Entropy as a High-Level Feature for XAI-Based Early Plant Stress Detection
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. The Use of Entropy and Max–Min Features as Universal, Explainable, and High-Level
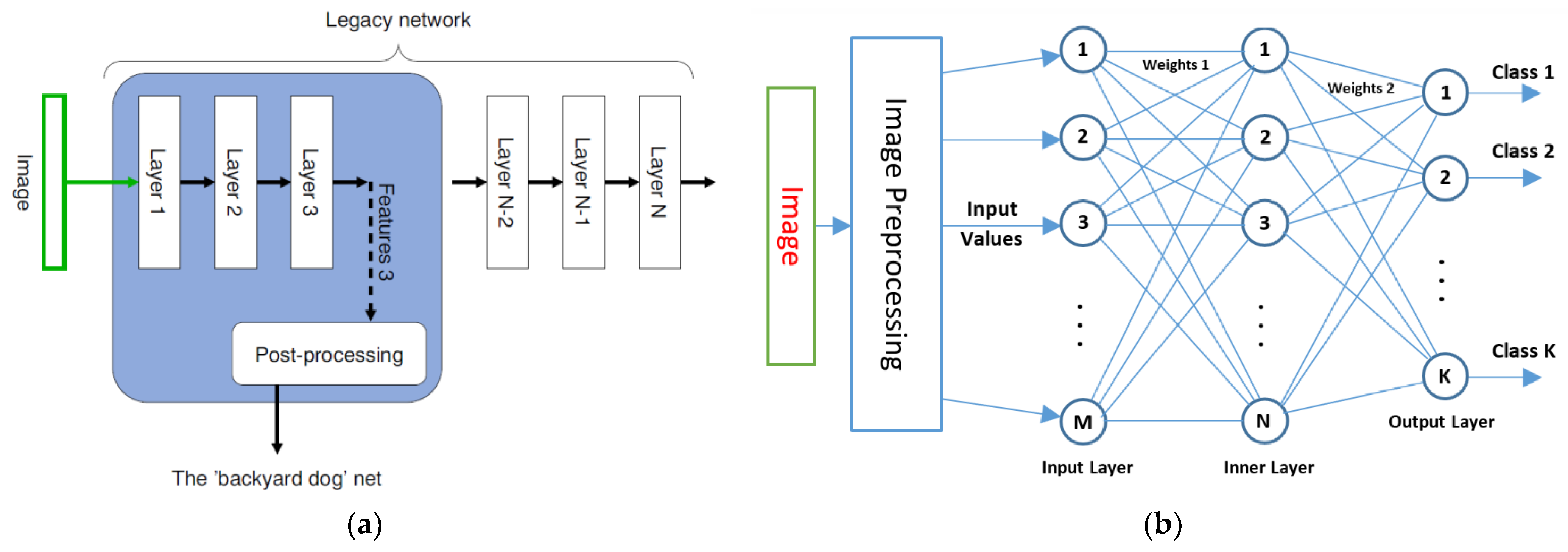
2.3. XAI-Based Classifier Description
Nw = M × N + N × K = 60 × 15 + 15 × 7 = 1005
2.4. Exploration Methods
3. Results
3.1. Significance of Each Feature from the Complete Feature Vector
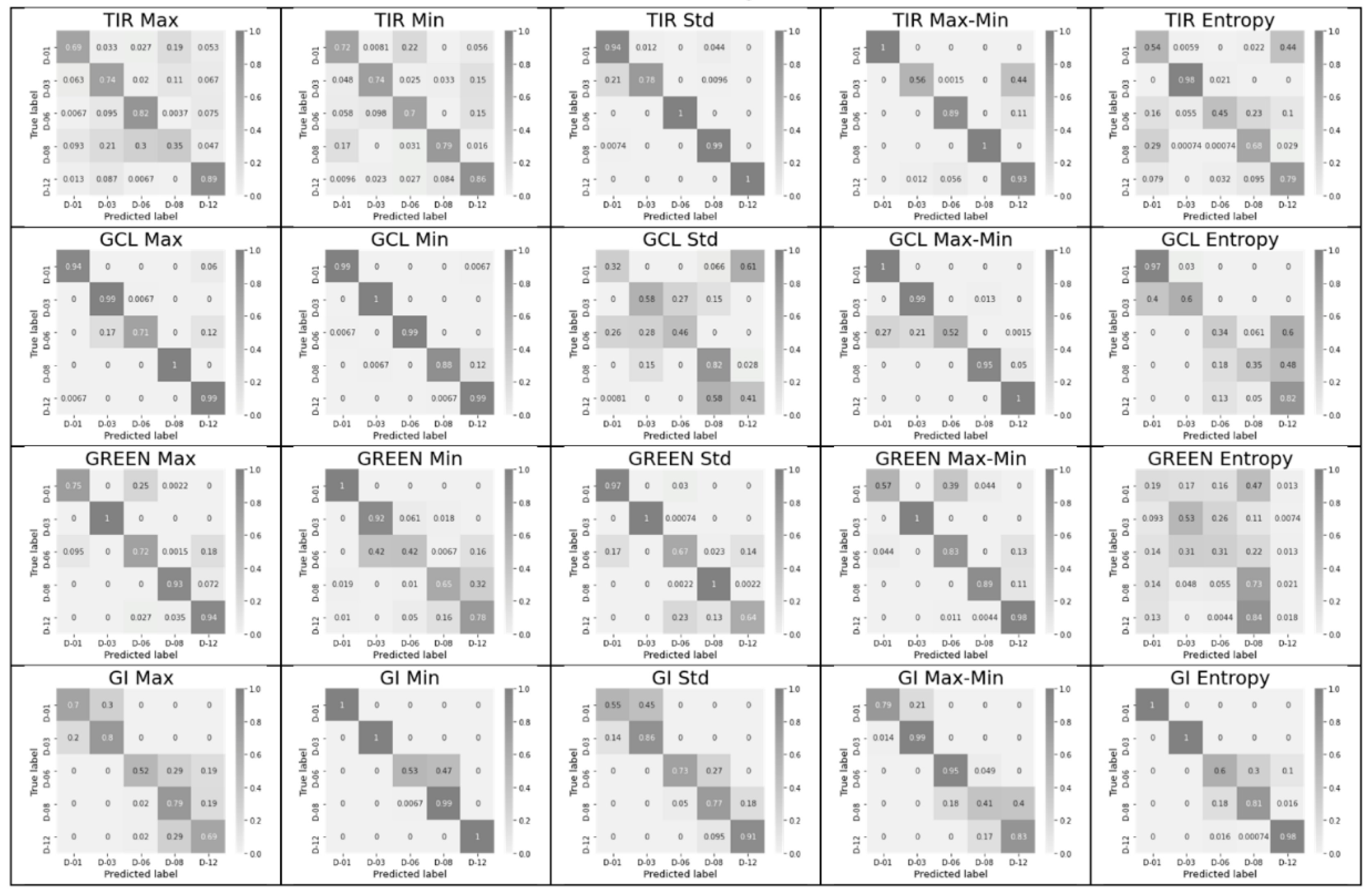
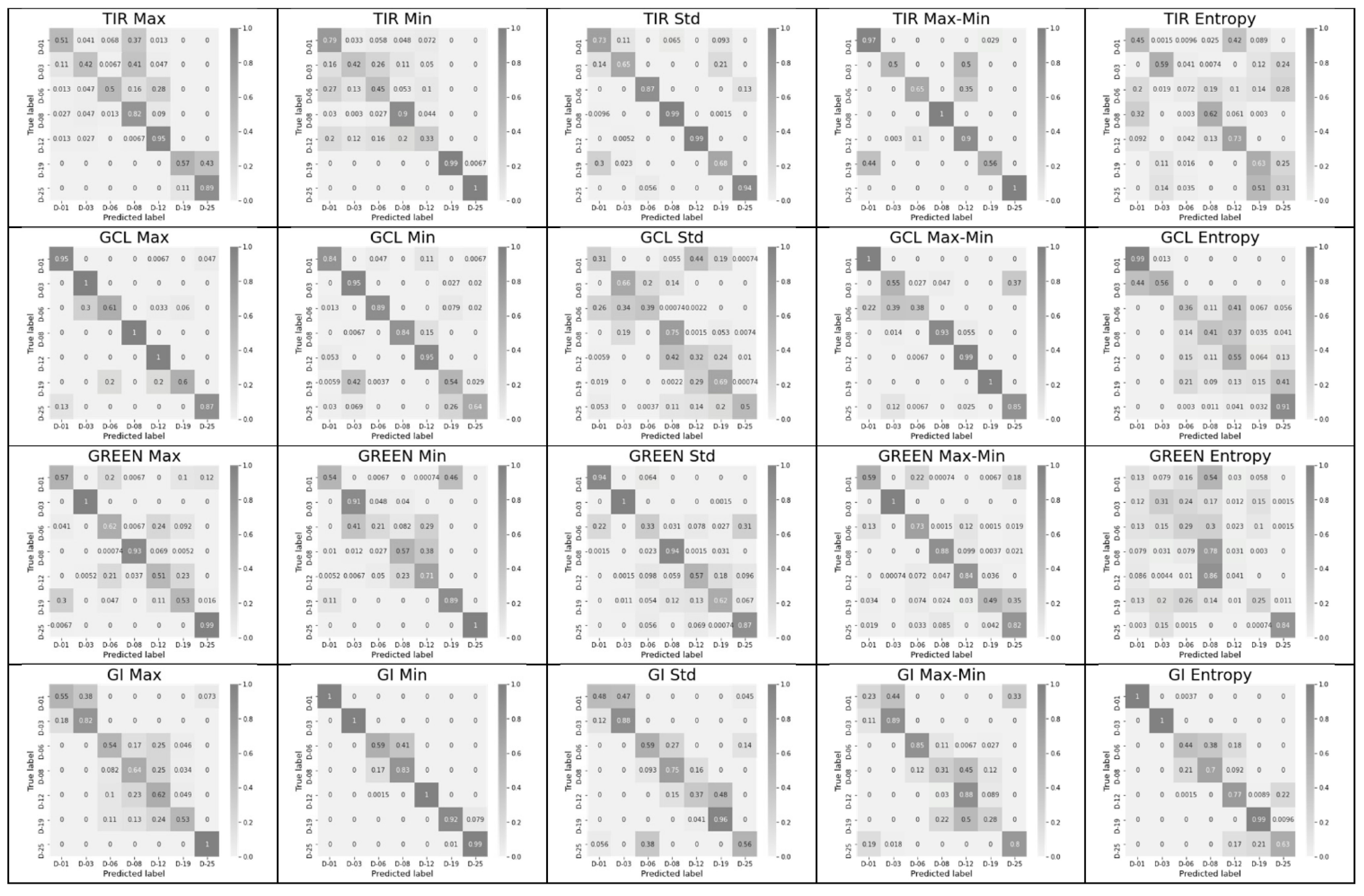
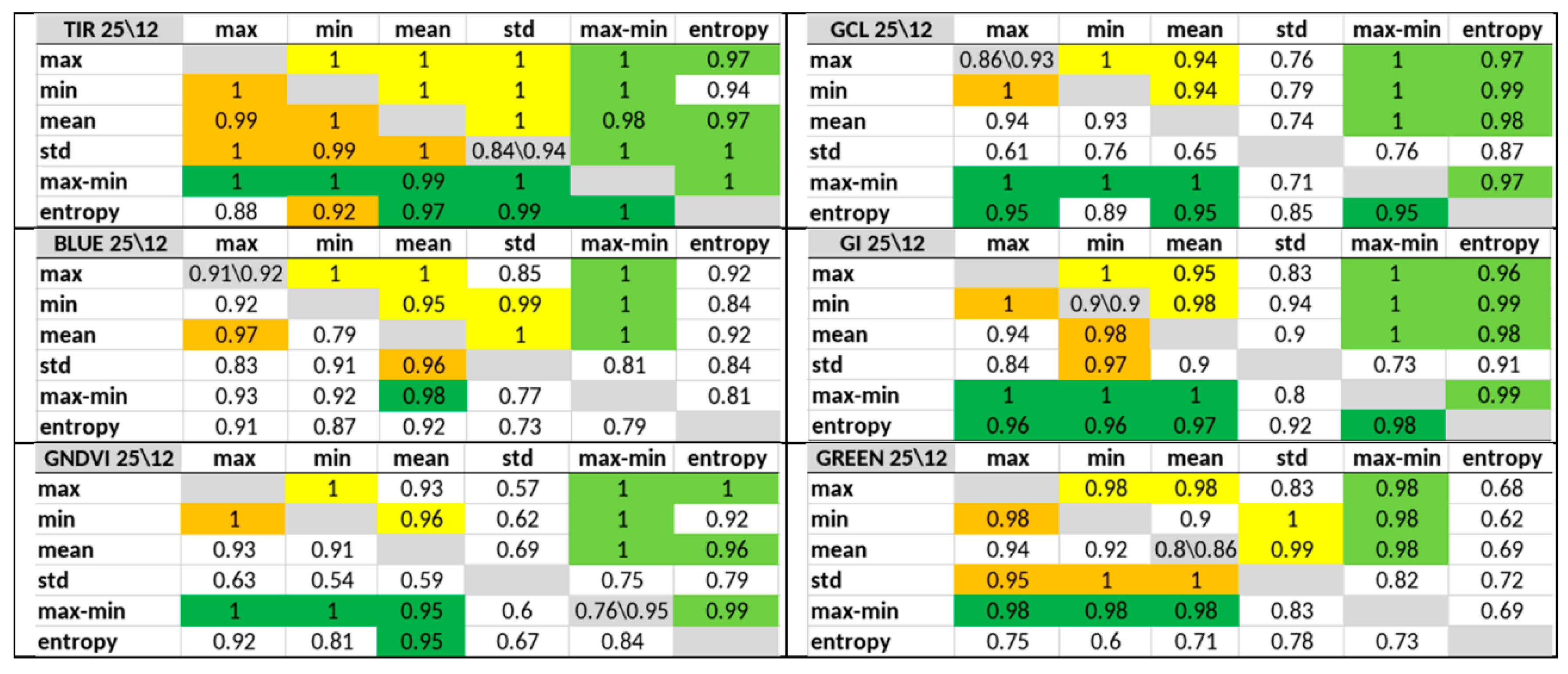
3.2. Significance of Feature Pairs within Each Index Separately and of the Entropy or Max–Min Presence in a Pair
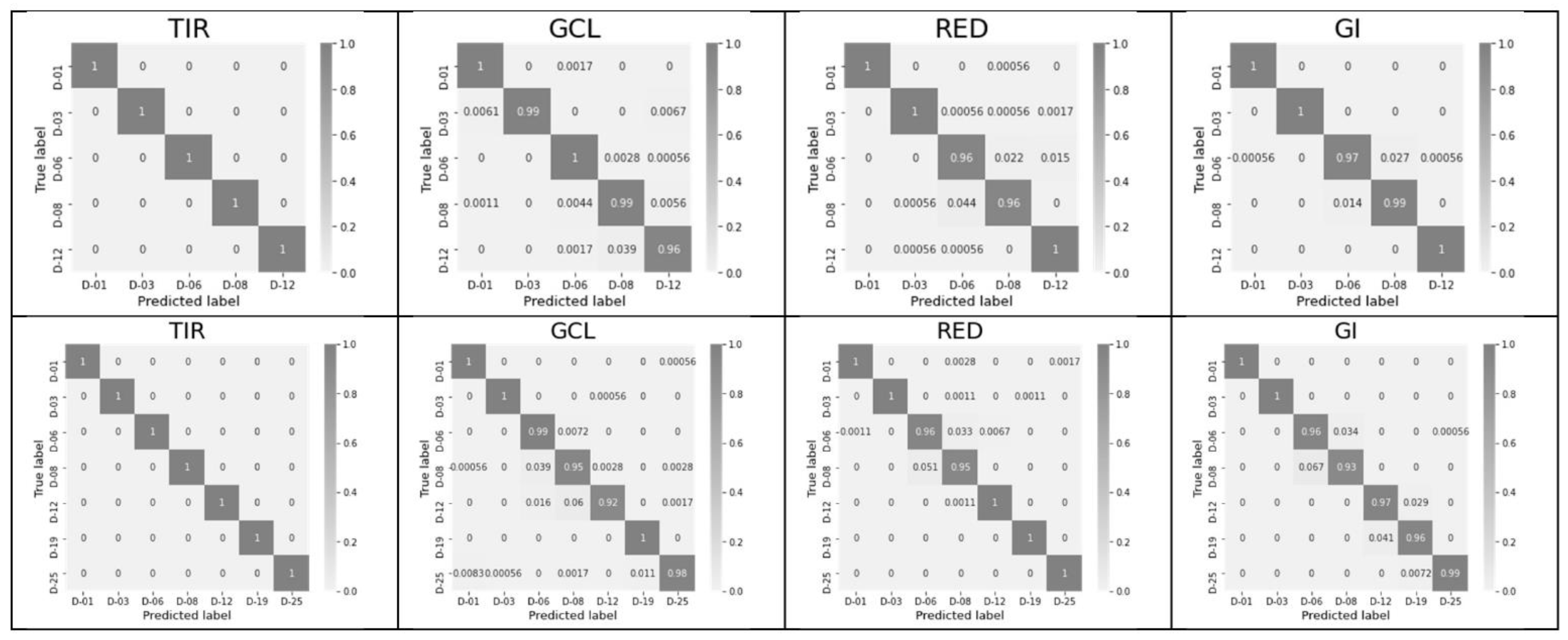
3.3. Significance of Excluding and Including All Six Features within Each Index and Using a Complete Feature Vector for All Indices Excluding or including TIR
4. Discussion
- (1)
- An SLP-classifier was built. The classifier structure was adjusted in terms of the number of neurons (N) used on the inner layer (according to the number of indices used), the length of the feature vector (M = m × N, where m = 1,2,…6), and the number of detected states (key days; K).
- (2)
- The classification accuracy of key days was determined individually for each of the 60 possible features and for their pair combinations within an index. Combinations that provide an accuracy of 1 (or near 1), as well as the number of such combinations for each of the indices, were determined, and indices and leaders were established.
- (3)
- We established that the involvement of the full-feature vector does not necessarily result in the maximum accuracy, and a set of at least two features and a few neurons in the inner layer is required to provide a solution to the problem.
- (4)
- We recommend to use the entropy as the main feature, as well as the max–min feature, which determines the number of states for which the entropy is calculated.
- (5)
- It is important to investigate not only the contribution (sensitivity) of individual features but also that of combinations of their minimal numbers inside indices. Therefore, the NDVI achieved the worst performance in index ranking when using all six features (Acc. = 0.77 for 25 days; see Table 3), but the best pair of NDVI features provided an Acc. = 0.97 for the 25-day range (see Table 2).
- (6)
- The use statistical features of the index image instead of the image itself as the MLP input and use of formalized key states as the output ensures the explainability of the SLP classifier as a whole and its high accuracy, representing a valuable XAI research tool.
- (7)
- Increasing the number of indices increases the robustness of the solution. As many as five index combinations may be required to ensure the robustness of solutions for smart farming applications.
5. Conclusions
- (1)
- Entropy can be used as a universal high-level explainable feature for classification, in particular for early detection of plant stress. The histogram of the single-channel image pixels belonging to any object of interest is the source for the entropy calculation.
- (2)
- The histogram width determined as max–min also can be used as a high-level explainable feature.
- (3)
- The entropy and the max–min features, in combination with the other histogram statistical features, should be used as the priority high-level input parameters of XAI neural networks (excluding pairs ‘max–min, std’, owing to their high correlation).
- −
- To replace the use of a complete set of statistical features of HSI-based indices;
- −
- To eliminate the need for thermal IR sensors for the early detection of plant stress;
- −
- To significantly reduce the requirements for sensors used in smart farming;
- −
- To eliminate the need for large datasets, energy, computational, and time resources for neural network training; and
- −
- For one-trial correction of AI systems [27].
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A

References
- Štrumbelj, E.; Kononenko, E. Explaining prediction models and individual predictions with feature contributions. Knowl. Inf. Syst. 2013, 41, 647–665. [Google Scholar] [CrossRef]
- Szegedy, C.; Zaremba, W.; Sutskever, I.; Bruna, J.; Erhan, D.; Goodfellow, I.; Fergus, R. Intriguing properties of neural networks. In Proceedings of the International Conference on Learning Representations, Banff, AB, Canada, 14–16 April 2014. [Google Scholar] [CrossRef]
- Wei, P.; Lu, Z.; Song, J. Variable importance analysis: A comprehensive review. Reliab. Eng. Syst. Saf. 2015, 142, 399–432. [Google Scholar] [CrossRef]
- Gorban, A.N.; Makarov, V.A.; Tyukin, I.Y. High-Dimensional Brain in a High-Dimensional World: Blessing of Dimensionality. Entropy 2020, 22, 82. [Google Scholar] [CrossRef] [PubMed]
- Hastie, T.; Tibshirani, R.; Friedman, J. The Elements of Statistical Learning. Data Mining, Reference, and Prediction; Springer Series in Statistics: New York, NY, USA, 2001; 764p. [Google Scholar]
- Ribeiro, M.T.; Singh, S.; Guestrin, C. “Why should i trust you?”: Explaining the predictions of any classifier. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 1135–1144. [Google Scholar] [CrossRef]
- Shrikumar, A.; Greenside, P.; Kundaje, A. Learning Important Features Through Propagating Activation Differences. arXiv 2017, arXiv:1704.02685. Available online: https://arxiv.org/abs/1704.02685 (accessed on 10 July 2022).
- Shrikumar, A.; Greenside, P.; Shcherbina, A.; Kundaje, A. Not Just a Black Box: Learning Important Features Through Propagating Activation Differences. arXiv 2016, arXiv:1605.01713. Available online: https://arxiv.org/abs/1605.01713 (accessed on 10 July 2022).
- Lundberg, S.M.; Lee, S.-I. A Unified Approach to Interpreting Model Predictio ns. In Proceedings of the 31st Conference on Neural Information Processing Systems (NeurIPS 2017), Long Beach, CA, USA, 4–9 December 2017; Available online: https://proceedings.neurips.cc/paper/2017/file/8a20a8621978632d76c43dfd28b67767-Paper.pdf (accessed on 10 July 2022).
- Zhao, Q.; Sheng, T.; Wang, Y.; Tang, Z.; Chen, Y.; Cai, L.; Ling, H. M2Det: A Single-Shot Object Detector Based on Multi-Level Feature Pyramid Network. In Proceedings of the AAAI Conference on Artificial Intelligence, Atlanta, Georgia, 8–12 October 2019; Volume 33, pp. 9259–9266. [Google Scholar]
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization. Int. J. Comput. Vis. 2020, 128, 336–359. [Google Scholar] [CrossRef]
- Albergante, L.; Mirkes, E.; Bac, J.; Chen, H.; Martin, A.; Faure, L.; Barillot, E.; Pinello, L.; Gorban, A.; Zinovyev, A. Robust and scalable learning of complex intrinsic dataset geometry via ElPiGraph. Entropy 2020, 22, 296. [Google Scholar] [CrossRef] [PubMed]
- Bac, J.; Mirkes, E.M.; Gorban, A.N.; Tyukin, I.; Zinovyev, A. Scikit-Dimension: A Python Package for Intrinsic Dimension Estimation. Entropy 2021, 23, 1368. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Yang, Y.; Pagnucco, M.; Song, Y. Explainability in Graph Neural Networks: An Experimental Survey. arXiv 2022, arXiv:2203.09258. [Google Scholar] [CrossRef]
- Ying, R.; Bourgeois, D.; You, J.; Zitnik, M.; Leskovec, J. GNNExplainer: Generating explanations for graph neural networks. NeurIPS 2019, 1, 1–13. [Google Scholar] [CrossRef]
- Luo, D.; Cheng, W.; Xu, D.; Yu, W.; Zong, B.; Chen, H.; Zhang, X. Parameterized explainer for graph neural network. NeurIPS 2020, 33, 19620–19631. [Google Scholar] [CrossRef]
- Vu, M.; Thai, M.T. PGM-Explainer: Probabilistic graphical model explanations for graph neural networks. In Proceedings of the NeurIPS 2020, Vancouver, BC, Canada, 6 December 2020; Available online: https://arxiv.org/abs/2010.05788 (accessed on 10 July 2022).
- Schlichtkrull, M.S.; De Cao, N.; Titov, I. Interpreting graph neural networks for NLP with differentiable edge masking. In Proceedings of the ICLR, Virtual Event, Austria, 3–7 May 2021. [Google Scholar] [CrossRef]
- Linardatos, P.; Papastefanopoulos, V.; Kotsiantis, S. Explainable AI: A Review of Machine Learning Interpretability Methods. Entropy 2021, 23, 18. [Google Scholar] [CrossRef] [PubMed]
- Pan, E.; Ma, Y.; Fan, F.; Mei, X.; Huang, J. Hyperspectral Image Classification across Different Datasets: A Generalization to Unseen Categories. Remote Sens. 2021, 13, 1672. [Google Scholar] [CrossRef]
- Dausset, J. Vegetation Indices for Chlorophyll (CI–MTCI–NDRE–ND705–ND550–mNDblue). Plant Phenotyping Vegetation Indices for Chlorophyll—Blog Hiphen (hiphen-plant.com). Available online: https://www.hiphen-plant.com/vegetation-indices-chlorophyll/3612/ (accessed on 10 July 2022).
- Jha, K.; Doshi, A.; Patel, P.; Shah, M. A comprehensive review on automation in agriculture using artificial intelligence. Artif. Intell. Agric. 2019, 2, 1–12. [Google Scholar] [CrossRef]
- Talaviya, T.; Shah, D.; Patel, N.; Yagnik, H.; Shah, M. Implementation of artificial intelligence in agriculture for optimisation of irrigation and application of pesticides and herbicides. Artif. Intell. Agric. 2020, 4, 58–73. [Google Scholar] [CrossRef]
- Pathan, M.; Patel, N.; Yagnik, H.; Shah, M. Artificial cognition for applications in smart agriculture: A comprehensive review. Artif. Intell. Agric. 2020, 4, 81–95. [Google Scholar] [CrossRef]
- Maximova, I.; Vasiliev, E.; Getmanskaya, A.; Kior, D.; Sukhov, V.; Vodeneev, V.; Turlapov, V. Study of XAI-capabilities for early diagnosis of plant drought. In Proceedings of the 2021 International Joint Conference on Neural Networks (IJCNN): International Joint Conference on Neural Networks, Shenzhen, China, 18–22 July 2021. [Google Scholar] [CrossRef]
- Dao, P.D.; He, Y.; Proctor, C. Plant drought impact detection using ultra-high spatial resolution hyperspectral images and machine learning. Int. J. Appl. Earth Obs. Geoinf. 2021, 102, 102364. [Google Scholar] [CrossRef]
- Gorban, A.N.; Burton, R.; Romanenko, I.; Tyukin, I.Y. One-trial correction of legacy AI systems and stochastic separation theorems. Inf. Sci. 2019, 484, 237–254. [Google Scholar] [CrossRef]
- Gorban, A.N.; Mirkes, E.M.; Tyukin, I.Y. How Deep Should be the Depth of Convolutional Neural Networks: A Backyard Dog Case Study. Cogn. Comput. 2020, 12, 388–397. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Jian Sun, J. Delving Deep into Rectifiers: Surpassing Human-Level Performance on ImageNet Classification. In Proceedings of the International Conference on Computer Vision, Las Condes, Chile, 11–18 December 2015. [Google Scholar] [CrossRef]
- Haralick, R.M. Pattern recognition with measurement space and spatial clustering for multiple image. Proc. IEEE 1969, 57, 654–665. [Google Scholar] [CrossRef]
| Index/Feature | Accuracy, 12 | Accuracy, 25 | Index/Feature | Accuracy, 12 | Accuracy, 25 |
|---|---|---|---|---|---|
| TIR/max | 0.70 | 0.67 | RED/max | 0.70 | 0.79 |
| TIR/min | 0.76 | 0.7 | RED/min | 0.87 | 0.82 |
| TIR/mean | 0.76 | 0.67 | RED/mean | 0.71 | 0.71 |
| TIR/std | 0.94 | 0.84 | RED/std | 0.6 | 0.54 |
| TIR/max–min | 0.88 | 0.8 | RED/max–min | 0.63 | 0.71 |
| TIR/entropy | 0.69 | 0.49 | RED/entropy | 0.76 | 0.59 |
| NDVI/max | 0.79 | 0.71 | GCL/max | 0.93 | 0.86 |
| NDVI/min | 0.61 | 0.50 | GCL/min | 0.97 | 0.81 |
| NDVI/mean | 0.8 | 0.68 | GCL/mean | 0.84 | 0.74 |
| NDVI/std | 0.69 | 0.65 | GCL/std | 0.52 | 0.52 |
| NDVI/max–min | 0.55 | 0.47 | GCL/max–min | 0.89 | 0.82 |
| NDVI/entropy | 0.55 | 0.57 | GCL/entropy | 0.61 | 0.56 |
| GNDVI/max | 0.81 | 0.72 | SIPI/max | 0.80 | 0.74 |
| GNDVI/min | 0.91 | 0.72 | SIPI/min | 0.83 | 0.84 |
| GNDVI/mean | 0.82 | 0.70 | SIPI/mean | 0.7 | 0.66 |
| GNDVI/std | 0.44 | 0.37 | SIPI/std | 0.67 | 0.66 |
| GNDVI/max–min | 0.95 | 0.76 | SIPI/max–min | 0.80 | 0.72 |
| GNDVI/entropy | 0.69 | 0.55 | SIPI/entropy | 0.61 | 0.55 |
| BLUE/max | 0.92 | 0.91 | NDblue/max | 0.91 | 0.71 |
| BLUE/min | 0.62 | 0.64 | NDblue/min | 0.58 | 0.51 |
| BLUE/mean | 0.79 | 0.73 | NDblue/mean | 0.60 | 0.59 |
| BLUE/std | 0.82 | 0.70 | NDblue/std | 0.8 | 0.6 |
| BLUE/max–min | 0.8 | 0.79 | NDblue/max–min | 0.93 | 0.74 |
| BLUE/entropy | 0.78 | 0.6 | NDblue/entropy | 0.58 | 0.42 |
| GREEN/max | 0.87 | 0.73 | GI/max | 0.70 | 0.67 |
| GREEN/min | 0.75 | 0.69 | GI/min | 0.90 | 0.90 |
| GREEN/mean | 0.86 | 0.80 | GI/mean | 0.81 | 0.79 |
| GREEN/std | 0.85 | 0.75 | GI/std | 0.76 | 0.66 |
| GREEN/max–min | 0.85 | 0.76 | GI/max–min | 0.80 | 0.61 |
| GREEN/entropy | 0.36 | 0.38 | GI/entropy | 0.88 | 0.79 |
| Combination of Features | Accuracy, 12 Days | Training Time (12 Days), s | Accuracy, 25 Days | Training Time (25 Days), s |
|---|---|---|---|---|
| GI/max, min | 1 | 0.19 | 1 | 0.31 |
| TIR/max–min, entropy | 1 | 0.20 | 1 | 0.34 |
| GCL/min, max–min | 1 | 0.23 | 1 | 0.32 |
| GI/min, max–min | 1 | 0.21 | 1 | 0.35 |
| GI/max, max–min | 1 | 0.21 | 1 | 0.36 |
| GCL/max, min | 1 | 0.29 | 1 | 0.32 |
| TIR/std, max–min | 1 | 0.19 | 1 | 0.46 |
| GI/mean, max–min | 1 | 0.23 | 1 | 0.43 |
| GCL/max, max–min | 1 | 0.31 | 1 | 0.38 |
| GCL/mean, max–min | 1 | 0.37 | 1 | 0.44 |
| TIR/max, min | 1 | 0.41 | 1 | 0.43 |
| TIR/max, max–min | 1 | 0.43 | 1 | 0.43 |
| TIR/min, mean | 1 | 0.41 | 1 | 0.48 |
| TIR/min, max–min | 1 | 0.40 | 1 | 0.50 |
| GNDVI/max, min | 1 | 0.42 | 1 | 0.57 |
| GNDVI/max, max–min | 1 | 0.42 | 1 | 0.58 |
| GNDVI/min, max–min | 1 | 0.41 | 1 | 0.65 |
| TIR/mean, std | 1 | 0.44 | 1 | 0.74 |
| TIR/max, std | 1 | 0.59 | 1 | 0.62 |
| GREEN/min, std | 1 | 0.55 | 1 | 0.79 |
| TIR/std, entropy | 1 | 0.38 | 0.99 | 0.60 |
| NDVI/max, mean | 0.96 | 0.8 | 0.97 | 0.87 |
| Index | Accuracy, 12 Days | Training Time (12 Days), s | Accuracy, 25 Days | Training Time (25 Days), s |
|---|---|---|---|---|
| TIR | 1 | 0.39 | 1 | 0.40 |
| GCL | 0.99 | 0.21 | 0.98 | 0.34 |
| RED | 0.98 | 0.63 | 0.99 | 0.81 |
| GI | 0.99 | 0.23 | 0.97 | 0.33 |
| BLUE | 0.97 | 0.66 | 0.95 | 0.67 |
| GNDVI | 0.96 | 0.35 | 0.93 | 0.54 |
| NDblue | 0.91 | 0.86 | 0.87 | 0.91 |
| GREEN | 0.82 | 0.74 | 0.95 | 0.89 |
| SIPI | 0.85 | 0.58 | 0.87 | 0.87 |
| NDVI | 0.79 | 0.49 | 0.77 | 0.58 |
| Combination of Indices | Accuracy (12 Days) | Training Time (12 Days), s | Accuracy (25 Days) | Training Time (25 Days), s |
|---|---|---|---|---|
| GCL, GI | 1 | 0.15 | 1 | 0.17 |
| BLUE, RED | 1 | 0.41 | 1 | 0.49 |
| GNDVI, GI | 1 | 0.13 | 0.99 | 0.30 |
| GNDVI, RED | 1 | 0.34 | 0.99 | 0.53 |
| GNDVI, NDblue | 0.97 | 0.55 | 1 | 0.54 |
| GREEN, GCL | 0.93 | 0.49 | 1 | 0.42 |
| GNDVI, GREEN | 0.9 | 0.67 | 1 | 0.49 |
| GNDVI, RED, GI | 1 | 0.15 | 1 | 0.19 |
| GNDVI, BLUE, GI | 1 | 0.15 | 1 | 0.21 |
| GNDVI, SIPI, GI | 1 | 0.15 | 1 | 0.25 |
| GNDVI, GREEN, GI | 1 | 0.27 | 1 | 0.38 |
| GREEN, RED, GI | 1 | 0.48 | 1 | 0.45 |
| GNDVI, NDblue, GI | 0.99 | 0.53 | 1 | 0.39 |
| GNDVI, BLUE, NDblue | 0.99 | 0.60 | 1 | 0.51 |
| GNDVI, GREEN, NDblue | 0.97 | 0.46 | 1 | 0.38 |
| GNDVI, GREEN, RED, SIPI | 0.99 | 0.46 | 1 | 0.55 |
| GREEN, RED, SIPI, NDblue | 1 | 0.52 | 0.99 | 0.67 |
| GREEN, RED, GCL, NDblue | 0.99 | 0.37 | 0.99 | 0.3 |
| Results of [26] | Our Results |
|---|---|
| A total of three of nine index images (CIRed-edge, mSR705, and SR [21,26]) indicated significant differences after 3 days of water treatment. Most of the indices were sensitive to drought-induced change after 6 days of the water treatment, with the exception of NDVI, ARI, and CCRI. | We used six statistical features instead of the image for each of 10 indices. We considered individual stress detection possibilities for each of 60 features and their pairs for two time ranges (12 and 25 days). All indices were able to detect plant stress states in both intervals with an accuracy 1 or near 1 (0.96). Entropy detected the earliest changes for all used indices, including the NDVI. |
| Indices cannot be used individually for quality diagnostics on all days. The best result (although not ideal) was achieved by a mixture of indices. | Indices can be employed for quality diagnostics on all days, using some feature pairs. The application of all six features is less productive than the use of pairs. Using a mixture of 2–5 indices guarantees an accuracy 1. |
| A near-ideal result was achieved by MLP with the use of average HSI signature curves and their derivatives, designated as DNN-Full and DNN-Deriv(atives) at the input. | SLP (practical case of MLP) is sufficient for plant stress classification with an accuracy 1. The employed properties of HSI signature derivatives practically equal to the properties of NDVI. |
| Owing to the use of hyperspectra as features, it was necessary to train the classifier for each plant type and possible irrigation condition. | Using the entropy and max–min features in a pair with another suitable statistical feature, we obtained the simplest and robust XAI-classifier, independent from plant type, irrigation conditions, temperature, and other external conditions. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lysov, M.; Maximova, I.; Vasiliev, E.; Getmanskaya, A.; Turlapov, V. Entropy as a High-Level Feature for XAI-Based Early Plant Stress Detection. Entropy 2022, 24, 1597. https://doi.org/10.3390/e24111597
Lysov M, Maximova I, Vasiliev E, Getmanskaya A, Turlapov V. Entropy as a High-Level Feature for XAI-Based Early Plant Stress Detection. Entropy. 2022; 24(11):1597. https://doi.org/10.3390/e24111597
Chicago/Turabian StyleLysov, Maxim, Irina Maximova, Evgeny Vasiliev, Alexandra Getmanskaya, and Vadim Turlapov. 2022. "Entropy as a High-Level Feature for XAI-Based Early Plant Stress Detection" Entropy 24, no. 11: 1597. https://doi.org/10.3390/e24111597
APA StyleLysov, M., Maximova, I., Vasiliev, E., Getmanskaya, A., & Turlapov, V. (2022). Entropy as a High-Level Feature for XAI-Based Early Plant Stress Detection. Entropy, 24(11), 1597. https://doi.org/10.3390/e24111597







