Exopolysaccharides Producing Lactic Acid Bacteria in Wine and Other Fermented Beverages: For Better or for Worse?
Abstract
1. Introduction
2. EPSs Produced and Biosynthetic Pathways
2.1. EPSs Produced by Wine and Cider LAB in Brief
2.2. EPS Localization
- WPS or wall polysaccharides, attached to the cell, covalently or not, but without forming a capsule.
- CPS (or capsular polysaccharides), most of the time linked to peptidoglycan, forming either a thick and cohesive (capsule) or a thin and cohesive (film) outer layer.
- Exocellular polysaccharides (or true EPSs), released into the environment surrounding the cell during planktonic growth. This kind of true EPS can also form a slime or a polymeric matrix during growth on solid media or biofilm formation.
2.3. Biosynthetic Pathways
- (i)
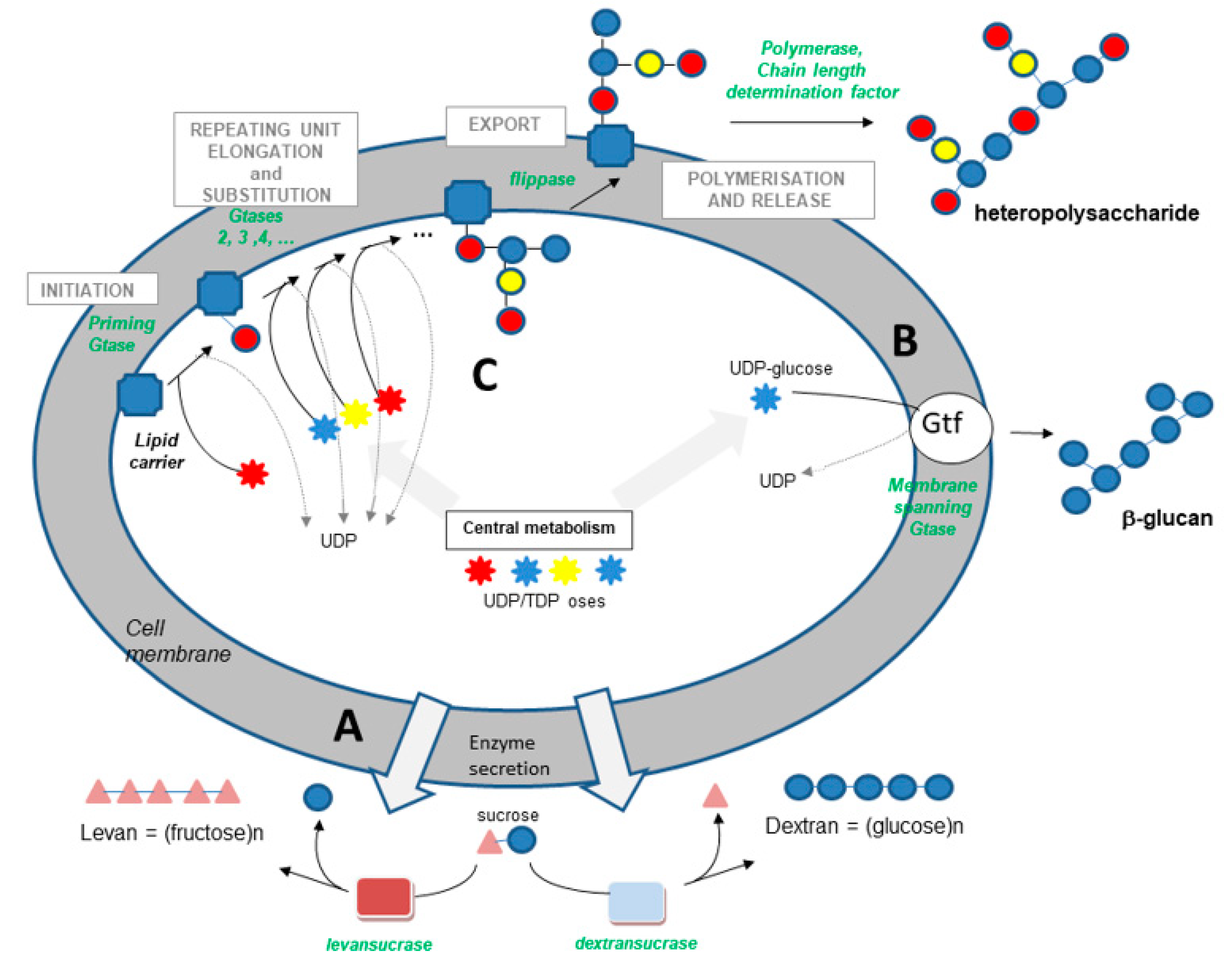
- Transglycosidases, which specifically use sucrose as a substrate (or glycansucrases), are classified into the CAZy GH-13, 68 and 70 families (www.cazy.org (accessed on 14 September 2021)) [56]. They catalyze the synthesis of homopolysaccharides made up of glucose or fructose, according to the following simplified reactions:
- (ii)
- The synthase pathway involves a single membrane-spanning enzyme, which alone carries out the initiation of synthesis, the elongation of the polymer (processive enzyme) and its export through the membrane (Figure 1). The β-glucan causing wine or cider ropiness is produced by an enzyme of this type, called Gtf. The role of Gtf in the synthesis of P. parvulus and O. oeni β-glucan was demonstrated in 2006 and 2008 [17,18]. Gtf is 32% identical to Tts, a synthase found in S. pneumoniae type 37, which produces a β-glucan with structure close to that produced by wine, beer and cider strains. This pneumococcal glucan is immunogenic in humans, as in mice, and was shown to be responsible for pneumococcal strain virulence [59].
- (iii)
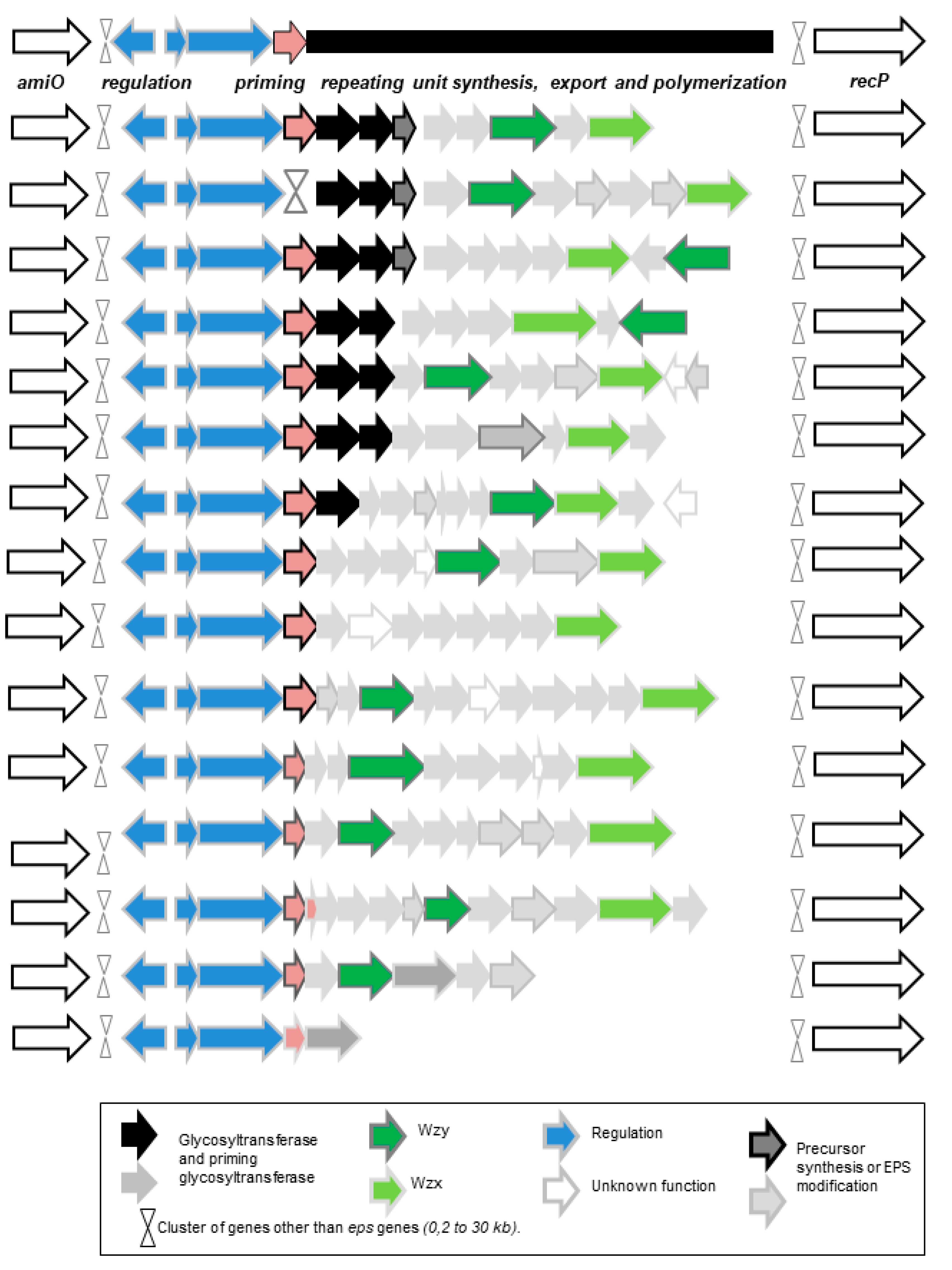
- The third pathway is the most complex (Figure 1). This pathway is sometimes called the Wzy-dependent pathway, based on the name given to the polymerase in E. coli [60]. This pathway has been perfectly characterized in Gram-negative bacteria and partially in Gram-positive bacteria [49,50,51,52,60]. The first step is the synthesis of a repeating oligosaccharidic unit through the transfer of monomers to a lipid transporter on the inner face of the cell membrane (Figure 1). This synthesis, carried out by a series of non-processive glycosyltransferases, is followed by the export of the repeating unit by a flippase (Wzx) and by the assembly of the exported repeating units by a polymerase attached to the external face of the cell membrane (Wzy). Regulating enzymes and factors modulate the chain length and the polymer release. The glycosyltransferase which initiates the synthesis of the repeat unit by transferring the first monomer to the lipid transporter is called the “priming glycosyltransferase”. Several priming glycosyltransferases are found in O. oeni and complement each other [61], ensuring EPS formation even in cases in which mutations inactivate one of the enzymes.
2.4. Genes Associated with EPS Synthesis
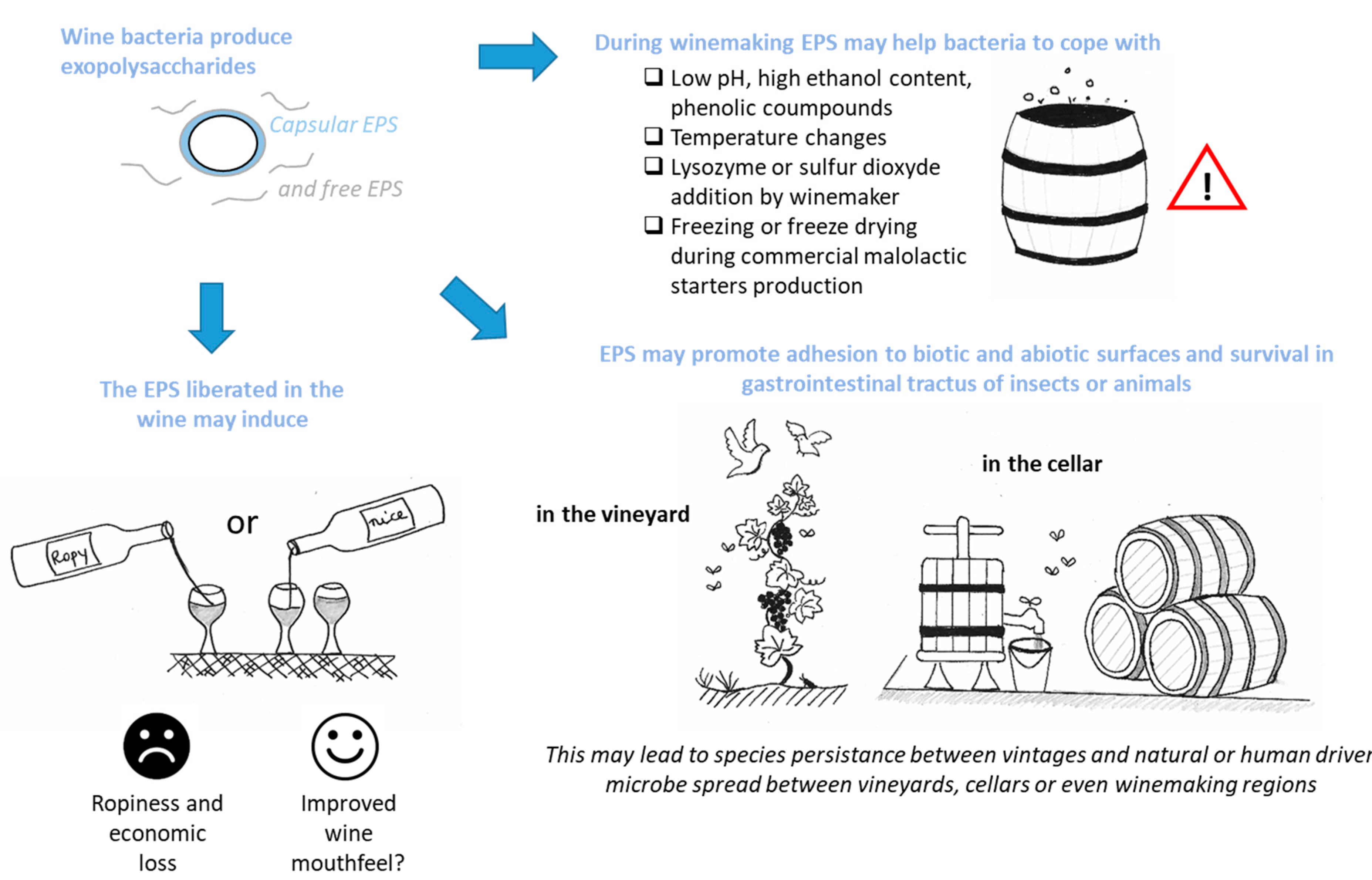
3. What Could Be the Consequences for the Winemaker or the Wine Lover?
3.1. EPSs for Bacteria Survival in Harsh Winemaking Conditions
3.2. EPSs, Biotic Interactions and Species Survival
3.3. EPSs and Bacterial Colonization of the Production Cellar
3.4. Beverage Spoilage and Possible Treatments
3.5. Bacterial EPSs and Wine Sensorial Properties
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- König, H.; Berkelmann-Löhnertz, B. Lactic Acid Bacteria. In Biology of Microorganisms on Grapes, in Must and in Wine; König, H., Unden, G., Fröhlich, J., Eds.; Springer: Cham, Switzerland, 2017; pp. 3–41. ISBN 978-3-319-60021-5. [Google Scholar]
- König, H.; Berkelmann-Löhnertz, B. Maintenance of Wine-Associated Microorganisms. In Biology of Microorganisms on Grapes, in Must and in Wine; König, H., Unden, G., Fröhlich, J., Eds.; Springer: Cham, Switzerland, 2017; pp. 549–571. ISBN 978-3-319-60020-8. [Google Scholar]
- Davis, C.R.; Wibowo, D.; Eschenbruch, R.; Lee, T.H.; Fleet, G.H. Practical Implications of Malolactic Fermentation: A Review. Am. J. Enol. Vitic. 1985, 36, 290–301. [Google Scholar]
- Lonvaud-Funel, A. Lactic Acid Bacteria in the Quality Improvement and Depreciation of Wine. Antonie Van Leeuwenhoek 1999, 76, 317–331. [Google Scholar] [CrossRef]
- Versari, A.; Parpinello, G.P.; Cattaneo, M. Leuconostoc oenos and Malolactic Fermentation in Wine: A Review. J. Ind. Microbiol. Biotechnol. 1999, 23, 447–455. [Google Scholar] [CrossRef]
- Sumby, K.M.; Grbin, P.R.; Jiranek, V. Microbial Modulation of Aromatic Esters in Wine: Current Knowledge and Future Prospects. Food Chem. 2010, 121, 1–16. [Google Scholar] [CrossRef]
- Sánchez, A.; Rodríguez, R.; Coton, M.; Coton, E.; Herrero, M.; García, L.A.; Díaz, M. Population Dynamics of Lactic Acid Bacteria during Spontaneous Malolactic Fermentation in Industrial Cider. Food Res. Int. 2010, 43, 2101–2107. [Google Scholar] [CrossRef]
- Sánchez, A.; Coton, M.; Coton, E.; Herrero, M.; García, L.A.; Díaz, M. Prevalent Lactic Acid Bacteria in Cider Cellars and Efficiency of Oenococcus oeni Strains. Food Microbiol. 2012, 32, 32–37. [Google Scholar] [CrossRef]
- Bartowsky, E.J. Oenococcus oeni and Malolactic Fermentation—Moving into the Molecular Arena. Aust. J. Grape Wine Res. 2005, 11, 174–187. [Google Scholar] [CrossRef]
- Lorentzen, M.P.G.; Lucas, P.M. Distribution of Oenococcus oeni Populations in Natural Habitats. Appl. Microbiol. Biotechnol. 2019, 103, 2937–2945. [Google Scholar] [CrossRef]
- Badotti, F.; Moreira, A.P.B.; Tonon, L.A.C.; de Lucena, B.T.L.; Fátima de Cássia, O.G.; Kruger, R.; Thompson, C.C.; de Morais, M.A.; Rosa, C.A.; Thompson, F.L. Oenococcus alcoholitolerans Sp. Nov., a Lactic Acid Bacteria Isolated from Cachaça and Ethanol Fermentation Processes. Antonie Van Leeuwenhoek 2014, 106, 1259–1267. [Google Scholar] [CrossRef]
- Endo, A.; Okada, S. Oenococcus kitaharae Sp. Nov., a Non-Acidophilic and Non-Malolactic-Fermenting Oenococcus Isolated from a Composting Distilled Shochu Residue. Int. J. Syst. Evol. Microbiol. 2006, 56, 2345–2348. [Google Scholar] [CrossRef]
- Cousin, F.J.; Le Guellec, R.; Schlusselhuber, M.; Dalmasso, M.; Laplace, J.-M.; Cretenet, M. Microorganisms in Fermented Apple Beverages: Current Knowledge and Future Directions. Microorganisms 2017, 5, 39. [Google Scholar] [CrossRef]
- Bartowsky, E.J. Oenococcus oeni and the Genomic Era. FEMS Microbiol. Rev. 2017, 41, S84–S94. [Google Scholar] [CrossRef]
- Dimopoulou, M.; Bardeau, T.; Ramonet, P.-Y.; Miot-Certier, C.; Claisse, O.; Doco, T.; Petrel, M.; Lucas, P.; Dols-Lafargue, M. Exopolysaccharides Produced by Oenococcus oeni: From Genomic and Phenotypic Analysis to Technological Valorization. Food Microbiol. 2016, 53, 10–17. [Google Scholar] [CrossRef]
- Dols Lafargue, M. Polysaccharide Production by Wine Lactic Acid Bacteria: Negative Trait or Potential Advantage? A Review. Appl. Microbiol. 2018, 4, 1–8. [Google Scholar] [CrossRef]
- Werning, M.L.; Ibarburu, I.; As, M.T.D.; Irastorza, A.; Navas, J.S.; Pez, P.L. Pediococcus Parvulus Gtf Gene Encoding the GTF Glycosyltransferase and Its Application for Specific PCR Detection of β-D-glucan–producing Bacteria in Foods and Beverages. J. Food Prot. 2006, 69, 161–169. [Google Scholar] [CrossRef]
- Dols-Lafargue, M.; Lee, H.Y.; Le Marrec, C.; Heyraud, A.; Chambat, G.; Lonvaud-Funel, A. Characterization of Gtf, a Glucosyltransferase Gene in the Genomes of Pediococcus parvulus and Oenococcus oeni, Two Bacterial Species Commonly Found in Wine. Appl. Environ. Microbiol. 2008, 74, 4079–4090. [Google Scholar] [CrossRef] [PubMed]
- Ibarburu, I.; Aznar, R.; Elizaquível, P.; García-Quintáns, N.; López, P.; Munduate, A.; Irastorza, A.; Dueñas, M.T. A Real-Time PCR Assay for Detection and Quantification of 2-Branched (1,3)-Beta-D-Glucan Producing Lactic Acid Bacteria in Cider. Int. J. Food Microbiol. 2010, 143, 26–31. [Google Scholar] [CrossRef]
- Duenas, M.; Irastorza, A.; Fernandez, K.; Bilbao, A. Heterofermentative Lactobacilli Causing Ropiness in Basque Country Ciders. J. Food Prot. 1995, 58, 76–80. [Google Scholar] [CrossRef]
- Llaubères, R.M.; Richard, B.; Lonvaud, A.; Dubourdieu, D.; Fournet, B. Structure of an Exocellular Beta-D-Glucan from Pediococcus sp., a Wine Lactic Bacteria. Carbohydr. Res. 1990, 203, 103–107. [Google Scholar] [CrossRef]
- Gindreau, E.; Walling, E.; Lonvaud-Funel, A. Direct Polymerase Chain Reaction Detection of Ropy Pediococcus damnosus Strains in Wine. J. Appl. Microbiol. 2001, 90, 535–542. [Google Scholar] [CrossRef]
- Walling, E.; Gindreau, E.; Lonvaud-Funel, A. A Putative Glucan Synthase Gene Dps Detected in Exopolysaccharide-Producing Pediococcus damnosus and Oenococcus oeni Strains Isolated from Wine and Cider. Int. J. Food Microbiol. 2005, 98, 53–62. [Google Scholar] [CrossRef] [PubMed]
- Llamas-Arriba, M.G.; Pérez-Ramos, A.; Puertas, A.I.; López, P.; Dueñas, M.T.; Prieto, A. Characterization of Pediococcus ethanolidurans CUPV141: A β-D-Glucan- and Heteropolysaccharide-Producing Bacterium. Front. Microbiol. 2018, 9, 2041. [Google Scholar] [CrossRef]
- Pittet, V.; Morrow, K.; Ziola, B. Ethanol Tolerance of Lactic Acid Bacteria, Including Relevance of the Exopolysaccharide Gene Gtf. J. Am. Soc. Brew. Chem. 2011, 69, 57–61. [Google Scholar] [CrossRef]
- Fraunhofer, M.E.; Geissler, A.J.; Wefers, D.; Bunzel, M.; Jakob, F.; Vogel, R.F. Characterization of β-Glucan Formation by Lactobacillus brevis TMW 1.2112 Isolated from Slimy Spoiled Beer. Int. J. Biol. Macromol. 2018, 107, 874–881. [Google Scholar] [CrossRef] [PubMed]
- Jakob, F.; Stahl, L.; Vogel, R.F. β-Glucan Formation Is a Selective Advantage for Beer-Spoiling Levilactobacillus brevis TMW 1.2112 during Planktonic Growth. Microbiol. Res. 2021, 243, 126648. [Google Scholar] [CrossRef]
- Garai-Ibabe, G.; Dueñas, M.T.; Irastorza, A.; Sierra-Filardi, E.; Werning, M.L.; López, P.; Corbí, A.L.; Fernández de Palencia, P. Naturally Occurring 2-Substituted (1,3)-Beta-D-Glucan Producing Lactobacillus suebicus and Pediococcus parvulus Strains with Potential Utility in the Production of Functional Foods. Bioresour. Technol. 2010, 101, 9254–9263. [Google Scholar] [CrossRef]
- Notararigo, S.; Nácher-Vázquez, M.; Ibarburu, I.; Werning, M.L.; de Palencia, P.F.; Dueñas, M.T.; Aznar, R.; López, P.; Prieto, A. Comparative Analysis of Production and Purification of Homo- and Hetero-Polysaccharides Produced by Lactic Acid Bacteria. Carbohydr. Polym. 2013, 93, 57–64. [Google Scholar] [CrossRef]
- Dueñas-Chasco, M.T.; Rodríguez-Carvajal, M.A.; Tejero-Mateo, P.; Espartero, J.L.; Irastorza-Iribas, A.; Gil-Serrano, A.M. Structural Analysis of the Exopolysaccharides Produced by Lactobacillus spp. G-77. Carbohydr. Res. 1998, 307, 125–133. [Google Scholar] [CrossRef]
- Koirala, P.; Maina, N.H.; Nihtilä, H.; Katina, K.; Coda, R. Brewers’ Spent Grain as Substrate for Dextran Biosynthesis by Leuconostoc pseudomesenteroides DSM20193 and Weissella confusa A16. Microb. Cell. Fact. 2021, 20, 23. [Google Scholar] [CrossRef]
- Zhou, Q.; Feng, F.; Yang, Y.; Zhao, F.; Du, R.; Zhou, Z.; Han, Y. Characterization of a Dextran Produced by Leuconostoc pseudomesenteroides XG5 from Homemade Wine. Int. J. Biol. Macromol. 2018, 107, 2234–2241. [Google Scholar] [CrossRef]
- Montersino, S.; Prieto, A.; Muñoz, R.; Rivas, B.D.L. Evaluation of Exopolysaccharide Production by Leuconostoc mesenteroides Strains Isolated from Wine. J. Food Sci. 2008, 73, M196–M199. [Google Scholar] [CrossRef] [PubMed]
- Dimopoulou, M.; Raffenne, J.; Claisse, O.; Miot-Sertier, C.; Iturmendi, N.; Moine, V.; Coulon, J.; Dols-Lafargue, M. Oenococcus oeni Exopolysaccharide Biosynthesis, a Tool to Improve Malolactic Starter Performance. Front. Microbiol. 2018, 9, 1276. [Google Scholar] [CrossRef]
- Dimopoulou, M.; Vuillemin, M.; Campbell-Sills, H.; Lucas, P.M.; Ballestra, P.; Miot-Sertier, C.; Favier, M.; Coulon, J.; Moine, V.; Doco, T.; et al. Exopolysaccharide (EPS) Synthesis by Oenococcus oeni: From Genes to Phenotypes. PLoS ONE 2014, 9, e98898. [Google Scholar] [CrossRef] [PubMed]
- Bastard, A.; Coelho, C.; Briandet, R.; Canette, A.; Gougeon, R.; Alexandre, H.; Guzzo, J.; Weidmann, S. Effect of Biofilm Formation by Oenococcus oeni on Malolactic Fermentation and the Release of Aromatic Compounds in Wine. Front. Microbiol. 2016, 7, 613. [Google Scholar] [CrossRef]
- Ibarburu, I.; Puertas, A.I.; Berregi, I.; Rodríguez-Carvajal, M.A.; Prieto, A.; Dueñas, M.T. Production and Partial Characterization of Exopolysaccharides Produced by Two Lactobacillus suebicus Strains Isolated from Cider. Int. J. Food Microbiol. 2015, 214, 54–62. [Google Scholar] [CrossRef] [PubMed]
- Dimopoulou, M.; Lonvaud-Funel, A.; Dols-Lafargue, M. Polysaccharide Production by Grapes Must and Wine Microorganisms. In Biology of Microorganisms on Grapes, in Must and in Wine; König, H., Unden, G., Fröhlich, J., Eds.; Springer International Publishing: Cham, Switzerland, 2017; pp. 293–314. ISBN 978-3-319-60021-5. [Google Scholar]
- Dueñas-Chasco, M.T.; Rodríguez-Carvajal, M.A.; Mateo, P.T.; Franco-Rodríguez, G.; Espartero, J.L.; Irastorza-Iribas, A.; Gil-Serrano, A.M. Structural Analysis of the Exopolysaccharide Produced by Pediococcus damnosus 2.6. Carbohydr. Res. 1997, 303, 453–458. [Google Scholar] [CrossRef]
- Puertas, A.I.; Ibarburu, I.; Elizaquivel, P.; Zuriarrain, A.; Berregi, I.; López, P.; Prieto, A.; Aznar, R.; Dueñas, M.T. Disclosing Diversity of Exopolysaccharide-Producing Lactobacilli from Spanish Natural Ciders. LWT 2018, 90, 469–474. [Google Scholar] [CrossRef]
- Dimopoulou, M.; Hazo, L.; Dols-Lafargue, M. Exploration of Phenomena Contributing to the Diversity of Oenococcus oeni Exopolysaccharides. Int. J. Food Microbiol. 2012, 153, 114–122. [Google Scholar] [CrossRef]
- Stivala, M.G.; Villecco, M.B.; Enriz, D.; Fernández, P.A. Effect of Phenolic Compounds on Viability of Wine Spoilage Lactic Acid Bacteria. A Structure-Activity Relationship Study. Am. J. Enol. Vitic. 2017, 68, 228–233. [Google Scholar] [CrossRef]
- Dols-Lafargue, M.; Gindreau, E.; Le Marrec, C.; Chambat, G.; Heyraud, A.; Lonvaud-Funel, A. Changes in Red Wine Soluble Polysaccharide Composition Induced by Malolactic Fermentation. J. Agric. Food Chem. 2007, 55, 9592–9599. [Google Scholar] [CrossRef]
- Ciezack, G.; Hazo, L.; Chambat, G.; Heyraud, A.; Lonvaud-Funel, A.; Dols-Lafargue, M. Evidence for Exopolysaccharide Production by Oenococcus oeni Strains Isolated from Non-Ropy Wines. J. Appl. Microbiol. 2010, 108, 499–509. [Google Scholar] [CrossRef]
- Ryan, P.M.; Ross, R.P.; Fitzgerald, G.F.; Caplice, N.M.; Stanton, C. Sugar-Coated: Exopolysaccharide Producing Lactic Acid Bacteria for Food and Human Health Applications. Food Funct. 2015, 6, 679–693. [Google Scholar] [CrossRef]
- Weidenmaier, C.; Peschel, A. Teichoic Acids and Related Cell-Wall Glycopolymers in Gram-Positive Physiology and Host Interactions. Nat. Rev. Microbiol. 2008, 6, 276–287. [Google Scholar] [CrossRef]
- Chapot-Chartier, M.-P. Interactions of the Cell-Wall Glycopolymers of Lactic Acid Bacteria with Their Bacteriophages. Front. Microbiol. 2014, 5, 236. [Google Scholar] [CrossRef]
- Fernández de Palencia, P.; Werning, M.L.; Sierra-Filardi, E.; Dueñas, M.T.; Irastorza, A.; Corbí, A.L.; López, P. Probiotic Properties of the 2-Substituted (1,3)-β-d-Glucan-Producing Bacterium Pediococcus parvulus 2.6. Appl. Environ. Microbiol. 2009, 75, 4887–4891. [Google Scholar] [CrossRef]
- Kolkman, M.A.; Morrison, D.A.; Van Der Zeijst, B.A.; Nuijten, P.J. The Capsule Polysaccharide Synthesis Locus of Streptococcus pneumoniae Serotype 14: Identification of the Glycosyl Transferase Gene Cps14E. J. Bacteriol. Res. 1996, 178, 3736–3741. [Google Scholar] [CrossRef][Green Version]
- Kolkman, M.A.; van der Zeijst, B.A.; Nuijten, P.J. Functional Analysis of Glycosyltransferases Encoded by the Capsular Polysaccharide Biosynthesis Locus of Streptococcus pneumoniae Serotype 14. J. Biol. Chem. 1997, 272, 19502–19508. [Google Scholar] [CrossRef] [PubMed]
- Stingele, F.; Lemoine, J.; Neeser, J.-R. Lactobacillus Helveticus Lh59 Secretes an Exopolysaccharide That Is Identical to the One Produced by Lactobacillus helveticus TN-4, a Presumed Spontaneous Mutant of Lactobacillus helveticus TY1–2. Carbohydr. Res. 1997, 302, 197–202. [Google Scholar] [CrossRef]
- Stingele, F.; Vincent, S.J.; Faber, E.J.; Newell, J.W.; Kamerling, J.P.; Neeser, J.R. Introduction of the Exopolysaccharide Gene Cluster from Streptococcus thermophilus Sfi6 into Lactococcus lactis MG1363: Production and Characterization of an Altered Polysaccharide. Mol. Microbiol. 1999, 32, 1287–1295. [Google Scholar] [CrossRef] [PubMed]
- De Vuyst, L.; De Vin, F.; Vaningelgem, F.; Degeest, B. Recent Developments in the Biosynthesis and Applications of Heteropolysaccharides from Lactic Acid Bacteria. Int. Dairy J. 2001, 11, 687–707. [Google Scholar] [CrossRef]
- Lebeer, S.; Verhoeven, T.L.A.; Francius, G.; Schoofs, G.; Lambrichts, I.; Dufrêne, Y.; Vanderleyden, J.; De Keersmaecker, S.C.J. Identification of a Gene Cluster for the Biosynthesis of a Long, Galactose-Rich Exopolysaccharide in Lactobacillus rhamnosus GG and Functional Analysis of the Priming Glycosyltransferase. Appl. Environ. Microbiol. 2009, 75, 3554–3563. [Google Scholar] [CrossRef]
- Monsan, P.; Bozonnet, S.; Albenne, C.; Joucla, G.; Willemot, R.-M.; Remaud-Siméon, M. Homopolysaccharides from Lactic Acid Bacteria. Int. Dairy J. 2001, 11, 675–685. [Google Scholar] [CrossRef]
- Lombard, V.; Golaconda Ramulu, H.; Drula, E.; Coutinho, P.M.; Henrissat, B. The Carbohydrate-Active Enzymes Database (CAZy) in 2013. Nucleic Acids Res. 2014, 42, D490–D495. [Google Scholar] [CrossRef] [PubMed]
- Moulis, C.; Joucla, G.; Harrison, D.; Fabre, E.; Potocki-Veronese, G.; Monsan, P.; Remaud-Simeon, M. Understanding the Polymerization Mechanism of Glycoside-Hydrolase Family 70 Glucansucrases. J. Biol. Chem. 2006, 281, 31254–31267. [Google Scholar] [CrossRef] [PubMed]
- Vuillemin, M.; Grimaud, F.; Claverie, M.; Rolland-Sabaté, A.; Garnier, C.; Lucas, P.; Monsan, P.; Dols-Lafargue, M.; Remaud-Siméon, M.; Moulis, C. A Dextran with Unique Rheological Properties Produced by the Dextransucrase from Oenococcus kitaharae DSM 17330. Carbohydr. Polym. 2018, 179, 10–18. [Google Scholar] [CrossRef] [PubMed]
- Llull, D.; Muñoz, R.; López, R.; García, E. A Single Gene (Tts) Located Outside the Cap Locus Directs the Formation of Streptococcus pneumoniae Type 37 Capsular Polysaccharide. Type 37 Pneumococci Are Natural, Genetically Binary Strains. J. Exp. Med. 1999, 190, 241–251. [Google Scholar] [CrossRef]
- Whitfield, C. Biosynthesis and Assembly of Capsular Polysaccharides in Escherichia coli. Annu. Rev. Biochem. 2006, 75, 39–68. [Google Scholar] [CrossRef]
- Dimopoulou, M.; Claisse, O.; Dutilh, L.; Miot-Sertier, C.; Ballestra, P.; Lucas, P.M.; Dols, M. Molecular Cloning, Expression and Characterization of Oenococcus oeni Priming Glycosyltransferases. Mol. Biotechnol. 2017, 59, 323–333. [Google Scholar] [CrossRef]
- Gänzle, M.G.; Zhang, C.; Monang, B.-S.; Lee, V.; Schwab, C. Novel Metabolites from Cereal-Associated Lactobacilli—Novel Functionalities for Cereal Products? Food Microbiol. 2009, 26, 712–719. [Google Scholar] [CrossRef]
- Lonvaud-Funel, A.; Joyeux, A. Antagonism between Lactic Acid Bacteria of Wines: Inhibition of Leuconostoc oenos by Lactobacillus plantarum and Pediococcus pentosaceus. Food Microbiol. 1993, 10, 411–419. [Google Scholar] [CrossRef]
- Pluvinet, A.; Charron-Bourgoin, F.; Morel, C.; Decaris, B. Polymorphism of Eps Loci in Streptococcus thermophilus: Sequence Replacement by Putative Horizontal Transfer in S. thermophilus IP6757. Int. Dairy J. 2004, 14, 627–634. [Google Scholar] [CrossRef]
- Bentley, S.D.; Aanensen, D.M.; Mavroidi, A.; Saunders, D.; Rabbinowitsch, E.; Collins, M.; Donohoe, K.; Harris, D.; Murphy, L.; Quail, M.A.; et al. Genetic Analysis of the Capsular Biosynthetic Locus from All 90 Pneumococcal Serotypes. PLoS Genet. 2006, 2, e31. [Google Scholar] [CrossRef]
- Milkman, R.; Jaeger, E.; McBride, R.D. Molecular Evolution of the Escherichia coli Chromosome. VI. Two Regions of High Effective Recombination. Genetics 2003, 163, 475–483. [Google Scholar] [CrossRef] [PubMed]
- Cieslewicz, M.J.; Chaffin, D.; Glusman, G.; Kasper, D.; Madan, A.; Rodrigues, S.; Fahey, J.; Wessels, M.R.; Rubens, C.E. Structural and Genetic Diversity of Group B Streptococcus Capsular Polysaccharides. Infect. Immun. 2005, 73, 3096–3103. [Google Scholar] [CrossRef]
- Mavroidi, A.; Aanensen, D.M.; Godoy, D.; Skovsted, I.C.; Kaltoft, M.S.; Reeves, P.R.; Bentley, S.D.; Spratt, B.G. Genetic Relatedness of the Streptococcus pneumoniae Capsular Biosynthetic Loci. J. Bacteriol. Res. 2007, 189, 7841–7855. [Google Scholar] [CrossRef] [PubMed]
- Kubota, T.; Itagaki, M.; Hoshino, C.; Nagata, M.; Morozumi, T.; Kobayashi, T.; Takagi, R.; Yoshie, H. Altered Gene Expression Levels of Matrix Metalloproteinases and Their Inhibitors in Periodontitis-Affected Gingival Tissue. J. Periodontol. 2008, 79, 166–173. [Google Scholar] [CrossRef] [PubMed]
- Ramírez-Castrillón, M.; Mendes, S.D.C.; Inostroza-Ponta, M.; Valente, P. (GTG)5 MSP-PCR Fingerprinting as a Technique for Discrimination of Wine Associated Yeasts? PLoS ONE 2014, 9, e105870. [Google Scholar] [CrossRef]
- Cotter, P.D.; Hill, C. Surviving the Acid Test: Responses of Gram-Positive Bacteria to Low PH. Microbiol. Mol. Biol. Rev. 2003, 67, 429–453. [Google Scholar] [CrossRef]
- Chowdhury, R.; Sahu, G.K.; Das, J. Stress Response in Pathogenic Bacteria. J. Biosci. 1996, 21, 149–160. [Google Scholar] [CrossRef]
- Renouf, V.; Falcou, M.; Miot-Sertier, C.; Perello, M.C.; De Revel, G.; Lonvaud-Funel, A. Interactions between Brettanomyces bruxellensis and Other Yeast Species during the Initial Stages of Winemaking. J. Appl. Microbiol. 2006, 100, 1208–1219. [Google Scholar] [CrossRef]
- El Khoury, M.; Campbell-Sills, H.; Salin, F.; Guichoux, E.; Claisse, O.; Lucas, P.M. Biogeography of Oenococcus oeni Reveals Distinctive but Nonspecific Populations in Wine-Producing Regions. Appl. Environ. Microbiol. 2017, 83, e02322-16. [Google Scholar] [CrossRef] [PubMed]
- Coulon, J.; Houlès, A.; Dimopoulou, M.; Maupeu, J.; Dols-Lafargue, M. Lysozyme Resistance of the Ropy Strain Pediococcus parvulus IOEB 8801 Is Correlated with Beta-Glucan Accumulation around the Cell. Int. J. Food Microbiol. 2012, 159, 25–29. [Google Scholar] [CrossRef] [PubMed]
- Caggianiello, G.; Kleerebezem, M.; Spano, G. Exopolysaccharides Produced by Lactic Acid Bacteria: From Health-Promoting Benefits to Stress Tolerance Mechanisms. Appl. Microbiol. Biotechnol. 2016, 100, 3877–3886. [Google Scholar] [CrossRef] [PubMed]
- Deveau, H.; Van Calsteren, M.-R.; Moineau, S. Effect of Exopolysaccharides on Phage-Host Interactions in Lactococcus lactis. Appl. Environ. Microbiol. 2002, 68, 4364–4369. [Google Scholar] [CrossRef] [PubMed]
- McCabe, O.; Spinelli, S.; Farenc, C.; Labbé, M.; Tremblay, D.; Blangy, S.; Oscarson, S.; Moineau, S.; Cambillau, C. The Targeted Recognition of Lactococcus lactis Phages to Their Polysaccharide Receptors. Mol. Microbiol. 2015, 96, 875–886. [Google Scholar] [CrossRef]
- Russo, P.; López, P.; Capozzi, V.; De Palencia, P.F.; Dueñas, M.T.; Spano, G.; Fiocco, D. Beta-Glucans Improve Growth, Viability and Colonization of Probiotic Microorganisms. Int. J. Mol. Sci. 2012, 13, 6026–6039. [Google Scholar] [CrossRef]
- Stack, H.M.; Kearney, N.; Stanton, C.; Fitzgerald, G.F.; Ross, R.P. Association of Beta-Glucan Endogenous Production with Increased Stress Tolerance of Intestinal Lactobacilli. Appl. Environ. Microbiol. 2010, 76, 500–507. [Google Scholar] [CrossRef]
- Arena, M.P.; Spano, G.; Fiocco, D. Beta-Glucans and Probiotics. Am. J. Immunol. 2017, 13, 34–44. [Google Scholar] [CrossRef]
- Llull, D.; Lopez, R.; Garcia, E. Genetic Bases and Medical Relevance of Capsular Polysaccharide Biosynthesis in Pathogenic Streptococci. Curr. Mol. Med. 2001, 1, 475–491. [Google Scholar] [CrossRef]
- Cooper, C.A.; Mainprize, I.L.; Nickerson, N.N. Genetic, Biochemical, and Structural Analyses of Bacterial Surface Polysaccharides. In Prokaryotic Systems Biology; Krogan, P., Nevan, J., Babu, P., Mohan, B., Eds.; Advances in Experimental Medicine and Biology; Springer International Publishing: Cham, Switzerland, 2015; pp. 295–315. ISBN 978-3-319-23603-2. [Google Scholar]
- Brooker, B.E. Ultrastructural Surface Changes Associated with Dextran Synthesis by Leuconostoc mesenteroides. J. Bacteriol. Res. 1977, 131, 288–292. [Google Scholar] [CrossRef]
- Nguyen, P.-T.; Nguyen, T.-T.; Bui, D.-C.; Hong, P.-T.; Hoang, Q.-K.; Nguyen, H.-T. Exopolysaccharide Production by Lactic Acid Bacteria: The Manipulation of Environmental Stresses for Industrial Applications. AIMS Microbiol. 2020, 6, 451–469. [Google Scholar] [CrossRef]
- Tada, S.; Katakura, Y.; Ninomiya, K.; Shioya, S. Fed-Batch Coculture of Lactobacillus kefiranofaciens with Saccharomyces cerevisiae for Effective Production of Kefiran. J. Biosci. Bioeng. 2007, 103, 557–562. [Google Scholar] [CrossRef]
- Polak-Berecka, M.; Waśko, A.; Paduch, R.; Skrzypek, T.; Sroka-Bartnicka, A. The Effect of Cell Surface Components on Adhesion Ability of Lactobacillus rhamnosus. Antonie Van Leeuwenhoek 2014, 106, 751–762. [Google Scholar] [CrossRef]
- Wang, C.; Mas, A.; Esteve-Zarzoso, B. The Interaction between Saccharomyces cerevisiae and Non-Saccharomyces Yeast during Alcoholic Fermentation Is Species and Strain Specific. Front. Microbiol. 2016, 7, 502. [Google Scholar] [CrossRef] [PubMed]
- da Silva, L.A.; Lopes Neto, J.H.P.; Cardarelli, H.R. Safety and Probiotic Functionality of Isolated Goat Milk Lactic Acid Bacteria. Ann. Microbiol. 2019, 69, 1497–1505. [Google Scholar] [CrossRef]
- Mah, T.-F.C.; O’Toole, G.A. Mechanisms of Biofilm Resistance to Antimicrobial Agents. Trends Microbiol. 2001, 9, 34–39. [Google Scholar] [CrossRef]
- Dimopoulou, M.; Kefalloniti, V.; Tsakanikas, P.; Papanikolaou, S.; Nychas, G.-J.E. Assessing the Biofilm Formation Capacity of the Wine Spoilage Yeast Brettanomyces bruxellensis through FTIR Spectroscopy. Microorganisms 2021, 9, 587. [Google Scholar] [CrossRef] [PubMed]
- Nácher-Vázquez, M.; Iturria, I.; Zarour, K.; Mohedano, M.L.; Aznar, R.; Pardo, M.Á.; López, P. Dextran Production by Lactobacillus Sakei MN1 Coincides with Reduced Autoagglutination, Biofilm Formation and Epithelial Cell Adhesion. Carbohydr. Polym. 2017, 168, 22–31. [Google Scholar] [CrossRef] [PubMed]
- Re, B.D.; Sgorbati, B.; Miglioli, M.; Palenzona, D. Adhesion, Autoaggregation and Hydrophobicity of 13 Strains of Bifidobacterium longum. Lett. Appl. Microbiol. 2000, 31, 438–442. [Google Scholar] [CrossRef]
- Pérez, P.F.; Minnaard, Y.; Disalvo, E.A.; De Antoni, G.L. Surface Properties of Bifidobacterial Strains of Human Origin. Appl. Environ. Microbiol. 1998, 64, 21–26. [Google Scholar] [CrossRef] [PubMed]
- van de Guchte, M.; Serror, P.; Chervaux, C.; Smokvina, T.; Ehrlich, S.D.; Maguin, E. Stress Responses in Lactic Acid Bacteria. Antonie Van Leeuwenhoek 2002, 82, 187–216. [Google Scholar] [CrossRef]
- Lonvaud-Funel, A.; Joyeux, A.; Desens, C. Inhibition of Malolactic Fermentation of Wines by Products of Yeast Metabolism. J. Sci. Food Agric. 1988, 44, 183–191. [Google Scholar] [CrossRef]
- Bockwoldt, J.A.; Stahl, L.; Ehrmann, M.A.; Vogel, R.F.; Jakob, F. Persistence and β-Glucan Formation of Beer-Spoiling Lactic Acid Bacteria in Wheat and Rye Sourdoughs. Food Microbiol. 2020, 91, 103539. [Google Scholar] [CrossRef]
- Delaherche, A.; Claisse, O.; Lonvaud-Funel, A. Detection and Quantification of Brettanomyces bruxellensis and “ropy” Pediococcus damnosus Strains in Wine by Real-Time Polymerase Chain Reaction. J. Appl. Microbiol. 2004, 97, 910–915. [Google Scholar] [CrossRef]
- Martínez Viedma, P.; Abriouel, H.; Sobrino López, A.; Ben Omar, N.; Lucas López, R.; Valdivia, E.; Martín Belloso, O.; Gálvez, A. Effect of Enterocin AS-48 in Combination with High-Intensity Pulsed-Electric Field Treatment against the Spoilage Bacterium Lactobacillus diolivorans in Apple Juice. Food Microbiol. 2009, 26, 491–496. [Google Scholar] [CrossRef]
- Pataro, G.; Barca, G.M.J.; Donsì, G.; Ferrari, G. On the Modeling of Electrochemical Phenomena at the Electrode-Solution Interface in a PEF Treatment Chamber: Methodological Approach to Describe the Phenomenon of Metal Release. J. Food Eng. 2015, 165, 34–44. [Google Scholar] [CrossRef]
- Van Wyk, S.; Silva, F.V.M.; Farid, M.M. Pulsed Electric Field Treatment of Red Wine: Inactivation of Brettanomyces and Potential Hazard Caused by Metal Ion Dissolution. Innov. Food Sci. Emerg. Technol. 2019, 52, 57–65. [Google Scholar] [CrossRef]
- Pérez-Serradilla, J.A.; de Castro, M.D.L. Role of Lees in Wine Production: A Review. Food Chem. 2008, 111, 447–456. [Google Scholar] [CrossRef] [PubMed]
- Vidal, S.; Williams, P.; Doco, T.; Moutounet, M.; Pellerin, P. The Polysaccharides of Red Wine: Total Fractionation and Characterization. Carbohydr. Polym. 2003, 54, 439–447. [Google Scholar] [CrossRef]
- Samant, S.K.; Singhal, R.S.; Kulkarni, P.R.; Rege, D.V. Protein-Polysaccharide Interactions: A New Approach in Food Formulations. Int. J. Food Sci. 1993, 28, 547–562. [Google Scholar] [CrossRef]
- Moine-Ledoux, V.; Dubourdieu, D. An Invertase Fragment Responsible for Improving the Protein Stability of Dry White Wines. J. Sci. Food Agric. 1999, 79, 537–543. [Google Scholar] [CrossRef]
- Taira, S.; Ono, M.; Matsumoto, N. Reduction of Persimmon Astringency by Complex Formation between Pectin and Tannins. Postharvest Biol. Technol. 1997, 12, 265–271. [Google Scholar] [CrossRef]
- Moine-Ledoux, V.; Dubourdieu, D. Role of yeast mannoproteins with regard to tartaric stabilization of wines. Bull. L’oiv 2002, 75, 471–482. [Google Scholar]
- Guise, R.; Filipe-Ribeiro, L.; Nascimento, D.; Bessa, O.; Nunes, F.M.; Cosme, F. Comparison between Different Types of Carboxylmethylcellulose and Other Oenological Additives Used for White Wine Tartaric Stabilization. Food Chem. 2014, 156, 250–257. [Google Scholar] [CrossRef]
- Martínez-Lapuente, L.; Guadalupe, Z.; Ayestarán, B. Properties of Wine Polysaccharides; IntechOpen: London, UK, 2019; ISBN 978-1-78984-072-8. [Google Scholar]
- Gonçalves, F.J.; Fernandes, P.A.R.; Wessel, D.F.; Cardoso, S.M.; Rocha, S.M.; Coimbra, M.A. Interaction of Wine Mannoproteins and Arabinogalactans with Anthocyanins. Food Chem. 2018, 243, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Soares, S.; Mateus, N.; de Freitas, V. Carbohydrates Inhibit Salivary Proteins Precipitation by Condensed Tannins. J. Agric. Food Chem. 2012, 60, 3966–3972. [Google Scholar] [CrossRef]
- Gonçalves, F.J.; Rocha, S.M.; Coimbra, M.A. Study of the Retention Capacity of Anthocyanins by Wine Polymeric Material. Food Chem. 2012, 134, 957–963. [Google Scholar] [CrossRef]
- Cameleyre, M.; Lytra, G.; Barbe, J.-C. Static Headspace Analysis Using Low-Pressure Gas Chromatography and Mass Spectrometry, Application to Determining Multiple Partition Coefficients: A Practical Tool for Understanding Red Wine Fruity Volatile Perception and the Sensory Impact of Higher Alcohols. Anal. Chem. 2018, 90, 10812–10818. [Google Scholar] [CrossRef] [PubMed]
- Muñoz-González, C.; Martín-Álvarez, P.J.; Moreno-Arribas, M.V.; Pozo-Bayón, M.Á. Impact of the Nonvolatile Wine Matrix Composition on the in Vivo Aroma Release from Wines. J. Agric. Food Chem. 2014, 62, 66–73. [Google Scholar] [CrossRef]
- Dufour, C.; Bayonove, C.L. Influence of Wine Structurally Different Polysaccharides on the Volatility of Aroma Substances in a Model System. J. Agric. Food Chem. 1999, 47, 671–677. [Google Scholar] [CrossRef] [PubMed]
| EPS Type | EPS Structure | Species | Niche | Implicated Genes | Consequences/Role of EPS | Reference |
|---|---|---|---|---|---|---|
| homopolysaccharides | β-glucan | Oenococcus oeni | wine, cider | gtf | ropy character, stress resistance | [17,18,19] |
| β-glucan | Pediococcus damnosus | cider | gtf | ropy character | [17,20,21,22,23] | |
| β-glucan | Pediococcus parvulus | cider, wine | gtf | ropy character, stress resistance | [18,23] | |
| β-glucan | Pediococcus ethanolidurans | cider | - | ropy character | [24] | |
| β-glucan | Pediococcus claussenii | beer | gtf | ropy character | [25,26] | |
| β-glucan | Lactobacillus brevis | beer | gtf2 | ropy character, ethanol tolerance, biofilm formation | [26,27] | |
| β-glucan | Lactobacillus diolivorans | cider | gtf | - | [17] | |
| β-glucan | Lactobacillus suebicus | cider | gtf | ropy character | [28,29] | |
| β-glucan | Lactobacillus spp. | cider | - | ropy character | [30] | |
| a-glucan | Leuconostoc pseudomesenteroides, Weissella confusa | beer | dsr | increased viscosity | [31] | |
| dextran | Leuconostoc pseudomesenteroides | homemade wine | - | - | [32] | |
| glucan and fructan | Leuconostoc mesenteroides | grape must and wine | Glucosyltransferase gene | more or less mucoid strains | [33] | |
| dextran and levan | Oenococcus oeni | wine | dsrO and levO | lyoprotective ability to freeze-drying process | [15,34,35] | |
| heteropolysaccharides | glucose, galactose, rhamnose | Oenococcus oeni | wine | eps cluster | aromatic complexity, biofilm formation, capsule, lyoprotective ability to freeze-drying | [15,34,35,36] |
| glucose, galactose, galactofuranose | Lactobacillus suebicus | cider | gtf | ropy character | [28,29] | |
| glucose, galactose, N-acetyl-glucosamine, phosphate | Lactobacillus suebicus | cider | eps cluster | ropy character | [37] | |
| glucose, galactose, glucosamine | Pediococcus ethanolidurans | cider | - | ropy character | [24] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dimopoulou, M.; Dols-Lafargue, M. Exopolysaccharides Producing Lactic Acid Bacteria in Wine and Other Fermented Beverages: For Better or for Worse? Foods 2021, 10, 2204. https://doi.org/10.3390/foods10092204
Dimopoulou M, Dols-Lafargue M. Exopolysaccharides Producing Lactic Acid Bacteria in Wine and Other Fermented Beverages: For Better or for Worse? Foods. 2021; 10(9):2204. https://doi.org/10.3390/foods10092204
Chicago/Turabian StyleDimopoulou, Maria, and Marguerite Dols-Lafargue. 2021. "Exopolysaccharides Producing Lactic Acid Bacteria in Wine and Other Fermented Beverages: For Better or for Worse?" Foods 10, no. 9: 2204. https://doi.org/10.3390/foods10092204
APA StyleDimopoulou, M., & Dols-Lafargue, M. (2021). Exopolysaccharides Producing Lactic Acid Bacteria in Wine and Other Fermented Beverages: For Better or for Worse? Foods, 10(9), 2204. https://doi.org/10.3390/foods10092204






