A Unique Trimeric Assembly of Human Dishevelled 1 PDZ Domain in Crystal: Implication of Homo- and Hetero-Oligomerization During Wnt Signaling Process
Abstract
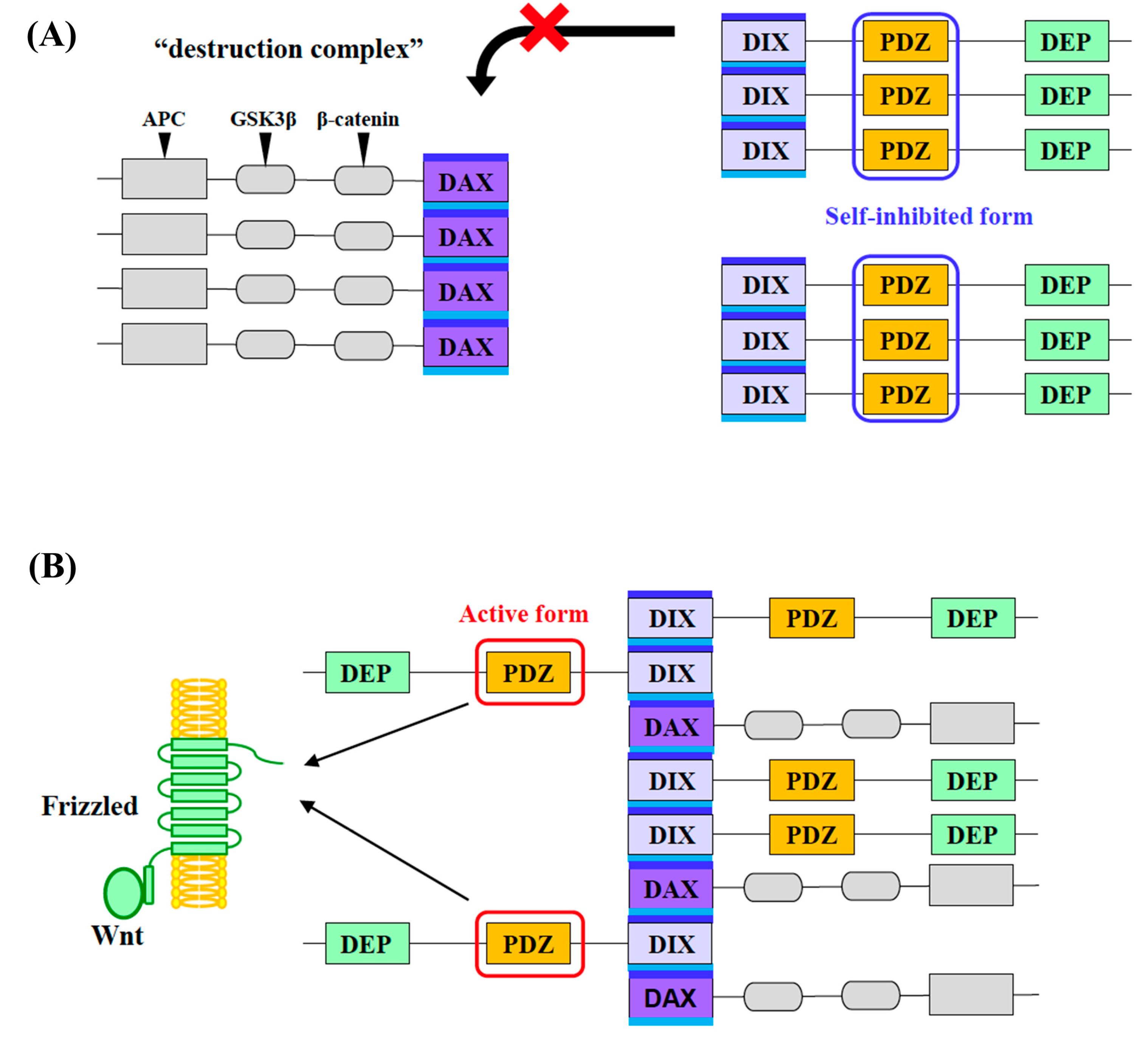
1. Introduction
2. Results
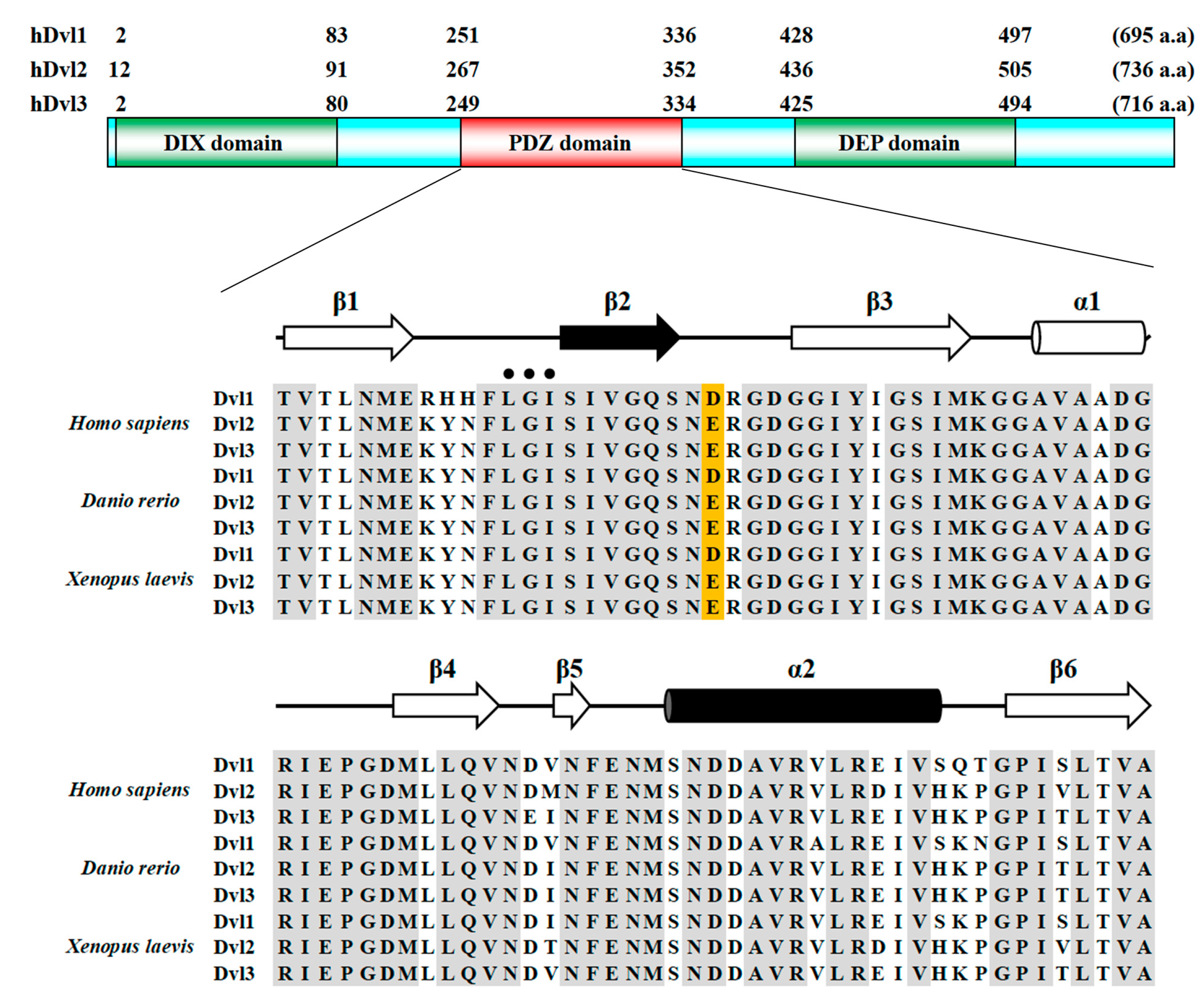
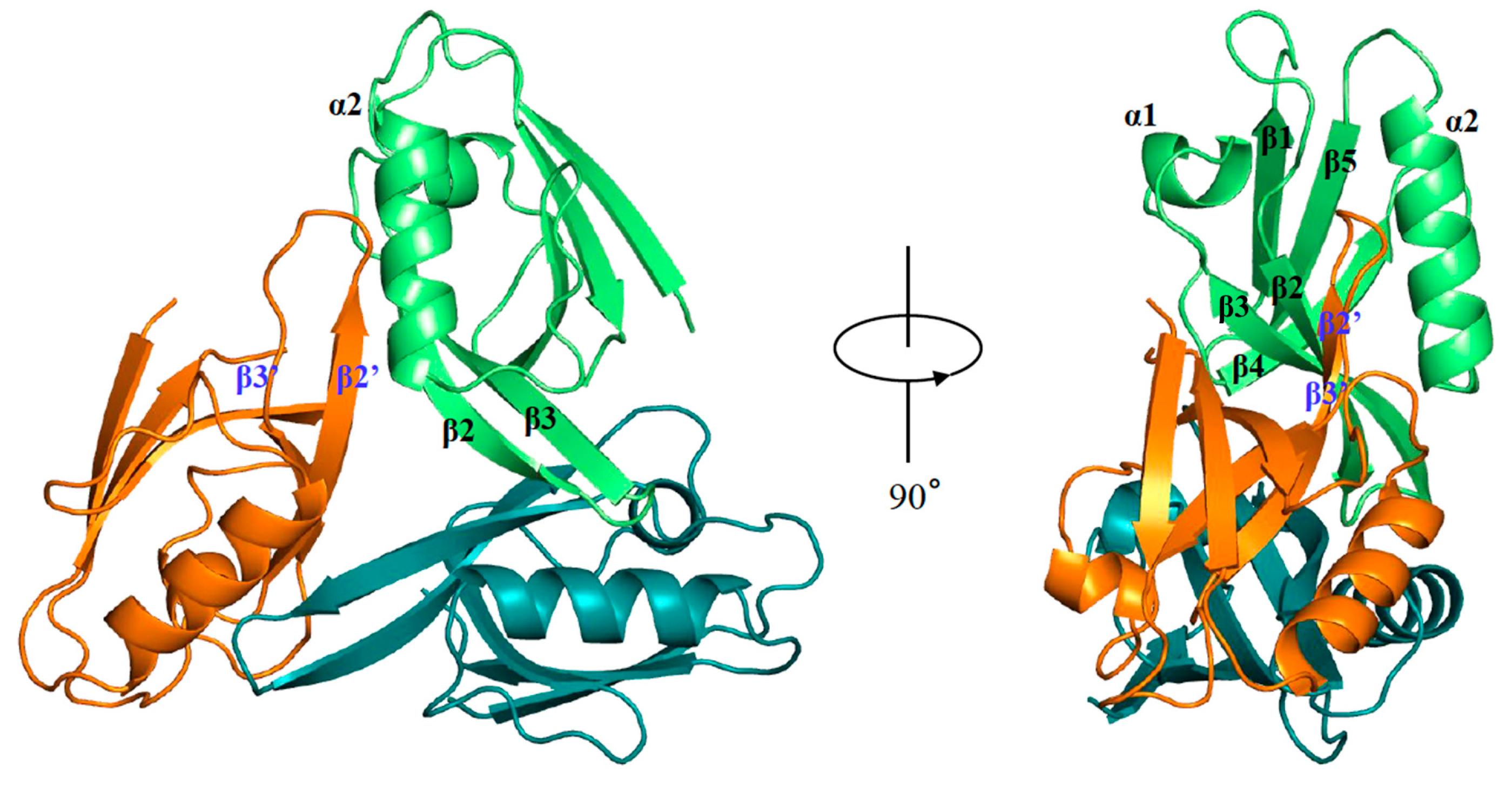
2.1. Overall Structural Arrangement of the PDZ Domain of hDvl1 in Crystal
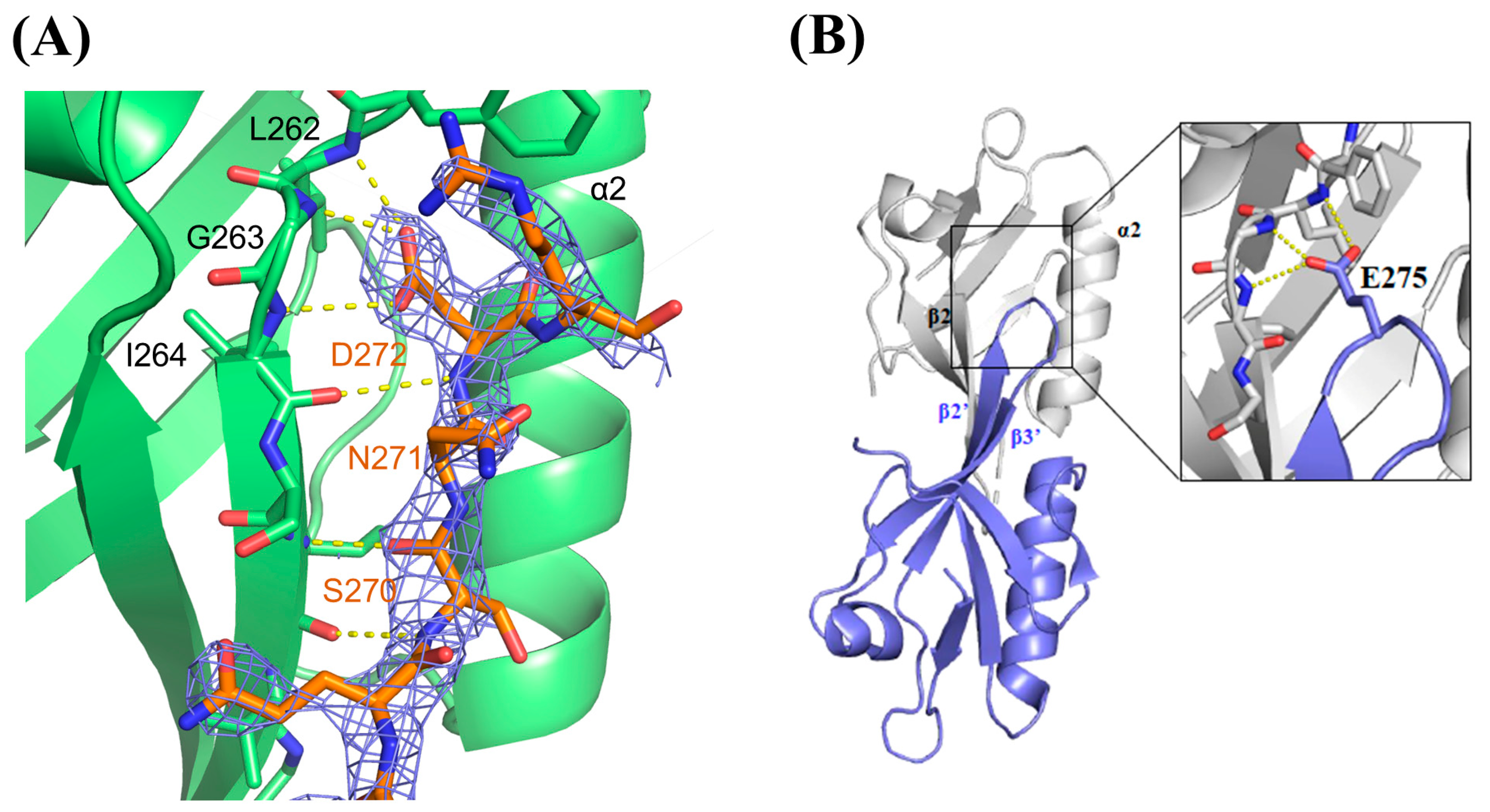
2.2. Dvl PDZ Domain Shows Symmetrical Trimer in Crystal Structure
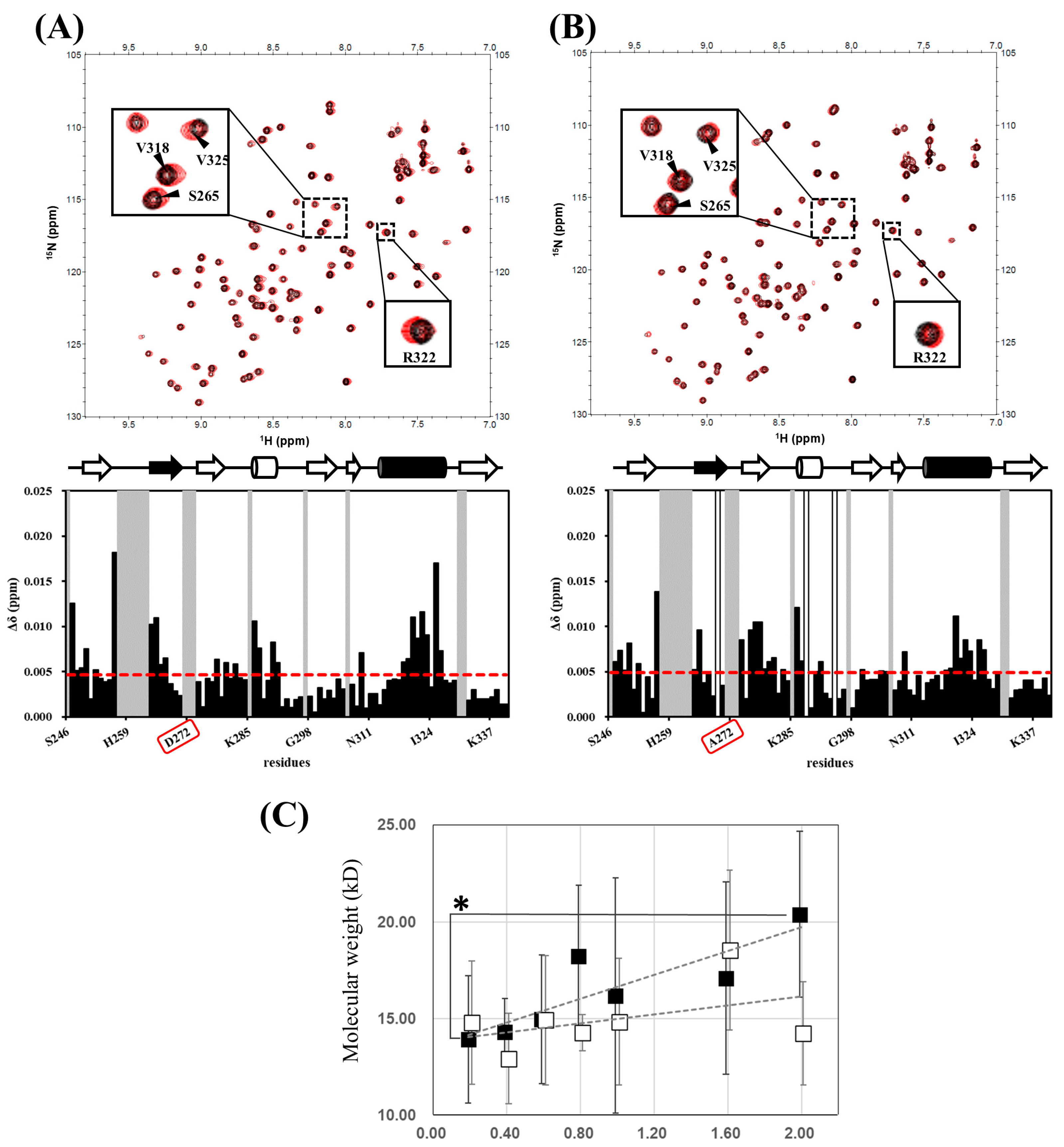
2.3. Equilibrium of hDvl1 PDZ Self-Assembly in Solution Confirmed by NMR and Dynamic Light Scattering Experiments
3. Discussion
4. Materials and Methods
4.1. Preparation of Protein Samples
4.2. Crystallization, Data Collection, Structure Determination, and Refinement
4.3. NMR Experiments
4.4. Dynamic Light Scattering
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| PDB | Protein Data Bank |
| Dvl | Dishevelled |
| PDZ | PSD-95, disks large, and zonula occludens-1 |
| DIX | Dvl/AXIN |
| DEP | Dvl/Egl-10/Pleckstrin |
| Fzd | Frizzled |
| LLPS | Liquid–liquid phase separation |
| NMR | Nuclear magnetic resonance |
| HSQC | Heteronuclear single quantum coherence spectroscopy |
| GSK | glycogen synthase kinase |
| APC | adenomatous polyposis coli |
References
- Logan, C.Y.; Nusse, R. The Wnt Signaling Pathway in Development and Disease. Annu. Rev. Cell Dev. Biol. 2004, 20, 781–810. [Google Scholar] [CrossRef]
- Clevers, H.; Nusse, R. Wnt/β-Catenin Signaling and Disease. Cell 2012, 149, 1192–1205. [Google Scholar] [CrossRef]
- Reya, T.; Clevers, H. Wnt Signalling in Stem Cells and Cancer. Nature 2005, 434, 843–850. [Google Scholar] [CrossRef] [PubMed]
- Clevers, H. Wnt/β-Catenin Signaling in Development and Disease. Cell 2006, 127, 469–480. [Google Scholar] [CrossRef]
- MacDonald, B.T.; Tamai, K.; He, X. Wnt/β-Catenin Signaling: Components, Mechanisms, and Diseases. Dev. Cell 2009, 17, 9–26. [Google Scholar] [CrossRef] [PubMed]
- Schwarz-Romond, T.; Metcalfe, C.; Bienz, M. Dynamic Recruitment of Axin by Dishevelled Protein Assemblies. J. Cell Sci. 2007, 120, 2402–2412. [Google Scholar] [CrossRef] [PubMed]
- Fiedler, M.; Mendoza-Topaz, C.; Rutherford, T.J.; Mieszczanek, J.; Bienz, M. Dishevelled Interacts with the DIX Domain Polymerization Interface of Axin to Interfere with Its Function in Down-Regulating β-Catenin. Proc. Natl. Acad. Sci. USA 2011, 108, 1937–1942. [Google Scholar] [CrossRef]
- Bienz, M. Signalosome Assembly by Domains Undergoing Dynamic Head-to-Tail Polymerization. Trends Biochem. Sci. 2014, 39, 487–495. [Google Scholar] [CrossRef]
- Smalley, M.J.; Signoret, N.; Robertson, D.; Tilley, A.; Hann, A.; Ewan, K.; Ding, Y.; Paterson, H.; Dale, T.C. Dishevelled (Dvl-2) Activates Canonical Wnt Signalling in the Absence of Cytoplasmic Puncta. J. Cell Sci. 2005, 118, 5279–5289. [Google Scholar] [CrossRef]
- Sear, R.P. Dishevelled: A Protein That Functions in Living Cells by Phase Separating. Soft Matter 2007, 3, 680. [Google Scholar] [CrossRef]
- Kang, K.; Shi, Q.; Wang, X.; Chen, Y.-G. Dishevelled Phase Separation Promotes Wnt Signalosome Assembly and Destruction Complex Disassembly. J. Cell Biol. 2022, 221. [Google Scholar] [CrossRef] [PubMed]
- Dai, D.-L.; Xie, C.; Zhong, L.-Y.; Liu, S.-X.; Zhang, L.-L.; Zhang, H.; Wu, X.-P.; Wu, Z.-M.; Kang, K.; Li, Y.; et al. AXIN1 Boosts Antiviral Response through IRF3 Stabilization and Induced Phase Separation. Signal Transduct. Target Ther. 2024, 9, 281. [Google Scholar] [CrossRef]
- Zhang, D.; Ni, Q.-Q.; Wang, S.-Y.; He, W.-F.; Hong, Z.-X.; Liu, H.-Y.; Chen, X.-H.; Chen, L.-J.; Han, F.-Y.; Zhang, L.-J.; et al. APC Mutations Disrupt β-Catenin Destruction Complex Condensates Organized by Axin Phase Separation. Cell. Mol. Life Sci. 2024, 81, 57. [Google Scholar] [CrossRef] [PubMed]
- Schaefer, K.N.; Peifer, M. Wnt/Beta-Catenin Signaling Regulation and a Role for Biomolecular Condensates. Dev. Cell 2019, 48, 429–444. [Google Scholar] [CrossRef]
- Bienz, M. Head-to-Tail Polymerization in the Assembly of Biomolecular Condensates. Cell 2020, 182, 799–811. [Google Scholar] [CrossRef] [PubMed]
- Nong, J.; Kang, K.; Shi, Q.; Zhu, X.; Tao, Q.; Chen, Y.-G. Phase Separation of Axin Organizes the β-Catenin Destruction Complex. J. Cell Biol. 2021, 220. [Google Scholar] [CrossRef]
- Shi, Q.; Kang, K.; Chen, Y.-G. Liquid–Liquid Phase Separation Drives the Β-catenin Destruction Complex Formation. BioEssays 2021, 43, 2100138. [Google Scholar] [CrossRef]
- Li, T.-M.; Ren, J.; Husmann, D.; Coan, J.P.; Gozani, O.; Chua, K.F. Multivalent Tumor Suppressor Adenomatous Polyposis Coli Promotes Axin Biomolecular Condensate Formation and Efficient β-Catenin Degradation. Sci. Rep. 2020, 10, 17425. [Google Scholar] [CrossRef]
- Schwarz-Romond, T.; Fiedler, M.; Shibata, N.; Butler, P.J.G.; Kikuchi, A.; Higuchi, Y.; Bienz, M. The DIX Domain of Dishevelled Confers Wnt Signaling by Dynamic Polymerization. Nat. Struct. Mol. Biol. 2007, 14, 484–492. [Google Scholar] [CrossRef]
- Kishida, S.; Yamamoto, H.; Hino, S.; Ikeda, S.; Kishida, M.; Kikuchi, A. DIX Domains of Dvl and Axin Are Necessary for Protein Interactions and Their Ability to Regulate β-Catenin Stability. Mol. Cell Biol. 1999, 19, 4414–4422. [Google Scholar] [CrossRef]
- Songyang, Z.; Fanning, A.S.; Fu, C.; Xu, J.; Marfatia, S.M.; Chishti, A.H.; Crompton, A.; Chan, A.C.; Andersen, J.M.; Cantley, L.C. Recognition of Unique Carboxyl-Terminal Motifs by Distinct PDZ Domains. Science 1997, 275, 73–77. [Google Scholar] [CrossRef]
- Wong, H.C.; Mao, J.; Nguyen, J.T.; Srinivas, S.; Zhang, W.; Liu, B.; Li, L.; Wu, D.; Zheng, J. Structural Basis of the Recognition of the Dishevelled DEP Domain in the Wnt Signaling Pathway. Nat. Struct. Biol. 2000, 7, 1178–1184. [Google Scholar] [CrossRef][Green Version]
- Madrzak, J.; Fiedler, M.; Johnson, C.M.; Ewan, R.; Knebel, A.; Bienz, M.; Chin, J.W. Ubiquitination of the Dishevelled DIX Domain Blocks Its Head-to-Tail Polymerization. Nat. Commun. 2015, 6, 6718. [Google Scholar] [CrossRef]
- Gammons, M.V.; Renko, M.; Johnson, C.M.; Rutherford, T.J.; Bienz, M. Wnt Signalosome Assembly by DEP Domain Swapping of Dishevelled. Mol. Cell 2016, 64, 92–104. [Google Scholar] [CrossRef]
- Wong, H.-C.; Bourdelas, A.; Krauss, A.; Lee, H.-J.; Shao, Y.; Wu, D.; Mlodzik, M.; Shi, D.-L.; Zheng, J. Direct Binding of the PDZ Domain of Dishevelled to a Conserved Internal Sequence in the C-Terminal Region of Frizzled. Mol. Cell 2003, 12, 1251–1260. [Google Scholar] [CrossRef] [PubMed]
- Hori, K.; Ajioka, K.; Goda, N.; Shindo, A.; Takagishi, M.; Tenno, T.; Hiroaki, H. Discovery of Potent Disheveled/Dvl Inhibitors Using Virtual Screening Optimized With NMR-Based Docking Performance Index. Front. Pharmacol. 2018, 9, 983. [Google Scholar] [CrossRef] [PubMed]
- Axelrod, J.D.; Miller, J.R.; Shulman, J.M.; Moon, R.T.; Perrimon, N. Differential Recruitment of Dishevelled Provides Signaling Specificity in the Planar Cell Polarity and Wingless Signaling Pathways. Genes Dev. 1998, 12, 2610–2622. [Google Scholar] [CrossRef]
- Rothbächer, U.; Laurent, M.N.; Deardorff, M.A.; Klein, P.S.; Cho, K.W.Y.; Fraser, S.E. Dishevelled Phosphorylation, Subcellular Localization and Multimerization Regulate Its Role in Early Embryogenesis. EMBO J. 2000, 19, 1010–1022. [Google Scholar] [CrossRef] [PubMed]
- Qi, J.; Lee, H.-J.; Saquet, A.; Cheng, X.-N.; Shao, M.; Zheng, J.J.; Shi, D.-L. Autoinhibition of Dishevelled Protein Regulated by Its Extreme C Terminus Plays a Distinct Role in Wnt/β-Catenin and Wnt/Planar Cell Polarity (PCP) Signaling Pathways. J. Biol. Chem. 2017, 292, 5898–5908. [Google Scholar] [CrossRef]
- Lee, H.-J.; Shi, D.-L.; Zheng, J.J. Conformational Change of Dishevelled Plays a Key Regulatory Role in the Wnt Signaling Pathways. Elife 2015, 4, e08142. [Google Scholar] [CrossRef][Green Version]
- Schubert, A.; Voloshanenko, O.; Ragaller, F.; Gmach, P.; Kranz, D.; Scheeder, C.; Miersch, T.; Schulz, M.; Trümper, L.; Binder, C.; et al. Superresolution Microscopy Localizes Endogenous Dvl2 to Wnt Signaling-Responsive Biomolecular Condensates. Proc. Natl. Acad. Sci. USA 2022, 119, e2122476119. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Banjade, S.; Cheng, H.-C.; Kim, S.; Chen, B.; Guo, L.; Llaguno, M.; Hollingsworth, J.V.; King, D.S.; Banani, S.F.; et al. Phase Transitions in the Assembly of Multivalent Signalling Proteins. Nature 2012, 483, 336–340. [Google Scholar] [CrossRef] [PubMed]
- Kan, W.; Enos, M.D.; Korkmazhan, E.; Muennich, S.; Chen, D.-H.; Gammons, M.V.; Vasishtha, M.; Bienz, M.; Dunn, A.R.; Skiniotis, G.; et al. Limited Dishevelled/Axin Oligomerization Determines Efficiency of Wnt/β-Catenin Signal Transduction. eLife 2020, 9, e55015. [Google Scholar] [CrossRef]
- Chavas, L.M.G.; Matsugaki, N.; Yamada, Y.; Hiraki, M.; Igarashi, N.; Suzuki, M.; Wakatsuki, S. Beamline AR-NW12A: High-Throughput Beamline for Macromolecular Crystallography at the Photon Factory. J. Synchrotron Radiat. 2012, 19, 450–454. [Google Scholar] [CrossRef]
- Kabsch, W. XDS. Acta Cryst. 2010, 66, 125–132. [Google Scholar] [CrossRef]
- Winn, M.D.; Ballard, C.C.; Cowtan, K.D.; Dodson, E.J.; Emsley, P.; Evans, P.R.; Keegan, R.M.; Krissinel, E.B.; Leslie, A.G.W.; McCoy, A.; et al. Overview of the CCP4 Suite and Current Developments. Acta Cryst. 2011, 67, 235–242. [Google Scholar]
- McCoy, A.J.; Grosse-Kunstleve, R.W.; Adams, P.D.; Winn, M.D.; Storoni, L.C.; Read, R.J. Phaser Crystallographic Software. J. Appl. Crystallogr. 2007, 40, 658–674. [Google Scholar] [CrossRef]
- Emsley, P.; Lohkamp, B.; Scott, W.G.; Cowtan, K. Features and Development of Coot. Acta Cryst. 2010, 66, 486–501. [Google Scholar] [CrossRef]
- Adams, P.D.; Afonine, P.V.; Bunkóczi, G.; Chen, V.B.; Davis, I.W.; Echols, N.; Headd, J.J.; Hung, L.W.; Kapral, G.J.; Grosse-Kunstleve, R.W.; et al. PHENIX: A Comprehensive Python-Based System for Macromolecular Structure Solution. Acta Cryst. 2010, 66, 213–221. [Google Scholar] [CrossRef]
- Chen, V.B.; Arendall, W.B.; Headd, J.J.; Keedy, D.A.; Immormino, R.M.; Kapral, G.J.; Murray, L.W.; Richardson, J.S.; Richardson, D.C. MolProbity: All-Atom Structure Validation for Macromolecular Crystallography. Acta Cryst. 2010, 66, 12–21. [Google Scholar] [CrossRef] [PubMed]
| Intramolecular | Intermolecular | |
|---|---|---|
| Interaction Type | PDZ: C-terminus | PDZ-PDZ |
| PDZ Binding Mode | canonical | non-canonical |
| Cellular Location | cytosol | puncta |
| Effective Concentration | below 12 µM | >1000 µM |
| Regulatory Trigger | post-translational modification | elevated concentration |
| Reference | Qi et. al. [29] | this study |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yasukochi, S.; Numoto, N.; Hori, K.; Tenno, T.; Hibino, E.; Ito, N.; Hiroaki, H. A Unique Trimeric Assembly of Human Dishevelled 1 PDZ Domain in Crystal: Implication of Homo- and Hetero-Oligomerization During Wnt Signaling Process. Molecules 2025, 30, 3538. https://doi.org/10.3390/molecules30173538
Yasukochi S, Numoto N, Hori K, Tenno T, Hibino E, Ito N, Hiroaki H. A Unique Trimeric Assembly of Human Dishevelled 1 PDZ Domain in Crystal: Implication of Homo- and Hetero-Oligomerization During Wnt Signaling Process. Molecules. 2025; 30(17):3538. https://doi.org/10.3390/molecules30173538
Chicago/Turabian StyleYasukochi, Shotaro, Nobutaka Numoto, Kiminori Hori, Takeshi Tenno, Emi Hibino, Nobutoshi Ito, and Hidekazu Hiroaki. 2025. "A Unique Trimeric Assembly of Human Dishevelled 1 PDZ Domain in Crystal: Implication of Homo- and Hetero-Oligomerization During Wnt Signaling Process" Molecules 30, no. 17: 3538. https://doi.org/10.3390/molecules30173538
APA StyleYasukochi, S., Numoto, N., Hori, K., Tenno, T., Hibino, E., Ito, N., & Hiroaki, H. (2025). A Unique Trimeric Assembly of Human Dishevelled 1 PDZ Domain in Crystal: Implication of Homo- and Hetero-Oligomerization During Wnt Signaling Process. Molecules, 30(17), 3538. https://doi.org/10.3390/molecules30173538








