Green Synthesized Copper Oxide Nanoparticles Ameliorate Defence and Antioxidant Enzymes in Lens culinaris
Abstract
1. Introduction
2. Materials and Methods
2.1. Collection and Identification of Plant Material
2.2. Preparation of Plant Extract
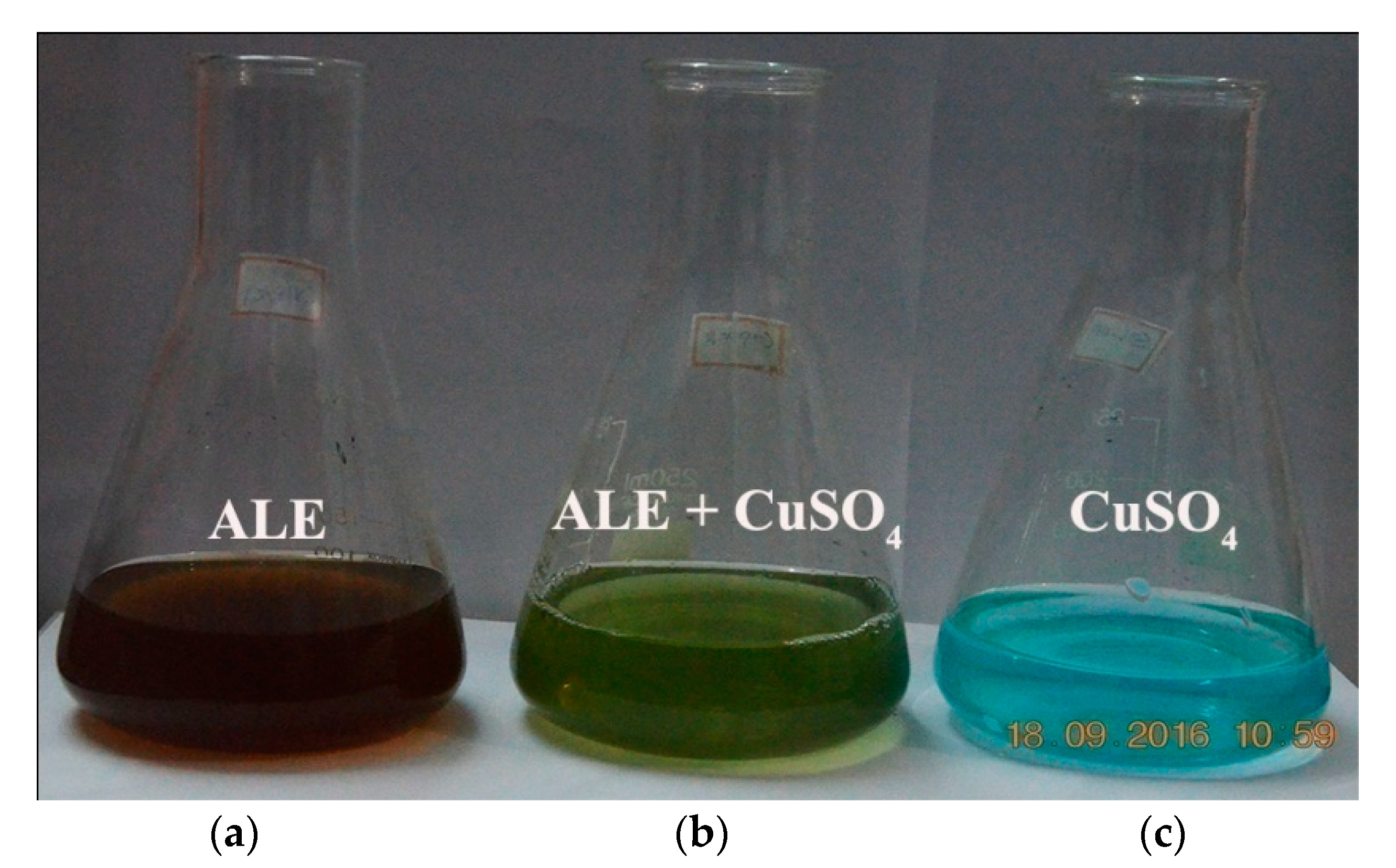
2.3. Green Synthesis of Copper Oxide Nanoparticles
2.4. Characterization of Copper Nanoparticles
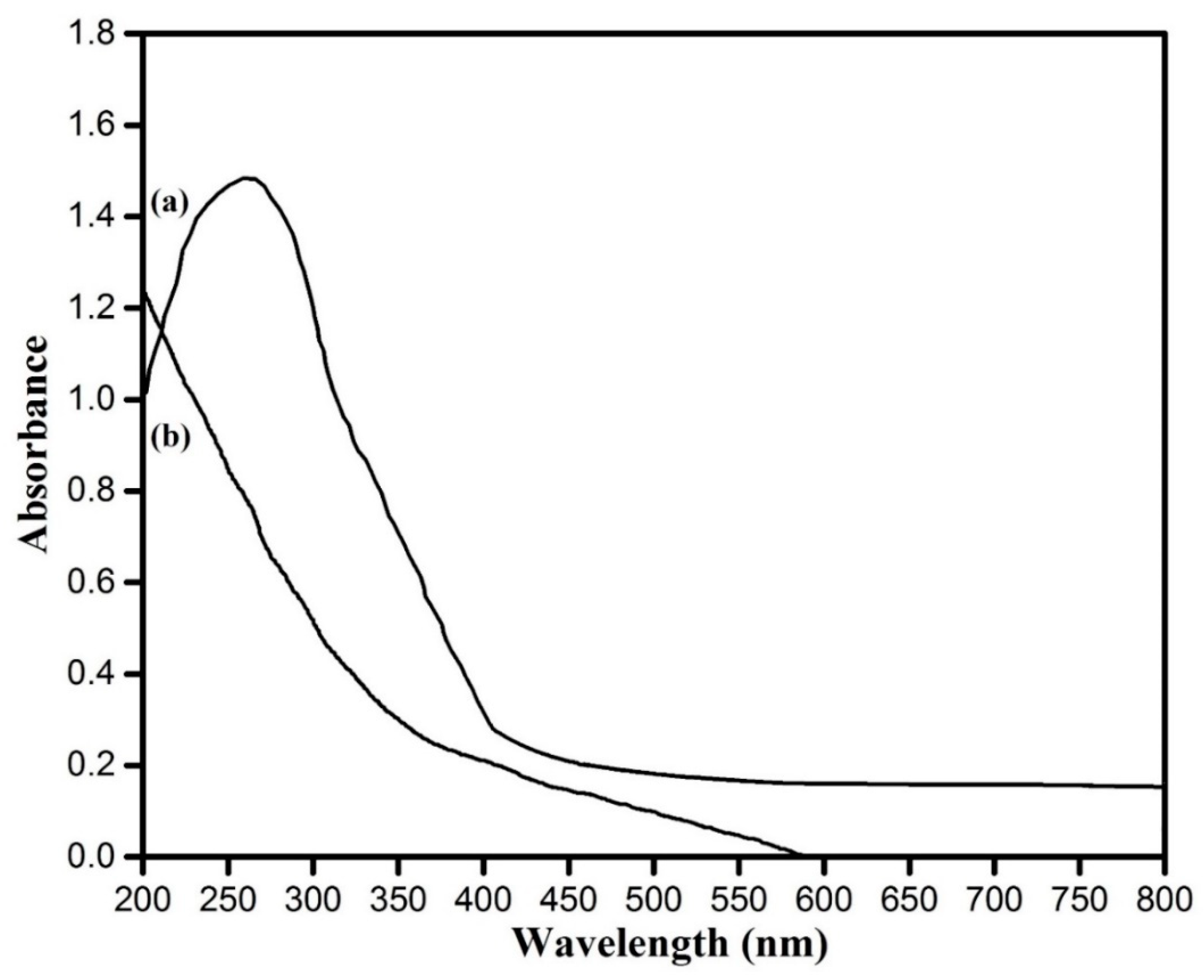
2.4.1. UV–Visible Spectroscopy of Synthesized Nanoparticles
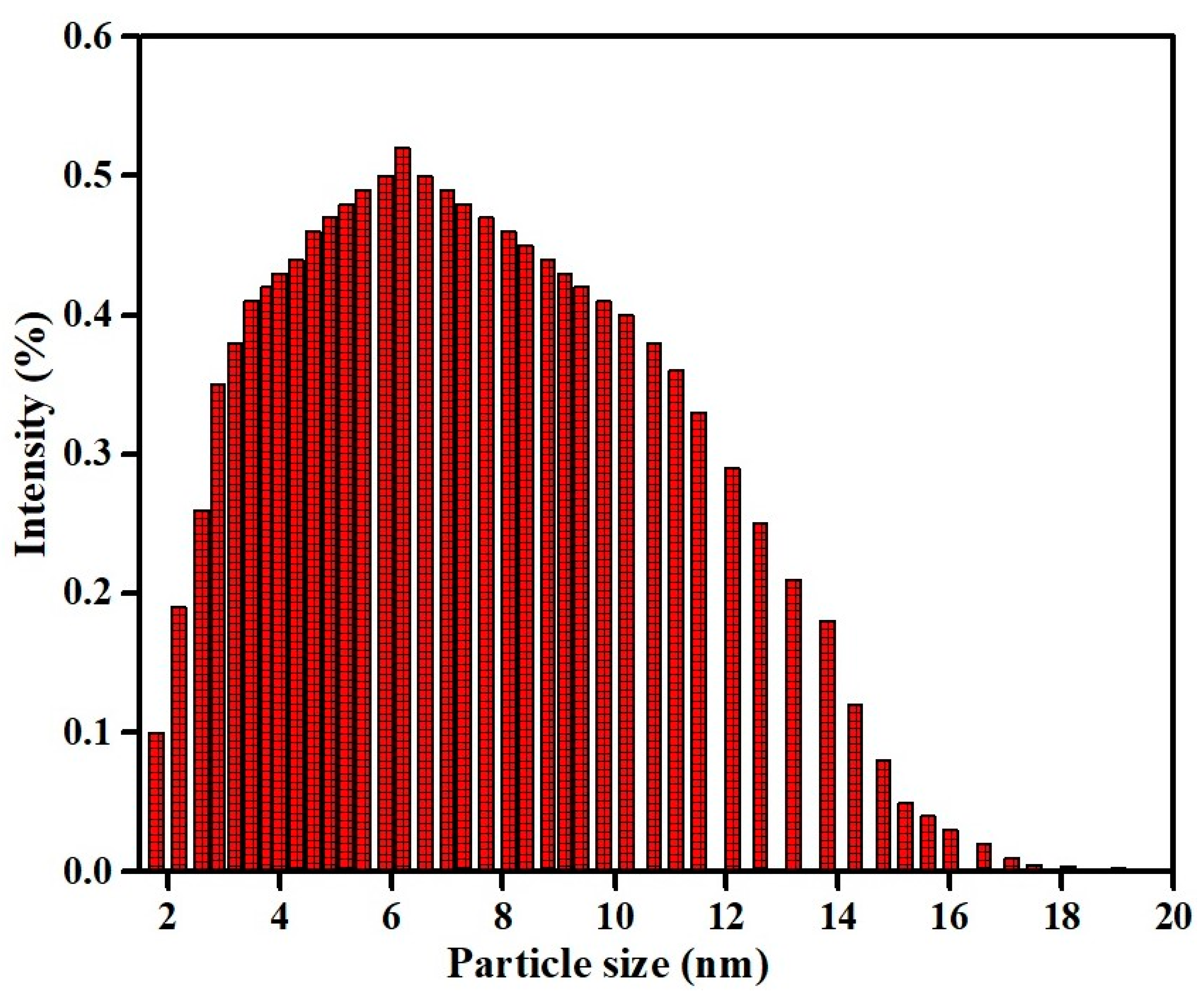
2.4.2. Particle Size Measurement by Dynamic Light Scattering (DLS) Experiment
2.4.3. Transmission Electron Microscopic (TEM) Observation of Nanoparticles
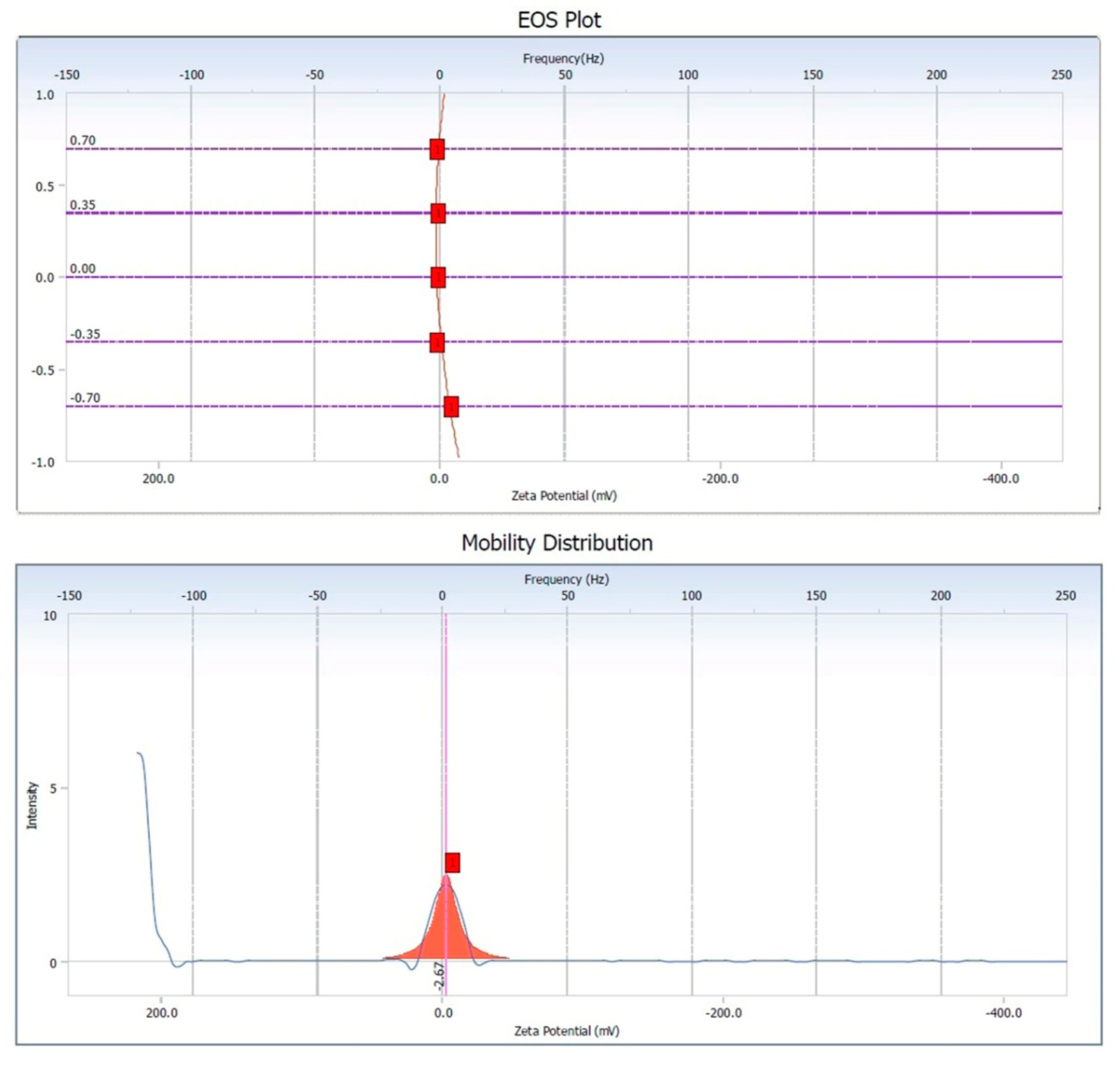
2.4.4. Zeta Potential Measurement
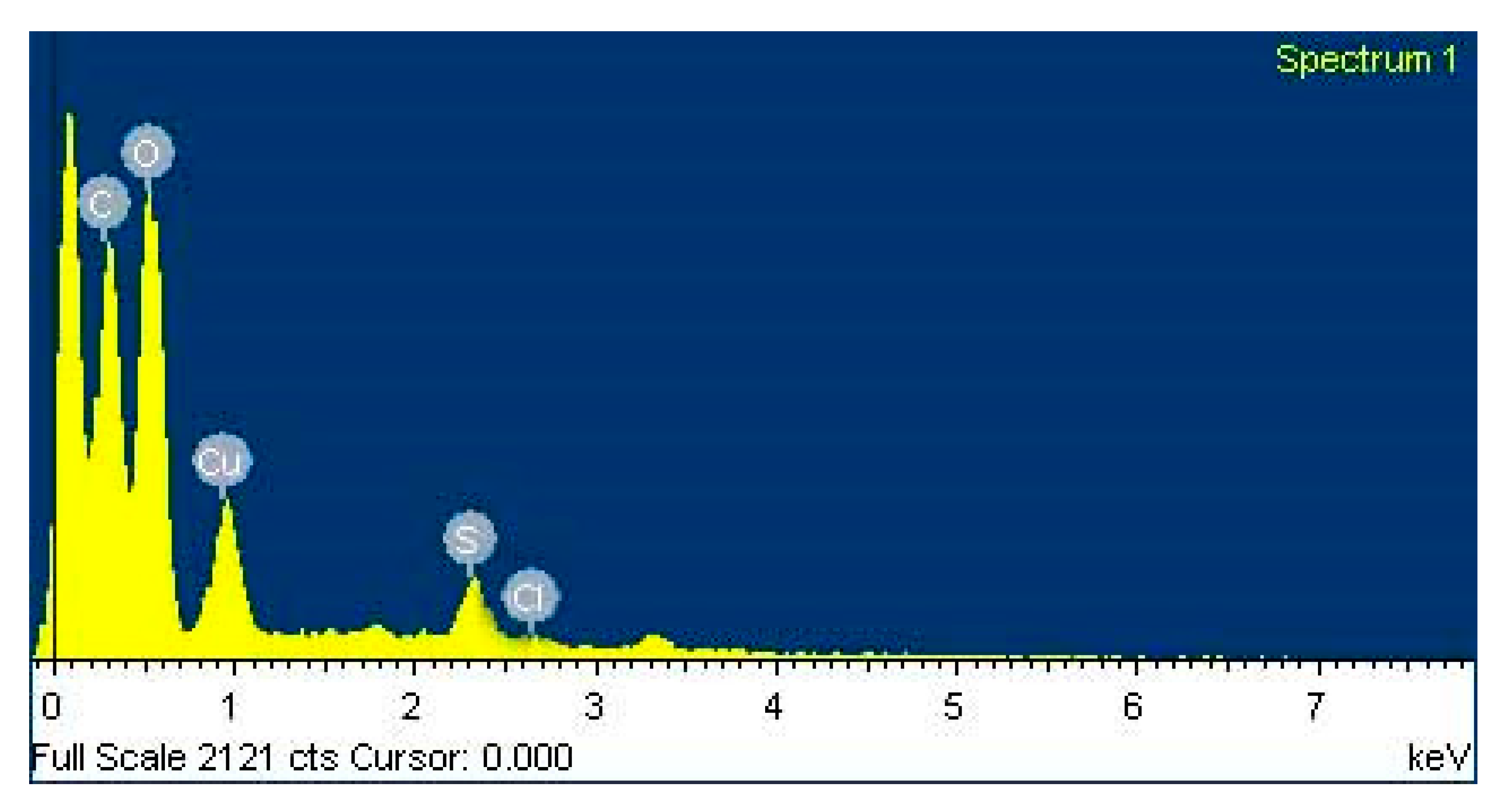
2.4.5. Investigation of Elemental Compositions of the Nanoparticles by Energy Dispersive Spectroscopy (EDX)
2.4.6. XRD Measurement
2.4.7. Fourier Transform Infrared Spectroscopic (FTIR) Analysis
2.5. Effects of CuONP on Seedling Germination and Seedling Development
2.6. Determination of Water Content (WC) and Relative Water Content (RWC)
2.7. Enzyme Extraction and Assays
2.7.1. Peroxidase Assay (PO)
2.7.2. Polyphenol Oxidase Assay (PPO)
2.7.3. Phenylalanine Ammonia Lyase (PAL)
2.7.4. β-1,3 glucanase Assay
2.7.5. Ascorbate Peroxidase Assay (APX)
2.7.6. Catalase Assay (CAT)
2.7.7. Superoxide Dismutase Assay (SOD)
2.8. Estimation of Total Protein Content
2.9. Estimation of Total Phenol Content
2.10. Estimation of Flavonoid Content
2.11. Determination of Lipid Peroxidation Rate
2.12. Estimation of Total Proline Content
2.13. Estimation of Nitric Oxide (NO)
2.14. Measurement of Reactive Oxygen Species (ROS)
2.15. In Vivo Detection of H2O2
2.16. Statistics
3. Results and Discussion
3.1. Characterization and Identification of the Plant Specimen
3.2. Production and Characterization of Copper Oxide Nanoparticles
3.3. UV–Visible Spectroscopic Analysis Copper Oxide Nanoparticles
3.4. Particle Size Measurement of Copper Oxide Nanoparticles
3.5. Zeta Potential of Copper Oxide Nanoparticles
3.6. EDX Observation of Copper Oxide Nanoparticles
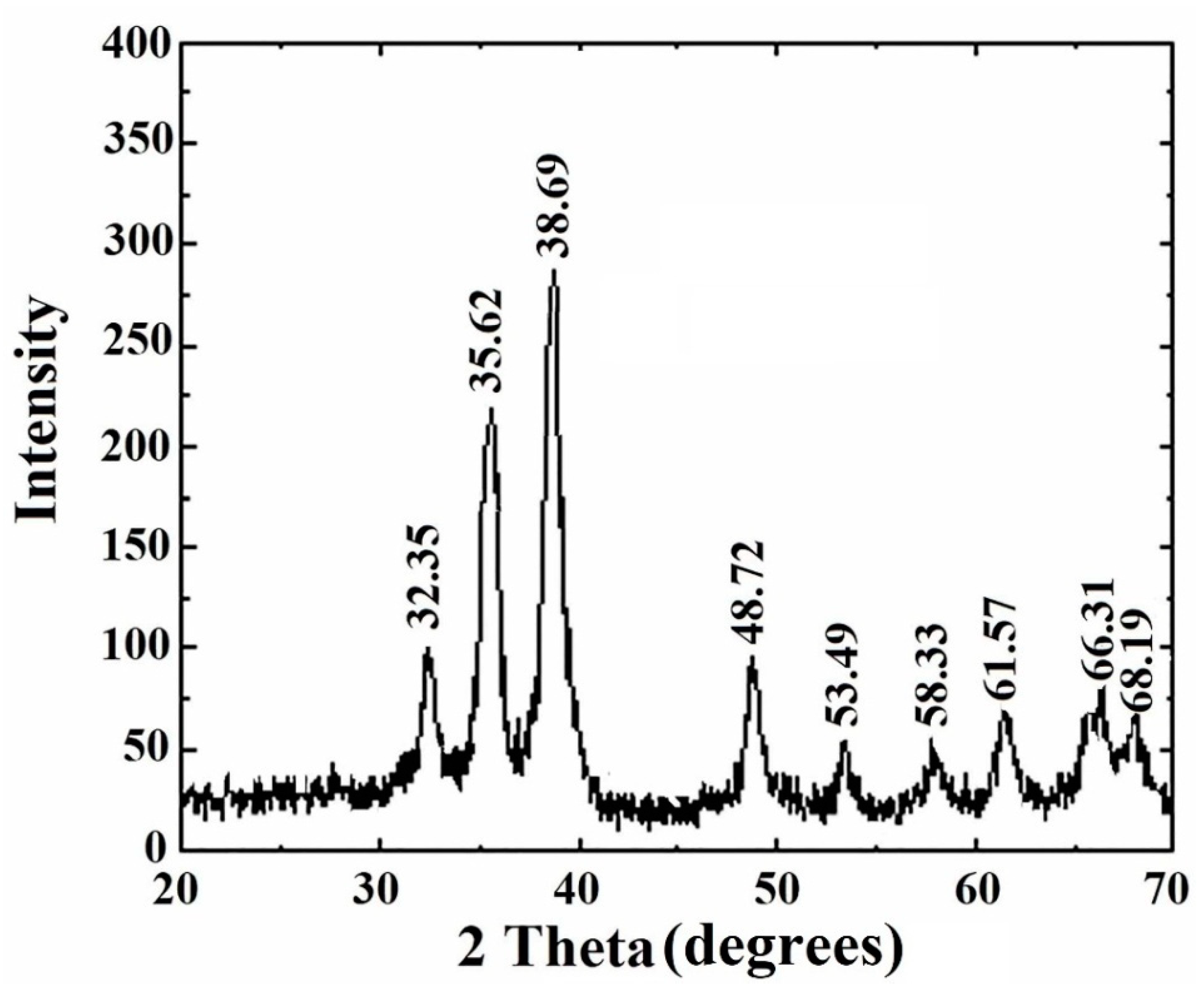
3.7. XRD Study of Copper Oxide Nanoparticles
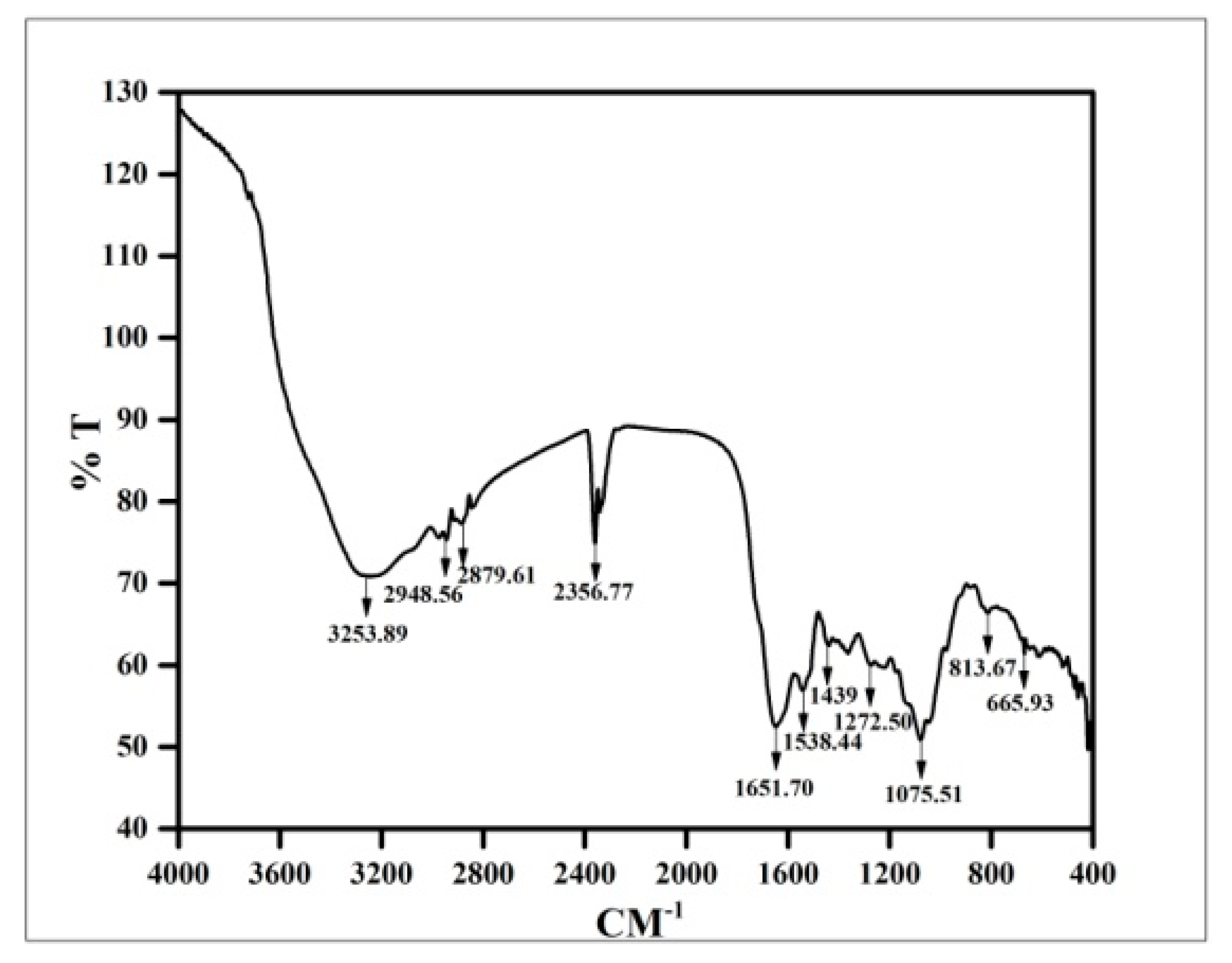
3.8. FTIR Analysis of Copper Oxide Nanoparticles
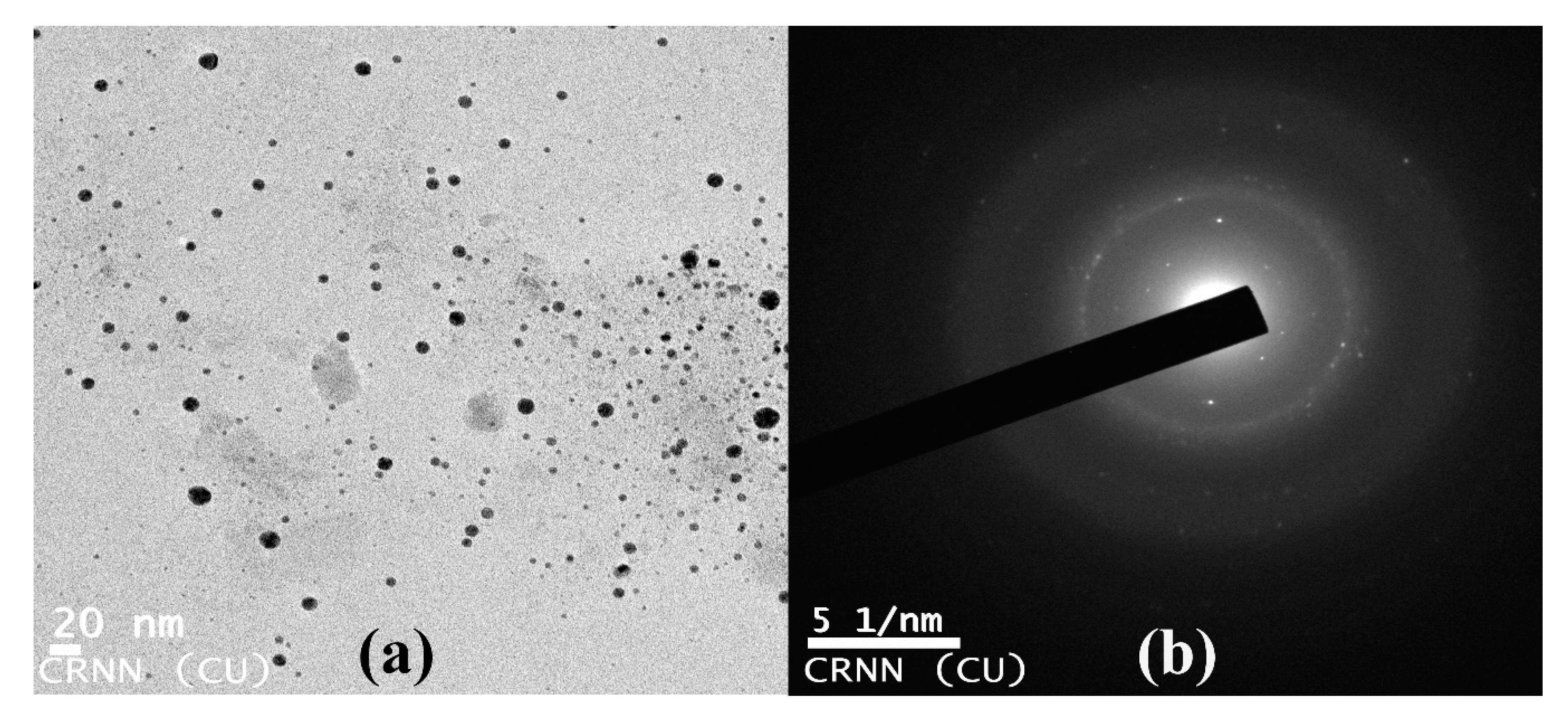
3.9. Transmission Electron Microscopy of Copper Oxide Nanoparticles
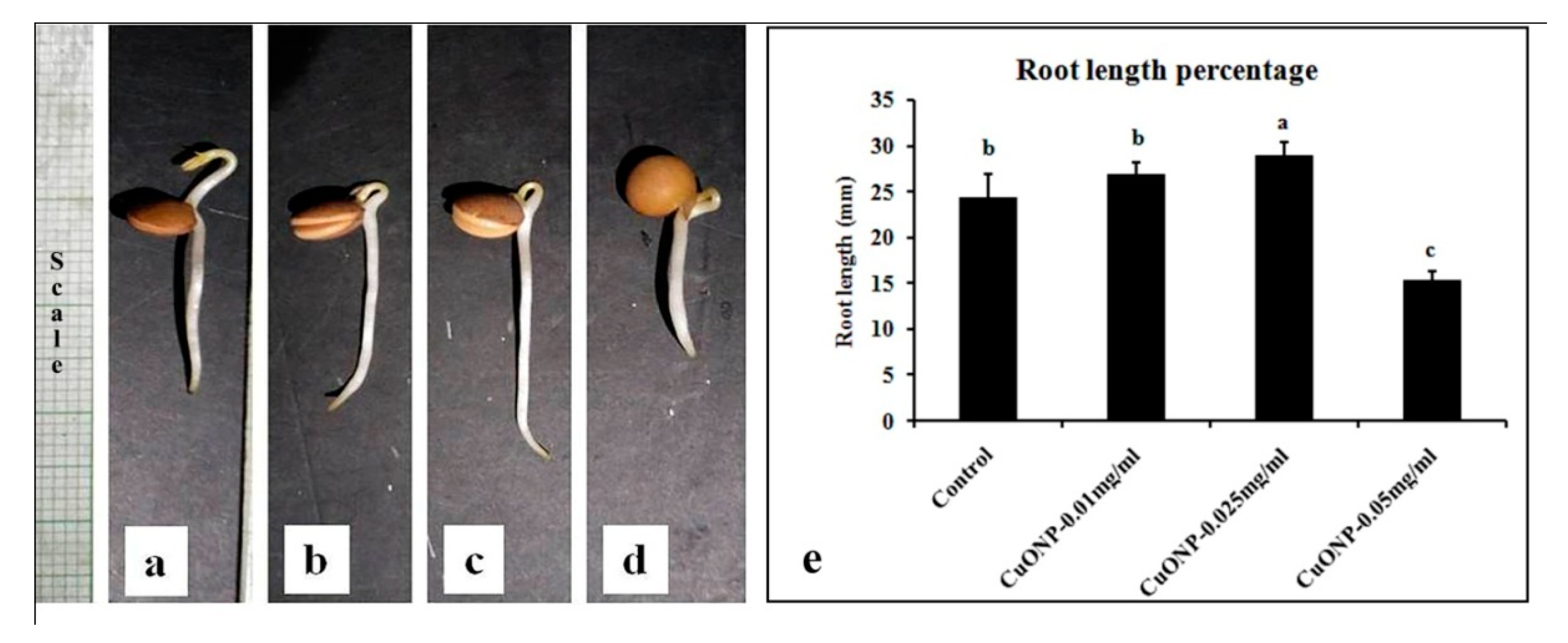
3.10. Effects of CuONPs on Seed Germination and Growth
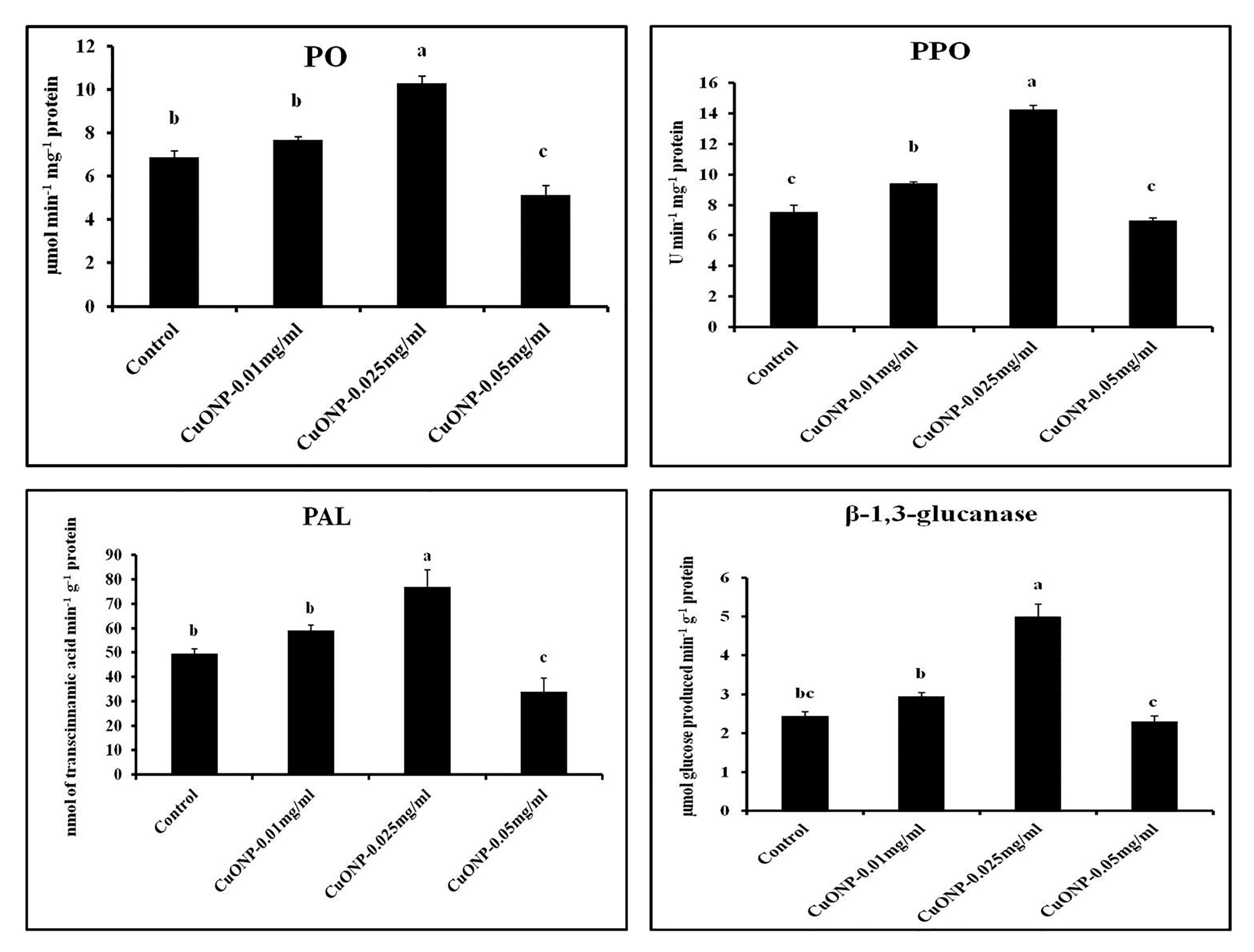
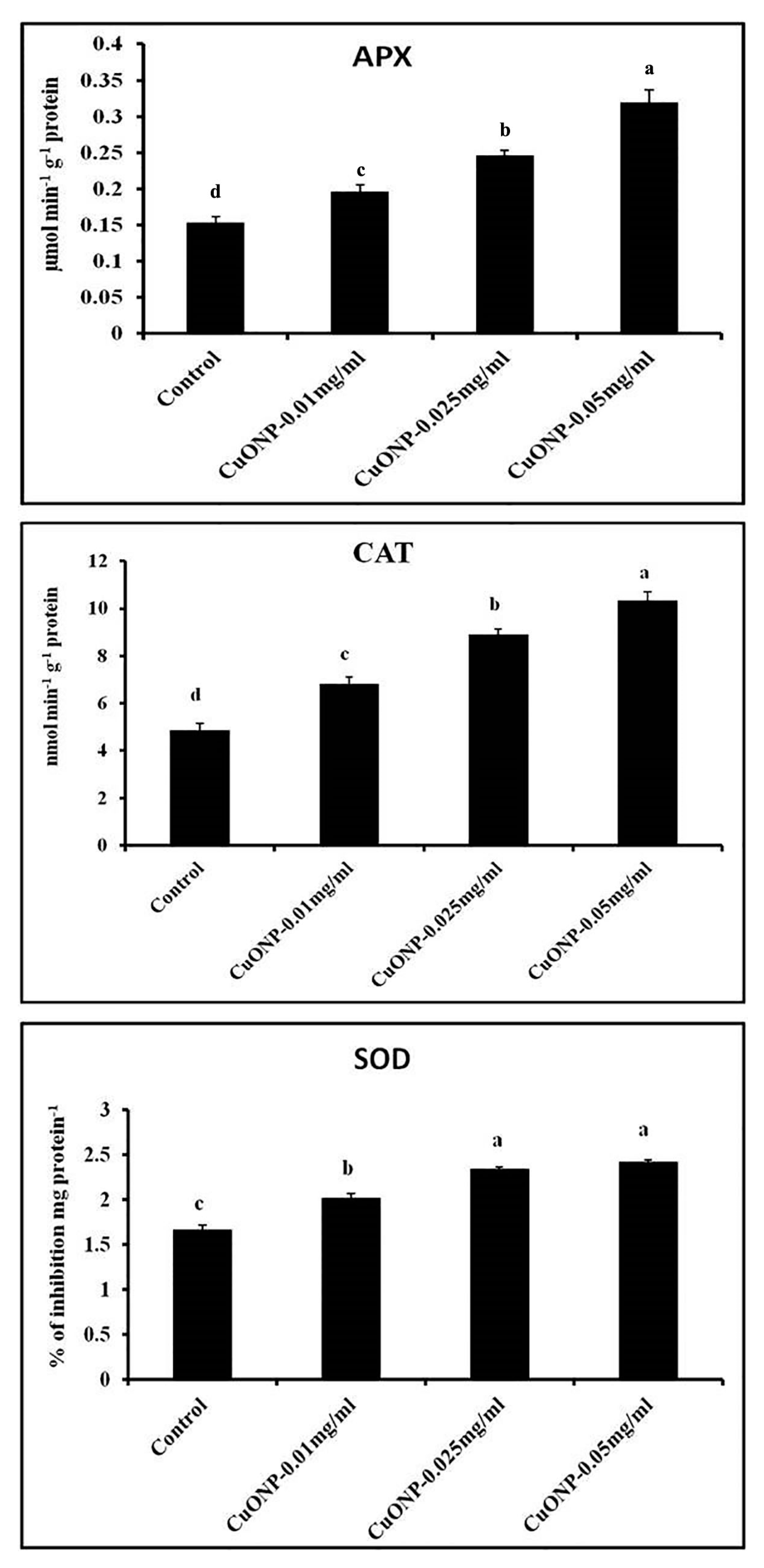
3.11. Effects of CuONPs on Defence Related and Antioxidative Enzymes
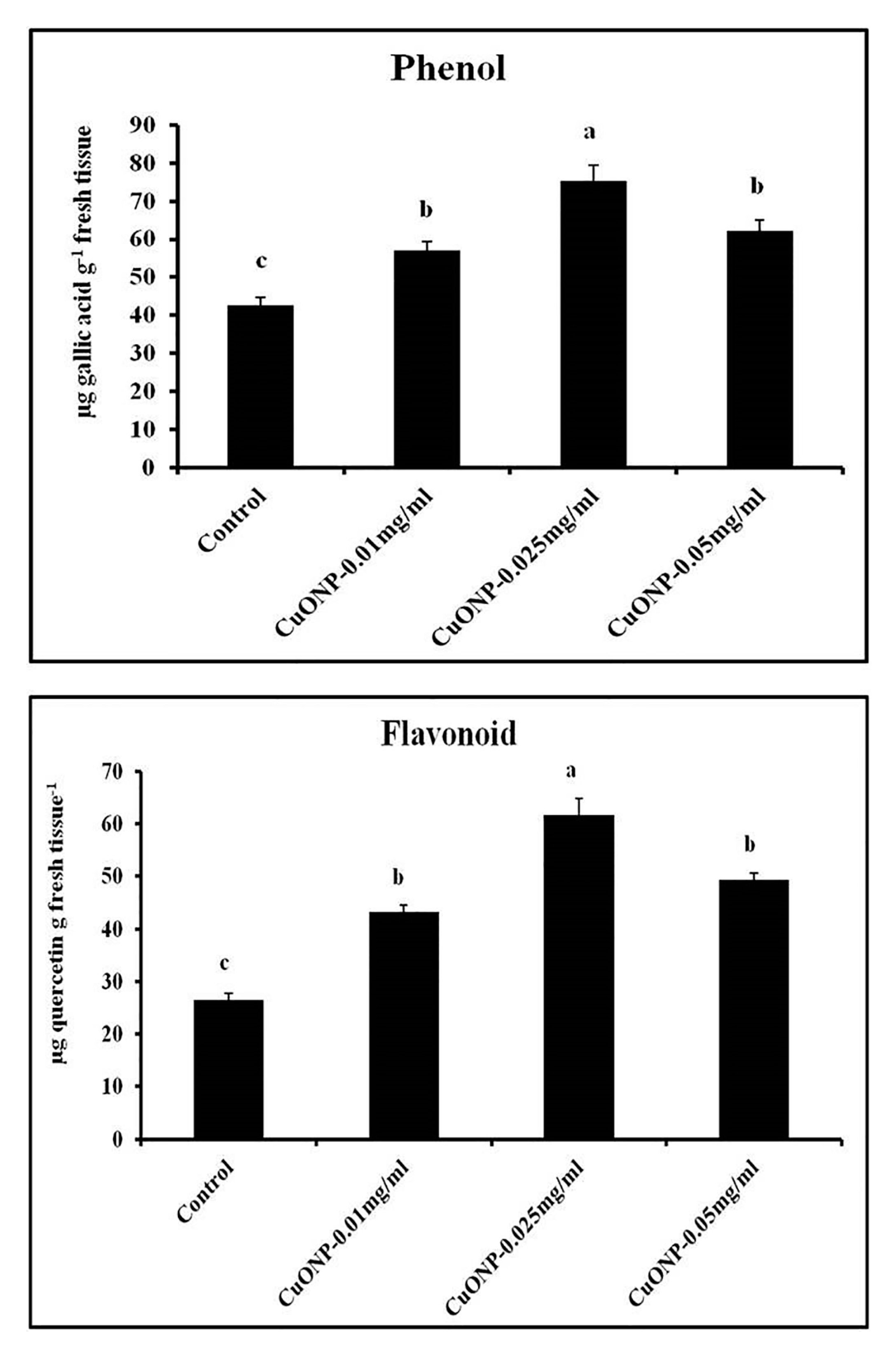
3.12. Effects of CuONPs on Phenol and Flavonoid Production
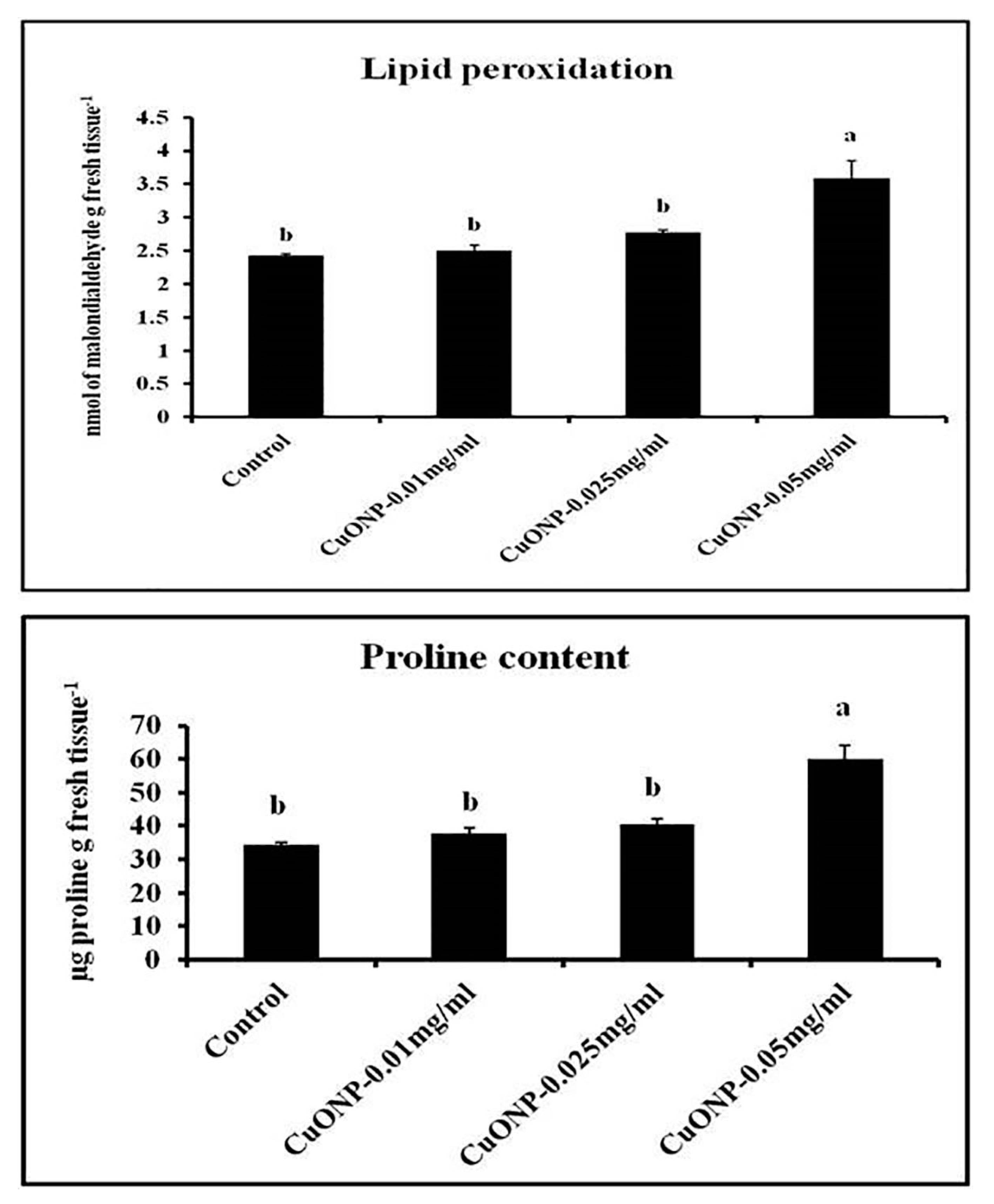
3.13. Effects of CuONPs on Lipid Peroxidation Rate and Proline Content
3.14. Effects of CuONPs on Nitric Oxide (NO) Production
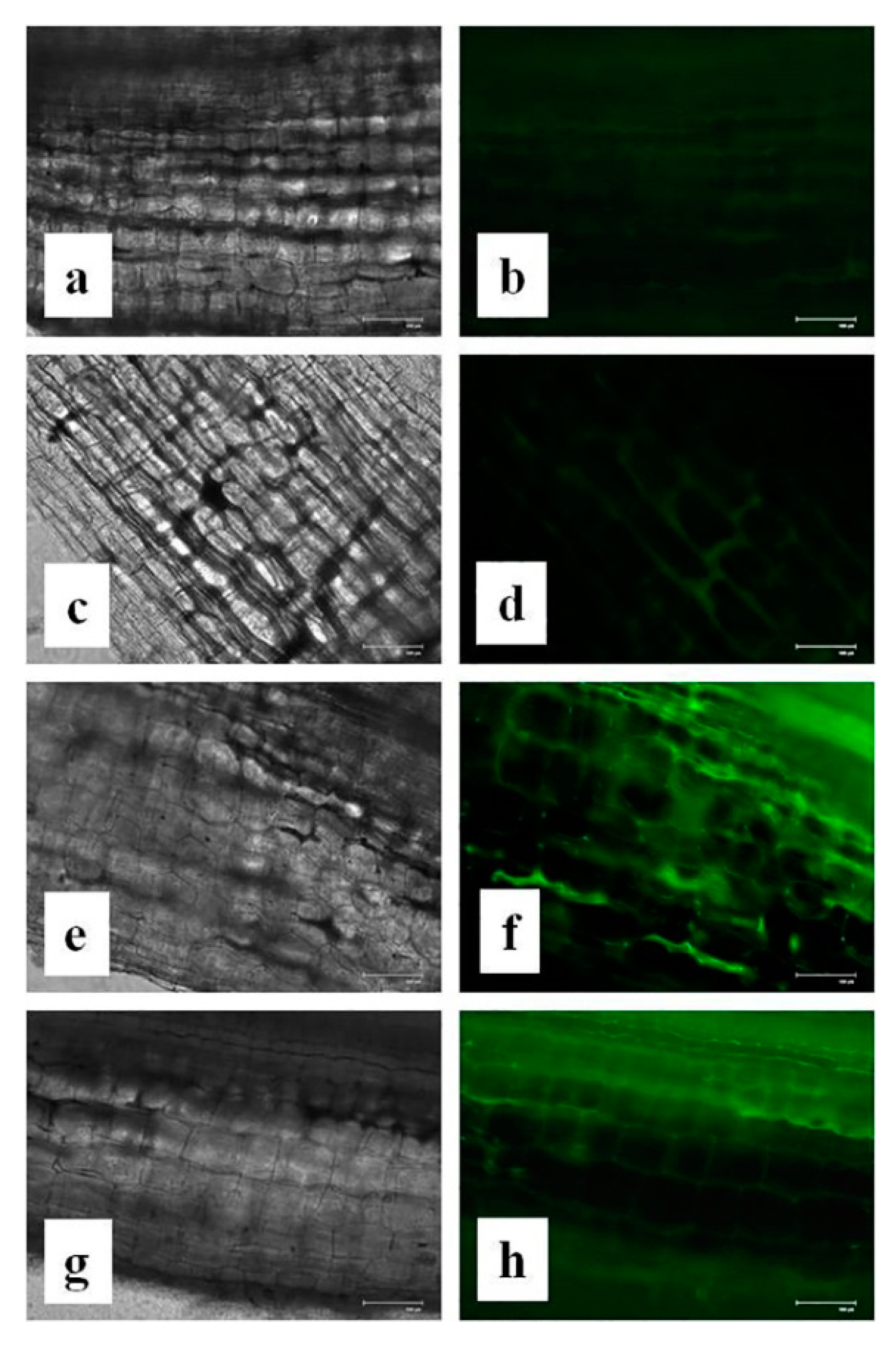
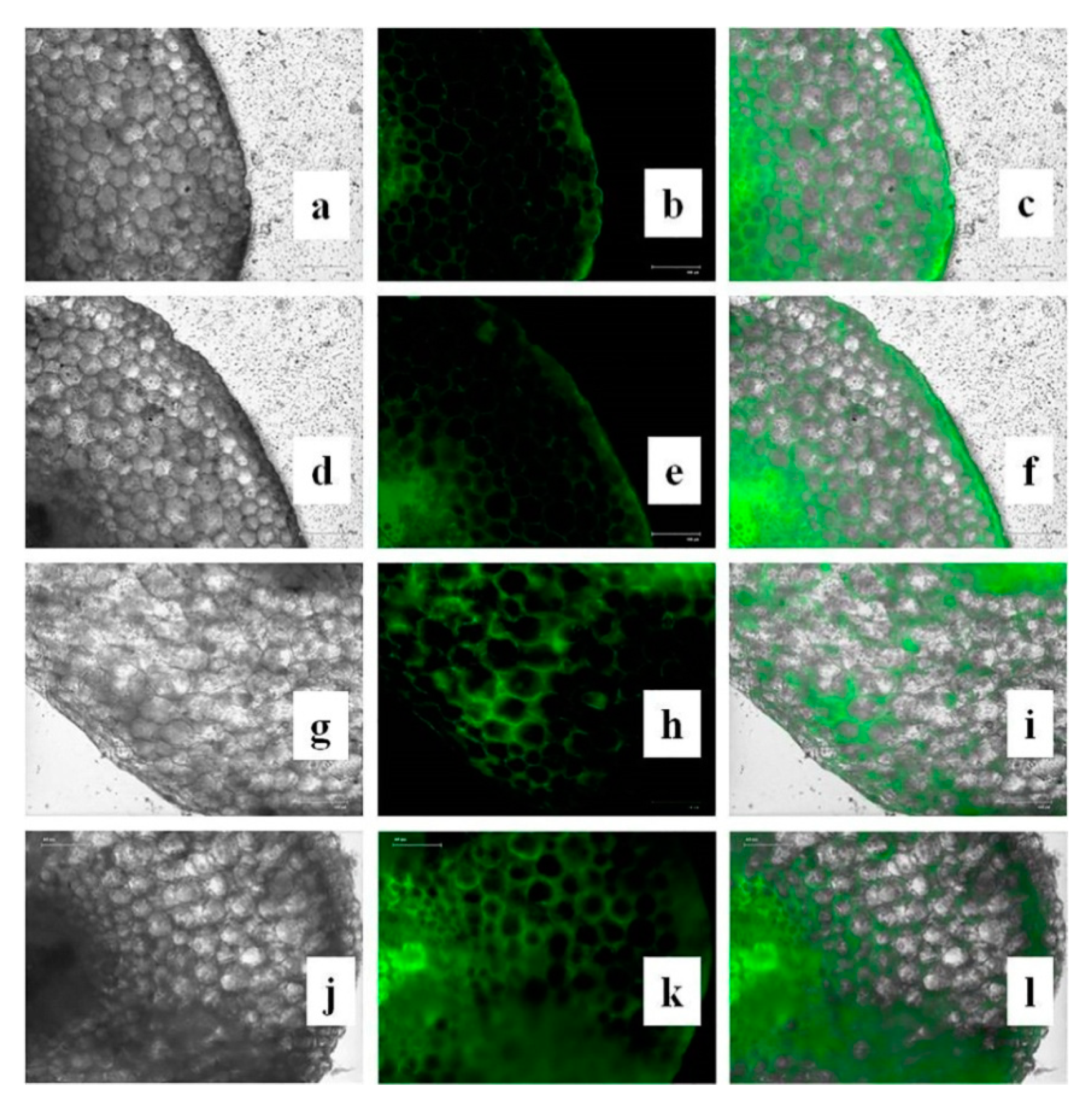
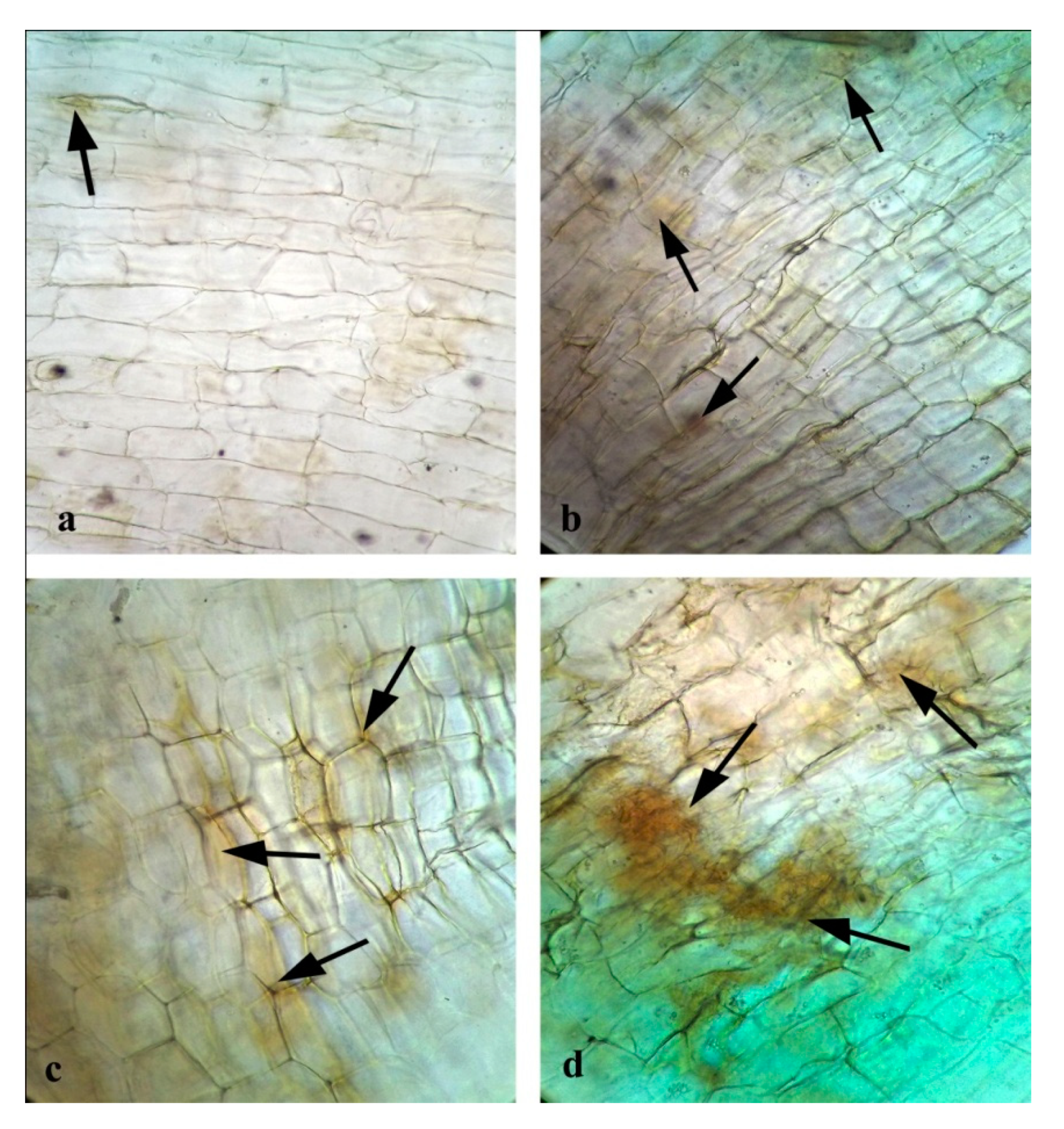
3.15. Effects of CuONPs on Reactive Oxygen Species (ROS) and H2O2 Production
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Passam, H.C.; Karapanos, I.C.; Bebeli, P.J.; Savvas, D. A Review of Recent Research on Tomato Nutrition, Breeding and Post-Harvest Technology with Reference to Fruit Quality. Eur. J. Plant Sci. Biotechnol. 2007, 1, 1–21. [Google Scholar]
- Shobha, G.; Moses, V.; Ananda, S. Biological Synthesis of Copper Nanoparticles and its impact—A Review. Int. J. Pharm. Sci. Invent. 2014, 3, 28–38. [Google Scholar]
- Yruela, I. Copper in plants. Braz. J. Plant Physiol. 2005, 17, 145–156. [Google Scholar] [CrossRef]
- Manceau, K.L.; Nagy, M.A.; Marcus, M.; Lanson, N.; Geoffroy, T.J.; Kirpichtchikova, T. Formation of metallic copper nanoparticles at the soil-root interface. Environ. Sci. Technol. 2008, 1, 1766–1772. [Google Scholar] [CrossRef]
- Filipe, V.; Hawe, A.; Jiskoot, W. Critical evaluation of Nanoparticle Tracking Analysis (NTA) by NanoSight for the measurement of nanoparticles and protein aggregates. Pharmaceut. Res. 2010, 27, 796–810. [Google Scholar] [CrossRef]
- Yedurkar, S.M.; Maurya, C.B.; Mahanwar, P.A. A Biological Approach for the Synthesis of Copper Oxide Nanoparticles by Ixora coccinea Leaf Extract. J. Mater. Environ. Sci. 2017, 8, 1173–1178. [Google Scholar]
- Saif, S.; Tahir, A.; Chen, Y. Green Synthesis of Iron Nanoparticles and Their Environmental Applications and Implications. Nanomaterials 2016, 6, 209. [Google Scholar] [CrossRef]
- Makarov, V.V.; Love, A.J.; Sinitsyna, O.V.; Makarova, S.S.; Yaminsky, I.V.; Taliansky, M.E.; Kalinina, N.O. “Green” nanotechnologies: Synthesis of metal nanoparticles using plants. Acta Nat. 2014, 6, 35–44. [Google Scholar] [CrossRef]
- Ahmed, S.; Ahmad, M.; Swami, B.L.; Ikram, S. A review on plants extract mediated synthesis of silver nanoparticles for antimicrobial applications: A green expertise. J. Adv. Res. 2016, 7, 17–28. [Google Scholar] [CrossRef]
- Küünal, S.; Rauwel, P.; Rauwel, E. Plant extract mediated synthesisof nanoparticles. In Emerging Applications of Nanoparticles and Architecture Nanostructures, 1st ed.; Barhoum, A., Makhlouf, A.S.H., Eds.; Elsevier: Cambridge, MA, USA, 2018; pp. 411–446. [Google Scholar] [CrossRef]
- Singh, V.; Patil, R.; Ananda, A.; Milani, P.; Gade, W. Biological Synthesis of Copper Oxide Nano Particles Using Escherichia coli. Curr. Nanosci. 2010, 6, 365–369. [Google Scholar] [CrossRef]
- Usha, R.; Prabu, E.; Palaniswamy, M.; Venil, C.K.; Rajendran, R. Synthesis of metal oxide nanoparticles by Streptomyces sp. for development of antimicrobial textiles. Glob. J. Biotechnol. Biochem. 2010, 5, 153–160. [Google Scholar]
- Abboud, Y.; Saffaj, T.; Chagraoui, A.; Bouari, E.; Tanane, K.B.; Ihssane, O. Biosynthesis, characterization and antimicrobial activity of copperoxide nanoparticles (CONPs) produced using brown alga extract (Bifurcaria bifurcata). Appl. Nanosci. 2013, 4, 571–576. [Google Scholar] [CrossRef]
- Honary, S.; Barabadi, H.; Fathabad, E.G.; Naghibi, F. Green synthesis of copper oxide nanoparticles using Penicillium aurantiogriseum, Penicillium citrinum and Penicillium wakasmanii. Digest J. Nanomater. Biostruct. 2012, 7, 999–1005. [Google Scholar]
- Salvadori, M.R.; Lepre, L.F.; Ando, R.A.; do Nascimento, C.A.O.; Corrêa, B. Biosynthesis and Uptake of Copper Nanoparticles by Dead Biomass of Hypocrea lixii isolated from the Metal Mine in the Brazilian Amazon Region. PLoS ONE 2013, 8, e80519. [Google Scholar] [CrossRef] [PubMed]
- Harne, S.; Sharma, A.; Dhaygude, M.; Joglekar, S.; Kodam, K.; Hudlikar, M. Novel route for rapid biosynthesis of copper nanoparticles using aqueous extract of Calotropis procera L. latex and their cytotoxicity on tumor cells. Colloids Surf. B 2012, 15, 284–288. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.-J.; Song, J.Y.; Kim, B.S. Biological synthesis of copper nanoparticles using Magnolia kobus leaf extract and their antibacterial activity. J. Chem. Technol. Biotechnol. 2013, 8, 1971–1977. [Google Scholar]
- Acharya, K.; Sarkar, J. Bryo-synthesis of gold nanoparticles. Int. J. Pharmaceut. Sci. Rev. Res. 2014, 29, 82–86. [Google Scholar]
- Baskaran, X.; Vigila, A.V.G.; Parimelazhagan, T.; Muralidhara-Rao, D.; Zhang, S. Biosynthesis, characterization, and evaluation of bioactivities of leaf extract-mediated biocompatible silver nanoparticles from an early tracheophyte, Pteristripartita Sw. Int. J. Nanomed. 2016, 11, 5789–5805. [Google Scholar] [CrossRef]
- Santhoshkumar, S.; Nagarajan, N. Biological synthesis of silver nanoparticles of Adiantum capillusveneris L. and their evaluation of antibacterial activity against human pathogenic bacteria. Int. J. Pharm. Sci. Res. 2014, 5, 5511–5518. [Google Scholar] [CrossRef]
- Britto, A.J.; Gracelin, D.H.S.; Jeya, P.B.; Kumar, R.; Molecular, P.; Nadu, T. Biogenic silver nanoparticles by Adiantum caudatum and their antibacterial activity. Int. J. Univer. Pharm. Life Sci. 2012, 2, 92–98. [Google Scholar]
- Sant, D.G.; Gujarathi, T.R.; Harne, S.R.; Ghosh, S.; Kitture, R.; Kale, S.; Chopade, B.K.; Pardesi, K.R. Adiantum philippense L. Frond Assisted Rapid Green Synthesis of Gold and Silver Nanoparticles. J. Nanopart. 2013, 1–9. [Google Scholar] [CrossRef]
- Kalita, S.; Kandimalla, R.; Sharma, K.K.; Kataki, A.C.; Deka, M.; Kotoky, J. Amoxicillin functionalized gold nanoparticles reverts MRSA resistance. Mater. Sci. Eng. C. 2016, 61, 720–727. [Google Scholar] [CrossRef] [PubMed]
- Şuţan, N.A.; Fierăscu, I.; Fierăscu, R.C.; Manolescu, D.Ş.; Soare, L.C. Comparative analytical characterization and in vitro cytogenotoxic activity evaluation of Asplenium scolopendrium L. leaves and rhizome extracts prior to and after Ag nanoparticles phytosynthesis. Ind. Crops Prod. 2016, 83, 379–386. [Google Scholar] [CrossRef]
- Koteswaramma, B.; Kamakshamma, J.; Varalaksmi, S. Biological synthesis of silver nanopartical from liquid extract of Actiniopteris radiata and evaluation of their antimicrobial activity. Int. J. Pharm. Biol. Sci. 2017, 8, 121–125. [Google Scholar]
- Ch, B.; Kunjiappan, S.; Bhattacharjee, C.; Chowdhury, R. Hepatoprotective and antioxidant effects of Azolla microphylla based gold nanoparticles against acetaminophen induced toxicity in a fresh water common carp fish (Cyprinus carpio L.) Biological activity of Azolla microphylla-based gold nanoparticles. Nanomed. J. 2015, 2, 88–110. [Google Scholar]
- Potter, D.M.; Baird, M.S. Carcinogenic effects of ptaquiloside in bracken fern and related compounds. Br. J. Cancer. 2000, 83, 914–920. [Google Scholar] [CrossRef]
- Johnson, M. Antibacterial Potentials of Adiantum Species against the UTI Pathogens. J. Microbiol. Exp. 2017, 4, 00104. [Google Scholar] [CrossRef]
- Ali, M.S.; Amin, M.R.; Kamal, C.M.I.; Hossain, M.A. In vitro antioxidant, cytotoxic, thrombolytic activities and phytochemical evaluation of methanol extract of the A. philippense L. leaves. Asian Pac. J. Trop. Biomed. 2013, 3, 464–469. [Google Scholar] [CrossRef]
- Paul, T.; Das, B.; Apte, K.G.; Banerjee, S.; Saxena, R.C. Evaluation of Anti-Hyperglycemic Activity of Adiantum philippense Linn, a Pteridophyte in Alloxan Induced Diabetic Rats. J. Diab. Metabol. 2012, 3, 1000226. [Google Scholar] [CrossRef]
- Sikarwar, R.; Pathak, B.; Jaiswal, A. Some unique ethnomedicinal perceptions of tribal communities of Chitrakoot, Madhya Pradesh. Int. J. Trad. Knowl. 2008, 7, 613–617. [Google Scholar]
- Pan, C.; Chen, Y.G.; Ma, X.Y.; Jiang, J.H.; He, F.; Zhang, Y. Phytochemical constituents and pharmacological activities of plants from the genus Adiantum: A review. Trop. J. Pharmaceut. Res. 2011, 10, 681–692. [Google Scholar] [CrossRef]
- Mithraja, M.; Marimuthu, J.; Mahesh, M.; Paul, Z. Inter–specific variation studies on the phyto–constituents of Christella and Adiantum using phytochemical methods. Asian Pac. J. Trop. Biomed. 2012, 2, S40–S45. [Google Scholar] [CrossRef]
- Chandra, S.; Chakraborty, N.; Dasgupta, A.; Sarkar, J.; Panda, K.; Acharya, K. Chitosan nanoparticles: A positive modulator of innate immune responses in plants. Sci. Rep. 2015, 5, 15195. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, J.; Chattopadhyay, D.; Patra, S.; Singh Deo, S.; Sinha, S.; Ghosh, M.; Mukherjee, A.; Acharya, K. Alternaria alternata Mediated Synthesis of Protein Capped Silver Nanoparticles And Their Genotoxic Activity. Digest J. Nanomat. Biostruct. 2011, 6, 563–573. [Google Scholar]
- Mollick, M.M.R.; Bhowmick, B.; Maity, D.; Mondal, D.; Bain, M.K.; Bankura, K.; Sarkar, J.; Rana, D.; Acharya, K.; Chattopadhyay, D. Green Synthesis of Silver Nanoparticles Using Paederia foetida L. Leaf Extract and Assessment of Their Antimicrobial Activities. Int. J. Green Nanotechnol. 2012, 4, 230–239. [Google Scholar] [CrossRef]
- Viet, P.V.; Nguyen, H.T.; Cao, T.M.; Hieu, L.V. Fusarium Antifungal Activities of Copper Nanoparticles Synthesized by a Chemical Reduction Method. J. Nanomat. 2016. [Google Scholar] [CrossRef]
- Shi, J.; Peng, C.; Yang, Y.; Yang, J.; Zhang, H.; Yuan, X.; Chen, Y.; Hu, T. Phytotoxicity and accumulation of copper oxide nanoparticles to the Cu-tolerant plant Elsholtzia splendens. Nanotoxicology 2014, 8, 179–188. [Google Scholar] [CrossRef]
- Nair, P.M.G.; Kim, S.-H.; Chung, I.M. Copper oxide nanoparticle toxicity in mung bean (Vignaradiata L.) seedlings: Physiological and molecular level responses of in vitro grown plants. Acta. Physiol. Plant. 2014, 36, 2947–2958. [Google Scholar] [CrossRef]
- Behera, M.; Giri, G. Green synthesis and characterization of cuprous oxide nanoparticles in presence of a bio-surfactant. Mat. Sci.-Pol. 2014, 32, 702–708. [Google Scholar] [CrossRef]
- Henglein, A. Physichochemical Properties of Small Metal Particles in Solution: “Microelectrode” Reactions, Chemisorption, Composite Metal Particles, and the Atom-to-Metal Transition. J. Phys. Chem. 1993, 97, 5457–5471. [Google Scholar] [CrossRef]
- Vigneshwaran, N.; Ashtaputre, N.M.; Varadarajan, P.V.; Nachane, R.P.; Paralikar, K.M.; Balasubramanya, R.H. Biological synthesis of silver nano particles using the fungus Aspergillus flavus. Mat. Lett. 2007, 61, 1413–1418. [Google Scholar] [CrossRef]
- Pal, S.; Tak, Y.; Song, J. Does the antibacterial activity of silver nanoparticles depend on the shape of the nanoparticle? A study of the Gram-negative bacterium Escherichia coli. Appl. Environ. Microbiol. 2007, 73, 1712–1720. [Google Scholar] [CrossRef]
- Saha, S.; Sarkar, J.; Chattopadhyay, D.; Patra, S.; Chakraborty, A.; Acharya, K. Production of silver nanoparticles by a phytopathogenic fungus Bipolaris nodulosa and its antimicrobial activity. Dig. J. Nanomater. Biostruc. 2010, 5, 887–895. [Google Scholar]
- Sastry, M.; Ahmad, A.; Khan, I.; Kumar, R. Biosynthesis of metal nanoparticles using fungi and actinomycete. Curr Sci. 2003, 85, 162–170. [Google Scholar]
- Sadowski, Z.; Maliszewska, I.H.; Grochowalska, B.; Polowczyk, I.; Kozlecki, T. Synthesis of silver nanoparticles using microorganisms. Mater. Sci. 2008, 26, 419–424. [Google Scholar]
- Balestri, M.; Bottega, S.; Spanὸ, C. Response of Pteris vittata to different cadmium treatments. Acta Physiol. Plant. 2014, 36, 767–775. [Google Scholar] [CrossRef]
- Hemada, H.M.; Klein, B.P. Effects of naturally occurring antioxidants on peroxidase activity of vegetable extracts. J. Food Sci. 1990, 55, 184–185. [Google Scholar] [CrossRef]
- Kumar, K.B.; Khan, P.A. Peroxidase and Polyphenoloxidase in excised ragi (Eleusine coracana cv PR 202) leaves during senescence. Ind. J. Exp. Bot. 1982, 20, 412–416. [Google Scholar]
- Dickerson, D.P.; Pascholati, S.F.; Gagerman, A.E.; Butler, L.G.; Nicholson, R.L. Phenylalanine ammonia-lyase and hydroxycinnamate, CoA ligase in maize mesocotyls inoculated with Helminthosporium maydis or Helminthosporium carbonum. Physol. Plant Pathol. 1984, 25, 111–123. [Google Scholar] [CrossRef]
- Pan, S.Q.; Ye, X.S.; Kuc, J. Association of β-1,3-glucanase activity and isoform pattern with systemic resistance to blue mould in tobacco induced by stem injection with Peronospora tabacina or leaf inoculation with tobacco mosaic virus. Physiol. Mol. Plant Pathol. 1991, 39, 25–39. [Google Scholar] [CrossRef]
- Nakano, Y.; Asada, K. Hydrogen peroxide is scavenged by ascorbate-specific peroxidase in spinach chloroplasts. Plant Cell Physiol. 1981, 22, 867–880. [Google Scholar]
- Cakmak, I.; Horst, J. Effect of aluminium on lipid peroxidation, superoxide dismutase, catalase, and peroxidase activities in root tips of soybean (Glycine max). Physiol. Plant. 1991, 83, 463–468. [Google Scholar] [CrossRef]
- Dhindsa, R.H.; Plumb-Dhindsa, R.; Thorpe, T.A. Leaf senescence correlated with increased level of membrane permeability, lipid peroxidation and decreased level of SOD and CAT. J. Exp. Bot. 1981, 32, 93–101. [Google Scholar] [CrossRef]
- Lowry, O.H.; Rosebrough, N.J.; Farr, A.L.; Randall, R.J. Protein measurement with the Folin phenol reagent. J. Biol. Chem. 1951, 193, 265–275. [Google Scholar] [PubMed]
- Zieslein, N.; Ben-Zaken, R. Peroxidase activity and presence of phenolic substances in peduncles in rose flowers. Plant Physiol. Biochem. 1993, 31, 333–339. [Google Scholar]
- Chang, C.; Yang, M.; Wen Hand Chem, J. Estimation of total flavonoid content in propolis by two complementary colorimetric methods. J. Food Drug Anal. 2002, 10, 178–182. [Google Scholar]
- Bates, L.S. Rapid determination of free proline for water-stress studies. Plant Soil. 1973, 39, 205–207. [Google Scholar] [CrossRef]
- Bartha, B.; Kolbert, Z.; Erdei, L. Nitric oxide production induced by heavy metals in Brassica juncea L. Czern and Pisum sativum L. Acta Biol. Szeged. 2005, 49, 9–12. [Google Scholar]
- Gupta, K.J.; Brotman, Y.; Mur, L.A. Localisation and Quantification of Reactive Oxygen Species and Nitric Oxide in Arabidopsis Roots in Response to Fungal Infection. Bio-Protocol 2014, 4, e1259. [Google Scholar] [CrossRef]
- Thordal-Christensen, H.; Zhang, Z.; Wei, Y.; Collinge, D.B. Subcellular localization of H2O2 in plants. H2O2 accumulation in papillae and hypersensitive response during the barley powdery mildew interaction. Plant J. 1997, 11, 1187–1194. [Google Scholar] [CrossRef]
- Hillis, D.M. Molecular Versus Morphological Approaches to Systematics. Ann. Rev. Eco. Syst. 1987, 18, 23–42. [Google Scholar] [CrossRef]
- Pallavi, G.; Virupaksha, G.K.L.; Chate, V.A. An Ethno-Pharmaco-Botanical Review of Hamsapadi—Adiantum lunulatum Burm. F. (A. Philippense Linn.). Int. J. Pharm. Biol. Arch. 2011, 2, 1668–1676. [Google Scholar]
- Kramer, K.U.; Green, P.S.; Peter, S.; Götz, E. Pteridophytes and Gymnosperms, 1st ed.; Springer: Berlin/Heidelberg, Germany, 2014; p. 404. [Google Scholar]
- Van den Boom, P.P.G. lichens and lichenicolous fungi of the azores (portugal), collected on sãomiguel and terceira with the descriptions of seven new species. Acta Bot. Hung. 2016, 58, 199–222. [Google Scholar] [CrossRef]
- Resmi, S.; Thomas, V.; Sreenivas, V.K. Anatomical studies of selected species of Adiantum L. in Kerala. South Ind. J. Biol. Sci. 2016, 2, 115–118. [Google Scholar] [CrossRef]
- Tryon, A.F.; Lugardon, B. Introduction. In Spores of the Pteridophyta; Springer: New York, NY, USA, 1991; pp. 1–26. [Google Scholar] [CrossRef]
- Chatterjee, A.; Khatua, S.; Acharya, K.; Sarkar, J. A green approach for the synthesis of antimicrobial bio-surfactant silver nanoparticles by using a fern. Dig. J. Nanomater. Biostruct. 2019, 14, 479–490. [Google Scholar]
- Kulkarni, V.D.; Kulkarni, P.S. Green Synthesis of Copper Nanoparticles Using Ocimum. Int. J. Chem. Stud. 2013, 1, 1–4. [Google Scholar]
- Padil, V.V.T.; Černík, M. Green Synthesis of Copper Oxide Nanoparticles Using Gum Karaya as a Biotemplate and Their Antibacterial Application. Int. J. Nanomed. 2013, 8, 889–898. [Google Scholar] [CrossRef]
- Cuevas, R.; Durán, N.; Diez, M.C.; Tortella, G.R.; Rubilar, O. Extracellular biosynthesis of copper and copper oxide nanoparticles by Stereumhirsutum, a native white-rot fungus from Chilean forests. J. Nanomater. 2015, 2015, 7. [Google Scholar] [CrossRef]
- Sarkar, J.; Dey, P.; Saha, S.; Acharya, K. Mycosynthesis of Selenium Nanoparticles. Micro Nano Lett. 2011, 6, 599–602. [Google Scholar] [CrossRef]
- Sundaramurthy, N.; Parthiban, C. Biosynthesis of Copper Oxide Nanoparticles Using Pyrus pyrifolia Leaf Extract and Evolve the Catalytic Activity. Int. Res. J. Eng. Technol. 2015, 2, 332–338. [Google Scholar]
- Sarkar, J.; Ray, S.; Chattopadhyay, D.; Laskar, A.; Acharya, K. Mycogenesis of gold nanoparticles using a phytopathogen Alternari aalternata. Bioproc. Biosys. Eng. 2012, 35, 637–643. [Google Scholar] [CrossRef]
- Okaiyeto, K.; Ojemaye, M.O.; Hoppe, H.; Mabinya, L.V.; Okoh, A.I. Phytofabrication of Silver/Silver Chloride Nanoparticles Using Aqueous Leaf Extract of Oederagenistifolia: Characterization and Antibacterial Potential. Molecules 2019, 24, 4382. [Google Scholar] [CrossRef]
- Kumar, P.P.N.V.; Shameem, U.; Kollu, P.; Kalyani, R.L.; Pammi, S.V.N. Green Synthesis of Copper Oxide Nanoparticles Using Aloe vera Leaf Extract and Its Antibacterial Activity Against Fish Bacterial Pathogens. BioNanoScience 2015, 5, 135–139. [Google Scholar] [CrossRef]
- Sanghi, R.; Verma, P.; Puri, S. Enzymatic formation of gold nanoparticles using Phanerochaete chrysosporium. Adv. Chem. Eng. Sci. 2011, 1, 154–162. [Google Scholar] [CrossRef]
- Kannan, R.R.; Stirk, W.A.; Van Staden, J. Synthesis of Silver Nanoparticles Using the Seaweed Codium capitatum P.C. Silva (Chlorophyceae). S. Afr. J. Bot. 2013, 86, 1–4. [Google Scholar] [CrossRef]
- Cai, S.; Singh, B.L. A Distinct Utility of the Amide III Infrared Band for Secondary Structure Estimation of Aqueous Protein Solutions Using Partial Least Squares Methods. Biochemistry 2004, 43, 2541–2549. [Google Scholar] [CrossRef] [PubMed]
- Hosseini, M.R.; Schaffie, M.; Pazouki, M.; Darezereshki, E.; Ranjbar, M. Biologically synthesized copper sulfide nanoparticles: Production and characterization. Mater. Sci. Semicond. Process. 2012, 15, 222–225. [Google Scholar] [CrossRef]
- Karthik, K.; Victor Jaya, N.; Kanagaraj, M.; Arumugam, S. Temperature-Dependent Magnetic Anomalies of CuO Nanoparticles. Solid State Commun. 2011, 151, 564–568. [Google Scholar] [CrossRef]
- Dwivedi, S.; AlKhedhairy, A.A.; Ahamed, M.; Musarrat, J. Biomimetic Synthesis of Selenium Nanospheres by Bacterial Strain JS-11 and Its Role as a Biosensor for Nanotoxicity Assessment: A Novel Se-Bioassay. PLoS ONE 2013, 8, e57404. [Google Scholar] [CrossRef]
- Mollick, M.M.R.; Bhowmick, B.; Mondal, D.; Maity, D.; Rana, D.; Dash, S.K.; Chattopadhyay, S.; Roy, S.; Sarkar, J.; Acharya, K.; et al. Anticancer (in vitro) and antimicrobial effect of gold nanoparticles synthesized using Abelmoschus esculentus (L.) pulp extract via a green route. RSC Adv. 2014, 4, 37838–37848. [Google Scholar] [CrossRef]
- Sarkar, J.; Mollick, M.M.R.; Chattopadhyay, D.; Acharya, K. An eco-friendly route of c-Fe2O3 nanoparticles formation and investigation of the mechanical properties of the HPMC-γ-Fe2O3 nanocomposites. Bioprocess Biosyst. Eng. 2017, 40, 351–359. [Google Scholar] [CrossRef]
- Nair, P.M.G.; Chung, I.M. Impact of copper oxide nanoparticles exposure on Arabidopsis thaliana growth, root system development, root lignification, and molecular level changes. Environ. Sci. Pol. Res. 2014, 21, 12709–12722. [Google Scholar] [CrossRef] [PubMed]
- Lamb, C.; Dixon, R.A. The oxidative burst in plant disease resistance. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1997, 48, 251–275. [Google Scholar] [CrossRef] [PubMed]
- Acharya, K.; Chakraborty, N.; Dutta, A.K.; Sarkar, S.; Acharya, R. Signaling role of nitric oxide in the induction of plant defense by exogenous application of abiotic inducers. Arch. Phytopathol. Plant Prot. 2011, 44, 1501–1511. [Google Scholar] [CrossRef]
- Chandra, S.; Chakraborty, N.; Chakraborty, A.; Rai, R.; Bera, B.; Acharya, K. Abiotic elicitor mediated improvement of innate immunity in Camellia sinensis. J. Plant Growth Regul. 2014, 33, 849–859. [Google Scholar] [CrossRef]
- Chakraborty, N.; Chandra, S.; Acharya, K. Biochemical basis of improvement of defense in tomato plant against Fusarium wilt by CaCl2. Physiol. Mol. Biol. Plants 2017, 23, 581–596. [Google Scholar] [CrossRef]
- Pellegrini, L.; Rohfritsch, O.; Fritig, B.; Legrand, M. Phenylalanine ammonia-lyase in tobacco. Molecular cloning and gene expression during the hypersensitive reaction to tobacco mosaic virus and the response to a fungal elicitor. Plant Physiol. 1994, 106, 877–886. [Google Scholar] [CrossRef]
- Whan, J.A.; Dann, E.K.; Smith, L.J.; Aitken, E.A.B. Acibenzolar-S-methyl-induced alteration of defence gene expression and enzyme activity in cotton infected with Fusarium oxysporum f. sp. vasinfectum. Physiol. Mol. Plant Pathol. 2008, 73, 175–182. [Google Scholar] [CrossRef]
- Najami, N.; Tibor, J.; Barriah, W.; Kayam, G.; Moshe, T.; Guy, M.; Volokita, M. Ascorbate peroxidase gene family in tomato: Its identification and characterization. Mol. Genet. Genom. 2008, 279, 171–182. [Google Scholar] [CrossRef]
- Shaw, A.K.; Hossain, Z. Impact of nano-CuO stress on rice (Oryza sativa L.) seedlings. Chemosphere 2013, 93, 906–991. [Google Scholar] [CrossRef]
- Trujillo-Reyes, J.; Majumdar, S.; Botez, C.E.; Peralta-Videa, J.R.; Gardea-Torresdey, J.L. Exposure studies of core–shell Fe/Fe3O4 and Cu/CuO NPs to lettuce (Lactuca sativa) plants: Are they a potential physiological and nutritional hazard? J. Hazard. Mate. 2014, 267, 255–263. [Google Scholar] [CrossRef] [PubMed]
- Biswas, S.K.; Pandey, N.K.; Rajik, M. Inductions of defense response in tomato against fusarium wilt through inorganic chemicals as inducers. J. Plant Pathol. Microbiol. 2012, 3, 1–7. [Google Scholar]
- Lin, C.; Kao, C. Effect of NaCl stress on H2O2 metabolism in rice leaves. Plant Growth Regul. 2000, 30, 151–155. [Google Scholar] [CrossRef]
- Hare, P.; Cress, W. Metabolic implications of stress- induced proline accumulation in plants. Plant Growth Regul. 1997, 21, 79–102. [Google Scholar] [CrossRef]
- Slama, I.; Ghnaya, T.; Hessini, K.; Messedi, D.; Savouré, A.; Abdelly, C. Comparative study of the effects of mannitol and PEG osmotic stress on growth and solute accumulation in Sesuvium portulacastrum. Environ. Exp. Bot. 2007, 61, 10–17. [Google Scholar] [CrossRef]
- Corpas, F.J.; Leterrier, M.; Valderrama, R.; Airaki, M.; Chaki, M.; Palma, J.M.; Barroso, J.B. Nitric oxide imbalance provokes a nitrosative response in plants under abiotic stress. Plant Sci. 2011, 181, 604–611. [Google Scholar] [CrossRef]
- Leterrier, M.; Valderrama, R.; Chaki, M.; Airaki, M.; Palma, J.M.; Barroso, J.B.; Corpas, F.J. Function of nitric oxide under environmental stress conditions. In Phytohormones and Abiotic Stress Tolerance in Plants, 1st ed.; Khan, N.A., Nazar, R., Iqbal, N., Anjum, N.A., Eds.; Springer: Berlin/Heidelberg, Germany, 2012; pp. 99–113. [Google Scholar] [CrossRef]
- Chakraborty, N.; Chandra, S.; Acharya, K. Sublethal Heavy Metal Stress Stimulates Innate Immunity in Tomato. Sci. World J. 2015, 2015, 208649. [Google Scholar] [CrossRef]
- Chakraborty, N.; Mukherjee, K.; Sarkar, A.; Acharya, K. Interaction between Bean and Colletotrichum gloeosporioides: Understanding Through a Biochemical Approach. Plants 2019, 8, 345. [Google Scholar] [CrossRef]
- Laspina, N.V.; Groppa, M.D.; Tomaro, M.L.; Benavides, M.P. Nitric oxide protects sunflower leaves against Cd-induced oxidative stress. Plant Sci. 2005, 169, 323–330. [Google Scholar] [CrossRef]
- Hasanuzzaman, M.; Nahar, K.; Alam, M.M.; Fujita, M. Exogenous nitric oxide alleviates high temperature induced oxidative stress in wheat (Triticum aestivum L.) seedlings by modulating the antioxidant defense and glyoxalase system. Aust. J. Crop Sci. 2012, 6, 1314–1323. [Google Scholar]
- Chakraborty, N.; Ghosh, S.; Chandra, S.; Sengupta, S.; Acharya, K. Abiotic elicitors mediated elicitation of innate immunity in tomato: An ex vivo comparison. Physiol. Mol. Biol. Plants. 2016, 22, 307–320. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, N.; Acharya, K. “NO way”! Says the plant to abiotic stress. Plant Gene 2017, 11, 99–105. [Google Scholar] [CrossRef]
- Repka, V. Elicitor-stimulated induction of defense mechanisms and defense gene activation in grapevine cell suspension cultures. Biol. Plant. 2001, 44, 555–565. [Google Scholar] [CrossRef]




| Sets | % Seed Germination | Seedling Vigor Index | RWC (%) |
|---|---|---|---|
| Control | 80.77 ± 1.82 b | 3661.04 ± 46.41 c | 87.39 ± 4.37 a |
| CuONP-0.01 mgmL−1 | 93.96 ± 1.75 a | 4269.4 ± 71.26 a | 88.38 ± 3.81 a |
| CuONP-0.025 mgmL−1 | 91.26 ± 1.31 a | 4168.43 ± 48.15 b | 84.37 ± 5.33 b |
| CuONP-0.05 mgmL−1 | 75.98 ± 2.41 c | 2541.36 ± 61.31 d | 81.28 ± 3.71 c |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sarkar, J.; Chakraborty, N.; Chatterjee, A.; Bhattacharjee, A.; Dasgupta, D.; Acharya, K. Green Synthesized Copper Oxide Nanoparticles Ameliorate Defence and Antioxidant Enzymes in Lens culinaris. Nanomaterials 2020, 10, 312. https://doi.org/10.3390/nano10020312
Sarkar J, Chakraborty N, Chatterjee A, Bhattacharjee A, Dasgupta D, Acharya K. Green Synthesized Copper Oxide Nanoparticles Ameliorate Defence and Antioxidant Enzymes in Lens culinaris. Nanomaterials. 2020; 10(2):312. https://doi.org/10.3390/nano10020312
Chicago/Turabian StyleSarkar, Joy, Nilanjan Chakraborty, Arindam Chatterjee, Avisek Bhattacharjee, Disha Dasgupta, and Krishnendu Acharya. 2020. "Green Synthesized Copper Oxide Nanoparticles Ameliorate Defence and Antioxidant Enzymes in Lens culinaris" Nanomaterials 10, no. 2: 312. https://doi.org/10.3390/nano10020312
APA StyleSarkar, J., Chakraborty, N., Chatterjee, A., Bhattacharjee, A., Dasgupta, D., & Acharya, K. (2020). Green Synthesized Copper Oxide Nanoparticles Ameliorate Defence and Antioxidant Enzymes in Lens culinaris. Nanomaterials, 10(2), 312. https://doi.org/10.3390/nano10020312





