Figure 1.
Workflow of the proposed pipeline. The green square from the input CFI represents the selected fiducial marker, which was implanted around the tumor to assist in determining the tumor location. In this study, we used the optical flow method to obtain the motion trajectory of the selected fiducial marker, which was used for comparison against the predicted tumor trajectory.
Figure 1.
Workflow of the proposed pipeline. The green square from the input CFI represents the selected fiducial marker, which was implanted around the tumor to assist in determining the tumor location. In this study, we used the optical flow method to obtain the motion trajectory of the selected fiducial marker, which was used for comparison against the predicted tumor trajectory.
Figure 2.
Gelatin cube preparation and image acquisition: (left) container with gelatin inside; (right) schematic diagram of image acquisition site, the gelatin container was placed at the ISO center.
Figure 2.
Gelatin cube preparation and image acquisition: (left) container with gelatin inside; (right) schematic diagram of image acquisition site, the gelatin container was placed at the ISO center.
Figure 3.
Captured X-ray images of the gelatin cube: (a) an X-ray fluoroscopic image captured by the SyncTraX system, grayscaled using CorrC2G; (b) clean image calculated by averaging 1800 images; (c) brightness contour map of the average image. The brightness contour map (c) shows that the brightness center of the image is shifted slightly to the left and does not overlap with the center of the image, which indicates that the central axis of the X-ray beam is offset. Since the gelatin container was placed at the ISO center and its shape is not circular, it is unlikely that this brightness deviation is caused by the container.
Figure 3.
Captured X-ray images of the gelatin cube: (a) an X-ray fluoroscopic image captured by the SyncTraX system, grayscaled using CorrC2G; (b) clean image calculated by averaging 1800 images; (c) brightness contour map of the average image. The brightness contour map (c) shows that the brightness center of the image is shifted slightly to the left and does not overlap with the center of the image, which indicates that the central axis of the X-ray beam is offset. Since the gelatin container was placed at the ISO center and its shape is not circular, it is unlikely that this brightness deviation is caused by the container.
Figure 4.
Noise characteristics in a gelatin cube fluoroscopic image (background pixels excluded). (a) Heatmap of degradation-induced noise probability and (b) amplitude, derived by computing the pixel-wise variance across 1800 subtracted images (individual frames minus the average of all frames) and normalizing it to a range. In (a), the mean noise probability within the central circular region is approximately 0.56, while that in the right peripheral arc region reaches about 0.83. In (b), the mean noise amplitude in the central circular region is approximately 2.7, increasing to around 7.2 in the right peripheral arc. Regions with lower grayscale values in the image, such as the cube’s edges, exhibit lower noise probability and amplitude, and the farther the area is from the center of the X-ray beam (note: the beam center, not the image center), the higher the noise probability and amplitude. Importantly, the empirical profile is not strictly monotonic: noise probability and amplitude increase with radial distance from the beam center until approximately one radius, after which a decline is observed. This decline becomes noticeable primarily on the right-hand side of the image, due to a slight leftward shift of the X-ray beam center, which exposes more of the far-right region beyond the effective beam axis. Here, the radius refers specifically to the radius of the circular foreground region in the image.
Figure 4.
Noise characteristics in a gelatin cube fluoroscopic image (background pixels excluded). (a) Heatmap of degradation-induced noise probability and (b) amplitude, derived by computing the pixel-wise variance across 1800 subtracted images (individual frames minus the average of all frames) and normalizing it to a range. In (a), the mean noise probability within the central circular region is approximately 0.56, while that in the right peripheral arc region reaches about 0.83. In (b), the mean noise amplitude in the central circular region is approximately 2.7, increasing to around 7.2 in the right peripheral arc. Regions with lower grayscale values in the image, such as the cube’s edges, exhibit lower noise probability and amplitude, and the farther the area is from the center of the X-ray beam (note: the beam center, not the image center), the higher the noise probability and amplitude. Importantly, the empirical profile is not strictly monotonic: noise probability and amplitude increase with radial distance from the beam center until approximately one radius, after which a decline is observed. This decline becomes noticeable primarily on the right-hand side of the image, due to a slight leftward shift of the X-ray beam center, which exposes more of the far-right region beyond the effective beam axis. Here, the radius refers specifically to the radius of the circular foreground region in the image.
![Bioengineering 12 01197 g004 Bioengineering 12 01197 g004]()
Figure 5.
(
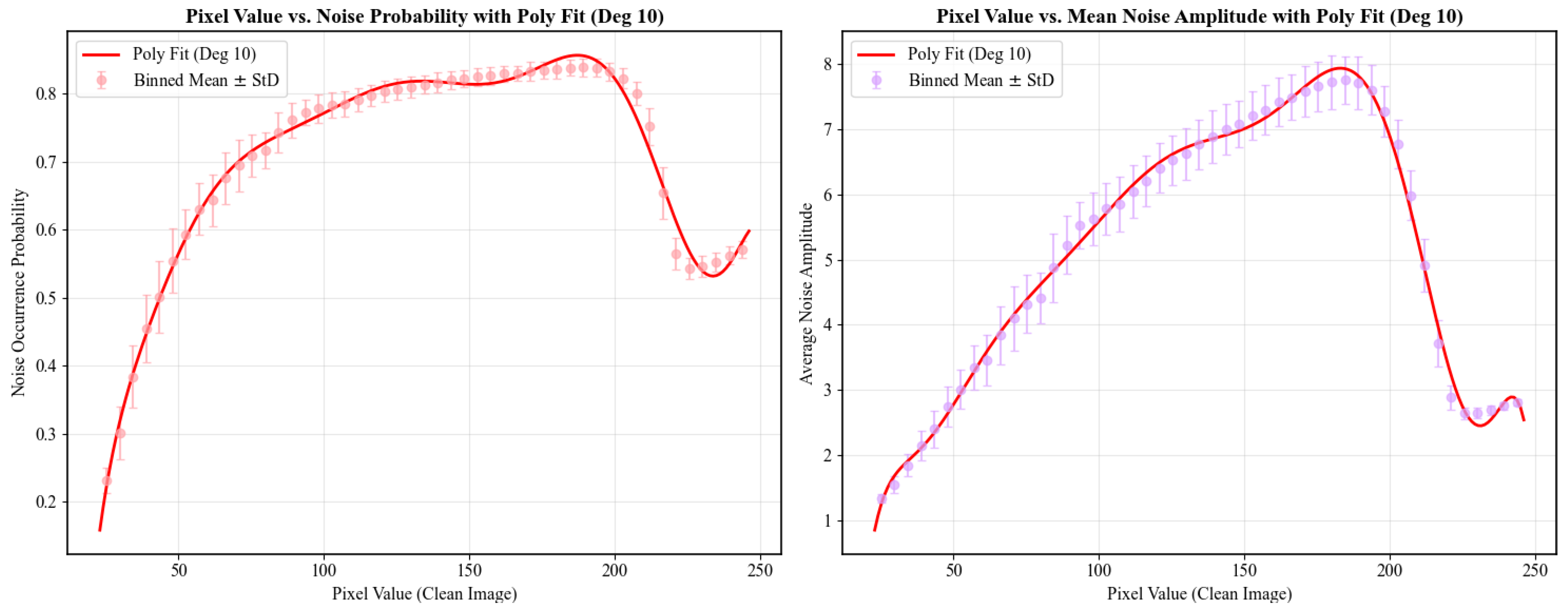
a) Relationship between pixel grayscale value and noise probability. (
b) Relationship between pixel grayscale value and noise amplitude. From a holistic perspective, the relationship can be broadly divided into four phases: rapid ascent phase (pixel values between 0 and 85), slow ascent phase (pixel values between 85 and 190, the average noise probability and amplitude are around 0.81 and 6.5, respectively), sharp decline phase (pixel values between 190 and 225), and recovery phase (pixel values between 225 and 255, the avearge noise probability and amplitude are around 0.56 and 2.75, respectively). The sharp decline phase is likely attributable to the clipping of high-amplitude noise fluctuations that exceed the upper grayscale limit of 255, while the recovery phase corresponds to the re-elevation of the overall noise level by low-amplitude noise. Given the presence of various types of noise [
35,
36] in X-ray imaging—such as quantum noise, electronic noise and structural noise from the imaging device—which exhibit different frequencies and amplitudes, this may account for the observed four-phase relationship.
Figure 5.
(
a) Relationship between pixel grayscale value and noise probability. (
b) Relationship between pixel grayscale value and noise amplitude. From a holistic perspective, the relationship can be broadly divided into four phases: rapid ascent phase (pixel values between 0 and 85), slow ascent phase (pixel values between 85 and 190, the average noise probability and amplitude are around 0.81 and 6.5, respectively), sharp decline phase (pixel values between 190 and 225), and recovery phase (pixel values between 225 and 255, the avearge noise probability and amplitude are around 0.56 and 2.75, respectively). The sharp decline phase is likely attributable to the clipping of high-amplitude noise fluctuations that exceed the upper grayscale limit of 255, while the recovery phase corresponds to the re-elevation of the overall noise level by low-amplitude noise. Given the presence of various types of noise [
35,
36] in X-ray imaging—such as quantum noise, electronic noise and structural noise from the imaging device—which exhibit different frequencies and amplitudes, this may account for the observed four-phase relationship.
![Bioengineering 12 01197 g005 Bioengineering 12 01197 g005]()
Figure 6.
Fitting results using a 10th-order polynomial.
Figure 6.
Fitting results using a 10th-order polynomial.
Figure 7.
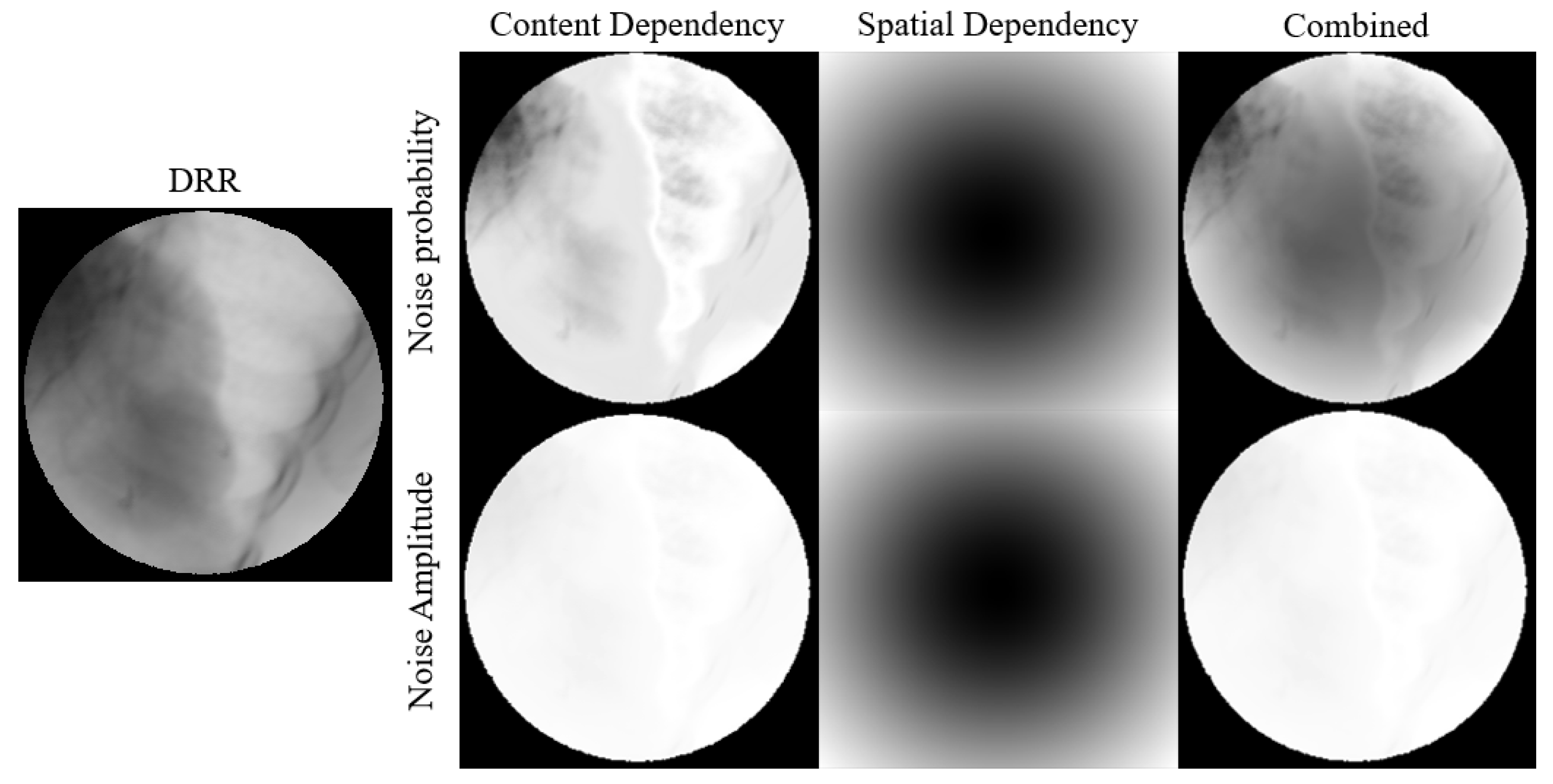
Visual representation of how pixel values and spatial position relationship affects noise probability and amplitude.
Figure 7.
Visual representation of how pixel values and spatial position relationship affects noise probability and amplitude.
Figure 8.
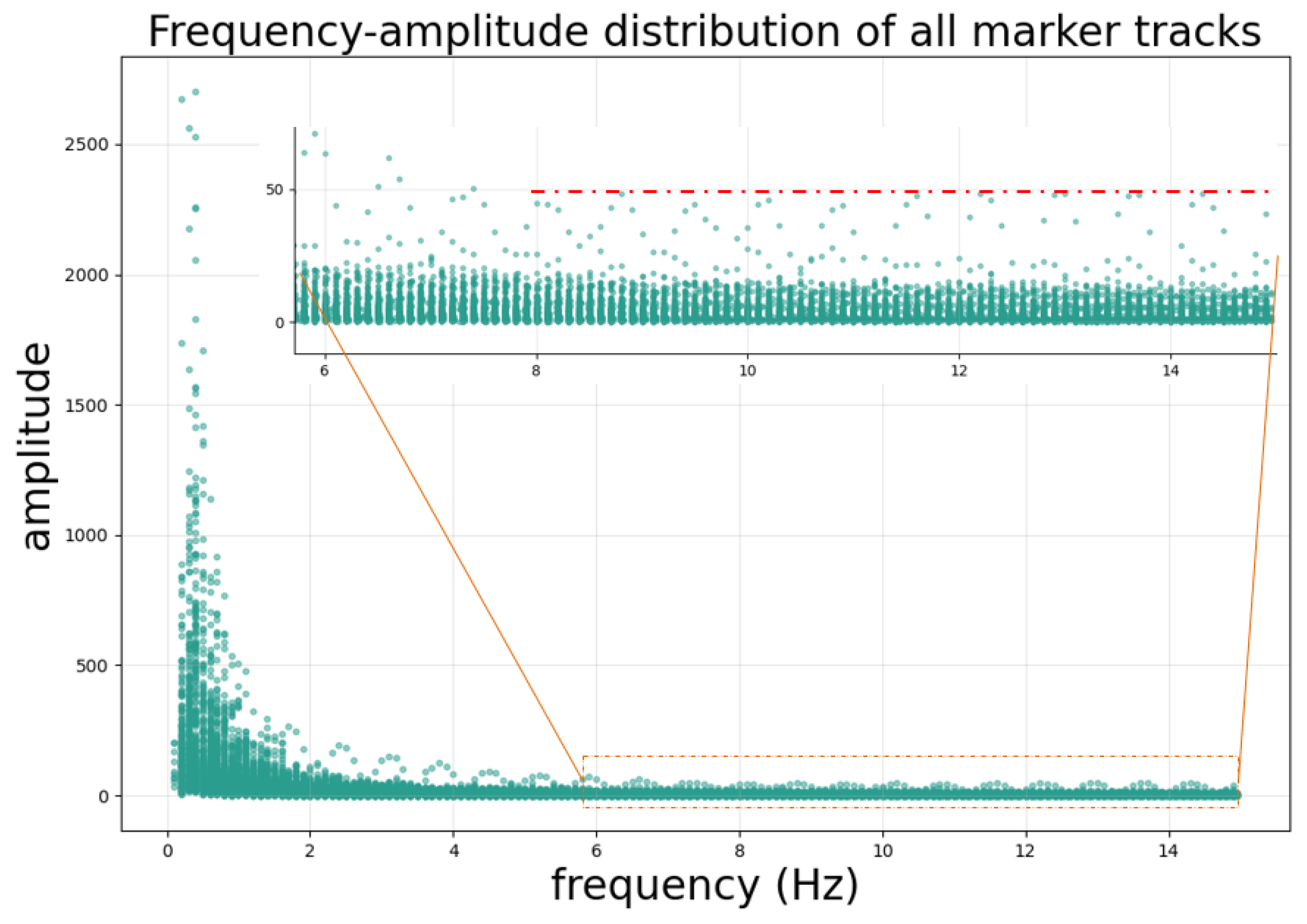
Frequency–amplitude distribution of the fiducial marker motion trajectory. Frequency analysis reveals the typical harmonic structure of respiratory motion, with a fundamental frequency of approximately 0.33 Hz (20 breaths/min) and multiple harmonics extending up to 15 Hz. Notably, the amplitude of harmonic components above 8 Hz remains stable at a low level (amplitude below 50 as indicated by the red dashed line). Based on this observation, we selected 8 Hz as the cutoff frequency to retain physiologically relevant motion components while filtering out trajectory jitter introduced by segmentation errors in individual frames, which corresponds to the high-frequency components.
Figure 8.
Frequency–amplitude distribution of the fiducial marker motion trajectory. Frequency analysis reveals the typical harmonic structure of respiratory motion, with a fundamental frequency of approximately 0.33 Hz (20 breaths/min) and multiple harmonics extending up to 15 Hz. Notably, the amplitude of harmonic components above 8 Hz remains stable at a low level (amplitude below 50 as indicated by the red dashed line). Based on this observation, we selected 8 Hz as the cutoff frequency to retain physiologically relevant motion components while filtering out trajectory jitter introduced by segmentation errors in individual frames, which corresponds to the high-frequency components.
Figure 9.
Fiducial marker expansion and matching method: (a) Illustration of the fiducial marker expansion process in CT data: the left panel shows the original CT slice with a red circle highlighting the target fiducial marker; the right panel shows the same slice after expanding the marker region into a spherical volume (5 mm radius, 3000 HU). (b) Fiducial marker matching between DRR and clinical fluoroscopic image: the upper row shows DRR images generated with identical geometric parameters to the actual imaging setup; the lower row shows the corresponding clinically acquired CFIs. The green circles indicate the identified target fiducial marker in the CFIs. Arrows in both DRR and CFIs point to other surrounding fiducial markers, demonstrating successful spatial correspondence between the expanded marker region in DRRs and the actual marker position in CFIs.
Figure 9.
Fiducial marker expansion and matching method: (a) Illustration of the fiducial marker expansion process in CT data: the left panel shows the original CT slice with a red circle highlighting the target fiducial marker; the right panel shows the same slice after expanding the marker region into a spherical volume (5 mm radius, 3000 HU). (b) Fiducial marker matching between DRR and clinical fluoroscopic image: the upper row shows DRR images generated with identical geometric parameters to the actual imaging setup; the lower row shows the corresponding clinically acquired CFIs. The green circles indicate the identified target fiducial marker in the CFIs. Arrows in both DRR and CFIs point to other surrounding fiducial markers, demonstrating successful spatial correspondence between the expanded marker region in DRRs and the actual marker position in CFIs.
Figure 10.
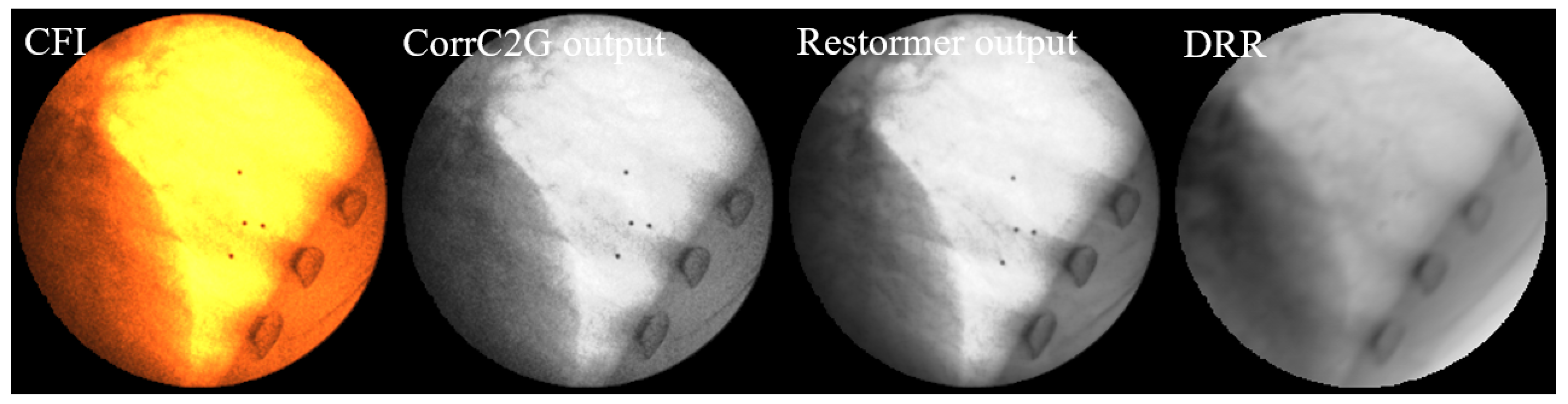
Comparison of image processing results at different stages. From (left) to (right): Original CFI; CorrC2G output; Restormer output; DRR: generated with identical imaging geometry. The CorrC2G algorithm preserves contrast information while Restormer effectively suppresses noise and blur, producing image characteristics closer to those of DRR images.
Figure 10.
Comparison of image processing results at different stages. From (left) to (right): Original CFI; CorrC2G output; Restormer output; DRR: generated with identical imaging geometry. The CorrC2G algorithm preserves contrast information while Restormer effectively suppresses noise and blur, producing image characteristics closer to those of DRR images.
Figure 11.
Visualization comparison of the image degradation model. From (left) to (right): DRR, degraded image with Gaussian noise, degraded image using the proposed method, and GFI. It can be seen that the degraded image generated by the proposed method can better simulate the noise pattern in GFI. It is notable that while the noise density is visibly reduced in the two highlighted areas of the GFI, the corresponding areas in our generated image, although also showing reduced noise, appear comparatively noisier. This discrepancy is because the corresponding areas in the source DRR are not as bright as those in the GFI, leading to a different noise expression after degradation.
Figure 11.
Visualization comparison of the image degradation model. From (left) to (right): DRR, degraded image with Gaussian noise, degraded image using the proposed method, and GFI. It can be seen that the degraded image generated by the proposed method can better simulate the noise pattern in GFI. It is notable that while the noise density is visibly reduced in the two highlighted areas of the GFI, the corresponding areas in our generated image, although also showing reduced noise, appear comparatively noisier. This discrepancy is because the corresponding areas in the source DRR are not as bright as those in the GFI, leading to a different noise expression after degradation.
Figure 12.
Visual comparison of denoising results using different methods on a sample X-ray fluoroscopic image. From (left) to (right): Grayscale fluoroscopic image (Input); Output from pre-trained DnCNN; Output from pre-trained Restormer (Restormer_Ori); Output from Restormer fine-tuned on baseline dataset (Restormer_Base); Output from Restormer fine-tuned on our proposed dataset (Restormer_Prop); Reference DRR. The results demonstrate that output from Restormer fine-tuned on our proposed dataset has better visual quality and closer resemblance to DRR-style images.
Figure 12.
Visual comparison of denoising results using different methods on a sample X-ray fluoroscopic image. From (left) to (right): Grayscale fluoroscopic image (Input); Output from pre-trained DnCNN; Output from pre-trained Restormer (Restormer_Ori); Output from Restormer fine-tuned on baseline dataset (Restormer_Base); Output from Restormer fine-tuned on our proposed dataset (Restormer_Prop); Reference DRR. The results demonstrate that output from Restormer fine-tuned on our proposed dataset has better visual quality and closer resemblance to DRR-style images.
Figure 13.
Focused comparison between the baseline method and our proposed approach. While both methods demonstrate effective noise suppression, the Restormer fine-tuned on our proposed dataset (left) preserves significantly more image details compared to the baseline method (right). This enhanced detail preservation contributes to better visual quality and a more accurate representation of anatomical structures.
Figure 13.
Focused comparison between the baseline method and our proposed approach. While both methods demonstrate effective noise suppression, the Restormer fine-tuned on our proposed dataset (left) preserves significantly more image details compared to the baseline method (right). This enhanced detail preservation contributes to better visual quality and a more accurate representation of anatomical structures.
Figure 14.
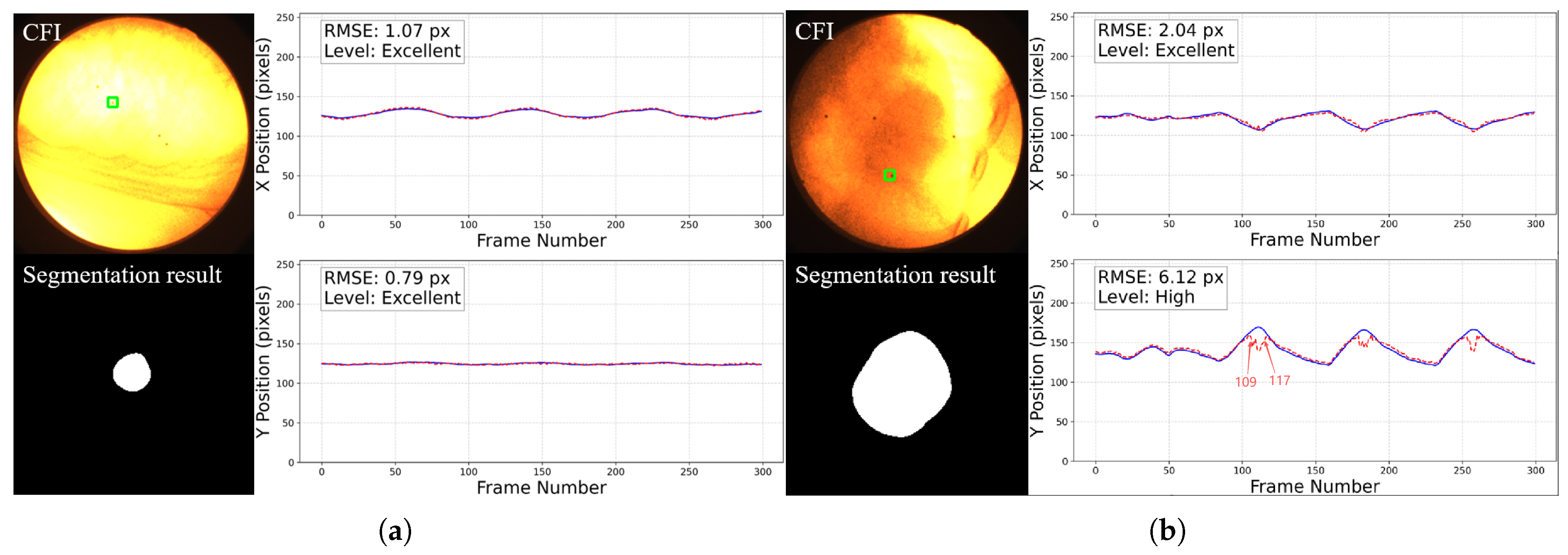
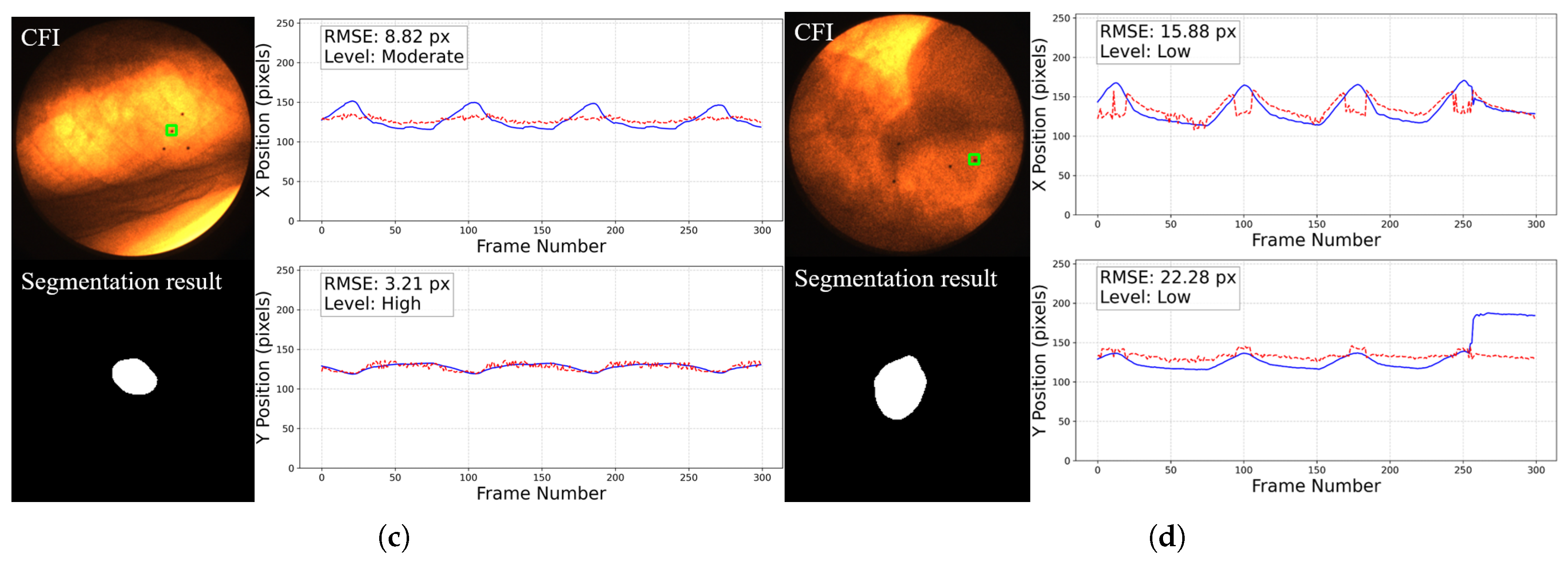
Representative cases of 2D motion trajectory comparison across different accuracy grades (LPF not applied). For each case: (Left) panel shows the workflow input CFI with a green square indicating the selected fiducial marker (top) and corresponding tumor segmentation result (bottom); (Right) panel displays the aligned temporal trajectories of tumor center (red dotted line) and fiducial marker (blue line) in image coordinates (x and y directions): (a) Excellent cases (50%) have near-perfect trajectory alignment. (b) High-accuracy cases (46%) show certain deviations in some frames (such as frame 109 to 117, we will discuss about it later) but the trajectory remains roughly aligned with the reference marker. This is common in cases where the image contrast is too poor, with severe noise and large tumor motion amplitude. (c) Moderate cases (3%) exhibit larger errors. (d) The only Low-graded case (1%) is caused by two factors: significant trajectory divergence in inspiratory frames and marker tracking error in the last 44 frames. The overall tumor detection rate is 100%.
Figure 14.
Representative cases of 2D motion trajectory comparison across different accuracy grades (LPF not applied). For each case: (Left) panel shows the workflow input CFI with a green square indicating the selected fiducial marker (top) and corresponding tumor segmentation result (bottom); (Right) panel displays the aligned temporal trajectories of tumor center (red dotted line) and fiducial marker (blue line) in image coordinates (x and y directions): (a) Excellent cases (50%) have near-perfect trajectory alignment. (b) High-accuracy cases (46%) show certain deviations in some frames (such as frame 109 to 117, we will discuss about it later) but the trajectory remains roughly aligned with the reference marker. This is common in cases where the image contrast is too poor, with severe noise and large tumor motion amplitude. (c) Moderate cases (3%) exhibit larger errors. (d) The only Low-graded case (1%) is caused by two factors: significant trajectory divergence in inspiratory frames and marker tracking error in the last 44 frames. The overall tumor detection rate is 100%.
![Bioengineering 12 01197 g014a Bioengineering 12 01197 g014a]()
![Bioengineering 12 01197 g014b Bioengineering 12 01197 g014b]()
Figure 15.
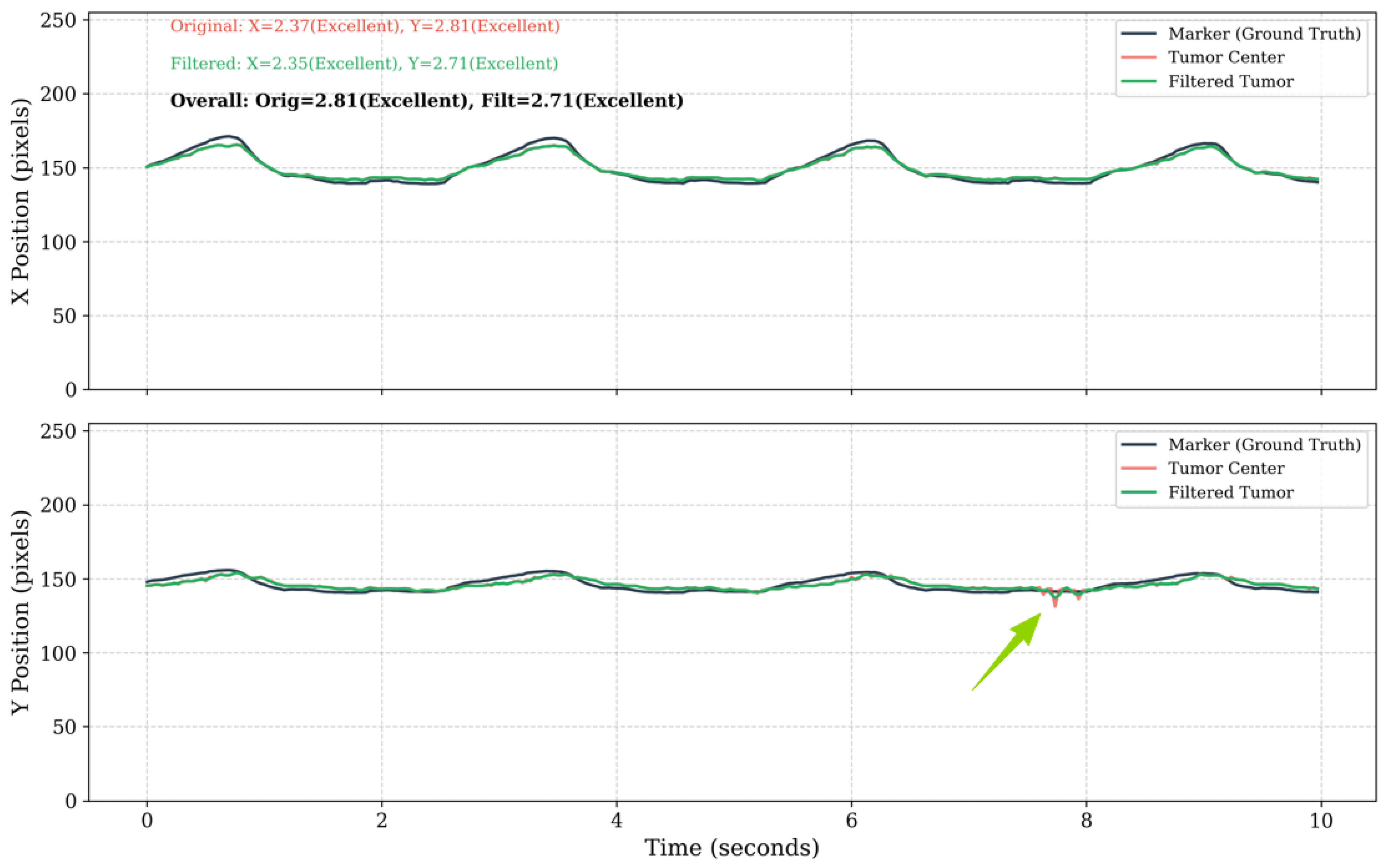
Comparison of filtered and unfiltered tumor trajectories. Unfiltered trajectory showing high-frequency jitter from segmentation error (red); Low-pass filtered trajectory (8 Hz cutoff) preserving the respiratory motion pattern while suppressing the jitter. The trajectory using the low-pass filter deviates less at the points indicated by the arrow.
Figure 15.
Comparison of filtered and unfiltered tumor trajectories. Unfiltered trajectory showing high-frequency jitter from segmentation error (red); Low-pass filtered trajectory (8 Hz cutoff) preserving the respiratory motion pattern while suppressing the jitter. The trajectory using the low-pass filter deviates less at the points indicated by the arrow.
Figure 16.
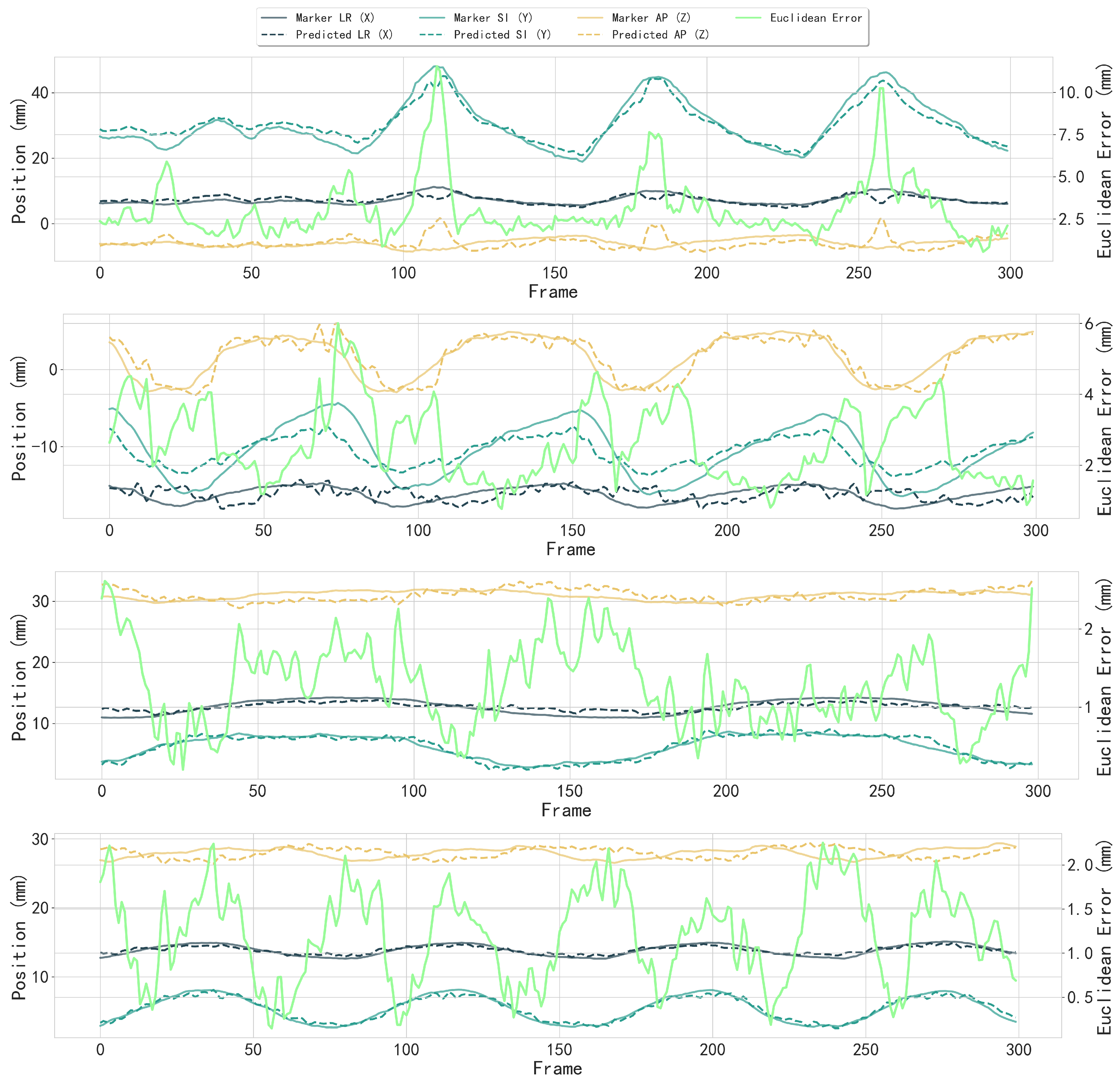
Partial results of 3D motion trajectory comparison between the fiducial marker (reference) and the tracked tumor center. Each subfigure shows individual trajectory components in LR (X), SI (Y), and AP (Z) directions with fiducial marker and detected tumor center; Euclidean distance error (bright green, right y-axis) between fiducial marker and tumor center across frames (x-axis). The median Euclidean error in all cases was 1.53 mm, with directional errors of mm (LR), mm (SI), and mm (AP).
Figure 16.
Partial results of 3D motion trajectory comparison between the fiducial marker (reference) and the tracked tumor center. Each subfigure shows individual trajectory components in LR (X), SI (Y), and AP (Z) directions with fiducial marker and detected tumor center; Euclidean distance error (bright green, right y-axis) between fiducial marker and tumor center across frames (x-axis). The median Euclidean error in all cases was 1.53 mm, with directional errors of mm (LR), mm (SI), and mm (AP).
Figure 17.
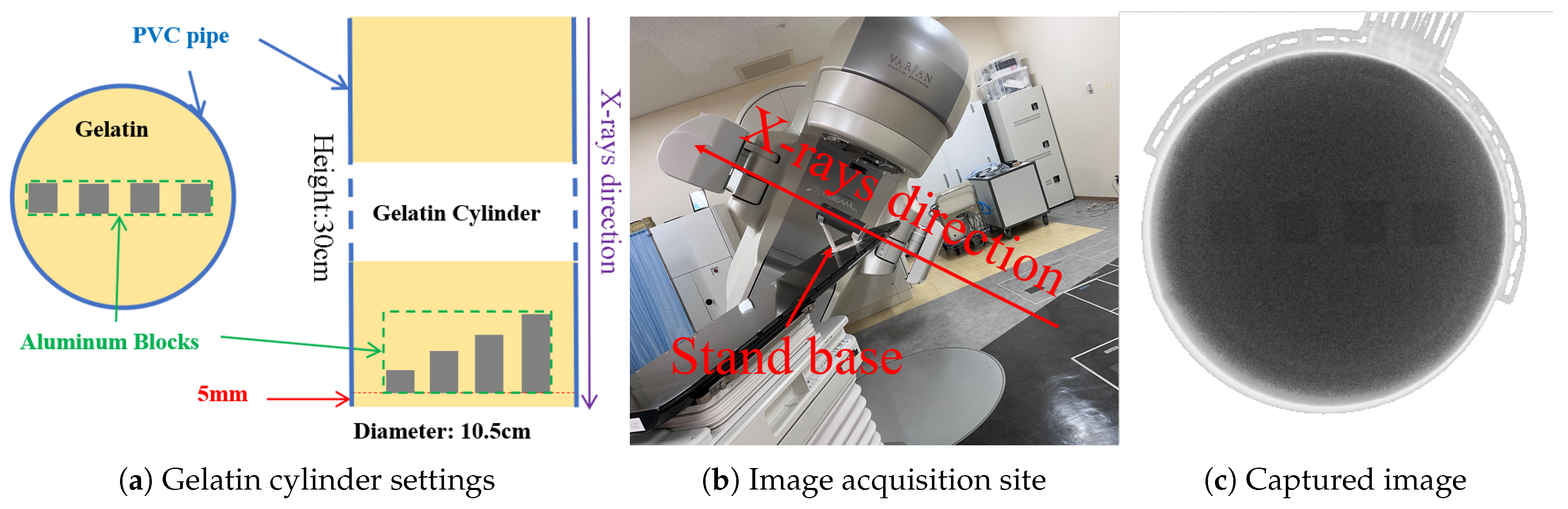
Gelatin preparation and image acquisition. The aluminum blocks are labeled `aluminum_1’ to `aluminum_4’ from left to right, with heights of 1, 2, 3, and 4 mm, respectively.
Figure 17.
Gelatin preparation and image acquisition. The aluminum blocks are labeled `aluminum_1’ to `aluminum_4’ from left to right, with heights of 1, 2, 3, and 4 mm, respectively.
Figure 18.
Noise characteristics in a gelatin cylinder fluoroscopic image: (a) Probability distribution map of degradation-induced noise, derived by computing the pixel-wise variance across 300 subtracted images (individual frames minus the average of all frames) and normalizing it to a [0, 1] range, where higher values indicate greater degradation likelihood. To account for the fact that the average image is not perfectly clean, a threshold of 5 grayscale values was applied, such that only differences exceeding this threshold were considered as noise. Square regions with embedded aluminum blocks show lower probabilities than gelatin areas, decreasing toward the image center within the circular field. (b) Heatmap of average noise amplitude across the image, calculated as the mean absolute difference between individual frames and the average image. The noise amplitude remains stable inside the gelatin area.
Figure 18.
Noise characteristics in a gelatin cylinder fluoroscopic image: (a) Probability distribution map of degradation-induced noise, derived by computing the pixel-wise variance across 300 subtracted images (individual frames minus the average of all frames) and normalizing it to a [0, 1] range, where higher values indicate greater degradation likelihood. To account for the fact that the average image is not perfectly clean, a threshold of 5 grayscale values was applied, such that only differences exceeding this threshold were considered as noise. Square regions with embedded aluminum blocks show lower probabilities than gelatin areas, decreasing toward the image center within the circular field. (b) Heatmap of average noise amplitude across the image, calculated as the mean absolute difference between individual frames and the average image. The noise amplitude remains stable inside the gelatin area.
Figure 19.
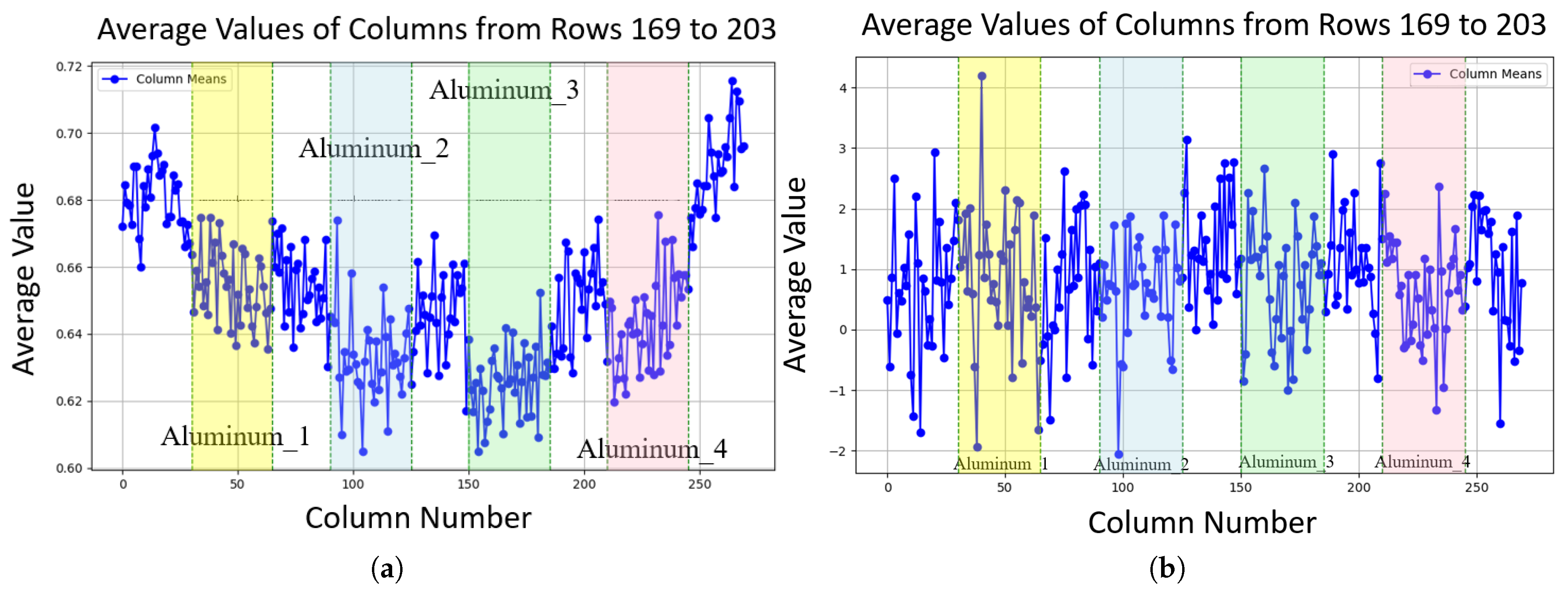
Line plots of noise characteristics along image columns, with edge pixels excluded to focus on internal variations: (a) Average probability along columns (rows 169 to 203 contain aluminum blocks); aluminum block regions exhibit lower probability (around 0.015 to 0.02 lower, depending on how close the pixel is to the center) compared to gelatin areas. (b) Average amplitude along columns (rows 169 to 203 contain aluminum blocks), showing stable amplitude in aluminum block regions compared to gelatin areas.
Figure 19.
Line plots of noise characteristics along image columns, with edge pixels excluded to focus on internal variations: (a) Average probability along columns (rows 169 to 203 contain aluminum blocks); aluminum block regions exhibit lower probability (around 0.015 to 0.02 lower, depending on how close the pixel is to the center) compared to gelatin areas. (b) Average amplitude along columns (rows 169 to 203 contain aluminum blocks), showing stable amplitude in aluminum block regions compared to gelatin areas.
Figure 20.
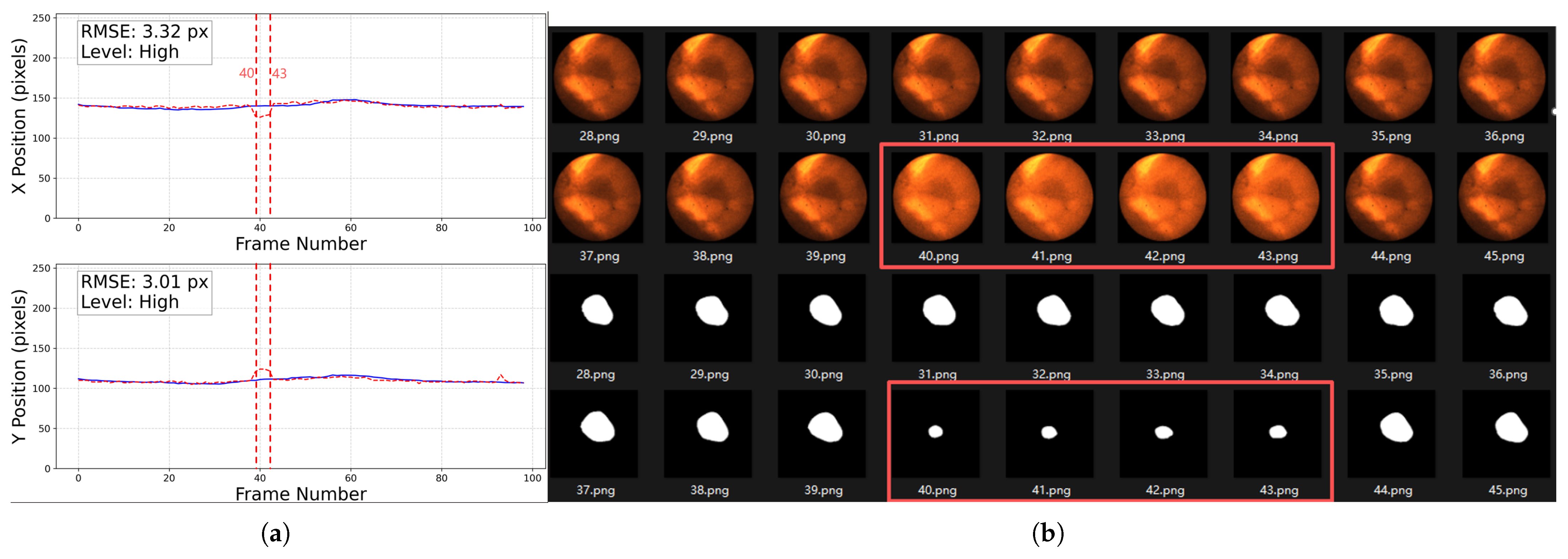
Abrupt variations in brightness and contrast. The left figure (a) shows a trajectory rated as High, the solid blue line represents the baseline trajectory, the dashed red line represents the predicted trajectory of the tumor center point. The tracking error markedly increases from frame 40 to 43 (as illustrated by the vertical dashed red lines). The right figure (b) presents the original CFIs and segmentation results, highlighting frames 40–43 (inside the red boxes) with strong brightness and contrast fluctuations. Note that the trajectory plot in (a) starts from frame 0, whereas the image filenames start from frame 1; therefore, frame 40 corresponds to position 39 in the trajectory plot.
Figure 20.
Abrupt variations in brightness and contrast. The left figure (a) shows a trajectory rated as High, the solid blue line represents the baseline trajectory, the dashed red line represents the predicted trajectory of the tumor center point. The tracking error markedly increases from frame 40 to 43 (as illustrated by the vertical dashed red lines). The right figure (b) presents the original CFIs and segmentation results, highlighting frames 40–43 (inside the red boxes) with strong brightness and contrast fluctuations. Note that the trajectory plot in (a) starts from frame 0, whereas the image filenames start from frame 1; therefore, frame 40 corresponds to position 39 in the trajectory plot.
Figure 21.
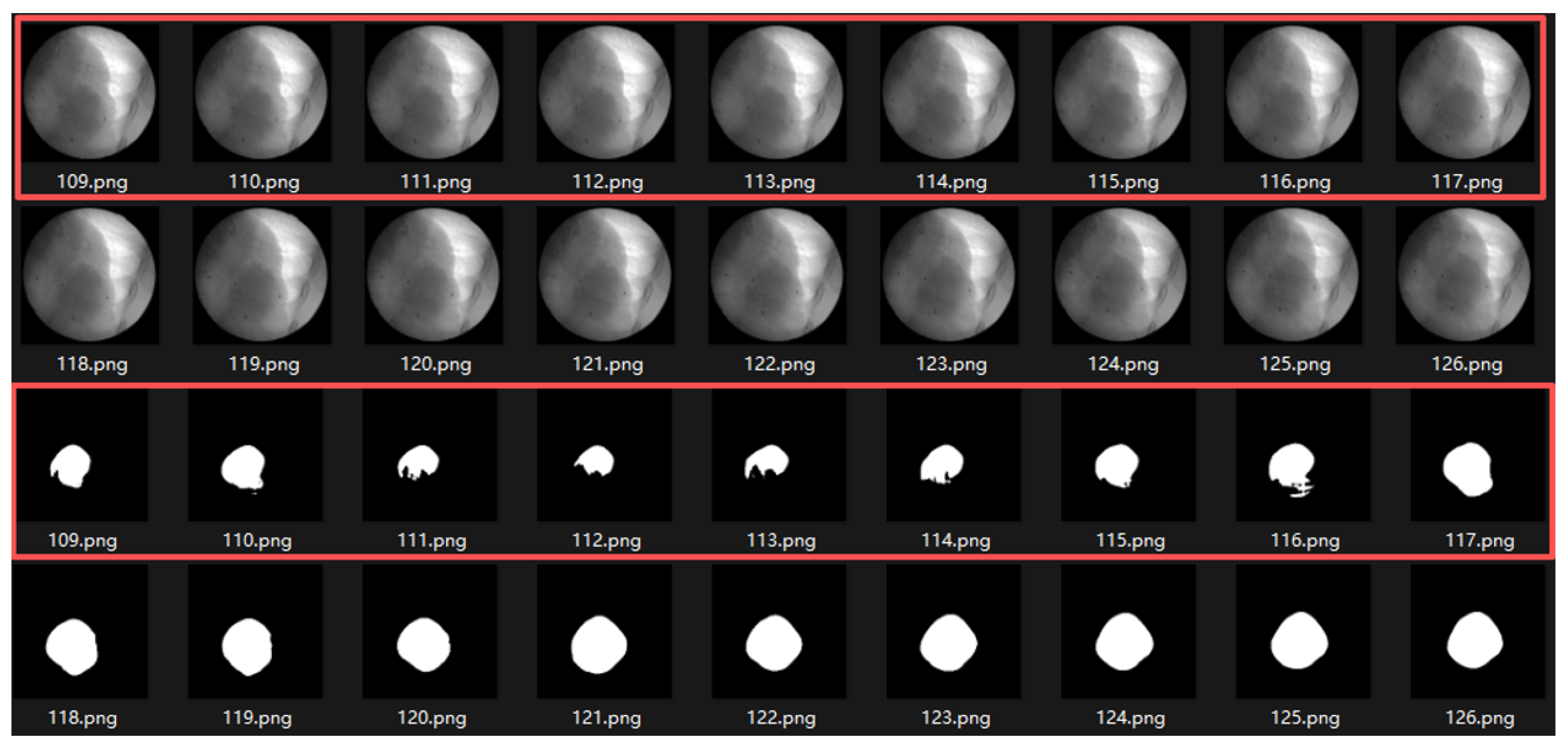
Motion-induced occlusion. During frames 109–117 (end of inhalation, inside the red boxes), the tumor enters the liver region and becomes partially occluded, resulting in blurred boundaries and decreased segmentation accuracy.
Figure 21.
Motion-induced occlusion. During frames 109–117 (end of inhalation, inside the red boxes), the tumor enters the liver region and becomes partially occluded, resulting in blurred boundaries and decreased segmentation accuracy.
Figure 22.
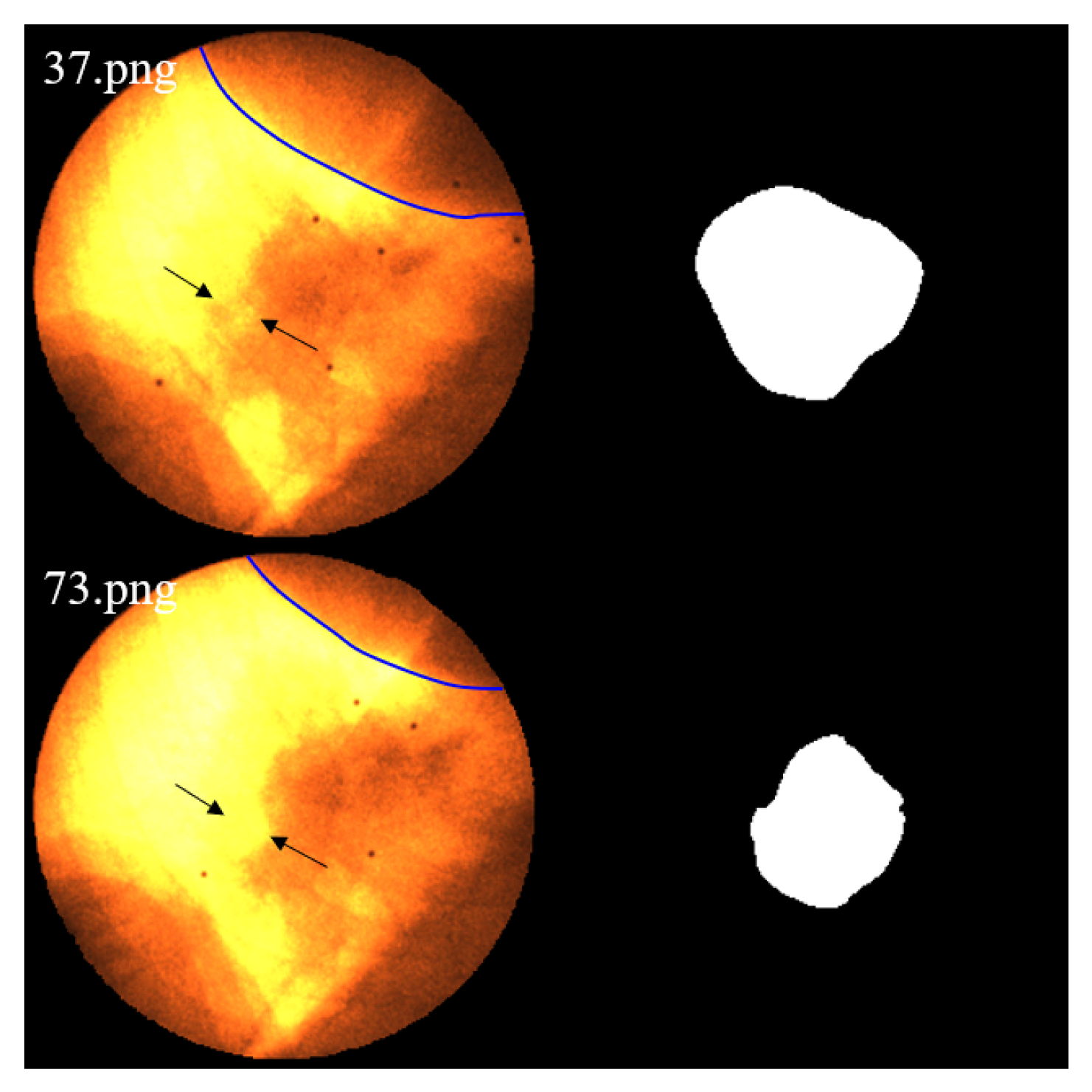
As lung expansion increases the proportion of lung regions in the image, the relatively low X-ray absorption of lung tissue leads to blurring of tumor-background boundaries and consequently degrades the segmentation performance. Image 37.png corresponds to the end-expiration phase, while 73.png shows the end-inspiration phase. The blue contour delineates the liver boundary. A marked reduction in liver area and a concurrent expansion of the pulmonary region are observed in 73.png. Additionally, the black arrows indicate regions where edges become blurred due to contrast variation.
Figure 22.
As lung expansion increases the proportion of lung regions in the image, the relatively low X-ray absorption of lung tissue leads to blurring of tumor-background boundaries and consequently degrades the segmentation performance. Image 37.png corresponds to the end-expiration phase, while 73.png shows the end-inspiration phase. The blue contour delineates the liver boundary. A marked reduction in liver area and a concurrent expansion of the pulmonary region are observed in 73.png. Additionally, the black arrows indicate regions where edges become blurred due to contrast variation.
Table 1.
Computer Configuration.
Table 1.
Computer Configuration.
| Component | Specification |
|---|
| CPU | Intel Core i9-9940X |
| Memory | 64 GB |
| GPU | GeForce RTX 3090 |
| GPU Memory | 24 GB GDDR6X |
Table 2.
Stochastic Strategy for Synthesizing Degraded Images.
Table 2.
Stochastic Strategy for Synthesizing Degraded Images.
| Parameter | Random Strategy/Distribution | Physical Significance |
|---|
| | Beam center offset |
| | Central flatness radius |
| | Base noise levels |
Table 3.
Training Configurations.
Table 3.
Training Configurations.
| Parameter | Original (Restormer) | Ours |
|---|
| Number of GPUs | 8 | 1 |
| Number of Workers per GPU | 8 | 4 |
| Batch Size per GPU | 8 | 4 |
| Minibatch Sizes (Progressive) | [8, 5, 4, 2, 1, 1] | [4, 3, 2, 2, 1, 1] |
| Max Patch Size (GT Size) | 384 | 320 |
| Patch Sizes (Progressive) | [160, 192, 256, 320, 384] | [128, 160, 192, 256, 320] |
| Max Minibatch (Validation) | 8 | 4 |
Table 4.
Grading Standard Based on 2D Trajectory Error.
Table 4.
Grading Standard Based on 2D Trajectory Error.
| Grade | E (Pixels) | Description |
|---|
| Excellent | <3 | Excellent performance in almost all frames. |
| High | 3 ≤ E < 8 | Few frames are not accurate enough. |
| Moderate | 8 ≤ E < 13 | Some frames had poor results. |
| Low | ≥13 | Poor accuracy in many frames. |
Table 5.
2D trajectory evaluation comparison. All compared methods were evaluated on the exact same dataset.
Table 5.
2D trajectory evaluation comparison. All compared methods were evaluated on the exact same dataset.
| Grade | E | Method 1 1 | Method 2 2 | Method 3 3 | Method 4 4 | Method 5 5 |
|---|
| Excellent | (0, 3] | 22.73% | 50% | 48% | 50% | 51% |
| High | (3, 8] | 39.9% | 45% | 47% | 46% | 45% |
| Moderate | (8, 13] | 22.73% | 4% | 4% | 3% | 3% |
| Low | | 14.64% | 1% | 1% | 1% | 1% |
Table 6.
3D trajectory evaluation comparison.
Table 6.
3D trajectory evaluation comparison.
| | Method 1 1 | Method 2 2 | Method 3 3 |
|---|
| LR | 0.98 ± 0.67 mm | 1.00 ± 0.69 mm | 0.98 ± 0.70 mm |
| SI | 1.14 ± 0.76 mm | 1.15 ± 0.77 mm | 1.09 ± 0.74 mm |
| AP | 1.34 ± 0.93 mm | 1.35 ± 0.91 mm | 1.34 ± 0.94 mm |
| MEE* | 1.62 mm | 1.64 mm | 1.53 mm |
| IQR* | 1.04 mm | 0.99 mm | 1.04 mm |
Table 7.
Performance trade-off analysis of the style transfer module.
Table 7.
Performance trade-off analysis of the style transfer module.
| Workflow Configuration | MEE (mm) | Processing Time (ms/Frame) |
|---|
| Without Style Transfer | 1.62 | 101.8 |
| With Style Transfer (Proposed) | 1.53 | 179.8 |
| Change | 5.6% performance improvement | Time increased by 76.6% |