Abstract
Reverse vaccinology (RV) is a computer-aided approach for vaccine development that identifies a subset of pathogen proteins as protective antigens (PAgs) or potential vaccine candidates. Machine learning (ML)-based RV is promising, but requires a dataset of PAgs (positives) and non-protective protein sequences (negatives). This study aimed to create an ML dataset, VPAgs-Dataset4ML, to predict viral PAgs based on PAgs obtained from Protegen. We performed seven steps to identify PAgs from the Protegen website and non-protective protein sequences from Universal Protein Resource (UniProt). The seven steps included downloading viral PAgs from Protegen, performing quality checks on PAgs using the standard BLASTp identity check ≤30% via MMseqs2, and computational steps running on Google Colaboratory and the Ubuntu terminal to retrieve and perform quality checks (similar to the PAgs) on non-protective protein sequences as negatives from UniProt. VPAgs-Dataset4ML contains 2145 viral protein sequences, with 210 PAgs in positive.fasta and 1935 non-protective protein sequences in negative.fasta. This dataset can be used to train ML models to predict antigens for various viral pathogens with the aim of developing effective vaccines.
Dataset: https://doi.org/10.17632/w78tyrjz4z.1
Dataset License: CC BY 4.0
Keywords:
viruses; antigens; machine learning; reverse vaccinology; vaccinology; vaccines; bioinformatics 1. Introduction
Infectious diseases are a significant public health concern, accounting for an average of ~177.07 deaths per 100,000 people worldwide annually [1,2]. Approximately 60% of all human infectious diseases have been identified, and 75% of the 30 new infectious diseases that have affected humanity over the last three decades are zoonotic [3,4]. The causative agents of these infections are disease-causing pathogens, such as viruses, bacteria, parasites, or fungi. The lack of a specific effective cure, treatment, and broadly protective vaccine makes combating these infections difficult. Viral pathogens, such as influenza type A (H1N1) virus, Ebola virus, human immunodeficiency virus (HIV), severe acute respiratory syndrome coronavirus 1 (SARS-CoV or SARS-CoV-1), and recently, SARS-CoV-2, have caused major pandemics and epidemics [5,6,7,8,9,10,11,12,13,14].
Vaccines are effective global public health interventions for controlling infectious diseases. Globally, vaccines save 386 million life years and 96 million disability-adjusted life years (DALYs), preventing nearly six million deaths annually [15]. Furthermore, vaccinations administered between 2021 and 2030 are expected to prevent an estimated 51 million deaths worldwide caused by various pathogens [16]. The Centers for Disease Control and Prevention (CDC) report that over 25 safe and effective vaccines are available to help prevent diseases [17]. Vaccines are available for viral diseases, such as influenza, hepatitis A, Ebola virus disease, COVID-19, Zika virus infection, polio, and smallpox [17,18]. However, more effective vaccines against these diseases are required to keep up with the emergence and spread of the viral variants. Additionally, despite advances in vaccination, no vaccines are available for numerous infectious diseases, including herpes, hepatitis C, and acquired immunodeficiency syndrome (AIDS) caused by HIV [18]. An innovative genome-based vaccine-design approach known as reverse vaccinology (RV), developed in the early 1990s, is promising for developing effective vaccines [19].
RV employs the computerized screening of pathogen protein sequences as the first step in vaccine development for selecting a subset of proteins as promising antigens, also known as potential vaccine candidates (PVCs) [20]. Following the identification of PVCs in this first step in the RV process, (i) the PVCs are purified and used to immunize animals, and (ii) the PVCs with experimentally proven protective potential and adjuvants are chosen for the clinical phase of vaccine development [20]. RV has three significant advantages over traditional vaccinology: (i) RV vaccine development takes 1–2 years, compared to 5–15 years in traditional vaccinology; (ii) the pathogen does not need to be grown in a laboratory; and (iii) all possible antigens, including those not expressed in vitro, can be identified [19]. The first RV study in 2000 selected and validated 28 immunogenic proteins in Group B meningococcus [21]. These proteins have been tested experimentally [21]. The Bexsero® vaccine, created using five of the 28 protein candidates, is licensed in Europe and the United States of America (USA) [22,23]. RV has been an active area of research since the success of Bexsero®, including the development and use of specialized bioinformatic tools known as RV prediction tools for vaccine candidate prediction.
RV prediction tools are computer software or online websites that can be broadly classified into two main categories based on their algorithmic approaches: rule-based filtering and machine learning (ML) [20,24]. Both types take pathogen protein sequences as inputs and output a subset as PVCs or non-PVCs [20].
Rule-based filtering is a traditional programming approach that uses predefined rules to filter input sequences and identify the PVCs [20]. On the other hand, ML is a data-driven approach that utilizes algorithms to identify patterns in a dataset of pathogen protein sequences [20]. The dataset used in ML is labeled as protective antigens (PAgs) for those sequences with experimental evidence (proven to generate a protective immune response in a laboratory animal model) and non-protective protein sequences otherwise. Supervised ML algorithms [25], such as Support Vector Machines (SVMs) [26,27], can be used to train models on a subset of the dataset (‘train set’) and then evaluate the model’s performance using a different subset of the data (‘test set’). An ML model is robust if it performs well on both the training and test sets. The PAgs are known as ‘positive data’ and non-protective protein sequences as ‘negative data’ in a binary classification ML setting. This binary classification setting was chosen because the aim was to try to distinguish between protective sequences and those that are not, similar to previous ML-based RV studies [24,28,29,30,31,32]. Other methods, such as cross-validation techniques, including k-fold cross-validation [33], can be used to evaluate the robustness of the model and prevent overfitting (where a model cannot generalize and instead fits closely to the training dataset). It is important to note that not all ML algorithms may be suitable for RV prediction, and choosing the correct algorithm depends on the specifics of the dataset and research goal. Furthermore, other methods, such as the filtering-based method, can be used alongside ML instead of depending on the research context.
Examples of rule-based filtering RV programs include (i) the new enhanced reverse vaccinology environment (NERVE) [34], (ii) Vaxign [35], (iii) Jenner-Predict [36], and (iv) VacSol [37]. The ML-based RV software includes (i) VaxiJen [28], (ii) ANTIGENpro [29], (iii) the model that Bowman et al. [30] developed and was revised by Heinson et al. [31], (iv) Antigenic [32], and (v) Vaxign-ML [24]. Vaxign2 [38] incorporates both the filtering-based method and ML.
ML is more powerful than the rule-based filtering approach for developing RV programs because ML establishes the rules from the data, whereas the programming used for rule-based filtering requires manual code rules. However, as previously stated, ML requires a training dataset of pathogen PAgs for positive data and non-protective protein sequences for negative data, which is difficult to obtain. Although PAgs may be found through a laborious manual curation of the literature, non-protective protein sequences (candidate proteins that failed preclinical testing) are not publicly available. To address this, a common approach is to artificially select a random set of proteins as non-protective protein sequences (non-PAgs) or to identify proteins with little similarity to all selected proteins as the ‘non-protective protein sequences’ and the PAgs [24,28,30,31].
Current ML-based RV programs were published between 2007 and 2020 and focused primarily on bacterial prediction. VaxiJen [28], ANTIGENpro [29], and Antigenic [32] are the only ML tools that predict viral PAgs. VaxiJen, published in 2007, includes pathogen-specific prediction models for viruses, bacteria, parasites, fungi, and tumors. VaxiJen’s authors predicted PAgs for their viral model using 200 viral protein sequence data (100 positives and 100 negatives) from the literature. However, the VaxiJen paper does not include the actual protein sequence data file(s) used to build the viral ML model. Only Swiss-Prot identifiers of the positive and negative data are provided in the paper’s supplementary Excel file.
Later, in 2010, Magnan et al. [29] developed a generic model, ANTIGENpro, for PAgs of any pathogen type. The authors used a dataset with a size of 1324 (576 PAgs and 748 non-protective protein sequences) to train the model. These included 100 viral PAgs referenced from VaxiJen in this dataset, and the remaining PAgs were from bacteria, fungi, and parasites. Their dataset contained no viral non-protective protein sequences (748 non-protective protein sequences were from non-viral pathogens). ANTIGENpro researchers, like the authors of VaxiJen, did not make their dataset public. In 2019, Rahman et al. [32] created a Random Forest-based generic model, Antigenic, for predicting the PAgs of any pathogen category. The dataset from ANTIGENpro’s study was used to create Antigenic. Because the dataset was not publicly available, the authors of Antigenic requested it from Magnan et al. [29], who provided it to them through private communication, as stated in their article. Although Rahman et al. [32] made this dataset publicly accessible, there are still no viral non-protective protein sequences (negatives) available for ML.
It is vital to create and publicize a dataset (including viral PAgs as positive and non-protective protein sequences as negatives) to predict viral PAgs for ML-based RV. This dataset can be incorporated into existing ML RV models [24,28,29,30,31,32] for potentially improved viral vaccine design or can be used by researchers to build robust new ML models. Such models can be used to predict novel PVCs, which may aid vaccinologists as a guide for vaccine development against various viral pathogens. Protegen (https://violinet.org/protegen/) (accessed on 9 November 2022) [39] is the only publicly accessible web-based PAg database, published in 2011. Protegen [39] contains 1371 manually curated PAgs from peer-reviewed articles in the literature for over 200 infectious diseases caused by different pathogens (bacteria, viruses, parasites, and fungi) and non-infectious diseases, including allergies, arthritis, cancer, and diabetes. Each of these collected PAgs is an antigen capable of eliciting a protective immune response, as demonstrated in at least one laboratory animal model. However, to our knowledge, no study has been published in which Protegen’s viral PAgs were used as a basis to create an ML dataset. Therefore, this study aimed to develop an ML dataset for predicting viral protective antigens (PAgs), called viral protective antigens (VPAgs) dataset for machine learning (VPAgs-Dataset4ML), for ML-based RV based on Protegen. Viral non-protective protein sequences were selected through computational steps from Universal Protein Resource (UniProt) [40]. Our contributions are listed in the context of the availability of datasets to predict viral PAgs for ML-based RV (Table 1).

Table 1.
Availability of datasets to predict viral protective antigens for machine learning-based reverse vaccinology.
The remainder of this paper is organized as follows. Section 2 provides an overview of the methods, including the seven-step workflow for creating the VPAgs-Dataset4ML. The VPAgs-Dataset4ML dataset is described in Section 3. Section 4 contains user notes for researchers who intend to use VPAgs-Dataset4ML. Finally, Section 5 concludes the study.
2. Methods
The VPAgs-Dataset4ML dataset was created using the following seven steps. The dataset creation process is illustrated in Figure 1. The complete source code of the scripts and outputs after each step, as mentioned in this section, is available in the GitHub repository: https://github.com/ZakiaSalod/VPAgs-Dataset4ML (accessed on 9 November 2022). The VPAgs-Dataset4ML dataset is publicly available to the research community on Mendeley Data at https://doi.org/10.17632/w78tyrjz4z.1 (accessed on 9 November 2022).
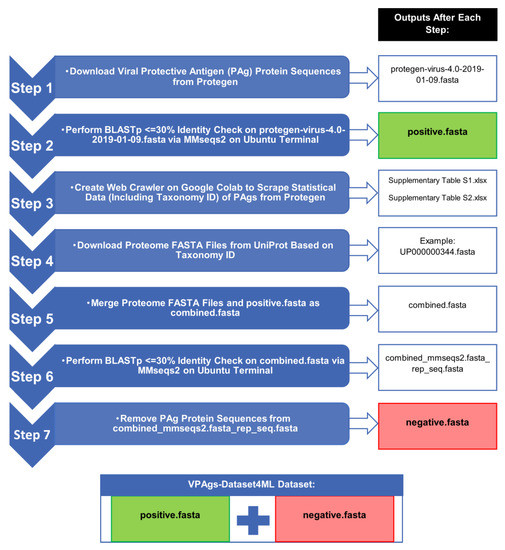
Figure 1.
Workflow of the seven steps for creating the VPAgs-Dataset4ML dataset (positive.fasta and negative.fasta). Abbreviations: PAg: protective antigen; BLASTp: basic local alignment search tool protein–protein; MMseqs2: many-against-many sequence; Google Colab: Google Colaboratory; VPAgs-Dataset4ML: viral protective antigens dataset for machine learning.
2.1. Step 1: Download Viral Protective Antigen (PAg) Protein Sequences from Protegen
We downloaded viral PAgs with experimental evidence (protegen-virus-4.0-2019-01-09.faa) from the Protegen database (https://violinet.org/protegen/download/index.php) (accessed on 1 September 2022) [39] as a FASTA file called protegen-virus-4.0-2019-01-09.fasta on 1 September 2022. As of 1 September 2022, Protegen contained 414 PAgs from 112 pathogenic viruses.
2.2. Step 2: Perform BLASTp ≤ 30% Identity Check on protegen-virus-4.0-2019-01-09.fasta via Many-Against-Many Sequence (MMseqs2) on the Ubuntu Terminal
The PAgs with a basic local alignment search tool (BLAST) [41] (specifically, BLASTp) identity check of over 30% similarity were deemed homologous proteins and eliminated from this analysis to remove any potential bias. The 30% cut-off is a standard bioinformatics test for homologous proteins [42]. We used Conda [43] to download the many-against-many sequence searching 2 (MMseqs2) [44] Python package and ran it on the Ubuntu terminal to access the BLAST. The FASTA file, protegen-virus-4.0-2019-01-09.fasta, was used as the input for MMseqs2, and we ran the following command on the Ubuntu terminal:
mmseqs easy-cluster protegen-virus-4.0-2019-01-09.fasta_rep_seq.fasta positive.fasta logs -c 0.5 --min-seq-id 0.3
Representative sequences in FASTA file format (positive.fasta_rep_seq.fasta) were output. We renamed positive.fasta_rep_seq.fasta as positive.fasta for better clarity as the final positive FASTA file. The final number of positive samples decreased from 414 out of 112 pathogenic viruses in protegen-virus-4.0-2019-01-09.fasta to 210 PAgs in positive.fasta from 81 pathogenic viruses.
2.2.1. A Brief Overview of BLAST
BLAST [41] is a bioinformatics algorithm for comparing a biological sequence, such as a deoxyribonucleic acid (DNA) or protein sequence, to a database of sequences. Several algorithms are available, including BLASTn (neucleotide–nucleotide), BLASTp (protein–protein), and BLASTx (translated nucleotide–protein). BLAST is commonly used to identify similar sequences in a database and can be used to infer functional and evolutionary relationships among sequences. The algorithm works by finding regions of high similarity between the query sequence and sequences in the database and then uses statistical methods to determine the significance of the similarities. In BLAST, there is also the option to ‘align two or more sequences’, which refers to the process of comparing multiple sequences to each other to identify regions of similarity. This option is particularly useful when preparing a dataset, such as the VPAgs-Dataset4ML, for RV. BLAST is one of the most widely used bioinformatics tools and is available in a variety of forms, including a command-line version, web-based version, and as a library for use in other software.
2.2.2. A Brief Overview of MMseqs2
MMseqs2 [44] is a bioinformatics tool used for analyzing and comparing large sets of protein or nucleotide sequences. The tool is an update to the original MMseqs [45] package and includes several improvements in terms of speed and accuracy. MMseqs2 can be used for tasks such as clustering, database searches, and profile–profile alignment, and creates sensitive and fast sequence searches. MMseqs2 is useful for preparing a dataset, such as VPAgs-Dataset4ML for RV, as it is a quicker method of conducting sequence similarity checks compared with using BLAST [41] directly. It is designed to be highly efficient, both in terms of memory usage and computational time, making it well suited for processing large-scale datasets. MMseqs2 is based on a novel indexing strategy and a parallelized algorithm that allows it to process millions of sequences in minutes. MMseqs2 software is open-source and can be run on a variety of platforms, including Linux, MacOS, and (as a beta version, via cygwin) Windows. To this end, although we had access to Ubuntu (a Linux distribution) and Windows platforms, we ran our MMseqs2 commands on Ubuntu for this study because the MMseqs2 implementation is stable on this platform, whereas the tool is in the beta version for Windows.
The -c parameter in the MMseqs2 command line was used to specify the level of clustering that should be performed on the input sequences. This sets the minimum coverage of the sequences in a cluster. Coverage is defined as the percentage of the length of the longest sequence in a cluster that has to be aligned. For example, in our study, we used a value of 0.5 for this -c parameter, which means that at least 50% of the longest sequence in a cluster must be aligned with the other sequences in the cluster for them to be considered as part of the same cluster. In other words, this parameter sets a threshold for the similarity of the sequences in a cluster, with higher values resulting in more stringent clustering (i.e., fewer, larger clusters) and lower values resulting in less stringent clustering (i.e., more, smaller clusters).
2.3. Step 3: Create Web Crawler on Google Colab to Scrape Statistical Data (Including Taxonomy ID) of PAgs from Protegen
We used Python version 3.7.15 to create a web crawler to scrape Protegen’s statistics webpage (https://violinet.org/protegen/stat.php) (accessed on 5 September 2022) to retrieve the statistical data of the positive.fasta file. Python script (VPAgs-Dataset4ML.ipynb) included the use of Python packages Pandas version 1.3.5 [46,47,48] for data processing and Beautiful Soup version 4.11.0 [49] for web scraping, and it ran in the browser on Google Research’s free Jupyter Notebook environment (.ipynb files) called Google Colaboratory (also known as ‘Colab’ for short) [50]. The output from this script is shown in Supplementary Tables S1 (shows data for all 414 viruses from Protegen) and S2 (shows data for 210 viruses as per Step 2). This result is also significant because it contains the ‘taxonomy id’ of the PAgs species, a requirement for the next steps in generating the negative dataset.
2.4. Step 4: Download Proteome FASTA Files from UniProt Based on Taxonomy ID
A set of ‘gold-standard’ non-protective protein sequences did not exist. As a result, negative samples were chosen based on their sequence dissimilarity to the PAgs (in positive.fasta) and the list of all sequences chosen as the negative samples, as described in previous ML-based RV prediction studies [24,28,30,31]. We downloaded the pathogen proteome (all protein sequences) FASTA files from the UniProt database [40], based on the distinct taxonomy ids listed in Supplementary Table S2. If the pan proteome of the taxonomy id was not available on UniProt, we chose a reference proteome, and as a last resort, we chose a proteome under ‘other proteome’ for the ‘taxonomy_id’. Briefly, the pan proteome is the entire set of proteins found in multiple strains of the same species [51]. The reference proteome is a collection of protein sequences that covers a wide range of species [51]. The ‘other proteome’ contains any sequences for that species that are available [51]. As we downloaded the proteomes, we updated the ‘uniprot_proteome_id’ column in Supplementary Table S2 with the proteome id values obtained from the UniProt downloaded files for tracking purposes. If no proteome was found in UniProt for a given taxonomy id, we noted it in Supplementary Table S2 by entering ‘FILE_NOT_FOUND_IN_UNIPROT’ in the ‘uniprot_proteome_id’ column. A total of 95 proteome FASTA files were downloaded, with 10 proteomes missing from UniProt.
2.5. Step 5: Merge Proteome FASTA Files and positive.fasta as combined.fasta
We placed the 95 proteome FASTA files and positive.fasta into the same directory in Ubuntu. Subsequently, we used the Ubuntu terminal to run the following command to combine all 96 files into a single file called combined.fasta, which was required for the next step:
cat *.fasta > combined.fasta
The combined.fasta file had 31,411 protein sequences.
2.6. Step 6: Perform BLASTp ≤30% Identity Check on combined.fasta via MMseqs2 on Ubuntu Terminal
Similar to Step 2 above, as part of the quality checks in selecting the negative dataset, we used the Ubuntu terminal to run the following command with combined.fasta as input for MMseqs2 to remove protein sequences in combined.fasta with more than 30% similarity to those in positive.fasta:
mmseqs easy-cluster combined.fasta combined_mmseqs2.fasta logs -c 0.5 --min-seq-id 0.3
The combined_mmseqs2.fasta_rep_seq.fasta representative sequence FASTA file was outputted with 2088 protein sequences. This step ensured that the protein sequences in combined.fasta that were very similar to those in positive.fasta were eliminated.
2.7. Step 7: Remove PAg Protein Sequences from combined_mmseqs2.fasta_rep_seq.fasta
On Google Colaboratory, we wrote a Python script (Biopython_Remove_Positives_(PAgs).ipynb) that used the Biopython [52] package to read in combined_mmseqs2.fasta_rep_seq.fasta and output a new FASTA file called negative.fasta, excluding 131 protein sequences that had the word ‘Protegen’ in combined_mmseqs2.fasta_rep_seq.fasta and 22 protein sequences that had matching entries in protegen-virus-4.0-2019-01-09.fasta (Biopython_Remove_Positives_(PAgs)_Helper.ipynb). This ensured the exclusion of all PAgs (positive). After removing 153 PAgs, the final negative samples in negative.fasta contained 1935 protein sequences.
The VPAgs-Dataset4ML dataset is composed of positive.fasta and negative.fasta. A description of the dataset is provided in Section 3.
2.8. Ethics and Permission
This study only used secondary data and did not include patient information. Therefore, no ethical approval was required for this data descriptor. Nevertheless, this study was part of a larger research project submitted to the Biomedical Research Ethics Committee (BREC) of the University of KwaZulu-Natal (UKZN) in Durban, KwaZulu-Natal, South Africa for ethical considerations. On 31 March 2022, the BREC granted exemption from the ethics review of this project.
3. Data Description
3.1. Quantitative Overview of VPAgs-Dataset4ML
There are 2145 protein sequences in the VPAgs-Dataset4ML dataset, made up of positive.fasta and negative.fasta. The positive.fasta file had 210 (VPAgs-Dataset4ML: 210/2145; 10%) protein sequences, and the negative.fasta file had 1935 (VPAgs-Dataset4ML: 1935/2145; 90%) protein sequences from various viral pathogens.
3.2. A Snapshot of VPAgs-Dataset4ML
Samples of the first five sequences were tabulated from positive.fasta (Table 2) and negative.fasta (Table 3). The five protein sequences in the respective tables are shown in the FASTA file format (Table 2 and Table 3). FASTA is a text-based bioinformatics standard format for displaying amino acid (protein) sequences. This file format depicts each protein sequence with a “>” symbol (greater-than-sign). The first line of every protein sequence in a FASTA file is the sequence header, starting with “>”, and the data that follow this symbol is a unique description of the sequence. In Table 2, for example, “>Protegen: 579|VO: VO_0011168|NP_056918.1 nucleocapsid protein [Measles morbillivirus]” is the first sequence, indicating the different identifiers of the sequence, for instance, the Protegen identifier, the protein name is “nucleocapsid protein”, and “[Measles morbillivirus]” is the name of the viral pathogen. Following this first line, the actual sequence of a standard one-letter character string is shown. The next sequence in Table 2 is: “>Protegen: 580|VO: VO_0011169|NP_056919.1 phosphoprotein [Measles morbillivirus]”, and so on, totaling five sequences.

Table 2.
Sample of the first five protein sequence data of protective antigens (PAgs) (positives) from the positive.fasta FASTA file.

Table 3.
Sample of the first five protein sequence data of non-protective protein sequences (negatives) from the negative.fasta FASTA file.
4. User Notes
4.1. Usage of the VPAgs-Dataset4ML Dataset
The VPAgs-Dataset4ML, which includes PAgs in positive.fasta and non-protective protein sequences in negative.fasta, can be used to train ML models to predict protective antigens (PVCs) for various viral pathogens. The positive data were PAgs, whereas the negative data were non-protective protein sequences. For ML, only the sequence data from the positive.fasta and negative.fasta files are required, and the sequence header line beginning with “>” can be ignored. The goal would be to predict the PAgs independent variable as ‘PVC’, denoted by a label (or dependent variable) set to ‘1’ (positive), and the non-protective protein sequences as ‘not-PVC’ as indicated by the label set as ‘0’ (negative). To do so, the positive.fasta and negative.fasta files should be read in a script. The Biopython [52] package in Python can be used to load these files into a Python script as the first step in the ML process. Researchers with ML expertise can then use Python ML tools, such as Tensorflow [53], scikit-learn [54], and PyTorch [55], to create and compare various ML models to predict viral protective antigens. In addition to these Python-based ML tools, the Weka [56], KNIME [57,58], Orange [59], dplyr [60], and caret [61] tools in the R [62] programming language are other options available to perform ML.
4.2. Imbalanced Data
As is common in medical datasets, VPAgs-Dataset4ML has a significant bias (referred to as ‘class imbalance’ in ML) toward one of the categories. Non-protective protein sequences accounted for 90% of the data (1935 of 2145), whereas PAgs accounted for 10% of the total dataset (210 of 2145). In ML, this may result in a high prediction accuracy driven by the majority category at the expense of the poor performance of the minority category. Therefore, anyone using this dataset should be aware of this issue and address it using standard data- or algorithm-level techniques. A sampling technique, such as random oversampling, the Synthetic Minority Over Sampling Technique (SMOTE) [63], or the Adaptive Synthetic Sampling Approach (ADASYN) [64] (with k-nearest neighbors default parameter k = 5, which may be amended as required) for oversampling, random undersampling, a clustering-based strategy [65], or Tomek link (T-link) [66] for undersampling can be used in the data-level approach. At the algorithmic level, an ML algorithm with balancing steps, such as Balanced Random Forest [67], or bagging classifiers with additional balancing, such as Over-Bagging [68], must be used.
5. Conclusions
This data descriptor study presented the VPAgs-Dataset4ML, a dataset created with quality checks based on experimentally validated viral PAgs from Protegen and computational steps conducted to select a representative set of non-protective protein sequences from proteomes taken from UniProt, based on the taxonomy id of the PAgs species. There were 2145 protein sequences in the VPAgs-Dataset4ML dataset (210/2145; 10% in positive.fasta and 1935/2145; 90% in negative.fasta). Researchers can incorporate the VPAgs-Dataset4ML into existing ML RV models, such as VaxiJen, which may enable improved vaccine design, or conduct new ML-based RV studies to predict viral PAgs. Although the VPAgs-Dataset4ML is imbalanced, we have provided suggestions in Section 4, pointing out that standard methods researchers could use to handle the imbalance to create more robust ML models. To the best of our knowledge, this is the first dataset made publicly available and ready to perform ML for predicting viral PAgs in RV, with the added benefit of providing details of all steps conducted and the code and outputs in each step of the dataset creation process. The PAgs constituted positive data, and the non-protective protein sequences were negative data for ML. The ML models developed using the VPAgs-Dataset4ML have implications for identifying potential vaccine candidates (PAgs) for various viral pathogens. Vaccinologists may use this information to guide the development of novel and effective vaccines that could assist in saving the lives of patients.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/data8020041/s1, Table S1: Statistical data of 414 PAgs from Protegen; Table S2: Statistical data of 210 PAgs from Protegen with UniProt Proteome ID.
Author Contributions
Conceptualization, Z.S.; methodology, Z.S.; software, Z.S.; validation, O.M., formal analysis, Z.S.; investigation, Z.S.; resources, Z.S. and O.M.; data curation, Z.S.; writing—original draft preparation, Z.S.; writing—review and editing, Z.S. and O.M.; visualization, Z.S.; supervision, O.M.; project administration, Z.S.; funding acquisition, Z.S. and O.M. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the National Research Foundation (NRF) of South Africa (grant number 130187) and College of Health Sciences (CHS) of the University of KwaZulu-Natal (UKZN) in Durban, KwaZulu-Natal, South Africa, grant number N/A. The APC was funded by Dr Ozayr Mahomed.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The data presented in this study is publicly available on Mendeley Data at https://doi.org/10.17632/w78tyrjz4z.1 (accessed on 9 November 2022).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Our World in Data. Death Rate from Infectious Diseases, 1990 to 2019. Available online: https://ourworldindata.org/grapher/infectious-disease-death-rates (accessed on 10 January 2023).
- Vos, T.; Lim, S.S.; Abbafati, C.; Abbas, K.M.; Abbasi, M.; Abbasifard, M.; Abbasi-Kangevari, M.; Abbastabar, H.; Abd-Allah, F.; Abdelalim, A. Global burden of 369 diseases and injuries in 204 countries and territories, 1990–2019: A systematic analysis for the Global Burden of Disease Study 2019. Lancet 2020, 396, 1204–1222. [Google Scholar] [CrossRef] [PubMed]
- Jones, K.E.; Patel, N.G.; Levy, M.A.; Storeygard, A.; Balk, D.; Gittleman, J.L.; Daszak, P. Global trends in emerging infectious diseases. Nature 2008, 451, 990–993. [Google Scholar] [CrossRef] [PubMed]
- Woolhouse, M.E.; Gowtage-Sequeria, S. Host range and emerging and reemerging pathogens. Emerg. Infect. Dis. 2005, 11, 1842. [Google Scholar] [CrossRef] [PubMed]
- Taubenberger, J.K.; Morens, D.M. 1918 Influenza: The mother of all pandemics. Rev. Biomed. 2006, 17, 69–79. [Google Scholar] [CrossRef]
- Frost, W.H. Statistics of Influenza Morbidity: With Special Reference to Certain Factors in Case Incidence and Case Fatality. Public Heal. Rep. 1896–1970 1920, 35, 584. [Google Scholar] [CrossRef]
- Johnson, N.P.A.S.; Mueller, J. Updating the Accounts: Global Mortality of the 1918–1920 “Spanish” Influenza Pandemic. Bull. Hist. Med. 2002, 76, 105–115. [Google Scholar] [CrossRef]
- World Health Organization. Ebola Virus Disease. Available online: https://www.who.int/news-room/fact-sheets/detail/ebola-virus-disease (accessed on 22 October 2022).
- Merson, M.H. The HIV–AIDS pandemic at 25—The global response. N. Engl. J. Med. 2006, 354, 2414–2417. [Google Scholar] [CrossRef]
- World Health Organization. HIV/AIDS. Available online: https://www.who.int/news-room/fact-sheets/detail/hiv-aids (accessed on 10 November 2022).
- Hon, E.; Li, A.; Nelson, E.; Leung, C. Severe Acute Respiratory Syndrome (SARS) In: Textbook of Paediatric Infectious Diseases, Feigin, R.D.; Cherry, J.D., Demmler, G.J., Kaplan, S., Eds.; Elsevier: Amsterdam, The Netherlands, 2003. [Google Scholar]
- World Health Organization. Summary of Probable SARS Cases with Onset of Illness from 1 November 2002 to 31 July 2003. Available online: https://www.who.int/publications/m/item/summary-of-probable-sars-cases-with-onset-of-illness-from-1-november-2002-to-31-july-2003 (accessed on 10 November 2022).
- World Health Organization. Consensus Document on the Epidemiology of Severe Acute Respiratory syndrome (SARS); World Health Organization: Geneva, Switzerland, 2003. [Google Scholar]
- Worldometers. COVID-19 Coronavirus Pandemic. Available online: https://www.worldometers.info/coronavirus/ (accessed on 10 November 2022).
- Ehreth, J. The global value of vaccination. Vaccine 2003, 21, 596–600. [Google Scholar] [CrossRef]
- Carter, A.; Msemburi, W.; Sim, S.Y.; AM Gaythorpe, K.; Lindstrand, A.; Hutubessy, R.C. Modeling the impact of vaccination for the immunization agenda 2030: Deaths averted due to vaccination against 14 pathogens in 194 countries from 2021–2030. Ann Hutubessy Raymond CW Model. Impact Vaccin. Immun. Agenda 2021, 2030, 1–41. [Google Scholar] [CrossRef]
- Centers for Disease Control and Prevention. Fast Facts on Global Immunization. Available online: https://www.cdc.gov/globalhealth/immunization/data/fast-facts.html#:~:text=Immunization%20Prevents%20Death%20Worldwide,save%20nearly%2019%20million%20lives (accessed on 5 November 2022).
- Koff, W.C.; Burton, D.R.; Johnson, P.R.; Walker, B.D.; King, C.R.; Nabel, G.J.; Ahmed, R.; Bhan, M.K.; Plotkin, S.A. Accelerating Next-Generation Vaccine Development for Global Disease Prevention. Science 2013, 340, 1232910. [Google Scholar] [CrossRef]
- Rappuoli, R. Reverse vaccinology. Curr. Opin. Microbiol. 2000, 3, 445–450. [Google Scholar] [CrossRef]
- Dalsass, M.; Brozzi, A.; Medini, D.; Rappuoli, R. Comparison of open-source reverse vaccinology programs for bacterial vaccine antigen discovery. Front. Immunol. 2019, 10, 113. [Google Scholar] [CrossRef]
- Pizza, M.; Scarlato, V.; Masignani, V.; Giuliani, M.M.; Aricò, B.; Comanducci, M.; Jennings, G.T.; Baldi, L.; Bartolini, E.; Capecchi, B.; et al. Identification of Vaccine Candidates Against Serogroup B Meningococcus by Whole-Genome Sequencing. Science 2000, 287, 1816–1820. [Google Scholar] [CrossRef] [PubMed]
- Folaranmi, T.; Rubin, L.; Martin, S.W.; Patel, M.; MacNeil, J.R. Use of serogroup B meningococcal vaccines in persons aged ≥10 years at increased risk for serogroup B meningococcal disease: Recommendations of the Advisory Committee on Immunization Practices, 2015. MMWR. Morb. Mortal. Wkly. Rep. 2015, 64, 608. [Google Scholar] [PubMed]
- Vernikos, G.; Medini, D. Bexsero® chronicle. Pathog. Glob. Health 2014, 108, 305–316. [Google Scholar] [CrossRef] [PubMed]
- Ong, E.; Wang, H.; Wong, M.U.; Seetharaman, M.; Valdez, N.; He, Y. Vaxign-ML: Supervised machine learning reverse vaccinology model for improved prediction of bacterial protective antigens. Bioinformatics 2020, 36, 3185–3191. [Google Scholar] [CrossRef] [PubMed]
- Kotsiantis, S.B.; Zaharakis, I.; Pintelas, P. Supervised machine learning: A review of classification techniques. Emerg. Artif. Intell. Appl. Comput. Eng. 2007, 160, 3–24. [Google Scholar]
- Vapnik, V.N. The Nature of Statistical Learning Theory; Springer Science & Business Media: Berlin/Heidelberg, Germany, 1995. [Google Scholar]
- Burges, C.J.C. A Tutorial on Support Vector Machines for Pattern Recognition. Data Min. Knowl. Discov. 1998, 2, 121–167. [Google Scholar] [CrossRef]
- Doytchinova, I.A.; Flower, D.R. VaxiJen: A server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinform. 2007, 8, 4. [Google Scholar] [CrossRef]
- Magnan, C.N.; Zeller, M.; Kayala, M.A.; Vigil, A.; Randall, A.; Felgner, P.L.; Baldi, P. High-throughput prediction of protein antigenicity using protein microarray data. Bioinformatics 2010, 26, 2936–2943. [Google Scholar] [CrossRef]
- Bowman, B.N.; McAdam, P.R.; Vivona, S.; Zhang, J.X.; Luong, T.; Belew, R.K.; Sahota, H.; Guiney, D.; Valafar, F.; Fierer, J.; et al. Improving reverse vaccinology with a machine learning approach. Vaccine 2011, 29, 8156–8164. [Google Scholar] [CrossRef] [PubMed]
- Heinson, A.I.; Gunawardana, Y.; Moesker, B.; Hume, C.C.D.; Vataga, E.; Hall, Y.; Stylianou, E.; McShane, H.; Williams, A.; Niranjan, M.; et al. Enhancing the Biological Relevance of Machine Learning Classifiers for Reverse Vaccinology. Int. J. Mol. Sci. 2017, 18, 312. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.S.; Rahman, K.; Saha, S.; Kaykobad, M.; Rahman, M.S. Antigenic: An improved prediction model of protective antigens. Artif. Intell. Med. 2019, 94, 28–41. [Google Scholar] [CrossRef]
- Kohavi, R. A Study of Cross-Validation and Bootstrap for Accuracy Estimation and Model Selection. In Proceedings of the The 1995 International Joint Conference, Montreal, QC, Canada, 20–25 August 1995; pp. 1137–1145. [Google Scholar]
- Vivona, S.; Bernante, F.; Filippini, F. NERVE: New enhanced reverse vaccinology environment. BMC Biotechnol. 2006, 6, 35. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Xiang, Z.; Mobley, H.L.T. Vaxign: The First Web-Based Vaccine Design Program for Reverse Vaccinology and Applications for Vaccine Development. J. Biomed. Biotechnol. 2010, 2010, 297505. [Google Scholar] [CrossRef] [PubMed]
- Jaiswal, V.; Chanumolu, S.K.; Gupta, A.; Chauhan, R.S.; Rout, C. Jenner-predict server: Prediction of protein vaccine candidates (PVCs) in bacteria based on host-pathogen interactions. BMC Bioinform. 2013, 14, 211. [Google Scholar] [CrossRef] [PubMed]
- Rizwan, M.; Naz, A.; Ahmad, J.; Naz, K.; Obaid, A.; Parveen, T.; Ahsan, M.; Ali, A. VacSol: A high throughput in silico pipeline to predict potential therapeutic targets in prokaryotic pathogens using subtractive reverse vaccinology. BMC Bioinform. 2017, 18, 106. [Google Scholar] [CrossRef]
- Ong, E.; Cooke, M.F.; Huffman, A.; Xiang, Z.; Wong, M.U.; Wang, H.; Seetharaman, M.; Valdez, N.; He, Y. Vaxign2: The second generation of the first Web-based vaccine design program using reverse vaccinology and machine learning. Nucleic Acids Res. 2021, 49, W671–W678. [Google Scholar] [CrossRef]
- Yang, B.; Sayers, S.; Xiang, Z.; He, Y. Protegen: A web-based protective antigen database and analysis system. Nucleic Acids Res. 2011, 39, D1073–D1078. [Google Scholar] [CrossRef]
- UniProt Consortium. The universal protein resource (UniProt). Nucleic Acids Res. 2007, 36, D190–D195. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Pearson, W.R. An introduction to sequence similarity (“homology”) searching. Curr. Protoc. Bioinform. 2013, 42, 3.1.1–3.1.8. [Google Scholar] [CrossRef]
- Anaconda Software Distribution. Conda. Available online: https://www.anaconda.com/ (accessed on 30 October 2022).
- Steinegger, M.; Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 2017, 35, 1026–1028. [Google Scholar] [CrossRef] [PubMed]
- Hauser, M.; Steinegger, M.; Söding, J. MMseqs software suite for fast and deep clustering and searching of large protein sequence sets. Bioinformatics 2016, 32, 1323–1330. [Google Scholar] [CrossRef] [PubMed]
- Reback, J.; McKinney, W.; Van Den Bossche, J.; Augspurger, T.; Cloud, P.; Klein, A.; Hawkins, S.; Roeschke, M.; Tratner, J.; She, C. pandas-dev/pandas: Pandas 1.0. 5. Zenodo 2020. [Google Scholar]
- McKinney, W. Data structures for Statistical Computing in Python. In Proceedings of the Proceedings of the 9th Python in Science Conference, Austin, TX, USA, 28 June–3 July 2010; pp. 51–56. [Google Scholar]
- McKinney, W. Pandas: A foundational Python library for data analysis and statistics. Python High Perform. Sci. Comput. 2011, 14, 1–9. [Google Scholar]
- Richardson, L. Beautiful Soup Documentation. Available online: https://www.crummy.com/software/BeautifulSoup/bs4/doc/ (accessed on 30 October 2022).
- Bisong, E. Google Colaboratory. In Building Machine Learning and Deep Learning Models on Google Cloud Platform: A Comprehensive Guide for Beginners; Apress: Berkeley, CA, USA, 2019; pp. 59–64. [Google Scholar]
- Apweiler, R.; Bairoch, A.; Wu, C.H.; Barker, W.C.; Boeckmann, B.; Ferro, S.; Gasteiger, E.; Huang, H.; Lopez, R.; Magrane, M. UniProt. Available online: https://www.uniprot.org/ (accessed on 11 March 2022).
- Cock, P.J.A.; Antao, T.; Chang, J.T.; Chapman, B.A.; Cox, C.J.; Dalke, A.; Friedberg, I.; Hamelryck, T.; Kauff, F.; Wilczynski, B.; et al. Biopython: Freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 2009, 25, 1422–1423. [Google Scholar] [CrossRef]
- Abadi, M.; Agarwal, A.; Barham, P.; Brevdo, E.; Chen, Z.; Citro, C.; Corrado, G.S.; Davis, A.; Dean, J.; Devin, M. TensorFlow: Large-Scale Machine Learning on Heterogeneous Systems. arXiv 2015, arXiv:1603.04467. [Google Scholar]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Paszke, A.; Gross, S.; Chintala, S.; Chanan, G.; Yang, E.; DeVito, Z.; Lin, Z.; Desmaison, A.; Antiga, L.; Lerer, A. Automatic differentiation in pytorch. NIPS 2017 Workshop Autodiff 2017. [Google Scholar]
- Frank, E.; Hall, M.A.; Witten, I.H. Online Appendix for “Data Mining: Practical Machine Learning Tools and Techniques, 4th ed.; Morgan Kaufmann: Burlington, MA, USA, 2016. [Google Scholar]
- Berthold, M.; Cebron, N.; Dill, F.; Gabriel, T.; Kötter, T.; Meinl, T.; Ohl, P.; Sieb, C.; Thiel, K.; Wiswedel, B. KNIME: The Konstanz Information Miner in Data Analysis, Machine Learning and Applications SE-38; Preisach, C., Burkhardt, H., Schmidt-Thieme, L., Decker, R., Eds.; Springer: Berlin/Heidelberg, Germany, 2008. [Google Scholar]
- Berthold, M.R.; Cebron, N.; Dill, F.; Gabriel, T.R.; Kötter, T.; Meinl, T.; Ohl, P.; Thiel, K.; Wiswedel, B. KNIME-the Konstanz information miner: Version 2.0 and beyond. AcM SIGKDD Explor. Newsl. 2009, 11, 26–31. [Google Scholar] [CrossRef]
- Demšar, J.; Curk, T.; Erjavec, A.; Gorup, Č.; Hočevar, T.; Milutinovič, M.; Možina, M.; Polajnar, M.; Toplak, M.; Starič, A. Orange: Data mining toolbox in Python. J. Mach. Learn. Res. 2013, 14, 2349–2353. [Google Scholar]
- Wickham, H.; François, R.; Henry, L.; Müller, K. dplyr: A grammar of data manipulation. R Package Version 0.4 2015, 3, 156. [Google Scholar]
- Kuhn, M. Building predictive models in R using the caret package. J. Stat. Softw. 2008, 28, 1–26. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing. Available online: https://www.R-project.org/ (accessed on 6 November 2022).
- Chawla, N.V.; Bowyer, K.W.; Hall, L.O.; Kegelmeyer, W.P. SMOTE: Synthetic minority over-sampling technique. J. Artif. Intell. Res. 2002, 16, 321–357. [Google Scholar] [CrossRef]
- He, H.; Bai, Y.; Garcia, E.A.; Li, S. ADASYN: Adaptive Synthetic Sampling Approach for Imbalanced Learning. In Proceedings of the 2008 IEEE International Joint Conference on Neural Networks (IEEE World Congress on Computational Intelligence), Hong Kong, China, 1–8 June 2008; pp. 1322–1328. [Google Scholar]
- Lin, W.-C.; Tsai, C.-F.; Hu, Y.-H.; Jhang, J.-S. Clustering-based undersampling in class-imbalanced data. Inf. Sci. 2017, 409-410, 17–26. [Google Scholar] [CrossRef]
- Tomek, I. An Experiment with The Edited Nearest-Nieghbor Rule. IEEE Trans. Syst. Man Cybern. 1976, 6, 448–452. [Google Scholar]
- Chen, C.; Liaw, A.; Breiman, L. Using random forest to learn imbalanced data. Univ. Calif. Berkeley 2004, 110, 24. [Google Scholar]
- Maclin, R.; Opitz, D. An empirical evaluation of bagging and boosting. AAAI/IAAI 1997, 1997, 546–551. [Google Scholar]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).