Investigating the Impact of Local Manipulations on Spontaneous and Evoked Brain Complexity Indices: A Large-Scale Computational Model
Abstract
1. Introduction
2. Materials and Methods
2.1. Model Simulations
2.1.1. Structural Connectivity
2.1.2. Neural Mass Model
2.1.3. Spontaneous Activity and Stimulation Protocol
2.1.4. Local Manipulation Modeling
2.1.5. Validation on a Larger Connectome (Hconn)
2.2. Analysis of Network Simulations
2.2.1. Firing Activity Analysis
2.2.2. Network Analysis
2.2.3. Complexity Indices
- -
- Lempel–Ziv Complexity
- -
- Perturbational Complexity Index
2.2.4. Complexity Topographies
2.2.5. Statistical Analysis
3. Results
3.1. Spontaneous Activity
Impact of Local Node Silencing on Spontaneous Complexity Measures
3.2. Evoked Activity
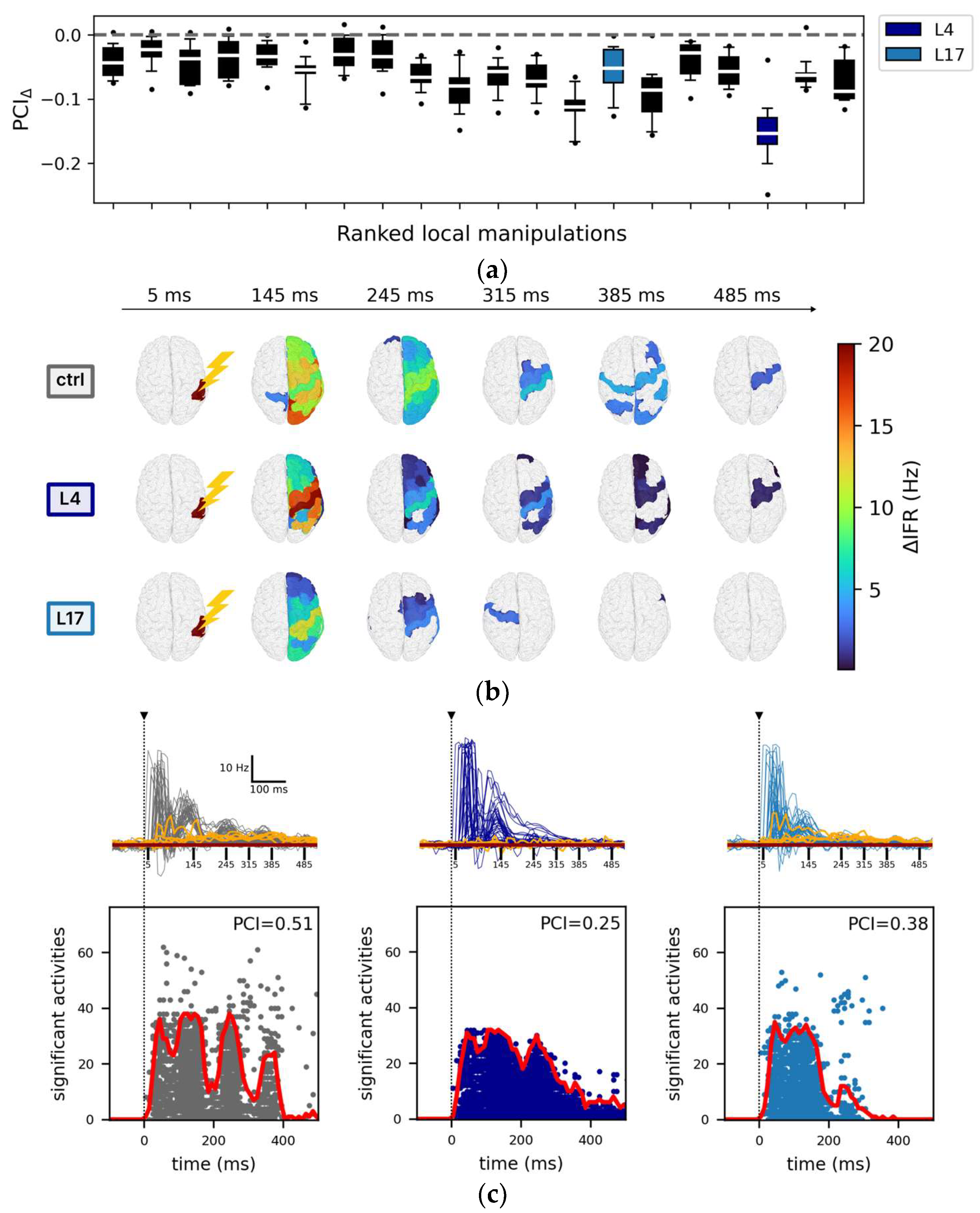
3.2.1. Local Node Silencing and PCI
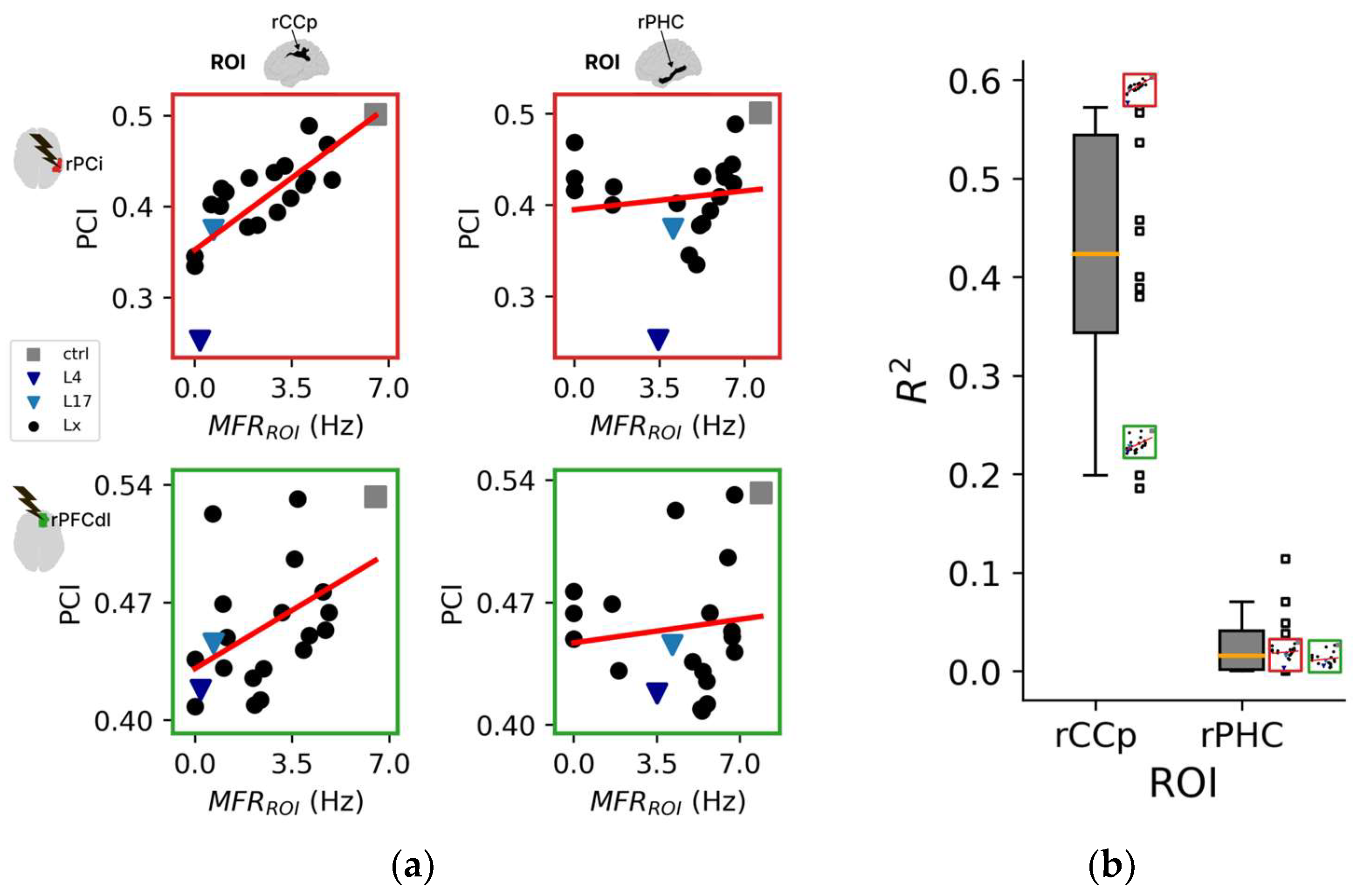
3.2.2. The Impact of Regional Silencing on PCI
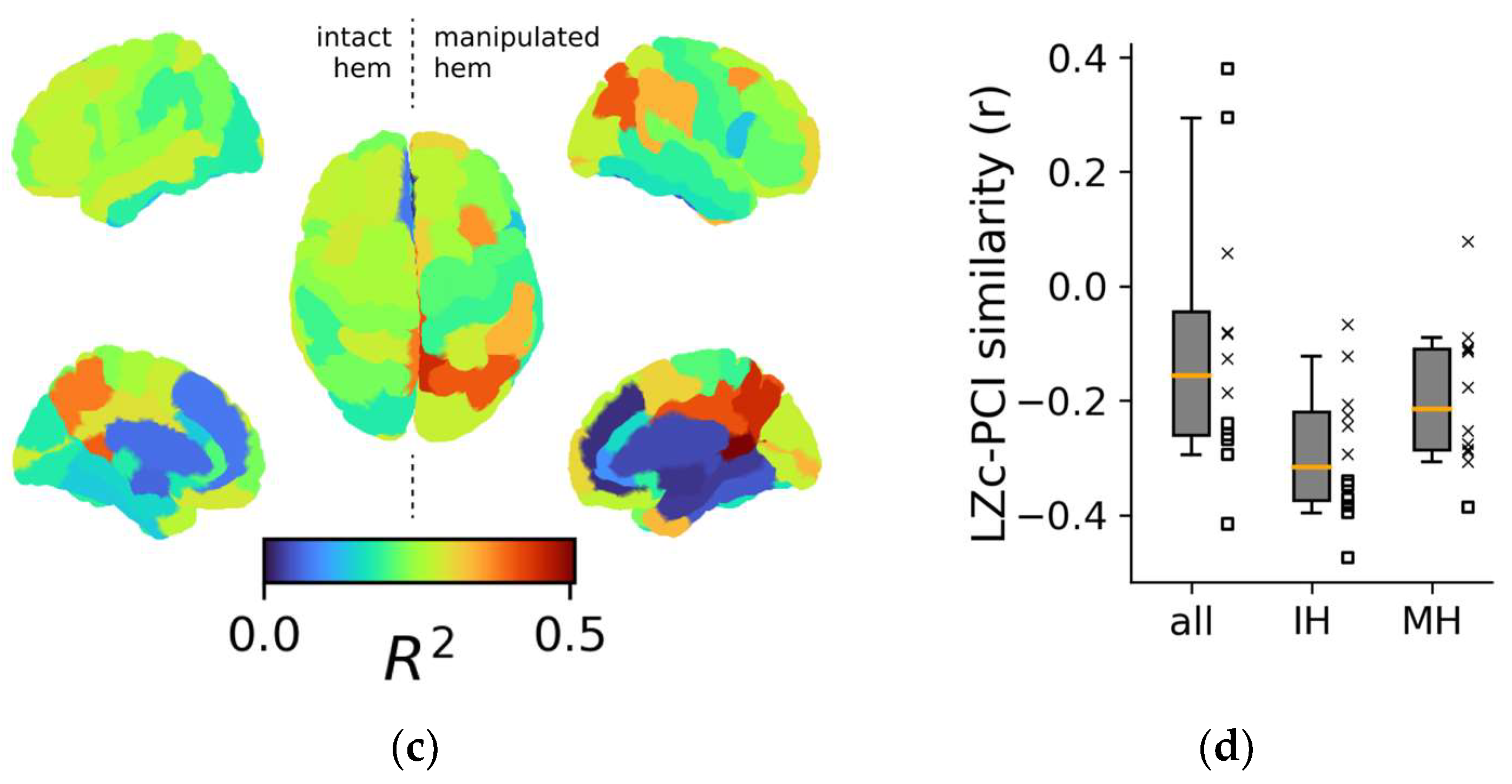
3.2.3. Assessing Regional Similarity between Spontaneous and Evoked Complexity Metrics
3.3. Assessing the Relationship between Complexity Measures and Graph Properties
4. Discussion
4.1. Impact of Local Silencing on Complexity Measures: The Role of Global Activity Levels and Network Dynamics/Interactions
4.2. Regional Aspects of Complexity Indices: The Role of Posterior Regions
4.3. Limitations and Future Directions
4.4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Tononi, G.; Sporns, O.; Edelman, G.M. A Measure for Brain Complexity: Relating Functional Segregation and Integration in the Nervous System. Proc. Natl. Acad. Sci. USA 1994, 91, 5033–5037. [Google Scholar] [CrossRef] [PubMed]
- Deco, G.; Tononi, G.; Boly, M.; Kringelbach, M.L. Rethinking Segregation and Integration: Contributions of Whole-Brain Modelling. Nat. Rev. Neurosci. 2015, 16, 430–439. [Google Scholar] [CrossRef]
- Bullmore, E.; Sporns, O. The Economy of Brain Network Organization. Nat. Rev. Neurosci. 2012, 13, 336–349. [Google Scholar] [CrossRef] [PubMed]
- Sporns, O.; Tononi, G.; Edelman, G.M. Connectivity and Complexity: The Relationship between Neuroanatomy and Brain Dynamics. Neural Netw. 2000, 13, 909–922. [Google Scholar] [CrossRef] [PubMed]
- Sporns, O. The Complex Brain: Connectivity, Dynamics, Information. Trends Cogn. Sci. 2022, 26, 1066–1067. [Google Scholar] [CrossRef] [PubMed]
- Sarasso, S.; Casali, A.G.; Casarotto, S.; Rosanova, M.; Sinigaglia, C.; Massimini, M. Consciousness and Complexity: A Consilience of Evidence. Neurosci. Conscious. 2021, 7, niab023. [Google Scholar] [CrossRef]
- Tononi, G.; Edelman, G.M. Consciousness and Complexity. Science 1998, 282, 1846–1851. [Google Scholar] [CrossRef]
- Hagmann, P. From Diffusion MRI to Brain Connectomics; EPFL: Lausanne, Switzerland, 2005. [Google Scholar]
- Sporns, O.; Tononi, G.; Kötter, R. The Human Connectome: A Structural Description of the Human Brain. PLoS Comput. Biol. 2005, 1, e42. [Google Scholar] [CrossRef]
- Van Essen, D.C.; Smith, S.M.; Barch, D.M.; Behrens, T.E.J.; Yacoub, E.; Ugurbil, K. The WU-Minn Human Connectome Project: An Overview. NeuroImage 2013, 80, 62–79. [Google Scholar] [CrossRef]
- Miller, K.L.; Alfaro-Almagro, F.; Bangerter, N.K.; Thomas, D.L.; Yacoub, E.; Xu, J.; Bartsch, A.J.; Jbabdi, S.; Sotiropoulos, S.N.; Andersson, J.L.R.; et al. Multimodal Population Brain Imaging in the UK Biobank Prospective Epidemiological Study. Nat. Neurosci. 2016, 19, 1523–1536. [Google Scholar] [CrossRef]
- Kaiser, M. Connectomes: From a Sparsity of Networks to Large-Scale Databases. Front. Neuroinform. 2023, 17, 1170337. [Google Scholar] [CrossRef] [PubMed]
- Sanz Leon, P.; Knock, S.; Woodman, M.; Domide, L.; Mersmann, J.; McIntosh, A.; Jirsa, V. The Virtual Brain: A Simulator of Primate Brain Network Dynamics. Front. Neuroinform. 2013, 7, 10. [Google Scholar] [CrossRef]
- Breakspear, M. Dynamic Models of Large-Scale Brain Activity. Nat. Neurosci. 2017, 20, 340–352. [Google Scholar] [CrossRef] [PubMed]
- Cakan, C.; Jajcay, N.; Obermayer, K. Neurolib: A Simulation Framework for Whole-Brain Neural Mass Modeling. Cogn. Comput. 2021, 15, 1132–1152. [Google Scholar] [CrossRef]
- Lord, L.-D.; Stevner, A.B.; Deco, G.; Kringelbach, M.L. Understanding Principles of Integration and Segregation Using Whole-Brain Computational Connectomics: Implications for Neuropsychiatric Disorders. Philos. Trans. R. Soc. A Math. Phys. Eng. Sci. 2017, 375, 20160283. [Google Scholar] [CrossRef]
- Casali, A.G.; Gosseries, O.; Rosanova, M.; Boly, M.; Sarasso, S.; Casali, K.R.; Casarotto, S.; Bruno, M.-A.; Laureys, S.; Tononi, G.; et al. A Theoretically Based Index of Consciousness Independent of Sensory Processing and Behavior. Sci. Transl. Med. 2013, 5, 198ra105. [Google Scholar] [CrossRef]
- Casarotto, S.; Comanducci, A.; Rosanova, M.; Sarasso, S.; Fecchio, M.; Napolitani, M.; Pigorini, A.G.; Casali, A.; Trimarchi, P.D.; Boly, M.; et al. Stratification of Unresponsive Patients by an Independently Validated Index of Brain Complexity. Ann. Neurol. 2016, 80, 718–729. [Google Scholar] [CrossRef]
- Timme, N.M.; Marshall, N.J.; Bennett, N.; Ripp, M.; Lautzenhiser, E.; Beggs, J.M. Criticality Maximizes Complexity in Neural Tissue. Front. Physiol. 2016, 7, 425. [Google Scholar] [CrossRef]
- Colombi, I.; Nieus, T.; Massimini, M.; Chiappalone, M. Spontaneous and Perturbational Complexity in Cortical Cultures. Brain Sci. 2021, 11, 1453. [Google Scholar] [CrossRef]
- Barbero-Castillo, A.; Mateos-Aparicio, P.; Porta, L.D.; Camassa, A.; Perez-Mendez, L.; Sanchez-Vives, M.V. Impact of GABAA and GABAB Inhibition on Cortical Dynamics and Perturbational Complexity during Synchronous and Desynchronized States. J. Neurosci. 2021, 41, 5029–5044. [Google Scholar] [CrossRef]
- Arena, A.; Comolatti, R.; Thon, S.; Casali, A.G.; Storm, J.F. General Anesthesia Disrupts Complex Cortical Dynamics in Response to Intracranial Electrical Stimulation in Rats. eNeuro 2021, 8. [Google Scholar] [CrossRef] [PubMed]
- Dasilva, M.; Camassa, A.; Navarro-Guzman, A.; Pazienti, A.; Perez-Mendez, L.; Zamora-López, G.; Mattia, M.; Sanchez-Vives, M.V. Modulation of Cortical Slow Oscillations and Complexity across Anesthesia Levels. NeuroImage 2021, 224, 117415. [Google Scholar] [CrossRef] [PubMed]
- Cavelli, M.L.; Mao, R.; Findlay, G.; Driessen, K.; Bugnon, T.; Tononi, G.; Cirelli, C. Sleep/Wake Changes in Perturbational Complexity in Rats and Mice. iScience 2023, 26, 106186. [Google Scholar] [CrossRef] [PubMed]
- Massimini, M.; Ferrarelli, F.; Huber, R.; Esser, S.K.; Singh, H.; Tononi, G. Breakdown of Cortical Effective Connectivity During Sleep. Science 2005, 309, 2228–2232. [Google Scholar] [CrossRef] [PubMed]
- Ragazzoni, A.; Pirulli, C.; Veniero, D.; Feurra, M.; Cincotta, M.; Giovannelli, F.; Chiaramonti, R.; Lino, M.; Rossi, S.; Miniussi, C. Vegetative versus Minimally Conscious States: A Study Using TMS-EEG, Sensory and Event-Related Potentials. PLoS ONE 2013, 8, e57069. [Google Scholar] [CrossRef] [PubMed]
- Sarasso, S.; D’Ambrosio, S.; Fecchio, M.; Casarotto, S.; Viganò, A.; Landi, C.; Mattavelli, G.; Gosseries, O.; Quarenghi, M.; Laureys, S.; et al. Local Sleep-like Cortical Reactivity in the Awake Brain after Focal Injury. Brain 2020, 143, 3672–3684. [Google Scholar] [CrossRef] [PubMed]
- Sinitsyn, D.O.; Poydasheva, A.G.; Bakulin, I.S.; Legostaeva, L.A.; Iazeva, E.G.; Sergeev, D.V.; Sergeeva, A.N.; Kremneva, E.I.; Morozova, S.N.; Lagoda, D.Y.; et al. Detecting the Potential for Consciousness in Unresponsive Patients Using the Perturbational Complexity Index. Brain Sci. 2020, 10, 917. [Google Scholar] [CrossRef]
- Usami, K.; Korzeniewska, A.; Matsumoto, R.; Kobayashi, K.; Hitomi, T.; Matsuhashi, M.; Kunieda, T.; Mikuni, N.; Kikuchi, T.; Yoshida, K.; et al. The Neural Tides of Sleep and Consciousness Revealed by Single-Pulse Electrical Brain Stimulation. Sleep 2019, 42, zsz050. [Google Scholar] [CrossRef]
- Spiegler, A.; Hansen, E.C.A.; Bernard, C.; McIntosh, A.R.; Jirsa, V.K. Selective Activation of Resting-State Networks Following Focal Stimulation in a Connectome-Based Network Model of the Human Brain. eNeuro 2016, 3. [Google Scholar] [CrossRef]
- Spiegler, A.; Abadchi, J.K.; Mohajerani, M.; Jirsa, V.K. In Silico Exploration of Mouse Brain Dynamics by Focal Stimulation Reflects the Organization of Functional Networks and Sensory Processing. Netw. Neurosci. 2020, 4, 807–851. [Google Scholar] [CrossRef]
- Momi, D.; Wang, Z.; Griffiths, J.D. TMS-Evoked Responses Are Driven by Recurrent Large-Scale Network Dynamics. Elife 2023, 12, e83232. [Google Scholar] [CrossRef] [PubMed]
- Goldman, J.S.; Kusch, L.; Aquilue, D.; Yalçınkaya, B.H.; Depannemaecker, D.; Ancourt, K.; Nghiem, T.-A.E.; Jirsa, V.; Destexhe, A. A Comprehensive Neural Simulation of Slow-Wave Sleep and Highly Responsive Wakefulness Dynamics. Front. Comput. Neurosci. 2023, 16, 1058957. [Google Scholar] [CrossRef] [PubMed]
- Cattani, A.; Galluzzi, A.; Fecchio, M.; Pigorini, A.; Mattia, M.; Massimini, M. Adaptation Shapes Local Cortical Reactivity: From Bifurcation Diagram and Simulations to Human Physiological and Pathological Responses. eNeuro 2023, 10. [Google Scholar] [CrossRef] [PubMed]
- Albert, R.; Jeong, H.; Barabási, A.-L. Error and Attack Tolerance of Complex Networks. Nature 2000, 406, 378–382. [Google Scholar] [CrossRef] [PubMed]
- Seguin, C.; Sporns, O.; Zalesky, A. Brain Network Communication: Concepts, Models and Applications. Nat. Rev. Neurosci. 2023, 24, 557–574. [Google Scholar] [CrossRef] [PubMed]
- Aerts, H.; Fias, W.; Caeyenberghs, K.; Marinazzo, D. Brain Networks under Attack: Robustness Properties and the Impact of Lesions. Brain 2016, 139, 3063–3083. [Google Scholar] [CrossRef]
- Rabuffo, G.; Lokossou, H.-A.; Li, Z.; Ziaee-Mehr, A.; Hashemi, M.; Quilichini, P.P.; Ghestem, A.; Arab, O.; Esclapez, M.; Verma, P.; et al. Probing the Mechanisms of Global Brain Reconfiguration after Local Manipulations. bioRxiv 2023, 2023, 2023-09. [Google Scholar]
- Alstott, J.; Breakspear, M.; Hagmann, P.; Cammoun, L.; Sporns, O. Modeling the Impact of Lesions in the Human Brain. PLOS Comput. Biol. 2009, 5, e1000408. [Google Scholar] [CrossRef]
- Hahn, G.; Zamora-López, G.; Uhrig, L.; Tagliazucchi, E.; Laufs, H.; Mantini, D.; Kringelbach, M.L.; Jarraya, B.; Deco, G. Signature of Consciousness in Brain-Wide Synchronization Patterns of Monkey and Human fMRI Signals. Neuroimage 2021, 226, 117470. [Google Scholar] [CrossRef]
- Koch, C.; Massimini, M.; Boly, M.; Tononi, G. Neural Correlates of Consciousness: Progress and Problems. Nat. Rev. Neurosci. 2016, 17, 307–321. [Google Scholar] [CrossRef]
- Sanders, R.D.; Mostert, N.; Lindroth, H.; Tononi, G.; Sleigh, J. Is Consciousness Frontal? Two Perioperative Case Reports That Challenge That Concept. Br. J. Anaesth. 2018, 121, 330–332. [Google Scholar] [CrossRef] [PubMed]
- Sanz-Leon, P.; Knock, S.A.; Spiegler, A.; Jirsa, V.K. Mathematical Framework for Large-Scale Brain Network Modeling in The Virtual Brain. NeuroImage 2015, 111, 385–430. [Google Scholar] [CrossRef] [PubMed]
- Schartner, M.; Seth, A.; Noirhomme, Q.; Boly, M.; Bruno, M.-A.; Laureys, S.; Barrett, A. Complexity of Multi-Dimensional Spontaneous EEG Decreases during Propofol Induced General Anaesthesia. PLoS ONE 2015, 10, e0133532. [Google Scholar] [CrossRef] [PubMed]
- Bullmore, E.; Sporns, O. Complex Brain Networks: Graph Theoretical Analysis of Structural and Functional Systems. Nat. Rev. Neurosci. 2009, 10, 186–198. [Google Scholar] [CrossRef]
- Woodman, M.M.; Pezard, L.; Domide, L.; Knock, S.A.; Sanz-Leon, P.; Mersmann, J.; McIntosh, A.R.; Jirsa, V. Integrating Neuroinformatics Tools in TheVirtualBrain. Front. Neuroinform. 2014, 8, 36. [Google Scholar] [CrossRef]
- Kunze, T.; Hunold, A.; Haueisen, J.; Jirsa, V.; Spiegler, A. Transcranial Direct Current Stimulation Changes Resting State Functional Connectivity: A Large-Scale Brain Network Modeling Study. NeuroImage 2016, 140, 174–187. [Google Scholar] [CrossRef]
- Hagmann, P.; Cammoun, L.; Gigandet, X.; Meuli, R.; Honey, C.J.; Wedeen, V.J.; Sporns, O. Mapping the Structural Core of Human Cerebral Cortex. PLOS Biol. 2008, 6, e159. [Google Scholar] [CrossRef]
- Breakspear, M.; Terry, J.R.; Friston, K.J. Modulation of Excitatory Synaptic Coupling Facilitates Synchronization and Complex Dynamics in a Biophysical Model of Neuronal Dynamics. Network 2003, 14, 703–732. [Google Scholar] [CrossRef]
- Honey, C.J.; Kötter, R.; Breakspear, M.; Sporns, O. Network Structure of Cerebral Cortex Shapes Functional Connectivity on Multiple Time Scales. Proc. Natl. Acad. Sci. USA 2007, 104, 10240–10245. [Google Scholar] [CrossRef]
- Honey, C.J.; Sporns, O.; Cammoun, L.; Gigandet, X.; Thiran, J.P.; Meuli, R.; Hagmann, P. Predicting Human Resting-State Functional Connectivity from Structural Connectivity. Proc. Natl. Acad. Sci. USA 2009, 106, 2035–2040. [Google Scholar] [CrossRef]
- Gollo, L.L.; Breakspear, M. The Frustrated Brain: From Dynamics on Motifs to Communities and Networks. Philos. Trans. R. Soc. B Biol. Sci. 2014, 369, 20130532. [Google Scholar] [CrossRef] [PubMed]
- Gollo, L.L.; Mirasso, C.; Sporns, O.; Breakspear, M. Mechanisms of Zero-Lag Synchronization in Cortical Motifs. PLOS Comput. Biol. 2014, 10, e1003548. [Google Scholar] [CrossRef] [PubMed]
- Gollo, L.L.; Zalesky, A.; Hutchison, R.M.; van den Heuvel, M.; Breakspear, M. Dwelling Quietly in the Rich Club: Brain Network Determinants of Slow Cortical Fluctuations. Philos. Trans. R. Soc. B Biol. Sci. 2015, 370, 20140165. [Google Scholar] [CrossRef] [PubMed]
- Roberts, J.A.; Friston, K.J.; Breakspear, M. Clinical Applications of Stochastic Dynamic Models of the Brain, Part I: A Primer. Biol. Psychiatry Cogn. Neurosci. Neuroimaging 2017, 2, 216–224. [Google Scholar] [CrossRef] [PubMed]
- Roberts, J.A.; Gollo, L.L.; Abeysuriya, R.G.; Roberts, G.; Mitchell, P.B.; Woolrich, M.W.; Breakspear, M. Metastable Brain Waves. Nat. Commun. 2019, 10, 1056. [Google Scholar] [CrossRef] [PubMed]
- Endo, H.; Hiroe, N.; Yamashita, O. Evaluation of Resting Spatio-Temporal Dynamics of a Neural Mass Model Using Resting fMRI Connectivity and EEG Microstates. Front. Comput. Neurosci. 2020, 13, 91. [Google Scholar] [CrossRef] [PubMed]
- Aquino, K.M.; Fulcher, B.; Oldham, S.; Parkes, L.; Gollo, L.; Deco, G.; Fornito, A. On the Intersection between Data Quality and Dynamical Modelling of Large-Scale fMRI Signals. NeuroImage 2022, 256, 119051. [Google Scholar] [CrossRef]
- Honey, C.J.; Sporns, O. Dynamical Consequences of Lesions in Cortical Networks. Hum. Brain Mapp. 2008, 29, 802–809. [Google Scholar] [CrossRef]
- Deco, G.; McIntosh, A.R.; Shen, K.; Hutchison, R.M.; Menon, R.S.; Everling, S.; Hagmann, P.; Jirsa, V.K. Identification of Optimal Structural Connectivity Using Functional Connectivity and Neural Modeling. J. Neurosci. 2014, 34, 7910–7916. [Google Scholar] [CrossRef]
- Adhikari, M.H.; Raja Beharelle, A.; Griffa, A.; Hagmann, P.; Solodkin, A.; McIntosh, A.R.; Small, S.L.; Deco, G. Computational Modeling of Resting-State Activity Demonstrates Markers of Normalcy in Children with Prenatal or Perinatal Stroke. J. Neurosci. 2015, 35, 8914–8924. [Google Scholar] [CrossRef]
- Tagliazucchi, E. The Signatures of Conscious Access and Its Phenomenology Are Consistent with Large-Scale Brain Communication at Criticality. Conscious. Cogn. 2017, 55, 136–147. [Google Scholar] [CrossRef] [PubMed]
- Eryilmaz, H.; Pax, M.; O’Neill, A.G.; Vangel, M.; Diez, I.; Holt, D.J.; Camprodon, J.A.; Sepulcre, J.; Roffman, J.L. Network Hub Centrality and Working Memory Performance in Schizophrenia. Schizophrenia 2022, 8, 76. [Google Scholar] [CrossRef] [PubMed]
- Friston, K.J.; Harrison, L.; Penny, W. Dynamic Causal Modelling. NeuroImage 2003, 19, 1273–1302. [Google Scholar] [CrossRef] [PubMed]
- Farnes, N.; Juel, B.E.; Nilsen, A.S.; Romundstad, L.G.; Storm, J.F. Increased Signal Diversity/Complexity of Spontaneous EEG, but Not Evoked EEG Responses, in Ketamine-Induced Psychedelic State in Humans. PLoS ONE 2020, 15, e0242056. [Google Scholar] [CrossRef] [PubMed]
- Ort, A.; Smallridge, J.W.; Sarasso, S.; Casarotto, S.; von Rotz, R.; Casanova, A.; Seifritz, E.; Preller, K.H.; Tononi, G.; Vollenweider, F.X. TMS-EEG and Resting-State EEG Applied to Altered States of Consciousness: Oscillations, Complexity, and Phenomenology. iScience 2023, 26. [Google Scholar] [CrossRef] [PubMed]
- Ferrarelli, F.; Massimini, M.; Sarasso, S.; Casali, A.; Riedner, B.A.; Angelini, G.; Tononi, G.; Pearce, R.A. Breakdown in Cortical Effective Connectivity during Midazolam-Induced Loss of Consciousness. Proc. Natl. Acad. Sci. USA 2010, 107, 2681–2686. [Google Scholar] [CrossRef] [PubMed]
- Bai, Y.; Xia, X.; Kang, J.; Yin, X.; Yang, Y.; He, J.; Li, X. Evaluating the Effect of Repetitive Transcranial Magnetic Stimulation on Disorders of Consciousness by Using TMS-EEG. Front. Neurosci. 2016, 10, 473. [Google Scholar] [CrossRef]
- King, J.-R.; Sitt, J.D.; Faugeras, F.; Rohaut, B.; El Karoui, I.; Cohen, L.; Naccache, L.; Dehaene, S. Information Sharing in the Brain Indexes Consciousness in Noncommunicative Patients. Curr. Biol. 2013, 23, 1914–1919. [Google Scholar] [CrossRef]
- Sitt, J.D.; King, J.-R.; El Karoui, I.; Rohaut, B.; Faugeras, F.; Gramfort, A.; Cohen, L.; Sigman, M.; Dehaene, S.; Naccache, L. Large Scale Screening of Neural Signatures of Consciousness in Patients in a Vegetative or Minimally Conscious State. Brain 2014, 137, 2258–2270. [Google Scholar] [CrossRef]
- Luppi, A.I.; Craig, M.M.; Pappas, I.; Finoia, P.; Williams, G.B.; Allanson, J.; Pickard, J.D.; Owen, A.M.; Naci, L.; Menon, D.K.; et al. Consciousness-Specific Dynamic Interactions of Brain Integration and Functional Diversity. Nat. Commun. 2019, 10, 4616. [Google Scholar] [CrossRef]
- Arena, A.; Juel, B.E.; Comolatti, R.; Thon, S.; Storm, J.F. Capacity for Consciousness under Ketamine Anaesthesia Is Selectively Associated with Activity in Posteromedial Cortex in Rats. Neurosci. Conscious. 2022, 2022, niac004. [Google Scholar] [CrossRef] [PubMed]
- Bensaid, S.; Modolo, J.; Merlet, I.; Wendling, F.; Benquet, P. COALIA: A Computational Model of Human EEG for Consciousness Research. Front. Syst. Neurosci. 2019, 13, 59. [Google Scholar] [CrossRef] [PubMed]
- Cabrera-Álvarez, J.; Doorn, N.; Maestú, F.; Susi, G. Modeling the Role of the Thalamus in Resting-State Functional Connectivity: Nature or Structure. PLoS Comput. Biol. 2023, 19, e1011007. [Google Scholar] [CrossRef] [PubMed]
- Næss, S.; Halnes, G.; Hagen, E.; Hagler, D.J.; Dale, A.M.; Einevoll, G.T.; Ness, T.V. Biophysically Detailed Forward Modeling of the Neural Origin of EEG and MEG Signals. NeuroImage 2021, 225, 117467. [Google Scholar] [CrossRef]
- Spiegler, A.; Jirsa, V. Systematic Approximations of Neural Fields through Networks of Neural Masses in the Virtual Brain. NeuroImage 2013, 83, 704–725. [Google Scholar] [CrossRef] [PubMed]
- Cakan, C.; Obermayer, K. Biophysically Grounded Mean-Field Models of Neural Populations under Electrical Stimulation. PLoS Comput. Biol. 2020, 16, e1007822. [Google Scholar] [CrossRef]
- Walter, W.G. The Electro-Encephalogram in Cases of Cerebral Tumour. Proc. R. Soc. Med. 1937, 30, 579–598. [Google Scholar] [CrossRef]
- Nuwer, M.R.; Jordan, S.E.; Ahn, S.S. Evaluation of Stroke Using EEG Frequency Analysis and Topographic Mapping. Neurology 1987, 37, 1153–1159. [Google Scholar] [CrossRef] [PubMed]
- Russo, S.; Pigorini, A.; Mikulan, E.; Sarasso, S.; Rubino, A.; Zauli, F.M.; Parmigiani, S.; d’Orio, P.; Cattani, A.; Francione, S.; et al. Focal Lesions Induce Large-Scale Percolation of Sleep-like Intracerebral Activity in Awake Humans. NeuroImage 2021, 234, 117964. [Google Scholar] [CrossRef]
- Tscherpel, C.; Dern, S.; Hensel, L.; Ziemann, U.; Fink, G.R.; Grefkes, C. Brain Responsivity Provides an Individual Readout for Motor Recovery after Stroke. Brain 2020, 143, 1873–1888. [Google Scholar] [CrossRef]
- D’Ambrosio, S.; Certo, F.; Bernardelli, L.; Pini, L.; Corbetta, M.; Pantoni, L.; Massimini, M.; Sarasso, S. Detecting Cortical Reactivity Alterations Induced by Structural Disconnection in Subcortical Stroke. Clin. Neurophysiol. 2023, 156, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Cakan, C.; Dimulescu, C.; Khakimova, L.; Obst, D.; Flöel, A.; Obermayer, K. Spatiotemporal Patterns of Adaptation-Induced Slow Oscillations in a Whole-Brain Model of Slow-Wave Sleep. Front. Comput. Neurosci. 2022, 15, 800101. [Google Scholar] [CrossRef] [PubMed]



| Label | Description |
|---|---|
| A1 | Primary auditory cortex |
| A2 | Secondary auditory cortex |
| Amyg | Amygdala |
| CCa | Anterior cingulate cortex |
| CCp | Posterior cingulate cortex |
| CCr | Retrosplenial cingulate cortex |
| CCs | Subgenual cingulate cortex |
| FEF | Frontal eye field |
| G | Gustatory area |
| HC | Hippocampal cortex |
| IA | Anterior insula |
| IP | Posterior insula |
| M1 | Primary motor area |
| PCi | Inferior parietal cortex |
| PCip | Cortex of intraparietal sulcus |
| PCm | Medial parietal cortex (i.e. precuneus) |
| PCs | Dorsal parietal cortex (superior parietal lobule) |
| PFCcl | Centrolateral prefrontal cortex |
| PFCdl | Dorsolateral prefrontal cortex |
| PFCdm | Dorsomedial prefrontal cortex |
| PFCm | Medial prefrontal cortex |
| PFCorb | Orbitofrontal cortex |
| PFCpol | Pole of prefrontal cortex |
| PFCvl | Ventrolateral prefrontal cortex |
| PHC | Parahippocampal cortex |
| PMCdl | Dorsolateral premotor cortex |
| PMCm | Medial premotor cortex (i.e. supplementary motor area) |
| PMCvl | Ventrolateral premotor cortex |
| S1 | Primary somatosensory cortex |
| S2 | Secondary somatosensory cortex |
| TCc | Central temporal cortex |
| TCi | Inferior temporal cortex |
| TCpol | Pole of temporal cortex |
| TCs | Superior temporal cortex |
| TCv | Ventral temporal cortex |
| V1 | Primary visual cortex |
| V2 | Secondary visual cortex |
| Local Manipulation Label | Nodes | Ranking (MFRΔ) |
|---|---|---|
| L1 | PCip, PCs, PCm | 10 |
| L2 | PFCcl, PFCdl, PMCvl | 7 |
| L3 | TCc, TCs, TCi | 4 |
| L4 | V2, V1, CCr | 18 |
| L5 | CCs, PFCorb, Amyg | 19 |
| L6 | CCa, CCp, CCr | 13 |
| L7 | M1, S1, PMCdl | 5 |
| L8 | PCm, PCip, CCp | 15 |
| L9 | PFCdm, CCa, PFCm | 11 |
| L10 | A1, A2, S2 | 16 |
| L11 | Amyg, PHC, CCs | 17 |
| L12 | HC, PHC, TCv | 2 |
| L13 | PFCpol, PFCm, PFCdm | 9 |
| L14 | PFCvl, G, PMCvl | 8 |
| L15 | TCpol, Amyg, PFCorb | 20 |
| L16 | CCa, PFCdm, PFCdl | 12 |
| L17 | IP, TCs, S2 | 14 |
| L18 | S1, M1, PCs | 3 |
| L19 | PFCdl, CCa, PFCcl | 6 |
| L20 | TCv, PHC, TCi | 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gaglioti, G.; Nieus, T.R.; Massimini, M.; Sarasso, S. Investigating the Impact of Local Manipulations on Spontaneous and Evoked Brain Complexity Indices: A Large-Scale Computational Model. Appl. Sci. 2024, 14, 890. https://doi.org/10.3390/app14020890
Gaglioti G, Nieus TR, Massimini M, Sarasso S. Investigating the Impact of Local Manipulations on Spontaneous and Evoked Brain Complexity Indices: A Large-Scale Computational Model. Applied Sciences. 2024; 14(2):890. https://doi.org/10.3390/app14020890
Chicago/Turabian StyleGaglioti, Gianluca, Thierry Ralph Nieus, Marcello Massimini, and Simone Sarasso. 2024. "Investigating the Impact of Local Manipulations on Spontaneous and Evoked Brain Complexity Indices: A Large-Scale Computational Model" Applied Sciences 14, no. 2: 890. https://doi.org/10.3390/app14020890
APA StyleGaglioti, G., Nieus, T. R., Massimini, M., & Sarasso, S. (2024). Investigating the Impact of Local Manipulations on Spontaneous and Evoked Brain Complexity Indices: A Large-Scale Computational Model. Applied Sciences, 14(2), 890. https://doi.org/10.3390/app14020890





