The Effect of 2′F-RNA on I-Motif Structure and Stability
Abstract
1. Introduction
2. Results
2.1. I-Motif Formation and Stability in 2′F-Riboc Containing Sequences
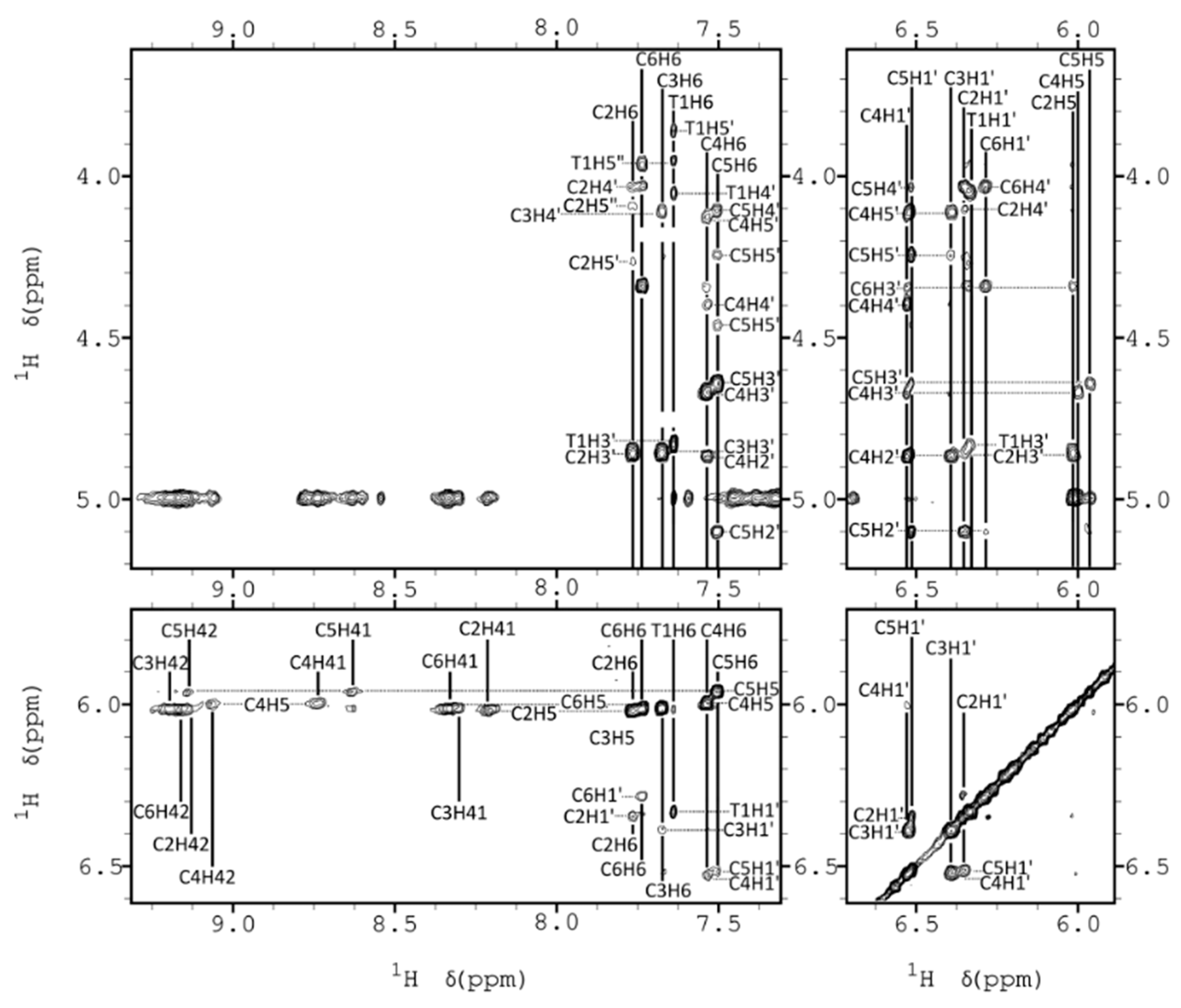
2.2. NMR Assignments
2.3. NMR Constraints and Structural Calculation
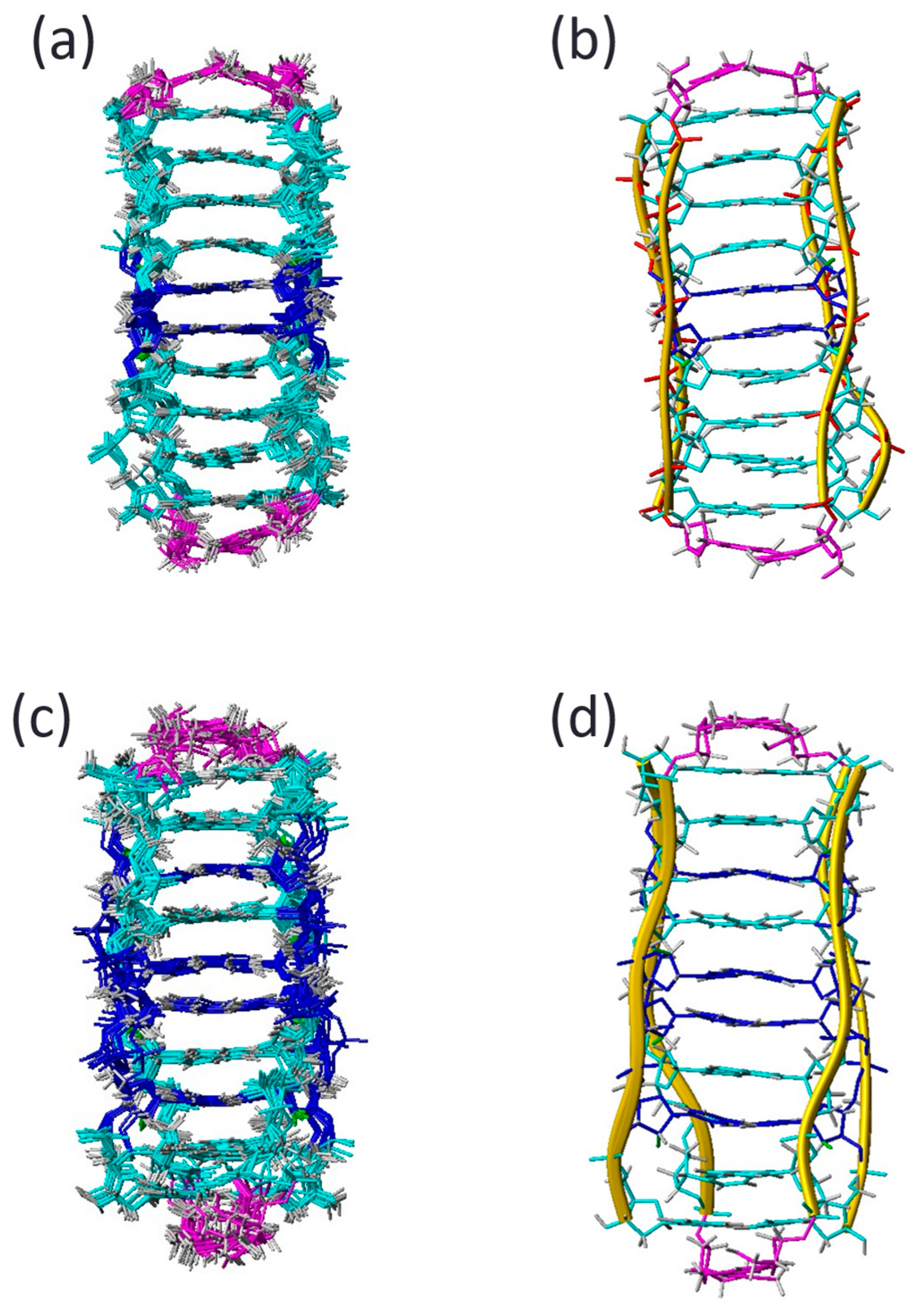
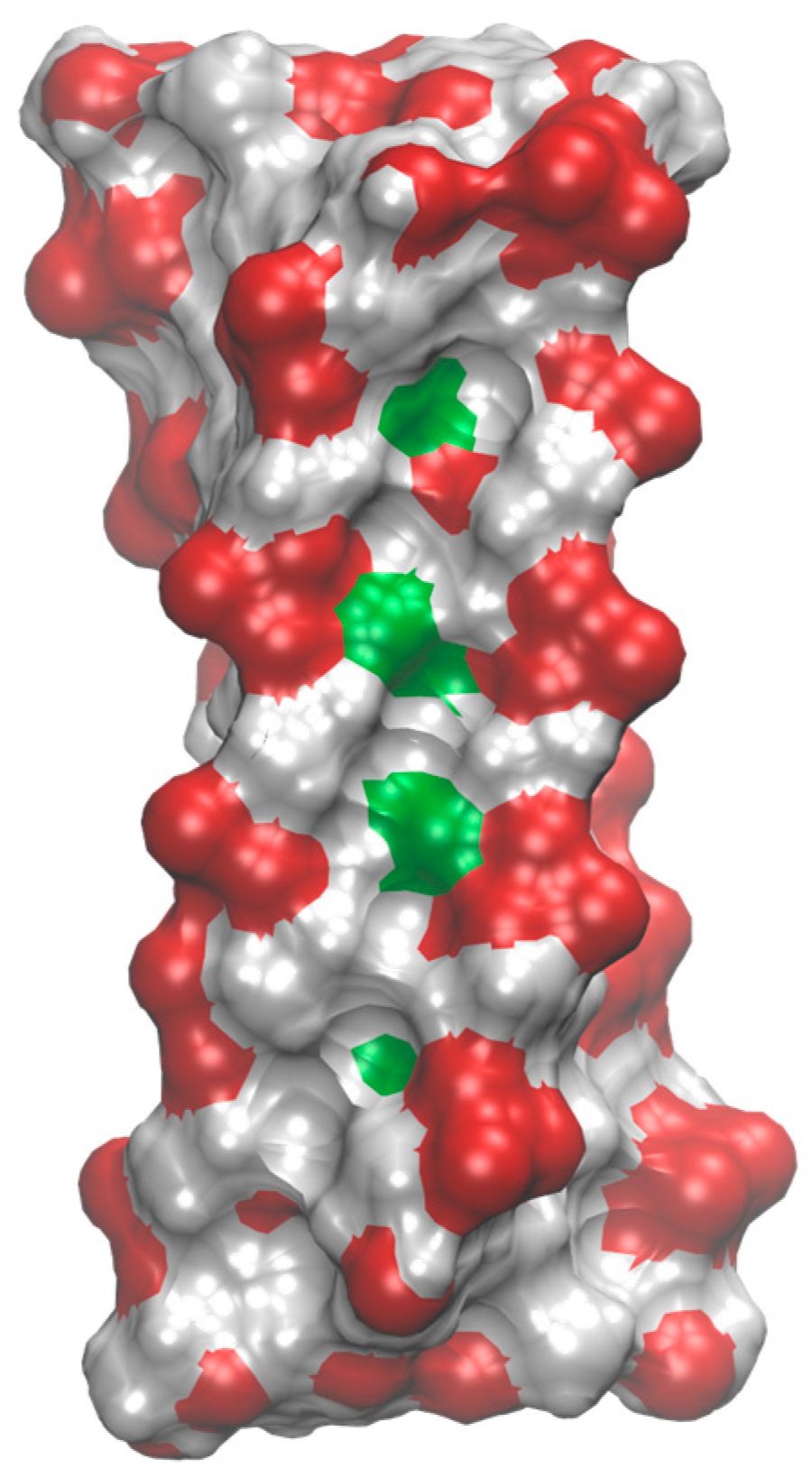
2.4. Structure Analysis
3. Discussion
4. Materials and Methods
4.1. Oligonucleotide Synthesis and Purification
4.2. Native Polyacrylamide Gel Electrophoresis (PAGE)
4.3. UV-Monitored Studies
4.4. NMR Experiments
4.5. Experimental NMR Constraints
4.6. Structure Determination and Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| 2′F-riboC | 2′-fluoro-ribo-cytosine |
| 2′F-araC | 2′-fluoro-arabino-cytosine |
| 2′F-RNA | Nucleic acids containing at least one 2′F-ribo-nucletides |
| 2′FANA | Nucleic acids containing at least one 2′F-arabino-nucletides |
| iFRNA | Imotif formed by 2′F-RNA |
| iFANA | Imotif formed by 2′FANA |
| iRNA | Imotif formed by RNA |
| iDNA | Imotif formed by DNA |
References
- Gehring, K.; Leroy, J.L.; Guéron, M. A Tetrameric DNA Structure with Protonated Cytosine-Cytosine Base Pairs. Nature 1993, 363, 561–565. [Google Scholar] [CrossRef]
- Abou Assi, H.; Garavís, M.; González, C.; Damha, M.J. i-Motif DNA: Structural Features and Significance to Cell Biology. Nucleic Acids Res. 2018, 46, 8038–8056. [Google Scholar] [CrossRef] [PubMed]
- Benabou, S.; Aviñó, A.; Eritja, R.; González, C.; Gargallo, R. Fundamental Aspects of the Nucleic Acid i-Motif Structures. RSC Adv. 2014, 4, 26956–26980. [Google Scholar] [CrossRef]
- Day, H.A.; Pavlou, P.; Waller, Z.A.E. i-Motif DNA: Structure, Stability and Targeting with Ligands. Bioorg. Med. Chem. 2014, 22, 4407–4418. [Google Scholar] [CrossRef] [PubMed]
- Tao, S.; Run, Y.; Monchaud, D.; Zhang, W. i-Motif DNA: Identification, Formation, and Cellular Functions. Trends Genet. 2024, 40, 853–867. [Google Scholar] [CrossRef]
- Sengupta, P.; Bose, D.; Chatterjee, S. The Molecular Tête-à-Tête Between G-Quadruplexes and the i-Motif in the Human Genome. ChemBioChem 2020, 22, 1517–1537. [Google Scholar] [CrossRef]
- Brown, S.L.; Kendrick, S. The i-Motif as a Molecular Target: More Than a Complementary DNA Secondary Structure. Pharmaceuticals 2021, 14, 96. [Google Scholar] [CrossRef]
- Deep, A.; Bhat, A.; Perumal, V.; Kumar, S. i-Motifs as Regulatory Switches: Mechanisms and Implications for Gene Expression. Mol. Ther. Nucleic Acids 2025, 36, 102474. [Google Scholar] [CrossRef]
- Brooks, T.A.; Kendrick, S.; Hurley, L. Making Sense of G-Quadruplex and i-Motif Functions in Oncogene Promoters. FEBS J. 2010, 277, 3459–3469. [Google Scholar] [CrossRef]
- Kendrick, S.; Kang, H.J.; Alam, M.P.; Madathil, M.M.; Agrawal, P.; Gokhale, V.; Yang, D.; Hecht, S.M.; Hurley, L.H. The Dynamic Character of the BCL2 Promoter i-Motif Provides a Mechanism for Modulation of Gene Expression by Compounds That Bind Selectively to the Alternative DNA Hairpin Structure. J. Am. Chem. Soc. 2014, 136, 4161–4171. [Google Scholar] [CrossRef]
- Zeraati, M.; Langley, D.B.; Schofield, P.; Moye, A.L.; Rouet, R.; Hughes, W.E.; Bryan, T.M.; Dinger, M.E.; Christ, D. i-Motif DNA Structures Are Formed in the Nuclei of Human Cells. Nat. Chem. 2018, 10, 631–637. [Google Scholar] [CrossRef] [PubMed]
- Boissieras, J.; Bonnet, H.; Susanto, M.F.; Gomez, D.; Granzhan, A.; Defrancq, E.; Dejeu, J. iMab Antibody Binds Single-Stranded Cytosine-Rich Sequences and Unfolds DNA i-Motifs. Nucleic Acids Res. 2023, 52, 8052–8062. [Google Scholar] [CrossRef] [PubMed]
- Amato, J.; Iaccarino, N.; Randazzo, A.; Novellino, E.; Pagano, B. Noncanonical DNA Secondary Structures as Drug Targets: The Prospect of the i-Motif. ChemMedChem 2014, 9, 2026–2030. [Google Scholar] [CrossRef]
- Bonnet, H.; Morel, M.; Devaux, A.; Boissieras, J.; Granzhan, A.; Elias, B.; Lavergne, T.; Dejeu, J.; Defrancq, E. Assessment of Presumed Small-Molecule Ligands of Telomeric i-DNA by Biolayer Interferometry (BLI). Chem. Commun. 2022, 58, 5116–5119. [Google Scholar] [CrossRef]
- Debnath, M.; Fatma, K.; Dash, J. Chemical Regulation of DNA i-Motifs for Nanobiotechnology and Therapeutics. Angew. Chem. Int. Ed. 2019, 131, 2968–2983. [Google Scholar] [CrossRef]
- Yatsunyk, L.A.; Mendoza, O.; Mergny, J.-L. “Nano-Oddities”: Unusual Nucleic Acid Assemblies for DNA-Based Nanostructures and Nanodevices. Acc. Chem. Res. 2014, 47, 1836–1844. [Google Scholar] [CrossRef]
- Dong, Y.; Yang, Z.; Liu, D. DNA Nanotechnology Based on i-Motif Structures. Acc. Chem. Res. 2014, 47, 1853–1860. [Google Scholar] [CrossRef]
- Pu, Q.; Lai, X.; Peng, Y.; Wu, Q. A Controllable DNA: Structural Features and Advanced Applications of i-Motif. Analyst 2025, 150, 1726–1740. [Google Scholar] [CrossRef] [PubMed]
- Luo, T.; Guan, Z.; Liu, Y.; Zhou, J.; Mergny, J.-L.; Cheng, M. i-Motif DNA Molecular Beacon for microRNA Detection. Nucleic Acids Res. 2025, 53, gkaf556. [Google Scholar] [CrossRef]
- Wright, E.P.; Huppert, J.L.; Waller, Z.A.E. Identification of Multiple Genomic DNA Sequences Which Form i-Motif Structures at Neutral pH. Nucleic Acids Res. 2017, 45, 2951–2959. [Google Scholar] [CrossRef] [PubMed]
- Fleming, A.M.; Ding, Y.; Rogers, R.A.; Zhu, J.; Zhu, J.; Burton, A.D.; Carlisle, C.B.; Burrows, C.J. 4n−1 Is a “Sweet Spot” in DNA i-Motif Folding of 2′-Deoxycytidine Homopolymers. J. Am. Chem. Soc. 2017, 139, 4682–4689. [Google Scholar] [CrossRef] [PubMed]
- Mir, B.; Serrano, I.; Buitrago, D.; Orozco, M.; Escaja, N.; González, C. Prevalent Sequences in the Human Genome Can Form Mini i-Motif Structures at Physiological pH. J. Am. Chem. Soc. 2017, 139, 13985–13988. [Google Scholar] [CrossRef]
- Berger, I.; Egli, M.; Rich, A. Inter-Strand C-H...O Hydrogen Bonds Stabilizing Four-Stranded Intercalated Molecules: Stereoelectronic Effects of O4’ in Cytosine-Rich DNA. Proc. Natl. Acad. Sci. USA 1996, 93, 12116–12121. [Google Scholar] [CrossRef]
- Pérez-Rentero, S.; Gargallo, R.; González, C.; Eritja, R. Modulation of the Stability of i-Motif Structures Using an Acyclic Threoninol Cytidine Derivative. RSC Adv. 2015, 5, 63278–63281. [Google Scholar] [CrossRef]
- Lannes, L.; Halder, S.; Krishnan, Y.; Schwalbe, H. Tuning the pH Response of i-Motif DNA Oligonucleotides. ChemBioChem 2015, 16, 1647–1656. [Google Scholar] [CrossRef]
- Pasternak, A.; Wengel, J. Modulation of i-Motif Thermodynamic Stability by the Introduction of UNA (Unlocked Nucleic Acid) Monomers. Bioorganic Med. Chem. Lett. 2011, 21, 752–755. [Google Scholar] [CrossRef] [PubMed]
- Snoussi, K.; Nonin-Lecomte, S.; Leroy, J.-L. The RNA i-Motif. J. Mol. Biol. 2001, 309, 139–153. [Google Scholar] [CrossRef] [PubMed]
- Collin, D.; Gehring, K. Stability of Chimeric DNA/RNA Cytosine Tetrads: Implications for i-Motif Formation by RNA. J. Am. Chem. Soc. 1998, 120, 4069–4072. [Google Scholar] [CrossRef]
- Kumar, N.; Petersen, M.; Maiti, S. Tunable c-MYC LNA i-Motif. Chem. Commun. 2009, 1532–1534. [Google Scholar] [CrossRef]
- Abou Assi, H.; Harkness, R.W.R.W.; Martin-Pintado, N.; Wilds, C.J.C.J.; Campos-Olivas, R.; Mittermaier, A.K.A.K.; González, C.; Damha, M.J.M.J. Stabilization of i-Motif Structures by 2’-β-Fluorination of DNA. Nucleic Acids Res. 2016, 44, 4998–5009. [Google Scholar] [CrossRef]
- Abou Assi, H.; El-Khoury, R.; González, C.; Damha, M.J. 2′-Fluoroarabinonucleic Acid Modification Traps G-Quadruplex and i-Motif Structures in Human Telomeric DNA. Nucleic Acids Res. 2017, 45, 11535–11546. [Google Scholar] [CrossRef] [PubMed]
- Robidoux, S.; Damha, M.J. D-2-Deoxyribose and D-Arabinose, but Not D-Ribose, Stabilize the Cytosine Tetrad (i-DNA) Structure. J. Biomol. Struct. Dyn. 1997, 15, 529–535. [Google Scholar] [CrossRef] [PubMed]
- Aviñó, A.; Dellafiore, M.; Gargallo, R.; González, C.; Iribarren, A.M.A.M.; Montserrat, J.; Eritja, R. Stabilization of Telomeric i-Motif Structures by (2′S)-2′-Deoxy-2′-C-Methylcytidine Residues. ChemBioChem 2017, 18, 1123–1128. [Google Scholar] [CrossRef]
- El-Khoury, R.; Macaluso, V.; Hennecker, C.; Mittermaier, A.K.; Orozco, M.; González, C.; Garavís, M.; Damha, M.J. i-Motif Folding Intermediates with Zero-Nucleotide Loops Are Trapped by 2′-Fluoroarabinocytidine via F···H and O···H Hydrogen Bonds. Commun. Chem. 2023, 6, 31. [Google Scholar] [CrossRef]
- Fenna, C.P.; Wilkinson, V.J.; Arnold, J.R.P.; Cosstick, R.; Fisher, J. The Effect of 2′-Fluorine Substitutions on DNA i-Motif Conformation and Stability. Chem. Commun. 2008, 3567–3569. [Google Scholar] [CrossRef] [PubMed]
- Bao, H.-L.; Masuzawa, T.; Oyoshi, T.; Xu, Y. Oligonucleotides DNA Containing 8-Trifluoromethyl-2′-Deoxyguanosine for Observing Z-DNA Structure. Nucleic Acids Res. 2020, 48, 7041–7051. [Google Scholar] [CrossRef]
- Baranowski, M.R.; Warminski, M.; Jemielity, J.; Kowalska, J. 5′-Fluoro(Di)Phosphate-Labeled Oligonucleotides Are Versatile Molecular Probes for Studying Nucleic Acid Secondary Structure and Interactions by 19F NMR. Nucleic Acids Res. 2020, 48, 8209–8224. [Google Scholar] [CrossRef]
- Rout, B.P.; Roy, S.; Srivatsan, S.G. 5-Fluoro-2′-Deoxyuridine as an Efficient 19F NMR Reporter for G-Quadruplex and i-Motif Structures. Bioorg. Med. Chem. Lett. 2025, 117, 130060. [Google Scholar] [CrossRef]
- El-Khoury, R.; Cabrero, C.; Movilla, S.; Kaur, H.; Friedland, D.; Domínguez, A.; Thorpe, J.D.; Roman, M.; Orozco, M.; González, C.; et al. Formation of Left-Handed Helices by C2′-Fluorinated Nucleic Acids Under Physiological Salt Conditions. Nucleic Acids Res. 2024, 52, 7414–7428. [Google Scholar] [CrossRef]
- Watts, J.K.; Martín-Pintado, N.; Gómez-Pinto, I.; Schwartzentruber, J.; Portella, G.; Orozco, M.; González, C.; Damha, M.J. Differential Stability of 2′F-ANA•RNA and ANA•RNA Hybrid Duplexes: Roles of Structure, Pseudohydrogen Bonding, Hydration, Ion Uptake and Flexibility. Nucleic Acids Res. 2010, 38, 2498–2511. [Google Scholar] [CrossRef]
- Martin-Pintado, N.; Deleavey, G.F.G.F.; Portella, G.; Campos-Olivas, R.; Orozco, M.; Damha, M.J.M.J.; González, C. Backbone FC-H···O Hydrogen Bonds in 2′F-Substituted Nucleic Acids. Angew. Chem. Int. Ed. 2013, 52, 12065–12068. [Google Scholar] [CrossRef]
- Martín-Pintado, N.; Yahyaee-Anzahaee, M.; Deleavey, G.F.; Portella, G.; Orozco, M.; Damha, M.J.; González, C. Dramatic Effect of Furanose C2′ Substitution on Structure and Stability: Directing the Folding of the Human Telomeric Quadruplex with a Single Fluorine Atom. J. Am. Chem. Soc. 2013, 135, 5344–5347. [Google Scholar] [CrossRef]
- Martín-Pintado, N.; Yahyaee-Anzahaee, M.; Campos-Olivas, R.; Noronha, A.M.; Wilds, C.J.; Damha, M.J.; González, C. The Solution Structure of Double Helical Arabino Nucleic Acids (ANA and 2′F-ANA): Effect of Arabinoses in Duplex-Hairpin Interconversion. Nucleic Acids Res. 2012, 40, 9329–9339. [Google Scholar] [CrossRef]
- Berger, I.; Egli, M. The Role of Backbone Oxygen Atoms in the Organization of Nucleic Acid Tertiary Structure: Zippers, Networks, Clamps, and C-H ··· O Hydrogen Bonds. Chem.—Eur. J. 1997, 3, 1400–1404. [Google Scholar] [CrossRef]
- Pallan, P.S.; Greene, E.M.; Jicman, P.A.; Pandey, R.K.; Manoharan, M.; Rozners, E.; Egli, M. Unexpected Origins of the Enhanced Pairing Affinity of 2’-Fluoro-Modified RNA. Nucleic Acids Res. 2011, 39, 3482–3495. [Google Scholar] [CrossRef]
- Manoharan, M.; Akinc, A.; Pandey, R.K.; Qin, J.; Hadwiger, P.; John, M.; Mills, K.; Charisse, K.; Maier, M.A.; Nechev, L.; et al. Unique Gene-Silencing and Structural Properties of 2′-Fluoro-Modified siRNAs. Angew. Chem. Int. Ed. 2011, 50, 2284–2288. [Google Scholar] [CrossRef]
- Khatik, S.Y.; Srivatsan, S.G. Environment-Sensitive Nucleoside Probe Unravels the Complex Structural Dynamics of i-Motif DNAs. Bioconj. Chem. 2022, 33, 1515–1526. [Google Scholar] [CrossRef] [PubMed]
- Wee, W.A.; Yum, J.H.; Hirashima, S.; Sugiyama, H.; Park, S. Synthesis and Application of a 19F-Labeled Fluorescent Nucleoside as a Dual-Mode Probe for i-Motif DNAs. RSC Chem. Biol. 2021, 2, 876–882. [Google Scholar] [CrossRef] [PubMed]
- Ishizuka, T.; Yamashita, A.; Asada, Y.; Xu, Y. Studying DNA g-Quadruplex Aptamer by 19F NMR. ACS Omega 2017, 2, 8843–8848. [Google Scholar] [CrossRef]
- Manna, S.; Sarkar, D.; Srivatsan, S.G. A Dual-App Nucleoside Probe Provides Structural Insights into the Human Telomeric Overhang in Live Cells. J. Am. Chem. Soc. 2018, 140, 12622–12633. [Google Scholar] [CrossRef] [PubMed]
- Bao, H.-L.; Liu, H.; Xu, Y. Hybrid-Type and Two-Tetrad Antiparallel Telomere DNA G-Quadruplex Structures in Living Human Cells. Nucleic Acids Res. 2019, 47, 4940–4947. [Google Scholar] [CrossRef] [PubMed]
- Viskova, P.; Krafcik, D.; Trantirek, L.; Foldynova-Trantirkova, S. In-Cell NMR Spectroscopy of Nucleic Acids in Human Cells. Curr. Protoc. Nucleic Acid Chem. 2019, 76, e71. [Google Scholar] [CrossRef]
- Viskova, P.; Istvankova, E.; Rynes, J.; Dzatko, S.; Loja, T.; Zivkovic, M.L.; Rigo, R.; El-Khoury, R.; Serrano-Chacón, I.; Damha, M.J.; et al. In cell NMR suggests that DNA i-motif levels are strongly depleted in living human cells. Nat. Comm. 2024, 15, 1992. [Google Scholar] [CrossRef]
- Goddard, D.T.; Kneller, G. SPARKY, 3rd ed.; University of California: San Francisco, CA, USA, 2000. [Google Scholar]
- Case, D.A.; Pearlman, D.A.; Caldwell, J.W.; Cheatham, T.E., III; Wang, J.; Ross, W.S.; Simmerling, C.L.; Darden, T.A.; Merz, K.M.; Stanton, R.V.; et al. AMBER 18; University of California: San Francisco, CA, USA, 2018. [Google Scholar]
- Darden, T.E.; York, D.; Pedersen, L. Particle Mesh Ewald: An N Log (N) Method for Ewald Sums in Large Systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]
- Ivani, I.; Dans, P.D.; Noy, A.; Pérez, A.; Faustino, I.; Hospital, A.; Walther, J.; Andrio, P.; Goñi, R.; Balaceanu, A.; et al. Parmbsc1: A Refined Force Field for DNA Simulations. Nat. Methods 2015, 13, 55–58. [Google Scholar] [CrossRef]
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 1983, 79, 926. [Google Scholar] [CrossRef]
- Lavery, R.; Sklenar, V. Curves+, Helical Analysis of Irregular Nucleic Acids; Laboratory of Theoretical Chemistry, CNRS: Paris, France, 1990. [Google Scholar]
- Lu, X.J.; Olson, W.K. 3DNA: A Versatile, Integrated Software System for the Analysis, Rebuilding and Visualization of Three-Dimensional Nucleic-Acid Structures. Nat. Protoc. 2008, 3, 1213–1227. [Google Scholar] [CrossRef]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual Molecular Dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Koradi, R.; Billeter, M.; Wüthrich, K. MOLMOL: A Program for Display and Analysis of Macromolecular Structures. J. Mol. Graphics 1996, 14, 29–32. [Google Scholar] [CrossRef] [PubMed]



| Name | Sequence (5′-3′) | T1/2 pH 5 (°C) | T1/2 pH 6 (°C) 2 |
|---|---|---|---|
| FR1 | dTCC(frC)CC | 55.9 | - |
| FR2 | dTCC(frC)(frC)C | 64.8 | - |
| FRfull | dT(frC)(frC)(frC)(frC)(frC) | - | - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ugedo, C.; Domínguez, A.; Gómez-Pinto, I.; Eritja, R.; González, C.; Aviñó, A. The Effect of 2′F-RNA on I-Motif Structure and Stability. Molecules 2025, 30, 3561. https://doi.org/10.3390/molecules30173561
Ugedo C, Domínguez A, Gómez-Pinto I, Eritja R, González C, Aviñó A. The Effect of 2′F-RNA on I-Motif Structure and Stability. Molecules. 2025; 30(17):3561. https://doi.org/10.3390/molecules30173561
Chicago/Turabian StyleUgedo, Cristina, Arnau Domínguez, Irene Gómez-Pinto, Ramon Eritja, Carlos González, and Anna Aviñó. 2025. "The Effect of 2′F-RNA on I-Motif Structure and Stability" Molecules 30, no. 17: 3561. https://doi.org/10.3390/molecules30173561
APA StyleUgedo, C., Domínguez, A., Gómez-Pinto, I., Eritja, R., González, C., & Aviñó, A. (2025). The Effect of 2′F-RNA on I-Motif Structure and Stability. Molecules, 30(17), 3561. https://doi.org/10.3390/molecules30173561










