Trident Nano-Indexing the Proteomics Table: Next-Version Clustering of Iron Carbide NPs and Protein Corona
Abstract
:1. Introduction
2. Results and Discussion
2.1. Identification and Quantification of Differentially Expressed Proteins
2.2. Differential Expression Based on Regulation
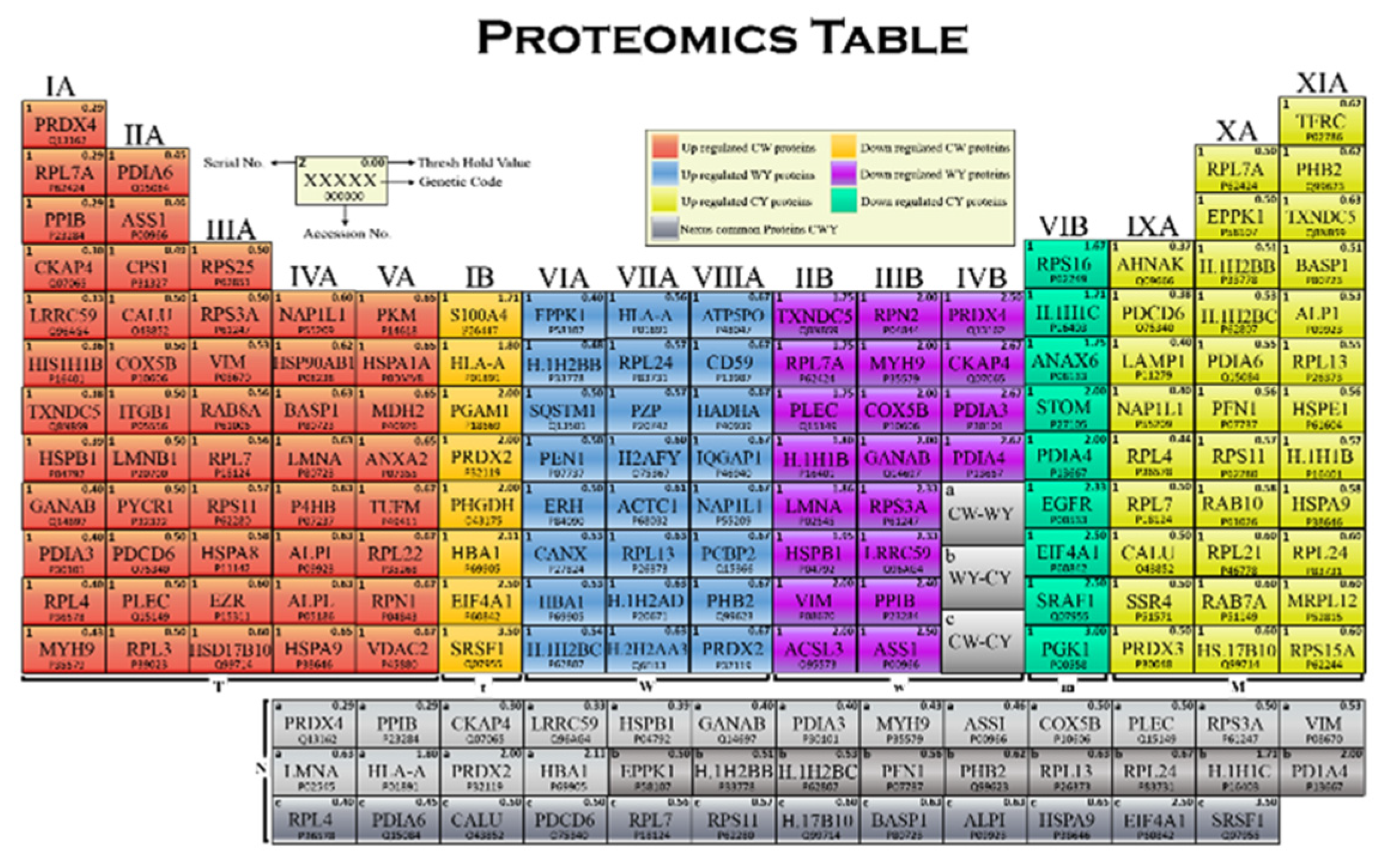
2.3. Proteomics Table
3. Materials and Methods
3.1. Green Synthesis of Iron Carbide NPs
3.2. Functionalization of Fe2C NPs
3.3. Cell Culture and NP Incubation
3.4. Sample Preparation
3.5. Protein Digestion and Peptide Formation
3.6. Cation Exchange Fractionation
3.7. Liquid Chromatography–Mass Spectrometry/Mass Spectrometry Analysis
3.8. Data Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Sample Availability
References
- Serban, K.A.; Pratte, K.A.; Bowler, R.P. Protein Biomarkers for COPD Outcomes. Chest 2021, 159, 2244–2253. [Google Scholar] [CrossRef] [PubMed]
- Hasan, M.; Yang, W.; Ju, Y.; Chu, X.; Wang, Y.; Deng, Y.; Mahmood, N.; Hou, Y. Biocompatibility of iron carbide and detection of metals ions signaling proteomic analysis via HPLC/ESI-Orbitrap. Nano Res. 2017, 10, 1912–1923. [Google Scholar] [CrossRef]
- Kaur, M.; Tiwari, S.; Jain, R. Protein based biomarkers for non-invasive COVID-19 detection. Sens. Bio-Sensing Res. 2020, 29, 100362. [Google Scholar] [CrossRef]
- Ueda, K. A proteome-wide catalog of extracellular vesicles, toward development of cancer liquid biopsy diagnostics. Cancer Sci. 2018, 109, 392. [Google Scholar]
- Hasan, M.; Mustafa, G.; Iqbal, J.; Ashfaq, M.; Mahmood, N. Quantitative proteomic analysis of HeLa cells in response to biocompatible Fe2C@C nanoparticles: 16O/18O-labelling & HPLC-ESI-orbit-trap profiling approach. Toxicol Res. 2018, 7, 84–92. [Google Scholar] [CrossRef]
- Zafar, A.; Jabbar, M.; Manzoor, Y.; Gulzar, H.; Hassan, S.G.; Nazir, M.A.; Ain-ul-Haq; Mustafa, G.; Sahar, R.; Masood, A.; et al. Quantifying Serum Derived Differential Expressed and Low Molecular Weight Protein in Breast Cancer patients. Protein Pept. Lett. 2020, 27, 658–673. [Google Scholar] [CrossRef]
- Iqbal, J.; Li, W.; Hasan, M.; Liu, K.; Awan, U.; Saeed, Y.; Zhang, Y.; Khan, A.M.; Shah, A.; Qing, H.; et al. Differential expression of specific cellular defense proteins in rat hypothalamus under simulated microgravity induced conditions: Comparative proteomics. Proteomics 2014, 14, 1424–1433. [Google Scholar] [CrossRef]
- Lu, C.; Urban, M.W. Stimuli-responsive polymer nano-science: Shape anisotropy, responsiveness, applications. Prog. Polym. Sci. 2018, 78, 24–46. [Google Scholar] [CrossRef]
- García-álvarez, R.; Vallet-Regí, M. Hard and soft protein corona of nanomaterials: Analysis and relevance. Nanomaterials 2021, 11, 888. [Google Scholar] [CrossRef]
- Kopac, T. Protein corona, understanding the nanoparticle–protein interactions and future perspectives: A critical review. Int. J. Biol. Macromol. 2021, 169, 290–301. [Google Scholar] [CrossRef]
- Saleh, D.A.; Shimoni, O.; Sosnik, A. Novel core-corona hybrid nanomaterials based on the conjugation of amphiphilic polymeric diblocks to the surface of multifunctional nanodiamond anchors. Mater. Today Chem. 2017, 3, 15–26. [Google Scholar] [CrossRef]
- Visalakshan, R.M.; García, L.E.G.; Benzigar, M.R.; Ghazaryan, A.; Simon, J.; Mierczynska-Vasilev, A.; Michl, T.D.; Vinu, A.; Mailänder, V.; Morsbach, S.; et al. The Influence of Nanoparticle Shape on Protein Corona Formation. Small. 2020, 16, 2000285. [Google Scholar] [CrossRef] [PubMed]
- Blume, J.E.; Manning, W.C.; Troiano, G.; Hornburg, D.; Figa, M.; Hesterberg, L.; Platt, T.L.; Zhao, X.; Cuaresma, R.A.; Everley, P.A.; et al. Rapid, deep and precise profiling of the plasma proteome with multi-nanoparticle protein corona. Nat. Commun. 2020, 11, 3662. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Xu, M.; Wang, R.; Yin, Y.; Lynch, I.; Liu, S. The Crucial Role of Environmental Coronas in Determining the Biological Effects of Engineered Nanomaterials. Small 2020, 16, 2003691. [Google Scholar] [CrossRef] [PubMed]
- Sebak, A.A.; Gomaa, I.E.O.; Elmeshad, A.N.; Farag, M.H.; Breitinger, U.; Breitinger, H.G.; Abdelkader, M.H. Distinct proteins in protein corona of nanoparticles represent a promising venue for endogenous targeting – part ii: In vitro and in vivo kinetics study. Int. J. Nanomed. 2020, 15, 9539. [Google Scholar] [CrossRef] [PubMed]
- Hasan, M.; Gulzar, H.; Zafar, A.; Haq, A.; Mustafa, G.; Tariq, T.; ul Khalid, A.; Mahmmod, A.; Shu, X.; Mahmood, N. Multiplexing surface anchored functionalized iron carbide nanoparticle: A low molecular weight proteome responsive nano-tracer. Colloids Surf. B Biointerfaces 2021, 203, 111746. [Google Scholar] [CrossRef]
- Albakova, Z.; Siam, M.K.S.; Sacitharan, P.K.; Ziganshin, R.H.; Ryazantsev, D.Y.; Sapozhnikov, A.M. Extracellular heat shock proteins and cancer: New perspectives. Transl. Oncol. 2021, 14, 100995. [Google Scholar] [CrossRef]
- Ignjatovic, V.; Geyer, P.E.; Palaniappan, K.K.; Chaaban, J.E.; Omenn, G.S.; Baker, M.S.; Deutsch, E.W.; Schwenk, J.M. Mass spectrometry-based plasma proteomics: Considerations from sample collection to achieving translational data. bioRxiv 2019. [Google Scholar] [CrossRef]
- Strojan, K.; Leonardi, A.; Bregar, V.B.; Križaj, I.; Svete, J.; Pavlin, M. Dispersion of nanoparticles in different media importantly determines the composition of their protein corona. PLoS ONE 2017, 12, e0169552. [Google Scholar] [CrossRef]
- Breznica, P.; Koliqi, R.; Daka, A. A review of the current understanding of nanoparticles protein corona composition. Med. Pharm. Rep. 2020, 93, 342–350. [Google Scholar] [CrossRef]
- Otun, K.O.; Yao, Y.; Liu, X.; Hildebrandt, D. Synthesis, structure, and performance of carbide phases in Fischer-Tropsch synthesis: A critical review. Fuel 2021, 296, 120689. [Google Scholar] [CrossRef]
- Zhang, Z.; Yin, H.; Yu, G.; He, S.; Kang, J.; Liu, Z.; Cheng, K.; Zhang, Q.; Wang, Y. Selective hydrogenation of CO2 and CO into olefins over Sodium- and Zinc-Promoted iron carbide catalysts. J. Catal. 2021, 395, 350–361. [Google Scholar] [CrossRef]
- Zafar, A.; Tariq, T.; Hasan, M.; Nazar, M.; Rasheed, M.N.; Mahmood, N.; Shu, X. Green-maturation of Cobalt-Oxide nano-sponges for reinforced bacterial apoptosis. Colloids Interface Sci. Commun. 2021, 45, 100531. [Google Scholar] [CrossRef]
- Hasan, M.; Ullah, I.; Zulfiqar, H.; Naeem, K.; Iqbal, A.; Gul, H.; Ashfaq, M.; Mahmood, N. Biological entities as chemical reactors for synthesis of nanomaterials: Progress, challenges and future perspective. Mater. Today Chem. 2018, 8, 13–28. [Google Scholar] [CrossRef]
- Hasan, M.; Zafar, A.; Shahzadi, I.; Luo, F.; Hassan, S.G.; Tariq, T.; Zehra, S.; Munawar, T.; Iqbal, F.; Shu, X. Fractionation of biomolecules in Withania coagulans extract for bioreductive nanoparticle synthesis, antifungal and biofilm activity. Molecules 2020, 25, 3478. [Google Scholar] [CrossRef] [PubMed]
- Saif, M.S.; Zafar, A.; Waqas, M.; Hassan, S.G.; Haq, A.; Tariq, T.; Batool, S.; Dilshad, M.; Hasan, M.; Shu, X. Phyto-reflexive Zinc Oxide Nano-Flowers synthesis: An advanced photocatalytic degradation and infectious therapy. J. Mater. Res. Technol. 2021, 13, 2375–2391. [Google Scholar] [CrossRef]
- Qasim, S.; Zafar, A.; Saif, M.S.; Ali, Z.; Nazar, M.; Waqas, M.; Haq, A.U.; Tariq, T.; Hassan, S.G.; Iqbal, F.; et al. Green synthesis of iron oxide nanorods using Withania coagulans extract improved photocatalytic degradation and antimicrobial activity. J. Photochem. Photobiol. B Biol. 2020, 204, 111784. [Google Scholar] [CrossRef]
- Zarei, M.; Aalaie, J. Profiling of nanoparticle–protein interactions by electrophoresis techniques. Anal. Bioanal. Chem. 2018, 411, 79–96. [Google Scholar] [CrossRef]
- Iqbal, J.; Li, W.; Hasan, M.; Li, Y.J.; Ullah, K.; Yun, W.; Awan, U.; Qing, H.; Deng, Y. Distortion of homeostatic signaling proteins by simulated microgravity in rat hypothalamus: A16O/18O-labeled comparative integrated proteomic approach. Proteomics 2014, 14, 262–273. [Google Scholar] [CrossRef]
- Iqbal, J.; Li, W.; Ullah, K.; Hasan, M.; Linna, G.; Awan, U.; Zhang, Y.; Batool, S.; Qing, H.; Deng, Y. Study of rat hypothalamic proteome by HPLC/ESI ion trap and HPLC/ESI-Q-TOF MS. Proteomics 2013, 13, 2455–2468. [Google Scholar] [CrossRef]
- Tenzer, S.; Docter, D.; Kuharev, J.; Musyanovych, A.; Fetz, V.; Hecht, R.; Schlenk, F.; Fischer, D.; Kiouptsi, K.; Reinhardt, C.; et al. Rapid formation of plasma protein corona critically affects nanoparticle pathophysiology. Nat. Nanotechnol. 2013, 8, 772–781. [Google Scholar] [CrossRef] [PubMed]
- Corchero, J.L.; Villaverde, A. Biomedical applications of distally controlled magnetic nanoparticles. Trends Biotechnol. 2009, 27, 468–476. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.; Zhao, H.; Luo, C.; Lei, T.; Zhang, M. Knockdown of ferritin heavy chain (FTH) inhibits the migration of prostate cancer through reducing S100A4, S100A2, and S100P expression. Transl. Cancer Res. 2020, 9, 5418–5429. [Google Scholar] [CrossRef] [PubMed]
- Cutolo, M.; Gotelli, E.; Montagna, P.; Tardito, S.; Paolino, S.; Pizzorni, C.; Sulli, A.; Smith, V.; Soldano, S. Nintedanib downregulates the transition of cultured systemic sclerosis fibrocytes into myofibroblasts and their pro-fibrotic activity. Arthritis Res. Ther. 2021, 23, 205. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Wang, Y.; Wu, S.; Yan, Y.; Hu, Y.; Zheng, Z.; Li, J.; Wu, W. Sulforaphane-cysteine inhibited migration and invasion via enhancing mitophagosome fusion to lysosome in human glioblastoma cells. Cell Death Dis. 2020, 11, 819. [Google Scholar] [CrossRef] [PubMed]
- Elsaadi, S.; Steiro, I.; Abdollahi, P.; Vandsemb, E.N.; Yang, R.; Slørdahl, T.S.; Rø, T.B.; Menu, E.; Sponaas, A.M.; Børset, M. Targeting phosphoglycerate dehydrogenase in multiple myeloma. Exp. Hematol. Oncol. 2021, 10, 3. [Google Scholar] [CrossRef]
- Li, M.; Wu, C.; Yang, Y.; Zheng, M.; Yu, S.; Wang, J.; Chen, L.; Li, H. 3-Phosphoglycerate dehydrogenase: A potential target for cancer treatment. Cell. Oncol. 2021, 44, 541–556. [Google Scholar] [CrossRef]
- Ali, A.; al Dhahouri, N.; Almesmari, F.S.A.; Fathalla, W.M.; al Jasmi, F. Characterization of etfdh and phgdh mutations in a patient with mild glutaric aciduria type ii and serine deficiency. Genes 2021, 12, 703. [Google Scholar] [CrossRef]
- Zhang, Y.; Yu, H.; Zhang, J.; Gao, H.; Wang, S.; Li, S.; Wei, P.; Liang, J.; Yu, G.; Wang, X.; et al. Cul4A-DDB1–mediated monoubiquitination of phosphoglycerate dehydrogenase promotes colorectal cancer metastasis via increased S-adenosylmethionine. J. Clin. Investig. 2021, 131, e146187. [Google Scholar] [CrossRef]
- Singh, C.; Benos, A.; Grenell, A.; Tran, V.; Hanna, D.; Anand-Apte, B.; Brunengraber, H.; Sears, J.E. The urea cycle is transcriptionally controlled by hypoxia-inducible factors. bioRxiv 2021. [Google Scholar] [CrossRef]
- Zhang, M.; Wang, S.; Sun, L.; Gan, L.; Lin, Y.; Shao, J.; Jiang, H.; Li, M. Ammonia induces changes in carbamoyl phosphate synthetase I and its regulation of glutamine synthesis and urea cycle in yellow catfish Pelteobagrus fulvidraco. Fish Shellfish Immunol. 2021, 120, 242–251. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.; Kanehira, K.; Taniguchi, A. PEGylated TiO2 nanoparticles mediated inhibition of cell migration via integrin beta 1. Sci. Technol. Adv. Mater. 2018, 19, 271–281. [Google Scholar] [CrossRef] [PubMed]
- Sang, S.; Zhang, C.; Shan, J. Pyrroline-5-Carboxylate Reductase 1 Accelerates the Migration and Invasion of Nonsmall Cell Lung Cancer In Vitro. Cancer Biotherapy Radiopharm. 2019, 34, 380–387. [Google Scholar] [CrossRef] [PubMed]
- Cai, F.; Miao, Y.; Liu, C.; Wu, T.; Shen, S.; Su, X.; Shi, Y. Pyrroline-5-carboxylate reductase 1 promotes proliferation and inhibits apoptosis in non-small cell lung cancer. Oncol. Lett. 2017, 15, 731–740. [Google Scholar] [CrossRef] [PubMed]
- Xiao, S.; Li, S.; Yuan, Z.; Zhou, L. Pyrroline-5-carboxylate reductase 1 (PYCR1) upregulation contributes to gastric cancer progression and indicates poor survival outcome. Ann. Transl. Med. 2020, 8, 937. [Google Scholar] [CrossRef]
- Ding, J.; Kuo, M.L.; Su, L.; Xue, L.; Luh, F.; Zhang, H.; Wang, J.; Lin, T.G.; Zhang, K.; Chu, P.; et al. Human mitochondrial pyrroline-5-carboxylate reductase 1 promotes invasiveness and impacts survival in breast cancers. Carcinogenesis 2017, 38, 519–531. [Google Scholar] [CrossRef]
- Takeuchi, I.; Nasukawa, T.; Sugimoto, R.; Takemura-Uchiyama, I.; Murakami, H.; Uchiyama, J. Analyses of propagation processes of Staphylococcus aureus bacteriophages S13′ and S25-3 in two different taxonomies by definitive screening design. Virus Res. 2021, 298, 198406. [Google Scholar] [CrossRef]
- Mohapatra, B.R. Characterization of β-mannanase extracted from a novel Streptomyces species Alg-S25 immobilized on chitosan nanoparticles. Biotechnol. Biotechnol. Equip. 2020, 35, 150–161. [Google Scholar] [CrossRef]
- Solarz, A.; Majcher-Maślanka, I.; Kryst, J.; Chocyk, A. A Search for Biomarkers of Early-life Stress-related Psychopathology: Focus on 70-kDa Heat Shock Proteins. Neuroscience 2021, 463, 238–253. [Google Scholar] [CrossRef]
- Orfanelli, T.; Giannopoulos, S.; Zografos, E.; Athanasiou, A.; Bongiovanni, A.M.; Doulaveris, G.; Moo, T.A.; LaPolla, D.; Bakoyiannis, C.N.; Theodoropoulos, G.E.; et al. Alterations of the 70 kDa heat shock protein (HSP70) and sequestosome-1 (p62) in women with breast cancer. Sci. Rep. 2021, 11, 22220. [Google Scholar] [CrossRef]
- González-Ruiz, L.; González-Moles, M.Á.; González-Ruiz, I.; Ruiz-Ávila, I.; Ayén, Á.; Ramos-García, P. An update on the implications of cyclin D1 in melanomas. Pigment Cell Melanoma Res. 2020, 33, 788–805. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Montalto, F.I.; de Amicis, F. Cyclin D1 in Cancer: A Molecular Connection for Cell Cycle Control, Adhesion and Invasion in Tumor and Stroma. Cells 2020, 9, 2648. [Google Scholar] [CrossRef] [PubMed]
- Maqbool, I.; Ponniresan, V.k.; Govindasamy, K.; Prasad, N.R. Understanding the survival mechanisms of Deinococcus radiodurans against oxidative stress by targeting thioredoxin reductase redox system. Arch Microbiol. 2020, 202, 2355–2366. [Google Scholar] [CrossRef] [PubMed]
- Pourmohammadi, K.; Abedi, E. Enzymatic modifications of gluten protein: Oxidative enzymes. Food Chem. 2021, 356, 129679. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Zhu, Y.; Li, X.; Zhang, X.; Yu, B. Identification of potential biomarkers and metabolic pathways based on integration of metabolomic and transcriptomic data in the development of breast cancer. Arch. Gynecol. Obstet. 2021, 303, 1599–1606. [Google Scholar] [CrossRef]
- Li, M.; Yu, W.; Ke, X.; Ye, P.; Peng, J.; Li, H. Downregulation of rab7 and caveolin-1 increases mmp-2 activity in renal tubular epithelial cells under hypoxic conditions. Open Med. 2021, 16, 1428–1437. [Google Scholar] [CrossRef]
- Xie, M.; Kobayashi, I.; Kiyoshima, T.; Nagata, K.; Ookuma, Y.; Fujiwara, H.; Sakai, H. In situ expression of ribosomal protein L21 in developing tooth germ of the mouse lower first molar. J. Mol. Histol. 2009, 40, 361–367. [Google Scholar] [CrossRef]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hasan, M.; Zafar, A.; Jabbar, M.; Tariq, T.; Manzoor, Y.; Ahmed, M.M.; Hassan, S.G.; Shu, X.; Mahmood, N. Trident Nano-Indexing the Proteomics Table: Next-Version Clustering of Iron Carbide NPs and Protein Corona. Molecules 2022, 27, 5754. https://doi.org/10.3390/molecules27185754
Hasan M, Zafar A, Jabbar M, Tariq T, Manzoor Y, Ahmed MM, Hassan SG, Shu X, Mahmood N. Trident Nano-Indexing the Proteomics Table: Next-Version Clustering of Iron Carbide NPs and Protein Corona. Molecules. 2022; 27(18):5754. https://doi.org/10.3390/molecules27185754
Chicago/Turabian StyleHasan, Murtaza, Ayesha Zafar, Maryum Jabbar, Tuba Tariq, Yasmeen Manzoor, Muhammad Mahmood Ahmed, Shahbaz Gul Hassan, Xugang Shu, and Nasir Mahmood. 2022. "Trident Nano-Indexing the Proteomics Table: Next-Version Clustering of Iron Carbide NPs and Protein Corona" Molecules 27, no. 18: 5754. https://doi.org/10.3390/molecules27185754
APA StyleHasan, M., Zafar, A., Jabbar, M., Tariq, T., Manzoor, Y., Ahmed, M. M., Hassan, S. G., Shu, X., & Mahmood, N. (2022). Trident Nano-Indexing the Proteomics Table: Next-Version Clustering of Iron Carbide NPs and Protein Corona. Molecules, 27(18), 5754. https://doi.org/10.3390/molecules27185754







