Auxin Information Processing; Partners and Interactions beyond the Usual Suspects
Abstract
1. Introduction
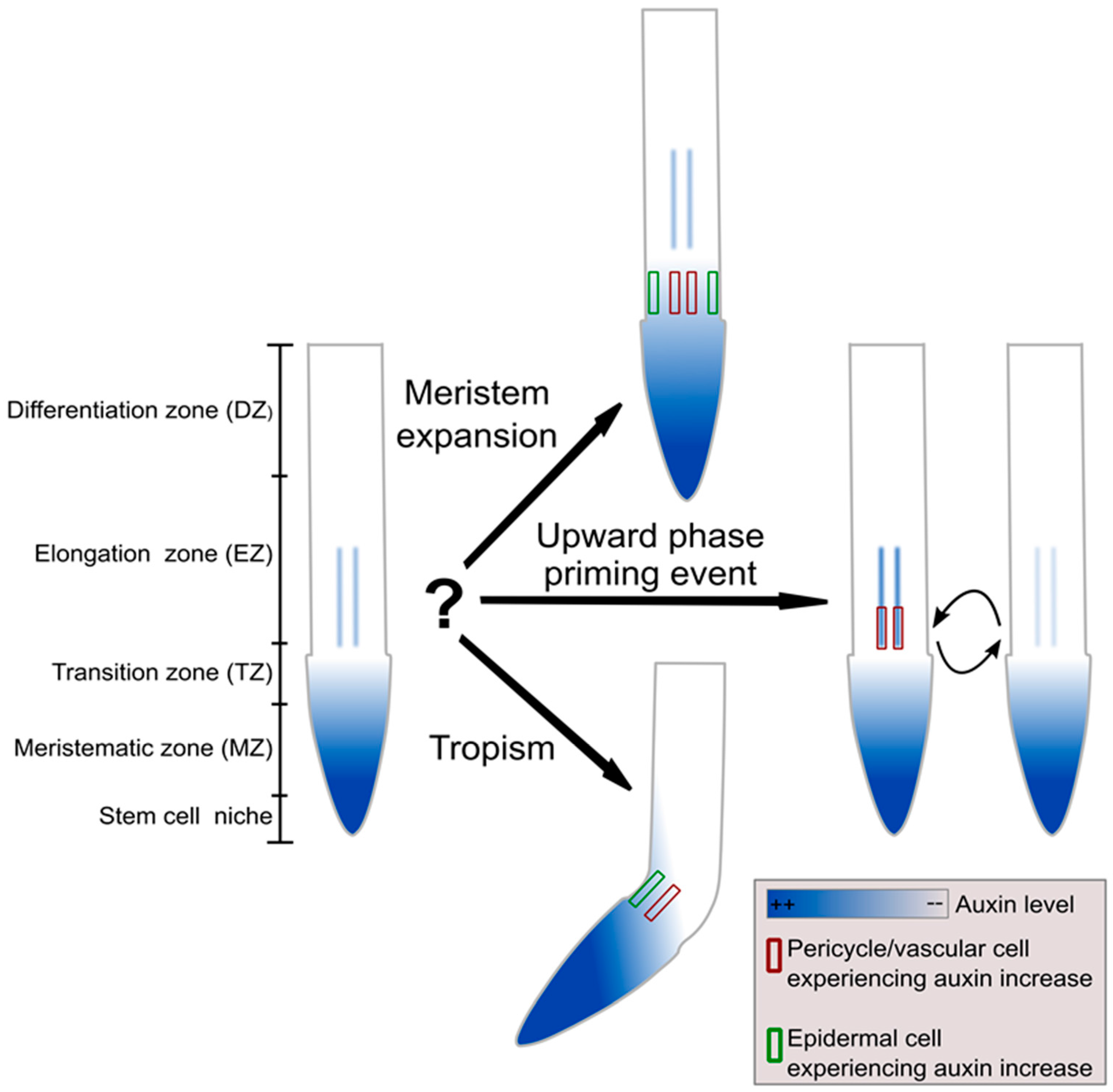
2. The Auxin–Plethora Division of Labour; A Separation of Timescales
3. Halotropism as a Case Study of a Graded-Signal Tropism; Auxin Computations in the Reflux Loop
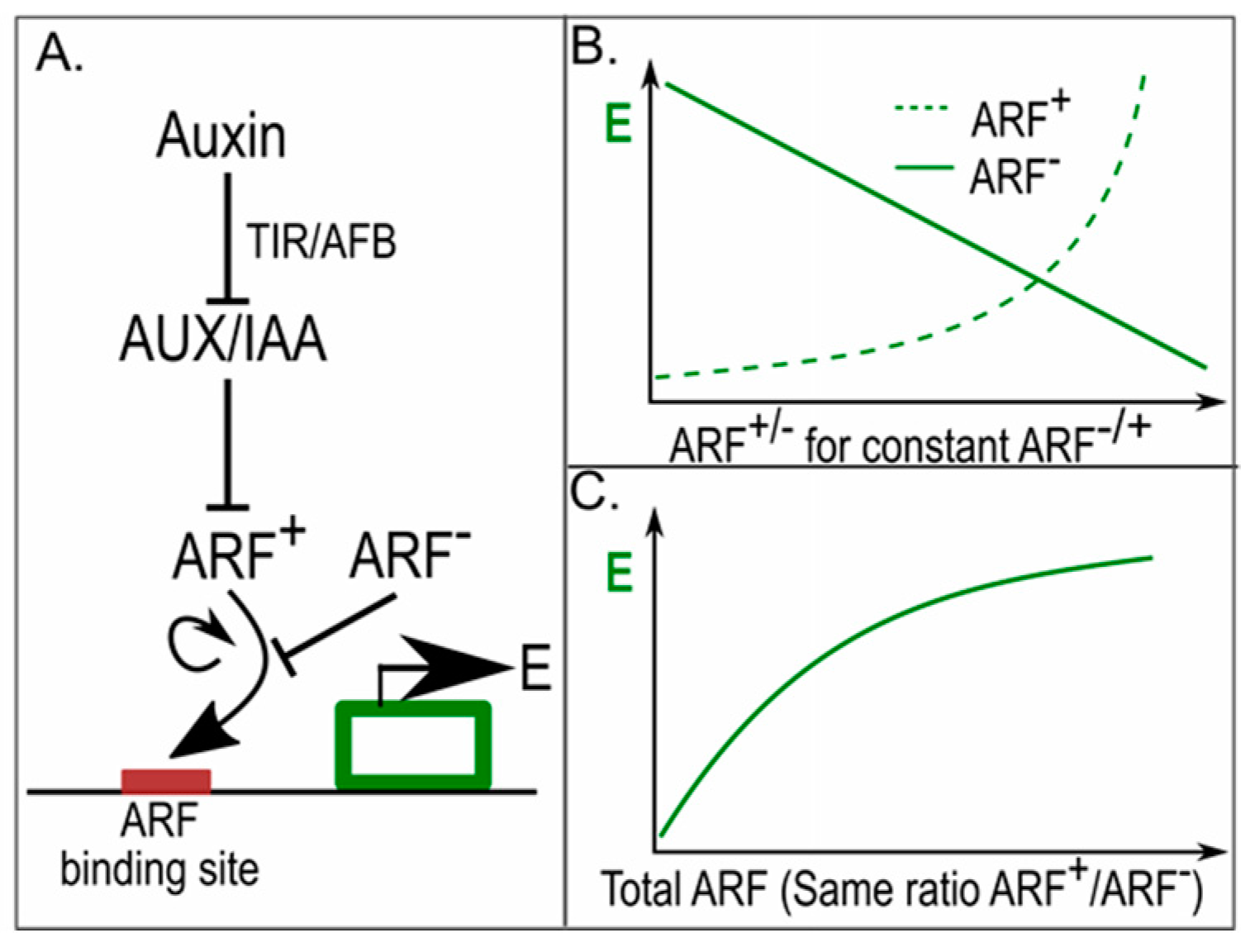
4. Auxin Signalling in Phyllotaxis; Same yet Different
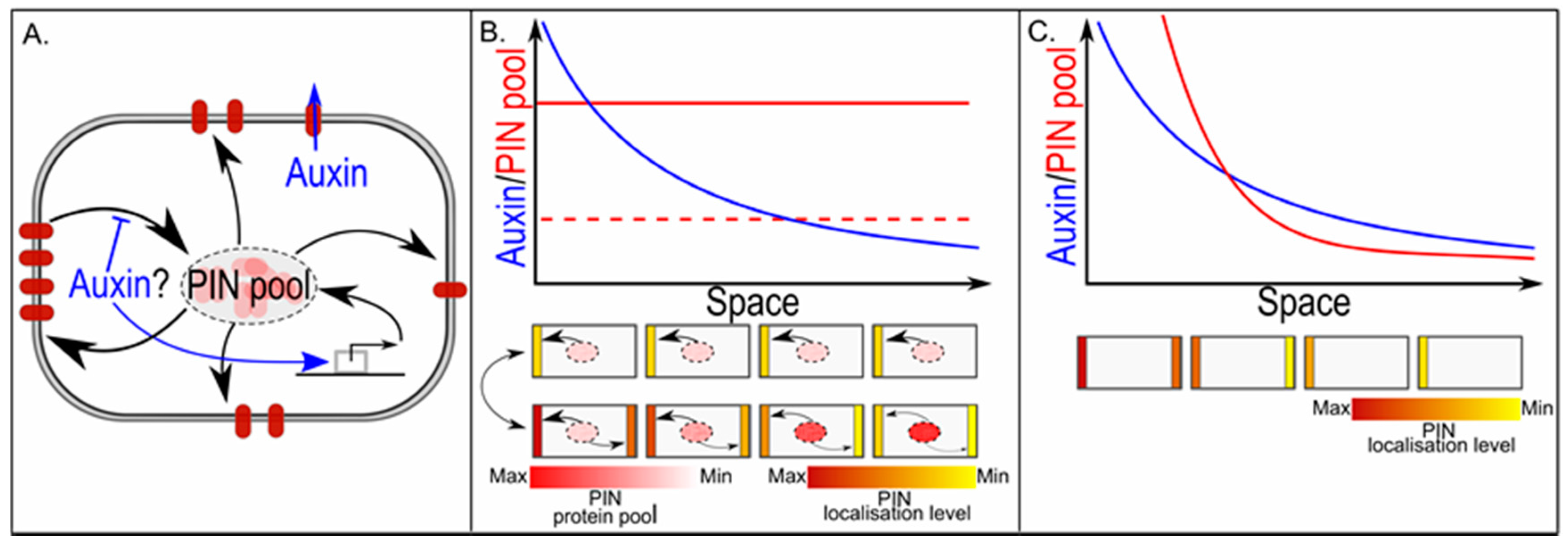
5. Pin Polarity in Gradients; Different yet Same
6. Conclusions and Outlook
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Saini, S.; Sharma, I.; Kaur, N.; Pati, P.K. Auxin: A master regulator in plant root development. Plant Cell Rep. 2013, 741–757. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.; Wang, S. Insights into auxin signaling in plant—Pathogen interactions. Front. Plant Sci. 2011, 2, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Kazan, K. Auxin and the integration of environmental signals into plant root development. Ann. Bot. 2013, 112, 1655–1665. [Google Scholar] [CrossRef] [PubMed]
- Dello Ioio, R.; Nakamura, K.; Moubayindin, L.; Perilli, S.; Taniguchi, M.; Morita, M.T.; Aoyama, T.; Costantino, P.; Sabatini, S. A genetic framework for genetic control of cell division and differentiation in the root meristem. Science 2008, 22, 1380–1384. [Google Scholar] [CrossRef] [PubMed]
- Grieneisen, V.A.; Xu, J.; Maree, A.F.M.; Hogeweg, P.; Scheres, B. Auxin transport is sufficient to generate a maximum and gradient guiding root growth. Nature 2007, 449, 1008–1013. [Google Scholar] [CrossRef] [PubMed]
- Petersson, S.V.; Johansson, A.I.; Kowalczyk, M.; Makoveychuk, A.; Wang, J.Y.; Moritz, T.; Grebe, M.; Benfey, P.N.; Sandberg, G.; Ljung, K. An auxin gradient and maximum in the Arabidopsis root apex shown by high-resolution cell-specific analysis of IAA distribution and synthesis. Plant Cell 2009, 21, 1659–1668. [Google Scholar] [CrossRef] [PubMed]
- Vernoux, T.; Kronenberger, J.; Grandjean, O.; Laufs, P.; Traas, J. PIN-FORMED 1 regulates cell fate at the periphery of the shoot apical meristem. Development 2000, 127, 5157–5165. [Google Scholar] [PubMed]
- Benková, E.; Michniewicz, M.; Sauer, M.; Teichmann, T.; Seifertová, D.; Jürgens, G.; Friml, J. Local, Efflux-Dependent Auxin Gradients as a Common Module for Plant Organ Formation. Cell 2003, 115, 591–602. [Google Scholar] [CrossRef]
- Heisler, M.G.; Ohno, C.; Das, P.; Sieber, P.; Reddy, G.V.; Long, J.A.; Meyerowitz, E.M. Patterns of auxin transport and gene expression during primordium development revealed by live imaging of the Arabidopsis inflorescence meristem. Curr. Biol. 2005, 15, 1899–1911. [Google Scholar] [CrossRef] [PubMed]
- De Smet, I.; Tetsumura, T.; de Rybel, B.; Frey, N.F.D.; Laplaze, L.; Casimiro, I.; Swarup, R.; Naudts, M.; Vanneste, S.; Audenaert, D.; et al. Auxin-dependent regulation of lateral root positioning in the basal meristem of Arabidopsis. Development 2007, 134, 681–690. [Google Scholar] [CrossRef] [PubMed]
- Sachs, T. The Control of the Patterned Differentiation of Vascular Tissues. Adv. Bot. Res. 1981, 9, 151–262. [Google Scholar]
- Sachs, T. Cell polarity and tissue patterning in plants. Development 1991, 113, 83–93. [Google Scholar]
- Friml, J.; Wiśniewska, J.; Benková, E.; Mendgen, K.; Palme, K. Lateral relocation of auxin efflux regulator PIN3 mediates tropism in Arabidopsis. Nature 2002, 415, 806–809. [Google Scholar] [CrossRef] [PubMed]
- Swarup, R.; Kramer, E.M.; Perry, P.; Knox, K.; Leyser, O.; Beemster, G.T.S.; Haseloff, J.; Bhalerao, R.; Bennett, M.J. Root gravitropism requires lateral root cap and epidermal cells for transport and response to a mobile auxin signal. Nat. Cell Biol. 2005, 7, 1057–1065. [Google Scholar] [CrossRef] [PubMed]
- Liscum, E.; Briggs, W.R. Mutations of Arabidopsis in potential transduction and response components of the phototropic signaling pathway. Plant Physiol. 1996, 112, 291–296. [Google Scholar] [CrossRef] [PubMed]
- Nacry, P.; Canivenc, G.; Muller, B.; Azmi, A.; van Onckelen, H.; Rossignol, M.; Doumas, P. A role for auxin redistribution in the responses of the root system architecture to phosphate starvation in Arabidopsis. Plant Physiol. 2005, 138, 2061–2074. [Google Scholar] [CrossRef] [PubMed]
- Krouk, G.; Mirowski, P.; LeCun, Y.; Shasha, D.E.; Coruzzi, G.M. Predictive network modeling of the high-resolution dynamic plant transcriptome in response to nitrate. Genome Biol. 2010, 11. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, S.B.; Hagen, G.; Guilfoyle, T. The roles of auxin response factor domains in auxin-responsive transcription. Plant Cell 2003, 15, 533–543. [Google Scholar] [CrossRef] [PubMed]
- Strader, L.C.; Zhao, Y. Auxin perception and downstream events. Curr. Opin. Plant Biol. 2016, 33, 8–14. [Google Scholar] [CrossRef] [PubMed]
- Mockaitis, K.; Estelle, M. Auxin Receptors and Plant Development: A New Signaling Paradigm. Annu. Rev. Cell Dev. Biol. 2008, 24, 55–80. [Google Scholar] [CrossRef] [PubMed]
- Liscum, E.; Reed, J.W. Genetics of Aux/IAA and ARF action in plant growth and development. Plant Mol. Biol. 2002, 49, 387–400. [Google Scholar] [CrossRef] [PubMed]
- Calderón Villalobos, L.I.A.; Lee, S.; De Oliveira, C.; Ivetac, A.; Brandt, W.; Armitage, L.; Sheard, L.B.; Tan, X.; Parry, G.; Mao, H.; et al. A combinatorial TIR1/AFB–Aux/IAA co-receptor system for differential sensing of auxin. Nat. Chem. Biol. 2012, 8, 477–485. [Google Scholar] [CrossRef] [PubMed]
- Shimizu-Mitao, Y.; Kakimoto, T. Auxin sensitivities of all arabidopsis Aux/IAAs for degradation in the presence of every TIR1/AFB. Plant Cell Physiol. 2014, 55, 1450–1459. [Google Scholar] [CrossRef] [PubMed]
- Weijers, D.; Benkova, E.; Jäger, K.E.; Schlereth, A.; Hamann, T.; Kientz, M.; Wilmoth, J.C.; Reed, J.W.; Jürgens, G. Developmental specificity of auxin response by pairs of ARF and Aux/IAA transcriptional regulators. EMBO J. 2005, 24, 1874–1885. [Google Scholar] [CrossRef] [PubMed]
- Rademacher, E.H.; Möller, B.; Lokerse, A.S.; Llavata-Peris, C.I.; van den Berg, W.; Weijers, D. A cellular expression map of the Arabidopsis AUXIN RESPONSE FACTOR gene family. Plant J. 2011, 68, 597–606. [Google Scholar] [CrossRef] [PubMed]
- Boer, D.R.; Freire-Rios, A.; van den Berg, W.A.M.; Saaki, T.; Manfield, I.W.; Kepinski, S.; López-Vidrieo, I.; Franco-Zorrilla, J.M.; de Vries, S.C.; Solano, R.; et al. Structural basis for DNA binding specificity by the auxin-dependent ARF transcription factors. Cell 2014, 156, 577–589. [Google Scholar] [CrossRef] [PubMed]
- De Rybel, B.; Vassileva, V.; Parizot, B.; Demeulenaere, M.; Grunewald, W.; Audenaert, D.; van Campenhout, J.; Overvoorde, P.; Jansen, L.; Vanneste, S.; et al. A novel Aux/IAA28 signaling cascade activates GATA23-dependent specification of lateral root founder cell identity. Curr. Biol. 2010, 20, 1697–1706. [Google Scholar] [CrossRef] [PubMed]
- Goh, T.; Joi, S.; Mimura, T.; Fukaki, H. The establishment of asymmetry in Arabidopsis lateral root founder cells is regulated by LBD16/ASL18 and related LBD/ASL proteins. Development 2012, 139, 883–893. [Google Scholar] [CrossRef] [PubMed]
- Lavenus, J.; Goh, T.; Roberts, I.; Guyomarc’h, S.; Lucas, M.; de Smet, I.; Fukaki, H.; Beeckman, T.; Bennett, M.; Laplaze, L. Lateral root development in Arabidopsis: Fifty shades of auxin. Trends Plant Sci. 2013, 18, 1360–1385. [Google Scholar] [CrossRef] [PubMed]
- Mähönen, A.P.; Tusscher ten, K.; Siligato, R.; Smetana, O.; Díaz-Triviño, S.; Salojärvi, J.; Wachsman, G.; Prasad, K.; Heidstra, R.; Scheres, B. PLETHORA gradient formation mechanism separates auxin responses. Nature 2014, 515, 125–129. [Google Scholar] [CrossRef] [PubMed]
- Adler, J. Chemotaxis in Bacteria. Science 1966, 153, 708–716. [Google Scholar] [CrossRef] [PubMed]
- Ishida, T.; Adachi, S.; Yoshimura, M.; Shimizu, K.; Umeda, M.; Sugimoto, K. Auxin modulates the transition from the mitotic cycle to the endocycle in Arabidopsis. Development 2010, 137, 63–71. [Google Scholar] [CrossRef] [PubMed]
- Perrot-Rechenmann, C. Cellular responses to auxin: Division versus expansion. Cold Spring Harb. Perspect. Biol. 2010, 2, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Evans, M.L.; Ishikawa, H. Responses of Arabidopsis roots to auxin studied with high temporal resolution: Comparison of wild type and auxin-response mutants. Planta 1994, 194, 215–222. [Google Scholar] [CrossRef]
- Mullen, J.L.; Ishikawa, H.; Evans, M.L. Analysis of changes in relative elemental growth rate patterns in the elongation zone of Arabidopsis roots upon gravistimulation. Planta 1998, 206, 598–603. [Google Scholar] [CrossRef] [PubMed]
- Galinha, C.; Hofhuis, H.; Luijten, M.; Willemsen, V.; Blilou, I.; Heidstra, R.; Scheres, B. PLETHORA proteins as dose-dependent master regulators of Arabidopsis root development. Nature 2007, 449, 1053–1057. [Google Scholar] [CrossRef] [PubMed]
- Aida, M.; Beis, D.; Heidstra, R.; Willemsen, V.; Blilou, I.; Galinha, C.; Nussaume, L.; Noh, Y.; Amasino, R.; Scheres, B.; et al. The PLETHORA Genes Mediate Patterning of the Arabidopsis Root Stem Cell Niche. Cell 2004, 119, 109–120. [Google Scholar] [CrossRef] [PubMed]
- Kawakami, Y.; Rasskin-gutman, D.; Izpisu, J.C.; Iban, M. Cell lineage transport: A mechanism for molecular gradient formation. Mol. Syst. Biol. 2006, 57, 1–12. [Google Scholar] [CrossRef]
- Shkolnik, D.; Krieger, G.; Nuriel, R.; Fromm, H. Hydrotropism: Root Bending Does Not Require Auxin Redistribution. Mol. Plant 2016, 9, 757–759. [Google Scholar] [CrossRef] [PubMed]
- Kutschera, U.; Briggs, W.R. Root phototropism: From dogma to the mechanism of blue light perception. Planta 2012, 235, 443–452. [Google Scholar] [CrossRef] [PubMed]
- Galvan-Ampudia, C.S.; Julkowska, M.M.; Darwish, E.; Gandullo, J.; Korver, R.A.; Brunoud, G.; Haring, M.A.; Munnik, T.; Vernoux, T.; Testerink, C. Halotropism is a response of plant roots to avoid a saline environment. Curr. Biol. 2013, 23, 2044–2050. [Google Scholar] [CrossRef] [PubMed]
- Eshel, A.; Beeckman, T. Plant Roots: The Hidden Half, 4th ed.; CRC Press: Roca Raton, FL, USA, 2013. [Google Scholar]
- Kleine-Vehn, J.; Ding, Z.; Jones, A.R.; Tasaka, M.; Morita, M.T.; Friml, J. Gravity-induced PIN transcytosis for polarization of auxin fluxes in gravity-sensing root cells. Proc. Natl. Acad. Sci. USA 2010, 107, 22344–22349. [Google Scholar] [CrossRef] [PubMed]
- Ottenschläger, I.; Wolff, P.; Wolverton, C.; Bhalerao, R.P.; Sandberg, G.; Ishikawa, H.; Evans, M.; Palme, K. Gravity-regulated differential auxin transport from columella to lateral root cap cells. Proc. Natl. Acad. Sci. USA 2003, 100, 2987–2991. [Google Scholar] [CrossRef] [PubMed]
- Monshausen, G.B.; Miller, N.D.; Murphy, A.S.; Gilroy, S. Dynamics of auxin-dependent Ca2+ and pH signaling in root growth revealed by integrating high-resolution imaging with automated computer vision-based analysis. Plant J. 2011, 65, 309–318. [Google Scholar] [CrossRef] [PubMed]
- Christie, J.M.; Murphy, A.S. Shoot phototropism in higher plants: New light through old concepts. Am. J. Bot. 2013, 100, 35–46. [Google Scholar] [CrossRef] [PubMed]
- Sun, F.; Zhang, W.; Hu, H.; Li, B.; Wang, Y.; Zhao, Y.; Li, K.; Liu, M.; Li, X. Salt Modulates Gravity Signaling Pathway to Regulate Growth Direction of Primary Roots in Arabidopsis. Plant Physiol. 2007, 146, 178–188. [Google Scholar] [CrossRef] [PubMed]
- Cassab, G.I.; Eapen, D.; Campos, M.E. Root hydrotropism: An update. Am. J. Bot. 2013, 100, 14–24. [Google Scholar] [CrossRef] [PubMed]
- Van den Berg, T.; Korver, R.A.; Testerink, C.; Ten Tusscher, K.H.W.J. Modeling halotropism: A key role for root tip architecture and reflux loop remodeling in redistributing auxin. Development 2016, 143, 3350–3362. [Google Scholar] [CrossRef] [PubMed]
- Okada, K.; Ueda, M.; Komaki, M.K.; Bell, C.J.; Shimura, Y. Requirement of the Auxin Polar Transport System in Early Stages of Arabidopsis Floral Bud Formation. Plant Cell Online 1991, 3, 677–684. [Google Scholar] [CrossRef] [PubMed]
- Vernoux, T.; Brunoud, G.; Farcot, E.; Morin, V.; van den Daele, H.; Legrand, J.; Oliva, M.; Das, P.; Larrieu, A.; Wells, D.; et al. The auxin signalling network translates dynamic input into robust patterning at the shoot apex. Mol. Syst. Biol. 2011, 7, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Ulmasov, T.; Hagen, G.; Guilfoyle, T.J. Dimerization and DNA binding of auxin response factors. Plant J. 1999, 19, 309–319. [Google Scholar] [CrossRef] [PubMed]
- Blilou, I.; Xu, J.; Wildwater, M.; Willemsen, V.; Paponov, I.; Friml, J.; Heidstra, R.; Aida, M.; Palme, K.; Scheres, B. The PIN auxin efflux facilitator network controls growth and patterning in Arabidopsis roots. Nature 2005, 433, 39–44. [Google Scholar] [CrossRef] [PubMed]
- Wisniewska, J.; Xu, J.; Seifertová, D.; Brewer, P.B.; Ruzicka, K.; Blilou, I.; Rouquie, D.; Benkova, E.; Scheres, B.; Friml, J. Polar PIN localization directs auxin flow in plants. Science 2006, 312, 883. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.; Wen, M.; Nagawa, S.; Fu, Y.; Chen, J.G.; Wu, M.J.; Perrot-Rechenmann, C.; Friml, J.; Jones, A.M.; Yang, Z. Cell surface- and Rho GTPase-based auxin signaling controls cellular interdigitation in Arabidopsis. Cell 2010, 143, 99–110. [Google Scholar] [CrossRef] [PubMed]
- Robert, S.; Kleine-Vehn, J.; Barbez, E.; Sauer, M.; Paciorek, T.; Baster, P.; Vanneste, S.; Zhang, J.; Simon, S.; Čovanová, M.; et al. ABP1 mediates auxin inhibition of clathrin-dependent endocytosis in arabidopsis. Cell 2010, 143, 111–121. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Zhang, Y.; Zhang, D.; Dai, X.; Estelle, M.; Zhao, Y. Auxin binding protein 1 (ABP1) is not required for either auxin signaling or Arabidopsis development. Proc. Natl. Acad. Sci. USA 2015, 112, 2275–2280. [Google Scholar] [CrossRef] [PubMed]
- Jásik, J.; Bokor, B.; Stuchlík, S.; Mičieta, K.; Turňa, J.; Schmelzer, E. Effects of auxins on PIN-FORMED2(PIN2) dynamics are not mediated by inhibiting PIN2 endocytosis. Plant Physiol. 2016, 172, 1019–1031. [Google Scholar] [CrossRef] [PubMed]
- Stoma, S.; Lucas, M.; Chopard, J.; Schaedel, M.; Traas, J.; Godin, C. Flux-based transport enhancement as a plausible unifying mechanism for auxin transport in meristem development. PLoS Comput. Biol. 2008, 4, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Heisler, M.G.; Jönsson, H. Modeling auxin transport and plant development. J. Plant Growth Regul. 2006, 25, 302–312. [Google Scholar] [CrossRef]
- Bayer, E.M.; Smith, R.S.; Mandel, T.; Nakayama, N.; Sauer, M.; Prusinkiewicz, P.; Kuhlemeier, C. Integration of transport-based models for phyllotaxis and midvein formation. Genes Dev. 2009, 23, 373–384. [Google Scholar] [CrossRef] [PubMed]
- Van Berkel, K.; de Boer, R.J.; Scheres, B.; ten Tusscher, K. Polar auxin transport: Models and mechanisms. Development 2013, 140, 2253–2268. [Google Scholar] [CrossRef] [PubMed]
- Vieten, A.; Vanneste, S.; Wisniewska, J.; Benková, E.; Benjamins, R.; Beeckman, T.; Luschnig, C.; Friml, J. Functional redundancy of PIN proteins is accompanied by auxin-dependent cross-regulation of PIN expression. Development 2005, 132, 4521–4531. [Google Scholar] [CrossRef] [PubMed]
- Laskowski, M.; ten Tusscher, K.H. Periodic Lateral Root Priming: What Makes It Tick? Plant Cell 2017, 29, 432–444. [Google Scholar] [CrossRef] [PubMed]
- Adler, J. Chemoreceptors in Bacteria. Science 1969, 166, 1588–1597. [Google Scholar] [CrossRef] [PubMed]
- Parkinson, J.S. Signal transduction schemes of bacteria. Cell 1993, 73, 857–871. [Google Scholar] [CrossRef]
- Navarro, L.; Dunoyer, P.; Jay, F.; Arnold, B.; Dharmasiri, N.; Estelle, M.; Voinnet, O.; Jones, J.D.G. A Plant miRNA Contributes to Antibacterial Resistance by Repressing Auxin Signaling. Science 2006, 312, 436–439. [Google Scholar] [CrossRef] [PubMed]
- Vert, G.; Walcher, C.L.; Chory, J.; Nemhauser, J.L. Integration of auxin and brassinosteroid pathways by Auxin Response Factor 2. Proc. Natl. Acad. Sci. USA 2008, 105, 9829–9834. [Google Scholar] [CrossRef] [PubMed]
- Antoniadi, I.; Plačková, L.; Simonovik, B.; Doležal, K.; Turnbull, C.; Ljung, K.; Novák, O. Cell-Type-Specific Cytokinin Distribution within the Arabidopsis Primary Root Apex. Plant Cell 2015, 27, 1955–1967. [Google Scholar] [CrossRef] [PubMed]
- Matsuzaki, Y.; Ogawa-Ohnishi, M.; Mori, A.; Matsubayashi, Y. Secreted Peptide Signals Required for Maintenance of Root Stem Cell Niche in Arabidopsis. Science 2010, 329, 1065–1067. [Google Scholar] [CrossRef] [PubMed]
- Yoon, E.K.; Yang, J.H.; Lim, J.; Kim, S.H.; Kim, S.; Lee, W.S. Auxin regulation of the microRNA390-dependent transacting small interfering RNA pathway in Arabidopsis lateral root development. Nucleic Acids Res. 2010, 38, 1382–1391. [Google Scholar] [CrossRef] [PubMed]
- Chandler, J.W.; Werr, W. Cytokinin-auxin crosstalk in cell type specification. Trends Plant Sci. 2015, 20, 291–300. [Google Scholar] [CrossRef] [PubMed]
- Cruz-Ramírez, A.; Díaz-Triviño, S.; Blilou, I.; Grieneisen, V.A.; Sozzani, R.; Zamioudis, C.; Miskolczi, P.; Nieuwland, J.; Benjamins, R.; Dhonukshe, P.; et al. A bistable circuit involving SCARECROW-RETINOBLASTOMA integrates cues to inform asymmetric stem cell division. Cell 2012, 150, 1002–1015. [Google Scholar] [CrossRef] [PubMed]
- Whitford, R.; Fernandez, A.; Tejos, R.; Pérez, A.C.; Kleine-Vehn, J.; Vanneste, S.; Drozdzecki, A.; Leitner, J.; Abas, L.; Aerts, M.; et al. GOLVEN Secretory Peptides Regulate Auxin Carrier Turnover during Plant Gravitropic Responses. Dev. Cell 2012, 22, 678–685. [Google Scholar] [CrossRef] [PubMed]
- Moubayidin, L.; Mambro, R.D.; Sozzani, R.; Pacifici, E.; Salvi, E.; Terpstra, I.; Bao, D.; Van Dijken, A.; Dello Ioio, R.; Perilli, S.; et al. Article Spatial Coordination between Stem Cell Activity and Cell Differentiation in the Root Meristem. Dev. Cell 2013, 26, 405–415. [Google Scholar] [CrossRef] [PubMed]


© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Van den Berg, T.; Ten Tusscher, K.H. Auxin Information Processing; Partners and Interactions beyond the Usual Suspects. Int. J. Mol. Sci. 2017, 18, 2585. https://doi.org/10.3390/ijms18122585
Van den Berg T, Ten Tusscher KH. Auxin Information Processing; Partners and Interactions beyond the Usual Suspects. International Journal of Molecular Sciences. 2017; 18(12):2585. https://doi.org/10.3390/ijms18122585
Chicago/Turabian StyleVan den Berg, Thea, and Kirsten H. Ten Tusscher. 2017. "Auxin Information Processing; Partners and Interactions beyond the Usual Suspects" International Journal of Molecular Sciences 18, no. 12: 2585. https://doi.org/10.3390/ijms18122585
APA StyleVan den Berg, T., & Ten Tusscher, K. H. (2017). Auxin Information Processing; Partners and Interactions beyond the Usual Suspects. International Journal of Molecular Sciences, 18(12), 2585. https://doi.org/10.3390/ijms18122585



