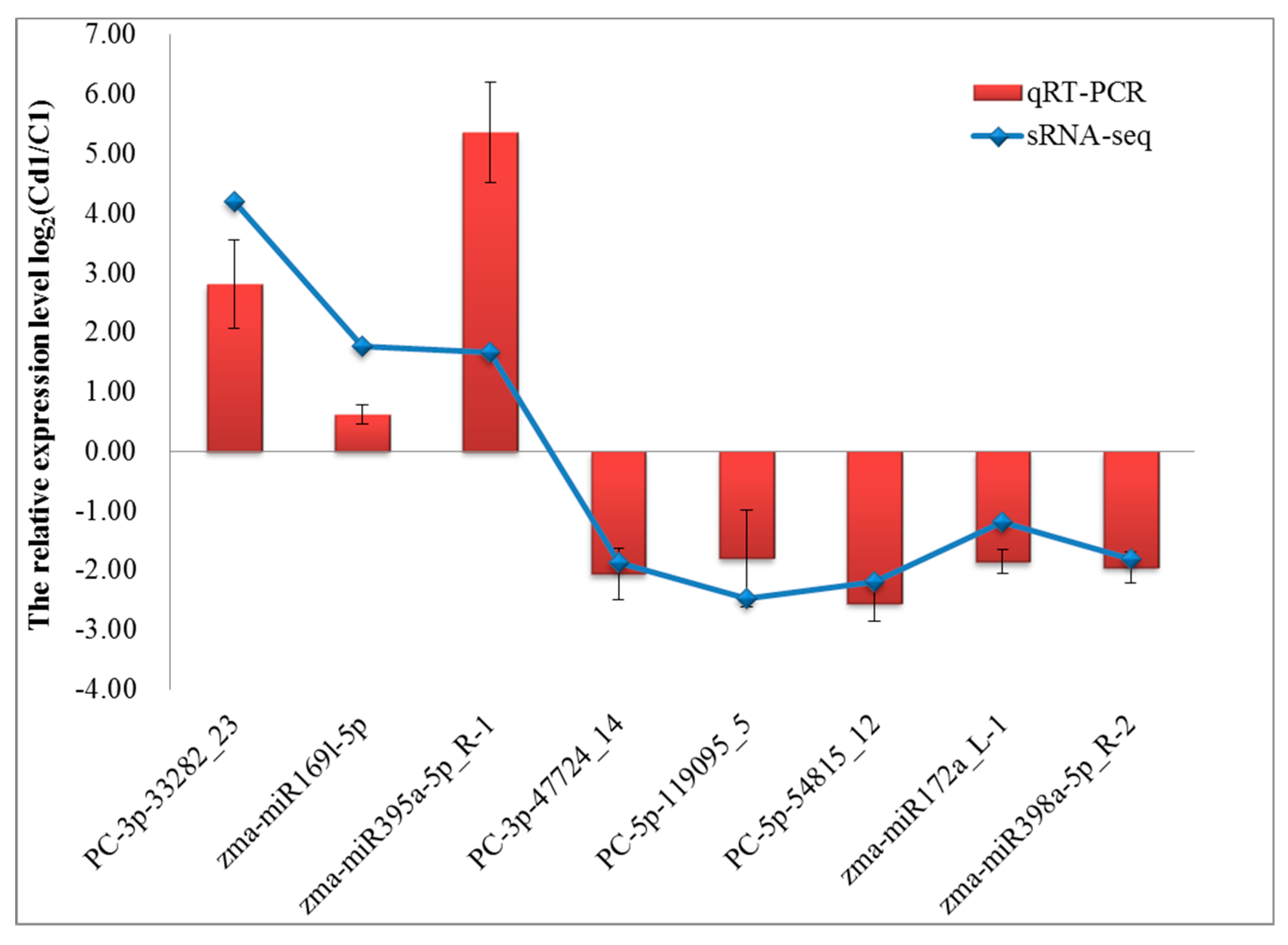
4.8. The Confirmation of sRNAs Expression by qRT-PCR
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Uraguchi, S.; Fujiwara, T. Cadmium transport and tolerance in rice: Perspectives for reducing grain cadmium accumulation. Rice 2012, 5, 5. [Google Scholar] [CrossRef] [PubMed]
- Kulik, A.; Anielska-Mazur, A.; Bucholc, M.; Koen, E.; Szymanska, K.; Zmienko, A.; Krzywinska, E.; Wawer, I.; McLoughlin, F.; Ruszkowski, D.; et al. SNF1-related protein kinases type 2 are involved in plant responses to cadmium stress. Plant Physiol. 2012, 160, 868–883. [Google Scholar] [CrossRef] [PubMed]
- He, F.; Liu, Q.; Zheng, L.; Cui, Y.; Shen, Z. RNA-seq analysis of rice roots reveals the involvement of post-transcriptional regulation in response to cadmium stress. Front. Plant Sci. 2015, 6, 1136. [Google Scholar] [CrossRef] [PubMed]
- Jian, H.; Yang, B.; Zhang, A.; Ma, J.; Ding, Y.; Chen, Z.; Li, J.; Xu, X.; Liu, L. Genome-wide identification of MicroRNAs in response to cadmium stress in oilseed rape (Brassica napus L.) using high-throughput sequencing. Int. J. Mol. Sci. 2018, 19, 1431. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zhang, H.; Srivastava, A.K.; Pan, Y.; Bai, J.; Fang, J.; Shi, H.; Zhu, J.K. Knockdown of rice MicroRNA166 confers drought resistance by causing leaf rolling and altering stem xylem development. Plant Physiol. 2018, 176, 2082–2094. [Google Scholar] [CrossRef]
- Zhang, B. MicroRNA: A new target for improving plant tolerance to abiotic stress. J. Exp. Bot. 2015, 66, 1749–1761. [Google Scholar] [CrossRef]
- Zhao, P.; Ding, D.; Zhang, F.; Zhao, X.; Xue, Y.; Li, W.; Fu, Z.; Li, H.; Tang, J. Investigating the molecular genetic basis of heterosis for internode expansion in maize by microRNA transcriptomic deep sequencing. Funct. Integr. Genom. 2015, 15, 261–270. [Google Scholar] [CrossRef]
- Xu, L.; Wang, Y.; Zhai, L.; Xu, Y.; Wang, L.; Zhu, X.; Gong, Y.; Yu, R.; Limera, C.; Liu, L. Genome-wide identification and characterization of cadmium-responsive microRNAs and their target genes in radish (Raphanus sativus L.) roots. J. Exp. Bot. 2013, 64, 4271–4287. [Google Scholar] [CrossRef]
- Kong, X.; Zhang, M.; Xu, X.; Li, X.; Li, C.; Ding, Z. System analysis of microRNAs in the development and aluminium stress responses of the maize root system. Plant Biotechnol. J. 2014, 12, 1108–1121. [Google Scholar] [CrossRef]
- Tang, M.; Mao, D.; Xu, L.; Li, D.; Song, S.; Chen, C. Integrated analysis of miRNA and mRNA expression profiles in response to Cd exposure in rice seedlings. BMC Genom. 2014, 15, 835. [Google Scholar] [CrossRef]
- Ding, Y.; Ye, Y.; Jiang, Z.; Wang, Y.; Zhu, C. MicroRNA390 is involved in cadmium tolerance and accumulation in rice. Front. Plant Sci. 2016, 7, 235. [Google Scholar] [CrossRef]
- Marin-Gonzalez, E.; Suarez-Lopez, P. “And yet it moves”: Cell-to-cell and long-distance signaling by plant microRNAs. Plant Sci. 2012, 196, 18–30. [Google Scholar] [CrossRef] [PubMed]
- Himber, C.; Dunoyer, P. The tracking of intercellular small RNA movement. Methods Mol. Biol. 2015, 1217, 275–281. [Google Scholar]
- Zhang, W.; Kollwig, G.; Stecyk, E.; Apelt, F.; Dirks, R.; Kragler, F. Graft-transmissible movement of inverted-repeat-induced siRNA signals into flowers. Plant J. 2014, 80, 106–121. [Google Scholar] [CrossRef] [PubMed]
- Khaldun, A.B.; Huang, W.; Lv, H.; Liao, S.; Zeng, S.; Wang, Y. Comparative profiling of miRNAs and target gene identification in distant-grafting between tomato and lycium (Goji berry). Front. Plant Sci. 2016, 7, 1475. [Google Scholar] [CrossRef] [PubMed]
- Pant, B.D.; Buhtz, A.; Kehr, J.; Scheible, W.R. MicroRNA399 is a long-distance signal for the regulation of plant phosphate homeostasis. Plant J. 2008, 53, 731–738. [Google Scholar] [CrossRef] [PubMed]
- Buhtz, A.; Pieritz, J.; Springer, F.; Kehr, J. Phloem small RNAs, nutrient stress responses, and systemic mobility. BMC Plant Biol. 2010, 10, 64. [Google Scholar] [CrossRef] [PubMed]
- McGarry, R.C.; Kragler, F. Phloem-mobile signals affecting flowers: Applications for crop breeding. Trends Plant Sci. 2013, 18, 198–206. [Google Scholar] [CrossRef]
- Puzey, J.R.; Karger, A.; Axtell, M.; Kramer, E.M. Deep annotation of Populus trichocarpa microRNAs from diverse tissue sets. PLoS ONE 2012, 7, e33034. [Google Scholar] [CrossRef]
- Zhao, Y.; Lin, S.; Qiu, Z.; Cao, D.; Wen, J.; Deng, X.; Wang, X.; Lin, J.; Li, X. MicroRNA857 Is involved in the regulation of secondary growth of vascular tissues in Arabidopsis. Plant Physiol. 2015, 169, 2539–2552. [Google Scholar] [CrossRef] [PubMed]
- Han, X.; Yin, H.; Song, X.; Zhang, Y.; Liu, M.; Sang, J.; Jiang, J.; Li, J.; Zhuo, R. Integration of small RNAs, degradome and transcriptome sequencing in hyperaccumulator Sedum alfredii uncovers a complex regulatory network and provides insights into cadmium phytoremediation. Plant Biotechnol. J. 2016, 14, 1470–1483. [Google Scholar] [CrossRef] [PubMed]
- Lian, C.; Yao, K.; Duan, H.; Li, Q.; Liu, C.; Yin, W.; Xia, X. Exploration of ABA responsive miRNAs reveals a new hormone signaling crosstalk pathway regulating root growth of Populus euphratica. Int. J. Mol. Sci. 2018, 19, 1481. [Google Scholar] [CrossRef]
- Li, W.; Jia, Y.; Liu, F.; Wang, F.; Fan, F.; Wang, J.; Zhu, J.; Xu, Y.; Zhong, W.; Yang, J. Integration analysis of small RNA and degradome sequencing reveals MicroRNAs responsive to Dickeya zeae in resistant rice. Int. J. Mol. Sci. 2019, 20, 222. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.F.; Zheng, Y.; Addo-Quaye, C.; Zhang, L.; Saini, A.; Jagadeeswaran, G.; Axtell, M.J.; Zhang, W.; Sunkar, R. Transcriptome-wide identification of microRNA targets in rice. Plant J. 2010, 62, 742–759. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Q.Y.; Yang, C.Y.; Ma, Q.B.; Li, X.P.; Dong, W.W.; Nian, H. Identification of wild soybean miRNAs and their target genes responsive to aluminum stress. BMC Plant Biol. 2012, 12, 182. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Qin, C.; Chen, Z.; Zuo, T.; Yang, X.; Zhou, H.; Xu, M.; Cao, S.; Shen, Y.; Lin, H.; et al. Identification of miRNAs and their target genes in developing maize ears by combined small RNA and degradome sequencing. BMC Genom. 2014, 15, 25. [Google Scholar] [CrossRef]
- Fei, Y.; Wang, R.; Li, H.; Liu, S.; Zhang, H.; Huang, J. DPMIND: Degradome-based plant miRNA-target interaction and network database. Bioinformatics 2018, 34, 1618–1620. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Liu, X.; Xu, B.; Zhao, N.; Yang, X.; Zhang, M. Identification of miRNAs and their targets using high-throughput sequencing and degradome analysis in cytoplasmic male-sterile and its maintainer fertile lines of Brassica juncea. BMC Genom. 2013, 14, 9. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.H.; Pan, X.P.; Cox, S.B.; Cobb, G.P.; Anderson, T.A. Evidence that miRNAs are different from other RNAs. Cell. Mol. Life Sci. 2006, 63, 246–254. [Google Scholar] [CrossRef] [PubMed]
- Guzman, F.; Almerao, M.P.; Korbes, A.P.; Christoff, A.P.; Zanella, C.M.; Bered, F.; Margis, R. Identification of potential miRNAs and their targets in Vriesea carinata (Poales, Bromeliaceae). Plant Sci. 2013, 210, 214–223. [Google Scholar] [CrossRef] [PubMed]
- Tian, T.; Liu, Y.; Yan, H.; You, Q.; Yi, X.; Du, Z.; Xu, W.; Su, Z. agriGO v2.0: A GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res. 2017, 45, W122–W129. [Google Scholar] [CrossRef]
- Usadel, B.; Poree, F.; Nagel, A.; Lohse, M.; Czedik-Eysenberg, A.; Stitt, M. A guide to using MapMan to visualize and compare Omics data in plants: A case study in the crop species, Maize. Plant Cell Environ. 2009, 32, 1211–1229. [Google Scholar] [CrossRef]
- Huang, D.; Koh, C.; Feurtado, J.A.; Tsang, E.W.; Cutler, A.J. MicroRNAs and their putative targets in Brassica napus seed maturation. BMC Genom. 2013, 14, 140. [Google Scholar] [CrossRef]
- Bhogale, S.; Mahajan, A.S.; Natarajan, B.; Rajabhoj, M.; Thulasiram, H.V.; Banerjee, A.K. MicroRNA156: A potential graft-transmissible microRNA that modulates plant architecture and tuberization in Solanum tuberosum ssp. andigena. Plant Physiol. 2014, 164, 1011–1027. [Google Scholar] [CrossRef]
- Yoo, B.C.; Kragler, F.; Varkonyi-Gasic, E.; Haywood, V.; Archer-Evans, S.; Lee, Y.M.; Lough, T.J.; Lucas, W.J. A systemic small RNA signaling system in plants. Plant Cell 2004, 16, 1979–2000. [Google Scholar] [CrossRef]
- Buhtz, A.; Springer, F.; Chappell, L.; Baulcombe, D.C.; Kehr, J. Identification and characterization of small RNAs from the phloem of Brassica napus. Plant J. 2008, 53, 739–749. [Google Scholar] [CrossRef]
- Varkonyi-Gasic, E.; Gould, N.; Sandanayaka, M.; Sutherland, P.; MacDiarmid, R.M. Characterisation of microRNAs from apple (Malus domestica ‘Royal Gala’) vascular tissue and phloem sap. BMC Plant Biol. 2010, 10, 159. [Google Scholar] [CrossRef]
- Martin, A.; Adam, H.; Diaz-Mendoza, M.; Zurczak, M.; Gonzalez-Schain, N.D.; Suarez-Lopez, P. Graft-transmissible induction of potato tuberization by the microRNA miR172. Development 2009, 136, 2873–2881. [Google Scholar] [CrossRef]
- Rodriguez-Celma, J.; Ceballos-Laita, L.; Grusak, M.A.; Abadia, J.; Lopez-Millan, A.F. Plant fluid proteomics: Delving into the xylem sap, phloem sap and apoplastic fluid proteomes. Biochim. Biophys. Acta 2016, 1864, 991–1002. [Google Scholar] [CrossRef] [PubMed]
- Fontanili, L.; Lancilli, C.; Suzui, N.; Dendena, B.; Yin, Y.G.; Ferri, A.; Ishii, S.; Kawachi, N.; Lucchini, G.; Fujimaki, S.; et al. Kinetic analysis of zinc/cadmium reciprocal competitions suggests a possible zn-insensitive pathway for root-to-shoot cadmium translocation in rice. Rice 2016, 9, 16. [Google Scholar] [CrossRef]
- Li, L.; He, Z.; Pandey, G.K.; Tsuchiya, T.; Luan, S. Functional cloning and characterization of a plant efflux carrier for multidrug and heavy metal detoxification. J. Biol. Chem. 2002, 277, 5360–5368. [Google Scholar] [CrossRef]
- Thieme, C.J.; Rojas-Triana, M.; Stecyk, E.; Schudoma, C.; Zhang, W.; Yang, L.; Minambres, M.; Walther, D.; Schulze, W.X.; Paz-Ares, J.; et al. Endogenous Arabidopsis messenger RNAs transported to distant tissues. Nat. Plants 2015, 1, 15025. [Google Scholar] [CrossRef]
- Ong, S.S.; Wickneswari, R. Characterization of microRNAs expressed during secondary wall biosynthesis in Acacia mangium. PLoS ONE 2012, 7, e49662. [Google Scholar] [CrossRef]
- Quan, M.; Wang, Q.; Phangthavong, S.; Yang, X.; Song, Y.; Du, Q.; Zhang, D. Association studies in Populus tomentosa reveal the genetic interactions of Pto-MIR156c and its targets in wood formation. Front. Plant Sci. 2016, 7, 1159. [Google Scholar] [CrossRef]
- Ding, Y.; Gong, S.; Wang, Y.; Wang, F.; Bao, H.; Sun, J.; Cai, C.; Yi, K.; Chen, Z.; Zhu, C. MicroRNA166 modulates cadmium tolerance and accumulation in rice. Plant Physiol. 2018, 177, 1691–1703. [Google Scholar] [CrossRef]
- Rogers, K.; Chen, X. Biogenesis, turnover, and mode of action of plant microRNAs. Plant Cell 2013, 25, 2383–2399. [Google Scholar] [CrossRef]
- Shao, J.F.; Xia, J.; Yamaji, N.; Shen, R.F.; Ma, J.F. Effective reduction of cadmium accumulation in rice grain by expressing OsHMA3 under the control of the OsHMA2 promoter. J. Exp. Bot. 2018, 69, 2743–2752. [Google Scholar] [CrossRef]
- Wu, T.Y.; Gruissem, W.; Bhullar, N.K. Facilitated citrate-dependent iron translocation increases rice endosperm iron and zinc concentrations. Plant Sci. 2018, 270, 13–22. [Google Scholar] [CrossRef] [PubMed]
- Cornu, J.Y.; Deinlein, U.; Horeth, S.; Braun, M.; Schmidt, H.; Weber, M.; Persson, D.P.; Husted, S.; Schjoerring, J.K.; Clemens, S. Contrasting effects of nicotianamine synthase knockdown on zinc and nickel tolerance and accumulation in the zinc/cadmium hyperaccumulator Arabidopsis halleri. New Phytol. 2015, 206, 738–750. [Google Scholar] [CrossRef]
- Bailey, K.J.; Leegood, R.C. Nitrogen recycling from the xylem in rice leaves: Dependence upon metabolism and associated changes in xylem hydraulics. J. Exp. Bot. 2016, 67, 2901–2911. [Google Scholar] [CrossRef]
- Nguyen, C.; Soulier, A.J.; Masson, P.; Bussiere, S.; Cornu, J.Y. Accumulation of Cd, Cu and Zn in shoots of maize (Zea mays L.) exposed to 0.8 or 20 nM Cd during vegetative growth and the relation with xylem sap composition. Environ. Sci. Pollut. Res. Int. 2016, 23, 3152–3164. [Google Scholar] [CrossRef] [PubMed]
- Axtell, M.J.; Meyers, B.C. Revisiting criteria for plant MicroRNA annotation in the era of big data. Plant Cell 2018, 30, 272–284. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Zhao, C.; Li, M.; Sun, W.; Liu, Y.; Xia, H.; Sun, M.; Li, A.; Li, C.; Zhao, S.; et al. Genome-wide identification of Thellungiella salsuginea microRNAs with putative roles in the salt stress response. BMC Plant Biol. 2013, 13, 180. [Google Scholar] [CrossRef] [PubMed]
- Meyers, B.C.; Axtell, M.J.; Bartel, B.; Bartel, D.P.; Baulcombe, D.; Bowman, J.L.; Cao, X.; Carrington, J.C.; Chen, X.; Green, P.J.; et al. Criteria for annotation of plant MicroRNAs. Plant Cell 2008, 20, 3186–3190. [Google Scholar] [CrossRef]
- Wang, Z.; Huang, J.; Sun, Z.; Zheng, B. Identification of microRNAs differentially expressed involved in male flower development. Funct. Integr. Genom. 2015, 15, 225–232. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Huang, R.; Sun, Z.; Zhang, T.; Huang, J. Identification and profiling of conserved and novel microRNAs involved in oil and oleic acid production during embryogenesis in Carya cathayensis Sarg. Funct. Integr. Genom. 2017, 17, 365–373. [Google Scholar] [CrossRef] [PubMed]
- Yin, F.; Gao, J.; Liu, M.; Qin, C.; Zhang, W.; Yang, A.; Xia, M.; Zhang, Z.; Shen, Y.; Lin, H.; et al. Genome-wide analysis of water-stress-responsive microRNA expression profile in tobacco roots. Funct. Integr. Genom. 2014, 14, 319–332. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Xu, L.; Wang, Y.; Shen, H.; Zhu, X.; Zhang, K.; Chen, Y.; Yu, R.; Limera, C.; Liu, L. Transcriptome-wide analysis of chromium-stress responsive microRNAs to explore miRNA-mediated regulatory networks in radish (Raphanus sativus L.). Sci. Rep. 2015, 5, 14024. [Google Scholar] [CrossRef]
- Jin, Q.; Xue, Z.; Dong, C.; Wang, Y.; Chu, L.; Xu, Y. Identification and characterization of microRNAs from tree peony (Paeonia ostii) and their response to copper stress. PLoS ONE 2015, 10, e0117584. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Shahid, M.Q.; Wu, J.; Wang, L.; Liu, X.; Lu, Y. Comparative small RNA analysis of pollen development in autotetraploid and diploid rice. Int. J. Mol. Sci. 2016, 17, 499. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.R.; Pathak, H.; Sharma, S.K.; Kala, Y.K.; Nirjal, M.K.; Singh, G.P.; Goswami, S.; Rai, R.D. Novel and conserved heat-responsive microRNAs in wheat (Triticum aestivum L.). Funct. Integr. Genom. 2015, 15, 323–348. [Google Scholar] [CrossRef] [PubMed]
- Dai, X.; Zhuang, Z.; Zhao, P.X. psRNATarget: A plant small RNA target analysis server (2017 release). Nucleic Acids Res. 2018, 46, W49–W54. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.J.; Ma, Y.K.; Chen, T.; Wang, M.; Wang, X.J. PsRobot: A web-based plant small RNA meta-analysis toolbox. Nucleic Acids Res. 2012, 40, W22–W28. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Zhang, X.; Liu, X.; Zhao, Y.; Wang, B.; Zhang, J. miRNA alterations are important mechanism in maize adaptations to low-phosphate environments. Plant Sci. 2016, 252, 103–117. [Google Scholar] [CrossRef] [PubMed]
| miRNA | c0 | C1 | Cd1 | Sequence (5′-3′) | Size | MFEI |
|---|---|---|---|---|---|---|
| cme-MIR156j-p3 | 27 | 22 | 52 | GCTCACTTCTCTTTCTGTCAGT | 22 | 0.90 |
| sof-MIR156-p3 | 324 | 470 | 780 | GCTCACTTCTCTCTCTGTCAGC | 22 | 1.00 |
| zma-MIR166n-p3 | 1179 | 856 | 835 | GCTGTCGTCGACCGGAGATC | 20 | 1.00 |
| zma-MIR169g-p3 | 8 | 6 | 10 | GGCGGTCTCCTTGGCTAGCC | 20 | 1.00 |
| zma-MIR171f-p3 | 30 | 49 | 41 | TGATTGAGCCGTGCCAATATC | 21 | 0.90 |
| sbi-MIR171h-p3 | 18 | 19 | 10 | TTGAGCCGCGTCAATATCTC | 20 | 1.10 |
| zma-MIR397b-p5 | 105 | 185 | 190 | TTGAGCGCAGCGTTGATGAGC | 21 | 0.90 |
| PC-5p-37430_20 | 14 | 16 | 16 | TCTCTTAAGGCTTGTTCGGA | 20 | 0.90 |
| PC-5p-27068_30 | 23 | 12 | 17 | ACCGGAGGAGGTTAGAGGAGC | 21 | 1.30 |
| PC-5p-14301_71 | 41 | 76 | 65 | GGTTTTAGCTTCAAGCCATCT | 21 | 0.90 |
| PC-5p-10912_100 | 30 | 52 | 52 | CCGGAAATACCCAATATCTTG | 21 | 1.00 |
| PC-3p-7571_159 | 50 | 103 | 60 | GGTGGCTTGTGGCTAAAACCA | 21 | 0.90 |
| PC-3p-65413_10 | 4 | 10 | 1 | GCTTTAAGGGATCTGTTGGAGA | 22 | 1.00 |
| PC-3p-52974_13 | 12 | 8 | 13 | AATGGTGCATTGACTTGGTC | 20 | 1.10 |
| PC-3p-49169_14 | 6 | 8 | 16 | TTTGTCAATTTAAGAACTAAAA | 22 | 1.80 |
| PC-3p-37537_20 | 88 | 62 | 77 | AATACTGAGCCGAATTGAAAT | 21 | 1.10 |
| PC-3p-33282_23 | 11 | 1 | 17 | GCATCCATTCTTGGCTAAGTG | 21 | 1.20 |
| PC-3p-18761_49 | 21 | 46 | 43 | GCCTGTATGCACTCTCGGTG | 20 | 0.90 |
| PC-3p-18578_50 | 17 | 22 | 20 | TTTATGATATGTTACTCTACT | 21 | 1.50 |
| PC-3p-10246_108 | 65 | 7 | 44 | CAGGCCTTCTTGGCTAAGCG | 20 | 0.90 |
| PC-3p-100706_6 | 8 | 8 | 15 | GGAGCTGCAAACACTCTGGT | 20 | 1.50 |
| osa-MIR1430-p5 | 58 | 34 | 63 | CTTAGCCAAGAATGGCTTGCCT | 22 | 1.00 |
| miRNA | C1 | Cd1 | log2FC | chr | Strand | Start | End | hairpinLen |
|---|---|---|---|---|---|---|---|---|
| Cd Upregulated | ||||||||
| PC-3p-10246_108 | 7 | 44 | 2.73 | chr3 | + | 229987606 | 229987776 | 167 |
| PC-3p-33282_23 | 1 | 17 | 4.20 | chr3 | − | 96704531 | 96704709 | 176 |
| zma-miR169l-5p | 31 | 107 | 1.77 | chr1 | + | 298277019 | 298277107 | 87 |
| zma-miR398a-3p | 31 | 73 | 1.23 | chr7 | + | 38540171 | 38540278 | 106 |
| zma-miR398a-3p | 31 | 73 | 1.23 | chr2 | + | 169527758 | 169527897 | 138 |
| Cd downregulated | ||||||||
| zma-miR164d-3p_R+1 | 6 | 1 | −2.47 | chr7 | − | 172723300 | 172723515 | 214 |
| PC-3p-74571_8 | 9 | 4 | −1.21 | chr2 | + | 22503757 | 22503904 | 142 |
| PC-5p-167816_4 | 5 | 1 | −2.21 | chr8 | − | 103189069 | 103189246 | 135 |
| PC-5p-395659_2 | 5 | 1 | −2.21 | chr1 | + | 296265246 | 296265464 | 216 |
| PC-5p-76360_8 | 5 | 1 | −2.21 | chr1 | − | 52464270 | 52464417 | 103 |
| PC-3p-65413_10 | 10 | 1 | −3.34 | chr6 | + | 161474760 | 161474953 | 129 |
| Gene IDs | Function | miRNA(s) $ | ||
|---|---|---|---|---|
| Abiotic Stress | ||||
| ZM2g053531 | Wound-responsive protein | miR164s | ||
| ZM2G129218 | DNAJ heat family protein | miR1432s | ||
| ZM2G134917 | DNAJ homolog1 chaperone | miR395s | ||
| ZM2G456000 | ERD, early-responsive to dehydration | miR396s | ||
| ZM2g042295 | SAM-dependent methyltransferase | PC-5p-87289_7 | ||
| Phytohormone | ||||
| ZM2G159399 | ZM2G081406 | AC207656.3_FG002 | auxin response factor | miR160s |
| ZM2G078274 | auxin response factor | miR6253s | ||
| ZM2g137451 | ZM5g848945 | ZM2G135978 | auxin signaling F-box | miR393s |
| ZM2g095786 | F-box protein FBX14 | MIR159s | ||
| ZM2G019799 | aldehyde oxidase3 | PC-3p-172602_4 | ||
| Secondary Metabolism | ||||
| ZM2G336337 | ZM2G388587 | ZM2G164467 | laccase | MIR397s |
| ZM2G094375 | ZM2G305526 | ZM2G400390 | ||
| ZM2G072780 | ZM2G072808 | ZM2G447271 | ||
| ZM2G145029 | isopentenyl pyrophosphate isomerase2 | PC-3p-158678_4 | ||
| Metal Handling, Chelation, and Storage | ||||
| ZM2G085301 | Major facilitator superfamily protein | miR166s | ||
| ZM2G166976 | Major facilitator superfamily protein | miR827s | ||
| ZM2G148937 | MATE efflux family protein | miR528s | ||
| ZM2G058032 | Heavy-metal-associated domain | miR399s | ||
| ZM2G407032 | ABC transporter I family member 1 | |||
| ZM2G000039 | SIT4 phosphatase-associated protein | miR171s | ||
| Transcription Factors | ||||
| ZM2G033356 | bHLH-transcription factor, bHLH130 | PC-3p-151371_4 | ||
| AC193786.3_FG005 | bHLH-transcription factor, Bhlh154 | MIR159s | ||
| ZM2G027960 | Zinc finger protein WIP6 | |||
| ZM2G399098 | AP2-EREBP-transcription factor 124 | |||
| ZM2G048450 | ZM2G111711 | WRKY-transcription factor | ||
| ZM2G093789 | ZM2G416652 | ZM2G167088 | MYB transcription factor | |
| ZM2G423833 | ZM2G075064 | ZM2G376684 | ||
| ZM2G046443 | ZM2G070523 | ZM2G139688 | ||
| ZM2G004090 | ZM2G028054 | MYB transcription factor | miR159s, miR319s | |
| ZM2G127490 | ZM2G171781 | MYB transcription factor | PC-5p-76360_8 | |
| ZM2G305856 | ZM2G096358 | MYB transcription factor | miR164s | |
| ZM2G139700 | ZM2G393433 | ZM2G114850 | NAC-transcription factor | |
| ZM2G063522 | ZM2G146380 | |||
| ZM2G003509 | ZM2G042250 | ZM2G178102 | Homeobox-transcription factor | miR166s |
| ZM2G469551 | ZM2G109987 | AC187157.4_FG005 | Homeobox-transcription factor | |
| Signaling | ||||
| ZM2g012584 | IQ-domain | miR164s | ||
| ZM2G104730 | Calcium-transporting ATPase 9 | miR169s | ||
| ZM2G107575 | calcineurin B-like1 | PC-3p-89447_7 | ||
| ZM2G312661 | Calcium-binding protein CML42 | PC-3p-327923_2, PC-3p-100706_6 | ||
| ZM2G174315 | CaM-binding heat-shock protein | PC-3p-513669_2 | ||
| ZM2G100454 | Protein kinase | PC-5p-442461_2 | ||
| ZM2G391794 | ZM2G061447 | ZM2G146346 | LRR receptor-like kinase | MIR159s |
| ZM2G304745 | LRR receptor-like kinase | miR390s | ||
| ZM2G145756 | ZM2G082522 | Protein kinase | miR167s | |
| miRNA | Target | psRNAtarget | DPMIND | Annotation (maizeGDB) | ||
|---|---|---|---|---|---|---|
| Exp * | UPE | miR | Deg $ | |||
| zma-miR394a-5p | ZM2G064954_T01 | 0 | 23.11 | zma-miR394a-5p | 2 | LOC103636344 F-box only protein |
| ZM2G119650_T01 | 0 | 22.97 | 1 | LOC100193727, F-box domain | ||
| zma-miR393b-5p_R-1 | ZM2G135978_T01 | 1 | 18.85 | zma-miR393b-5p | 1 | transport inhibitor response 1-like |
| zma-miR390a-5p | ZM2G155490_T01 | 2 | 9.90 | zma-miR390a-5p | 1 | GRMZM2G155490 |
| ZM2G304745_T01 | 1 | 21.54 | 1 | LOC103648480 LRR receptor-like kinase | ||
| zma-miR827-5p_L+1 | ZM2G044788_T01 | 2.5 | 20.66 | zma-miR827-5p | 1 | LOC100274914 |
| zma-miR160f-5p_1ss21GA | AC207656.3_FGT002 | 0 | 24.06 | zma-miR160f-5p | 1 | arftf19, ARF-transcription factor |
| ZM2G081406_T01 | 1 | 22.15 | 1 | arftf15 | ||
| ZM2G159399_T01 | 0 | 21.94 | 2 | arftf17 | ||
| zma-miR159a-3p_R-1 | ZM2G028054_T03 | 1.5 | 16.12 | zma-miR159a-3p | 1 | myb74 transcription factor |
| zma-miR319a-3p_R+1 | 1 | 16.03 | zma-miR319a-3p | 2 | ||
| zma-miR159a-3p_R-1 | ZM2G139688_T01 | 2 | 17.18 | zma-miR159a-3p | 3 | myb138 |
| gma-miR171m_1ss21AC | ZM2G098800_T01 | 0.5 | 23.32 | zma-miR171m-3p | 1 | gras80 transcription factor |
| sbi-MIR171h-p3 | 0.5 | 23.32 | 1 | |||
| osa-MIR171a-p3 | 0 | 23.32 | zma-miR171n-3p | 1 | ||
| zma-MIR171f-p3 | 0.5 | 23.52 | zma-miR171f-3p | 3 | ||
| zma-miR166a-3p | ZM2G109987_T04 | 1 | 24.23 | zma-miR166a-3p | 2 | rld1, rolled leaf, Homeobox |
| osa-miR166a-3p_1ss21CT | ZM2G042250_T04 | 23.53 | zma-miR166c-3p | rld2 | ||
| zma-miR166l-3p | ZM2G469551_T02 | 19.01 | zma-miR166l-3p | hb69, Homeobox-transcription factor | ||
| lus-miR169a_R-1 | ZM2G000686_T06 | 2 | 18.96 | zma-miR169h | 6 | nfya1 nuclear transcription factor Y |
| zma-miR169a-5p_R-1 | ZM2G040349_T01 | 2 | 18.79 | zma-miR169a-5p | 2 | ca2p4, NFY/CCAAT-HAP2-transcription factor |
| zma-miR169f-5p_R-1_1ss1TG | ZM2G091964_T02 | 2.5 | 20.96 | zma-miR169f-5p | 7 | ca2p16 |
| zma-miR169i-5p_R-1 | ZM2G038303_T01 | 2 | 16.87 | zma-miR169i-5p | 6 | ca2p15 |
| zma-miR169l-5p | ZM5G857944_T03 | 2.5 | 17.47 | zma-miR169l-5p | 6 | ca2p13 |
| zma-miR169o-5p_R-1 | ZM5G853836_T01 | 2.5 | 20.94 | zma-miR169o-5p | 4 | ca2p5 |
| osa-miR169b_R+1 | ZM5G857944_T03 | 2 | 17.47 | zma-miR169c-5p | 6 | ca2p13 |
| ZM2G067624_T02 | 1 | 16.26 | 7 | sbp29, squamosa promoter binding protein | ||
| bdi-miR156b-5p_R+2 | ZM2G097275_T04 | 21.87 | zma-miR156i-5p | 1 | sbp27 | |
| osa-miR156a_R+1 | ZM2G113779_T01 | 14.23 | zma-miR156a-5p | 5 | sbp13 | |
| zma-miR156a-5p | ZM2G126018_T01 | 18.45 | zma-miR156a-5p | 1 | sbp23 | |
| zma-miR156j-5p_R-1 | ZM2G126827_T01 | 22.70 | zma-miR156j-5p | 1 | sbp12 | |
| zma-miR156k-5p | ZM2G156621_T01 | 22.70 | zma-miR156k-5p | 1 | sbp31 | |
| zma-miR529-5p | ZM2G307588_T01 | 16.69 | zma-miR529-5p | 1 | tsh4, tassel sheath4, SBP | |
| ZM2G371033_T01 | 18.52 | 3 | sbp18 | |||
| ZM5G878561_T01 | 19.13 | 3 | sbp22 | |||
| ZM2G460544_T01 | 20.77 | 1 | ub3, unbranched3, SBP | |||
| zma-miR529-5p | ZM2G160917_T01 | 0.5 | 22.12 | zma-miR529-5p | 1 | ub2, unbranched2 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, B.; Cheng, D.; Chen, Z.; Zhang, M.; Zhang, G.; Jiang, M.; Tan, M. Bioinformatic Exploration of the Targets of Xylem Sap miRNAs in Maize under Cadmium Stress. Int. J. Mol. Sci. 2019, 20, 1474. https://doi.org/10.3390/ijms20061474
Wang B, Cheng D, Chen Z, Zhang M, Zhang G, Jiang M, Tan M. Bioinformatic Exploration of the Targets of Xylem Sap miRNAs in Maize under Cadmium Stress. International Journal of Molecular Sciences. 2019; 20(6):1474. https://doi.org/10.3390/ijms20061474
Chicago/Turabian StyleWang, Baoxiang, Dan Cheng, Ziyan Chen, Manman Zhang, Guoqiang Zhang, Mingyi Jiang, and Mingpu Tan. 2019. "Bioinformatic Exploration of the Targets of Xylem Sap miRNAs in Maize under Cadmium Stress" International Journal of Molecular Sciences 20, no. 6: 1474. https://doi.org/10.3390/ijms20061474
APA StyleWang, B., Cheng, D., Chen, Z., Zhang, M., Zhang, G., Jiang, M., & Tan, M. (2019). Bioinformatic Exploration of the Targets of Xylem Sap miRNAs in Maize under Cadmium Stress. International Journal of Molecular Sciences, 20(6), 1474. https://doi.org/10.3390/ijms20061474






