Due to scheduled maintenance work on our servers, there may be short service disruptions on this website between 11:00 and 12:00 CEST on March 28th.
Journal Description
DNA
DNA
is an international, peer-reviewed, open access journal on DNA and DNA-related technologies published quarterly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within ESCI (Web of Science) and other databases.
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 36 days after submission; acceptance to publication is undertaken in 14.7 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: APC discount vouchers, optional signed peer review, and reviewer names published annually in the journal.
Latest Articles
The Improvements and Applications of Prime Editing
DNA 2026, 6(1), 16; https://doi.org/10.3390/dna6010016 - 20 Mar 2026
Abstract
►
Show Figures
Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)/Cas9, a genome-editing technology pioneered in 2012, enables the precise correction of deleterious mutations or disruption of disease-causing genes through targeted double-strand breaks (DSBs), offering potential for treating genetic diseases. However, CRISPR/Cas9 can cause off-target cleavage at
[...] Read more.
Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)/Cas9, a genome-editing technology pioneered in 2012, enables the precise correction of deleterious mutations or disruption of disease-causing genes through targeted double-strand breaks (DSBs), offering potential for treating genetic diseases. However, CRISPR/Cas9 can cause off-target cleavage at non-specific DNA sites, leading to unintended insertions or deletions (indels), which limit its safety and applicability despite ongoing improvements in specificity. Recently, prime editing (PE), an advanced CRISPR-derived technology, has been employed with a Cas9 nickase (Cas9n) fused with a reverse transcriptase and a prime editing guide RNA (pegRNA) to enable precise insertions, deletions, and transversions without inducing DSBs, thus reducing risks of indels and chromosomal aberrations. Furthermore, ongoing optimizations, such as improved pegRNA design and enhanced editing efficiency, have expanded the applications of PE in medical therapeutics, agriculture, and fundamental research. This review summarizes recent advancements in the PE system, including optimized pegRNA designs and enzyme engineering for enhanced efficiency and specificity, alongside novel delivery methods. It also evaluates cutting-edge delivery strategies, such as adeno-associated virus (AAV) vectors, lipid nanoparticles (LNPs) and novel extracellular vesicle (EV)-based systems, and explores PE applications in vitro and in vivo, including disease modeling and therapeutic gene correction.
Full article
Open AccessArticle
An Integrative Evolutionary–Genomic Analysis Reveals the Factors That Shape the Sexual Diversity and Molecular Specificity of Gametophytic Self-Incompatibility in Prunus Species
by
Shashi N. Goonetilleke and Michelle G. Wirthensohn
DNA 2026, 6(1), 15; https://doi.org/10.3390/dna6010015 (registering DOI) - 13 Mar 2026
Abstract
►▼
Show Figures
Background: Gametophytic self-incompatibility (GSI) controlled by a multi-allelic S-locus, is inferred to have evolved before the spilt of the Rosidae and Asteridae. In Rosaceae, molecular characterisation of the genera Prunus and Malus reveals that different numbers of genes determine GSI specificity. In
[...] Read more.
Background: Gametophytic self-incompatibility (GSI) controlled by a multi-allelic S-locus, is inferred to have evolved before the spilt of the Rosidae and Asteridae. In Rosaceae, molecular characterisation of the genera Prunus and Malus reveals that different numbers of genes determine GSI specificity. In Prunus, one pistil-expressed (female) gene and one pollen (male) gene encode a series of stylar RNase (S-RNase) alleles and series of S-haplotype-specific F-box (SFB) alleles, respectively, thereby determining the female and male specificity. In contrast, in Malus, GSI specificity is controlled by one pistil gene and multiple pollen genes, known as SFB-brothers (SFBBs), which encode a series of S-RNase and SFBB alleles, respectively, within the S-locus, to determine female and male specificity. Despite these advances, the molecular mechanisms of these two genera remain largely unknown, and it is still uncertain how GSI originated or which factors shape the orientation, evolution, and function of the S-locus. Methods: Therefore, in this study, we applied a holistic integrative approach combining analyses of gene distribution, phylogenetic inference, biogeographic history, selective pressures, co-evolution, and protein interaction networks across three Prunus genomes (P. dulcis, P. persica, and P. avium) to elucidate the evolutionary forces driving sexual diversity and molecular specificity of GSI within the Rosaceae. Results: Our results indicated that rapid diversification of the Prunus S-locus was due to the repeated duplication events in the SFB, SLF, and S-RNase genes producing both functional and non-functional duplicates. Conclusions: In Rosaceae, diversity of S-locus mechanisms is shaped by lineage-specific selection, functional divergence, co-evolution of pistil- and pollen-expressed components, dynamic protein-interaction networks, geological history and climatic change.
Full article

Graphical abstract
Open AccessReview
Genetic Susceptibility in Sudden Infant Death Syndrome (SIDS): A Narrative Review of Emerging Evidence
by
Eteesha Rao and Srinivas Annavarapu
DNA 2026, 6(1), 14; https://doi.org/10.3390/dna6010014 - 5 Mar 2026
Abstract
Sudden Infant Death Syndrome (SIDS) continues to be one of the most challenging and tragic causes of infant mortality in developed countries. While public health interventions have reduced its prevalence, the underlying mechanisms contributing to SIDS remain largely unclear. The biological basis of
[...] Read more.
Sudden Infant Death Syndrome (SIDS) continues to be one of the most challenging and tragic causes of infant mortality in developed countries. While public health interventions have reduced its prevalence, the underlying mechanisms contributing to SIDS remain largely unclear. The biological basis of SIDS is widely believed to be multifactorial in nature, involving inherited genetic vulnerabilities, including mutations in cardiac ion channels and genes associated with brainstem serotonin function, metabolic enzymes, and inflammatory mediators. This review presents a comprehensive analysis of genetic studies relating to SIDS, incorporating recent findings from molecular autopsies, genome-wide association studies and functional assays. It also explores how gene–environment interactions, polygenic risk scores, and multi-omic strategies are reshaping our understanding of this complex condition. The review aims to integrate recent insights from molecular autopsy, genomic profiling, and gene–environment interactions to offer a framework for better risk assessment and the stratification of vulnerable infants who could benefit from targeted clinical and public health interventions.
Full article
Open AccessReview
Recognition Mechanism of Complementary Nucleobases and Sequences in DNA and RNA: Interplay of Watson–Crick Hydrogen Bond Formation and Base Stacking Interactions
by
Masayuki Takahashi and Bengt Nordén
DNA 2026, 6(1), 13; https://doi.org/10.3390/dna6010013 - 4 Mar 2026
Abstract
►▼
Show Figures
A/T(U) and G/C nucleobase pair formation in DNA and RNA is crucial to numerous fundamental biological processes, including replication, transcription, and translation. The specificity of A/T(U) and G/C base pairing is used for the recognition of complementary sequences in medical and biotechnological applications,
[...] Read more.
A/T(U) and G/C nucleobase pair formation in DNA and RNA is crucial to numerous fundamental biological processes, including replication, transcription, and translation. The specificity of A/T(U) and G/C base pairing is used for the recognition of complementary sequences in medical and biotechnological applications, such as PCR, nucleic acid drugs, and CRISPR–Cas9-based gene editing. It is essential to understand and predict fidelity of biological reactions, avoiding off-target binding, in order to improve the accuracy and efficacy of applications. In particular, recognition mechanisms of complementary bases or whole sequences must be understood in detail. Despite the prevailing view that Watson–Crick hydrogen bonding is a primary mechanism for complementary base recognition, several experiments have shown that DNA polymerase does not require hydrogen bonding to select complementary bases. Other factors, such as the shape and geometric fitting of the bases and the base stacking, also appear to be crucially involved in the selection. E.g., artificial bases lacking the ability to form hydrogen bonds can still be recognized by DNA polymerase solely based on base-pair geometry. However, hydrogen bonding also contributes importantly to recognition. The accuracy of selecting a complementary nucleobase or sequence varies depending on reactions, suggesting the co-existence of multiple selection mechanisms. This review provides an overview of biological processes and applications involving base pairing and discusses the molecular mechanism underlying complementary base recognition.
Full article

Figure 1
Open AccessReview
Genetic Architecture, Developmental Mechanisms and Genomic Applications in Left Ventricular Non-Compaction Cardiomyopathy (LVNC)
by
Luis Elias Martínez-Tittonel, Florin Radu Ciorba, Xavier Bayona-Huguet and Edgardo Kaplinsky
DNA 2026, 6(1), 12; https://doi.org/10.3390/dna6010012 - 2 Mar 2026
Abstract
Left ventricular noncompaction cardiomyopathy (LVNC) is characterised by a two-layered ventricular wall with prominent trabeculations and deep recesses adjacent to a thinned compact layer. The phenotype spans from incidental findings to severe heart failure and malignant arrhythmias. More than 190 genes belonging to
[...] Read more.
Left ventricular noncompaction cardiomyopathy (LVNC) is characterised by a two-layered ventricular wall with prominent trabeculations and deep recesses adjacent to a thinned compact layer. The phenotype spans from incidental findings to severe heart failure and malignant arrhythmias. More than 190 genes belonging to sarcomeric, cytoskeletal, mitochondrial, transcriptional and signalling pathways have been implicated, although only a subset reaches high gene disease validity in contemporary frameworks. Objectives: (i) Delineate the validated genetic landscape of LVNC; (ii) integrate developmental biology with cardiac genomics; (iii) translate genotype knowledge into diagnostic, prognostic and therapeutic guidance; (iv) outline a research agenda for precision cardiology. Methods: A narrative, pathway-oriented review of human and experimental studies (2000–July 2024). Results: Thirty-two genes meet definitive/strong validity thresholds and cluster in five biological networks. Oligogenic constellations account for ~4% of probands in recent cohorts. Imaging correlates (especially quantitative trabecular complexity and diffuse fibrosis metrics) provide complementary risk information. Conclusions: LVNC represents a convergence phenotype triggered by perturbations across developmental and structural networks; clinical management benefits from integrated genomics–imaging workflows and mechanism-informed trial design.
Full article
Open AccessArticle
Erimin: A Pipeline to Identify Bacterial Strain Specific Primers
by
Margaritis Tsifintaris, Paraskevi Koutra, Pavlos Tsiartas, Panagiotis Repanas, Sotirios Touliopoulos, Grigorios Nelios, Anastasia Anastasiadou, Georgia Tamouridou, Anastasios Nikolaou and Ilias Tsochantaridis
DNA 2026, 6(1), 11; https://doi.org/10.3390/dna6010011 - 25 Feb 2026
Abstract
►▼
Show Figures
Background/Objectives: Strain-level detection of bacteria is essential for applications such as diagnostics, food safety, and microbial monitoring. While 16S rRNA gene sequencing provides genus- or species-level resolution, it cannot reliably discriminate closely related strains. Whole-genome sequencing (WGS) offers high-resolution strain differentiation but remains
[...] Read more.
Background/Objectives: Strain-level detection of bacteria is essential for applications such as diagnostics, food safety, and microbial monitoring. While 16S rRNA gene sequencing provides genus- or species-level resolution, it cannot reliably discriminate closely related strains. Whole-genome sequencing (WGS) offers high-resolution strain differentiation but remains impractical for routine detection due to cost and analytical complexity. This study aims to enable the translation of WGS data into accurate and cost-effective strain-specific PCR assays. Methods: We developed Erimin, a modular, shell-based bioinformatics pipeline for the automated identification of strain-specific genomic regions from short-read WGS data. Erimin systematically analyzes all available reference genomes for a given bacterial species in combination with sequencing data from a target strain. The workflow integrates reference-based read alignment, extraction of unmapped reads, de novo assembly, contig filtering and validation, genome annotation, and in silico PCR primer design and specificity evaluation. Results: Erimin was applied to Lactiplantibacillus pentosus whole-genome sequencing data to identify genomic regions specific to strain L33 through comparative analysis against a comprehensive set of reference genome assemblies representing multiple Lactiplantibacillus species. These regions were used for in silico PCR primer design and computational specificity assessment against non-target bacterial genomes, supporting discrimination of closely related strains. Conclusions: Erimin provides a structured computational approach for identifying strain-specific genomic regions from WGS data and for supporting the in silico design of PCR primers. This framework facilitates strain-level discrimination using targeted molecular assays.
Full article

Graphical abstract
Open AccessArticle
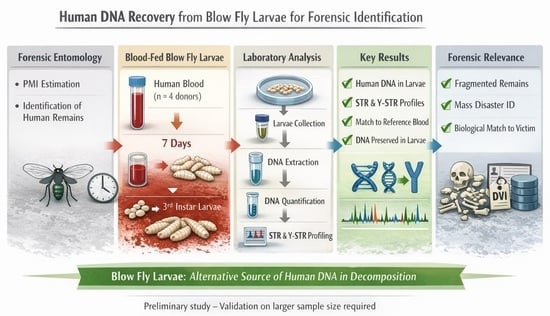
Assessment of Human DNA and Y-DNA Quantity in Blow Fly Larvae Cultured on Human Blood: Evaluation of Utility for Identification and Forensic Reporting in Sexual Offense Cases
by
Dagmara Lisman, Ilona Savochka, Ewelina Tarnawska and Andrzej Ossowski
DNA 2026, 6(1), 10; https://doi.org/10.3390/dna6010010 - 13 Feb 2026
Abstract
►▼
Show Figures
Background/Objectives: Forensic entomology remains an underutilized discipline within forensic medicine, particularly in Poland, where it is primarily applied to post-mortem interval (PMI) estimation. Recent studies indicate that insect-derived material may also hold value in the identification of human remains. Methods: In this pilot
[...] Read more.
Background/Objectives: Forensic entomology remains an underutilized discipline within forensic medicine, particularly in Poland, where it is primarily applied to post-mortem interval (PMI) estimation. Recent studies indicate that insect-derived material may also hold value in the identification of human remains. Methods: In this pilot study, we assess whether blow fly larvae fed on human blood retain amplifiable human DNA, including Y-DNA. Larvae were reared on blood obtained from four volunteers and collected at the third instar stage seven days after oviposition. Human DNA quantification, degradation assessment, and STR/Y-STR profiling were performed. Results: Despite the deliberately small, exploratory sample size, all larval samples yielded complete and concordant STR and, where applicable, Y-STR profiles matching the respective reference donors. Conclusions: These preliminary findings indicate the potential utility of larvae as an alternative biological substrate in forensic contexts, particularly when conventional tissues are unavailable or degraded. However, the results should be interpreted cautiously and require validation in larger, systematically controlled studies before any routine forensic application can be recommended.
Full article
Graphical abstract
Open AccessReview
Mechanisms of Fork Destabilization Under Hydroxyurea: The Interplay of ROS, Checkpoints, and Replisome Integrity
by
Srinivasu Karri and Chuanhe Yu
DNA 2026, 6(1), 9; https://doi.org/10.3390/dna6010009 - 9 Feb 2026
Abstract
►▼
Show Figures
Faithful DNA replication is essential for genome stability but is constantly challenged by metabolic and oxidative stresses. Hydroxyurea (HU), a widely used antiproliferative drug, is traditionally known to inhibit ribonucleotide reductase and deplete dNTP pools. Recent studies, especially in Saccharomyces cerevisiae, reveal that
[...] Read more.
Faithful DNA replication is essential for genome stability but is constantly challenged by metabolic and oxidative stresses. Hydroxyurea (HU), a widely used antiproliferative drug, is traditionally known to inhibit ribonucleotide reductase and deplete dNTP pools. Recent studies, especially in Saccharomyces cerevisiae, reveal that HU-induced replication stress also arises from reactive oxygen species (ROS), which oxidize DNA, impair iron–sulfur-dependent replication enzymes, and disrupt replisome function. These combined effects promote helicase–polymerase uncoupling, accumulation of RPA-coated ssDNA, and activation of the Mec1–Rad53 (ATR–CHK1) checkpoint, leading to strand-specific changes such as PCNA unloading and reduced lagging-strand synthesis. When protective pathways are overwhelmed, HU-treated forks collapse, generating chromosome breaks and genome instability. This review summarizes current understanding of how HU remodels replication forks through both ROS-dependent and ROS-independent pathways and highlights emerging insights into how these mechanisms influence genome stability and may be exploited for therapeutic benefit.
Full article

Graphical abstract
Open AccessArticle
Using the SIRAH Force-Field to Model Interactions Between Short DNA Duplexes
by
Romina Ruberto, Enrico Smargiassi and Giorgio Pastore
DNA 2026, 6(1), 8; https://doi.org/10.3390/dna6010008 - 2 Feb 2026
Abstract
►▼
Show Figures
Background/Objectives: In recent years, short DNA duplexes have been studied as promising self-assembling systems and versatile building blocks for DNA-based nanotechnologies. Numerical simulations of colloidal systems incorporating such components require, as an input ingredient, reliable yet simplified force-fields capable of capturing the essential
[...] Read more.
Background/Objectives: In recent years, short DNA duplexes have been studied as promising self-assembling systems and versatile building blocks for DNA-based nanotechnologies. Numerical simulations of colloidal systems incorporating such components require, as an input ingredient, reliable yet simplified force-fields capable of capturing the essential features of duplex-duplex interactions. Methods: We employed the coarse-grained SIRAH force field under an implicit solvent approximation to investigate the interactions between a pair of short, rigid double-stranded DNA (dsDNA) duplexes. We investigated the effect of duplex size by employing duplexes of 8 and 20 base pairs. Results: Using this realistic coarse-grained model, we obtained detailed insights into how the interaction force depends on the relative positions and orientations of the duplexes, as well as on salt concentration. Conclusions: Our findings provide a foundational step toward the systematic development of simplified, yet qualitatively accurate model potentials for DNA-based colloidal systems. Beyond nanotechnology, the short-range interaction features captured here are also relevant to biological contexts, including chromatin compaction, homologous recombination, and DNA repair.
Full article

Graphical abstract
Open AccessReview
Mitochondrial DNA Alterations in HPV-Related Cancers: Emerging Insights and Future Directions
by
Muharrem Okan Cakir, Melis Selek, Guldide Kayhan, Betul Yilmaz, Mustafa Ozdogan and Gholam Hossein Ashrafi
DNA 2026, 6(1), 7; https://doi.org/10.3390/dna6010007 - 2 Feb 2026
Abstract
►▼
Show Figures
Human papillomavirus (HPV) infection is a leading cause of cervical cancer and a significant contributor to anogenital and oropharyngeal malignancies worldwide. While the oncogenic functions of HPV oncoproteins E6 and E7 in disrupting nuclear tumor suppressor pathways are well established, their influence on
[...] Read more.
Human papillomavirus (HPV) infection is a leading cause of cervical cancer and a significant contributor to anogenital and oropharyngeal malignancies worldwide. While the oncogenic functions of HPV oncoproteins E6 and E7 in disrupting nuclear tumor suppressor pathways are well established, their influence on mitochondrial biology has only recently emerged as a critical facet of HPV-driven carcinogenesis. This review synthesizes current evidence on the qualitative and quantitative alterations of mitochondrial DNA (mtDNA) and their functional consequences in HPV-associated cancers. We discuss how E6 and E7 modulate mitochondrial dynamics, bioenergetics, and redox balance, contributing to metabolic reprogramming, resistance to apoptosis, and adaptation to tumor microenvironmental stress. We also examine the clinical significance of mtDNA mutations, deletions, and copy number variations as potential biomarkers for diagnosis, prognosis, and therapy response. Advances in multi-omics approaches, high-throughput sequencing, and patient-derived organoid models have accelerated the exploration of mitochondria as therapeutic targets. Integrating mitochondrial profiling into HPV-related cancer research holds promise for identifying novel metabolic vulnerabilities and guiding the development of mitochondria-directed treatment strategies.
Full article

Graphical abstract
Open AccessReview
Advancing Liver Cancer Treatment Through Dynamic Genomics and Systems Biology: A Path Toward Personalized Oncology
by
Giovanni Colonna
DNA 2026, 6(1), 6; https://doi.org/10.3390/dna6010006 - 21 Jan 2026
Abstract
►▼
Show Figures
This review aims to provide a broad, multidisciplinary perspective on how dynamic genomics and systems biology are transforming modern healthcare, with a focus on cancer especially liver cancer (HCC). It explains how integrating multi-omics technologies such as genomics, transcriptomics, proteomics, interactomics, metabolomics, and
[...] Read more.
This review aims to provide a broad, multidisciplinary perspective on how dynamic genomics and systems biology are transforming modern healthcare, with a focus on cancer especially liver cancer (HCC). It explains how integrating multi-omics technologies such as genomics, transcriptomics, proteomics, interactomics, metabolomics, and spatial transcriptomics deepens our understanding of the complex tumor environment. These innovations enable precise patient stratification based on molecular, spatial, and functional tumor characteristics, allowing for personalized treatment plans. Emphasizing the role of regulatory networks and cell-specific pathways, the review shows how mapping these networks using multi-omics data can predict resistance, identify therapeutic targets, and aid in the development of targeted therapies. The approach shifts from standard, uniform treatments to flexible, real-time strategies guided by technologies such as liquid biopsies and wearable biosensors. A case study showcases the benefits of personalized therapy, which integrates epigenetic modifications, checkpoint inhibitors, and ongoing multi-omics monitoring in a patient with HCC. Future innovations, such as cloud-based genomic ecosystems, federated learning for privacy, and AI-driven data analysis, are also discussed to enhance decision-making and outcomes. The review underscores a move toward predictive and preventive healthcare by integrating layered data into clinical workflows. It reviews ongoing clinical trials using advanced molecular and immunological techniques for HCC. Overall, it promotes a systemic, technological, and spatial approach to cancer treatment, emphasizing the importance of experimental, biochemical–functional, and biophysical data-driven insights in personalizing medicine.
Full article

Figure 1
Open AccessReview
Genetic Basis of Familial Cancer Risk: A Narrative Review
by
Eman Fares Sabik
DNA 2026, 6(1), 5; https://doi.org/10.3390/dna6010005 - 13 Jan 2026
Abstract
►▼
Show Figures
Familial cancers are caused by inherited mutations in specific genes that regulate cell growth, division, and repair. Approximately 5–10% of all cancer cases have a hereditary component, where germline mutations in certain genes increase an individual’s susceptibility to developing cancer. Two major categories
[...] Read more.
Familial cancers are caused by inherited mutations in specific genes that regulate cell growth, division, and repair. Approximately 5–10% of all cancer cases have a hereditary component, where germline mutations in certain genes increase an individual’s susceptibility to developing cancer. Two major categories of genes are involved in cancer development: tumour suppressor genes and oncogenes. Both play critical roles in regulating normal cell behaviour, and when mutated, they can contribute to uncontrolled cell proliferation and tumour formation. In addition to genetic mutations, epigenetic alterations also play a significant role in familial cancer. Epigenetics refers to changes in gene expression due to DNA methylation, histone modifications, and the dysregulation of non-coding RNAs without alter the underlying DNA sequence. Familial cancer syndromes follow various inheritance patterns, including autosomal dominant, autosomal recessive, X-linked, and mitochondrial inheritance, each with distinct characteristics. Identifying genetic mutations associated with familial cancers is a cornerstone of genetic counselling, which helps individuals and families navigate the complex intersection of genetics, cancer risk, and prevention. Early identification of mutations enables personalized strategies for risk reduction, early detection, and, when applicable, targeted treatment options, ultimately improving patient outcomes.
Full article

Figure 1
Open AccessArticle
The Effect of Fatty Acid-Binding Protein 3 Exposure on Endothelial Transcriptomics
by
Hien C. Nguyen, Aman Singh, Christina A. Castellani, Mohammad Qadura and Krishna K. Singh
DNA 2026, 6(1), 4; https://doi.org/10.3390/dna6010004 - 8 Jan 2026
Abstract
►▼
Show Figures
Background: Fatty acid-binding protein 3 (FABP3) is released in circulation following myocardial infarction, and an increased level of circulatory FABP3 has also been reported in peripheral artery disease patients, exposing endothelial cells to higher levels of FABP3. Recently, loss of endothelial FABP3 was
[...] Read more.
Background: Fatty acid-binding protein 3 (FABP3) is released in circulation following myocardial infarction, and an increased level of circulatory FABP3 has also been reported in peripheral artery disease patients, exposing endothelial cells to higher levels of FABP3. Recently, loss of endothelial FABP3 was shown to protect endothelial cells against inflammation-induced endothelial dysfunction; however, the effect of FABP3 exposure on endothelial cells is unknown. Accordingly, to study the effect of FABP3 exposure on endothelial cells, we performed transcriptomic profiling following recombinant human FABP3 (rhFABP3) treatment of endothelial cells. Methods: Cultured human endothelial cells were treated with either a vehicle or rhFABP3 (50 ng/mL, 6 h); then, RNA sequencing was performed. Gene expression analysis followed by gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses was performed to identify differentially expressed genes and affected cellular functions and pathways. Results: Differential gene expression analysis revealed kinesin family member 26b (KIF26B) to be the most upregulated and survival of motor neuron 2 (SMN2) to be the most downregulated genes in rhFABP3-treated compared to vehicle-treated endothelial cells. Most of the differentially expressed genes were associated with endothelial cell motility, immune response, and angiogenesis. GO and KEGG analyses indicated that rhFABP3 exposure impacts several crucial pathways, predominantly “Regulation of leukocyte mediated cytotoxicity” and “Natural killer cell mediated cytotoxicity”, suggesting its involvement in endothelial cell physiology and response mechanisms to cardiovascular stress. Conclusions: This is the first study to evaluate rhFABP3-induced transcriptomics in human endothelial cells. Our data reveal novel genes and pathways affected by the exposure of endothelial cells to FABP3. Further research is necessary to validate these findings and fully understand FABP3’s role in endothelial biology and in cardiovascular diseases like myocardial infarction and peripheral artery disease.
Full article

Figure 1
Open AccessArticle
A Preliminary Machine Learning Assessment of Oxidation-Reduction Potential and Classical Sperm Parameters as Predictors of Sperm DNA Fragmentation Index
by
Emmanouil D. Oikonomou, Efthalia Moustakli, Athanasios Zikopoulos, Stefanos Dafopoulos, Ermioni Prapa, Antonis-Marios Gkountis, Athanasios Zachariou, Agni Pantou, Nikolaos Giannakeas, Konstantinos Pantos, Alexandros T. Tzallas and Konstantinos Dafopoulos
DNA 2026, 6(1), 3; https://doi.org/10.3390/dna6010003 - 8 Jan 2026
Cited by 1
Abstract
►▼
Show Figures
Background/Objectives: Traditional semen analysis techniques frequently result in incorrect male infertility diagnoses, despite advancements in assisted reproductive technology (ART). Reduced fertilization potential, decreased embryo development, and lower pregnancy success rates are associated with elevated DNA Fragmentation Index (DFI), which has been proposed as
[...] Read more.
Background/Objectives: Traditional semen analysis techniques frequently result in incorrect male infertility diagnoses, despite advancements in assisted reproductive technology (ART). Reduced fertilization potential, decreased embryo development, and lower pregnancy success rates are associated with elevated DNA Fragmentation Index (DFI), which has been proposed as a diagnostic indicator of sperm DNA integrity. Improving reproductive outcomes requires incorporating DFI into predictive models due to its diagnostic importance. Methods: In this study, semen samples were stratified into low and high DFI groups across two datasets: the “Reference” dataset (162 samples) containing sperm motility (A, B, and C), total sperm count, and morphology percentage, and the “ORP” dataset (37 samples) with the same features plus oxidation-reduction potential (ORP). We trained and evaluated four machine learning (ML) models—Logistic Regression, Support Vector Machines (SVM), Bernoulli Naive Bayes (BNB), and Random Forest (RF)- using three feature subsets and three preprocessing techniques (Robust Scaling, Min-Max Scaling, and Standard Scaling). Results: Feature subset selection had a significant impact on model performance, with the full feature set (X_all) yielding the best results, and the combination of Robust and MinMax scaling forming the most effective preprocessing pipeline. Conclusions: ORP proved to be a critical feature, enhancing model generalization and prediction performance. These findings suggest that data enrichment, particularly with ORP, could enable the development of ML frameworks that improve prognostic precision and patient outcomes in ART.
Full article

Graphical abstract
Open AccessReview
Dermatogenomic Insights into Systemic Diseases: Implications for Primary and Preventive Medicine
by
Yu Xuan Jin, David Alexandru Anton, Ming Yuan Zhou, Amir Pourghadiri and Chaocheng Liu
DNA 2026, 6(1), 2; https://doi.org/10.3390/dna6010002 - 6 Jan 2026
Abstract
►▼
Show Figures
The emerging field of dermatogenomics, which examines visible dermatologic phenotypes alongside their polygenic factors, offers insights for early disease recognition and initiation of preventative measures. This review explores key dermatologic manifestations serving as clinical markers of systemic diseases, emphasizing cardiovascular, autoimmune, neuropsychiatric, metabolic/endocrine,
[...] Read more.
The emerging field of dermatogenomics, which examines visible dermatologic phenotypes alongside their polygenic factors, offers insights for early disease recognition and initiation of preventative measures. This review explores key dermatologic manifestations serving as clinical markers of systemic diseases, emphasizing cardiovascular, autoimmune, neuropsychiatric, metabolic/endocrine, and cancer-related conditions. Importantly, the pathogenesis of certain skin conditions including psoriasis, atopic dermatitis, vitiligo, and hidradenitis suppurativa is linked to systemic disease through shared genetic and epigenetic mechanisms. The diagnostic markers for these integumentary diseases are discussed alongside their shared mechanisms to systemic diseases, highlighting the clinical manifestation typically seen in primary care settings. This narrative review integrates dermatology with genomics, primary care, preventative care, public health, and internal medicine perspectives, underscoring the importance of an interdisciplinary and collaborative approach to patient care. Lastly, this review advocates for standardized dermatogenomic screening thresholds, inclusivity and expansion of genomic datasets, and the leverage of artificial intelligence and multi-omic technologies in preventative healthcare.
Full article

Figure 1
Open AccessArticle
Integrating Genome Mining and Untargeted Metabolomics to Uncover the Chemical Diversity of Streptomyces galbus I339, a Strain from the Unique Brazilian Caatinga Biome
by
Edson Alexandre Nascimento-Silva, André Luiz Leocádio de Souza Matos, Thalisson Amorim de Souza, Anauara Lima e Silva, Lucas Silva Abreu, Monalisa Mota Merces, Renata Priscila Almeida Silva, Ubiratan Ribeiro da Silva Filho, Adrielly Silva Albuquerque de Andrade, Josean Fechine Tavares, Celso José Bruno de Oliveira, Patrícia Emilia Naves Givisiez, Demetrius Antonio Machado de Araújo, Valnês da Silva Rodrigues-Junior and Samuel Paulo Cibulski
DNA 2026, 6(1), 1; https://doi.org/10.3390/dna6010001 - 24 Dec 2025
Abstract
►▼
Show Figures
Background/Objectives: The escalating antimicrobial resistance crisis underscores the urgent need to explore underexplored ecological niches as reservoirs of novel bioactive compounds. The Brazilian Caatinga, a unique semi-arid biome, represents a promising reservoir for microbial discovery. Methods: In this study, we report
[...] Read more.
Background/Objectives: The escalating antimicrobial resistance crisis underscores the urgent need to explore underexplored ecological niches as reservoirs of novel bioactive compounds. The Brazilian Caatinga, a unique semi-arid biome, represents a promising reservoir for microbial discovery. Methods: In this study, we report the polyphasic characterization of Streptomyces galbus I339, a strain isolated from Caatinga soil. Whole-genome sequencing and phylogenomic analysis confirmed its taxonomic identity. In silico mining of the genome was conducted to assess biosynthetic potential. This genetic promise was experimentally validated through an integrated metabolomic approach, including liquid chromatography-tandem mass spectrometry (LC-MS/MS), nuclear magnetic resonance (NMR) spectroscopy, and gas chromatography-mass spectrometry (GC-MS) profiling. The anti-mycobacterial activity of the crude extract was evaluated against Mycobacterium tuberculosis. Results: The strain S. galbus I339 possesses a 7.55 Mbp genome with a high GC content (73.17%). Genome mining uncovered a remarkable biosynthetic potential, with 45 biosynthetic gene clusters (BGCs) predicted, including those for known antibiotics like actinomycins, as well as numerous orphan clusters. Genome mining uncovered a remarkable biosynthetic potential, with 45 biosynthetic gene clusters (BGCs) predicted, including those for known antibiotics like actinomycins, as well as numerous orphan clusters. Metabolomic analyses confirmed the production of actinomycins and identified abundant diketopiperazines. Furthermore, the crude extract exhibited antimycobacterial activity, with a potent MIC of 0.625 µg/mL. Conclusions: The convergence of genomic and metabolomic data not only validates the expression of a fraction of this strain’s biosynthetic arsenal but also highlights a significant untapped potential, with the majority of BGCs remaining silent under the tested conditions. Our work establishes S. galbus I339 as a compelling candidate for biodiscovery and underscores the value of integrating genomics and metabolomics to unlock the chemical diversity of microbes from extreme environments.
Full article

Graphical abstract
Open AccessEditor’s ChoiceReview
Nutrigenomics and Epigenetic Regulation in Poultry: DNA-Based Mechanisms Linking Diet to Performance and Health
by
Muhammad Naeem and Arjmand Fatima
DNA 2025, 5(4), 60; https://doi.org/10.3390/dna5040060 - 18 Dec 2025
Cited by 1
Abstract
In animals and humans, nutrients influence signaling cascades, transcriptional programs, chromatin dynamics, and mitochondrial function, collectively shaping traits related to growth, immunity, reproduction, and stress resilience. This review synthesizes evidence supporting nutrient-mediated regulation of DNA methylation, histone modifications, non-coding RNAs, and mitochondrial biogenesis,
[...] Read more.
In animals and humans, nutrients influence signaling cascades, transcriptional programs, chromatin dynamics, and mitochondrial function, collectively shaping traits related to growth, immunity, reproduction, and stress resilience. This review synthesizes evidence supporting nutrient-mediated regulation of DNA methylation, histone modifications, non-coding RNAs, and mitochondrial biogenesis, and emphasizes their integration within metabolic and developmental pathways. Recent advances in epigenome-wide association studies (EWAS), single-cell multi-omics, and systems biology approaches have revealed how diet composition and timing can reprogram gene networks, sometimes across generations. Particular attention is given to central metabolic regulators (e.g., PPARs, mTOR) and to interactions among methyl donors, fatty acids, vitamins, and trace elements that maintain genomic stability and metabolic homeostasis. Nutrigenetic evidence further shows how genetic polymorphisms (SNPs) in loci such as IGF-1, MSTN, PPARs, and FASN alter nutrient responsiveness and influence traits like feed efficiency, body composition, and egg quality, information that can be exploited via marker-assisted or genomic selection. Mitochondrial DNA integrity and oxidative capacity are key determinants of feed conversion and energy efficiency, while dietary antioxidants and mitochondria-targeted nutrients help preserve bioenergetic function. The gut microbiome acts as a co-regulator of host gene expression through metabolite-mediated epigenetic effects, linking diet, microbial metabolites (e.g., SCFAs), and host genomic responses via the gut–liver axis. Emerging tools such as whole-genome and transcriptome sequencing, EWAS, integrated multi-omics, and CRISPR-based functional studies are transforming the field and enabling DNA-informed precision nutrition. Integrating genetic, epigenetic, and molecular data will enable genotype-specific feeding strategies, maternal and early-life programming, and predictive models that enhance productivity, health, and sustainability in poultry production. Translating these molecular insights into practice offers pathways to enhance animal welfare, reduce environmental impact, and shift nutrition from empirical feeding toward mechanistically informed precision approaches.
Full article
(This article belongs to the Special Issue Epigenetics and Environmental Exposures)
►▼
Show Figures

Figure 1
Open AccessEditor’s ChoiceReview
The CB2 Receptor in Immune Regulation and Disease: Genetic Architecture, Epigenetic Control, and Emerging Therapeutic Strategies
by
Hilal Kalkan and Nicolas Flamand
DNA 2025, 5(4), 59; https://doi.org/10.3390/dna5040059 - 11 Dec 2025
Abstract
►▼
Show Figures
The cannabinoid receptor type 2 (CB2) is increasingly recognized as a crucial regulator of neuroimmune balance in the brain. In addition to its well-established role in immunity, the CB2 receptor has been identified in specific populations of neurons and glial
[...] Read more.
The cannabinoid receptor type 2 (CB2) is increasingly recognized as a crucial regulator of neuroimmune balance in the brain. In addition to its well-established role in immunity, the CB2 receptor has been identified in specific populations of neurons and glial cells throughout various brain regions, and its expression is dynamically increased during inflammatory and neuropathological conditions, positioning it as a potential non-psychoactive target for modifying neurological diseases. The expression of the CB2 gene (CNR2) is finely tuned by epigenetic processes, including promoter CpG methylation, histone modifications, and non-coding RNAs, which regulate receptor availability and signaling preferences in response to stress, inflammation, and environmental factors. CB2 signaling interacts with TRP channels (such as TRPV1), nuclear receptors (PPARγ), and orphan G Protein-Coupled Receptors (GPCRs, including GPR55 and GPR18) within the endocannabinoidome (eCBome), influencing microglial characteristics, cytokine production, and synaptic activity. We review how these interconnected mechanisms affect neurodegenerative and neuropsychiatric disorders, underscore the species- and cell-type-specificities that pose challenges for translation, and explore emerging strategies, including selective agonists, positive allosteric modulators, and biased ligands, that leverage the signaling adaptability of the CB2 receptor while reducing central effects mediated by the CB1 receptor. This focus on the neuro-centric perspective repositions the CB2 receptor as an epigenetically informed, context-dependent hub within the eCBome, making it a promising candidate for precision therapies in conditions featuring neuroinflammation.
Full article

Figure 1
Open AccessEditor’s ChoiceArticle
First Metagenomic Shotgun Sequencing Report on the Microbiome of Local Goat and Sheep Raw Milk in Benin for Dairy Valorization
by
Yvette Adje, Philippe Sessou, Konstantinos Tegopoulos, Yaovi Mahuton Gildas Hounmanou, Nikistratos Siskos, Ioanna Farmakioti, Paulin Azokpota, Souaïbou Farougou, Lamine Baba-Moussa, George Skavdis and Maria E. Grigoriou
DNA 2025, 5(4), 58; https://doi.org/10.3390/dna5040058 - 4 Dec 2025
Cited by 1
Abstract
►▼
Show Figures
Background/Objectives: Goat and sheep farming is an important agro-economic resource in Benin. However, their milk is both underutilized and insufficiently characterized, which limits the development of innovative dairy products and raises concerns about its safety. Against this backdrop, our pioneering study set
[...] Read more.
Background/Objectives: Goat and sheep farming is an important agro-economic resource in Benin. However, their milk is both underutilized and insufficiently characterized, which limits the development of innovative dairy products and raises concerns about its safety. Against this backdrop, our pioneering study set out to investigate, for the first time in Benin and using an advanced metagenomic approach, the microbial diversity present in goat and sheep raw milk. The aim was to lay the groundwork for safer and more efficient dairy valorization. Methods: To achieve this, metagenomic DNA was extracted from 20 pooled milk samples representing both animal species, followed by shotgun sequencing. Results: Analyses revealed seven dominant phyla: Bacillota (17.44–27.23%), Pseudomonadota (12.39–15.55%), Campylobacterota (3.65–5.29%), Actinomycetota (1.47–6.03%), Spirochaetota (1.14–2.02%), Apicomplexa (0.28–0.50%), and Bacteroidota (0.17–0.22%) in the raw milk of both species. However, their proportions differ. Bacillota, which was the most dominant in both types of milk, was significantly more abundant in goat (27.23 ± 5.33) than in sheep milk (17.44 ± 8.44). In sheep milk, Enterobacteriaceae (11.36 ± 5.79) were the most predominant family, followed by Streptococcaceae (5.57 ± 2.29) and Staphylococcaceae (4.51 ± 3.63). Goat milk, on the other hand, presents a different hierarchy. Streptococcaceae (6.65 ± 2.19) and Staphylococcaceae (6.43 ± 2.33) were the most abundant families, surpassing Enterobacteriaceae (5.33 ± 1.66). The genus Escherichia was the most abundant in sheep milk (6.18 ± 5.33). The genera Staphylococcus (4.50 ± 3.63) and Streptococcus (5.05 ± 1.98) were also present. In contrast, in goat milk, the genera Streptococcus (6.54 ± 2.35) and Staphylococcus (6.42 ± 2.32) were the most dominant, while the average abundance of Escherichia was much lower (1.98 ± 1.28). In terms of species, Sheep milk was dominated by Escherichia coli (6.14 ± 5.28) and Staphylococcus aureus (5.17 ± 2.28) while Klebsiella pneumoniae (2.82 ± 1.72), Streptococcus pneumoniae (1.92 ± 1.36), and Campylobacter coli (1.52 ± 1.27) were also found. In addition to a relatively high abundance of Staphylococcus aureus (6.40 ± 2.45), goat milk was characterized by the presence of Corynebacterium praerotentium (5.32 ± 2.28) and Clostridium perfringens (3.39 ± 2.09). Additional pathogens identified included Clostridioides difficile (1.17–2.00%), Clostridium botulinum (0.27–0.43%), Listeria monocytogenes, Mycobacterium tuberculosis, Helicobacter pylori (0.36–0.62%), Salmonella enterica (0.22–0.26%). As for fungi, Ascomycota were predominant, with the presence of Aspergillus fumigatus, Saccharomyces cerevisiae, Trichophyton mentagrophytes, and Candida auris. Moreover, lactic acid bacteria with technological interest such as Oenococcus oeni (0.60–0.97%), Levilactobacillus namurensis (0.25–0.44%), Lactobacillus agrestimuris, and Lacticaseibacillus rhamnosus were also detected. Conclusions: These findings provide essential insights into the technological potential and health risks associated with these milks, which are key to developing safer and more efficient local dairy value chains.
Full article

Graphical abstract
Open AccessEditor’s ChoiceReview
CRISPR-Based Transcriptional Regulation: Technologies, Applications, and Future Directions
by
Mira A. Srinivasa and Mario Escobar
DNA 2025, 5(4), 57; https://doi.org/10.3390/dna5040057 - 1 Dec 2025
Cited by 1
Abstract
CRISPR-based transcriptional regulation technologies, including CRISPR activation (CRISPRa) and CRISPR interference (CRISPRi), offer precise and programmable control over gene expression, representing a major advance in gene and epigenetic therapy. CRISPRa uses nuclease-inactive Cas proteins fused to transcriptional activators to upregulate target genes, while
[...] Read more.
CRISPR-based transcriptional regulation technologies, including CRISPR activation (CRISPRa) and CRISPR interference (CRISPRi), offer precise and programmable control over gene expression, representing a major advance in gene and epigenetic therapy. CRISPRa uses nuclease-inactive Cas proteins fused to transcriptional activators to upregulate target genes, while CRISPRi employs repressor domains for gene silencing. Preclinical studies have demonstrated the efficacy of CRISPRa/i in models of metabolic, neurological, muscular, and oncological diseases. Notably, CRISPRi-based therapies have entered clinical trials for conditions like hepatitis B and muscular dystrophy, showing encouraging safety and efficacy profiles. Despite ongoing challenges related to delivery efficiency, immunogenicity, and off-target activity, innovations in protein engineering and guide RNA design are rapidly enhancing the precision and safety of these technologies. Overall, CRISPRa and CRISPRi are poised to transform the treatment of genetic and epigenetic disorders, with continued optimization expected to accelerate their clinical adoption and broaden their therapeutic impact.
Full article
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Agriculture, DNA, Genes, IJMS, Plants, Crops
Genetic Engineering in Agriculture, 2nd Edition
Topic Editors: Amy Klocko, Jianjun Chen, Haiwei LuDeadline: 30 April 2026
Topic in
Animals, CIMB, DNA, Genes, IJMS, Biology
Advances in Molecular Genetics and Breeding of Cattle, Sheep, and Goats, 2nd Edition
Topic Editors: Xiukai Cao, Hui Li, Huitong ZhouDeadline: 31 December 2026
Topic in
Agriculture, Agronomy, Crops, Genes, Plants, DNA
Vegetable Breeding, Genetics and Genomics, 2nd Volume
Topic Editors: Padma Nimmakayala, Georgia Ntatsi, Umesh K. Reddy, Yan R. TomasonDeadline: 5 February 2027
Topic in
Brain Sciences, CIMB, Epigenomes, Genes, IJMS, DNA
Genetics and Epigenetics of Substance Use Disorders
Topic Editors: Aleksandra Suchanecka, Anna Maria Grzywacz, Kszysztof ChmielowiecDeadline: 14 April 2027

Conferences
Special Issues
Special Issue in
DNA
Epigenetics and Environmental Exposures
Guest Editors: Benoît Miotto, Tina Bianco-MiottoDeadline: 25 May 2026
Special Issue in
DNA
Molecular Structure and Dynamics of DNA/RNA Helices
Guest Editor: Mauricio Alcolea PalafoxDeadline: 15 July 2026











