An Extensive Assessment of Network Embedding in PPI Network Alignment
Abstract
:1. Introduction
2. Related Surveys and Differences
3. Network Alignment
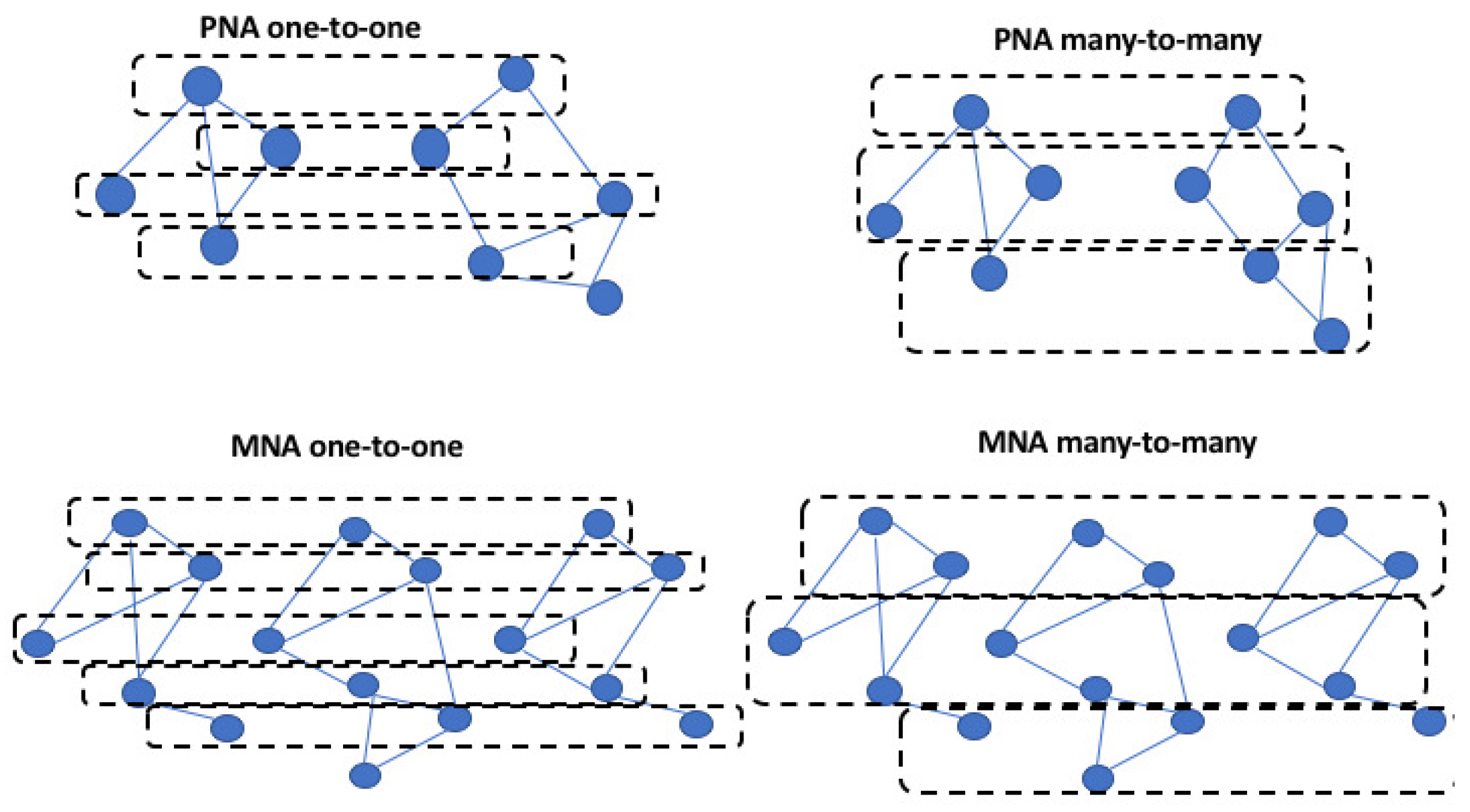
3.1. Global or Local Alignment
3.2. Pairwise or Multiple Alignment
3.3. Quality Evaluation of Network Alignments
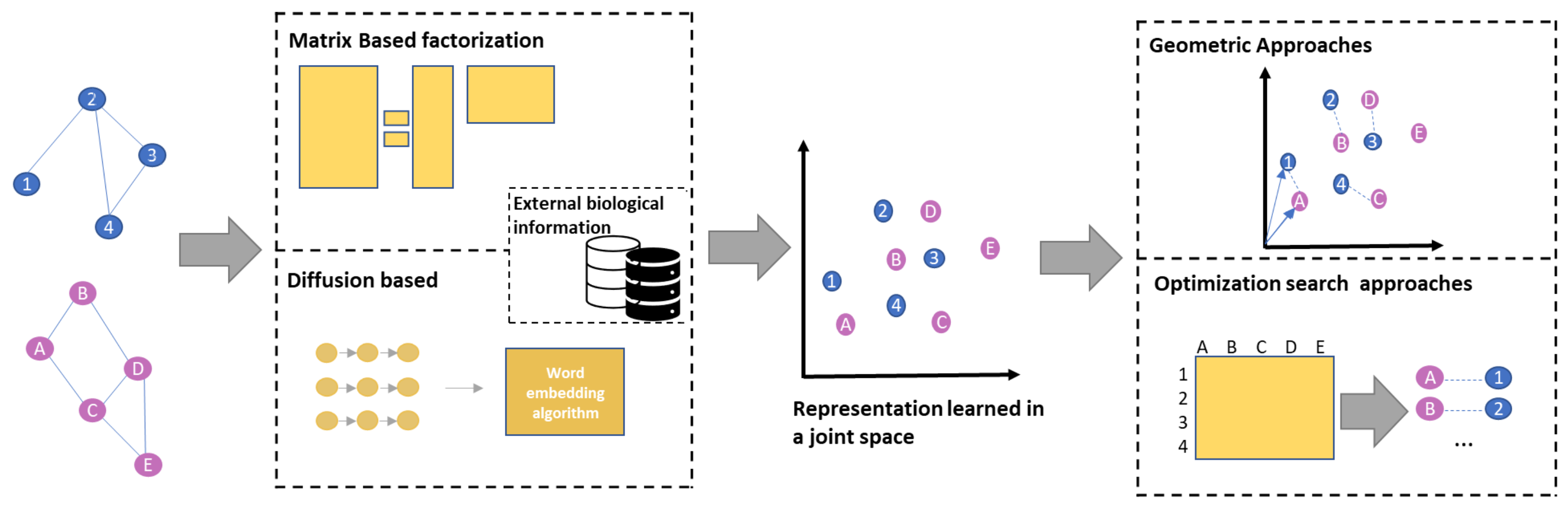
4. Representation Learning
Network Embedding for Network Alignment
5. Discussion
Qualitative Comparison of NE-NA
6. Conclusions and Future Works
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Barabási, A.L. Network science. Philos. Trans. R. Soc. Math. Phys. Eng. Sci. 2013, 371, 20120375. [Google Scholar] [CrossRef] [PubMed]
- Milano, M.; Guzzi, P.H.; Cannataro, M. Glalign: A novel algorithm for local network alignment. IEEE/ACM Trans. Comput. Biol. Bioinform. 2018, 16, 1958–1969. [Google Scholar] [CrossRef] [PubMed]
- Milano, M.; Hayes, W.; Veltri, P.; Cannataro, M.; Guzzi, P.H. SL-GLAlign: Improving local alignment of biological networks through simulated annealing. Netw. Model. Anal. Health Informat. Bioinform. 2020, 9, 10. [Google Scholar] [CrossRef]
- Meng, L.; Striegel, A.; Milenković, T. Local versus global biological network alignment. Bioinformatics 2016, 32, 3155–3164. [Google Scholar] [CrossRef] [PubMed]
- Bengio, Y.; Courville, A.; Vincent, P. Representation learning: A review and new perspectives. IEEE Trans. Pattern Anal. Mach. Intell. 2013, 35, 1798–1828. [Google Scholar] [CrossRef]
- Heimann, M.; Koutra, D. On generalizing neural node embedding methods to multi-network problems. In Proceedings of the KDD MLG Workshop, Halifax, NS, Canada, 14 August 2017. [Google Scholar]
- Nelson, W.; Zitnik, M.; Wang, B.; Leskovec, J.; Goldenberg, A.; Sharan, R. To embed or not: Network embedding as a paradigm in computational biology. Front. Genet. 2019, 10, 381. [Google Scholar] [CrossRef] [Green Version]
- Moyano, L. Learning network representations. Eur. Phys. J. Spec. Top. 2017, 226, 499–518. [Google Scholar] [CrossRef]
- Hamilton, W.L.; Ying, R.; Leskovec, J. Representation learning on graphs: Methods and applications. arXiv 2017, arXiv:1709.05584. [Google Scholar]
- Zhang, D.; Yin, J.; Zhu, X.; Zhang, C. Network representation learning: A survey. IEEE Trans. Big Data 2018, 6, 3–28. [Google Scholar] [CrossRef] [Green Version]
- Chen, H.; Perozzi, B.; Al-Rfou, R.; Skiena, S. A tutorial on network embeddings. arXiv 2018, arXiv:1808.02590. [Google Scholar]
- Goyal, P.; Ferrara, E. Graph embedding techniques, applications, and performance: A survey. Knowl. Based Syst. 2018, 151, 78–94. [Google Scholar] [CrossRef] [Green Version]
- Cai, H.; Zheng, V.W.; Chang, K.C.C. A comprehensive survey of graph embedding: Problems, techniques, and applications. IEEE Trans. Knowl. Data Eng. 2018, 30, 1616–1637. [Google Scholar] [CrossRef] [Green Version]
- Yue, X.; Wang, Z.; Huang, J.; Parthasarathy, S.; Moosavinasab, S.; Huang, Y.; Lin, S.; Zhang, W.; Zhang, P.; Sun, H. Graph Embedding on Biomedical Networks: Methods, Applications, and Evaluations. Bioinformatics 2019, 36, 1241–1251. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kobler, J.; Schöning, U.; Torán, J. The Graph Isomorphism Problem: Its Structural Complexity; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2012. [Google Scholar]
- Saraph, V.; Milenković, T. MAGNA: Maximizing accuracy in global network alignment. Bioinformatics 2014, 30, 2931–2940. [Google Scholar] [CrossRef] [Green Version]
- Liao, C.S.; Lu, K.; Baym, M.; Singh, R.; Berger, B. IsoRankN: Spectral methods for global alignment of multiple protein networks. Bioinformatics 2009, 25, i253–i258. [Google Scholar] [CrossRef] [Green Version]
- Kuchaiev, O.; Milenković, T.; Memišević, V.; Hayes, W.; Pržulj, N. Topological network alignment uncovers biological function and phylogeny. J. R. Soc. Interface 2010, 7, 1341–1354. [Google Scholar] [CrossRef] [Green Version]
- Milenković, T.; Ng, W.L.; Hayes, W.; Pržulj, N. Optimal network alignment with graphlet degree vectors. Cancer Informat. 2010, 9, CIN-S4744. [Google Scholar] [CrossRef]
- Kuchaiev, O.; Pržulj, N. Integrative network alignment reveals large regions of global network similarity in yeast and human. Bioinformatics 2011, 27, 1390–1396. [Google Scholar] [CrossRef]
- Memišević, V.; Pržulj, N. C-GRAAL: Common-neighbors-based global GRA ph AL ignment of biological networks. Integr. Biol. 2012, 4, 734–743. [Google Scholar] [CrossRef]
- Malod-Dognin, N.; Pržulj, N. L-GRAAL: Lagrangian graphlet-based network aligner. Bioinformatics 2015, 31, 2182–2189. [Google Scholar] [CrossRef] [Green Version]
- Pržulj, N. Biological network comparison using graphlet degree distribution. Bioinformatics 2007, 23, e177–e183. [Google Scholar] [CrossRef] [PubMed]
- Patro, R.; Kingsford, C. Global network alignment using multiscale spectral signatures. Bioinformatics 2012, 28, 3105–3114. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, Y.; Crawford, J.; Tang, J.; Milenković, T. Simultaneous optimization of both node and edge conservation in network alignment via WAVE. In Proceedings of the International Workshop on Algorithms in Bioinformatics, Atlanta, GA, USA, 10–12 September 2015; Springer: Berlin/Heidelberg, Germany, 2015; pp. 16–39. [Google Scholar]
- Vijayan, V.; Saraph, V.; Milenković, T. MAGNA++: Maximizing Accuracy in Global Network Alignment via both node and edge conservation. Bioinformatics 2015, 31, 2409–2411. [Google Scholar] [CrossRef] [PubMed]
- Mamano, N.; Hayes, W. SANA: Simulated Annealing Network Alignment Applied to Biological Networks. arXiv 2016, arXiv:1607.02642. [Google Scholar]
- Malod-Dognin, N.; Ban, K.; Pržulj, N. Unified Alignment of Protein-Protein Interaction Networks. Sci. Rep. 2017, 7, 953. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meng, L.; Striegel, A.; Milenkovic, T. IGLOO: Integrating global and local biological network alignment. arXiv 2016, arXiv:1604.06111. [Google Scholar]
- Sharan, R.; Ideker, T. Modeling cellular machinery through biological network comparison. Nat. Biotechnol. 2006, 24, 427–433. [Google Scholar] [CrossRef]
- Pache, R.A.; Ceol, A.; Aloy, P. NetAligner, a network alignment server to compare complexes, pathways and whole interactomes. Nucleic Acids Res. 2012, 40, W157–W161. [Google Scholar] [CrossRef]
- Ciriello, G.; Mina, M.; Guzzi, P.H.; Cannataro, M.; Guerra, C. AlignNemo: A local network alignment method to integrate homology and topology. PLoS ONE 2012, 7, e38107. [Google Scholar] [CrossRef]
- Mina, M.; Guzzi, P.H. Improving the robustness of local network alignment: Design and extensive assessmentof a markov clustering-based approach. Comput. Biol. Bioinformat. IEEE/ACM Trans. 2014, 11, 561–572. [Google Scholar] [CrossRef]
- Enright, A.J.; Van Dongen, S.; Ouzounis, C.A. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 2002, 30, 1575–1584. [Google Scholar] [CrossRef] [PubMed]
- Stelzl, U.; Worm, U.; Lalowski, M.; Haenig, C.; Brembeck, F.H.; Goehler, H.; Stroedicke, M.; Zenkner, M.; Schoenherr, A.; Koeppen, S.; et al. A human protein-protein interaction network: A resource for annotating the proteome. Cell 2005, 122, 957–968. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vijayan, V.; Milenković, T. Multiple network alignment via multiMAGNA++. IEEE/ACM Trans. Comput. Biol. Bioinform. 2017, 15, 1669–1682. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ibragimov, R.; Malek, M.; Baumbach, J.; Guo, J. Multiple graph edit distance: Simultaneous topological alignment of multiple protein-protein interaction networks with an evolutionary algorithm. In Proceedings of the 2014 Annual Conference on Genetic and Evolutionary Computation, Vancouver, BC, Canada, 12–16 July 2014; pp. 277–284. [Google Scholar]
- Hu, J.; Reinert, K. LocalAli: An evolutionary-based local alignment approach to identify functionally conserved modules in multiple networks. Bioinformatics 2014, 31, 363–372. [Google Scholar] [CrossRef]
- Sahraeian, S.M.E.; Yoon, B.J. SMETANA: Accurate and scalable algorithm for probabilistic alignment of large-scale biological networks. PLoS ONE 2013, 8, e67995. [Google Scholar] [CrossRef]
- Gligorijević, V.; Malod-Dognin, N.; Pržulj, N. Fuse: Multiple network alignment via data fusion. Bioinformatics 2016, 32, 1195–1203. [Google Scholar] [CrossRef] [Green Version]
- Hu, J.; Kehr, B.; Reinert, K. NetCoffee: A fast and accurate global alignment approach to identify functionally conserved proteins in multiple networks. Bioinformatics 2014, 30, 540–548. [Google Scholar] [CrossRef] [Green Version]
- Alkan, F.; Erten, C. BEAMS: Backbone extraction and merge strategy for the global many-to-many alignment of multiple PPI networks. Bioinformatics 2014, 30, 531–539. [Google Scholar] [CrossRef] [Green Version]
- Vijayan, V.; Gu, S.; Krebs, E.T.; Meng, L.; Milenkovic, T. Pairwise Versus Multiple Global Network Alignment. IEEE Access 2020, 8, 41961–41974. [Google Scholar] [CrossRef]
- Mikolov, T.; Sutskever, I.; Chen, K.; Corrado, G.S.; Dean, J. Distributed representations of words and phrases and their compositionality. In Proceedings of the Advances in Neural Information Processing Systems, Lake Tahoe, NV, USA, 5–8 December 2013; 2013; pp. 3111–3119. [Google Scholar]
- Perozzi, B.; Al-Rfou, R.; Skiena, S. Deepwalk: Online learning of social representations. In Proceedings of the 20th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, New York, NY, USA, 24–27 August 2014; pp. 701–710. [Google Scholar]
- Grover, A.; Leskovec, J. node2vec: Scalable feature learning for networks. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 855–864. [Google Scholar]
- Chu, X.; Fan, X.; Yao, D.; Zhu, Z.; Huang, J.; Bi, J. Cross-Network Embedding for Multi-Network Alignment. In Proceedings of the The World Wide Web Conference, WWW ’19, San Francisco, CA, USA, 13 May 2019; Association for Computing Machinery: New York, NY, USA, 2019; pp. 273–284. [Google Scholar]
- Derr, T.; Karimi, H.; Liu, X.; Xu, J.; Tang, J. Deep Adversarial Network Alignment. arXiv 2019, arXiv:1902.10307. [Google Scholar]
- Chen, C.; Xie, W.; Xu, T.; Rong, Y.; Huang, W.; Ding, X.; Huang, Y.; Huang, J. Unsupervised Adversarial Graph Alignment with Graph Embedding. arXiv 2019, arXiv:1907.00544. [Google Scholar]
- Fan, J.; Cannistra, A.; Fried, I.; Lim, T.; Schaffner, T.; Crovella, M.; Hescott, B.; Leiserson, M.D.M. Functional protein representations from biological networks enable diverse cross-species inference. Nucleic Acids Res. 2019, 47, e51. [Google Scholar] [CrossRef] [PubMed]
- Zhou, D.; Schölkopf, B. A regularization framework for learning from graph data. In Proceedings of the ICML 2004 Workshop on Statistical Relational Learning and Its Connections to Other Fields (SRL 2004), Banff, AB, Canada, 8 July 2004; pp. 132–137. [Google Scholar]
- Kuhn, H.W. The Hungarian method for the assignment problem. Nav. Res. Logist. Q. 1955, 2, 83–97. [Google Scholar] [CrossRef] [Green Version]
- Gao, J.; Tian, L.; Lv, T.; Wang, J.; Song, B.; Hu, X. Protein2Vec: Aligning Multiple PPI Networks with Representation Learning. IEEE/ACM Trans. Comput. Biol. Bioinform. 2019, 18, 240–249. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, L.F.; Saverese, P.H.; Figueiredo, D.R. struc2vec: Learning node representations from structural identity. In Proceedings of the 23rd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, Halifax, NS, Canada, 13–17 August 2017; pp. 385–394. [Google Scholar]
- Liu, Y.; Ding, H.; Chen, D.; Xu, J. Novel geometric approach for global alignment of PPI networks. In Proceedings of the Thirty-First AAAI Conference on Artificial Intelligence, San Francisco, CA, USA, 4–9 February 2017. [Google Scholar]
- Shaw, B.; Jebara, T. Structure preserving embedding. In Proceedings of the 26th Annual International Conference on Machine Learning, Montreal, QC, Canada, 14–18 June 2009; pp. 937–944. [Google Scholar]
- Heimann, M.; Shen, H.; Safavi, T.; Koutra, D. REGAL: Representation Learning-based Graph Alignment. arXiv 2018, arXiv:1802.06257. [Google Scholar]
- Bentley, J.L. Multidimensional binary search trees used for associative searching. Commun. ACM 1975, 18, 509–517. [Google Scholar] [CrossRef]
- Chen, X.; Heimann, M.; Vahedian, F.; Koutra, D. Consistent Network Alignment with Node Embedding. arXiv 2020, arXiv:2005.04725. [Google Scholar]
- Grave, E.; Joulin, A.; Berthet, Q. Unsupervised alignment of embeddings with wasserstein procrustes. In Proceedings of the The 22nd International Conference on Artificial Intelligence and Statistics, Okinawa, Japan, 16–18 April 2019; pp. 1880–1890. [Google Scholar]
- Qiu, J.; Dong, Y.; Ma, H.; Li, J.; Wang, K.; Tang, J. Network embedding as matrix factorization: Unifying deepwalk, line, pte, and node2vec. In Proceedings of the Eleventh ACM International Conference on Web Search and Data Mining, Los Angeles, CA, USA, 5–9 February 2018; pp. 459–467. [Google Scholar]
| Algorithm | GNA or LNA | PNA or MNA | One-to-One or Many-to-Many | Evaluation Measures |
|---|---|---|---|---|
| GRAAL | GNA | PNA | One-to-one | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| H-GRAAL | GNA | PNA | One-to-one | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| MI-GRAAL | GNA | PNA | One-to-one | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| C-GRAAL | GNA | PNA | One-to-one | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| L-GRAAL | GNA | PNA | One-to-one | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| IsoRank | GNA | PNA | One-to-one | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| GHOST | GNA | PNA | One-to-one | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| WAVE | GNA | PNA | One-to-one | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| MAGNA | GNA | PNA | One-to-one | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| MAGNA++ | GNA | PNA | One-to-one | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| SANA | GNA | PNA | One-to-one | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| IGLOO | GNA | PNA | One-to-one | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| NetworkBLAST | LNA | PNA | Many-to-many | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| NetAligner | LNA | PNA | Many-to-many | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| AlignNemo | LNA | PNA | Many-to-many | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| AlignMCL | LNA | PNA | Many-to-many | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| LocalAli | LNA | PNA | Many-to-many | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| GLAlign | LNA | PNA | Many-to-many | P-NC, R-NC, F-NC, GS3, NCV-S3, GO correctness, P-PF, R-PF, F-PF |
| MultiMAGNA++ | GNA | MNA | One-to-one | NCV-MNC, NCV-CIQ, LCCS, MNE, GC, P-PF, R-PF, F-PF |
| GEDEVO-M | GNA | MNA | One-to-one | NCV-MNC, NCV-CIQ, LCCS, MNE, GC, P-PF, R-PF, F-PF |
| IsoRankN | GNA | MNA | Many-to-many | NCV-MNC, NCV-CIQ, LCCS, MNE, GC, P-PF, R-PF, F-PF |
| SMETANA | GNA | MNA | Many-to-many | NCV-MNC, NCV-CIQ, LCCS, MNE, GC, P-PF, R-PF, F-PF |
| LocalAli | LNA | MNA | Many-to-many | NCV-MNC, NCV-CIQ, LCCS, MNE, GC, P-PF, R-PF, F-PF |
| NetCoffee | GNA | MNA | Many-to-many | NCV-MNC, NCV-CIQ, LCCS, MNE, GC, P-PF, R-PF, F-PF |
| BEAMS | GNA | MNA | Many-to-many | NCV-MNC, NCV-CIQ, LCCS, MNE, GC, P-PF, R-PF, F-PF |
| Algorithm | PNA or MNA | Output | Embedding Approach | Alignment Approach | External Biological Sources | Comparison Metrics |
|---|---|---|---|---|---|---|
| REGAL | MNA | Matching node list | Matrix factorization | Geo | Domain-agnostic | AA, -AA, Runtime |
| GeoAlign | PNA | Matching node list | Matrix Factorization | Geo | Network simplex Sequence similarity | ICS, SPE, MNE, COI |
| MUNK | PNA | MUNK similarity score matrix (nxm) | Matrix factorization | Opti | Sequence homologs GO annotations | GO Con, K-fs AUPR |
| Protein2Vec | MNA | Similarity ranking | Diffusion method | Opti | BLASTP | ME, MNE, EC, ICS, MNS |
| CONE-ALIGN | PNA | Alignment matrix | Matrix decomposition | Geo | Domain-agnostic | AA |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Milano, M.; Zucco, C.; Settino, M.; Cannataro, M. An Extensive Assessment of Network Embedding in PPI Network Alignment. Entropy 2022, 24, 730. https://doi.org/10.3390/e24050730
Milano M, Zucco C, Settino M, Cannataro M. An Extensive Assessment of Network Embedding in PPI Network Alignment. Entropy. 2022; 24(5):730. https://doi.org/10.3390/e24050730
Chicago/Turabian StyleMilano, Marianna, Chiara Zucco, Marzia Settino, and Mario Cannataro. 2022. "An Extensive Assessment of Network Embedding in PPI Network Alignment" Entropy 24, no. 5: 730. https://doi.org/10.3390/e24050730
APA StyleMilano, M., Zucco, C., Settino, M., & Cannataro, M. (2022). An Extensive Assessment of Network Embedding in PPI Network Alignment. Entropy, 24(5), 730. https://doi.org/10.3390/e24050730








