Simple Summary
Salmonella is an important pathogen in both livestock and humans, therefore much research has been conducted aiming to improve control methods and consequently reduce prevalence. Salmonella monitoring systems have been implemented in swine slaughter plants across Germany to test meat juices or blood of finisher animals being processed. The farms are placed into categories according to the level of Salmonella positivity, which then dictate actions required to be taken by the farmer as well as any price penalties applied to the meat sold. Since the implementation of this monitoring system, the distribution of farms across categories has not drastically altered, indicating that the national Salmonella situation is not necessarily improving. More action seems to be required at the breeding and multiplying levels of production rather than simply monitoring the finisher farms. Improving the diagnostic methods at these early stages of production in sows and nursery pigs, and consequently the control of Salmonella, would aid improvement of the Salmonella situation across the whole pig stock across Germany, including finishing farms. Sock and swab sampling has been demonstrated as a simple, cheap and effective sampling method for other enteric pathogens and therefore may be a useful tool for the diagnosis of Salmonella on swine farms. Availability of an easily accessible diagnostic method for Salmonella may increase investigations into the pathogen on sow farms and consequently improve the control methods implemented and reduce Salmonella positivity at slaughter.
Abstract
Salmonellosis is the second most reported gastrointestinal infection in humans after campylobacteriosis and a common cause of foodborne outbreaks in the European Union (EU). In addition to consumption of contaminated animal-based foods, such as poultry, beef and eggs, pork is an important source of human salmonellosis outbreaks; therefore, Salmonella (S.) control should start in the early stages of pig production. To be able to implement effective control measures to reduce the risk of pigs being infected by Salmonella, it is important to identify the serovars circulating on farm within the different stages of production, including as early as sow and piglet breeding. The aim of the present study was to assess the Salmonella status of sow farms either producing their own finishers or delivering piglets to fattening farms with a known high serological prevalence identified within the QS Salmonella monitoring system. Overall, 97 (92.4%) of 105 investigated piglet-producing farms across Germany tested positive in at least one sample. Salmonella was detected in 38.2% of the sock and 27.1% of the environmental swab samples. S. Typhimurium was the most frequent serovar. In conclusion, sock and environmental swab samples are well suited for non-invasive Salmonella detection in different production units in farrowing farms. To establish a holistic Salmonella control program, all age classes of pig production should be sampled to enable intervention and implementation of countermeasures at an early stage if necessary.
1. Introduction
Salmonella (S.) is a Gram-negative, facultative anaerobic rod-shaped bacterium, which can cause diarrhea and septicemia [1]. More than 2500 serovars [2] with different virulence levels and host adaptations exist [3]. According to the EFSA European Union One Health 2020 Zoonoses Report published in 2021, Salmonella remains one of the most frequently reported causative agents of foodborne outbreaks [4]. The most common sources of salmonellosis outbreaks are eggs, egg products and pig meat, with the most commonly involved serovar being S. Enteritidis, followed by S. Typhimurium and its monophasic variants [5]. Due to the leading role of pigs as a source of Salmonella introduction into the food chain, different control programs for Salmonella in pigs have been implemented in Europe [6]. A huge amount of research has been conducted worldwide, focusing on the epidemiology and control of Salmonella in pigs [7,8,9,10,11]. The EFSA Annual Report 2008 highlighted the impact of the Salmonella status of breeding herds, demonstrating that countries with a low Salmonella prevalence in breeding herds tended to have relatively lower prevalence in slaughter pigs compared to countries with high breeding herd prevalence [12]. Salmonella positivity in fatteners was also impacted by the prevalence of Salmonella in the breeding herd [12,13], emphasizing the importance of the control of breeding herds supplying fatteners.
Salmonella diagnosis is often very challenging due to shedding pattern variation between different serovars and differences in the infectious dose received by the animal [14,15], as well as the intermittent shedding of carriers [16]. In Germany, a non-governmental serological Salmonella monitoring system in pigs intended for human consumption was introduced by the German Quality Assurance System (QS) (https://www.q-s.de/en/, accessed on 3 December 2022) in 2002 [17]. The Salmonella status of a fattening farm is determined quarterly by testing either meat juice/blood samples taken at the slaughterhouse or blood samples taken from fatteners up to 14 days before slaughter [18]. Every year, 60 samples per farm (15 per quarter) are examined with one of three validated Salmonella ELISA test kits, using a standardized cut-off at OD 40% [17]. Based on the antibody determination of these samples, the farms are divided into three categories based on the risk of Salmonella introduction into the slaughterhouse: category one (low risk) 0–20% positive samples; category two (medium risk) 20–40%; and category three (high risk) more than 40% positive samples. Farms in category three must identify sources of Salmonella entry and initiate control measures in consultation with their farm veterinarian. Since the introduction of Salmonella monitoring, a decrease in category-three pig farms has been observed from 5.8% in 2005 to 1.6% in 2020 [19].
Various studies have shown that direct contact with infected animals and the introduction of already infected piglets into nurseries [20,21] and fattening units [8,11,13,21,22] present a greater risk of infection than contaminated feed [23] or the environment. Therefore, control methods should aim to reduce the presence of Salmonella in all early stages of production (gilts, sows, piglets) to help to decrease the risk of infection in fatteners and consequently the risk of introduction of the pathogen into the food chain [13,24,25]. Often, animals on Salmonella-infected farms do not show clinical signs, and therefore it is important to use a sensitive sampling method to identify infected breeding herds and prevent purchases of asymptomatic carriers [22,26,27]. Additionally, the sampling method must be practically and economically feasible to ensure it can be used on a large scale. Although serology on sows could be valuable to identify the Salmonella prevalence within a sow herd [28,29], blood samples do not give information about the serovars involved or the spread of the pathogen on the farm, in addition to having possible welfare implications. For direct detection of Salmonella on-farm, bacteriological examination or direct PCR methods using fecal samples and different types of environmental samples have been described, including surface wipes and sock swabs [9,10,11,22,30]. However, large numbers of fecal samples are required to detect low shedding rates within groups of animals [8,11,22,29].
Sock and environmental swab samples have already been described as a reliable diagnostic method in nursery pigs, guiding clinical decisions for treatment and prevention [31], and have been proven to be a suitable method for monitoring farms and detecting Salmonella in poultry [32,33,34]. Furthermore, in a study including 101 fattening and 41 farrow-to-finish farms in the north-west of Germany, environmental sampling was proven to provide an adequate basis for Salmonella control in farms with a high seroprevalence. Significantly more areas tested positive compared to farms with a low seroprevalence [30].
In contrast to individual fecal samples, the sock and swab method does not give information about the Salmonella prevalence within a group of animals. However, using one swab and one sock within a compartment means that only a small number of samples are required per farm. Low shedding rates of sows, shown to often be below 10%, would require a high number of individual fecal samples. To detect one shedding sow in a 1000-sow herd with 10% Salmonella prevalence with a 95% confidence level, 28 samples would be necessary [35,36]. In a longitudinal follow-up trial in a Salmonella control program in the UK, 60 individual samples in each epidemiological group (dry sows, lactating sows, weaners, etc.) were required each visit to address a 5% prevalence presuming 100% sensitivity [24]. Using pooled fecal samples lowers the number of samples needed [37,38]. In another longitudinal approach, sock samples led to a higher sensitivity than pooled fecal samples [9]. Environmental swabs were found to be an ideal method for identifying residual Salmonella in areas which are usually not accessible by animals within the compartments, such as, the undersides of troughs and window sills covered with dust [30].
The aim of the present study was to assess the Salmonella status of sow farms either producing their own finishers or delivering piglets to fattening farms. All downstream fattening units had a known high serological prevalence identified within the QS Salmonella monitoring system. The upstream sow herds were selected for sampling depending on the serological status of the associated fattening herd. As only some federal states in Germany are offering facultative Salmonella monitoring of sow farms, the sow farms in this study were sampled for Salmonella for the first time. Previous studies have shown the viability and sensitivity of sock and environmental swab samples for individual farms with different serological statuses [9,11,30,39,40]. However, this large-scale study aimed to determine whether sock samples from pens and environmental swab samples from reachable surfaces are suitable to identify areas contaminated with Salmonella in sow farms and the associated nursery, which are linked to fattening farms with a high serological Salmonella prevalence. Additionally, it was assessed whether a minimum of 16 samples per farm is a suitable number of samples to detect Salmonella on-farm. As in a recent publication, it was concluded, that sock sampling alone would be sufficient to determine the status of a herd [9], within this trial, we investigated, if both types of samples are needed to give a good overview of the farm. Additionally, it was aimed to identify the serovars present on the farms to be able to implement appropriate control measures.
2. Materials and Methods
2.1. Study Farms
A total of 105 farms across Germany, including two farrowing farms, 71 farrow–nursery farms and 32 farrow-to-finish farms with sow numbers ranging from 50 to 10,000 were sampled within this study. The evaluation period was from January 2015 to July 2021. All sow farms selected for the study were linked to fattening units with a known high Salmonella (S.) seroprevalence.
2.2. Sampling
The number of sock and environmental swab samples taken on each farm depended on the size of the farm and the production cycle. To cover all production sites, one sock sample and one environmental sample each were taken in two farrowing rooms (around farrowing and shortly before weaning), in the insemination room, in gestation rooms (pens with newly grouped sows), the gilt quarantine room and in three rooms in the nursery (beginning, middle and end of nursery), resulting in a minimum of 16 samples taken per farm. In farms containing more than one compartment filled with the respective age group/stage of production, all rooms belonging to these ages were sampled. Empty, cleaned and disinfected compartments already prepared for the arrival of new pigs were sampled if accessible during the farm visits. This was performed in farms using a routine protocol for cleaning and disinfection, including mechanical removal of dirt before soaking with water and cleaning with a high-pressure washer followed by drying and disinfection. To increase the probability of finding pigs shedding Salmonella, groups were sampled preferentially after a stressful period, e.g., after they had been moved from the mating pen to a holding pen.
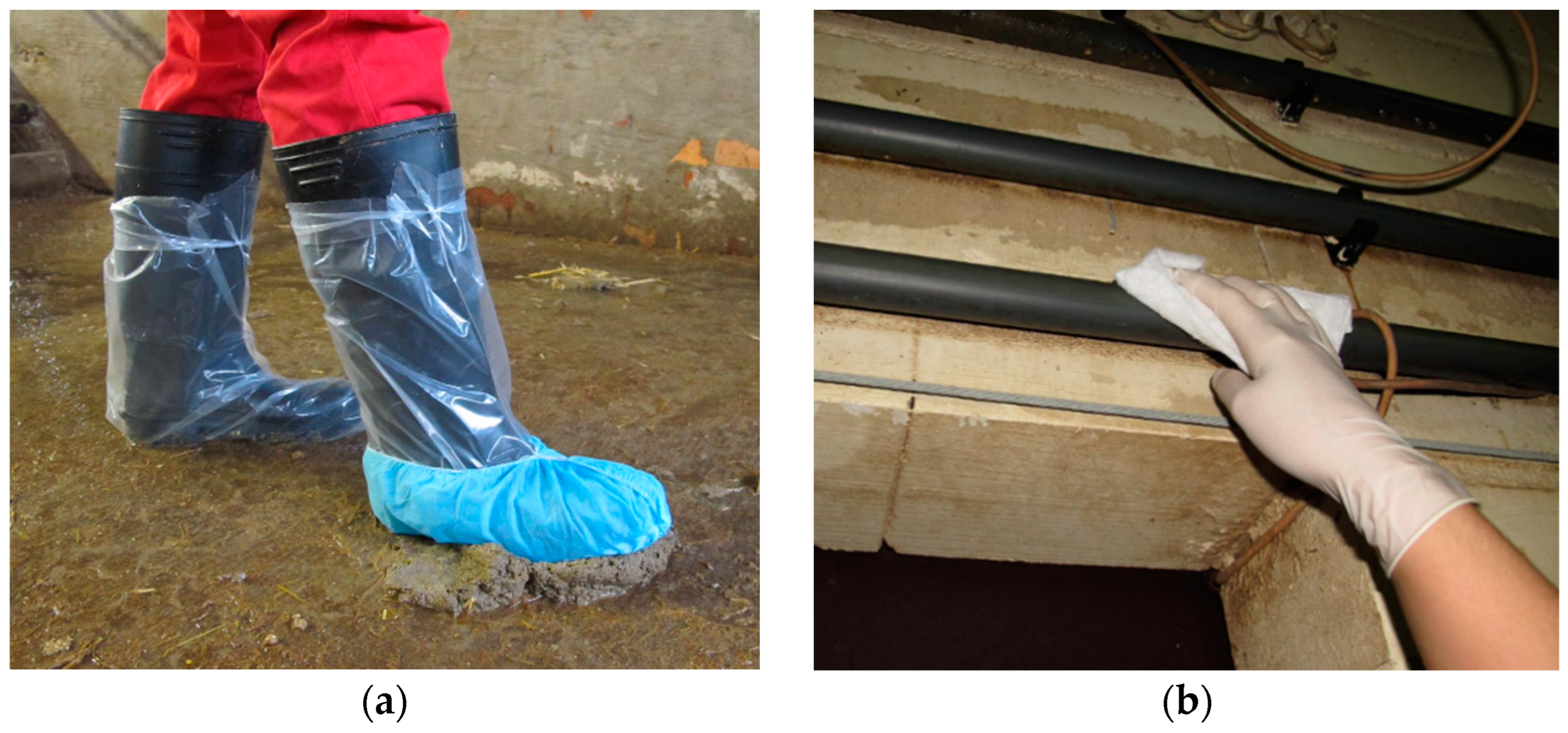
All materials used for the sampling were dry, clean and individually packed by Sterimatic, Stroud, UK. The sample kits contained one pair of non-sterile gloves (lightly powdered latex examination gloves, Covetrus Germany), a dry synthetic dust cloth (Swiffer®, Procter & Gamble Service GmbH, Crailsheim, Germany), one pair of plastic overshoes (PE boot cover transparent, WIROS Wilfried Rosbach GmbH, Willich, Germany) one sock (“Hygostar” polypropylene nonwoven overshoes, Franz Mensch GmbH, Buchlohe, Germany) and two plastic bags. Sampling was carried out by one of four trained and experienced people. To prevent cross contamination, a new pair of gloves was used for each sample collection. After sampling, the socks and swabs were placed into individual plastic bags, which were firmly closed and individually marked. When collecting the sock sample, each boot was first covered with the plastic overshoe before entering the pen. The sock was placed over one plastic shoe to collect fecal material from the floor of each pen within the sampled compartment (Figure 1a). All pens within a sampled compartment were sampled in a standardized way. As much fecal material was stepped in as possible within each pen. It was walked along the walls, into the corners, around the water supply and the trough. While sampling the pens, it was important not to contact the walkways to ensure the sample only reflected the shedding of the animals within the pens. In the cleaned and disinfected compartments, the focus was on stepping into obvious dirty spots or, if present, into dead flies and rodent feces. One environmental swab was used per room to sample sedimented dust on water pipes and feeding lines (a minimum of 3 m at different locations) (Figure 1b), as well as walls above the belly button height, surfaces of air inlets, ventilation systems and window sills.

Figure 1.
(a) Sock sampling; (b) Environmental swabs.
All samples were shipped without extra cooling by an overnight courier to the laboratory.
2.3. Bacteriology
Socks and environmental swabs were examined bacteriologically at the Institute for Microbiology at the University of Veterinary Medicine in Hanover (Stiftung Tierärztliche Hochschule Hannover, Institut für Mikrobiologie, Bischofsholer Damm 15, Gebäude 126, 30173 Hannover, Germany). All samples were enriched non-selectively in 225 mL of buffered peptone water to yield a tenfold dilution (BPW; Oxoid, Thermo Fisher, Wesel, Germany) and incubated for 18 h at 37 °C. From this non-selective pre-enrichment, 1 mL was transferred to 8 mL of tetrathionate brilliant green bile broth (TBG; VWR, Darmstadt, Germany) and 0.1 mL was transferred to 10 mL of Rappaport-Vassiliadis Soy Broth (RVS; Oxoid, Germany). Both tetrathionate brilliant green bile broth and Rappaport-Vassiliadis Soy Broth acted as selective liquid media. All samples were incubated for 24 h at 42 °C and then streaked on Oxoid Brilliance Salmonella Agar. The plates were inspected for growth of typical purple colonies after 24 h at 37 °C. One typical colony per sample was subcultured on non-selective sheep blood agar (Oxoid, Germany). Colonies were then submitted to an in-house PCR that identified S. Typhimurium (STM) (including the monophasic variant of STM (mSTM)), S. Enteritidis (SE) and S. Dublin. Other serovars produce a S. enterica ssp. enterica-specific and/or a genus-specific signal in this multiplex PCR [41]. Further characterization via slide agglutination for a selection of relevant O- and H-specific antigens (Sifin Antisera, Berlin, Germany) was completed if isolates were only identified as S. enterica ssp. enterica and/or S. genus by PCR. These antisera included anti-O4, anti-O5, anti-O9, anti-group C, anti-group E; anti-Hi, anti-H2, anti-Hf, anti-Hg, anti-Hm, anti-Hp, anti-Hq, anti-Hs and anti-Ht). A farm was classified as Salmonella-positive if at least one sample tested positive. For farm analysis, the samples were grouped according to their positivity for STM, Salmonella Enteritidis (SE), S. Derby (SD) or for other serovars. The full identification process was performed only for STM, SE and SD, whereas the other isolates were only identified within their serogroup.
2.4. Statistical Analysis
The data were tested pairwise between the groups with a two-sided Wilcoxon–Mann–Whitney U-Test, alpha = 0.05. Tests were performed per region (East/West Germany) and per farm size (1–299 sows, 300–999 sows and more than 1000 sows) and additionally considering the difference in positivity within each area of the farm (farrowing, nursery, other areas). For the binominal distribution of samples within each area, the confidence interval (CI) was set at 95%. To evaluate a possible difference in positivity of sock and environmental swabs, a McNemar test was used, with p < 0.05 showing a statistical difference between frequencies of positive samples between the two types of samples. Additionally, the Kappa Index was calculated to obtain a measure of the agreement of these two methods (<0.01: no agreement; values between 0.1 and 0.4: weak agreement; values between 0.41 and 0.60: clear agreement, values between 0.61 and 0.80: strong agreement) [42]. The analysis was performed with the validated program Testimate Version 6.5 from IDV Datenanalyse und Versuchsplanung, Gauting, Germany.
3. Results
In total, 105 farms were included in the study. At farm level, 97/105 farms (92.4%) from all over Germany tested positive for Salmonella in at least one sample. Evaluating samples geographically showed that 27/30 (90%) of the farms in eastern Germany tested positive and 70/75 (93.3%) of the farms in western Germany tested positive.
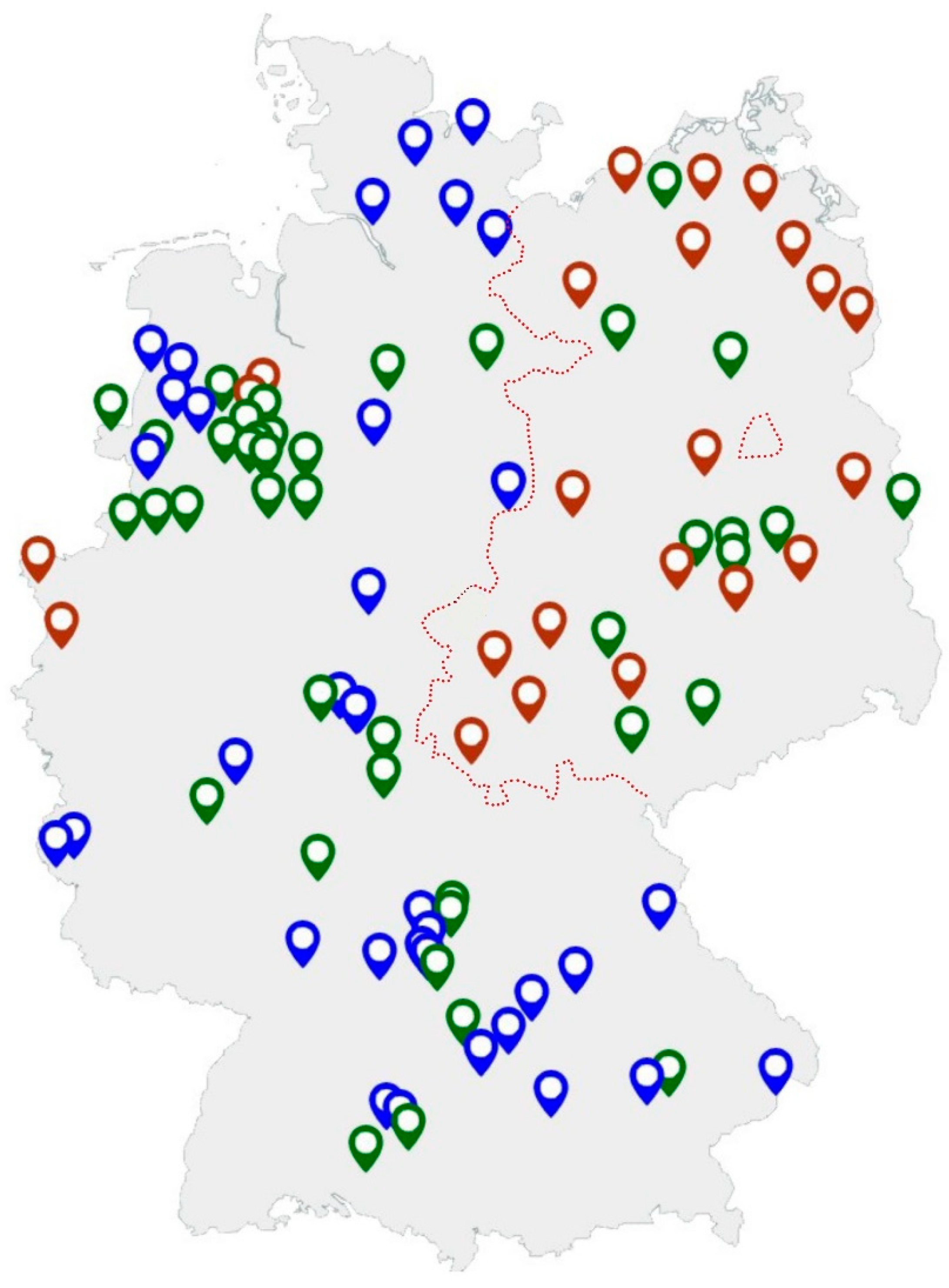
In summary, 36/37 (97.3%) farms keeping one to 299 sows and 36/42 (85.7%) farms with 300–999 sows tested positive. Nearly all farms (22/23) with over 1000 sows were found to be positive (95.5%). For three farms, all of which were positive for Salmonella, no information about the farm size was available. All farms with <300 sows were located in western areas of Germany (n = 37), while no farms of this size were present in eastern areas of Germany. There were no statistical differences between farm size, region and positivity for Salmonella. Figure 2 gives an overview of the location of the tested farms.

Figure 2.
Location of tested sow farms (blue: farms with up to 299 sows, green: farms with 300–999 sows, red: farms with >1000 sows). The border of former East and West Germany until 1989 is indicated by the red line.
The number of samples taken per production area is depicted in Table 1. In total, 2355 environmental and sock swabs with an average of 22 samples/farm were analyzed (detailed information about the number of samples taken per farm is shown in the Supplementary Material Table S1). In total, 779 samples were identified as positive (33.1%). Most samples (n = 970) were taken from the nursery, followed by 662 samples from the farrowing area. A further 452 samples were taken from other areas, including gestation, gilt quarantine and insemination areas. Cleaned and disinfected compartments and boot washers were available on 23 and 15 farms, respectively.

Table 1.
Overview of samples (n = 2355) taken from different areas in 105 German sow farms (CI = confidence interval).
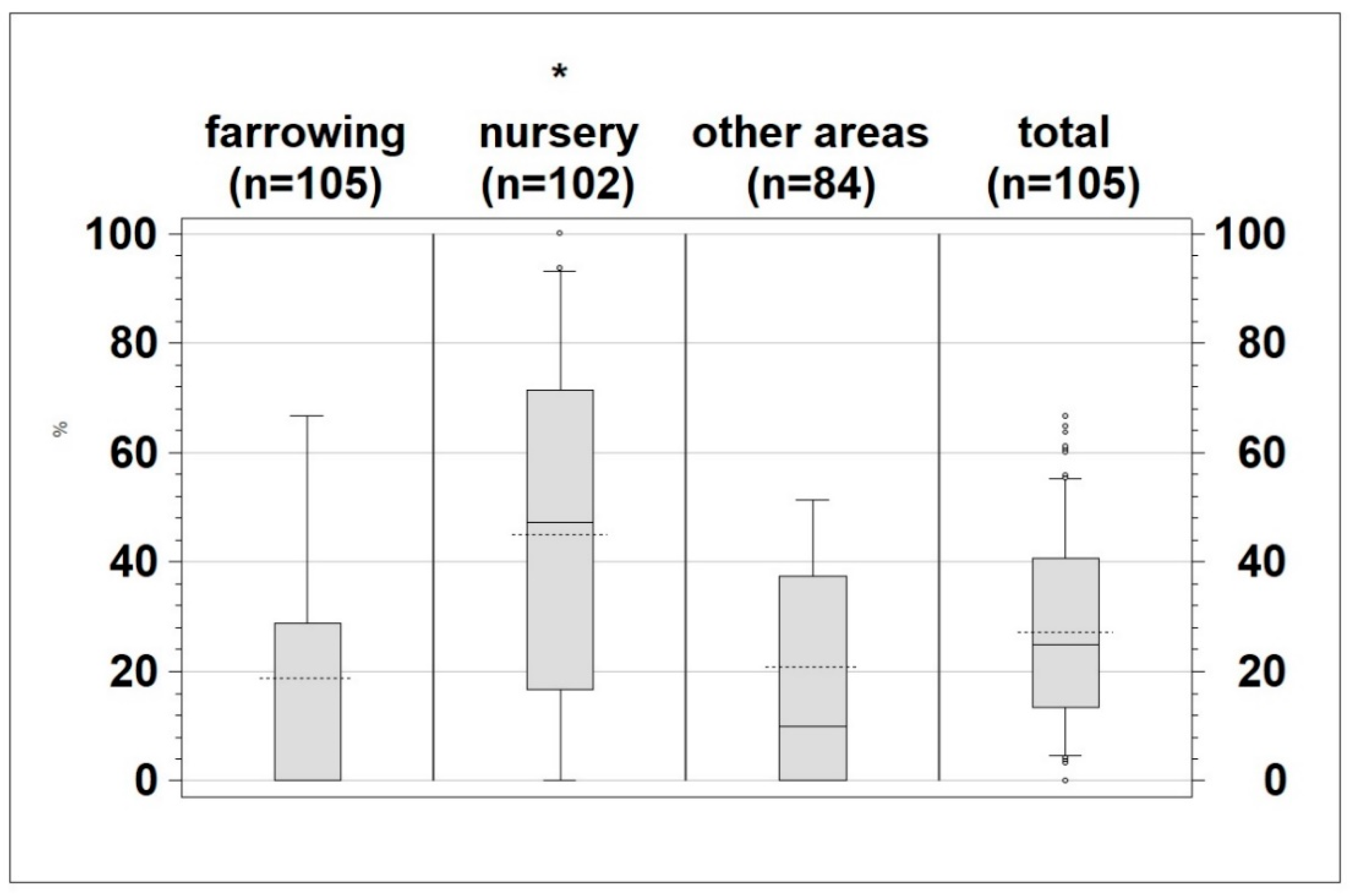
The proportion of positive samples (mean 27.2%) at herd level is shown in Figure 3. The number of samples that tested positive within the different production areas differed for the individual farms. Nevertheless, a significantly higher (p < 0.0001) proportion of samples from the nursery were positive for Salmonella (mean 45%) compared to those taken from farrowing (mean 18.7%) and other areas (mean 20.86%) (Figure 3).
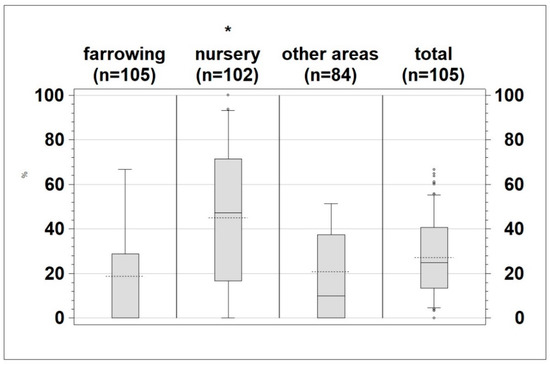
Figure 3.
Percentage of positive samples per sampled area (farrowing, nursery, other areas and total) * p < 0.0001, dotted line: mean; whiskers: 10th and 90th percentile.
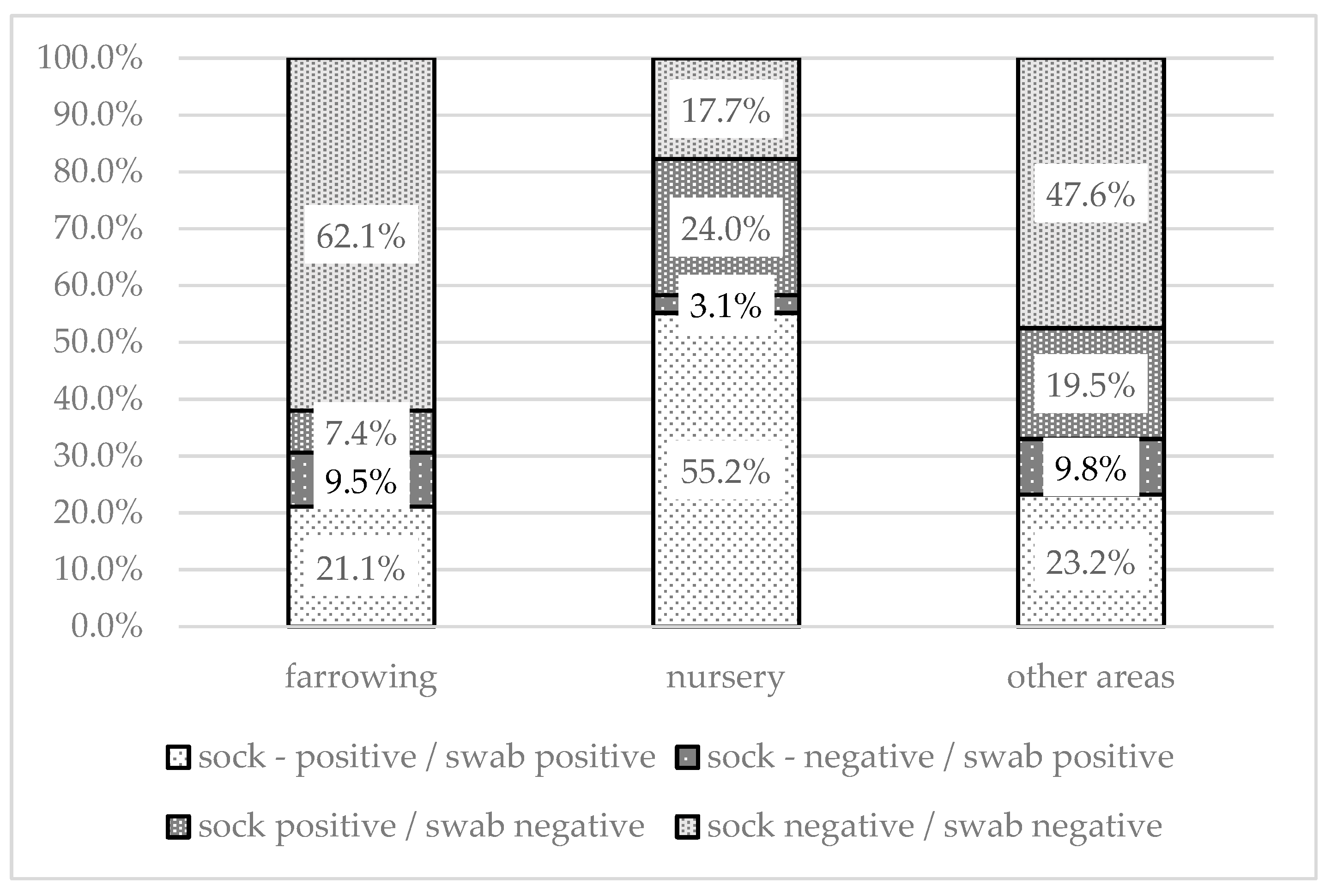
The proportion of different combinations of positive or negative swabs and environmental samples varied for different sampling areas (Figure 4). Both positive sock and swab samples were found most often in the nursery (55.2%). A combination of positive swabs and negative socks was the least commonly recorded. Negative results for both sock and swab samples were highest in farrowing areas (62.1%). In 24% of the nurseries, only the socks were positive; this was significantly higher (p < 0.001) than the proportion of nurseries in which only swabs were positive (3.1%). The results of the statistical analysis are shown in Table 2.
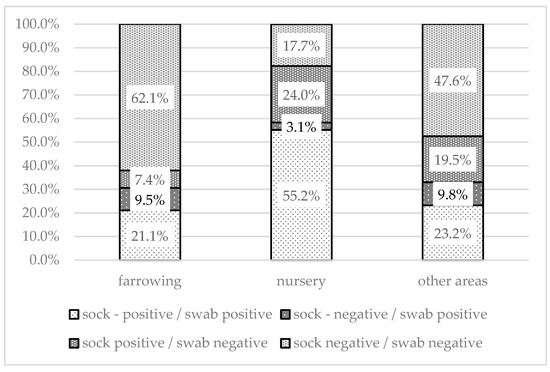
Figure 4.
Comparison of positivity of sample by method (sock swab/environmental swab) within each sampled area.

Table 2.
Distribution of frequencies with McNemar test and Cohen’s Kappa values reflecting agreement between sock swabs and environmental swabs within the different sampled areas (farrowing, nursery, other areas).
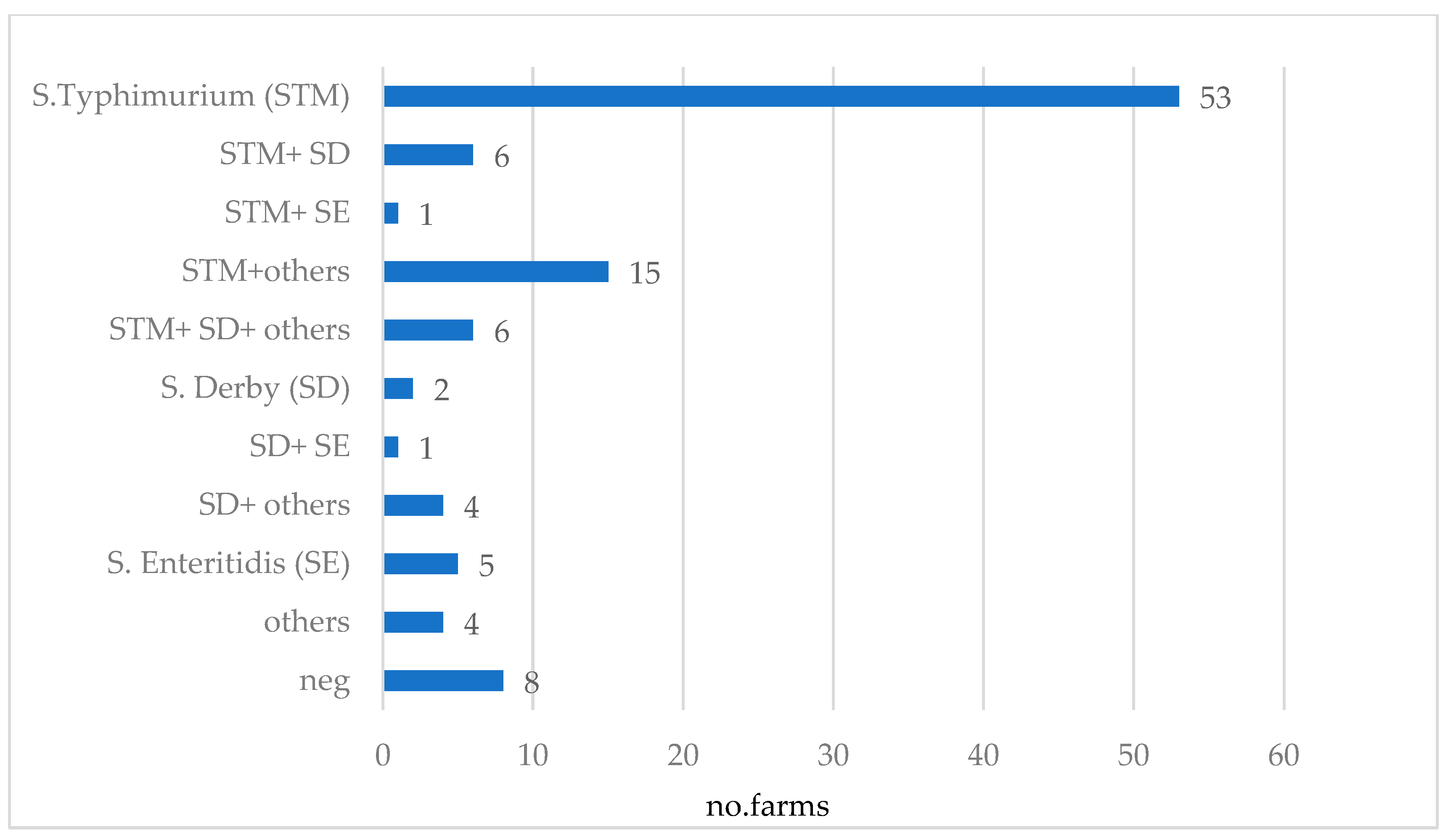
The distribution of detected serovars across the 105 farms is presented in Figure 5. STM was found on 77.1% (n = 81) of the farms and in 50% (n = 53), it was the only serovar detected. Detection of STM together with SD occurred in 11.4% (n = 12) of farms, while the detection rate of STM combined with other serovars was 15.2% (n = 16). SD could be detected on 18.1% (n = 19) of the farms, and in 1.9% (n = 2) of cases, it was the only serovar detected. Salmonella Enteritidis (SE) alone occurred on 4.8% (n = 5) of farms, and in combination with other serovars in 1.9% (n = 2) of farms. Positivity for other serovars alone was reported for 3.8% (n = 4) of farms, and on 7.6% (n = 8) of farms, no Salmonella could be detected. A complete overview of the serovars and serogroups detected on each farm is provided in the Supplementary Material Table S1.
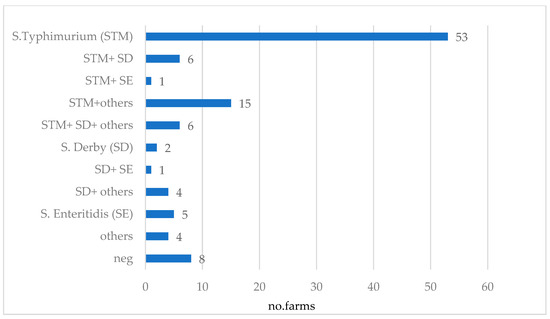
Figure 5.
Combinations of Salmonella serovars detected in 105 German sow farms via bacteriological examination of sock and environmental swab samples (others: S. group B (other than STM and SD), S. group C and S. group E and non-typable isolates given the limited set of sera implemented in this study).
Evaluation of individual farms and their different production areas shows that 97/105 (92.3%) were positive for at least one serovar. While in 9 farms (8.6%) only one serovar was found in other areas, and nursery and farrowing were negative for Salmonella, in 19 farms (18.1%), only the nursery was positive and one serovar was identified. On 31 (29.5%) farms, different serovars were detected in different positive areas of the farm. (Supplementary Material Table S1).
4. Discussion
This study aimed to identify Salmonella (S.) on 105 piglet-producing farms in Germany, which may be the source of Salmonella introduction into the linked fattening farms. The farms involved were all delivering pigs to fattening farms with high serological prevalence, as identified via the obligatory German serological monitoring system [17], or were experiencing clinical issues due to Salmonella. These included severe, hard-to-treat clinical signs, reduced daily weight gain or an unspecific increased number of losses and runts [1]. Before sampling, farm Salmonella status was unknown. This initial sampling was carried out to gain insights into the groups of animals shedding Salmonella and the environmental load. Identification of serovars present on the farms was also an aim of the study. Sock and environmental swab samples were used to identify Salmonella serovars present on farm, with sock sampling being reflective of bacteria shed by the animals [9] and environmental swabs giving an overview of the Salmonella contamination in the environment [30]. We also aimed to identify the areas of production with the highest shedding and environmental pressure, as shown by the number of positive samples, and determine if both sample materials are really needed to identify the production areas at risk within a farm.
Salmonella was detected in 92.4% (n = 97) of the studied farms, with a farm being classified as positive when just one sample was positive for Salmonella. In our study, the chance of identifying Salmonella was independent of the farm size, which is contrary to other publications where a relationship between the Salmonella positivity and the size of the farm or group size was reported [43]. Previously, STM shedding of breeding pigs was found to be associated with herd size and the number of pigs in the pens [25]. In contrast, we found no significant difference regarding the positivity within each production area within the farms when comparing farm size and region. The high positivity rate and the lack of variation in farm size and region within this study could be due to herd selection being based on linkage to fattening farms with a known high serological prevalence or the presence of clinical signs with suspicion of Salmonella involvement. This is supported by previous authors demonstrating the direct link between serologically positive fatteners and the Salmonella prevalence of breeding herds [12,13].
The eight farms that had negative results might have truly been negative; however, a false negative result may occur due to either a low sample size that was not sufficient to detect a very low prevalence or the introduction of Salmonella into the fattening farms via other routes, such as feed, rodents or other piglet origin [23,25,44]. Another possible explanation could be infection of animals with a host-adapted Salmonella, mainly S. Choleraesuis (SCS), which may have led to a strong immune response and therefore a high serological prevalence in fatteners. Unlike most of the other serovars, SCS is seldom detected in the environment [1]; however, the pathogen can be found during necropsy using suitable tissue for isolation, including that from the lungs, especially the cranioventral portion of the caudal lobe, mesenteric lymph nodes, spleen, liver, and kidneys [45]. As a next step, farms testing negative should be sampled a second time to validate the result. If this result is confirmed, in these cases the risk of infected piglets being introduced into fattening farms is very low. Thus, the hygienic measures in these fattening farms seem to have most impact on the Salmonella status of their fattening pigs before slaughter.
The highest number of positive samples (p < 0.0001) was found in the nurseries. This supports previous findings [8,11,29] reporting the highest detection rates in the nursery. Especially important are the findings of Bernad-Roche et al. in 2021 [21], where they demonstrated that the serovars detected in weaners and within a given nursery remained the same over a long period of time (>150 days), and were found even later in gilt rearing units. The high number of positive samples in the nursery in our study supports the hypothesis that these piglets are a source of Salmonella for the linked fattening farms, particularly as they can become subclinically infected and act as active carriers of Salmonella [21].
Our study underlines the risk of suckling piglets becoming infected in the farrowing units. The mean of samples in the farrowing areas that were positive for Salmonella was 18.9%, with a maximum of 70% of samples positive in some farms, which is much higher than reported by Hollmann et al. 2022, who in the peripartal period only found a few boot swabs positive for Salmonella [11]. Other authors also reported prevalence below 10% in three Belgian sow farms using individual fecal samples [22], similar to another study where just a low amount of environmental samples taken in 12 Salmonella-suspected identified farms were positive in culture [46]. Our data underline, that environmental sock and swab samples taken in the farrowing area can indicate shedding of sows, shedding of previously infected suckling piglets or a contaminated environment. It has been shown that weaning piglets can be infected and positive in the mesenterial lymph nodes [20,21]. Therefore, the farrowing area should be sampled in a thorough manner, especially the environment, to assess the risk for the first infection of piglets as early as the suckling period. Even if vaccination is successful at reducing shedding of the sows [24,47], farrowing houses are difficult to clean and disinfect [48] and therefore can be a source of constant reintroduction of the pathogen into the nurseries.
In 23 farms it was possible to take samples in cleaned and disinfected compartments. In 14 (60.9%) of these farms, allegedly cleaned and disinfected areas were positive for Salmonella, leading to 35.9% (n = 23) positive samples. Sampling of some of these compartments led to detection of visible contamination (old feed in feeders, droplets of feces in the corners or dust in the water/feed pipes), which indicates the need for improvement of hygiene measures. Residual dirt on drinkers and feeders in cleaned and disinfected compartments have been shown to be a source of reinfection of pigs [49]. In poultry, there is evidence for contaminated feeders and drinkers being reservoirs for Salmonella in production systems [50,51]. Thorough cleaning prior to disinfection to remove any organic matter is vital in ensuring the effectiveness of the disinfectant is not reduced [52]. Non-adequate room temperature, especially one that is too cold, and the pH of the solution prepared for disinfection can affect the potency of a disinfectant [53]. It is important to (i) ensure proper drying out of a room before disinfection, (ii) ensure compatibility of used detergents and disinfectant and (iii) ensure the correct final concentration of the disinfectant [53].
In pig production, and mainly in nursery and fattening, similar to poultry production, “all-in/all-out” management is commonly used to break infection chains and minimize the risk of young animals being infected by older animals [54,55]. Walkways and any crossing of paths on walkways can also have an important role in transmission of Salmonella between different age groups and production areas. In this study, the walkways in 38 out of 74 farms, and the boot washers in 10 of 15 farms tested positive. As Salmonella can be sustained for up to 50 months in the environment, e.g., in dust [56], and up to 24 weeks in manure [57], reinfection by contact with contaminated walkways is a high risk which must be addressed in Salmonella control programs [9].
The findings of this study can support decisions for the best sampling strategy in a sow farm. Whereas other authors concluded that sock samples would be a good sampling strategy as a standalone method [9,11], the results of our study indicate that, besides the nursery, where sock samples were positive significantly more often than environmental swabs, both samples should be taken at each sampling point. Although the statistical analysis for the farrowing area showed that there was strong agreement between the two methods, indicating one of the methods alone would be enough, in almost 10% of cases (9.5%), a positive result would have been missed had no swab been taken. Therefore, we conclude that in farrowing, the use of both methods is advisable. The same can be recommended for the other areas. This clearly demonstrates the importance of assessing environmental contamination in these areas with a swab, as well as the impact of shedding animals, by using a sock sample [27]. Acids in the feed and specific feeding strategies, including an increase in particle sizes, as well as vaccination against Salmonella, can reduce Salmonella shedding [58,59] and might consequently decrease the detection rate of Salmonella in feces and thus in sock samples. Especially in these cases, environmental swabs are extremely important for detecting environmental contamination (Salmonella reservoirs) and therefore potential risk of reinfection.
To implement sustainable Salmonella programs within a production chain, it is important to assess the serovars involved. In this study, 77.1% of the farms were positive for STM followed by 18.1% for S. Derby, either alone or in combination with other serovars. These findings reflect the situation found in European pig production over the last few years [60], considering STM is currently the most frequently reported serovar isolated from pigs [26,27,61] and also the most relevant Salmonella reported in human cases related to pork or pork meat products [5]. In 35% of the farms (n = 37), more than one serovar could be detected. Within the serogroups C, D and E, no further analysis on the individual serotype was performed. These serogroups include serotypes such as S. Infantis, S. Rissen, S. Ohio and S. London, which are also regularly detected either on-farm [21,22,62] or in lymph nodes at slaughter [5,20,21]. Therefore, a conclusion on the exact number of different serovars present on each farm and within each production area cannot be given. In recent studies, several different serotypes within a serogroup were detected on one farm [20,21,24]. SCS, also belonging to serogroup C, is mainly found in the organs of acutely diseased pigs, and is not commonly found in the environment [1,45]. Therefore, no further analysis to identify SCS was performed within this study. On one of the farrow-to-finish farms that tested negative in socks and environmental swabs, further examinations of acutely diseased pigs showed that SCS could be found from Salmonella enrichment of organs (spleen, kidneys, lung and liver). The farm did not observe salmonellosis outbreaks but remained in category three within the QS Salmonella monitoring system for years. This shows that, in addition to environmental sampling, in some cases necropsies should be performed to confirm the Salmonella involved.
Using the results of farm sampling, further categorization of Salmonella status for each farm can be performed. If a very low number of samples are positive within a sampling period, the relevance for the farm must be discussed. This was the case for eight farms within this study, where only one environmental sample was positive for an exotic Salmonella serovar. In such cases, further sampling should be conducted prior to implementing a Salmonella control program. However, for farms with high positivity rates, especially in the nursery, control measures should be implemented. In our study, the median of positive samples within the nursery was 47.2% and the lower quartile was 16.7%, showing that most farms within the study need to implement control measures for Salmonella. Due to the high positivity rate in cleaned and disinfected compartments, the focus should be on the improvement of hygiene measurements. As STM was the predominantly detected serovar, in addition to hygiene measures, vaccination can help to reduce STM prevalence on-farm. Vaccination of sows using a live vaccine with a double attenuated STM strain (Salmoporc®, Ceva Santé Animale, Libourne, France) has been shown to reduce shedding around farrowing, and consequently reduce infection of piglets [9,24,47,63]. Vaccination of piglets leads to a reduction in STM shedding and prevalence at slaughter [64,65,66]. Autogenous vaccines, custom-made from the specific serovar(s) identified on the farm, can be employed when commercial vaccines are not available or effective against the Salmonella serovar of concern [67]. Except for vaccination, evidence of different serovars does not require different control measures. In addition to the implementation of strategies to reduce shedding of Salmonella by infected sows and piglets by feeding strategies [68], the use of acids [59,69] or improvement of housing [12], it is important to determine the residual sources of Salmonella, which could, for example, be contaminated feed/water pipes or rodents.
Within our study, no Salmonella could be detected on eight farms which underlines the results of a trial in the Netherlands, where it was possible to reduce the Salmonella prevalence to below the detection level [9]. As a follow-up, comparing the management systems of positive and negative farms could result in valuable information on the management risk factors with most impact on the Salmonella status of the farm.
In addition to the use in an initial evaluation of the status of the sow farms, sock and environmental swab samples are valuable in determining the success of implemented control measures [9,10,11]. Pooled fecal samples have already been shown to be more sensitive than individual samples, with 20 samples per pool identified as being the most sensitive indicator [37]. Socks taken within the pens were shown to be more sensitive than individual samples [11], aiding the demonstration of a successful reduction in the on-farm prevalence in longitudinal sampling parallel to implemented measures against Salmonella [9]. Previous studies have shown that the time needed to reduce Salmonella prevalence at farm level is months rather than weeks [9,24]; therefore, sampling intervals of every 3–6 months are sufficient. The monitoring of effective cleaning and disinfection should be performed on a regular basis and only requires the use of one sock and one swab sample per compartment.
5. Conclusions
In the present study, sock and environmental swab samples proved to be useful as non-invasive and easy-to-apply methods to screen sow farms for the presence of Salmonella, being suitable for all farm sizes. Our results underline the need to include breeding and multiplying levels into monitoring and control systems, as this allows the involved Salmonella serovars to be identified in early stages, before they are introduced into the fattening units and later into the food chain. Additionally, it could be underlined that there is a huge demand for thorough control of cleaning and disinfection to interrupt the Salmonella infection cycle. The use of sock and environmental swabs in combination proved to be crucial as the detection rates of each sampling method varied.
Sock and environmental swabs are useful for validating the implementation of Salmonella control measures. They can be used to verify the effects of various Salmonella reduction measures, such as cleaning and disinfection, optimizing walking routes, changing boots and vaccination [9,11].
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/ani13061031/s1. Table S1: Individual results of Salmonella serovars detected on each farm within each area including information about farm size, type of farm, no. of samples and date of sampling.
Author Contributions
Conceptualization: I.H.-P. and K.L.-J.; methodology: J.R. and K.L.-J.; formal analysis: I.H.-P., K.L.-J. and C.W.; investigation: K.L.-J., M.V. and C.W.; data curation: K.L.-J.; writing—original draft preparation: K.L.-J., S.H. and M.V.; writing—review and editing: I.H.-P., J.R. and C.G.; visualization: K.L.-J. and C.W.; supervision: I.H.-P.; project administration: K.L.-J. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Ethical review and approval were waived for this study as the samples were taken from the environment and no animals were involved in the sampling process.
Informed Consent Statement
Informed consent was obtained from all subjects involved in the study.
Data Availability Statement
Data are available from corresponding author upon request.
Acknowledgments
The authors thank all the veterinarians and farmers who supported this investigation.
Conflicts of Interest
The authors declare no conflict of interest. This study did not use or evaluate any commercial product.
References
- Zimmerman, J.J.; Karriker, L.A.; Ramirez, A.; Schwartz, K.J.; Stevenson, G.W.; Jianqiang, Z. (Eds.) Diseases of Swine, 11th ed.; Wiley-Blackwell/American Association of Swine Veterinarians: Hoboken, NJ, USA, 2019; ISBN 978-1-119-35090-3. [Google Scholar]
- Vo, A.T.T.; van Duijkeren, E.; Fluit, A.C.; Heck, M.E.O.C.; Verbruggen, A.; Maas, H.M.E.; Gaastra, W. Distribution of Salmonella Enterica Serovars from Humans, Livestock and Meat in Vietnam and the Dominance of Salmonella Typhimurium Phage Type 90. Vet. Microbiol. 2006, 113, 153–158. [Google Scholar] [CrossRef] [PubMed]
- Basler, C.; Nguyen, T.-A.; Anderson, T.C.; Hancock, T.; Behravesh, C.B. Outbreaks of Human Salmonella Infections Associated with Live Poultry, United States, 1990–2014. Emerg. Infect. Dis. 2016, 22, 1705–1711. [Google Scholar] [CrossRef] [PubMed]
- European Food Safety Authority; European Centre for Disease Prevention and Control The European Union One Health 2020 Zoonoses Report. EFSA Journal 2021, 19, e06971. [CrossRef]
- European Food Safety Authority and European Centre for Disease Prevention and Control (EFSA and ECDC) The European Union One Health 2018 Zoonoses Report. EFSA Journal 2019, 17, e05926. [CrossRef]
- Correia-Gomes, C.; Leonard, F.; Graham, D. Description of Control Programmes for Salmonella in Pigs in Europe. Progress to Date? J. Food Saf. 2021, 41, e12916. [Google Scholar] [CrossRef]
- dos Santos Bersot, L.; Quintana Cavicchioli, V.; Viana, C.; Konrad Burin, R.C.; Camargo, A.C.; de Almeida Nogueira Pinto, J.P.; Nero, L.A.; Destro, M.T. Prevalence, Antimicrobial Resistance, and Diversity of Salmonella along the Pig Production Chain in Southern Brazil. Pathogens 2019, 8, 204. [Google Scholar] [CrossRef]
- Weaver, T.; Valcanis, M.; Mercoulia, K.; Sait, M.; Tuke, J.; Kiermeier, A.; Hogg, G.; Pointon, A.; Hamilton, D.; Billman-Jacobe, H. Longitudinal Study of Salmonella 1,4,[5],12:I:-Shedding in Five Australian Pig Herds. Prev. Vet. Med. 2017, 136, 19–28. [Google Scholar] [CrossRef]
- van der Wolf, P.; Meijerink, M.; Libbrecht, E.; Tacken, G.; Gijsen, E.; Lillie-Jaschniski, K.; Schüller, V. Salmonella Typhimurium Environmental Reduction in a Farrow-to-Finish Pig Herd Using a Live Attenuated Salmonella Typhimurium Vaccine. Porc. Health Manag. 2021, 7, 43. [Google Scholar] [CrossRef]
- Lynch, H.; Walia, K.; Leonard, F.C.; Lawlor, P.G.; Manzanilla, E.G.; Grant, J.; Duffy, G.; Gardiner, G.E.; Cormican, M.; King, J.; et al. Salmonella in Breeding Pigs: Shedding Pattern, Transmission of Infection and the Role of Environmental Contamination in Irish Commercial Farrow-to-Finish Herds. Zoonoses Public Health 2018, 65, e196–e206. [Google Scholar] [CrossRef]
- Hollmann, I.; Lingens, J.B.; Wilke, V.; Homann, C.; Teich, K.; Buch, J.; Chuppava, B.; Visscher, C. Epidemiological Study on Salmonella Prevalence in Sow Herds Using Direct and Indirect Detection Methods. Microorganisms 2022, 10, 1532. [Google Scholar] [CrossRef]
- Hill, A.; Simons, R.; Ramnial, V.; Tennant, J.; Denman, S.; Cheney, T.; Snary, E.; Swart, A.; Evers, E.; Nauta, M.; et al. Quantitative Microbiological Risk Assessment on Salmonella in Slaughter and Breeder Pigs: Final Report. EFS3 2010, 7, 46E. [Google Scholar] [CrossRef]
- Hill, A.A.; Simons, R.R.L.; Kelly, L.; Snary, E.L. A Farm Transmission Model for Salmonella in Pigs, Applicable to E.U. Member States: Farm Transmission Model for Salmonella in Pigs to Assist E.U. Member States. Risk Anal. 2016, 36, 461–481. [Google Scholar] [CrossRef]
- Ivanek, R.; Österberg, J.; Gautam, R.; Sternberg Lewerin, S. Salmonella Fecal Shedding and Immune Responses Are Dose- and Serotype- Dependent in Pigs. PLoS ONE 2012, 7, e34660. [Google Scholar] [CrossRef]
- Österberg, J.; Sternberg Lewerin, S.; Wallgren, P. Patterns of Excretion and Antibody Responses of Pigs Inoculated with Salmonella Derby and Salmonella Cubana. Vet. Rec. 2009, 165, 404–408. [Google Scholar] [CrossRef]
- Pires, A.F.A.; Funk, J.A.; Bolin, C.A. Longitudinal Study of Salmonella Shedding in Naturally Infected Finishing Pigs. Epidemiol. Infect. 2013, 141, 1928–1936. [Google Scholar] [CrossRef]
- Merle, R.; Kösters, S.; May, T.; Portsch, U.; Blaha, T.; Kreienbrock, L. Serological Salmonella Monitoring in German Pig Herds: Results of the Years 2003–2008. Prev. Vet. Med. 2011, 99, 229–233. [Google Scholar] [CrossRef]
- Busch, H. Guideline Salmonella Monitoring Pigs. 17. Available online: https://www.q-s.de/monitoring-programmes/salmonella-monitoring.html#documents-pig-farming (accessed on 3 December 2022).
- QS Qualität und Sicherheit GmbH Anzahl Betriebe Mit Erhöhtem Salmonellenrisiko So Gering Wie Noch Nie. Available online: https://www.q-s.de/pressemeldungen/anzahl-betriebe-erhoehtes-salmonellenrisiko-gering.html (accessed on 3 December 2022).
- Casanova-Higes, A.; Marín-Alcalá, C.M.; Andrés-Barranco, S.; Cebollada-Solanas, A.; Alvarez, J.; Mainar-Jaime, R.C. Weaned Piglets: Another Factor to Be Considered for the Control of Salmonella Infection in Breeding Pig Farms. Vet. Res. 2019, 50, 45. [Google Scholar] [CrossRef] [PubMed]
- Bernad-Roche, M.; Casanova-Higes, A.; Marín-Alcalá, C.M.; Cebollada-Solanas, A.; Mainar-Jaime, R.C. Salmonella Infection in Nursery Piglets and Its Role in the Spread of Salmonellosis to Further Production Periods. Pathogens 2021, 10, 123. [Google Scholar] [CrossRef] [PubMed]
- Nollet, N.; Houf, K.; Dewulf, J.; De Kruif, A.; De Zutter, L.; Maes, D. Salmonella in Sows: A Longitudinal Study in Farrow-to-Finish Pig Herds. Vet. Res. 2005, 36, 645–656. [Google Scholar] [CrossRef] [PubMed]
- Parker, E.M.; Parker, A.J.; Short, G.; O’Connor, A.M.; Wittum, T.E. Salmonella Detection in Commercially Prepared Livestock Feed and the Raw Ingredients and Equipment Used to Manufacture the Feed: A Systematic Review and Meta-Analysis. Prev. Vet. Med. 2022, 198, 105546. [Google Scholar] [CrossRef]
- Davies, R.; Gosling, R.J.; Wales, A.D.; Smith, R.P. Use of an Attenuated Live Salmonella Typhimurium Vaccine on Three Breeding Pig Units: A Longitudinal Observational Field Study. Comp. Immunol. Microbiol. Infect. Dis. 2016, 46, 7–15. [Google Scholar] [CrossRef]
- Correia-Gomes, C.; Economou, T.; Mendonça, D.; Vieira-Pinto, M.; Niza-Ribeiro, J. Assessing Risk Profiles for Salmonella Serotypes in Breeding Pig Operations in Portugal Using a Bayesian Hierarchical Model. BMC Vet. Res. 2012, 8, 226. [Google Scholar] [CrossRef] [PubMed]
- Bonardi, S. Salmonella in the Pork Production Chain and Its Impact on Human Health in the European Union. Epidemiol. Infect. 2017, 145, 1513–1526. [Google Scholar] [CrossRef] [PubMed]
- Visscher, C.F.; Klein, G.; Verspohl, J.; Beyerbach, M.; Stratmann-Selke, J.; Kamphues, J. Serodiversity and Serological as Well as Cultural Distribution of Salmonella on Farms and in Abattoirs in Lower Saxony, Germany. Int. J. Food Microbiol. 2011, 146, 44–51. [Google Scholar] [CrossRef] [PubMed]
- Vangroenweghe, F. Relationship between Serological Status of Sows and the Assignment as Salmonella Risk Farm in the Belgian Salmonella Control Program. In Proceedings of the 21st IPVS Congress, Vancouver, BC, Canada, 18–21 July 2010; p. 177. [Google Scholar]
- Kranker, S.; Alban, L.; Boes, J.; Dahl, J. Longitudinal Study of Salmonella Enterica Serotype Typhimurium Infection in Three Danish Farrow-to-Finish Swine Herds. J. Clin. Microbiol. 2003, 41, 2282–2288. [Google Scholar] [CrossRef]
- Gotter, V.; Blaha, T.; Klein, G. A Case-Control Study on the Occurrence of Salmonella spp. in the Environment of Pigs. Epidemiol. Infect. 2012, 140, 150–156. [Google Scholar] [CrossRef]
- Pedersen, K.S.; Okholm, E.; Johansen, M.; Angen, Ø.; Jorsal, S.E.; Nielsen, J.P.; Bækbo, P. Clinical Utility and Performance of Sock Sampling in Weaner Pig Diarrhoea. Prev. Vet. Med. 2015, 120, 313–320. [Google Scholar] [CrossRef]
- Buhr, R.J.; Richardson, L.J.; Cason, J.A.; Cox, N.A.; Fairchild, B.D. Comparison of Four Sampling Methods for the Detection of Salmonella in Broiler Litter. Poult. Sci. 2007, 86, 21–25. [Google Scholar] [CrossRef]
- Davies, R.H.; Wales, A.D. Investigations into Salmonella Contamination in Poultry Feedmills in the United Kingdom: Salmonella in Poultry Feedmills. J. Appl. Microbiol. 2010, 109, 1430–1440. [Google Scholar] [CrossRef]
- Gradel, K.O.; Andersen, J.; Madsen, M. Comparisons of Sampling Procedures and Time of Sampling for the Detection of Salmonella in Danish Infected Chicken Flocks Raised in Floor Systems. Acta Veter-Scand. 2002, 43, 10. [Google Scholar]
- Cannon, R.M.; Roe, R.T. Livestock Disease Surveys: A Field Manual for Veterinarians; repr.; Australian Government Pub. Service: Canberra, Australia, 1986; ISBN 978-0-644-02101-2.
- Humphry, R.W.; Cameron, A.; Gunn, G.J. A Practical Approach to Calculate Sample Size for Herd Prevalence Surveys. Prev. Vet. Med. 2004, 65, 173–188. [Google Scholar] [CrossRef]
- Arnold, M.E.; Cook, A.; Davies, R. A Modelling Approach to Estimate the Sensitivity of Pooled Faecal Samples for Isolation of Salmonella in Pigs. J. R. Soc. Interface. 2005, 2, 365–372. [Google Scholar] [CrossRef][Green Version]
- Sanchez, J.; Dohoo, I.R.; Christensen, J.; Rajic, A. Factors Influencing the Prevalence of Salmonella Spp. in Swine Farms: A Meta-Analysis Approach. Prev. Vet. Med. 2007, 81, 148–177. [Google Scholar] [CrossRef] [PubMed]
- van der Wolf, P.; Koenders, K.; Gotter, V. Analysis of Sock, Swab and Fecal Samples for Presence of Salmonella from Sow Farms in The Netherlands. In Proceedings of the 9th European Symposium of Porcine Health Management, Prague, Czech Republic, 3–5 May 2017; p. 218. [Google Scholar]
- Belœil, P.-A.; Fravalo, P.; Fablet, C.; Jolly, J.-P.; Eveno, E.; Hascoet, Y.; Chauvin, C.; Salvat, G.; Madec, F. Risk Factors for Salmonella Enterica Subsp. Enterica Shedding by Market-Age Pigs in French Farrow-to-Finish Herds. Prev. Vet. Med. 2004, 63, 103–120. [Google Scholar] [CrossRef]
- Park, S.H.; Kim, H.J.; Cho, W.H.; Kim, J.H.; Oh, M.H.; Kim, S.H.; Lee, B.K.; Ricke, S.C.; Kim, H.Y. Identification of Salmonella Enterica Subspecies I, Salmonella Enterica Serovars Typhimurium, Enteritidis and Typhi Using Multiplex PCR. FEMS Microbiol. Lett. 2009, 301, 137–146. [Google Scholar] [CrossRef]
- Hedderich, J.; Sachs, L. Angewandte Statistik: Methodensammlung mit R; 16., überarbeitete und erweiterte Auflage; Springer Spektrum: Berlin, Germany, 2018; ISBN 978-3-662-56657-2. [Google Scholar]
- García-Feliz, C.; Carvajal, A.; Collazos, J.Á.; Rubio, P. Herd-Level Risk Factors for Faecal Shedding of Salmonella Enterica in Spanish Fattening Pigs. Prev. Vet. Med. 2009, 91, 130–136. [Google Scholar] [CrossRef] [PubMed]
- Davies, P.R.; Scott Hurd, H.; Funk, J.A.; Fedorka-Cray, P.J.; Jones, F.T. The Role of Contaminated Feed in the Epidemiology and Control of Salmonella Enterica in Pork Production. Foodborne Pathog. Dis. 2004, 1, 202–215. [Google Scholar] [CrossRef]
- Kolb, J.R.; Hoffman, L.J. Salmonella Choleraesuis as a Cause of Respiratory Disease in Growing and Finishing Swine. Iowa State Univ. Vet. 1990, 2, 66–68. [Google Scholar]
- Buch, J.-M.; Visscher, C.; Schulte zu Sundern, A.; Schulte-Wülwer, J.; Deermann, A.; Holling, C. Prevalence of Salmonella by Serological and Direct Detection Methods in Piglets from Inconspicuous, Conspicuous, and Vaccinated Sow Herds. Animals 2019, 10, 29. [Google Scholar] [CrossRef]
- Smith, R.P.; Andres, V.; Martelli, F.; Gosling, B.; Marco-Jimenez, F.; Vaughan, K.; Tchorzewska, M.; Davies, R. Maternal Vaccination as a Salmonella> Typhimurium Reduction Strategy on Pig Farms. J. Appl. Microbiol. 2018, 124, 274–285. [Google Scholar] [CrossRef] [PubMed]
- Owen, J.M. Disinfection of Farrowing Pens. Rev. Sci. Tech. OIE 1995, 14, 381–391. [Google Scholar] [CrossRef] [PubMed]
- Rho, M.-J.; Chung, M.-S.; Lee, J.-H.; Park, J. Monitoring of Microbial Hazards at Farms, Slaughterhouses, and Processing Lines of Swine in Korea. J. Food Prot. 2001, 64, 1388–1391. [Google Scholar] [CrossRef] [PubMed]
- Hoover, N.; Kenney, P.; Amick, J.; Hypes, W. Preharvest Sources of Salmonella Colonization in Turkey Production. Poult. Sci. 1997, 76, 1232–1238. [Google Scholar] [CrossRef]
- Renwick, S.A.; Irwin, R.J.; Clarke, R.C.; McNab, W.B.; Poppe, C.; McEwen, S.A. Epidemiological Associations between Characteristics of Registered Broiler Chicken Flocks in Canada and the Salmonella Culture Status of Floor Litter and Drinking Water. Can. Vet. J. 1992, 33, 449–458. [Google Scholar]
- Mannion, C.; Leonard, F.C.; Lynch, P.B.; Egan, J. Efficacy of Cleaning and Disinfection on Pig Farms in Ireland. Vet. Rec. 2007, 161, 371–375. [Google Scholar] [CrossRef] [PubMed]
- Gosling, R.J.; Mawhinney, I.; Vaughan, K.; Davies, R.H.; Smith, R.P. Efficacy of Disinfectants and Detergents Intended for a Pig Farm Environment Where Salmonella Is Present. Vet. Microbiol. 2017, 204, 46–53. [Google Scholar] [CrossRef]
- Jayaraman, B.; Nyachoti, C.M. Husbandry Practices and Gut Health Outcomes in Weaned Piglets: A Review. Anim. Nutr. 2017, 3, 205–211. [Google Scholar] [CrossRef]
- Lo Fo Wong, D.M.A.; Dahl, J.; Stege, H.; van der Wolf, P.J.; Leontides, L.; von Altrock, A.; Thorberg, B.M. Herd-Level Risk Factors for Subclinical Salmonella Infection in European Finishing-Pig Herds. Prev. Vet. Med. 2004, 62, 253–266. [Google Scholar] [CrossRef]
- Robertson, M.H. Survival of S. Typhimurium in Floor Dust—A Possible Reservoir of Infection in Institutions. Public Health 1972, 87, 39–45. [Google Scholar] [CrossRef]
- Schilling, T.; Hoelzle, K.; Philipp, W.; Hoelzle, L.E. Survival of Salmonella Typhimurium, Listeria Monocytogenes, and ESBL Carrying Escherichia Coli in Stored Anaerobic Biogas Digestates in Relation to Different Biogas Input Materials and Storage Temperatures. Agriculture 2022, 12, 67. [Google Scholar] [CrossRef]
- Lynch, H.; Leonard, F.C.; Walia, K.; Lawlor, P.G.; Duffy, G.; Fanning, S.; Markey, B.K.; Brady, C.; Gardiner, G.E.; Argüello, H. Investigation of In-Feed Organic Acids as a Low Cost Strategy to Combat Salmonella in Grower Pigs. Prev. Vet. Med. 2017, 139, 50–57. [Google Scholar] [CrossRef] [PubMed]
- Calveyra, J.C.; Nogueira, M.G.; Kich, J.D.; Biesus, L.L.; Vizzotto, R.; Berno, L.; Coldebella, A.; Lopes, L.; Morés, N.; Lima, G.J.M.M.; et al. Effect of Organic Acids and Mannanoligosaccharide on Excretion of Salmonella Typhimurium in Experimentally Infected Growing Pigs. Res. Vet. Sci. 2012, 93, 46–47. [Google Scholar] [CrossRef]
- Ferrari, R.G.; Rosario, D.K.A.; Cunha-Neto, A.; Mano, S.B.; Figueiredo, E.E.S.; Conte-Junior, C.A. Worldwide Epidemiology of Salmonella Serovars in Animal-Based Foods: A Meta-Analysis. Appl. Environ. Microbiol. 2019, 85, e00591-19. [Google Scholar] [CrossRef]
- Nollet, N.; Maes, D.; De Zutter, L.; Duchateau, L.; Houf, K.; Huysmans, K.; Imberechts, H.; Geers, R.; de Kruif, A.; Van Hoof, J. Risk Factors for the Herd-Level Bacteriologic Prevalence of Salmonella in Belgian Slaughter Pigs. Prev. Vet. Med. 2004, 65, 63–75. [Google Scholar] [CrossRef] [PubMed]
- Davison, H.C.; Smith, R.P.; Pascoe, S.J.S.; Sayers, A.R.; Davies, R.H.; Weaver, J.P.; Kidd, S.A.; Dalziel, R.W.; Evans, S.J. Prevalence, Incidence and Geographical Distribution of Serovars of Salmonella on Dairy Farms in England and Wales. Vet. Rec. 2005, 157, 703–711. [Google Scholar] [CrossRef]
- Peeters, L.; Dewulf, J.; Boyen, F.; Brossé, C.; Vandersmissen, T.; Rasschaert, G.; Heyndrickx, M.; Cargnel, M.; Pasmans, F.; Maes, D. Influence of Different Vaccination Strategies against Salmonella Typhimurium in Pig Farms on the Number of Carriers in Ileocecal Lymph Nodes. In Proceedings of the International Conference on the Epidemiology and Control of Biological, Chemical and Physical Hazards in Pigs and Pork; Iowa State University, Ames, IA, USA, 21–24 August 2017; Digital Press: Porto, Portugal, 2017; pp. 105–106. [Google Scholar]
- Lindner, T.; Springer, S.; Selbitz, H.-J. The Use of a Salmonella Typhimurium Live Vaccine to Control Salmonella Typhimurium in Fattening Pigs in Field and Effects on Serological Surveillance. In Proceedings of the International Conference on the Epidemiology and Control of Biological, Chemical and Physical Hazards in Pigs and Pork; Iowa State University, Ames, IA, USA, 1 January 2007; Digital Press: Verona, Italy, 2007; pp. 237–239. [Google Scholar]
- De Ridder, L.; Maes, D.; Dewulf, J.; Pasmans, F.; Boyen, F.; Haesebrouck, F.; Méroc, E.; Butaye, P.; Van der Stede, Y. Evaluation of Three Intervention Strategies to Reduce the Transmission of Salmonella Typhimurium in Pigs. Vet. J. 2013, 197, 613–618. [Google Scholar] [CrossRef] [PubMed]
- Peeters, L.; Dewulf, J.; Boyen, F.; Brossé, C.; Vandersmissen, T.; Rasschaert, G.; Heyndrickx, M.; Cargnel, M.; Mattheus, W.; Pasmans, F.; et al. Bacteriological Evaluation of Vaccination against Salmonella Typhimurium with an Attenuated Vaccine in Subclinically Infected Pig Herds. Prev. Vet. Med. 2020, 182, 104687. [Google Scholar] [CrossRef]
- Bearson, S.M.D. Salmonella in Swine: Prevalence, Multidrug Resistance, and Vaccination Strategies. Annu. Rev. Anim. Biosci. 2022, 10, 373–393. [Google Scholar] [CrossRef]
- Visscher, C.F.; Winter, P.; Verspohl, J.; Stratmann-Selke, J.; Upmann, M.; Beyerbach, M.; Kamphues, J. Effects of Feed Particle Size at Dietary Presence of Added Organic Acids on Caecal Parameters and the Prevalence of Salmonella in Fattening Pigs on Farm and at Slaughter. J. Anim. Physiol. Anim. Nutr. 2009, 93, 423–430. [Google Scholar] [CrossRef]
- Creus, E.; Pérez, J.F.; Peralta, B.; Baucells, F.; Mateu, E. Effect of Acidified Feed on the Prevalence of Salmonella in Market-Age Pigs. Zoonoses Public Health 2007, 54, 314–319. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).