IESS-FusionNet: Physiologically Inspired EEG-EMG Fusion with Linear Recurrent Attention for Infantile Epileptic Spasms Syndrome Detection
Abstract
1. Introduction
- We present IESS-FusionNet, an end-to-end multimodal framework that achieves accurate fusion of EEG and EMG for automated IESS detection.
- We introduce a unified Unimodal Encoder that jointly captures multi-scale frequency, spatial topology, local morphology, and global temporal dynamics of non-stationary biosignals in an efficient hierarchical design.
- We propose Cross Time-Mixing, a linear recurrent attention mechanism that enables dynamic, physiologically plausible, and bidirectional integration of EEG and EMG sequences.
2. Methods
2.1. Overall Architecture
2.2. Unimodal Encoder
2.2.1. Time-Frequency Decomposition
2.2.2. Spatio-Temporal Feature Extraction
2.2.3. Global Sequence Modeling
2.3. Cross-Modal Fusion
2.4. Classifier
3. Clinical Dataset
3.1. Data Source
3.2. Data Preprocessing
4. Experimental Results
4.1. Implementation Details
4.2. Evaluation Metrics
4.3. Comparative Performance
4.4. Ablation Study on Unimodal Encoder Components
4.5. Computational Efficiency Analysis
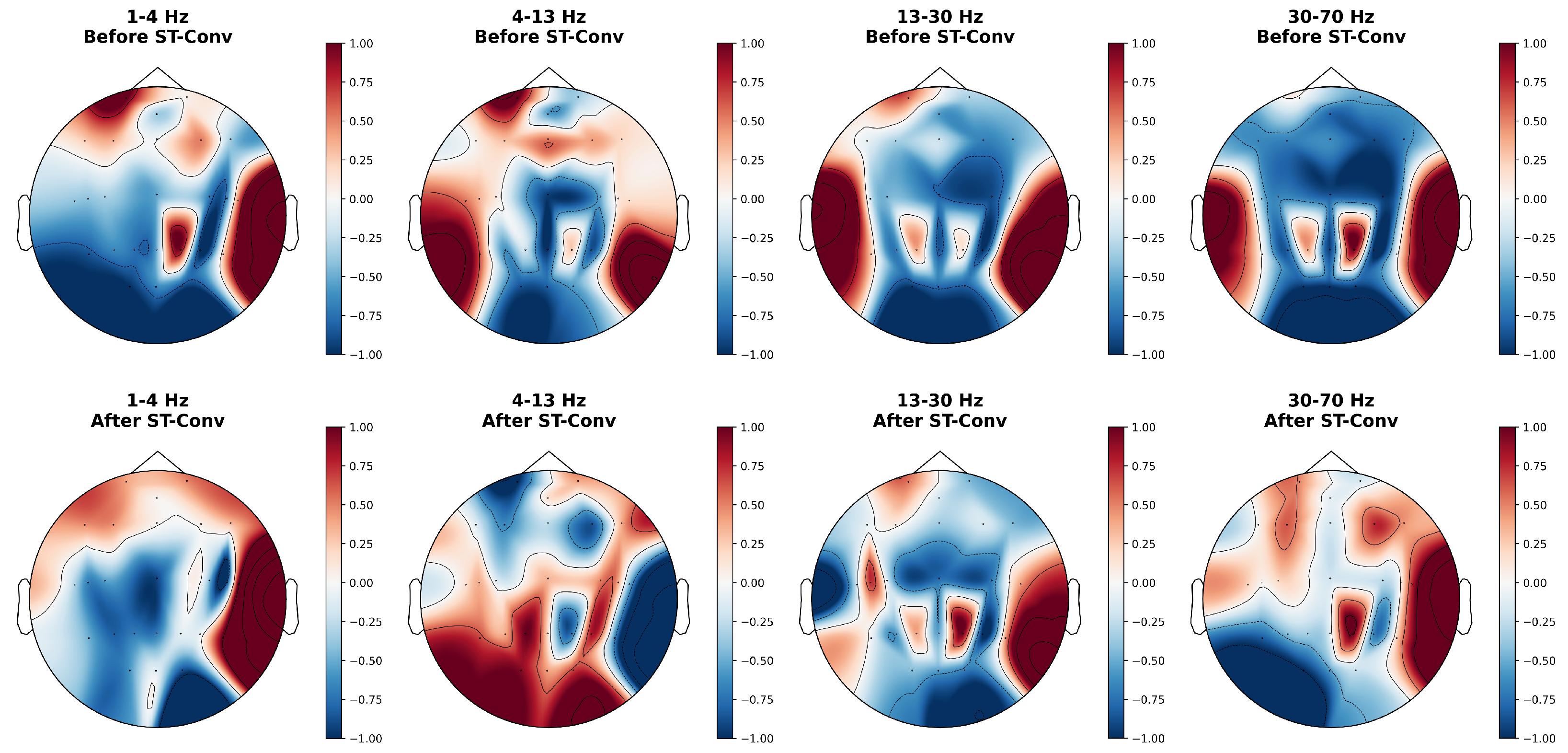
4.6. Feature Visualization
5. Discussion
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
References
- Pavone, P.; Striano, P.; Falsaperla, R.; Pavone, L.; Ruggieri, M. Infantile spasms syndrome, West syndrome and related phenotypes: What we know in 2013. Brain Dev. 2014, 36, 739–751. [Google Scholar] [CrossRef]
- Specchio, N.; Wirrell, E.C.; Scheffer, I.E.; Nabbout, R.; Riney, K.; Samia, P.; Guerreiro, M.; Gwer, S.; Zuberi, S.M.; Wilmshurst, J.M.; et al. International League Against Epilepsy classification and definition of epilepsy syndromes with onset in childhood: Position paper by the ILAE Task Force on Nosology and Definitions. Epilepsia 2022, 63, 1398–1442. [Google Scholar] [CrossRef]
- Romero Milà, B.; Remakanthakurup Sindhu, K.; Mytinger, J.R.; Shrey, D.W.; Lopour, B.A. EEG biomarkers for the diagnosis and treatment of infantile spasms. Front. Neurol. 2022, 13, 960454. [Google Scholar] [CrossRef] [PubMed]
- Riikonen, R.S. Favourable prognostic factors with infantile spasms. Eur. J. Paediatr. Neurol. 2010, 14, 13–18. [Google Scholar] [CrossRef] [PubMed]
- Primec, Z.R.; Stare, J.; Neubauer, D. The risk of lower mental outcome in infantile spasms increases after three weeks of hypsarrhythmia duration. Epilepsia 2006, 47, 2202–2205. [Google Scholar] [CrossRef]
- Napuri, S.; Le Gall, E.; Dulac, O.; Chaperon, J.; Riou, F. Factors associated with treatment lag in infantile spasms. Dev. Med. Child Neurol. 2010, 52, 1164–1166. [Google Scholar] [CrossRef] [PubMed]
- Hussain, S.A.; Lay, J.; Cheng, E.; Weng, J.; Sankar, R.; Baca, C.B. Recognition of infantile spasms is often delayed: The ASSIST study. J. Pediatr. 2017, 190, 215–221. [Google Scholar] [CrossRef]
- Teplan, M. Fundamentals of EEG measurement. Meas. Sci. Rev. 2002, 2, 1–11. [Google Scholar]
- Watanabe, K.; Negoro, T.; Aso, K.; Matsumoto, A. Reappraisal of interictal electroencephalograms in infantile spasms. Epilepsia 1993, 34, 679–685. [Google Scholar] [CrossRef]
- Oostenveld, R.; Praamstra, P. The five percent electrode system for high-resolution EEG and ERP measurements. Clin. Neurophysiol. 2001, 112, 713–719. [Google Scholar] [CrossRef]
- Cao, J.; Chen, Y.; Zheng, R.; Cui, X.; Jiang, T.; Gao, F. DSMN-ESS: Dual-stream multitask network for epilepsy syndrome classification and seizure detection. IEEE Trans. Instrum. Meas. 2023, 72, 1–12. [Google Scholar] [CrossRef]
- Guo, Y.; Jiang, X.; Tao, L.; Meng, L.; Dai, C.; Long, X.; Wan, F.; Zhang, Y.; Van Dijk, J.; Aarts, R.M.; et al. Epileptic seizure detection by cascading isolation forest-based anomaly screening and EasyEnsemble. IEEE Trans. Neural Syst. Rehabil. Eng. 2022, 30, 915–924. [Google Scholar] [CrossRef]
- Zheng, R.; Feng, Y.; Wang, T.; Cao, J.; Wu, D.; Jiang, T.; Gao, F. Scalp EEG functional connection and brain network in infants with West syndrome. Neural Netw. 2022, 153, 76–86. [Google Scholar] [CrossRef]
- Zhang, F.; Li, P.; Hou, Z.G.; Lu, Z.; Chen, Y.; Li, Q.; Tan, M. sEMG-based continuous estimation of joint angles of human legs by using BP neural network. Neurocomputing 2012, 78, 139–148. [Google Scholar] [CrossRef]
- Xi, X.; Sun, Z.; Hua, X.; Yuan, C.; Zhao, Y.B.; Miran, S.M.; Luo, Z.; Lü, Z. Construction and analysis of cortical–muscular functional network based on EEG-EMG coherence using wavelet coherence. Neurocomputing 2021, 438, 248–258. [Google Scholar] [CrossRef]
- Siddiqui, M.K.; Morales-Menendez, R.; Huang, X.; Hussain, N. A review of epileptic seizure detection using machine learning classifiers. Brain Inform. 2020, 7, 5. [Google Scholar] [CrossRef]
- Shen, M.; Wen, P.; Song, B.; Li, Y. An EEG based real-time epilepsy seizure detection approach using discrete wavelet transform and machine learning methods. Biomed. Signal Process. Control 2022, 77, 103820. [Google Scholar] [CrossRef]
- Wei, B.; Zhao, X.; Shi, L.; Xu, L.; Liu, T.; Zhang, J. A deep learning framework with multi-perspective fusion for interictal epileptiform discharges detection in scalp electroencephalogram. J. Neural Eng. 2021, 18, 0460b3. [Google Scholar] [CrossRef]
- Pan, Y.; Zhou, X.; Dong, F.; Wu, J.; Xu, Y.; Zheng, S. Epileptic seizure detection with hybrid time-frequency EEG input: A deep learning approach. Comput. Math. Methods Med. 2022, 2022, 8724536. [Google Scholar] [CrossRef]
- Liu, G.; Tian, L.; Wen, Y.; Yu, W.; Zhou, W. Cosine convolutional neural network and its application for seizure detection. Neural Netw. 2024, 174, 106267. [Google Scholar] [CrossRef]
- Song, Y.; Zheng, Q.; Liu, B.; Gao, X. EEG conformer: Convolutional transformer for EEG decoding and visualization. IEEE Trans. Neural Syst. Rehabil. Eng. 2022, 31, 710–719. [Google Scholar] [CrossRef]
- Miao, Z.; Zhao, M.; Zhang, X.; Ming, D. LMDA-Net: A lightweight multi-dimensional attention network for general EEG-based brain-computer interfaces and interpretability. NeuroImage 2023, 276, 120209. [Google Scholar] [CrossRef]
- Yan, S.; Fan, C.; Zhang, H.; Yang, X.; Tao, J.; Lv, Z. Darnet: Dual attention refinement network with spatiotemporal construction for auditory attention detection. Adv. Neural Inf. Process. Syst. 2024, 37, 31688–31707. [Google Scholar]
- Zhu, R.; Pan, W.X.; Liu, J.X.; Shang, J.L. Epileptic seizure prediction via multidimensional transformer and recurrent neural network fusion. J. Transl. Med. 2024, 22, 895. [Google Scholar] [CrossRef]
- Lu, G.; Peng, J.; Huang, B.; Gao, C.; Stefanov, T.; Hao, Y.; Chen, Q. SlimSeiz: Efficient Channel-Adaptive Seizure Prediction Using a Mamba-Enhanced Network. In Proceedings of the 2025 IEEE International Symposium on Circuits and Systems (ISCAS), Qingdao, China, 26–27 October 2025; pp. 1–5. [Google Scholar]
- Verma, G.K.; Tiwary, U.S. Multimodal fusion framework: A multiresolution approach for emotion classification and recognition from physiological signals. NeuroImage 2014, 102, 162–172. [Google Scholar] [CrossRef]
- Al-Quraishi, M.S.; Elamvazuthi, I.; Tang, T.B.; Al-Qurishi, M.; Parasuraman, S.; Borboni, A. Multimodal fusion approach based on EEG and EMG signals for lower limb movement recognition. IEEE Sens. J. 2021, 21, 27640–27650. [Google Scholar] [CrossRef]
- Kim, S.; Shin, D.Y.; Kim, T.; Lee, S.; Hyun, J.K.; Park, S.M. Enhanced recognition of amputated wrist and hand movements by deep learning method using multimodal fusion of electromyography and electroencephalography. Sensors 2022, 22, 680. [Google Scholar] [CrossRef]
- Bhatlawande, S.; Shilaskar, S.; Pramanik, S.; Sole, S. Multimodal emotion recognition based on the fusion of vision, EEG, ECG, and EMG signals. Int. J. Electr. Comput. Eng. Syst. 2024, 15, 41–58. [Google Scholar] [CrossRef]
- Wu, D.; Zhang, W.; Jiang, L.; Zhang, L.; Vidal, P.P.; Wang, D.; Cao, J.; Jiang, T. Optimization of EEG-EMG Fusion Network for West Syndrome Seizure Detection Based on Enhanced Artificial Rabbit Algorithm. IEEE Trans. Instrum. Meas. 2024, 73, 4010113. [Google Scholar] [CrossRef]
- Cui, R.; Chen, W.; Li, M. Emotion recognition using cross-modal attention from EEG and facial expression. Knowl.-Based Syst. 2024, 304, 112587. [Google Scholar] [CrossRef]
- Sitnikova, E.; Hramov, A.E.; Koronovsky, A.A.; Van Luijtelaar, G. Sleep spindles and spike–wave discharges in EEG: Their generic features, similarities and distinctions disclosed with Fourier transform and continuous wavelet analysis. J. Neurosci. Methods 2009, 180, 304–316. [Google Scholar] [CrossRef]
- Wu, F.; Mai, W.; Tang, Y.; Liu, Q.; Chen, J.; Guo, Z. Learning spatial-spectral-temporal EEG representations with deep attentive-recurrent-convolutional neural networks for pain intensity assessment. Neuroscience 2022, 481, 144–155. [Google Scholar] [CrossRef]
- Gu, A.; Dao, T. Mamba: Linear-time sequence modeling with selective state spaces. arXiv 2023, arXiv:2312.00752. [Google Scholar] [CrossRef]
- Peng, B.; Alcaide, E.; Anthony, Q.; Albalak, A.; Arcadinho, S.; Biderman, S.; Cao, H.; Cheng, X.; Chung, M.; Grella, M.; et al. Rwkv: Reinventing rnns for the transformer era. arXiv 2023, arXiv:2305.13048. [Google Scholar] [CrossRef]
- Zhang, B.; Sennrich, R. Root mean square layer normalization. Adv. Neural Inf. Process. Syst. 2019, 32. [Google Scholar]
- Asano, E.; Juhász, C.; Shah, A.; Muzik, O.; Chugani, D.C.; Shah, J.; Sood, S.; Chugani, H.T. Origin and propagation of epileptic spasms delineated on electrocorticography. Epilepsia 2005, 46, 1086–1097. [Google Scholar] [CrossRef]



| ID | Gender | Age | Seizure Count | Seizure Time (s) |
|---|---|---|---|---|
| a | Female | 1y6m | 156 | 205.1 |
| c | Male | 10m | 62 | 148.3 |
| d | Female | 11m | 86 | 87.1 |
| f | Male | 10m | 357 | 488.4 |
| g | Male | 2y4m | 29 | 25.5 |
| h | Female | 11m | 52 | 60.1 |
| i | Female | 1y5m | 468 | 580.7 |
| l | Female | 5m | 338 | 297.6 |
| m | Male | 3y | 139 | 139.8 |
| n | Male | 4y | 254 | 449.1 |
| Attribute | Value |
|---|---|
| Total Seizure Events | 1941 |
| Total Seizure Duration | 2481.7 s |
| Shortest/Longest Duration | 0.4 s/9.2 s |
| Number of Recordings | 129 EEG-EMG recordings |
| Total Recording Duration | 630 min |
| Acquisition Device | Compumedics Grael |
| Sampling Rate | 1024 Hz |
| Electrode Type | Disk electrodes |
| Number of Electrodes | 25 (EEG), 4 (EMG) |
| Placement System | International 10–20 system |
| Recording Environment | Hospital epilepsy monitoring ward |
| Method | Acc (%) | Spe (%) | Sen (%) |
|---|---|---|---|
| SOTA Methods Comparison | |||
| EEGConformer [21] | 77.5 ± 2.1 | 82.1 ± 1.6 | 72.9 ± 4.9 |
| CosCNN [20] | 81.5 ± 0.8 | 88.7 ± 1.8 | 74.4 ± 2.2 |
| LMDANET [22] | 84.3 ± 1.0 | 91.4 ± 1.1 | 77.2 ± 1.3 |
| DARNet [23] | 81.2 ± 0.9 | 86.7 ± 2.3 | 75.7 ± 2.6 |
| IESS-FusionNet | 89.5 ± 0.7 | 90.7 ± 1.4 | 88.3 ± 2.3 |
| Fusion Strategy Comparison | |||
| Concatenation | 87.3 ± 1.2 | 92.7 ± 0.7 | 81.8 ± 3.1 |
| Averaging | 88.2 ± 0.8 | 90.6 ± 2.1 | 85.8 ± 1.9 |
| Cross-Attention | 88.1 ± 1.2 | 89.9 ± 0.9 | 86.3 ± 2.2 |
| Cross Time-Mixing | 89.5 ± 0.7 | 90.7 ± 1.4 | 88.3 ± 2.3 |
| Modality Comparison | |||
| EEG-only | 86.9 ± 1.6 | 88.3 ± 3.5 | 85.6 ± 1.5 |
| EMG-only | 62.4 ± 0.7 | 76.7 ± 5.8 | 48.1 ± 7.0 |
| EEG + EMG | 89.5 ± 0.7 | 90.7 ± 1.4 | 88.3 ± 2.3 |
| Configuration | Acc (%) | Spe (%) | Sen (%) |
|---|---|---|---|
| Full Encoder | 89.5 ± 0.7 | 90.7 ± 1.4 | 88.3 ± 2.3 |
| w/o CWT | 81.8 ± 1.7 | 89.3 ± 1.7 | 74.3 ± 2.3 |
| w/o ST-Conv | 85.6 ± 1.2 | 88.2 ± 2.5 | 83.0 ± 0.7 |
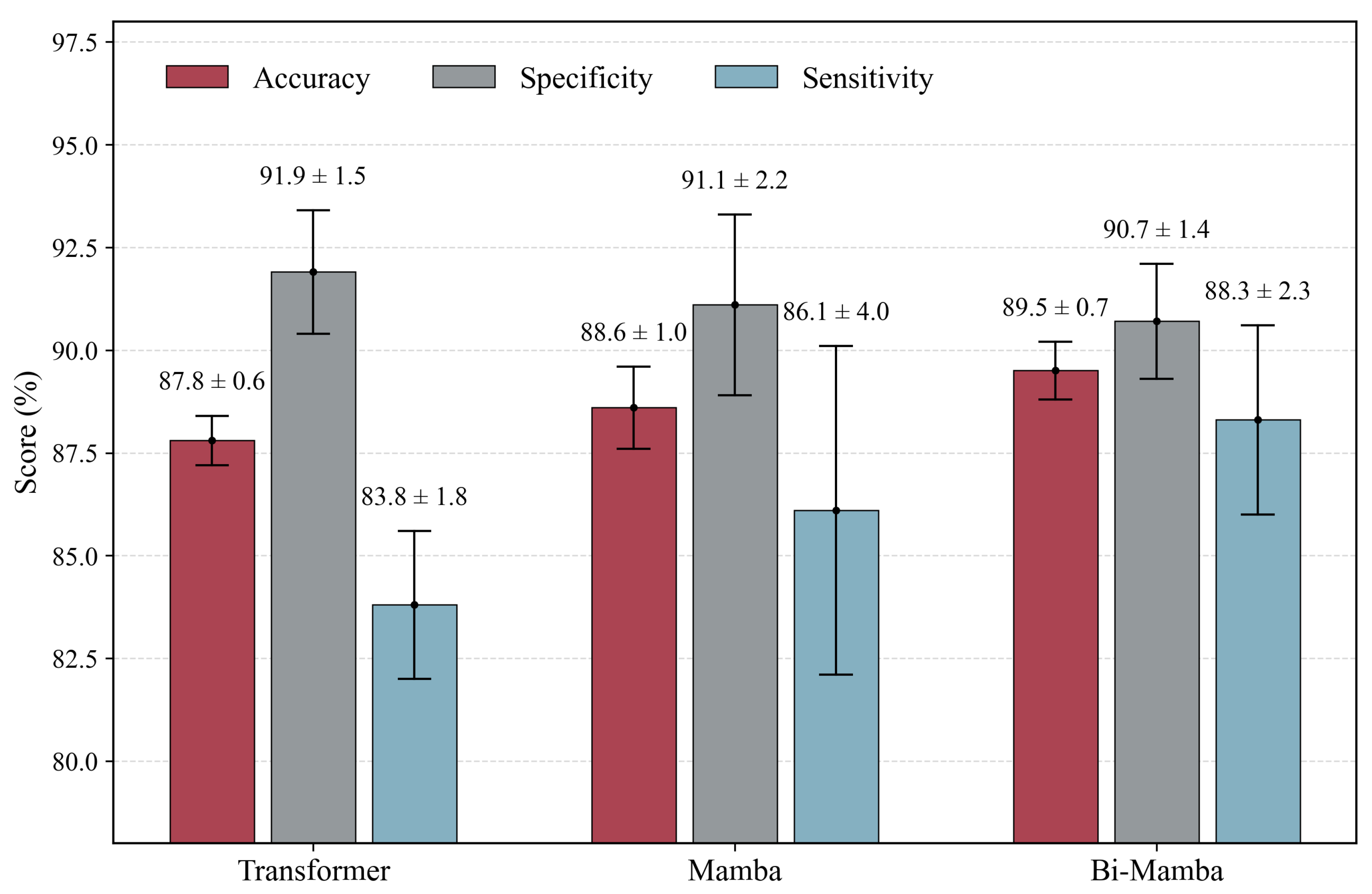
| w/o Bi-Mamba | 87.7 ± 2.2 | 91.9 ± 1.0 | 83.6 ± 4.4 |
| Component | Params (M) | FLOPs (G) |
|---|---|---|
| Global Sequence Modeling | ||
| Transformer | 0.78 | 0.80 |
| Mamba | 0.17 | 0.08 |
| Bi-Mamba | 0.25 | 0.22 |
| Cross-Modal Fusion | ||
| Cross-Attention | 0.58 | 1.43 |
| Cross Time-Mixing | 0.23 | 0.75 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Feng, J.; Liu, Z.; Shen, L.; Luo, X.; Chen, Y.; Li, L.; Zhang, T. IESS-FusionNet: Physiologically Inspired EEG-EMG Fusion with Linear Recurrent Attention for Infantile Epileptic Spasms Syndrome Detection. Bioengineering 2026, 13, 57. https://doi.org/10.3390/bioengineering13010057
Feng J, Liu Z, Shen L, Luo X, Chen Y, Li L, Zhang T. IESS-FusionNet: Physiologically Inspired EEG-EMG Fusion with Linear Recurrent Attention for Infantile Epileptic Spasms Syndrome Detection. Bioengineering. 2026; 13(1):57. https://doi.org/10.3390/bioengineering13010057
Chicago/Turabian StyleFeng, Junyuan, Zhenzhen Liu, Linlin Shen, Xiaoling Luo, Yan Chen, Lin Li, and Tian Zhang. 2026. "IESS-FusionNet: Physiologically Inspired EEG-EMG Fusion with Linear Recurrent Attention for Infantile Epileptic Spasms Syndrome Detection" Bioengineering 13, no. 1: 57. https://doi.org/10.3390/bioengineering13010057
APA StyleFeng, J., Liu, Z., Shen, L., Luo, X., Chen, Y., Li, L., & Zhang, T. (2026). IESS-FusionNet: Physiologically Inspired EEG-EMG Fusion with Linear Recurrent Attention for Infantile Epileptic Spasms Syndrome Detection. Bioengineering, 13(1), 57. https://doi.org/10.3390/bioengineering13010057







