SARS-CoV-2: An Update on Potential Antivirals in Light of SARS-CoV Antiviral Drug Discoveries
Abstract
:1. Introduction
2. SARS-CoV
3. SARS-CoV Antivirals
3.1. Entry Inhibitors (Antibodies, Recombinant Proteins, Peptides, and Small Molecule Antivirals)
3.1.1. Antibodies
3.1.2. S Protein-Binding Molecules
A. Lectins
B. Emodin
3.1.3. Fusion Inhibitors
3.1.4. ACE2 Inhibitors
3.1.5. ACE2-Derived Peptides and Soluble Recombinant ACE2
3.1.6. Inhibitors of Endosomal Acidification, Endocytosis, Cathepsin L, and TMPRSS2
3.2. Protease Inhibitors
3.2.1. 3CLpro Inhibitors
3.2.2. PLpro Inhibitors
3.2.3. Cellular Protease Inhibitors
3.3. Viral Helicase Inhibitors
3.4. Replication and Transcription Inhibitors
3.5. Miscellaneous Agents
4. SARS-CoV-2
5. SARS-CoV-2 Antivirals
5.1. Entry Inhibitors (Antibodies and Other Antivirals)
5.1.1. Antibodies
5.1.2. Soluble Recombinant ACE2
5.1.3. Inhibitors of Endosomal Acidification, Cathepsin L, and TMPRSS2
5.1.4. Fusion Inhibitors
5.2. Replication and Transcription Inhibitors
5.3. Protease Inhibitors
5.4. Miscellaneous Agents
6. Conclusions
Funding
Conflicts of Interest
References
- Woo, P.C.; Huang, Y.; Lau, S.K.; Yuen, K.Y. Coronavirus genomics and bioinformatics analysis. Viruses 2010, 2, 1804–1820. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Woo, P.C.; Lau, S.K.; Lam, C.S.; Lau, C.C.; Tsang, A.K.; Lau, J.H.; Bai, R.; Teng, J.L.; Tsang, C.C.; Wang, M.; et al. Discovery of seven novel Mammalian and avian coronaviruses in the genus deltacoronavirus supports bat coronaviruses as the gene source of alphacoronavirus and betacoronavirus and avian coronaviruses as the gene source of gammacoronavirus and deltacoronavirus. J. Virol. 2012, 86, 3995–4008. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lim, Y.X.; Ng, Y.L.; Tam, J.P.; Liu, D.X. Human Coronaviruses: A Review of Virus-Host Interactions. Diseases 2016, 4, 26. [Google Scholar] [CrossRef]
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Inglesby, T.V. Public Health Measures and the Reproduction Number of SARS-CoV-2. JAMA 2020. [Google Scholar] [CrossRef]
- Li, W.; Shi, Z.; Yu, M.; Ren, W.; Smith, C.; Epstein, J.H.; Wang, H.; Crameri, G.; Hu, Z.; Zhang, H.; et al. Bats are natural reservoirs of SARS-like coronaviruses. Science 2005, 310, 676–679. [Google Scholar] [CrossRef]
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N.; et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2020, 395, 565–574. [Google Scholar] [CrossRef] [Green Version]
- Drosten, C.; Gunther, S.; Preiser, W.; van der Werf, S.; Brodt, H.R.; Becker, S.; Rabenau, H.; Panning, M.; Kolesnikova, L.; Fouchier, R.A.; et al. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N. Engl. J. Med. 2003, 348, 1967–1976. [Google Scholar] [CrossRef]
- Rota, P.A.; Oberste, M.S.; Monroe, S.S.; Nix, W.A.; Campagnoli, R.; Icenogle, J.P.; Penaranda, S.; Bankamp, B.; Maher, K.; Chen, M.H.; et al. Characterization of a novel coronavirus associated with severe acute respiratory syndrome. Science 2003, 300, 1394–1399. [Google Scholar] [CrossRef] [Green Version]
- Marra, M.A.; Jones, S.J.; Astell, C.R.; Holt, R.A.; Brooks-Wilson, A.; Butterfield, Y.S.; Khattra, J.; Asano, J.K.; Barber, S.A.; Chan, S.Y.; et al. The Genome sequence of the SARS-associated coronavirus. Science 2003, 300, 1399–1404. [Google Scholar] [CrossRef] [Green Version]
- Siu, Y.L.; Teoh, K.T.; Lo, J.; Chan, C.M.; Kien, F.; Escriou, N.; Tsao, S.W.; Nicholls, J.M.; Altmeyer, R.; Peiris, J.S.; et al. The M, E, and N structural proteins of the severe acute respiratory syndrome coronavirus are required for efficient assembly, trafficking, and release of virus-like particles. J. Virol. 2008, 82, 11318–11330. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xiao, X.; Chakraborti, S.; Dimitrov, A.S.; Gramatikoff, K.; Dimitrov, D.S. The SARS-CoV S glycoprotein: Expression and functional characterization. Biochem. Biophys. Res. Commun. 2003, 312, 1159–1164. [Google Scholar] [CrossRef]
- Li, W.; Moore, M.J.; Vasilieva, N.; Sui, J.; Wong, S.K.; Berne, M.A.; Somasundaran, M.; Sullivan, J.L.; Luzuriaga, K.; Greenough, T.C.; et al. Angiotensin-converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature 2003, 426, 450–454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wong, S.K.; Li, W.; Moore, M.J.; Choe, H.; Farzan, M. A 193-amino acid fragment of the SARS coronavirus S protein efficiently binds angiotensin-converting enzyme 2. J. Biol. Chem. 2004, 279, 3197–3201. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Simmons, G.; Zmora, P.; Gierer, S.; Heurich, A.; Pohlmann, S. Proteolytic activation of the SARS-coronavirus spike protein: Cutting enzymes at the cutting edge of antiviral research. Antivir. Res. 2013, 100, 605–614. [Google Scholar] [CrossRef]
- Bosch, B.J.; Bartelink, W.; Rottier, P.J. Cathepsin L functionally cleaves the severe acute respiratory syndrome coronavirus class I fusion protein upstream of rather than adjacent to the fusion peptide. J. Virol. 2008, 82, 8887–8890. [Google Scholar] [CrossRef] [Green Version]
- Huang, I.C.; Bosch, B.J.; Li, F.; Li, W.; Lee, K.H.; Ghiran, S.; Vasilieva, N.; Dermody, T.S.; Harrison, S.C.; Dormitzer, P.R.; et al. SARS coronavirus, but not human coronavirus NL63, utilizes cathepsin L to infect ACE2-expressing cells. J. Biol. Chem. 2006, 281, 3198–3203. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Y.; Vedantham, P.; Lu, K.; Agudelo, J.; Carrion, R., Jr.; Nunneley, J.W.; Barnard, D.; Pohlmann, S.; McKerrow, J.H.; Renslo, A.R.; et al. Protease inhibitors targeting coronavirus and filovirus entry. Antivir. Res. 2015, 116, 76–84. [Google Scholar] [CrossRef]
- Matsuyama, S.; Nagata, N.; Shirato, K.; Kawase, M.; Takeda, M.; Taguchi, F. Efficient activation of the severe acute respiratory syndrome coronavirus spike protein by the transmembrane protease TMPRSS2. J. Virol. 2010, 84, 12658–12664. [Google Scholar] [CrossRef] [Green Version]
- Masters, P.S. The molecular biology of coronaviruses. Adv. Virus Res. 2006, 66, 193–292. [Google Scholar] [CrossRef]
- Brian, D.A.; Baric, R.S. Coronavirus genome structure and replication. Curr. Top. Microbiol. Immunol. 2005, 287, 1–30. [Google Scholar] [CrossRef] [Green Version]
- Li, G.; Chen, X.; Xu, A. Profile of specific antibodies to the SARS-associated coronavirus. N. Engl. J. Med. 2003, 349, 508–509. [Google Scholar] [CrossRef]
- Subbarao, K.; McAuliffe, J.; Vogel, L.; Fahle, G.; Fischer, S.; Tatti, K.; Packard, M.; Shieh, W.J.; Zaki, S.; Murphy, B. Prior infection and passive transfer of neutralizing antibody prevent replication of severe acute respiratory syndrome coronavirus in the respiratory tract of mice. J. Virol. 2004, 78, 3572–3577. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Keng, C.T.; Zhang, A.; Shen, S.; Lip, K.M.; Fielding, B.C.; Tan, T.H.; Chou, C.F.; Loh, C.B.; Wang, S.; Fu, J.; et al. Amino acids 1055 to 1192 in the S2 region of severe acute respiratory syndrome coronavirus S protein induce neutralizing antibodies: Implications for the development of vaccines and antiviral agents. J. Virol. 2005, 79, 3289–3296. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, H.; Wang, G.; Li, J.; Nie, Y.; Shi, X.; Lian, G.; Wang, W.; Yin, X.; Zhao, Y.; Qu, X.; et al. Identification of an antigenic determinant on the S2 domain of the severe acute respiratory syndrome coronavirus spike glycoprotein capable of inducing neutralizing antibodies. J. Virol. 2004, 78, 6938–6945. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- He, Y.; Zhou, Y.; Liu, S.; Kou, Z.; Li, W.; Farzan, M.; Jiang, S. Receptor-binding domain of SARS-CoV spike protein induces highly potent neutralizing antibodies: Implication for developing subunit vaccine. Biochem. Biophys. Res. Commun. 2004, 324, 773–781. [Google Scholar] [CrossRef]
- Bisht, H.; Roberts, A.; Vogel, L.; Bukreyev, A.; Collins, P.L.; Murphy, B.R.; Subbarao, K.; Moss, B. Severe acute respiratory syndrome coronavirus spike protein expressed by attenuated vaccinia virus protectively immunizes mice. Proc. Natl. Acad. Sci. USA 2004, 101, 6641–6646. [Google Scholar] [CrossRef] [Green Version]
- He, Y.; Lu, H.; Siddiqui, P.; Zhou, Y.; Jiang, S. Receptor-binding domain of severe acute respiratory syndrome coronavirus spike protein contains multiple conformation-dependent epitopes that induce highly potent neutralizing antibodies. J. Immunol. 2005, 174, 4908–4915. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Yang, F.; Li, Y.H.; Li, W.H.; Tu, X.M.; Wei, Q.; Zhu, H.; Liu, L.; Wang, H.; Qin, C.; et al. Potent neutralization antibody elicited in mice by SARS-associated coronavirus spike protein S1 domain. Zhonghua Shi Yan He Lin Chuang Bing Du Xue Za Zhi 2004, 18, 258–260. [Google Scholar]
- Hofmann, H.; Hattermann, K.; Marzi, A.; Gramberg, T.; Geier, M.; Krumbiegel, M.; Kuate, S.; Uberla, K.; Niedrig, M.; Pohlmann, S. S protein of severe acute respiratory syndrome-associated coronavirus mediates entry into hepatoma cell lines and is targeted by neutralizing antibodies in infected patients. J. Virol. 2004, 78, 6134–6142. [Google Scholar] [CrossRef] [Green Version]
- Cao, Z.; Liu, L.; Du, L.; Zhang, C.; Jiang, S.; Li, T.; He, Y. Potent and persistent antibody responses against the receptor-binding domain of SARS-CoV spike protein in recovered patients. Virol. J. 2010, 7, 299. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cheng, Y.; Wong, R.; Soo, Y.O.; Wong, W.S.; Lee, C.K.; Ng, M.H.; Chan, P.; Wong, K.C.; Leung, C.B.; Cheng, G. Use of convalescent plasma therapy in SARS patients in Hong Kong. Eur. J. Clin. Microbiol. Infect. Dis. 2005, 24, 44–46. [Google Scholar] [CrossRef] [PubMed]
- Soo, Y.O.; Cheng, Y.; Wong, R.; Hui, D.S.; Lee, C.K.; Tsang, K.K.; Ng, M.H.; Chan, P.; Cheng, G.; Sung, J.J. Retrospective comparison of convalescent plasma with continuing high-dose methylprednisolone treatment in SARS patients. Clin. Microbiol. Infect. 2004, 10, 676–678. [Google Scholar] [CrossRef] [Green Version]
- Sui, J.; Li, W.; Murakami, A.; Tamin, A.; Matthews, L.J.; Wong, S.K.; Moore, M.J.; Tallarico, A.S.; Olurinde, M.; Choe, H.; et al. Potent neutralization of severe acute respiratory syndrome (SARS) coronavirus by a human mAb to S1 protein that blocks receptor association. Proc. Natl. Acad. Sci. USA 2004, 101, 2536–2541. [Google Scholar] [CrossRef] [Green Version]
- ter Meulen, J.; Bakker, A.B.; van den Brink, E.N.; Weverling, G.J.; Martina, B.E.; Haagmans, B.L.; Kuiken, T.; de Kruif, J.; Preiser, W.; Spaan, W.; et al. Human monoclonal antibody as prophylaxis for SARS coronavirus infection in ferrets. Lancet 2004, 363, 2139–2141. [Google Scholar] [CrossRef] [Green Version]
- Berry, J.D.; Jones, S.; Drebot, M.A.; Andonov, A.; Sabara, M.; Yuan, X.Y.; Weingartl, H.; Fernando, L.; Marszal, P.; Gren, J.; et al. Development and characterisation of neutralising monoclonal antibody to the SARS-coronavirus. J. Virol. Methods 2004, 120, 87–96. [Google Scholar] [CrossRef] [PubMed]
- Gubbins, M.J.; Plummer, F.A.; Yuan, X.Y.; Johnstone, D.; Drebot, M.; Andonova, M.; Andonov, A.; Berry, J.D. Molecular characterization of a panel of murine monoclonal antibodies specific for the SARS-coronavirus. Mol. Immunol. 2005, 42, 125–136. [Google Scholar] [CrossRef]
- van den Brink, E.N.; Ter Meulen, J.; Cox, F.; Jongeneelen, M.A.; Thijsse, A.; Throsby, M.; Marissen, W.E.; Rood, P.M.; Bakker, A.B.; Gelderblom, H.R.; et al. Molecular and biological characterization of human monoclonal antibodies binding to the spike and nucleocapsid proteins of severe acute respiratory syndrome coronavirus. J. Virol. 2005, 79, 1635–1644. [Google Scholar] [CrossRef] [Green Version]
- Greenough, T.C.; Babcock, G.J.; Roberts, A.; Hernandez, H.J.; Thomas, W.D., Jr.; Coccia, J.A.; Graziano, R.F.; Srinivasan, M.; Lowy, I.; Finberg, R.W.; et al. Development and characterization of a severe acute respiratory syndrome-associated coronavirus-neutralizing human monoclonal antibody that provides effective immunoprophylaxis in mice. J. Infect. Dis. 2005, 191, 507–514. [Google Scholar] [CrossRef]
- He, Y.; Zhu, Q.; Liu, S.; Zhou, Y.; Yang, B.; Li, J.; Jiang, S. Identification of a critical neutralization determinant of severe acute respiratory syndrome (SARS)-associated coronavirus: Importance for designing SARS vaccines. Virology 2005, 334, 74–82. [Google Scholar] [CrossRef] [Green Version]
- Sui, J.; Li, W.; Roberts, A.; Matthews, L.J.; Murakami, A.; Vogel, L.; Wong, S.K.; Subbarao, K.; Farzan, M.; Marasco, W.A. Evaluation of human monoclonal antibody 80R for immunoprophylaxis of severe acute respiratory syndrome by an animal study, epitope mapping, and analysis of spike variants. J. Virol. 2005, 79, 5900–5906. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tripp, R.A.; Haynes, L.M.; Moore, D.; Anderson, B.; Tamin, A.; Harcourt, B.H.; Jones, L.P.; Yilla, M.; Babcock, G.J.; Greenough, T.; et al. Monoclonal antibodies to SARS-associated coronavirus (SARS-CoV): Identification of neutralizing and antibodies reactive to S, N, M and E viral proteins. J. Virol. Methods 2005, 128, 21–28. [Google Scholar] [CrossRef] [PubMed]
- Duan, J.; Yan, X.; Guo, X.; Cao, W.; Han, W.; Qi, C.; Feng, J.; Yang, D.; Gao, G.; Jin, G. A human SARS-CoV neutralizing antibody against epitope on S2 protein. Biochem. Biophys. Res. Commun. 2005, 333, 186–193. [Google Scholar] [CrossRef] [PubMed]
- Lip, K.M.; Shen, S.; Yang, X.; Keng, C.T.; Zhang, A.; Oh, H.L.; Li, Z.H.; Hwang, L.A.; Chou, C.F.; Fielding, B.C.; et al. Monoclonal antibodies targeting the HR2 domain and the region immediately upstream of the HR2 of the S protein neutralize in vitro infection of severe acute respiratory syndrome coronavirus. J. Virol. 2006, 80, 941–950. [Google Scholar] [CrossRef] [Green Version]
- Roberts, A.; Thomas, W.D.; Guarner, J.; Lamirande, E.W.; Babcock, G.J.; Greenough, T.C.; Vogel, L.; Hayes, N.; Sullivan, J.L.; Zaki, S.; et al. Therapy with a severe acute respiratory syndrome-associated coronavirus-neutralizing human monoclonal antibody reduces disease severity and viral burden in golden Syrian hamsters. J. Infect. Dis. 2006, 193, 685–692. [Google Scholar] [CrossRef]
- Coughlin, M.; Lou, G.; Martinez, O.; Masterman, S.K.; Olsen, O.A.; Moksa, A.A.; Farzan, M.; Babcook, J.S.; Prabhakar, B.S. Generation and characterization of human monoclonal neutralizing antibodies with distinct binding and sequence features against SARS coronavirus using XenoMouse. Virology 2007, 361, 93–102. [Google Scholar] [CrossRef] [Green Version]
- Coughlin, M.M.; Babcook, J.; Prabhakar, B.S. Human monoclonal antibodies to SARS-coronavirus inhibit infection by different mechanisms. Virology 2009, 394, 39–46. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Elshabrawy, H.A.; Coughlin, M.M.; Baker, S.C.; Prabhakar, B.S. Human monoclonal antibodies against highly conserved HR1 and HR2 domains of the SARS-CoV spike protein are more broadly neutralizing. PLoS ONE 2012, 7, e50366. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sharon, N.; Lis, H. History of lectins: From hemagglutinins to biological recognition molecules. Glycobiology 2004, 14, 53R–62R. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mitchell, C.A.; Ramessar, K.; O’Keefe, B.R. Antiviral lectins: Selective inhibitors of viral entry. Antivir. Res. 2017, 142, 37–54. [Google Scholar] [CrossRef]
- O’Keefe, B.R.; Giomarelli, B.; Barnard, D.L.; Shenoy, S.R.; Chan, P.K.; McMahon, J.B.; Palmer, K.E.; Barnett, B.W.; Meyerholz, D.K.; Wohlford-Lenane, C.L.; et al. Broad-spectrum in vitro activity and in vivo efficacy of the antiviral protein griffithsin against emerging viruses of the family Coronaviridae. J. Virol. 2010, 84, 2511–2521. [Google Scholar] [CrossRef] [Green Version]
- Kumaki, Y.; Wandersee, M.K.; Smith, A.J.; Zhou, Y.; Simmons, G.; Nelson, N.M.; Bailey, K.W.; Vest, Z.G.; Li, J.K.; Chan, P.K.; et al. Inhibition of severe acute respiratory syndrome coronavirus replication in a lethal SARS-CoV BALB/c mouse model by stinging nettle lectin, Urtica dioica agglutinin. Antivir. Res. 2011, 90, 22–32. [Google Scholar] [CrossRef] [PubMed]
- Ho, T.Y.; Wu, S.L.; Chen, J.C.; Li, C.C.; Hsiang, C.Y. Emodin blocks the SARS coronavirus spike protein and angiotensin-converting enzyme 2 interaction. Antivir. Res. 2007, 74, 92–101. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, S.; Wang, K.; Yu, W.; Sun, B.; Schwarz, W. Emodin inhibits current through SARS-associated coronavirus 3a protein. Antivir. Res. 2011, 90, 64–69. [Google Scholar] [CrossRef]
- Liu, S.; Xiao, G.; Chen, Y.; He, Y.; Niu, J.; Escalante, C.R.; Xiong, H.; Farmar, J.; Debnath, A.K.; Tien, P.; et al. Interaction between heptad repeat 1 and 2 regions in spike protein of SARS-associated coronavirus: Implications for virus fusogenic mechanism and identification of fusion inhibitors. Lancet 2004, 363, 938–947. [Google Scholar] [CrossRef] [Green Version]
- Sainz, B., Jr.; Mossel, E.C.; Gallaher, W.R.; Wimley, W.C.; Peters, C.J.; Wilson, R.B.; Garry, R.F. Inhibition of severe acute respiratory syndrome-associated coronavirus (SARS-CoV) infectivity by peptides analogous to the viral spike protein. Virus Res. 2006, 120, 146–155. [Google Scholar] [CrossRef]
- Coleman, C.M.; Sisk, J.M.; Mingo, R.M.; Nelson, E.A.; White, J.M.; Frieman, M.B. Abelson Kinase Inhibitors Are Potent Inhibitors of Severe Acute Respiratory Syndrome Coronavirus and Middle East Respiratory Syndrome Coronavirus Fusion. J. Virol. 2016, 90, 8924–8933. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yi, L.; Li, Z.; Yuan, K.; Qu, X.; Chen, J.; Wang, G.; Zhang, H.; Luo, H.; Zhu, L.; Jiang, P.; et al. Small molecules blocking the entry of severe acute respiratory syndrome coronavirus into host cells. J. Virol. 2004, 78, 11334–11339. [Google Scholar] [CrossRef] [Green Version]
- Huentelman, M.J.; Zubcevic, J.; Hernandez Prada, J.A.; Xiao, X.; Dimitrov, D.S.; Raizada, M.K.; Ostrov, D.A. Structure-based discovery of a novel angiotensin-converting enzyme 2 inhibitor. Hypertension 2004, 44, 903–906. [Google Scholar] [CrossRef] [Green Version]
- Han, D.P.; Penn-Nicholson, A.; Cho, M.W. Identification of critical determinants on ACE2 for SARS-CoV entry and development of a potent entry inhibitor. Virology 2006, 350, 15–25. [Google Scholar] [CrossRef] [Green Version]
- Hofmann, H.; Geier, M.; Marzi, A.; Krumbiegel, M.; Peipp, M.; Fey, G.H.; Gramberg, T.; Pohlmann, S. Susceptibility to SARS coronavirus S protein-driven infection correlates with expression of angiotensin converting enzyme 2 and infection can be blocked by soluble receptor. Biochem. Biophys. Res. Commun. 2004, 319, 1216–1221. [Google Scholar] [CrossRef] [PubMed]
- Al-Bari, M.A. Chloroquine analogues in drug discovery: New directions of uses, mechanisms of actions and toxic manifestations from malaria to multifarious diseases. J. Antimicrob. Chemother. 2015, 70, 1608–1621. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vincent, M.J.; Bergeron, E.; Benjannet, S.; Erickson, B.R.; Rollin, P.E.; Ksiazek, T.G.; Seidah, N.G.; Nichol, S.T. Chloroquine is a potent inhibitor of SARS coronavirus infection and spread. Virol. J. 2005, 2, 69. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- de Wilde, A.H.; Jochmans, D.; Posthuma, C.C.; Zevenhoven-Dobbe, J.C.; van Nieuwkoop, S.; Bestebroer, T.M.; van den Hoogen, B.G.; Neyts, J.; Snijder, E.J. Screening of an FDA-approved compound library identifies four small-molecule inhibitors of Middle East respiratory syndrome coronavirus replication in cell culture. Antimicrob. Agents Chemother. 2014, 58, 4875–4884. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Keyaerts, E.; Vijgen, L.; Maes, P.; Neyts, J.; Van Ranst, M. In vitro inhibition of severe acute respiratory syndrome coronavirus by chloroquine. Biochem. Biophys. Res. Commun. 2004, 323, 264–268. [Google Scholar] [CrossRef] [PubMed]
- Elshabrawy, H.A.; Fan, J.; Haddad, C.S.; Ratia, K.; Broder, C.C.; Caffrey, M.; Prabhakar, B.S. Identification of a broad-spectrum antiviral small molecule against severe acute respiratory syndrome coronavirus and Ebola, Hendra, and Nipah viruses by using a novel high-throughput screening assay. J. Virol. 2014, 88, 4353–4365. [Google Scholar] [CrossRef] [Green Version]
- Adedeji, A.O.; Severson, W.; Jonsson, C.; Singh, K.; Weiss, S.R.; Sarafianos, S.G. Novel inhibitors of severe acute respiratory syndrome coronavirus entry that act by three distinct mechanisms. J. Virol. 2013, 87, 8017–8028. [Google Scholar] [CrossRef] [Green Version]
- Shah, P.P.; Wang, T.; Kaletsky, R.L.; Myers, M.C.; Purvis, J.E.; Jing, H.; Huryn, D.M.; Greenbaum, D.C.; Smith, A.B., 3rd; Bates, P.; et al. A small-molecule oxocarbazate inhibitor of human cathepsin L blocks severe acute respiratory syndrome and ebola pseudotype virus infection into human embryonic kidney 293T cells. Mol. Pharmacol. 2010, 78, 319–324. [Google Scholar] [CrossRef]
- Kawase, M.; Shirato, K.; van der Hoek, L.; Taguchi, F.; Matsuyama, S. Simultaneous treatment of human bronchial epithelial cells with serine and cysteine protease inhibitors prevents severe acute respiratory syndrome coronavirus entry. J. Virol. 2012, 86, 6537–6545. [Google Scholar] [CrossRef] [Green Version]
- Zhao, H.; Zhou, J.; Zhang, K.; Chu, H.; Liu, D.; Poon, V.K.; Chan, C.C.; Leung, H.C.; Fai, N.; Lin, Y.P.; et al. A novel peptide with potent and broad-spectrum antiviral activities against multiple respiratory viruses. Sci. Rep. 2016, 6, 22008. [Google Scholar] [CrossRef]
- Chen, X.P.; Li, G.H.; Tang, X.P.; Xiong, Y.; Chen, X.J.; Cao, Y. Lack of severe acute respiratory syndrome in 19 AIDS patients hospitalized together. J. Acquir. Immune. Defic. Syndr. 2003, 34, 242–243. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.P.; Cao, Y. Consideration of highly active antiretroviral therapy in the prevention and treatment of severe acute respiratory syndrome. Clin. Infect. Dis. 2004, 38, 1030–1032. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chu, C.M.; Cheng, V.C.; Hung, I.F.; Wong, M.M.; Chan, K.H.; Chan, K.S.; Kao, R.Y.; Poon, L.L.; Wong, C.L.; Guan, Y.; et al. Role of lopinavir/ritonavir in the treatment of SARS: Initial virological and clinical findings. Thorax 2004, 59, 252–256. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chan, K.S.; Lai, S.T.; Chu, C.M.; Tsui, E.; Tam, C.Y.; Wong, M.M.; Tse, M.W.; Que, T.L.; Peiris, J.S.; Sung, J.; et al. Treatment of severe acute respiratory syndrome with lopinavir/ritonavir: A multicentre retrospective matched cohort study. Hong Kong Med. J. 2003, 9, 399–406. [Google Scholar] [PubMed]
- Yamamoto, N.; Yang, R.; Yoshinaka, Y.; Amari, S.; Nakano, T.; Cinatl, J.; Rabenau, H.; Doerr, H.W.; Hunsmann, G.; Otaka, A.; et al. HIV protease inhibitor nelfinavir inhibits replication of SARS-associated coronavirus. Biochem. Biophys. Res. Commun. 2004, 318, 719–725. [Google Scholar] [CrossRef]
- Zhang, X.W.; Yap, Y.L. Old drugs as lead compounds for a new disease? Binding analysis of SARS coronavirus main proteinase with HIV, psychotic and parasite drugs. Bioorg. Med. Chem. 2004, 12, 2517–2521. [Google Scholar] [CrossRef]
- Ghibelli, L.; Mengoni, F.; Lichtner, M.; Coppola, S.; De Nicola, M.; Bergamaschi, A.; Mastroianni, C.; Vullo, V. Anti-apoptotic effect of HIV protease inhibitors via direct inhibition of calpain. Biochem. Pharmacol. 2003, 66, 1505–1512. [Google Scholar] [CrossRef]
- Andre, P.; Groettrup, M.; Klenerman, P.; de Giuli, R.; Booth, B.L., Jr.; Cerundolo, V.; Bonneville, M.; Jotereau, F.; Zinkernagel, R.M.; Lotteau, V. An inhibitor of HIV-1 protease modulates proteasome activity, antigen presentation, and T cell responses. Proc. Natl. Acad. Sci. USA 1998, 95, 13120–13124. [Google Scholar] [CrossRef] [Green Version]
- Equils, O.; Shapiro, A.; Madak, Z.; Liu, C.; Lu, D. Human immunodeficiency virus type 1 protease inhibitors block toll-like receptor 2 (TLR2)- and TLR4-Induced NF-kappaB activation. Antimicrob. Agents Chemother. 2004, 48, 3905–3911. [Google Scholar] [CrossRef] [Green Version]
- Piccinini, M.; Rinaudo, M.T.; Chiapello, N.; Ricotti, E.; Baldovino, S.; Mostert, M.; Tovo, P.A. The human 26S proteasome is a target of antiretroviral agents. AIDS 2002, 16, 693–700. [Google Scholar] [CrossRef] [Green Version]
- Chen, L.; Gui, C.; Luo, X.; Yang, Q.; Gunther, S.; Scandella, E.; Drosten, C.; Bai, D.; He, X.; Ludewig, B.; et al. Cinanserin is an inhibitor of the 3C-like proteinase of severe acute respiratory syndrome coronavirus and strongly reduces virus replication in vitro. J. Virol. 2005, 79, 7095–7103. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hamill, P.; Hudson, D.; Kao, R.Y.; Chow, P.; Raj, M.; Xu, H.; Richer, M.J.; Jean, F. Development of a red-shifted fluorescence-based assay for SARS-coronavirus 3CL protease: Identification of a novel class of anti-SARS agents from the tropical marine sponge Axinella corrugata. Biol. Chem. 2006, 387, 1063–1074. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wen, C.C.; Kuo, Y.H.; Jan, J.T.; Liang, P.H.; Wang, S.Y.; Liu, H.G.; Lee, C.K.; Chang, S.T.; Kuo, C.J.; Lee, S.S.; et al. Specific plant terpenoids and lignoids possess potent antiviral activities against severe acute respiratory syndrome coronavirus. J. Med. Chem. 2007, 50, 4087–4095. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ryu, Y.B.; Park, S.J.; Kim, Y.M.; Lee, J.Y.; Seo, W.D.; Chang, J.S.; Park, K.H.; Rho, M.C.; Lee, W.S. SARS-CoV 3CLpro inhibitory effects of quinone-methide triterpenes from Tripterygium regelii. Bioorg. Med. Chem. Lett. 2010, 20, 1873–1876. [Google Scholar] [CrossRef]
- Park, J.Y.; Kim, J.H.; Kwon, J.M.; Kwon, H.J.; Jeong, H.J.; Kim, Y.M.; Kim, D.; Lee, W.S.; Ryu, Y.B. Dieckol, a SARS-CoV 3CL(pro) inhibitor, isolated from the edible brown algae Ecklonia cava. Bioorg. Med. Chem. 2013, 21, 3730–3737. [Google Scholar] [CrossRef]
- Park, J.Y.; Ko, J.A.; Kim, D.W.; Kim, Y.M.; Kwon, H.J.; Jeong, H.J.; Kim, C.Y.; Park, K.H.; Lee, W.S.; Ryu, Y.B. Chalcones isolated from Angelica keiskei inhibit cysteine proteases of SARS-CoV. J. Enzyme Inhib. Med. Chem. 2016, 31, 23–30. [Google Scholar] [CrossRef] [Green Version]
- Jo, S.; Kim, S.; Shin, D.H.; Kim, M.S. Inhibition of SARS-CoV 3CL protease by flavonoids. J. Enzyme Inhib. Med. Chem. 2020, 35, 145–151. [Google Scholar] [CrossRef] [Green Version]
- Ramajayam, R.; Tan, K.P.; Liu, H.G.; Liang, P.H. Synthesis and evaluation of pyrazolone compounds as SARS-coronavirus 3C-like protease inhibitors. Bioorg. Med. Chem. 2010, 18, 7849–7854. [Google Scholar] [CrossRef]
- Ratia, K.; Pegan, S.; Takayama, J.; Sleeman, K.; Coughlin, M.; Baliji, S.; Chaudhuri, R.; Fu, W.; Prabhakar, B.S.; Johnson, M.E.; et al. A noncovalent class of papain-like protease/deubiquitinase inhibitors blocks SARS virus replication. Proc. Natl. Acad. Sci. USA 2008, 105, 16119–16124. [Google Scholar] [CrossRef] [Green Version]
- Cho, J.K.; Curtis-Long, M.J.; Lee, K.H.; Kim, D.W.; Ryu, H.W.; Yuk, H.J.; Park, K.H. Geranylated flavonoids displaying SARS-CoV papain-like protease inhibition from the fruits of Paulownia tomentosa. Bioorg. Med. Chem. 2013, 21, 3051–3057. [Google Scholar] [CrossRef]
- Ghosh, A.K.; Takayama, J.; Aubin, Y.; Ratia, K.; Chaudhuri, R.; Baez, Y.; Sleeman, K.; Coughlin, M.; Nichols, D.B.; Mulhearn, D.C.; et al. Structure-based design, synthesis, and biological evaluation of a series of novel and reversible inhibitors for the severe acute respiratory syndrome-coronavirus papain-like protease. J. Med. Chem. 2009, 52, 5228–5240. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ghosh, A.K.; Takayama, J.; Rao, K.V.; Ratia, K.; Chaudhuri, R.; Mulhearn, D.C.; Lee, H.; Nichols, D.B.; Baliji, S.; Baker, S.C.; et al. Severe acute respiratory syndrome coronavirus papain-like novel protease inhibitors: Design, synthesis, protein-ligand X-ray structure and biological evaluation. J. Med. Chem. 2010, 53, 4968–4979. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Park, J.Y.; Jeong, H.J.; Kim, J.H.; Kim, Y.M.; Park, S.J.; Kim, D.; Park, K.H.; Lee, W.S.; Ryu, Y.B. Diarylheptanoids from Alnus japonica inhibit papain-like protease of severe acute respiratory syndrome coronavirus. Biol. Pharm. Bull. 2012, 35, 2036–2042. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cheng, K.W.; Cheng, S.C.; Chen, W.Y.; Lin, M.H.; Chuang, S.J.; Cheng, I.H.; Sun, C.Y.; Chou, C.Y. Thiopurine analogs and mycophenolic acid synergistically inhibit the papain-like protease of Middle East respiratory syndrome coronavirus. Antivir. Res. 2015, 115, 9–16. [Google Scholar] [CrossRef]
- Lin, M.H.; Moses, D.C.; Hsieh, C.H.; Cheng, S.C.; Chen, Y.H.; Sun, C.Y.; Chou, C.Y. Disulfiram can inhibit MERS and SARS coronavirus papain-like proteases via different modes. Antivir. Res. 2018, 150, 155–163. [Google Scholar] [CrossRef] [PubMed]
- Perrin, B.J.; Huttenlocher, A. Calpain. Int. J. Biochem. Cell Biol. 2002, 34, 722–725. [Google Scholar] [CrossRef]
- Wang, K.K. Calpain and caspase: Can you tell the difference?, by kevin K.W. Wang: Vol. 23, pp. 20–26. Trends Neurosci. 2000, 23, 59. [Google Scholar] [CrossRef]
- Neumar, R.W.; Xu, Y.A.; Gada, H.; Guttmann, R.P.; Siman, R. Cross-talk between calpain and caspase proteolytic systems during neuronal apoptosis. J. Biol. Chem. 2003, 278, 14162–14167. [Google Scholar] [CrossRef] [Green Version]
- Teranishi, F.; Liu, Z.Q.; Kunimatsu, M.; Imai, K.; Takeyama, H.; Manabe, T.; Sasaki, M.; Okamoto, T. Calpain is involved in the HIV replication from the latently infected OM10.1 cells. Biochem. Biophys. Res. Commun. 2003, 303, 940–946. [Google Scholar] [CrossRef]
- DeBiasi, R.L.; Edelstein, C.L.; Sherry, B.; Tyler, K.L. Calpain inhibition protects against virus-induced apoptotic myocardial injury. J. Virol. 2001, 75, 351–361. [Google Scholar] [CrossRef] [Green Version]
- Barnard, D.L.; Hubbard, V.D.; Burton, J.; Smee, D.F.; Morrey, J.D.; Otto, M.J.; Sidwell, R.W. Inhibition of severe acute respiratory syndrome-associated coronavirus (SARSCoV) by calpain inhibitors and beta-D-N4-hydroxycytidine. Antivir. Chem. Chemother. 2004, 15, 15–22. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tanner, J.A.; Zheng, B.J.; Zhou, J.; Watt, R.M.; Jiang, J.Q.; Wong, K.L.; Lin, Y.P.; Lu, L.Y.; He, M.L.; Kung, H.F.; et al. The adamantane-derived bananins are potent inhibitors of the helicase activities and replication of SARS coronavirus. Chem. Biol. 2005, 12, 303–311. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, N.; Tanner, J.A.; Wang, Z.; Huang, J.D.; Zheng, B.J.; Zhu, N.; Sun, H. Inhibition of SARS coronavirus helicase by bismuth complexes. Chem. Commun. (Camb) 2007, 4413–4415. [Google Scholar] [CrossRef] [PubMed]
- Yu, M.S.; Lee, J.; Lee, J.M.; Kim, Y.; Chin, Y.W.; Jee, J.G.; Keum, Y.S.; Jeong, Y.J. Identification of myricetin and scutellarein as novel chemical inhibitors of the SARS coronavirus helicase, nsP13. Bioorg. Med. Chem. Lett. 2012, 22, 4049–4054. [Google Scholar] [CrossRef] [PubMed]
- Adedeji, A.O.; Singh, K.; Kassim, A.; Coleman, C.M.; Elliott, R.; Weiss, S.R.; Frieman, M.B.; Sarafianos, S.G. Evaluation of SSYA10-001 as a replication inhibitor of severe acute respiratory syndrome, mouse hepatitis, and Middle East respiratory syndrome coronaviruses. Antimicrob. Agents Chemother. 2014, 58, 4894–4898. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, X.; Liu, Y.; Weiss, S.; Arnold, E.; Sarafianos, S.G.; Ding, J. Molecular model of SARS coronavirus polymerase: Implications for biochemical functions and drug design. Nucleic Acids Res. 2003, 31, 7117–7130. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Taylor, R.; Kotian, P.; Warren, T.; Panchal, R.; Bavari, S.; Julander, J.; Dobo, S.; Rose, A.; El-Kattan, Y.; Taubenheim, B.; et al. BCX4430—A broad-spectrum antiviral adenosine nucleoside analog under development for the treatment of Ebola virus disease. J. Infect. Public Health 2016, 9, 220–226. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Warren, T.K.; Wells, J.; Panchal, R.G.; Stuthman, K.S.; Garza, N.L.; Van Tongeren, S.A.; Dong, L.; Retterer, C.J.; Eaton, B.P.; Pegoraro, G.; et al. Protection against filovirus diseases by a novel broad-spectrum nucleoside analogue BCX4430. Nature 2014, 508, 402–405. [Google Scholar] [CrossRef] [Green Version]
- Julander, J.G.; Siddharthan, V.; Evans, J.; Taylor, R.; Tolbert, K.; Apuli, C.; Stewart, J.; Collins, P.; Gebre, M.; Neilson, S.; et al. Efficacy of the broad-spectrum antiviral compound BCX4430 against Zika virus in cell culture and in a mouse model. Antivir. Res. 2017, 137, 14–22. [Google Scholar] [CrossRef] [Green Version]
- Julander, J.G.; Bantia, S.; Taubenheim, B.R.; Minning, D.M.; Kotian, P.; Morrey, J.D.; Smee, D.F.; Sheridan, W.P.; Babu, Y.S. BCX4430, a novel nucleoside analog, effectively treats yellow fever in a Hamster model. Antimicrob. Agents Chemother. 2014, 58, 6607–6614. [Google Scholar] [CrossRef] [Green Version]
- Cinatl, J.; Morgenstern, B.; Bauer, G.; Chandra, P.; Rabenau, H.; Doerr, H.W. Glycyrrhizin, an active component of liquorice roots, and replication of SARS-associated coronavirus. Lancet 2003, 361, 2045–2046. [Google Scholar] [CrossRef] [Green Version]
- De Clercq, E. Antiviral drugs in current clinical use. J. Clin. Virol. 2004, 30, 115–133. [Google Scholar] [CrossRef] [PubMed]
- Cameron, C.E.; Castro, C. The mechanism of action of ribavirin: Lethal mutagenesis of RNA virus genomes mediated by the viral RNA-dependent RNA polymerase. Curr. Opin. Infect. Dis. 2001, 14, 757–764. [Google Scholar] [CrossRef] [PubMed]
- Hong, Z.; Cameron, C.E. Pleiotropic mechanisms of ribavirin antiviral activities. Prog. Drug Res. 2002, 59, 41–69. [Google Scholar] [CrossRef]
- Chen, F.; Chan, K.H.; Jiang, Y.; Kao, R.Y.; Lu, H.T.; Fan, K.W.; Cheng, V.C.; Tsui, W.H.; Hung, I.F.; Lee, T.S.; et al. In vitro susceptibility of 10 clinical isolates of SARS coronavirus to selected antiviral compounds. J. Clin. Virol. 2004, 31, 69–75. [Google Scholar] [CrossRef]
- Tan, E.L.; Ooi, E.E.; Lin, C.Y.; Tan, H.C.; Ling, A.E.; Lim, B.; Stanton, L.W. Inhibition of SARS coronavirus infection in vitro with clinically approved antiviral drugs. Emerg. Infect. Dis. 2004, 10, 581–586. [Google Scholar] [CrossRef] [Green Version]
- Koren, G.; King, S.; Knowles, S.; Phillips, E. Ribavirin in the treatment of SARS: A new trick for an old drug? CMAJ 2003, 168, 1289–1292. [Google Scholar]
- Morgenstern, B.; Michaelis, M.; Baer, P.C.; Doerr, H.W.; Cinatl, J., Jr. Ribavirin and interferon-beta synergistically inhibit SARS-associated coronavirus replication in animal and human cell lines. Biochem. Biophys. Res. Commun. 2005, 326, 905–908. [Google Scholar] [CrossRef]
- Zhao, Z.; Zhang, F.; Xu, M.; Huang, K.; Zhong, W.; Cai, W.; Yin, Z.; Huang, S.; Deng, Z.; Wei, M.; et al. Description and clinical treatment of an early outbreak of severe acute respiratory syndrome (SARS) in Guangzhou, PR China. J. Med. Microbiol. 2003, 52, 715–720. [Google Scholar] [CrossRef]
- Mazzulli, T.; Farcas, G.A.; Poutanen, S.M.; Willey, B.M.; Low, D.E.; Butany, J.; Asa, S.L.; Kain, K.C. Severe acute respiratory syndrome-associated coronavirus in lung tissue. Emerg. Infect. Dis. 2004, 10, 20–24. [Google Scholar] [CrossRef]
- Peiris, J.S.; Chu, C.M.; Cheng, V.C.; Chan, K.S.; Hung, I.F.; Poon, L.L.; Law, K.I.; Tang, B.S.; Hon, T.Y.; Chan, C.S.; et al. Clinical progression and viral load in a community outbreak of coronavirus-associated SARS pneumonia: A prospective study. Lancet 2003, 361, 1767–1772. [Google Scholar] [CrossRef] [Green Version]
- Barnard, D.L.; Day, C.W.; Bailey, K.; Heiner, M.; Montgomery, R.; Lauridsen, L.; Winslow, S.; Hoopes, J.; Li, J.K.; Lee, J.; et al. Enhancement of the infectivity of SARS-CoV in BALB/c mice by IMP dehydrogenase inhibitors, including ribavirin. Antivir. Res. 2006, 71, 53–63. [Google Scholar] [CrossRef] [PubMed]
- Booth, C.M.; Matukas, L.M.; Tomlinson, G.A.; Rachlis, A.R.; Rose, D.B.; Dwosh, H.A.; Walmsley, S.L.; Mazzulli, T.; Avendano, M.; Derkach, P.; et al. Clinical features and short-term outcomes of 144 patients with SARS in the greater Toronto area. JAMA 2003, 289, 2801–2809. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Knowles, S.R.; Phillips, E.J.; Dresser, L.; Matukas, L. Common adverse events associated with the use of ribavirin for severe acute respiratory syndrome in Canada. Clin. Infect. Dis 2003, 37, 1139–1142. [Google Scholar] [CrossRef] [Green Version]
- Ferron, F.; Subissi, L.; Silveira De Morais, A.T.; Le, N.T.T.; Sevajol, M.; Gluais, L.; Decroly, E.; Vonrhein, C.; Bricogne, G.; Canard, B.; et al. Structural and molecular basis of mismatch correction and ribavirin excision from coronavirus RNA. Proc. Natl. Acad. Sci. USA 2018, 115, E162–E171. [Google Scholar] [CrossRef] [Green Version]
- Mulangu, S.; Dodd, L.E.; Davey, R.T., Jr.; Tshiani Mbaya, O.; Proschan, M.; Mukadi, D.; Lusakibanza Manzo, M.; Nzolo, D.; Tshomba Oloma, A.; Ibanda, A.; et al. A Randomized, Controlled Trial of Ebola Virus Disease Therapeutics. N. Engl. J. Med. 2019, 381, 2293–2303. [Google Scholar] [CrossRef]
- Warren, T.K.; Jordan, R.; Lo, M.K.; Ray, A.S.; Mackman, R.L.; Soloveva, V.; Siegel, D.; Perron, M.; Bannister, R.; Hui, H.C.; et al. Therapeutic efficacy of the small molecule GS-5734 against Ebola virus in rhesus monkeys. Nature 2016, 531, 381–385. [Google Scholar] [CrossRef]
- Sheahan, T.P.; Sims, A.C.; Graham, R.L.; Menachery, V.D.; Gralinski, L.E.; Case, J.B.; Leist, S.R.; Pyrc, K.; Feng, J.Y.; Trantcheva, I.; et al. Broad-spectrum antiviral GS-5734 inhibits both epidemic and zoonotic coronaviruses. Sci. Transl. Med. 2017, 9. [Google Scholar] [CrossRef] [Green Version]
- Lu, X.; Pan, J.; Tao, J.; Guo, D. SARS-CoV nucleocapsid protein antagonizes IFN-beta response by targeting initial step of IFN-beta induction pathway, and its C-terminal region is critical for the antagonism. Virus Genes 2011, 42, 37–45. [Google Scholar] [CrossRef] [Green Version]
- Akerstrom, S.; Mirazimi, A.; Tan, Y.J. Inhibition of SARS-CoV replication cycle by small interference RNAs silencing specific SARS proteins, 7a/7b, 3a/3b and S. Antivir. Res. 2007, 73, 219–227. [Google Scholar] [CrossRef]
- He, M.L.; Zheng, B.J.; Chen, Y.; Wong, K.L.; Huang, J.D.; Lin, M.C.; Yuen, K.Y.; Sung, J.J.; Kung, H.F. Development of interfering RNA agents to inhibit SARS-associated coronavirus infection and replication. Hong Kong Med. J. 2009, 15, 28–31. [Google Scholar] [PubMed]
- Wilson, L.; Gage, P.; Ewart, G. Hexamethylene amiloride blocks E protein ion channels and inhibits coronavirus replication. Virology 2006, 353, 294–306. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- He, R.; Adonov, A.; Traykova-Adonova, M.; Cao, J.; Cutts, T.; Grudesky, E.; Deschambaul, Y.; Berry, J.; Drebot, M.; Li, X. Potent and selective inhibition of SARS coronavirus replication by aurintricarboxylic acid. Biochem. Biophys. Res. Commun. 2004, 320, 1199–1203. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.J.; Jan, J.T.; Chen, C.M.; Hsieh, H.P.; Hwang, D.R.; Liu, H.W.; Liu, C.Y.; Huang, H.W.; Chen, S.C.; Hong, C.F.; et al. Inhibition of severe acute respiratory syndrome coronavirus replication by niclosamide. Antimicrob. Agents Chemother. 2004, 48, 2693–2696. [Google Scholar] [CrossRef] [Green Version]
- Stadler, K.; Ha, H.R.; Ciminale, V.; Spirli, C.; Saletti, G.; Schiavon, M.; Bruttomesso, D.; Bigler, L.; Follath, F.; Pettenazzo, A.; et al. Amiodarone alters late endosomes and inhibits SARS coronavirus infection at a post-endosomal level. Am. J. Respir. Cell Mol. Biol. 2008, 39, 142–149. [Google Scholar] [CrossRef] [PubMed]
- Knoops, K.; Kikkert, M.; Worm, S.H.; Zevenhoven-Dobbe, J.C.; van der Meer, Y.; Koster, A.J.; Mommaas, A.M.; Snijder, E.J. SARS-coronavirus replication is supported by a reticulovesicular network of modified endoplasmic reticulum. PLoS Biol. 2008, 6, e226. [Google Scholar] [CrossRef]
- Lundin, A.; Dijkman, R.; Bergstrom, T.; Kann, N.; Adamiak, B.; Hannoun, C.; Kindler, E.; Jonsdottir, H.R.; Muth, D.; Kint, J.; et al. Targeting membrane-bound viral RNA synthesis reveals potent inhibition of diverse coronaviruses including the middle East respiratory syndrome virus. PLoS Pathog. 2014, 10, e1004166. [Google Scholar] [CrossRef] [Green Version]
- Rider, T.H.; Zook, C.E.; Boettcher, T.L.; Wick, S.T.; Pancoast, J.S.; Zusman, B.D. Broad-spectrum antiviral therapeutics. PLoS ONE 2011, 6, e22572. [Google Scholar] [CrossRef] [Green Version]
- Dattilo, J.B.; Makhoul, R.G. The role of nitric oxide in vascular biology and pathobiology. Ann. Vasc. Surg. 1997, 11, 307–314. [Google Scholar] [CrossRef]
- Lane, T.E.; Paoletti, A.D.; Buchmeier, M.J. Disassociation between the in vitro and in vivo effects of nitric oxide on a neurotropic murine coronavirus. J. Virol. 1997, 71, 2202–2210. [Google Scholar] [CrossRef] [Green Version]
- Pope, M.; Marsden, P.A.; Cole, E.; Sloan, S.; Fung, L.S.; Ning, Q.; Ding, J.W.; Leibowitz, J.L.; Phillips, M.J.; Levy, G.A. Resistance to murine hepatitis virus strain 3 is dependent on production of nitric oxide. J. Virol. 1998, 72, 7084–7090. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reiss, C.S.; Komatsu, T. Does nitric oxide play a critical role in viral infections? J. Virol. 1998, 72, 4547–4551. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Keyaerts, E.; Vijgen, L.; Chen, L.; Maes, P.; Hedenstierna, G.; Van Ranst, M. Inhibition of SARS-coronavirus infection in vitro by S-nitroso-N-acetylpenicillamine, a nitric oxide donor compound. Int. J. Infect. Dis. 2004, 8, 223–226. [Google Scholar] [CrossRef] [Green Version]
- Chen, L.; Liu, P.; Gao, H.; Sun, B.; Chao, D.; Wang, F.; Zhu, Y.; Hedenstierna, G.; Wang, C.G. Inhalation of nitric oxide in the treatment of severe acute respiratory syndrome: A rescue trial in Beijing. Clin. Infect. Dis. 2004, 39, 1531–1535. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pfefferle, S.; Schopf, J.; Kogl, M.; Friedel, C.C.; Muller, M.A.; Carbajo-Lozoya, J.; Stellberger, T.; von Dall’Armi, E.; Herzog, P.; Kallies, S.; et al. The SARS-coronavirus-host interactome: Identification of cyclophilins as target for pan-coronavirus inhibitors. PLoS Pathog. 2011, 7, e1002331. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Clipstone, N.A.; Crabtree, G.R. Identification of calcineurin as a key signalling enzyme in T-lymphocyte activation. Nature 1992, 357, 695–697. [Google Scholar] [CrossRef] [PubMed]
- Fruman, D.A.; Klee, C.B.; Bierer, B.E.; Burakoff, S.J. Calcineurin phosphatase activity in T lymphocytes is inhibited by FK 506 and cyclosporin A. Proc. Natl. Acad. Sci. USA 1992, 89, 3686–3690. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- de Wilde, A.H.; Falzarano, D.; Zevenhoven-Dobbe, J.C.; Beugeling, C.; Fett, C.; Martellaro, C.; Posthuma, C.C.; Feldmann, H.; Perlman, S.; Snijder, E.J. Alisporivir inhibits MERS- and SARS-coronavirus replication in cell culture, but not SARS-coronavirus infection in a mouse model. Virus Res. 2017, 228, 7–13. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.H.; Tao, P.Z.; Liu, Y.Z.; Jiang, J.D. Geldanamycin, a ligand of heat shock protein 90, inhibits the replication of herpes simplex virus type 1 in vitro. Antimicrob. Agents Chemother. 2004, 48, 867–872. [Google Scholar] [CrossRef] [Green Version]
- Balzarini, J.; Keyaerts, E.; Vijgen, L.; Egberink, H.; De Clercq, E.; Van Ranst, M.; Printsevskaya, S.S.; Olsufyeva, E.N.; Solovieva, S.E.; Preobrazhenskaya, M.N. Inhibition of feline (FIPV) and human (SARS) coronavirus by semisynthetic derivatives of glycopeptide antibiotics. Antivir. Res. 2006, 72, 20–33. [Google Scholar] [CrossRef]
- Sen, G.C. Viruses and interferons. Annu. Rev. Microbiol. 2001, 55, 255–281. [Google Scholar] [CrossRef] [PubMed]
- Nelson, M.; Rubio, R.; Lazzarin, A.; Romanova, S.; Luetkemeyer, A.; Conway, B.; Molina, J.M.; Xu, D.; Srinivasan, S.; Portsmouth, S. Safety and Efficacy of Pegylated Interferon Lambda, Ribavirin, and Daclatasvir in HCV and HIV-Coinfected Patients. J. Interferon Cytokine Res. 2017, 37, 103–111. [Google Scholar] [CrossRef] [PubMed]
- Cinatl, J.; Morgenstern, B.; Bauer, G.; Chandra, P.; Rabenau, H.; Doerr, H.W. Treatment of SARS with human interferons. Lancet 2003, 362, 293–294. [Google Scholar] [CrossRef]
- Haagmans, B.L.; Kuiken, T.; Martina, B.E.; Fouchier, R.A.; Rimmelzwaan, G.F.; van Amerongen, G.; van Riel, D.; de Jong, T.; Itamura, S.; Chan, K.H.; et al. Pegylated interferon-alpha protects type 1 pneumocytes against SARS coronavirus infection in macaques. Nat. Med. 2004, 10, 290–293. [Google Scholar] [CrossRef] [PubMed]
- Cheng, V.C.; Chan, J.F.; To, K.K.; Yuen, K.Y. Clinical management and infection control of SARS: Lessons learned. Antivir. Res. 2013, 100, 407–419. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Baltina, L.A. Chemical modification of glycyrrhizic acid as a route to new bioactive compounds for medicine. Curr. Med. Chem. 2003, 10, 155–171. [Google Scholar] [CrossRef]
- Shibata, S. A drug over the millennia: Pharmacognosy, chemistry, and pharmacology of licorice. Yakugaku Zasshi 2000, 120, 849–862. [Google Scholar] [CrossRef]
- Lin, J.C. Mechanism of action of glycyrrhizic acid in inhibition of Epstein-Barr virus replication in vitro. Antivir. Res. 2003, 59, 41–47. [Google Scholar] [CrossRef]
- Pompei, R.; Flore, O.; Marccialis, M.A.; Pani, A.; Loddo, B. Glycyrrhizic acid inhibits virus growth and inactivates virus particles. Nature 1979, 281, 689–690. [Google Scholar] [CrossRef]
- Crance, J.M.; Scaramozzino, N.; Jouan, A.; Garin, D. Interferon, ribavirin, 6-azauridine and glycyrrhizin: Antiviral compounds active against pathogenic flaviviruses. Antivir. Res. 2003, 58, 73–79. [Google Scholar] [CrossRef]
- Sasaki, H.; Takei, M.; Kobayashi, M.; Pollard, R.B.; Suzuki, F. Effect of glycyrrhizin, an active component of licorice roots, on HIV replication in cultures of peripheral blood mononuclear cells from HIV-seropositive patients. Pathobiology 2002, 70, 229–236. [Google Scholar] [CrossRef]
- Cherng, J.M.; Lin, H.J.; Hsu, Y.H.; Hung, M.S.; Lin, J.C. A quantitative bioassay for HIV-1 gene expression based on UV activation: Effect of glycyrrhizic acid. Antivir. Res. 2004, 62, 27–36. [Google Scholar] [CrossRef] [PubMed]
- O’Brian, C.A.; Ward, N.E.; Vogel, V.G. Inhibition of protein kinase C by the 12-O-tetradecanoylphorbol-13-acetate antagonist glycyrrhetic acid. Cancer Lett. 1990, 49, 9–12. [Google Scholar] [CrossRef]
- Harada, S.; Karino, A.; Shimoyama, Y.; Shamsa, F.; Ohtsuki, K. Identification of glycyrrhizin-binding protein kinase as casein kinase II and characterization of its associated phosphate acceptors in mouse liver. Biochem. Biophys. Res. Commun. 1996, 227, 102–109. [Google Scholar] [CrossRef] [PubMed]
- Yi, H.; Nakashima, I.; Isobe, K. Enhancement of nitric oxide production from activated macrophages by glycyrrhizin. Am. J. Chin. Med. 1996, 24, 271–278. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.Y.; Jan, J.T.; Ma, S.H.; Kuo, C.J.; Juan, H.F.; Cheng, Y.S.; Hsu, H.H.; Huang, H.C.; Wu, D.; Brik, A.; et al. Small molecules targeting severe acute respiratory syndrome human coronavirus. Proc. Natl. Acad. Sci. USA 2004, 101, 10012–10017. [Google Scholar] [CrossRef] [Green Version]
- Li, S.Y.; Chen, C.; Zhang, H.Q.; Guo, H.Y.; Wang, H.; Wang, L.; Zhang, X.; Hua, S.N.; Yu, J.; Xiao, P.G.; et al. Identification of natural compounds with antiviral activities against SARS-associated coronavirus. Antivir. Res. 2005, 67, 18–23. [Google Scholar] [CrossRef]
- Kaul, D. An overview of coronaviruses including the SARS-2 coronavirus—Molecular biology, epidemiology and clinical implications. Curr. Med. Res. Pract. 2020. [Google Scholar] [CrossRef]
- Helmy, Y.A.; Fawzy, M.; Elaswad, A.; Sobieh, A.; Kenney, S.P.; Shehata, A.A. The COVID-19 Pandemic: A Comprehensive Review of Taxonomy, Genetics, Epidemiology, Diagnosis, Treatment, and Control. J. Clin. Med. 2020, 9, 1225. [Google Scholar] [CrossRef]
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [Green Version]
- Andersen, K.G.; Rambaut, A.; Lipkin, W.I.; Holmes, E.C.; Garry, R.F. The proximal origin of SARS-CoV-2. Nat. Med. 2020, 26, 450–452. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lau, S.K.P.; Luk, H.K.H.; Wong, A.C.P.; Li, K.S.M.; Zhu, L.; He, Z.; Fung, J.; Chan, T.T.Y.; Fung, K.S.C.; Woo, P.C.Y. Possible Bat Origin of Severe Acute Respiratory Syndrome Coronavirus 2. Emerg. Infect. Dis. 2020, 26. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Z.; Peng, F.; Xu, B.; Zhao, J.; Liu, H.; Peng, J.; Li, Q.; Jiang, C.; Zhou, Y.; Liu, S.; et al. Risk factors of critical & mortal COVID-19 cases: A systematic literature review and meta-analysis. J. Infect. 2020. [Google Scholar] [CrossRef]
- Ashour, H.M.; Elkhatib, W.F.; Rahman, M.M.; Elshabrawy, H.A. Insights into the Recent 2019 Novel Coronavirus (SARS-CoV-2) in Light of Past Human Coronavirus Outbreaks. Pathogens 2020, 9, 186. [Google Scholar] [CrossRef] [Green Version]
- Li, B.; Li, X.; Wang, Y.; Han, Y.; Wang, Y.; Wang, C.; Zhang, G.; Jin, J.; Jia, H.; Fan, F.; et al. Diagnostic value and key features of computed tomography in Coronavirus Disease 2019. Emerg. Microbes Infect. 2020, 9, 787–793. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, A.; Peng, Y.; Huang, B.; Ding, X.; Wang, X.; Niu, P.; Meng, J.; Zhu, Z.; Zhang, Z.; Wang, J.; et al. Genome Composition and Divergence of the Novel Coronavirus (2019-nCoV) Originating in China. Cell Host Microbe 2020, 27, 325–328. [Google Scholar] [CrossRef] [Green Version]
- Wan, Y.; Shang, J.; Graham, R.; Baric, R.S.; Li, F. Receptor Recognition by the Novel Coronavirus from Wuhan: An Analysis Based on Decade-Long Structural Studies of SARS Coronavirus. J. Virol. 2020, 94. [Google Scholar] [CrossRef] [Green Version]
- Wrapp, D.; Wang, N.; Corbett, K.S.; Goldsmith, J.A.; Hsieh, C.L.; Abiona, O.; Graham, B.S.; McLellan, J.S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263. [Google Scholar] [CrossRef] [Green Version]
- Wang, Q.; Zhang, Y.; Wu, L.; Niu, S.; Song, C.; Zhang, Z.; Lu, G.; Qiao, C.; Hu, Y.; Yuen, K.Y.; et al. Structural and Functional Basis of SARS-CoV-2 Entry by Using Human ACE2. Cell 2020, 181, 894–904. [Google Scholar] [CrossRef]
- Tai, W.; He, L.; Zhang, X.; Pu, J.; Voronin, D.; Jiang, S.; Zhou, Y.; Du, L. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: Implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell Mol. Immunol. 2020, 17, 613–620. [Google Scholar] [CrossRef] [Green Version]
- Othman, H.; Bouslama, Z.; Brandenburg, J.T.; da Rocha, J.; Hamdi, Y.; Ghedira, K.; Srairi-Abid, N.; Hazelhurst, S. Interaction of the spike protein RBD from SARS-CoV-2 with ACE2: Similarity with SARS-CoV, hot-spot analysis and effect of the receptor polymorphism. Biochem Biophys. Res. Commun. 2020, 527, 702–708. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Kruger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280. [Google Scholar] [CrossRef] [PubMed]
- Coutard, B.; Valle, C.; de Lamballerie, X.; Canard, B.; Seidah, N.G.; Decroly, E. The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antivir. Res. 2020, 176, 104742. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Yuan, Q.; Wang, H.; Liu, W.; Liao, X.; Su, Y.; Wang, X.; Yuan, J.; Li, T.; Li, J.; et al. Antibody responses to SARS-CoV-2 in patients of novel coronavirus disease 2019. Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef] [PubMed]
- Okba, N.M.A.; Muller, M.A.; Li, W.; Wang, C.; GeurtsvanKessel, C.H.; Corman, V.M.; Lamers, M.M.; Sikkema, R.S.; de Bruin, E.; Chandler, F.D.; et al. Severe Acute Respiratory Syndrome Coronavirus 2-Specific Antibody Responses in Coronavirus Disease 2019 Patients. Emerg. Infect. Dis. 2020, 26. [Google Scholar] [CrossRef]
- Ou, X.; Liu, Y.; Lei, X.; Li, P.; Mi, D.; Ren, L.; Guo, L.; Guo, R.; Chen, T.; Hu, J.; et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat. Commun. 2020, 11, 1620. [Google Scholar] [CrossRef] [Green Version]
- Shen, C.; Wang, Z.; Zhao, F.; Yang, Y.; Li, J.; Yuan, J.; Wang, F.; Li, D.; Yang, M.; Xing, L.; et al. Treatment of 5 Critically Ill Patients With COVID-19 With Convalescent Plasma. JAMA 2020. [Google Scholar] [CrossRef]
- Tian, X.; Li, C.; Huang, A.; Xia, S.; Lu, S.; Shi, Z.; Lu, L.; Jiang, S.; Yang, Z.; Wu, Y.; et al. Potent binding of 2019 novel coronavirus spike protein by a SARS coronavirus-specific human monoclonal antibody. Emerg. Microbes Infect. 2020, 9, 382–385. [Google Scholar] [CrossRef] [Green Version]
- Wang, C.; Li, W.; Drabek, D.; Okba, N.M.A.; van Haperen, R.; Osterhaus, A.; van Kuppeveld, F.J.M.; Haagmans, B.L.; Grosveld, F.; Bosch, B.J. A human monoclonal antibody blocking SARS-CoV-2 infection. Nat. Commun. 2020, 11, 2251. [Google Scholar] [CrossRef]
- Yi, C.; Sun, X.; Ye, J.; Ding, L.; Liu, M.; Yang, Z.; Lu, X.; Zhang, Y.; Ma, L.; Gu, W.; et al. Key residues of the receptor binding motif in the spike protein of SARS-CoV-2 that interact with ACE2 and neutralizing antibodies. Cell Mol. Immunol. 2020. [Google Scholar] [CrossRef]
- Wu, Y.; Wang, F.; Shen, C.; Peng, W.; Li, D.; Zhao, C.; Li, Z.; Li, S.; Bi, Y.; Yang, Y.; et al. A noncompeting pair of human neutralizing antibodies block COVID-19 virus binding to its receptor ACE2. Science 2020. [Google Scholar] [CrossRef] [PubMed]
- Monteil, V.; Kwon, H.; Prado, P.; Hagelkruys, A.; Wimmer, R.A.; Stahl, M.; Leopoldi, A.; Garreta, E.; Hurtado Del Pozo, C.; Prosper, F.; et al. Inhibition of SARS-CoV-2 Infections in Engineered Human Tissues Using Clinical-Grade Soluble Human ACE2. Cell 2020. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Cao, R.; Zhang, L.; Yang, X.; Liu, J.; Xu, M.; Shi, Z.; Hu, Z.; Zhong, W.; Xiao, G. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. 2020, 30, 269–271. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Cao, R.; Xu, M.; Wang, X.; Zhang, H.; Hu, H.; Li, Y.; Hu, Z.; Zhong, W.; Wang, M. Hydroxychloroquine, a less toxic derivative of chloroquine, is effective in inhibiting SARS-CoV-2 infection in vitro. Cell Discov. 2020, 6, 16. [Google Scholar] [CrossRef] [Green Version]
- Multicenter Collaboration Group of Department of Science and Technology of Guangdong Province and Health Commission of Guangdong Province for Chloroquine in the Treatment of Novel Coronavirus Pneumonia. Expert consensus on chloroquine phosphate for the treatment of novel coronavirus pneumonia. Zhonghua Jie He He Hu Xi Za Zhi 2020, 43, 185–188. [Google Scholar] [CrossRef]
- Cortegiani, A.; Ingoglia, G.; Ippolito, M.; Giarratano, A.; Einav, S. A systematic review on the efficacy and safety of chloroquine for the treatment of COVID-19. J. Crit. Care 2020, 57, 279–283. [Google Scholar] [CrossRef]
- Gao, J.; Tian, Z.; Yang, X. Breakthrough: Chloroquine phosphate has shown apparent efficacy in treatment of COVID-19 associated pneumonia in clinical studies. Biosci. Trends 2020, 14, 72–73. [Google Scholar] [CrossRef] [Green Version]
- Gautret, P.; Lagier, J.C.; Parola, P.; Hoang, V.T.; Meddeb, L.; Mailhe, M.; Doudier, B.; Courjon, J.; Giordanengo, V.; Vieira, V.E.; et al. Hydroxychloroquine and azithromycin as a treatment of COVID-19: Results of an open-label non-randomized clinical trial. Int. J. Antimicrob. Agents 2020, 105949. [Google Scholar] [CrossRef]
- Rosenberg, E.S.; Dufort, E.M.; Udo, T.; Wilberschied, L.A.; Kumar, J.; Tesoriero, J.; Weinberg, P.; Kirkwood, J.; Muse, A.; DeHovitz, J.; et al. Association of Treatment With Hydroxychloroquine or Azithromycin With In-Hospital Mortality in Patients With COVID-19 in New York State. JAMA 2020. [Google Scholar] [CrossRef]
- Hoffmann, M.; Schroeder, S.; Kleine-Weber, H.; Muller, M.A.; Drosten, C.; Pohlmann, S. Nafamostat mesylate blocks activation of SARS-CoV-2: New treatment option for COVID-19. Antimicrob. Agents Chemother. 2020. [Google Scholar] [CrossRef] [Green Version]
- Zeng, L.Y.; Yang, J.; Liu, S. Investigational hemagglutinin-targeted influenza virus inhibitors. Expert Opin. Investig. Drugs 2017, 26, 63–73. [Google Scholar] [CrossRef] [PubMed]
- Ledford, H. Hopes rise for coronavirus drug remdesivir. Nature 2020. [Google Scholar] [CrossRef] [PubMed]
- Elfiky, A.A. Anti-HCV, nucleotide inhibitors, repurposing against COVID-19. Life Sci. 2020, 248, 117477. [Google Scholar] [CrossRef] [PubMed]
- Mifsud, E.J.; Hayden, F.G.; Hurt, A.C. Antivirals targeting the polymerase complex of influenza viruses. Antivir. Res. 2019, 169, 104545. [Google Scholar] [CrossRef]
- Delang, L.; Abdelnabi, R.; Neyts, J. Favipiravir as a potential countermeasure against neglected and emerging RNA viruses. Antivir. Res. 2018, 153, 85–94. [Google Scholar] [CrossRef]
- Cai, Q.; Yang, M.; Liu, D.; Chen, J.; Shu, D.; Xia, J.; Liao, X.; Gu, Y.; Cai, Q.; Yang, Y.; et al. Experimental Treatment with Favipiravir for COVID-19: An Open-Label Control Study. Engineering (Beijing) 2020. [Google Scholar] [CrossRef]
- Lim, J.; Jeon, S.; Shin, H.Y.; Kim, M.J.; Seong, Y.M.; Lee, W.J.; Choe, K.W.; Kang, Y.M.; Lee, B.; Park, S.J. Case of the Index Patient Who Caused Tertiary Transmission of COVID-19 Infection in Korea: The Application of Lopinavir/Ritonavir for the Treatment of COVID-19 Infected Pneumonia Monitored by Quantitative RT-PCR. J. Korean Med. Sci. 2020, 35, e79. [Google Scholar] [CrossRef]
- Cao, B.; Wang, Y.; Wen, D.; Liu, W.; Wang, J.; Fan, G.; Ruan, L.; Song, B.; Cai, Y.; Wei, M.; et al. A Trial of Lopinavir-Ritonavir in Adults Hospitalized with Severe Covid-19. N. Engl. J. Med. 2020, 382, 1787–1799. [Google Scholar] [CrossRef]
- Chen, Y.W.; Yiu, C.B.; Wong, K.Y. Prediction of the SARS-CoV-2 (2019-nCoV) 3C-like protease (3CL (pro)) structure: Virtual screening reveals velpatasvir, ledipasvir, and other drug repurposing candidates. F1000Res 2020, 9, 129. [Google Scholar] [CrossRef] [Green Version]
- Cinatl, J., Jr.; Michaelis, M.; Scholz, M.; Doerr, H.W. Role of interferons in the treatment of severe acute respiratory syndrome. Expert Opin. Biol. Ther. 2004, 4, 827–836. [Google Scholar] [CrossRef]
- Felsenstein, S.; Herbert, J.A.; McNamara, P.S.; Hedrich, C.M. COVID-19: Immunology and treatment options. Clin. Immunol. 2020, 215, 108448. [Google Scholar] [CrossRef] [PubMed]
- Mascarenhas, J.; Mughal, T.I.; Verstovsek, S. Biology and clinical management of myeloproliferative neoplasms and development of the JAK inhibitor ruxolitinib. Curr. Med. Chem. 2012, 19, 4399–4413. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Richardson, P.; Griffin, I.; Tucker, C.; Smith, D.; Oechsle, O.; Phelan, A.; Stebbing, J. Baricitinib as potential treatment for 2019-nCoV acute respiratory disease. Lancet 2020, 395, e30–e31. [Google Scholar] [CrossRef] [Green Version]
- Macrae, F.L.; Peacock-Young, B.; Bowman, P.; Baker, S.R.; Quested, S.; Linton, E.; Hillmen, P.; Griffin, M.; Munir, T.; Payne, D.; et al. Patients with paroxysmal nocturnal hemoglobinuria demonstrate a prothrombotic clotting phenotype which is improved by complement inhibition with eculizumab. Am. J. Hematol. 2020. [Google Scholar] [CrossRef]
- Ellithi, M.; Shahid, M.; Abdullah, H.M.; Bleeker, J. Complement C3 mutation causing atypical hemolytic uremic syndrome successfully treated with eculizumab. Hematol. Transfus. Cell Ther. 2020. [Google Scholar] [CrossRef]
- Patriquin, C.J.; Kuo, K.H.M. Eculizumab and Beyond: The Past, Present, and Future of Complement Therapeutics. Transfus. Med. Rev. 2019, 33, 256–265. [Google Scholar] [CrossRef]
- Wood, A.J.T.; Vassallo, A.; Summers, C.; Chilvers, E.R.; Conway-Morris, A. C5a anaphylatoxin and its role in critical illness-induced organ dysfunction. Eur. J. Clin. Investig. 2018, 48, e13028. [Google Scholar] [CrossRef] [Green Version]
- Biggioggero, M.; Crotti, C.; Becciolini, A.; Favalli, E.G. Tocilizumab in the treatment of rheumatoid arthritis: An evidence-based review and patient selection. Drug Des. Devel. Ther. 2019, 13, 57–70. [Google Scholar] [CrossRef] [Green Version]
- Burmester, G.R.; Lin, Y.; Patel, R.; van Adelsberg, J.; Mangan, E.K.; Graham, N.M.; van Hoogstraten, H.; Bauer, D.; Ignacio Vargas, J.; Lee, E.B. Efficacy and safety of sarilumab monotherapy versus adalimumab monotherapy for the treatment of patients with active rheumatoid arthritis (MONARCH): A randomised, double-blind, parallel-group phase III trial. Ann. Rheum. Dis. 2017, 76, 840–847. [Google Scholar] [CrossRef]
- Fajgenbaum, D.C. Novel insights and therapeutic approaches in idiopathic multicentric Castleman disease. Hematol. Am. Soc. Hematol. Educ. Program. 2018, 2018, 318–325. [Google Scholar] [CrossRef] [Green Version]
- Chen, R.; Chen, B. Siltuximab (CNTO 328): A promising option for human malignancies. Drug Des. Dev. Ther. 2015, 9, 3455–3458. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Verdecchia, P.; Cavallini, C.; Spanevello, A.; Angeli, F. The pivotal link between ACE2 deficiency and SARS-CoV-2 infection. Eur. J. Intern. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Vaduganathan, M.; Vardeny, O.; Michel, T.; McMurray, J.J.V.; Pfeffer, M.A.; Solomon, S.D. Renin-Angiotensin-Aldosterone System Inhibitors in Patients with Covid-19. N. Engl. J. Med. 2020, 382, 1653–1659. [Google Scholar] [CrossRef] [PubMed]
- Pachetti, M.; Marini, B.; Benedetti, F.; Giudici, F.; Mauro, E.; Storici, P.; Masciovecchio, C.; Angeletti, S.; Ciccozzi, M.; Gallo, R.C.; et al. Emerging SARS-CoV-2 mutation hot spots include a novel RNA-dependent-RNA polymerase variant. J. Transl. Med. 2020, 18, 179. [Google Scholar] [CrossRef] [Green Version]
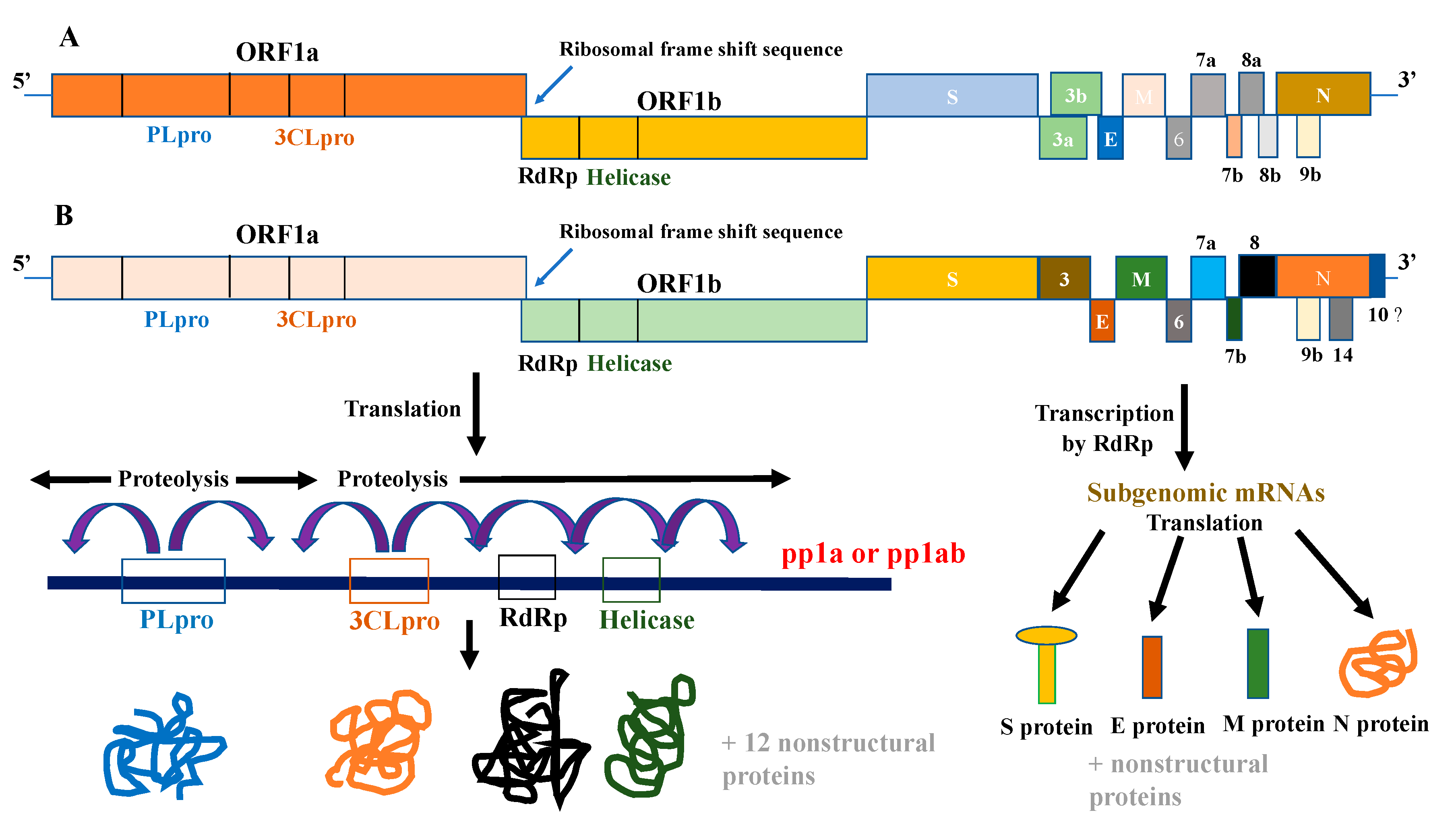
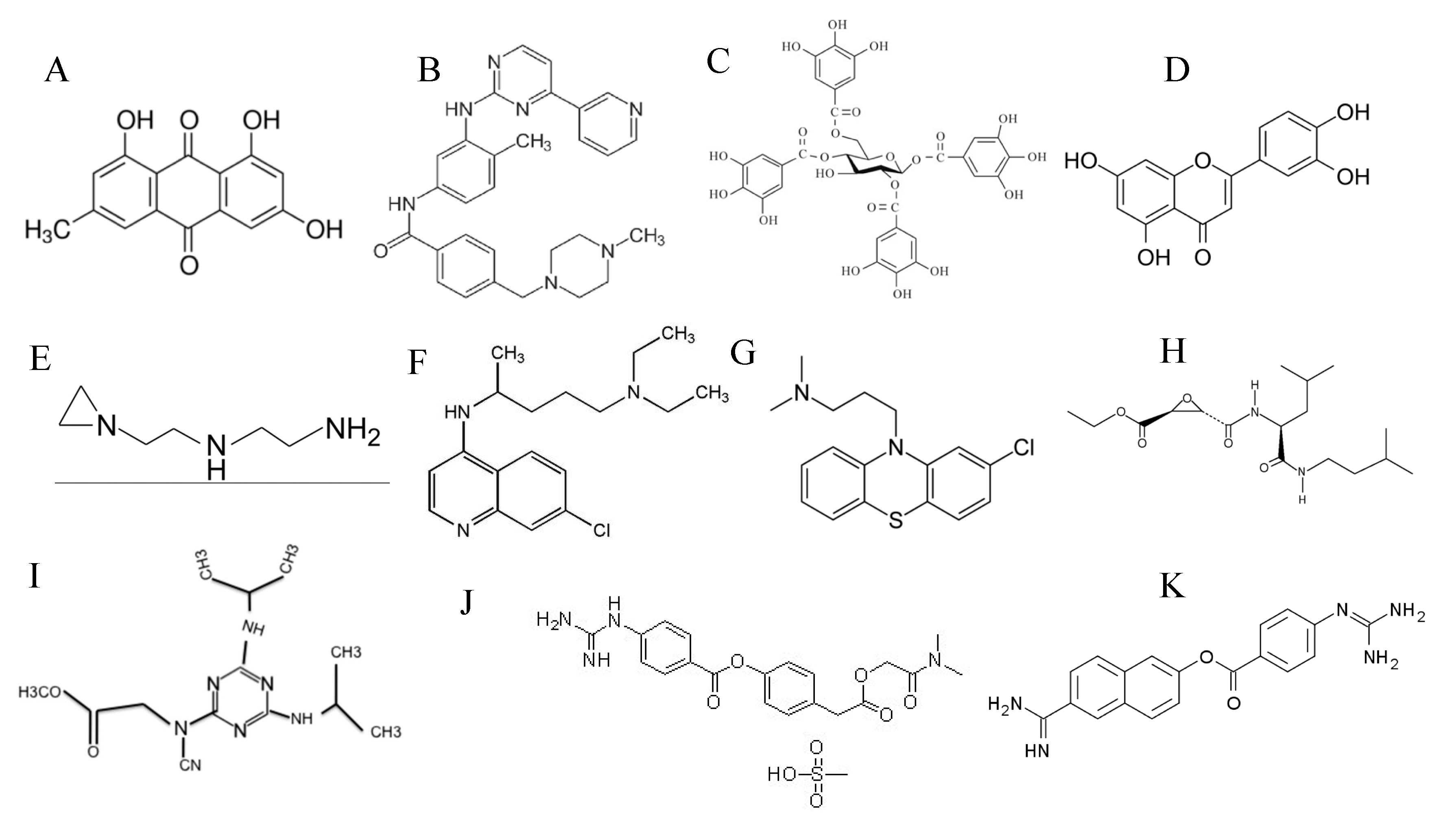
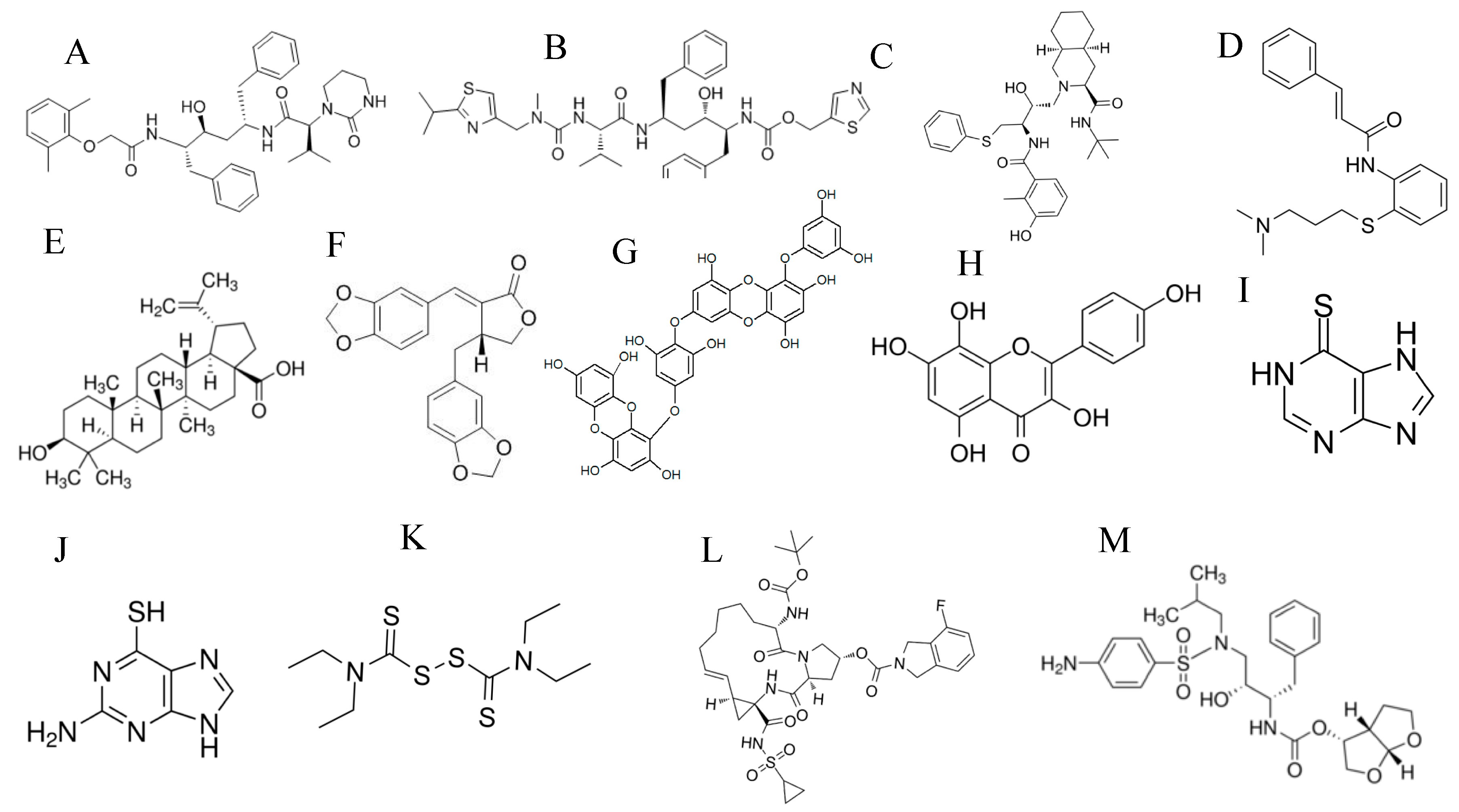
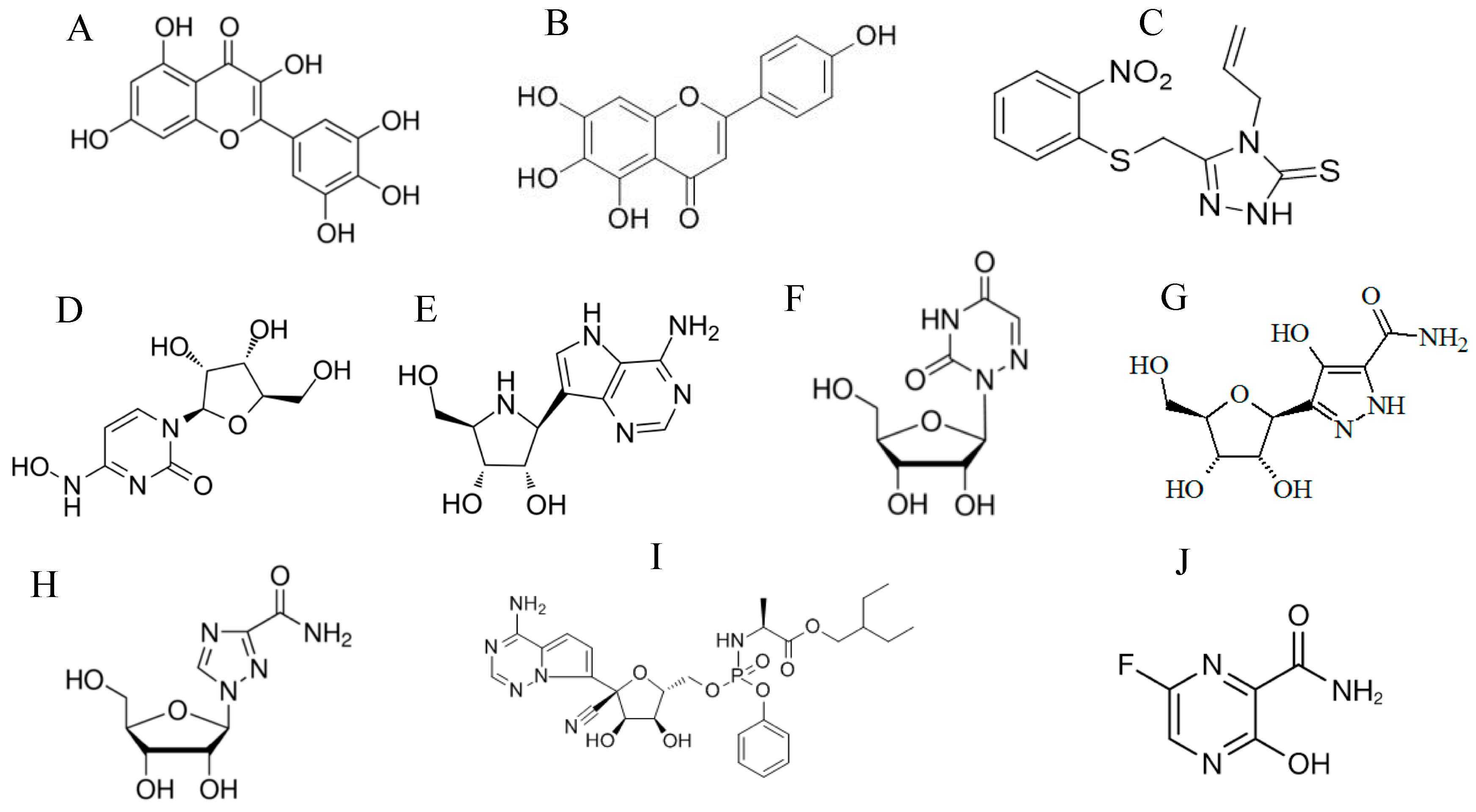
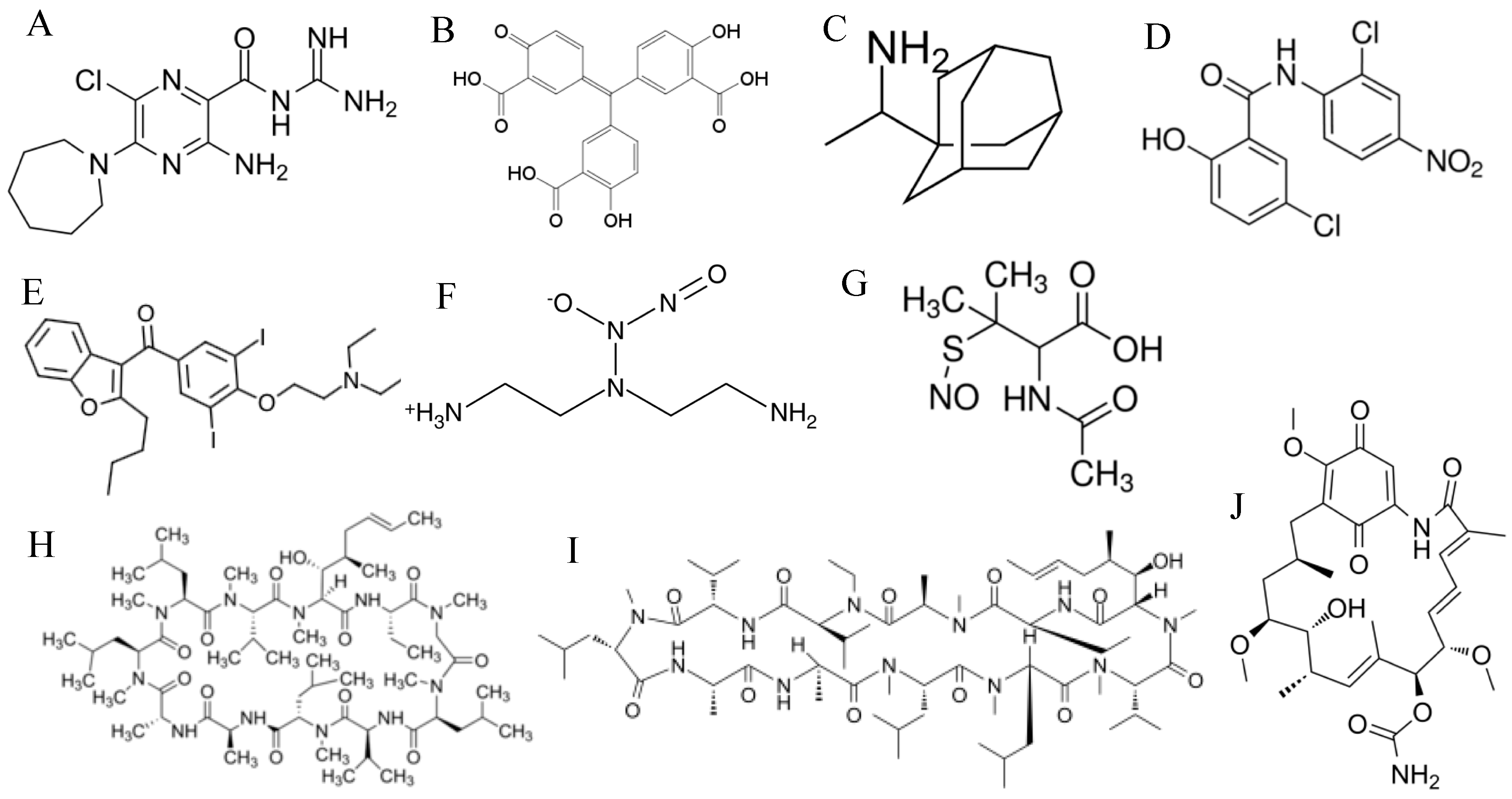
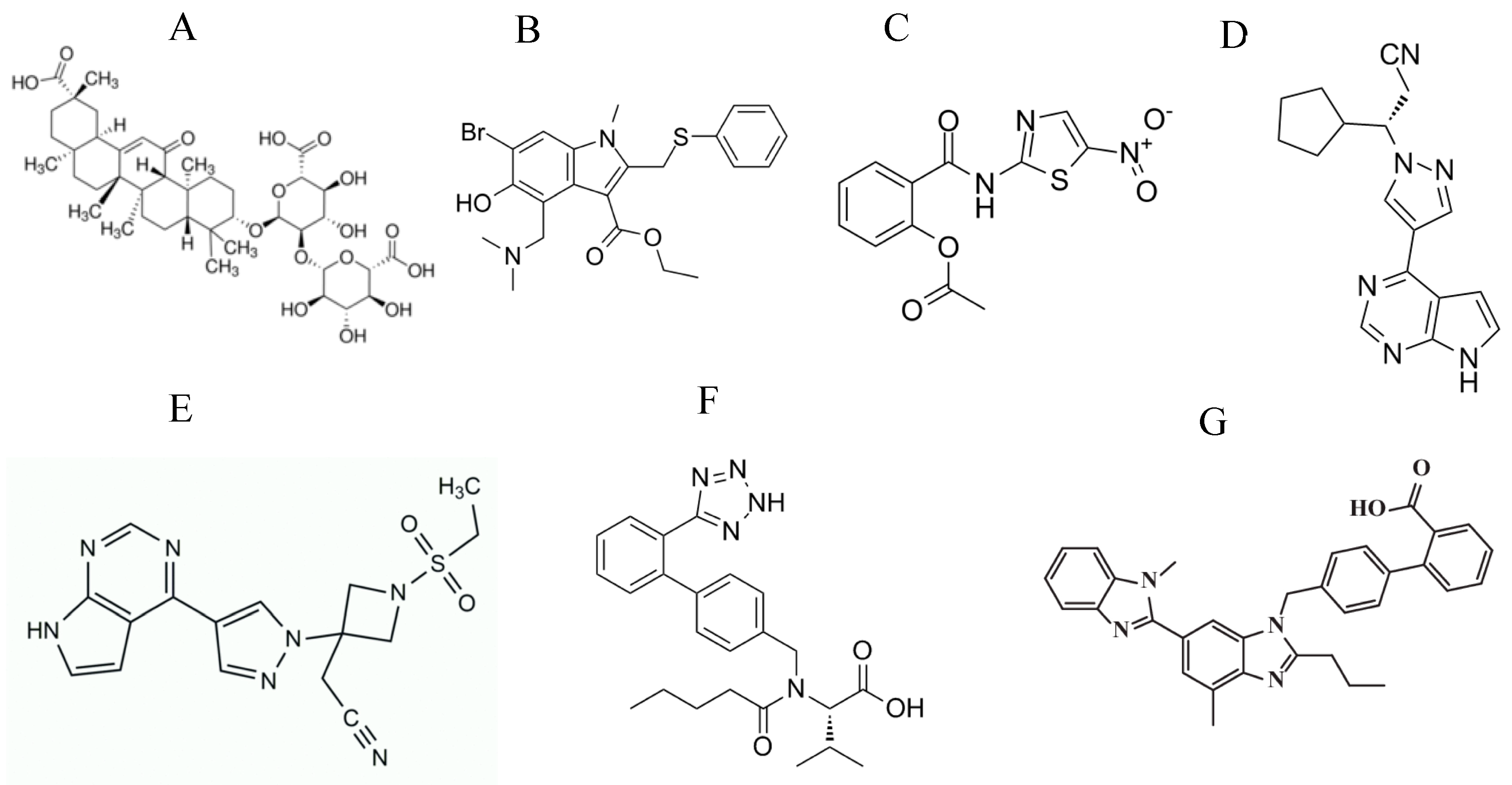
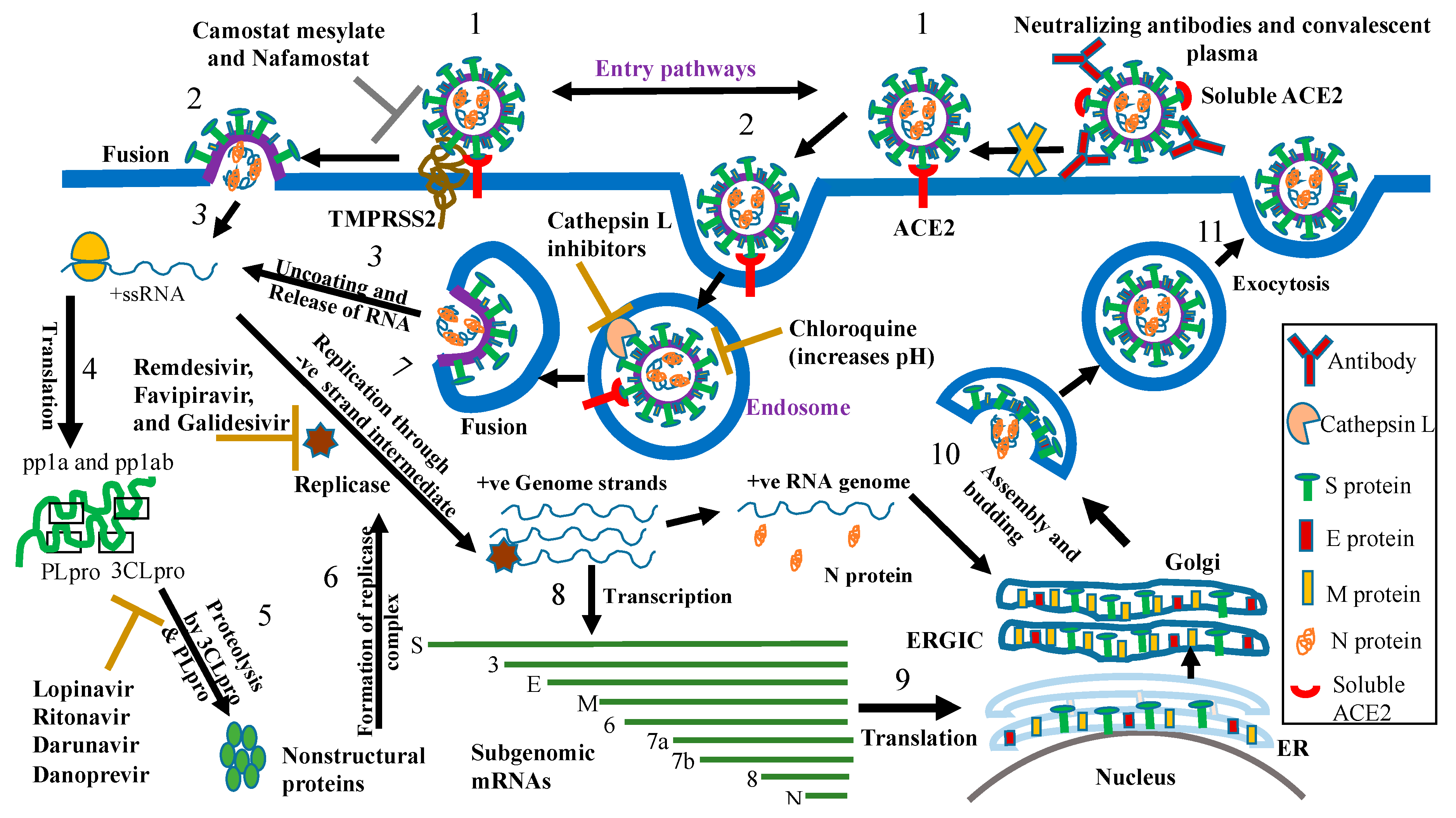
| Target of Antivirals | Example/s | Mechanism of Action |
|---|---|---|
| Viral Entry | ||
| • Spike (S) protein | Monoclonal Antibodies such as S230.15, m396, S109.8 S227.14, S230.15, 80R scFv, CR3022 CR3014, 33G4 35B5, 30F9, 4D4, IF8, 5E9, and B1 scFv | Most of the antibodies bind to S1 domain of SARS-CoV S protein and block its binding to ACE2 on target cells. Antibodies, such as 4D4, bind to regions other than RBD and block a post-attachment step in viral entry. Others (1F8, 5E9, and B1) bind to S2 domain and inhibit viral envelope-cell membrane fusion |
| Convalescent plasma | Contains neutralizing antibodies that bind to different regions on SARS-CoV S protein and inhibit viral entry by different mechanisms | |
| Lectins: Griffithsin (a lectin isolated from the red alga Griffithsia spp.) and Urtica dioica agglutinin (UDA) (a lectin isolated from Urtica dioca) | Bind to SARS-CoV S protein and inhibit binding of SARS-CoV to ACE2 | |
| Synthetic peptides derived from ACE2 and soluble ACE2 ectodomain | Bind to SARS-CoV S protein and inhibit attachment to target cells | |
| Synthetic peptides derived from HR1 or HR2 regions of S2 domain e.g. CP-1 peptide. | Bind to S2 domain and block fusion of viral envelope with cell membrane | |
| Imatinib (Abl kinase inhibitor) | Blocks fusion of viral envelope with cell membrane | |
| Chinese herbal medicine molecules tetra-O-galloyl-beta-D-glucose (TGG) and luteolin | Bind to S2 domain and inhibit fusion | |
| Emodin (a plant anthraquinone) | Binds to SARS-CoV S protein and blocks its interaction with ACE2 | |
| • Angiotensin-converting enzyme 2 (ACE2) | N-(2-aminoethyl)-1-aziridineethanamine (NAAE) | Inhibits ACE2 activity and blocks fusion |
| • Clathrin-mediated endocytosis | Chlorpromazine | Inhibits viral entry by blocking clathrin-mediated endocytosis of viral particles |
| • Endosomes | Chloroquine and P9 peptide | Raise the pH of the endosome and thus inhibit viral entry by blocking cathepsin L-mediated cleavage of S protein |
| • Cathepsin L | Small molecules such as E-64D and 5705213 | Inhibit cathepsin L and thus inhibits cathepsin L-mediated cleavage of S protein |
| • Transmembrane Serine Protease 2 (TMPRSS2) | Camostat mesylate | Blocks TMPRSS2 and thus inhibits processing of SARS-CoV S protein and viral entry at plasma membrane |
| RNA-dependent RNA polymerase (RdRp) | Nucleoside analogs: β-D-N4-hydroxycitidine, galidesivir also known as BCX4430 or immucillin-A, and remdesivir | Inhibit RdRp leading to premature termination of RNA synthesis |
| Nucleoside analogs: 6-azauridine and pyrazofurin | Inhibit orotidine monophosphate decarboxylase which leads to depletion of pyrimidine nucleosides pool | |
| Nucleoside analog: ribavirin | Ribavirin gets incorporated into the newly synthesized RNA chain resulting in viral lethal mutations. It is also believed to inhibit cellular inosine monophosphate dehydrogenase resulting in depletion of intracellular GTP which leads to inhibition of mRNA cap synthesis | |
| Viral helicase | Bananins | Inhibit ATPase activity of viral helicase and thus blocks viral replication |
| Bismuth complexes | Inhibit ATPase and unwinding activities of viral helicase | |
| Falvonoids such as myricetin and scutellarein | Inhibit ATPase activity of viral helicase. | |
| Small molecule SSYA10-001 | Inhibits unwinding activity of viral helicase | |
| Viral proteases | ||
| • Picornavirus 3-chymotrypsin-like protease (3CLpro) | HIV-1 protease inhibitors; lopinavir/ritonavir and nelfinavir | One of the mechanisms is inhibition of 3CLpro activity which leads to inhibition of processing of SARS-CoV polyproteins and blockage of viral replication. Other mechanisms involve inhibition of the cellular enzymes; calpains |
| Cinanserin (a serotonin antagonist), esculetin-4-carboxylic acid ethyl ester (a marine natural product; coumarin derivative), plant-derived compounds such as betulinic acid (sesquiterpenoid) and savinin (lignoid), quinone-methide trit-erpenes isolated from Tripterygium regelii, dieckol (a phlorotannin) isolated from brown algae Ecklonia cava, alkylated chalcones from Angelica keiskei, flavonoids such as herbacetin, rhoifolin and pectolinarin, and pyrazolone compounds | Inhibit 3CLpro and thus inhibit viral replication | |
| • Papain-like protease 2 (PLpro) | Diarylheptanoids from Alnus japonica and geranylated flavonoids from Paulownia tree, 6-mercaptopurine (6-MP), 6-thioguanine (6-TG), and disulfiram | Inhibit viral replication by blocking PLpro and processing of viral polyproteins |
| Cellular proteases: Calpain | Z-Val-Phe-Ala-CHO (calpain inhibitor III) and Val-Leu-CHO (calpain inhibitor VI) | Inhibit calpains and thus inhibit viral replication and virus-induced apoptosis |
| Miscellaneous Antivirals | E, M, N, ORF3a, ORF7a, and ORF7b siRNAs | Inhibit viral replication by inducing the breakdown of viral mRNAs |
| Hexamethylene amiloride | Blocks E protein cationic-selective ion channel activity | |
| Aurintricarboxylic acid | Inhibits SARS-CoV attachment to target cells and inhibits viral replication | |
| Rimantadine | Blocks E protein cationic-selective ion channel activity. | |
| Niclosamide | Inhibits replication of SARS-CoV. Mechanism is not well known | |
| Amiodarone | Inhibits post-entry step in viral life cycle | |
| K22 | Inhibits formation of double membrane vesicles | |
| dsRNA-activated caspase oligomerizer (DRACO) | Induces apoptosis in cells that are infected with SARS-CoV by recognizing viral dsRNA intermediates using its viral dsRNA-binding domain | |
| NO donors: 2,2′-(hydroxynitrosihydrazino)bis-ethamine (DETA NONOate) and S-nitroso-N-acetylpenicillamine (SNAP) | Dissociate to produce NO which is antiviral | |
| Cyclosporine A and its analog alisporivir | Form a complex with cyclophilins which leads to inhibition of viral replication. Also, the complex with cyclophilins inhibits calcineurin which blocks T cell activation and subsequent T cell-mediated tissue damage | |
| Geldanamycin (an antibiotic) | Binds to and inhibits heat shock protein 90 (Hsp90) which inhibits SARS-CoV replication | |
| Interferon alpha and beta | Viral RNA degradation, inhibition of protein synthesis, and viral assembly | |
| Glycyrrhizin (triterpenoid saponin glycoside from licorice root) | Inhibits adsorption of viral particles on target cells and induces production of NO. Other antiviral mechanisms need to be explored |
| Target of Antivirals | Example/s | Mechanism of Action |
|---|---|---|
| Viral entry | ||
| • S protein | Convalescent plasma | Contains neutralizing antibodies that block binding of SARS-CoV-2 S protein to ACE2 |
| Monoclonal antibodies (47D11, HA001, B38, H4, and CR3022) | Neutralize SARS-CoV-2 by binding to S protein and blocking attachment to ACE2 | |
| Umifenovir (is currently tested in clinical trials against SARS-CoV-2) | An anti-influenza drug that inhibits fusion of viral envelope with cellular membranes | |
| Soluble ACE2 | Binds to SARS-CoV-2 S protein and inhibits attachment to target cells | |
| • Endosomes | Chloroquine and hydroxychloroquine (currently tested in clinical trials) | Raise the pH of the endosomes and thus inhibit viral entry by blocking cathepsin L-mediated cleavage of S protein |
| • Cathepsin L | Small molecule E-64D and 5705213 | Inhibit cathepsin L and thus inhibits cathepsin L-mediated cleavage of S protein |
| • TMPRSS2 | Camostat mesylate (is currently evaluated in a clinical trial) and nafamostat meslylate | Block processing of SARS-CoV-2 S protein by TMPRSS2 and thus inhibits viral entry at plasma membrane |
| RdRp | Remdesivir (is currently evaluated in clinical trials), penciclovir, favipiravir (favipiravir plus interferon alpha and favipiravir plus baloxavir marboxil are currently tested in clinical trials), and galidesivir (is currently tested in clinical trials) | Inhibit RdRp leading to premature termination of RNA synthesis |
| Viral Proteases • 3CLpro | HIV protease inhibitors: lopinavir, ritonavir, darunavir and lopinavir/ritonavir combination (are currently tested in clinical trials) | Inhibition of 3CLpro activity which leads to inhibition of processing of SARS-CoV-2 polyproteins and blockage of viral replication |
| HCV protease inhibitor: danoprevir (is currently tested in a clinical trial combined with ritonavir) | Inhibition of 3CLpro activity | |
| Miscellaneous Agents | Interferon alpha and beta (is currently tested in a clinical trial combined with ribavirin and in another trial combined with lopinavir/ritonavir) | Viral RNA degradation, inhibition of protein synthesis, and viral assembly |
| Nitazoxanide | Enhances host innate interferon responses | |
| Ruxolitinib (is currently tested in clinical trials alone or combined with Simvastatin) and baricitinib | Inhibit JAK1/2 signaling and thus downregulate immune responses that are responsible for tissue damage. Barcitinib may block viral entry | |
| Eculizumab antibody (is currently tested in clinical trials) | Downregulates inflammation by binding to the complement component C5 and inhibits its cleavage by C5 convertase into C5a and C5b | |
| Tocilizumab (is currently tested in clinical trials alone or combined with favipiravir) and sarilumab | Monoclonal antibodies that block IL-6R and thus downregulate inflammation that is associated with COVID-19 disease | |
| Siltuximab (is currently tested in clinical trials) | An anti-IL-6 antibody that ameliorates inflammation and downregulates damaging immune responses. | |
| Valsartan and telmisartan (are currently tested in clinical trials) | Are angiotensin II receptor blockers that protect against harmful effects of angiotensin II in COVID-19 patients |
© 2020 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Elshabrawy, H.A. SARS-CoV-2: An Update on Potential Antivirals in Light of SARS-CoV Antiviral Drug Discoveries. Vaccines 2020, 8, 335. https://doi.org/10.3390/vaccines8020335
Elshabrawy HA. SARS-CoV-2: An Update on Potential Antivirals in Light of SARS-CoV Antiviral Drug Discoveries. Vaccines. 2020; 8(2):335. https://doi.org/10.3390/vaccines8020335
Chicago/Turabian StyleElshabrawy, Hatem A. 2020. "SARS-CoV-2: An Update on Potential Antivirals in Light of SARS-CoV Antiviral Drug Discoveries" Vaccines 8, no. 2: 335. https://doi.org/10.3390/vaccines8020335
APA StyleElshabrawy, H. A. (2020). SARS-CoV-2: An Update on Potential Antivirals in Light of SARS-CoV Antiviral Drug Discoveries. Vaccines, 8(2), 335. https://doi.org/10.3390/vaccines8020335




