Segmentation-Based vs. Regression-Based Biomarker Estimation: A Case Study of Fetus Head Circumference Assessment from Ultrasound Images
Abstract
:1. Introduction
2. Related Works
2.1. Fetus Head Circumference Estimation
2.2. Segmentation-Free Approaches for Biomarker Estimation
3. Methodological Framework
3.1. Head Circumference Estimation Based on Segmentation
3.1.1. CNN Segmentation Model
3.1.2. Post-Processing of Segmentation Results
3.1.3. HC Computation Based on Segmentation Results
3.2. Head Circumference Estimation Using Regression CNN
3.2.1. Regression CNN Model
3.2.2. Loss Functions
3.3. Model Configuration
4. Experimental Settings
4.1. Dataset and Pre-Processing
4.2. Experiment Configuration
4.3. Evaluation Metrics
5. Results and Discussion
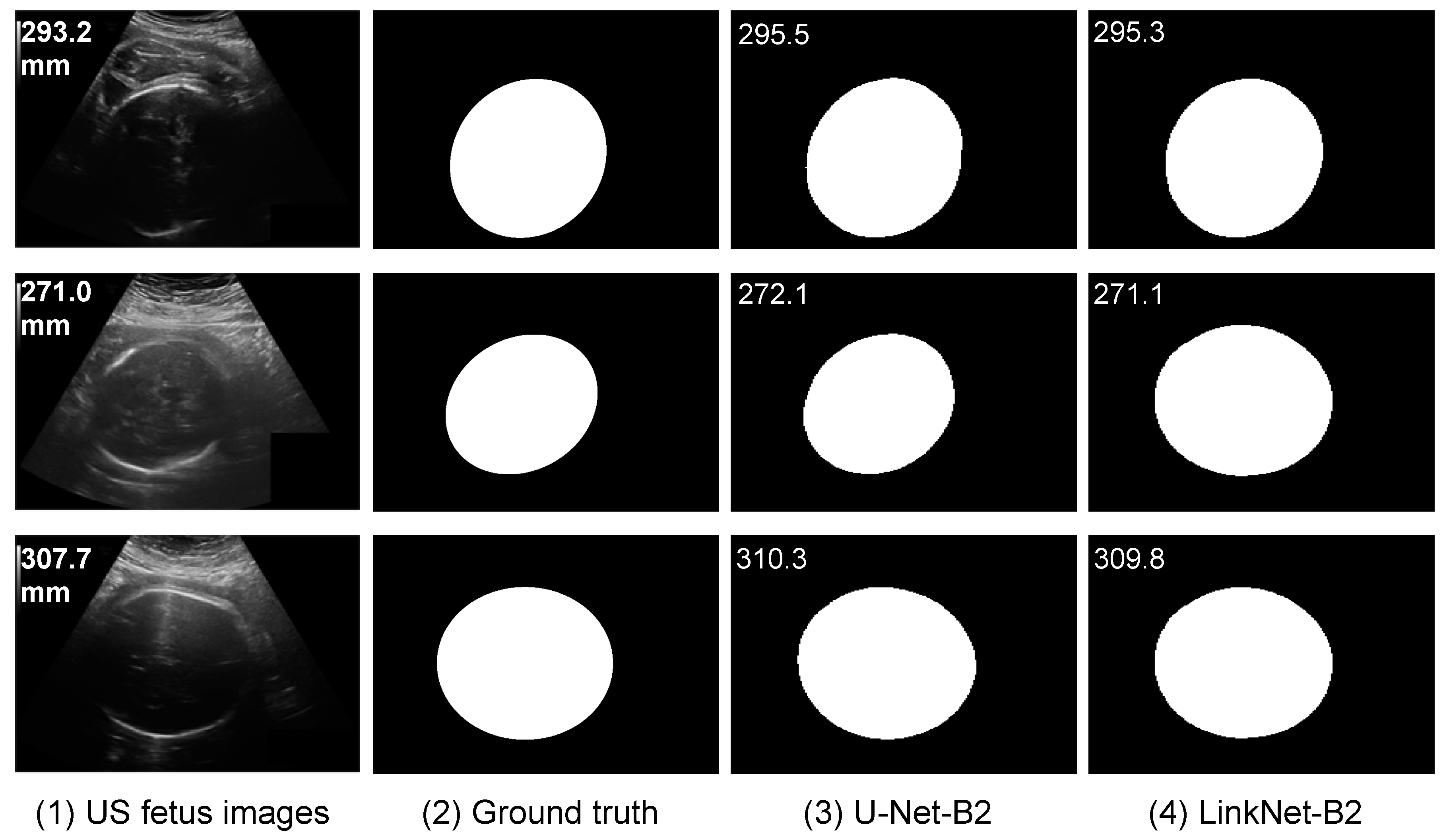
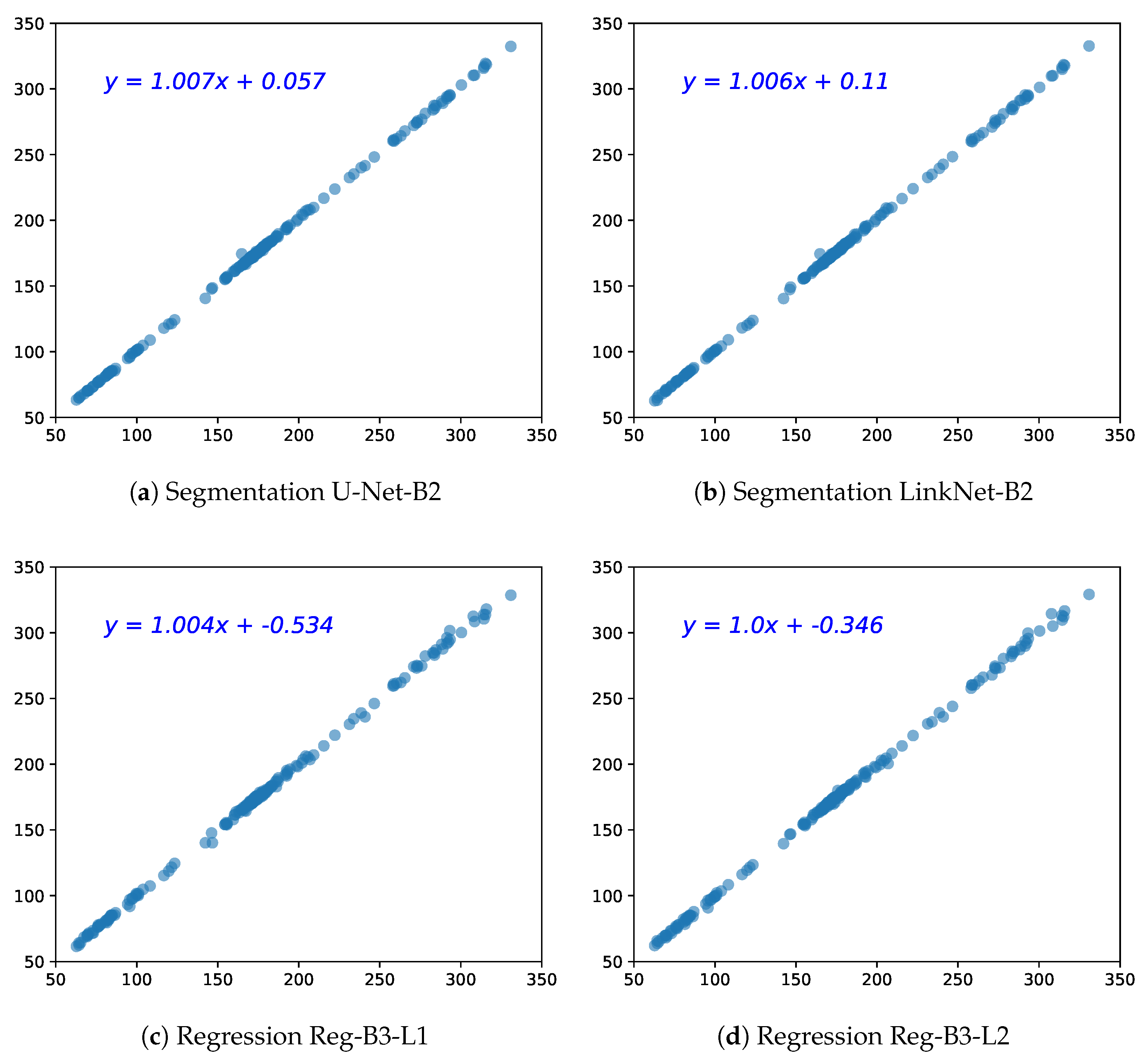
5.1. HC Estimation Based on Segmentation
5.2. HC Estimation Based on Regression CNN
5.3. Interpretability of Regression CNN Result
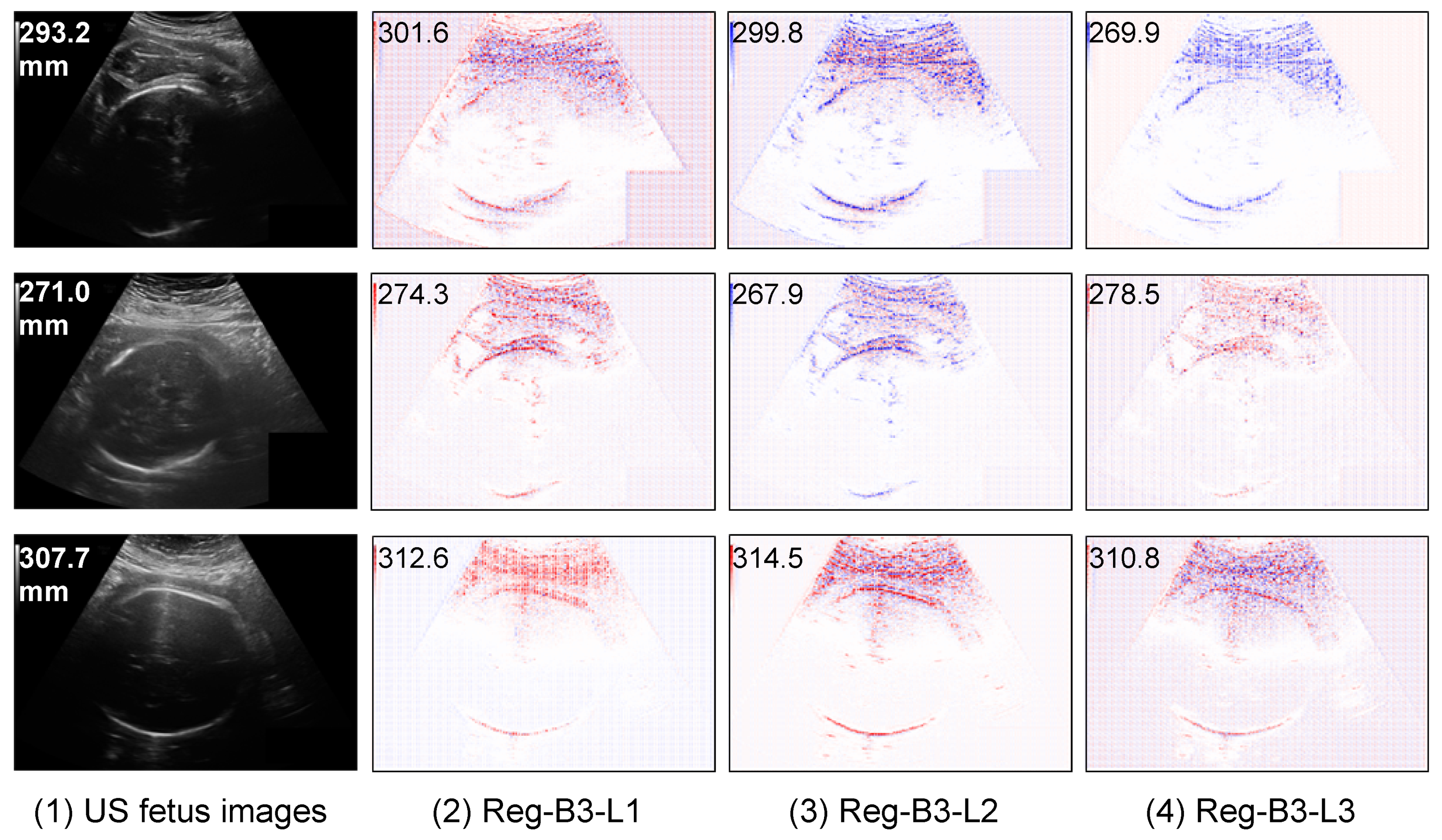
5.3.1. Saliency Maps of Regression CNN Results on HC
5.3.2. Saliency Maps on Outlier Analysis
5.4. Comparison of Segmentation CNN vs. Regression CNN
5.5. Memory Usage and Computational Efficiency
5.6. Comparison of HC Estimation with State-of-the-Art
6. Conclusions and Future Work
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| HC | Head circumference |
| US | Ultrasound |
| CT | Computed tomography |
| MR | Magnetic resonance |
| CNN | Convolutional neural networks |
| pp | post processing |
| EF | Ellipse fitting |
| MAE | Mean absolute error |
| MSE | Mean square error |
| HL | Huber loss |
| DI | Dice index |
| HD | Hausdorff distance |
| ASSD | Average symmetric surface distance |
| PMAE | Percentage mean absolute error |
| LRP | Layer-wise relevance propagation |
References
- Van den Heuvel, T.L.A.; de Bruijn, D.; de Korte, C.L.; Ginneken, B.V. Automated measurement of fetal head circumference using 2D ultrasound images [Data set]. Zenodo 2018. [Google Scholar] [CrossRef]
- Sarris, I.; Ioannou, C.; Chamberlain, P.; Ohuma, E.; Roseman, F.; Hoch, L.; Altman, D.; Papageorghiou, A.; INTERGROWTH-21st. Intra-and interobserver variability in fetal ultrasound measurements. Ultrasound Obstet. Gynecol. 2012, 39, 266–273. [Google Scholar] [CrossRef] [PubMed]
- Petitjean, C.; Dacher, J.N. A review of segmentation methods in short axis cardiac MR images. Med. Image Anal. 2011, 15, 169–184. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Blanc-Durand, P.; Campedel, L.; Mule, S.; Jegou, S.; Luciani, A.; Pigneur, F.; Itti, E. Prognostic value of anthropometric measures extracted from whole-body CT using deep learning in patients with non-small-cell lung cancer. Eur. Radiol. 2020, 13, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Zhen, X.; Wang, Z.; Islam, A.; Bhaduri, M.; Chan, I.; Li, S. Direct volume estimation without segmentation. In Proceedings of the SPIE Medical Imaging 2015, Orlando, FL, USA, 21–26 February 2015. [Google Scholar]
- Hussain, M.A.; Hamarneh, G.; O’Connell, T.W.; Mohammed, M.F.; Abugharbieh, R. Segmentation-free estimation of kidney volumes in CT with dual regression forests. In International Workshop on Machine Learning in Medical Imaging; Springer: Athens, Greece, 2016; pp. 156–163. [Google Scholar]
- Pang, S.; Leung, S.; Nachum, I.B.; Feng, Q.; Li, S. Direct automated quantitative measurement of spine via cascade amplifier regression network. In International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer: Granada, Spain, 2018; pp. 940–948. [Google Scholar]
- Luo, G.; Dong, S.; Wang, W.; Wang, K.; Cao, S.; Tam, C.; Zhang, H.; Howey, J.; Ohorodnyk, P.; Li, S. Commensal correlation network between segmentation and direct area estimation for bi-ventricle quantification. Med. Image Anal. 2020, 59, 101591. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Yang, G.; Zhao, S.; Zhang, Y.; Ghista, D.; Zhang, H.; Li, S. Direct quantification of coronary artery stenosis through hierarchical attentive multi-view learning. IEEE Trans. Med. Imaging 2020, 39, 4322–4334. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Petitjean, C.; Lopez, P.; Ainouz, S. Direct estimation of fetal head circumference from ultrasound images based on regression CNN. In Medical Imaging with Deep Learning; PMLR: Montreal, QC, Canada, 2020. [Google Scholar]
- Li, J.; Wang, Y.; Lei, B.; Cheng, J.Z.; Qin, J.; Wang, T.; Li, S.; Ni, D. Automatic fetal head circumference measurement in ultrasound using random forest and fast ellipse fitting. IEEE J. Biomed. Health Inform. 2017, 22, 215–223. [Google Scholar] [CrossRef] [PubMed]
- Lu, W.; Tan, J.; Floyd, R. Automated fetal head detection and measurement in ultrasound images by iterative randomized hough transform. Ultrasound Med. Biol. 2005, 31, 929–936. [Google Scholar] [CrossRef] [PubMed]
- Jardim, S.M.; Figueiredo, M.A. Segmentation of fetal ultrasound images. Ultrasound Med. Biol. 2005, 31, 243–250. [Google Scholar] [CrossRef] [PubMed]
- Van den Heuvel, T.L.A.; de Bruijn, D.; de Korte, C.L.; Ginneken, B.V. Automated measurement of fetal head circumference using 2D ultrasound images. PLoS ONE 2018, 13, e0200412. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.P.; Lee, S.M.; Kwon, J.Y.; Park, Y.; Kim, K.C.; Seo, J.K. Automatic evaluation of fetal head biometry from ultrasound images using machine learning. Physiol. Meas. 2019, 40, 065009. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Budd, S.; Sinclair, M.; Khanal, B.; Matthew, J.; Lloyd, D.; Gomez, A.; Toussaint, N.; Robinson, E.C.; Kainz, B. Confident Head Circumference Measurement from Ultrasound with Real-time Feedback for Sonographers. In MICCAI; Springer: Cham, Switzerland, 2019; pp. 683–691. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer: Munich, Germany, 2015; pp. 234–241. [Google Scholar]
- Sobhaninia, Z.; Rafiei, S.; Emami, A.; Karimi, N.; Najarian, K.; Samavi, S.; Soroushmehr, S.R. Fetal Ultrasound Image Segmentation for Measuring Biometric Parameters Using Multi-Task Deep Learning. In Proceedings of the 41st Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Berlin, Germany, 23–27 July 2019; pp. 6545–6548. [Google Scholar]
- Fiorentino, M.C.; Moccia, S.; Capparuccini, M.; Giamberini, S.; Frontoni, E. A regression framework to head-circumference delineation from US fetal images. Comput. Methods Programs Biomed. 2021, 198, 105771. [Google Scholar] [CrossRef] [PubMed]
- Moccia, S.; Fiorentino, M.C.; Frontoni, E. Mask-R2 CNN: A distance-field regression version of Mask-RCNN for fetal-head delineation in ultrasound images. Int. J. Comput. Assist. Radiol. Surg. 2021, 16, 1711–1718. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Liang, W.; Wang, Y.; Li, S.; Pei, M. 3D head pose estimation with convolutional neural network trained on synthetic images. In Proceedings of the IEEE International Conference on Image Processing (ICIP), Phoenix, AZ, USA, 25–28 September 2016; pp. 1289–1293. [Google Scholar]
- Sun, Y.; Wang, X.; Tang, X. Deep convolutional network cascade for facial point detection. In Proceedings of the IEEE CVPR, Portland, OR, USA, 23–28 June 2013; pp. 3476–3483. [Google Scholar]
- Toshev, A.; Szegedy, C. Deeppose: Human pose estimation via deep neural networks. In Proceedings of the IEEE CVPR, Columbus, OH, USA, 23–28 June 2014; pp. 1653–1660. [Google Scholar]
- Zhou, Z.; Siddiquee, M.M.R.; Tajbakhsh, N.; Liang, J. Unet++: A nested u-net architecture for medical image segmentation. In Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support; Springer: Granada, Spain, 2018; pp. 3–11. [Google Scholar]
- Jha, D.; Riegler, M.A.; Johansen, D.; Halvorsen, P.; Johansen, H.D. Doubleu-net: A deep convolutional neural network for medical image segmentation. In Proceedings of the 2020 IEEE 33rd International symposium on computer-based medical systems (CBMS), Rochester, MN, USA, 28–30 July 2020; pp. 558–564. [Google Scholar]
- Lin, T.Y.; Dollár, P.; Girshick, R.; He, K.; Hariharan, B.; Belongie, S. Feature pyramid networks for object detection. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 2117–2125. [Google Scholar]
- Chaurasia, A.; Culurciello, E. Linknet: Exploiting encoder representations for efficient semantic segmentation. In Proceedings of the 2017 IEEE Visual Communications and Image Processing (VCIP), St. Petersburg, FL, USA, 10–13 December 2017; pp. 1–4. [Google Scholar]
- Zhao, H.; Shi, J.; Qi, X.; Wang, X.; Jia, J. Pyramid scene parsing network. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 2881–2890. [Google Scholar]
- Wacker, J.; Ladeira, M.; Nascimento, J. Transfer Learning for Brain Tumor Segmentation. In International MICCAI Brainlesion Workshop; Springer: Cham, Switzerland, 2020. [Google Scholar]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv 2015, arXiv:1409.1556. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Tan, M.; Le, Q. Efficientnet: Rethinking model scaling for convolutional neural networks. In Proceedings of the International Conference on Machine Learning, Long Beach, CA, USA, 10–15 June 2019; pp. 6105–6114. [Google Scholar]
- Ma, J.; Chen, J.; Ng, M.; Huang, R.; Li, Y.; Li, C.; Yang, X.; Martel, A.L. Loss odyssey in medical image segmentation. Med. Image Anal. 2021, 71, 102035. [Google Scholar] [CrossRef] [PubMed]
- Barnard, R.W.; Pearce, K.; Schovanec, L. Inequalities for the perimeter of an ellipse. J. Math. Anal. Appl. 2001, 260, 295–306. [Google Scholar] [CrossRef] [Green Version]
- Huang, G.; Liu, Z.; Van Der Maaten, L.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 4700–4708. [Google Scholar]
- Chollet, F. Xception: Deep learning with depthwise separable convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 1251–1258. [Google Scholar]
- Howard, A.G.; Zhu, M.; Chen, B.; Kalenichenko, D.; Wang, W.; Weyand, T.; Andreetto, M.; Adam, H. Mobilenets: Efficient convolutional neural networks for mobile vision applications. arXiv 2017, arXiv:1704.04861. [Google Scholar]
- Szegedy, C.; Vanhoucke, V.; Ioffe, S.; Shlens, J.; Wojna, Z. Rethinking the inception architecture for computer vision. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 2818–2826. [Google Scholar]
- Deng, J.; Dong, W.; Socher, R.; Li, L.J.; Li, K.; Fei-Fei, L. Imagenet: A large-scale hierarchical image database. In Proceedings of the 2009 IEEE Conference on Computer Vision and Pattern Recognition, Miami, FL, USA, 20–25 June 2009; pp. 248–255. [Google Scholar]
- Lathuilière, S.; Mesejo, P.; Alameda-Pineda, X.; Horaud, R. A comprehensive analysis of deep regression. IEEE Trans. Pattern Anal. Mach. Intell. 2019, 42, 2065–2081. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yakubovskiy, P. Segmentation Models, GitHub. 2019. Available online: https://github.com/qubvel/segmentation_models (accessed on 31 January 2020).
- Zhang, J.; Petitjean, C.; Yger, F.; Ainouz, S. Explainability for regression CNN in fetal head circumference estimation from ultrasound images. In Interpretable and Annotation-Efficient Learning for Medical Image Computing; Springer: Lima, Peru, 2020; pp. 73–82. [Google Scholar]
- Bach, S.; Binder, A.; Montavon, G.; Klauschen, F.; Müller, K.R.; Samek, W. On pixel-wise explanations for non-linear classifier decisions by layer-wise relevance propagation. PLoS ONE 2015, 10, e0130140. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morch, N.J.; Kjems, U.; Hansen, L.K.; Svarer, C.; Law, I.; Lautrup, B.; Strother, S.; Rehm, K. Visualization of neural networks using saliency maps. In Proceedings of the IEEE International Conference on Neural Networks, Perth, WA, Australia, 27 November–1 December 1995; Volume 4, pp. 2085–2090. [Google Scholar]
- Dobrescu, A.; Valerio Giuffrida, M.; Tsaftaris, S.A. Understanding deep neural networks for regression in leaf counting. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops, Long Beach, CA, USA, 16–20 June 2019. [Google Scholar]






| Segmentation Models | # Parameters (M) | Regression Models | # Parameters (M) |
|---|---|---|---|
| Original U-Net | 31.06 | Reg-B1 | 15.15 |
| U-Net-B1, B2, B3 | 23.75, 32.51, 14.23 | Reg-B2 | 23.63 |
| DoubleU-Net | 29.29 | Reg-B3 | 76.73 |
| U-Net++ B1, B2, B3 | 24.15, 34.34, 16.03 | Reg-B4 | 70.04 |
| FPN-B1, B2, B3 | 17.59, 26.89, 10.77 | Reg-B5 | 20.91 |
| LinkNet-B1, B2, B3 | 20.32, 28.73, 10.15 | Reg-B6 | 3.26 |
| PSPNet-B1, B2, B3 | 21.55, 17.99, 9.41 | Reg-B7 | 21.82 |
| Method | DI ↑ (%) | HD ↓ (mm) | ASSD ↓ (mm) | MAE ↓ (mm) w/o pp | MAE (mm) w pp | MAE (px) w/o pp | MAE (px) w pp | PMAE ↓ (%) w/o pp | PMAE (%) w pp |
|---|---|---|---|---|---|---|---|---|---|
| U-Net-original | 98.5 ± | 1.56 ± | 0.35 ± | 1.55 ± | 1.23 ± | 11.83 ± | 9.11 ± | 1.04 ± | 0.75 ± |
| DoubleU-Net | 98.7 ± | 1.14 ± | 0.29 ± | 2.60 ± | 2.59 ± | 18.94 ± | 18.76 ± | 1.58 ± | 1.56 ± |
| U-Net-B1 | 98.6 ± | 1.16 ± | 0.31 ± | 1.31 ± | 1.21 ± | 9.99 ± | 8.98 ± | 0.85 ± | 0.74 ± |
| U-Net-B2 | 98.8 ± | 1.09 ± | 0.27 ± | 1.16 ± | 1.08 ± | 8.69 ± | 7.87 ± | 0.74 ± | 0.65 ± |
| U-Net-B3 | 98.7 ± | 1.11 ± | 0.29 ± | 1.34 ± | 1.32 ± | 10.23 ± | 9.94 ± | 0.86 ± | 0.84 ± |
| U-Net++ B1 | 98.5 ± | 1.29 ± | 0.31 ± | 2.03 ± | 1.3 ± | 16.95 ± | 9.92 ± | 1.51 ± | 0.87 ± |
| U-Net++ B2 | 98.7 ± | 1.24 ± | 0.29 ± | 1.74 ± | 1.15 ± | 12.65 ± | 8.63 ± | 1.16 ± | 0.72 ± |
| U-Net++ B3 | 98.7 ± | 1.17 ± | 0.29 ± | 2.32 ± | 1.19 ± | 19.08 ± | 8.91 ± | 1.57 ± | 0.76 ± |
| FPN-B1 | 98.6 ± | 1.28 ± | 0.32 ± | 1.44 ± | 1.29 ± | 11.17 ± | 9.70 ± | 0.99 ± | 0.80 ± |
| FPN-B2 | 98.7 ± | 1.18 ± | 0.30 ± | 1.38 ± | 1.26 ± | 10.35 ± | 9.19 ± | 1.90 ± | 0.76 ± |
| FPN-B3 | 98.7 ± 1 | 1.19 ± | 0.30 ± | 1.46 ± | 1.39 ± | 11.09 ± | 10.33 ± | 0.94 ± | 0.86 ± |
| LinkNet-B1 | 98.6 ± | 1.31 ± | 0.33 ± | 1.46 ± | 1.32 ± | 11.32 ± | 9.91 ± | 0.98 ± | 0.83 ± |
| LinkNet-B2 | 98.7 ± | 1.12 ± | 0.30 ± | 1.19 ± | 1.15 ± | 8.86 ± | 8.45 ± | 0.73 ± | 0.69 ± |
| LinkNet-B3 | 98.6 ± 1 | 1.15 ± | 0.31 ± | 1.37 ± | 1.29 ± | 10.55 ± | 9.70 ± | 0.89 ± | 0.79 ± |
| PSPNet-B1 | 98.6 ± | 2.01 ± | 0.38 ± | 3.07 ± | 1.32 ± | 22.38 ± | 9.84 ± | 2.21 ± | 0.81 ± |
| PSPNet-B2 | 98.8 ± | 1.42 ± | 0.31 ± | 1.66 ± | 1.20 ± | 11.98 ± | 8.75 ± | 1.07 ± | 0.72 ± |
| PSPNet-B3 | 98.7 ± | 1.12 ± | 0.32 ± | 1.38 ± | 1.29 ± | 10.59 ± | 9.64 ± | 0.93 ± | 0.81 ± |
| Model | MAE (mm) | MAE (px) | PMAE (%) |
|---|---|---|---|
| Reg-B1-L1 | 3.04 ± | 22.41 ± | 1.94 ± |
| Reg-B2-L1 | 3.24 ± | 24.11 ± | 2.14 ± |
| Reg-B3-L1 | 1.83 ± | 13.57 ± | 1.17 ± |
| Reg-B4-L1 | 12.59 ± | 93.63 ± | 8.68 ± |
| Reg-B5-L1 | 2.96 ± | 22.39 ± | 1.89 ± |
| Reg-B6-L1 | 3.23 ± | 24.29 ± | 2.13 ± |
| Reg-B7-L1 | 3.34 ± | 26.04 ± | 2.28 ± |
| Reg-B1-L2 | 3.16 ± | 23.83 ± | 2.13 ± |
| Reg-B2-L2 | 3.73 ± | 28.41 ± | 2.55 ± |
| Reg-B3-L2 | 2.35 ± | 17.32 ± | 1.53 ± |
| Reg-B4-L2 | 5.69 ± | 43.54 ± | 3.87 ± |
| Reg-B5-L2 | 3.12 ± | 23.77 ± | 1.99 ± |
| Reg-B6-L2 | 4.68 ± | 35.39 ± | 3.10 ± |
| Reg-B7-L2 | 4.33 ± | 32.29 ± | 2.87 ± |
| Reg-B1-L3 | 3.37 ± | 25.75 ± | 2.33 ± |
| Reg-B2-L3 | 3.12 ± | 24.03 ± | 2.11 ± |
| Reg-B3-L3 | 2.78 ± | 20.62 ± | 1.79 ± |
| Reg-B4-L3 | 9.15 ± | 70.49 ± | 6.20 ± |
| Reg-B5-L3 | 3.40 ± | 26.08 ± | 2.19 ± |
| Reg-B6-L3 | 4.30 ± | 32.48 ± | 2.86 ± |
| Reg-B7-L3 | 6.29 ± | 48.39 ± | 4.33 ± |
| Metrics | MAE (mm) | MAE (px) | PMAE (%) |
|---|---|---|---|
| Methods | Segmentation-based methods | ||
| U-Net-B2 | 1.08 ± 1.25 | 7.87 ± 7.51 | 0.65 ± 0.68 |
| LinkNet-B2 | 1.15 ± 1.32 | 8.45 ± 8.39 | 0.69 ± 0.77 |
| Segmentation-free methods | |||
| Reg-B3-L1 | 1.83 ± 2.11 | 13.57 ± 13.53 | 1.17 ± 1.43 |
| Reg-B3-L2 | 2.35 ± 2.74 | 17.32 ± 17.95 | 1.53 ± 2.02 |
| Methods | Train (s/Epoch) | Predict (s/Test Set) | Mem-M (GB) | Mem-P (GB) |
|---|---|---|---|---|
| Segmentation-based methods | ||||
| U-Net-B2 | 29 | 68.26 | 3.06 | 1.84 |
| DoubleU-Net | 70 | 114.21 | 7.21 | 2.40 |
| U-Net++-B2 | 68 | 172.45 | 7.26 | 2.34 |
| FPN-B2 | 44 | 101.30 | 5.47 | 2.04 |
| LinkNet-B2 | 30 | 80.36 | 3.82 | 1.90 |
| PSPNet-B2 | 88 | 225.38 | 11.06 | 4.04 |
| Segmentation-free method | ||||
| Reg-B1-L1 | 17 | 30.86 | 0.96 | 1.36 |
| Reg-B2-L1 | 20 | 48.28 | 2.31 | 1.73 |
| Reg-B3-L1 | 38 | 36.95 | 2.29 | 2.68 |
| Reg-B4-L1 | 21 | 65.55 | 3.01 | 1.69 |
| Reg-B5-L1 | 35 | 51.78 | 2.15 | 1.67 |
| Reg-B6-L1 | 14 | 18.71 | 1.03 | 1.14 |
| Reg-B7-L1 | 17 | 22.55 | 1.09 | 1.60 |
| Metrics | MAE (mm) | DI (%) |
|---|---|---|
| Methods | Segmentation-based methods | |
| Budd et al. [16] | 1.81 ± | 98.20 ± |
| Sobhaninia et al. [18] | 2.12 ± | 96.84 ± |
| Fiorentino et al. [19] | 1.90 ± | 97.75 ± |
| Moccia et al. [20] | 1.95 ± | 97.90 ± |
| U-Net-B2 (Proposed) | 1.08 ± | 98.80 ± |
| Segmentation-free methods | ||
| Reg-B3-L1 (Proposed) | 1.83 ± | N/A |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, J.; Petitjean, C.; Ainouz, S. Segmentation-Based vs. Regression-Based Biomarker Estimation: A Case Study of Fetus Head Circumference Assessment from Ultrasound Images. J. Imaging 2022, 8, 23. https://doi.org/10.3390/jimaging8020023
Zhang J, Petitjean C, Ainouz S. Segmentation-Based vs. Regression-Based Biomarker Estimation: A Case Study of Fetus Head Circumference Assessment from Ultrasound Images. Journal of Imaging. 2022; 8(2):23. https://doi.org/10.3390/jimaging8020023
Chicago/Turabian StyleZhang, Jing, Caroline Petitjean, and Samia Ainouz. 2022. "Segmentation-Based vs. Regression-Based Biomarker Estimation: A Case Study of Fetus Head Circumference Assessment from Ultrasound Images" Journal of Imaging 8, no. 2: 23. https://doi.org/10.3390/jimaging8020023
APA StyleZhang, J., Petitjean, C., & Ainouz, S. (2022). Segmentation-Based vs. Regression-Based Biomarker Estimation: A Case Study of Fetus Head Circumference Assessment from Ultrasound Images. Journal of Imaging, 8(2), 23. https://doi.org/10.3390/jimaging8020023








