Transcriptional Regulation of Ripening in Chili Pepper Fruits (Capsicum spp.)
Abstract
:1. Introduction
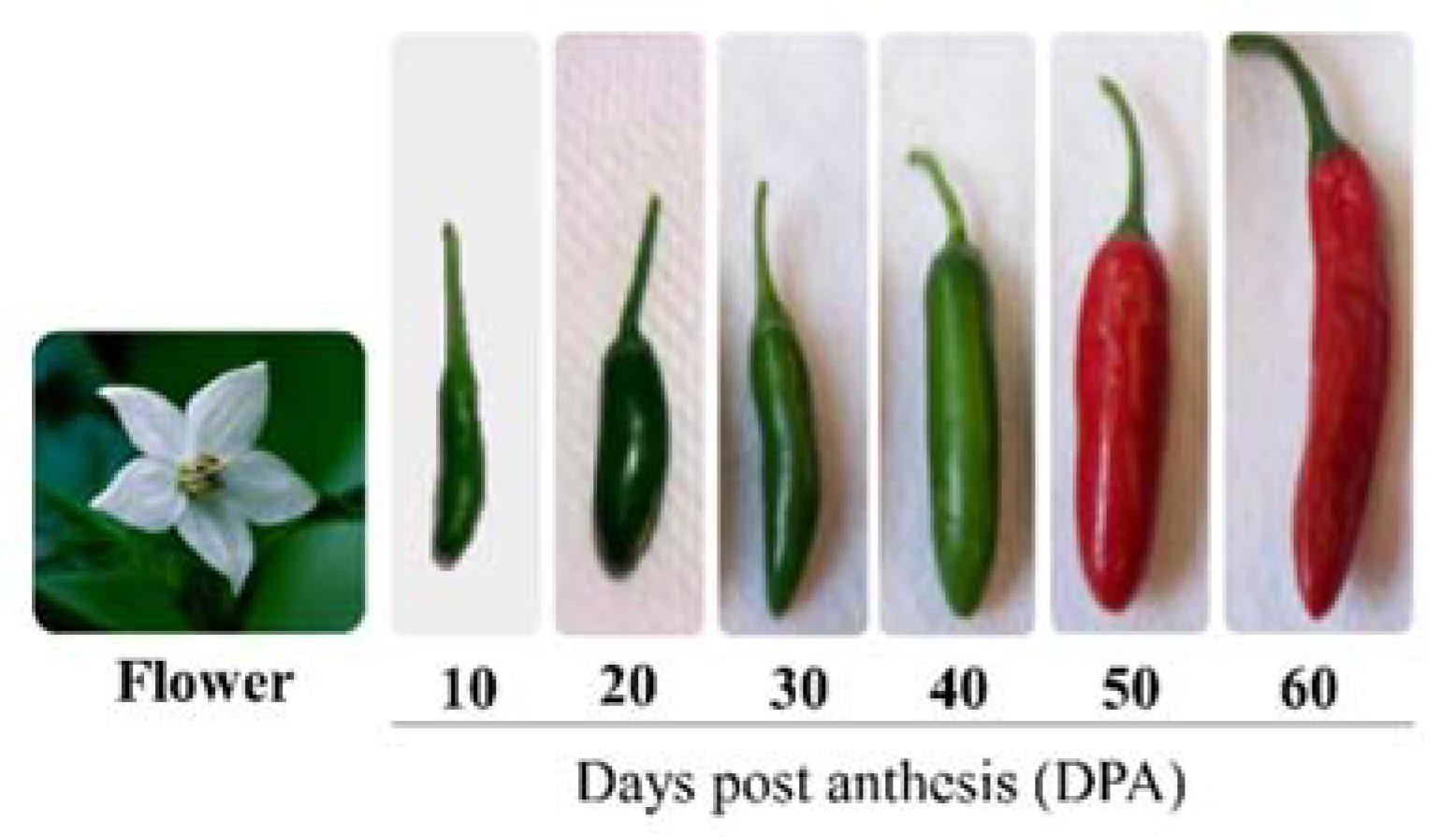
2. Chili Pepper Ripening Regulation Process
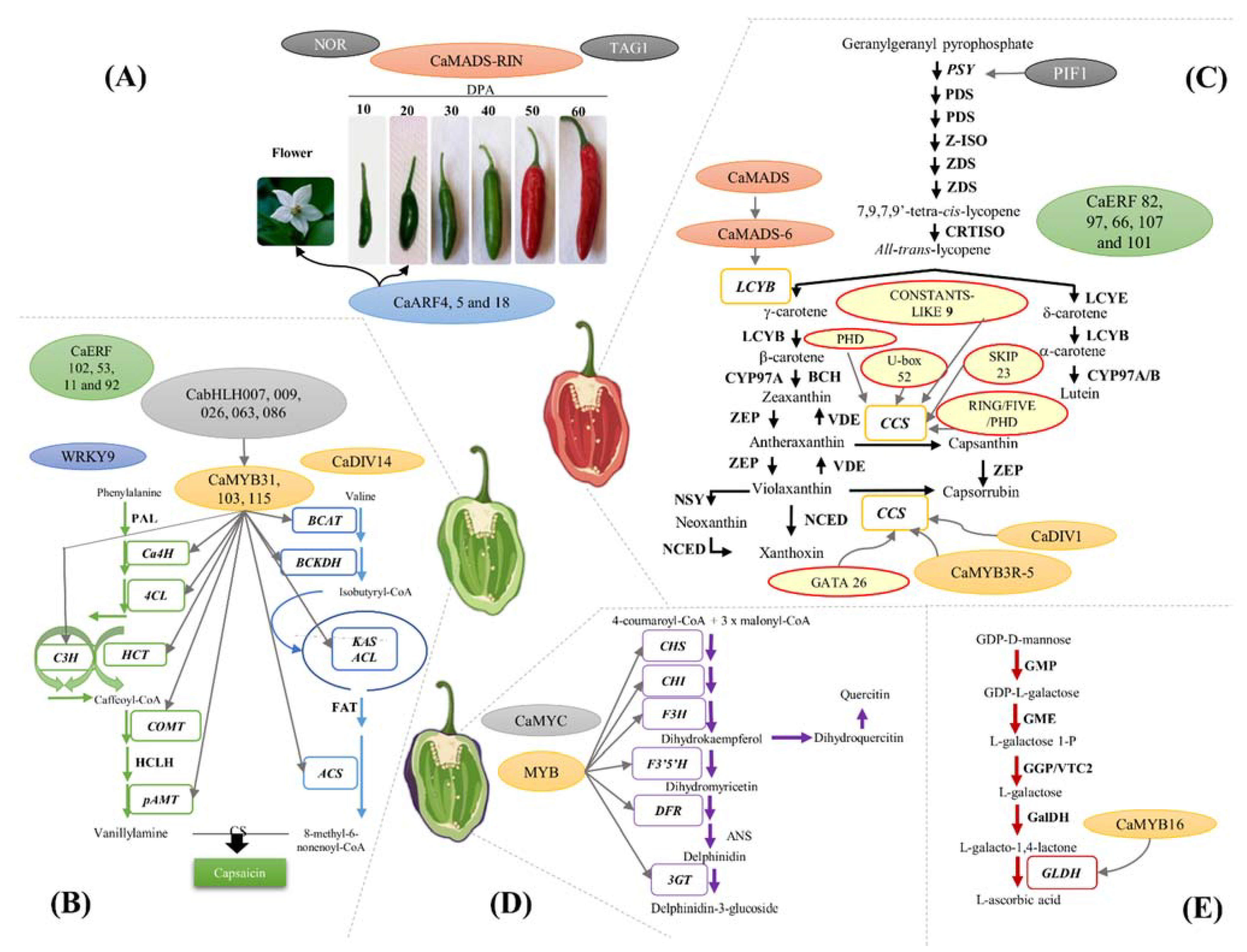
3. Transcriptional Regulation of Capsaicinoids Biosynthesis
4. Transcriptional Regulation of Carotenoid Biosynthesis
5. Transcriptional Regulation of the Anthocyanin Biosynthetic Pathway
6. Transcriptional Regulation of the Ascorbic Acid (AsA) Biosynthetic Pathway
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Carrizo García, C.; Barfuss, M.H.; Sehr, E.M.; Barboza, G.E.; Samuel, R.; Moscone, E.A.; Ehrendorfer, F. Phylogenetic relationships, diversification and expansion of chili peppers (Capsicum, Solanaceae). Ann. Bot. 2016, 118, 35–51. [Google Scholar] [CrossRef] [Green Version]
- Jarret, R.L.; Barboza, G.E.; Costa Batista, F.R.D.; Berke, T.; Chou, Y.-Y.; Hulse-Kemp, A.; Ochoa-Alejo, N.; Tripodi, P.; Veres, A.; Garcia, C.C.; et al. Capsicum—An Abbreviated Compendium. J. Am. Soc. Hortic. Sci. 2019, 144, 3–22. [Google Scholar] [CrossRef] [Green Version]
- Dias, G.B.; Gomes, V.M.; Moraes, T.M.; Zottich, U.P.; Rabelo, G.R.; Carvalho, A.O.; Moulin, M.; Goncalves, L.S.; Rodrigues, R.; Da Cunha, M. Characterization of Capsicum species using anatomical and molecular data. Genet. Mol. Res. 2013, 12, 6488–6501. [Google Scholar] [CrossRef]
- Tripodi, P.; Kumar, S. The Capsicum Crop: An introduction. In The Capsicum Genome; Ramchiary, N., Kole, C., Eds.; Springer International Publishing: Basel, Switzaerland, 2019. [Google Scholar] [CrossRef]
- Chamikara, M.D.M.; Dissanayake, D.R.R.P.; Ishan, M.; Sooriyapathirana, S.D.S.S. Dietary, anticancer and medicinal properties of the phytochemicals in chili pepper (Capsicum spp.). Ceylon J. Sci. 2016, 45, 5–20. [Google Scholar] [CrossRef] [Green Version]
- Batiha, G.E.; Alqahtani, A.; Ojo, O.A.; Shaheen, H.M.; Wasef, L.; Elzeiny, M.; Ismail, M.; Shalaby, M.; Murata, T.; Zaragoza-Bastida, A.; et al. Biological properties, bioactive constituents, and pharmacokinetics of some Capsicum spp. and capsaicinoids. Int. J. Mol. Sci. 2020, 21, 5179. [Google Scholar] [CrossRef] [PubMed]
- Cervantes-Hernandez, F.; Alcala-Gonzalez, P.; Martinez, O.; Ordaz-Ortiz, J.J. Placenta, pericarp, and seeds of tabasco chili pepper fruits show a contrasting diversity of bioactive metabolites. Metabolites 2019, 9, 206. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wahyuni, Y.; Ballester, A.-R.; Sudarmonowati, E.; Bino, R.J.; Bovy, A.G. Secondary metabolites of Capsicum species and their importance in the human diet. J. Nat. Prod. 2013, 76, 783–793. [Google Scholar] [CrossRef]
- Baenas, N.; Belovic, M.; Ilic, N.; Moreno, D.A.; Garcia-Viguera, C. Industrial use of pepper (Capsicum annum L.) derived products: Technological benefits and biological advantages. Food Chem. 2019, 274, 872–885. [Google Scholar] [CrossRef]
- Antonio, A.S.; Wiedemann, L.S.M.; Veiga Junior, V.F. The genus Capsicum: A phytochemical review of bioactive secondary metabolites. RSC Adv. 2018, 8, 25767–25784. [Google Scholar] [CrossRef] [Green Version]
- De Sá Mendes, N.; Branco de Andrade Gonçalves, É.C. The role of bioactive components found in peppers. Trends Food Sci. Technol. 2020, 99, 229–243. [Google Scholar] [CrossRef]
- Giovannoni, J. Molecular biology of fruit maturation and ripening. Annu. Rev. Plant. Physiol. Plant Mol. Biol. 2001, 52, 725–749. [Google Scholar] [CrossRef] [PubMed]
- Giovannoni, J.J. Genetic regulation of fruit development and ripening. Plant Cell 2004, 16, S170–S180. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bouzayen, M.; Latché, A.; Nath, P.; Pech, J.C. Mechanism of fruit ripening. In Plant Developmental Biology—Biotechnological Perspectives: Volume 1; Pua, E.C., Davey, M.R., Eds.; Springer: Berlin/Heidelberg, Germany, 2010; pp. 319–339. [Google Scholar] [CrossRef]
- Paul, V.; Pandey, R.; Srivastava, G.C. The fading distinctions between classical patterns of ripening in climacteric and non-climacteric fruit and the ubiquity of ethylene-An overview. J. Food Sci. Technol. 2012, 49, 1–21. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jang, Y.K.; Jung, E.S.; Lee, H.A.; Choi, D.; Lee, C.H. Metabolomic characterization of hot pepper (Capsicum annuum “CM334”) during fruit development. J. Agric. Food Chem. 2015, 63, 9452–9460. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Hou, X.; Qi, N.; Liu, H.; Li, Y.; Huang, D.; Wang, C.; Liao, W. Insight into ripening-associated transcription factors in tomato: A review. Sci. Hort. 2021, 288, 110363. [Google Scholar] [CrossRef]
- Kim, S.; Park, M.; Yeom, S.I.; Kim, Y.M.; Lee, J.M.; Lee, H.A.; Seo, E.; Choi, J.; Cheong, K.; Kim, K.T.; et al. Genome sequence of the hot pepper provides insights into the evolution of pungency in Capsicum species. Nat. Genet. 2014, 46, 270–278. [Google Scholar] [CrossRef]
- Lee, S.; Chung, E.J.; Joung, Y.H.; Choi, D. Non-climacteric fruit ripening in pepper: Increased transcription of EIL-like genes normally regulated by ethylene. Funct. Integr. Genomics 2010, 10, 135–146. [Google Scholar] [CrossRef]
- Razo-Mendivil, F.G.; Hernandez-Godínez, F.; Hayano-Kanashiro, C.; Martínez, O. Transcriptomic analysis of a wild and a cultivated varieties of Capsicum annuum over fruit development and ripening. PLoS ONE 2021, 16, e0256319. [Google Scholar] [CrossRef]
- Villavicencio, L.; Blankenship, S.M.; Sanders, D.C.; Swallow, W.H. Ethylene and carbon dioxide production in detached fruit of selected pepper cultivars. J. Am. Soc. Hortic. Sci. 1999, 124, 402–406. [Google Scholar] [CrossRef] [Green Version]
- Aizat, W.M.; Able, J.A.; Stangoulis, J.C.R.; Able, A.J. Proteomic analysis during capsicum ripening reveals differential expression of ACC oxidase isoform 4 and other candidates. Funct. Plant Biol. 2013, 40, 1115–1128. [Google Scholar] [CrossRef] [Green Version]
- Aizat, W.M.; Able, J.A.; Stangoulis, J.C.; Able, A.J. Characterisation of ethylene pathway components in non-climacteric Capsicum. BMC Plant Biol. 2013, 13, 191. [Google Scholar] [CrossRef] [Green Version]
- Chen, G.; Alexander, L.; Grierson, D. Constitutive expression of EIL-like transcription factor partially restores ripening in the ethylene-insensitive Nr tomato mutant. J. Exp. Bot. 2004, 55, 1491–1497. [Google Scholar] [CrossRef] [Green Version]
- Dubey, M.; Jaiswal, V.; Rawoof, A.; Kumar, A.; Nitin, M.; Chhapekar, S.S.; Kumar, N.; Ahmad, I.; Islam, K.; Brahma, V.; et al. Identification of genes involved in fruit development/ripening in Capsicum and development of functional markers. Genomics 2019, 111, 1913–1922. [Google Scholar] [CrossRef] [PubMed]
- Vrebalov, J.; Ruezinsky, D.; Padmanabhan, V.; White, R.; Medrano, D.; Drake, R.; Schuch, W.; Giovannoni, J. A MADS-box gene necessary for fruit ripening at the tomato ripening-inhibitor (rin) locus. Science 2002, 296, 343–346. [Google Scholar] [CrossRef] [PubMed]
- Vrebalov, J.; Pan, I.L.; Arroyo, A.J.; McQuinn, R.; Chung, M.; Poole, M.; Rose, J.; Seymour, G.; Grandillo, S.; Giovannoni, J.; et al. Fleshy fruit expansion and ripening are regulated by the Tomato SHATTERPROOF gene TAGL1. Plant Cell 2009, 21, 3041–3062. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Manning, K.; Tör, M.; Poole, M.; Hong, Y.; Thompson, A.J.; King, G.J.; Giovannoni, J.J.; Seymour, G.B. A naturally occurring epigenetic mutation in a gene encoding an SBP-box transcription factor inhibits tomato fruit ripening. Nat. Genet. 2006, 38, 948–952. [Google Scholar] [CrossRef] [PubMed]
- Powell, A.; Nguyen, C.; Hill, T.; Cheng, K.L.; Figueroa-Balderas, R.; Aktas, H.; Ashrafi, H.; Pons, C.; Fernández-Muñoz, R.; Vicente, A.; et al. Uniform ripening encodes a Golden 2-like transcription factor regulating tomato fruit chloroplast development. Science 2012, 336, 1711–1715. [Google Scholar] [CrossRef] [Green Version]
- Lin, Z.; Hong, Y.; Yin, M.; Li, C.; Zhang, K.; Grierson, D. A tomato HD-Zip homeobox protein, LeHB-1, plays an important role in floral organogenesis and ripening. Plant J. 2008, 55, 301–310. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sung, S.-K.; Moon, Y.-H.; Chung, J.-E.; Lee, S.-Y.; Park, H.G.; An, G. Characterization of MADS box genes from hot pepper. Mol. Cells 2001, 11, 352–359. [Google Scholar] [PubMed]
- Ng, M.; Yanofsky, M.F. Function and evolution of the plant MADS-box gene family. Nat. Rev. Genet. 2001, 2, 186–195. [Google Scholar] [CrossRef] [PubMed]
- Mizukami, Y.; Huang, H.; Tudor, M.; Hu, Y.; Ma, H. Functional domains of the floral regulator AGAMOUS: Characterization of the DNA binding domain and analysis of dominant negative mutations. Plant Cell 1996, 8, 831–845. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Riechmann, J.L.; Krizek, B.A.; Meyerowitz, E.M. Dimerization specificity of Arabidopsis MADS domain homeotic proteins APETALA1, APETALA3, PISTILLATA, and AGAMOUS. Proc. Natl. Acad. Sci. USA 1996, 93, 4793–4798. [Google Scholar] [CrossRef] [Green Version]
- Fan, H.-Y.; Hu, Y.; Tudor, M.; Ma, H. Specific interactions between the K domains of AG and AGLs, members of the MADS domain family of DNA binding proteins. Plant J. 1997, 12, 999–1010. [Google Scholar] [CrossRef]
- Yang, Y.; Fanning, L.; Jack, T. The K domain mediates heterodimerization of the Arabidopsis floral organ identity proteins, APETALA3 and PISTILLATA. Plant J. 2003, 33, 47–59. [Google Scholar] [CrossRef] [PubMed]
- Dong, T.; Chen, G.; Tian, S.; Xie, Q.; Yin, W.; Zhang, Y.; Hu, Z. A non-climacteric fruit gene CaMADS-RIN regulates fruit ripening and ethylene biosynthesis in climacteric fruit. PLoS ONE 2014, 9, e95559. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.; Zhan, Y.; Feng, X.; Huang, Z.A.; Sun, C. Identification and expression profiling of the auxin response factors in Capsicum annuum L. under abiotic stress and hormone treatments. Int. J. Mol. Sci. 2017, 18, 2719. [Google Scholar] [CrossRef] [Green Version]
- Liu, R.; Song, J.; Liu, S.; Chen, C.; Zhang, S.; Wang, J.; Xiao, Y.; Cao, B.; Lei, J.; Zhu, Z. Genome-wide identification of the Capsicum bHLH transcription factor family: Discovery of a candidate regulator involved in the regulation of species-specific bioactive metabolites. BMC Plant Biol. 2021, 21, 262. [Google Scholar] [CrossRef]
- Lu, B.-Y.; Cheng, G.-X.; Zhang, Z.; Sun, J.-T.; Ali, M.; Jia, Q.-L.; Luo, D.-X.; Gong, Z.-H.; Li, D.-W. CaMYC, a novel transcription factor, regulates anthocyanin biosynthesis in color-leaved pepper (Capsicum annuum L.). J. Plant Growth Regul. 2018, 38, 574–585. [Google Scholar] [CrossRef]
- Ohme-Takagi, M.; Shinshi, H. Ethylene-inducible DNA binding proteins that interact with an ethylene-responsive element. Plant Cell 1995, 7, 173–182. [Google Scholar] [CrossRef] [Green Version]
- Song, J.; Chen, C.; Zhang, S.; Wang, J.; Huang, Z.; Chen, M.; Cao, B.; Zhu, Z.; Lei, J. Systematic analysis of the Capsicum ERF transcription factor family: Identification of regulatory factors involved in the regulation of species-specific metabolites. BMC Genom. 2020, 21, 573. [Google Scholar] [CrossRef]
- Jin, H.; Martin, C. Multifunctionality and diversity within the plant MYB-gene family. Plant Mol. Biol. 1999, 41, 577–585. [Google Scholar] [CrossRef] [PubMed]
- Arce-Rodriguez, M.L.; Ochoa-Alejo, N. An R2R3-MYB transcription factor regulates capsaicinoid biosynthesis. Plant Physiol. 2017, 174, 1359–1370. [Google Scholar] [CrossRef] [Green Version]
- Sun, B.; Zhu, Z.; Chen, C.; Chen, G.; Cao, B.; Chen, C.; Lei, J. Jasmonate-inducible R2R3-MYB transcription factor regulates capsaicinoid biosynthesis and stamen development in Capsicum. J. Agric. Food Chem. 2019, 67, 10891–10903. [Google Scholar] [CrossRef] [PubMed]
- Sun, B.; Zhou, X.; Chen, C.; Chen, C.; Chen, K.; Chen, M.; Liu, S.; Chen, G.; Cao, B.; Cao, F.; et al. Coexpression network analysis reveals an MYB transcriptional activator involved in capsaicinoid biosynthesis in hot peppers. Hortic. Res. 2020, 7, 162. [Google Scholar] [CrossRef] [PubMed]
- Arce-Rodriguez, M.L.; Martinez, O.; Ochoa-Alejo, N. Genome-wide Identification and analysis of the MYB transcription factor gene family in chili pepper (Capsicum spp.). Int. J. Mol. Sci. 2021, 22, 2229. [Google Scholar] [CrossRef]
- Aguilar-Barragán, A.; Ochoa-Alejo, N. Virus-induced silencing of MYB and WD40 transcription factor genes affects the accumulation of anthocyanins in chilli pepper fruit. Biol. Plant. 2014, 58, 567–574. [Google Scholar] [CrossRef]
- Li, Q.-H.; Yang, S.-P.; Yu, Y.-N.; Khan, A.; Feng, P.-L.; Ali, M.; Shao, D.-K.; Wang, Y.-Y.; Zhang, R.-X.; Gai, W.-X.; et al. Comprehensive transcriptome-based characterization of differentially expressed genes involved in carotenoid biosynthesis of different ripening stages of Capsicum. Sci. Hort. 2021, 288, 110311. [Google Scholar] [CrossRef]
- Ulmasov, T.; Hagen, G.; Guilfoyle, T.J. Activation and repression of transcription by auxin-response factors. Proc. Natl. Acad. Sci. USA 1999, 96, 5844–5849. [Google Scholar] [CrossRef] [Green Version]
- Guilfoyle, T.J.; Hagen, G. Auxin response factors. Curr. Opin. Plant Biol. 2007, 10, 453–560. [Google Scholar] [CrossRef]
- Cheng, Y.; Ahammed, G.J.; Yu, J.; Yao, Z.; Ruan, M.; Ye, Q.; Li, Z.; Wang, R.; Feng, K.; Zhou, G.; et al. Putative WRKYs associated with regulation of fruit ripening revealed by detailed expression analysis of the WRKY gene family in pepper. Sci. Rep. 2016, 6, 39000. [Google Scholar] [CrossRef] [Green Version]
- Rushton, P.J.; Somssich, I.E.; Ringler, P.; Shen, Q.J. WRKY transcription factors. Trends Plant Sci. 2010, 15, 247–258. [Google Scholar] [CrossRef]
- Zuo, J.; Wang, Y.; Zhu, B.; Luo, Y.; Wang, Q.; Gao, L. Network analysis of noncoding RNAs in pepper provides insights into fruit ripening control. Sci. Rep. 2019, 9, 8734. [Google Scholar] [CrossRef]
- Thiele, R.; Mueller-Seitz, E.; Petz, M. Chili pepper fruits: Presumed precursors of fatty acids characteristic for capsaicinoids. J. Agric. Food Chem. 2008, 56, 4219–4224. [Google Scholar] [CrossRef] [PubMed]
- Chapa-Oliver, A.M.; Mejia-Teniente, L. Capsaicin: From plants to a cancer-suppressing agent. Molecules 2016, 21, 931. [Google Scholar] [CrossRef] [Green Version]
- Naves, E.R.; de Avila Silva, L.; Sulpice, R.; Araujo, W.L.; Nunes-Nesi, A.; Peres, L.E.P.; Zsogon, A. Capsaicinoids: Pungency beyond Capsicum. Trends Plant Sci. 2019, 24, 109–120. [Google Scholar] [CrossRef] [PubMed]
- Vázquez-Espinosa, M.; Olguín-Rojas, J.A.; Fayos, O.; González-de-Peredo, A.V.; Espada-Bellido, E.; Ferreiro-González, M.; Barroso, C.G.; Barbero, G.F.; Garcés-Claver, A.; Palma, M. Influence of fruit ripening on the total and individual capsaicinoids and capsiate content in naga jolokia peppers (Capsicum chinense Jacq.). Agronomy 2020, 10, 252. [Google Scholar] [CrossRef] [Green Version]
- Arce-Rodriguez, M.L.; Ochoa-Alejo, N. Biochemistry and molecular biology of capsaicinoid biosynthesis: Recent advances and perspectives. Plant Cell Rep. 2019, 38, 1017–1030. [Google Scholar] [CrossRef] [PubMed]
- Burgos-Valencia, E.; Echevarría-Machado, I.; Narváez-Zapata, J.A.; Martínez-Estévez, M. Gene expression related to the capsaicinoids biosynthesis in the Capsicum genus: Molecular and transcriptomic studies. Braz. J. Bot. 2020, 43, 201–212. [Google Scholar] [CrossRef]
- Tanaka, Y.; Watachi, M.; Nemoto, W.; Goto, T.; Yoshida, Y.; Yasuba, K.I.; Ohno, S.; Doi, M. Capsaicinoid biosynthesis in the pericarp of chili pepper fruits is associated with a placental septum-like transcriptome profile and tissue structure. Plant Cell Rep. 2021, 40, 1859–1874. [Google Scholar] [CrossRef]
- Aza-Gonzalez, C.; Nunez-Palenius, H.G.; Ochoa-Alejo, N. Molecular biology of capsaicinoid biosynthesis in chili pepper (Capsicum spp.). Plant Cell Rep. 2011, 30, 695–706. [Google Scholar] [CrossRef]
- Zhang, Z.X.; Zhao, S.N.; Liu, G.F.; Huang, Z.M.; Cao, Z.M.; Cheng, S.H.; Lin, S.S. Discovery of putative capsaicin biosynthetic genes by RNA-Seq and digital gene expression analysis of pepper. Sci. Rep. 2016, 6, 34121. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Du, H.; Zhang, L.; Liu, L.; Tang, X.F.; Yang, W.J.; Wu, Y.M.; Huang, Y.B.; Tang, Y.X. Biochemical and molecular characterization of plant MYB transcription factor family. Biochemistry 2009, 74, 1–11. [Google Scholar] [CrossRef]
- Ambawat, S.; Sharma, P.; Yadav, N.R.; Yadav, R.C. MYB transcription factor genes as regulators for plant responses: An overview. Physiol. Mol. Biol. Plants 2013, 19, 307–321. [Google Scholar] [CrossRef] [Green Version]
- Han, K.; Jang, S.; Lee, J.H.; Lee, D.G.; Kwon, J.K.; Kang, B.C. A MYB transcription factor is a candidate to control pungency in Capsicum annuum. Theor. Appl. Genet. 2019, 132, 1235–1246. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Sun, B.; Cai, W.; Zhou, X.; Mao, Y.; Chen, C.; Wei, J.; Cao, B.; Chen, C.; Chen, G.; et al. Natural variations in the MYB transcription factor MYB31 determine the evolution of extremely pungent peppers. New Phytol. 2019, 223, 922–938. [Google Scholar] [CrossRef]
- Kondo, F.; Hatakeyama, K.; Sakai, A.; Minami, M.; Nemoto, K.; Matsushima, K. The pungent-variable sweet chili pepper ‘Shishito’ (Capsicum annuum) provides insights regarding the relationship between pungency, the number of seeds, and gene expression involving capsaicinoid biosynthesis. Mol. Genet. Genom. 2021, 296, 591–603. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Liu, Y.; Tang, B.; Dai, X.; Xie, L.; Liu, F.; Zou, X. Genome-wide identification and capsaicinoid biosynthesis-related expression analysis of the R2R3-MYB gene family in Capsicum annuum L. Front. Genet. 2020, 11, 598183. [Google Scholar] [CrossRef] [PubMed]
- Galego, L.; Almeida, J. Role of DIVARICATA in the control of dorsoventral asymmetry in Antirrhinum flowers. Genes Dev. 2002, 16, 880–891. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nakano, T.; Suzuki, K.; Fujimura, T.; Shinshi, H. Genome-wide analysis of the ERF Gene Family in Arabidopsis and rice. Plant Physiol. 2006, 140, 411–432. [Google Scholar] [CrossRef] [Green Version]
- Huang, P.-Y.; Catinot, J.; Zimmerli, L. Ethylene response factors in Arabidopsis immunity. J. Exp. Bot. 2016, 67, 1231–1241. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Keyhaninejad, N.; Curry, J.; Romero, J.; O’Connell, M.A. Fruit specific variability in capsaicinoid accumulation and transcription of structural and regulatory genes in Capsicum fruit. Plant Sci. 2014, 215–216, 59–68. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Toledo-Ortiz, G.; Huq, E.; Quail, P.H. The Arabidopsis Basic/Helix-Loop-Helix transcription factor Family. Plant Cell 2003, 15, 1749–1770. [Google Scholar] [CrossRef] [Green Version]
- Guzman, I.; Bosland, P.W.; O’Connell, M.A. Heat, color, and flavor compounds in Capsicum fruit. In The Biological Activity of Phytochemicals. Recent Advances in Phytochemistry; Springer: New York, NY, USA, 2011; Volume 41, pp. 109–126. [Google Scholar] [CrossRef]
- Rodriguez-Concepcion, M.; Avalos, J.; Bonet, M.L.; Boronat, A.; Gomez-Gomez, L.; Hornero-Mendez, D.; Limon, M.C.; Meléndez-Martínez, A.J.; Olmedilla-Alonso, B.; Palou, A.; et al. A global perspective on carotenoids: Metabolism, biotechnology, and benefits for nutrition and health. Progr. Lipid Res. 2018, 70, 62–93. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rosas-Saavedra, C.; Stange, C. Biosynthesis of carotenoids in plants: Enzymes and color. In Carotenoids in Nature: Biosynthesis, Regulation and Function; Stange, C., Ed.; Springer International Publishing: Cham, Switzerland, 2016; pp. 35–69. [Google Scholar] [CrossRef]
- Villa-Rivera, M.G.; Ochoa-Alejo, N. Chili pepper carotenoids: Nutraceutical properties and mechanisms of action. Molecules 2020, 25, 5573. [Google Scholar] [CrossRef]
- Wahyuni, Y.; Ballester, A.-R.; Sudarmonowati, E.; Bino, R.J.; Bovy, A.G. Metabolite biodiversity in pepper (Capsicum) fruits of thirty-two diverse accessions: Variation in health-related compounds and implications for breeding. Phytochemistry 2011, 72, 1358–1370. [Google Scholar] [CrossRef] [PubMed]
- Mohd Hassan, N.; Yusof, N.A.; Yahaya, A.F.; Mohd Rozali, N.N.; Othman, R. Carotenoids of Capsicum fruits: Pigment profile and health-promoting functional attributes. Antioxidants 2019, 8, 469. [Google Scholar] [CrossRef] [Green Version]
- Brand, A.; Borovsky, Y.; Hill, T.; Rahman, K.A.; Bellalou, A.; Van Deynze, A.; Paran, I. CaGLK2 regulates natural variation of chlorophyll content and fruit color in pepper fruit. Theor. Appl. Genet. 2014, 127, 2139–2148. [Google Scholar] [CrossRef]
- Borovsky, Y.; Monsonego, N.; Mohan, V.; Shabtai, S.; Kamara, I.; Faigenboim, A.; Hill, T.; Chen, S.; Stoffel, K.; Van Deynze, A.; et al. The zinc-finger transcription factor CcLOL1 controls chloroplast development and immature pepper fruit color in Capsicum chinense and its function is conserved in tomato. Plant J. 2019, 99, 41–55. [Google Scholar] [CrossRef]
- Hurtado-Hernandez, H.; Smith, P.G. Inheritance of mature fruit color in Capsicum annuum L. J. Hered. 1985, 76, 211–213. [Google Scholar] [CrossRef]
- Jeong, H.B.; Jang, S.J.; Kang, M.Y.; Kim, S.; Kwon, J.K.; Kang, B.C. Candidate gene analysis reveals that the fruit color locus C1 corresponds to PRR2 in pepper (Capsicum frutescens). Front. Plant Sci. 2020, 11, 399. [Google Scholar] [CrossRef]
- Pan, Y.; Bradley, G.; Pyke, K.; Ball, G.; Lu, C.; Fray, R.; Marshall, A.; Jayasuta, S.; Baxter, C.; van Wijk, R.; et al. Network inference analysis identifies an APRR2-like gene linked to pigment accumulation in tomato and pepper fruits. Plant Physiol. 2013, 161, 1476–1485. [Google Scholar] [CrossRef] [Green Version]
- Makino, S.; Kiba, T.; Imamura, A.; Hanaki, N.; Nakamura, A.; Suzuki, T.; Taniguchi, M.; Ueguchi, C.; Sugiyama, T.; Mizuno, T. Genes encoding pseudo-response regulators: Insight into His-to-Asp phosphorelay and circadian rhythm in Arabidopsis thaliana. Plant Cell Physiol. 2000, 41, 791–803. [Google Scholar] [CrossRef] [Green Version]
- Sun, T.; Yuan, H.; Cao, H.; Yazdani, M.; Tadmor, Y.; Li, L. Carotenoid metabolism in plants: The role of plastids. Mol. Plant. 2018, 11, 58–74. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Camara, B.; Brangeon, J. Carotenoid metabolism during chloroplast to chromoplast transformation in Capsicum annuum fruit. Planta 1981, 151, 359–364. [Google Scholar] [CrossRef] [PubMed]
- Hornero-Méndez, D.; Mínguez-Mosquera, M.I. Xanthophyll esterification accompanying carotenoid overaccumulation in chromoplast of Capsicum annuum ripening fruits is a constitutive process and useful for ripeness index. J. Agric. Food Chem. 2000, 48, 1617–1622. [Google Scholar] [CrossRef] [PubMed]
- Ytterberg, A.J.; Peltier, J.B.; van Wijk, K.J. Protein profiling of plastoglobules in chloroplasts and chromoplasts. A surprising site for differential accumulation of metabolic enzymes. Plant Physiol. 2006, 140, 984–997. [Google Scholar] [CrossRef] [Green Version]
- Siddique, M.A.; Grossmann, J.; Gruissem, W.; Baginsky, S. Proteome analysis of bell pepper (Capsicum annuum L.) Chromoplasts. Plant Cell Physiol. 2006, 47, 1663–1673. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.Q.; Yang, Y.; Fei, Z.; Yuan, H.; Fish, T.; Thannhauser, T.W.; Mazourek, M.; Kochian, L.V.; Wang, X.; Li, L. Proteomic analysis of chromoplasts from six crop species reveals insights into chromoplast function and development. J. Exp. Bot. 2013, 64, 949–961. [Google Scholar] [CrossRef] [Green Version]
- Gómez-García, M.D.R.; Ochoa-Alejo, N. Biochemistry and molecular biology of carotenoid biosynthesis in chili peppers (Capsicum spp.). Int. J. Mol. Sci. 2013, 14, 19025–19053. [Google Scholar] [CrossRef] [Green Version]
- Sathasivam, R.; Radhakrishnan, R.; Kim, J.K.; Park, S.U. An update on biosynthesis and regulation of carotenoids in plants. S. Afr. J. Bot. 2021, 140, 290–302. [Google Scholar] [CrossRef]
- Martínez-López, L.A.; Ochoa-Alejo, N.; Martinez, O. Dynamics of the chili pepper transcriptome during fruit development. BMC Genom. 2014, 15, 143. [Google Scholar] [CrossRef] [Green Version]
- Camara, B.; Bardat, F.; Moneger, R. Sites of biosynthesis of carotenoids in Capsicum chromoplasts. Eur. J. Biochem. 1982, 127, 255–258. [Google Scholar] [CrossRef] [PubMed]
- Cazzonelli, C.I.; Pogson, B.J. Source to sink: Regulation of carotenoid biosynthesis in plants. Trends Plant Sci. 2010, 15, 266–274. [Google Scholar] [CrossRef]
- Stanley, L.; Yuan, Y.W. Transcriptional regulation of carotenoid biosynthesis in plants: So many regulators, so little consensus. Front. Plant Sci. 2019, 10, 1017. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Toledo-Ortiz, G.; Huq, E.; Rodriguez-Concepcion, M. Direct regulation of phytoene synthase gene expression and carotenoid biosynthesis by phytochrome-interacting factors. Proc. Natl. Acad. Sci. USA 2010, 107, 11626–11631. [Google Scholar] [CrossRef] [Green Version]
- Huq, E.; Al-Sady, B.; Hudson, M.; Kim, C.; Apel, K.; Quail, P.H. Phytochrome-interacting factor 1 is a critical bHLH regulator of chlorophyll biosynthesis. Science 2004, 305, 1937–1941. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bou-Torrent, J.; Toledo-Ortiz, G.; Ortiz-Alcaide, M.; Cifuentes-Esquivel, N.; Halliday, K.J.; Martinez-Garcia, J.F.; Rodriguez-Concepcion, M. Regulation of carotenoid biosynthesis by shade relies on specific subsets of antagonistic transcription factors and cofactors. Plant Physiol. 2015, 169, 1584–1594. [Google Scholar] [CrossRef] [Green Version]
- Grassi, S.; Piro, G.; Lee, J.M.; Zheng, Y.; Fei, Z.; Dalessandro, G.; Giovannoni, J.J.; Lenucci, M.S. Comparative genomics reveals candidate carotenoid pathway regulators of ripening watermelon fruit. BMC Genom. 2013, 14, 781. [Google Scholar] [CrossRef] [Green Version]
- Hou, B.Z.; Li, C.L.; Han, Y.Y.; Shen, Y.Y. Characterization of the hot pepper (Capsicum frutescens) fruit ripening regulated by ethylene and ABA. BMC Plant Biol. 2018, 18, 162. [Google Scholar] [CrossRef]
- Zhu, F.; Luo, T.; Liu, C.; Wang, Y.; Yang, H.; Yang, W.; Zheng, L.; Xiao, X.; Zhang, M.; Xu, R.; et al. An R2R3-MYB transcription factor represses the transformation of alpha- and beta-branch carotenoids by negatively regulating expression of CrBCH2 and CrNCED5 in flavedo of Citrus reticulate. New Phytol. 2017, 216, 178–192. [Google Scholar] [CrossRef] [Green Version]
- Lu, S.; Zhang, Y.; Zhu, K.; Yang, W.; Ye, J.; Chai, L.; Xu, Q.; Deng, X. The citrus transcription factor CsMADS6 modulates carotenoid metabolism by directly regulating carotenogenic genes. Plant Physiol. 2018, 176, 2657–2676. [Google Scholar] [CrossRef] [Green Version]
- Aza-González, C.; Núñez-Palenius, H.; Ochoa-Alejo, N. Molecular biology of chili pepper anthocyanin biosynthesis. Rev. Soc. Quím. Méx. 2012, 56, 93–98. [Google Scholar] [CrossRef]
- Khoo, H.E.; Azlan, A.; Tang, S.T.; Lim, S.M. Anthocyanidins and anthocyanins: Colored pigments as food, pharmaceutical ingredients, and the potential health benefits. Food Nutr. Res. 2017, 61, 1361779. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ramsay, N.A.; Glover, B.J. MYB–bHLH–WD40 protein complex and the evolution of cellular diversity. Trends Plant Sci. 2005, 10, 63–70. [Google Scholar] [CrossRef] [PubMed]
- Stommel, J.R.; Lightbourn, G.J.; Winkel, B.S.; Griesbach, R.J. Transcription factor families regulate the anthocyanin biosynthetic pathway in Capsicum annuum. J. Am. Soc. Hortic. Sci. 2009, 134, 244–251. [Google Scholar] [CrossRef] [Green Version]
- Smith, T.F.; Gaitatzes, C.; Saxena, K.; Neer, E.J. The WD repeat: A common architecture for diverse functions. Trends Biochem. Sci. 1999, 24, 181–185. [Google Scholar] [CrossRef]
- Howard, L.R.; Smith, R.T.; Wagner, A.B.; Villalon, B.; Burns, E.E. Provitamin A and ascorbic acid content of fresh pepper cultivars (Capsicum annuum) and processed jalapeños. J. Food Sci. 1994, 59, 362–365. [Google Scholar] [CrossRef]
- Campos, M.R.S.; Gómez, K.R.; Ordoñez, Y.M.; Ancona, D.B. Polyphenols, ascorbic acid and carotenoids contents and antioxidant properties of habanero pepper (Capsicum chinense) fruit. Food Nutr. Sci. 2013, 04, 47–54. [Google Scholar] [CrossRef] [Green Version]
- Njus, D.; Kelley, P.M.; Tu, Y.-J.; Schlegel, H.B. Ascorbic acid: The chemistry underlying its antioxidant properties. Free Radic. Biol. Med. 2020, 159, 37–43. [Google Scholar] [CrossRef]
- Smirnoff, N. Vitamin C. In Biosynthesis of Vitamins in Plants Part B; Rébeillé, F., Douce, R., Eds.; Academic Press: Cambridge, MA, USA, 2011; Volume 59, pp. 107–177. [Google Scholar] [CrossRef]
- Mousavi, S.; Bereswill, S.; Heimesaat, M.M. Immunomodulatory and Antimicrobial Effects of Vitamin C. Eur. J. Microbiol. Immunol. 2019, 9, 73–79. [Google Scholar] [CrossRef]
- Chaitanya, N.C.; Muthukrishnan, A.; Krishnaprasad, C.M.S.; Sanjuprasanna, G.; Pillay, P.; Mounika, B. An insight and update on the analgesic properties of vitamin C. J. Pharm. Bioallied Sci. 2018, 10, 119–125. [Google Scholar] [CrossRef]
- Colunga Biancatelli, R.M.L.; Berrill, M.; Catravas, J.D.; Marik, P.E. Quercetin and vitamin C: An experimental, synergistic therapy for the prevention and treatment of SARS-CoV-2 related disease (COVID-19). Front. Immunol. 2020, 11, 1451. [Google Scholar] [CrossRef]
- Vissers, M.C.M.; Das, A.B. Potential mechanisms of action for vitamin C in cancer: Reviewing the evidence. Front. Physiol. 2018, 9, 809. [Google Scholar] [CrossRef] [Green Version]
- Khadi, B.; Goud, J.V.; Patil, V. Variation in ascorbic acid and mineral content in fruits of some varieties of chilli (Capsicum annuum L.). Plant Foods Hum. Nutr. 1987, 37, 9–15. [Google Scholar] [CrossRef]
- Kumar, O.; Tata, S. Ascorbic acid contents in chili peppers (Capsicum L.). Not. Sci. Biol. 2009, 1, 50–52. [Google Scholar] [CrossRef] [Green Version]
- Alos, E.; Rodrigo, M.J.; Zacarias, L. Transcriptomic analysis of genes involved in the biosynthesis, recycling and degradation of L-ascorbic acid in pepper fruits (Capsicum annuum L.). Plant Sci. 2013, 207, 2–11. [Google Scholar] [CrossRef]
- Gómez-García, M.D.R.; Ochoa-Alejo, N. Predominant role of the L-galactose pathway in L-ascorbic acid biosynthesis in fruits and leaves of the Capsicum annuum L. chili pepper. Braz. J. Bot. 2015, 39, 157–168. [Google Scholar] [CrossRef]
- Smirnoff, N. Ascorbic acid metabolism and functions: A comparison of plants and mammals. Free Radic. Biol. Med. 2018, 122, 116–129. [Google Scholar] [CrossRef]
- Yahia, E.M.; Contreras-Padilla, M.; Gonazalez-Aguilar, G. Ascorbic acid content in relation to ascorbic acid oxidase activity and polyamine content in tomato and bell pepper fruits during development, maturation and senescence. LWT-Food Sci. Technol. 2001, 34, 452–457. [Google Scholar] [CrossRef] [Green Version]
- Barata-Soares, A.D.; Gomez, M.L.P.A.; Mesquita, C.H.D.; Lajolo, F.M. Ascorbic acid biosynthesis: A precursor study on plants. Braz. J. Plant Physiol. 2004, 16, 147–154. [Google Scholar] [CrossRef]
- Chiaiese, P.; Corrado, G.; Minutolo, M.; Barone, A.; Errico, A. Transcriptional regulation of ascorbic acid during fruit ripening in pepper (Capsicum annuum) varieties with low and high antioxidants content. Plants 2019, 8, 206. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, F.; Hu, Y.; Vannozzi, A.; Wu, K.; Cai, H.; Qin, Y.; Mullis, A.; Lin, Z.; Zhang, L. The WRKY transcription factor family in model plants and crops. Crit. Rev. Plant Sci. 2017, 36, 311–335. [Google Scholar] [CrossRef]
- Wen, X.; Wang, J.; Zhang, D.; Ding, Y.; Ji, X.; Tan, Z.; Wang, Y. Reverse Chromatin Immunoprecipitation (R-ChIP) enables investigation of the upstream regulators of plant genes. Commun. Biol. 2020, 3, 770. [Google Scholar] [CrossRef] [PubMed]
| Transcription Factor | Consensus Motif | Gene | Function during Ripening | Reference |
|---|---|---|---|---|
| ARF (Auxin-response factor) | TGTCTC TGTCCC TGTCAC | CaARF4, CaARF5, CaARF18 | Flower and fruit development | [38] |
| bHLH (basic Helix-Loop-Helix) | E-box CANNTG | CabHLH007, CabHLH009, CabHLH026, CabHLH063 CabHLH086 | Capsaicinoid biosynthesis regulation | [39] |
| CabHLH009 CabHLH032 CabHLH048 CabHLH095 CabHLH100 | Carotenoid accumulation | |||
| CaMYC | Anthocyanin biosynthesis | [40] | ||
| ERF | GCC box TAAGAGCCGCC | CaERF102, CaERF53, CaERF111, CaERF92 | Capsaicinoid biosynthesis | [41,42] |
| CaERF82, CaERF97, CaERF66, CaERF107, CaERF101 | Carotenoid biosynthesis | [42] | ||
| MADS box | CArG | CaMADS CaMADS6 CaMADS-RIN | Ethylene signaling | [25,31,37] |
| MYB | TAACTAAC | CaMYB31 CaMYB48 CaMYB108 CaMYB103 CaMYB115 CaDIV14 | Capsaicinoid biosynthesis and accumulation | [43,44,45,46,47] |
| CaMYB3R-5 CaDIV1 | Carotenoid biosynthesis Putative regulator of the CCS gene | [47] | ||
| MYB | Anthocyanin biosynthesis | [48] | ||
| CaMYB16 | Ascorbic acid biosynthesis | [47] | ||
| F-box protein SKIP23 | Carotenoid biosynthesis. Putative regulators of the CCS gene | [49] | ||
| GATA transcription factor 26 | ||||
| U-box domain-containing protein 52 | ||||
| zinc finger family FYVE/PHD-type | ||||
| RING/FYVE/PHD-type | ||||
| CONSTANS-LIKE 9 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Villa-Rivera, M.G.; Ochoa-Alejo, N. Transcriptional Regulation of Ripening in Chili Pepper Fruits (Capsicum spp.). Int. J. Mol. Sci. 2021, 22, 12151. https://doi.org/10.3390/ijms222212151
Villa-Rivera MG, Ochoa-Alejo N. Transcriptional Regulation of Ripening in Chili Pepper Fruits (Capsicum spp.). International Journal of Molecular Sciences. 2021; 22(22):12151. https://doi.org/10.3390/ijms222212151
Chicago/Turabian StyleVilla-Rivera, Maria Guadalupe, and Neftalí Ochoa-Alejo. 2021. "Transcriptional Regulation of Ripening in Chili Pepper Fruits (Capsicum spp.)" International Journal of Molecular Sciences 22, no. 22: 12151. https://doi.org/10.3390/ijms222212151
APA StyleVilla-Rivera, M. G., & Ochoa-Alejo, N. (2021). Transcriptional Regulation of Ripening in Chili Pepper Fruits (Capsicum spp.). International Journal of Molecular Sciences, 22(22), 12151. https://doi.org/10.3390/ijms222212151