Single Nucleotide Polymorphisms in Genes Encoding Toll-Like Receptors 7 and 8 and Their Association with Proviral Load of SRLVs in Goats of Polish Carpathian Breed
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals and Sample Preparation
2.2. SNP Identification–Variant Calling
2.3. SNP Genotyping
2.4. Poviral Load Quantification
2.5. Statistical Analysis
3. Results
3.1. Serology and qPCR
3.2. TLR7 Gene-Detected SNPs, Allele and Haplotype Frequency
3.3. TLR8 Gene-Detected SNPs, Allele and Haplotype Frequency
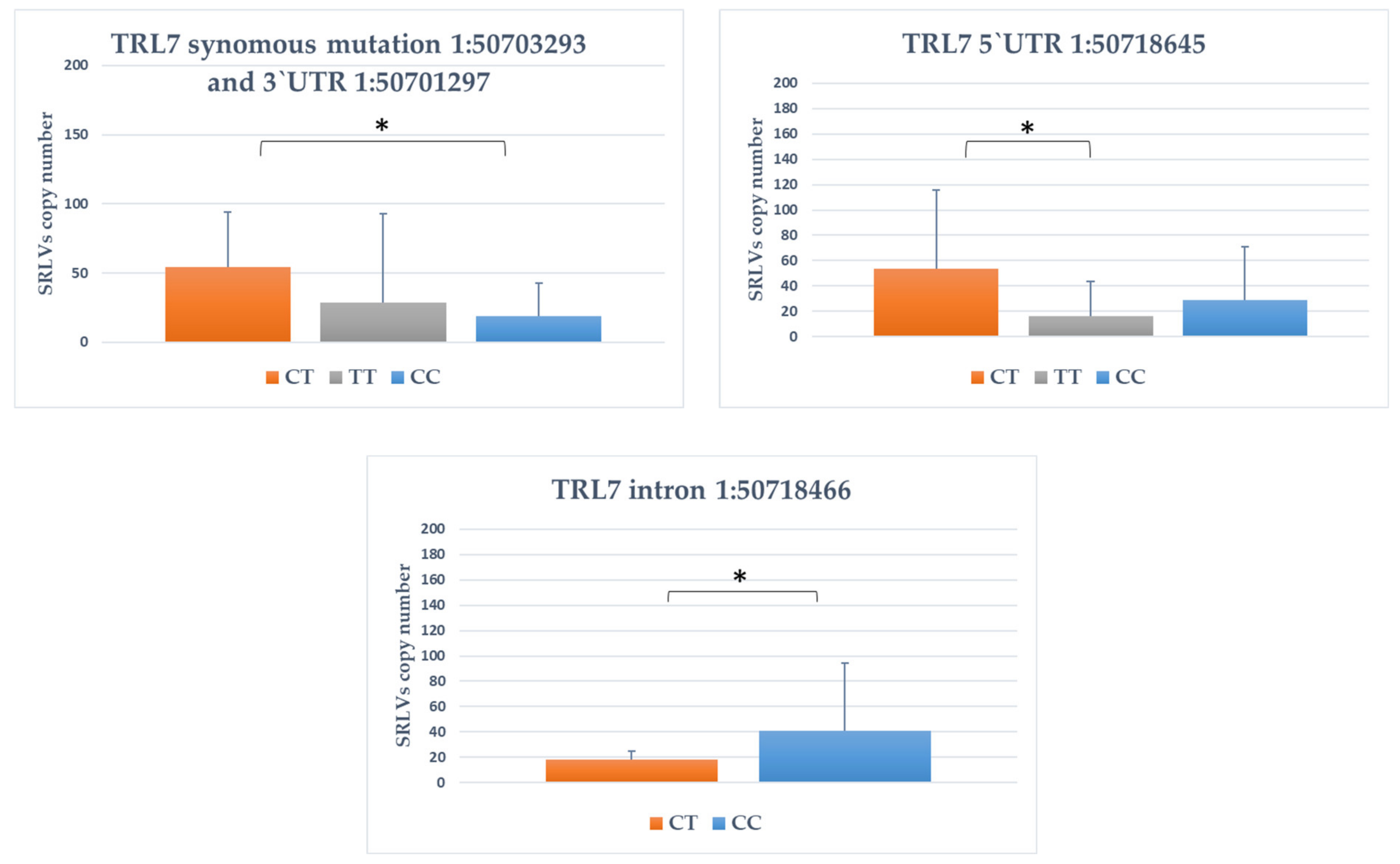
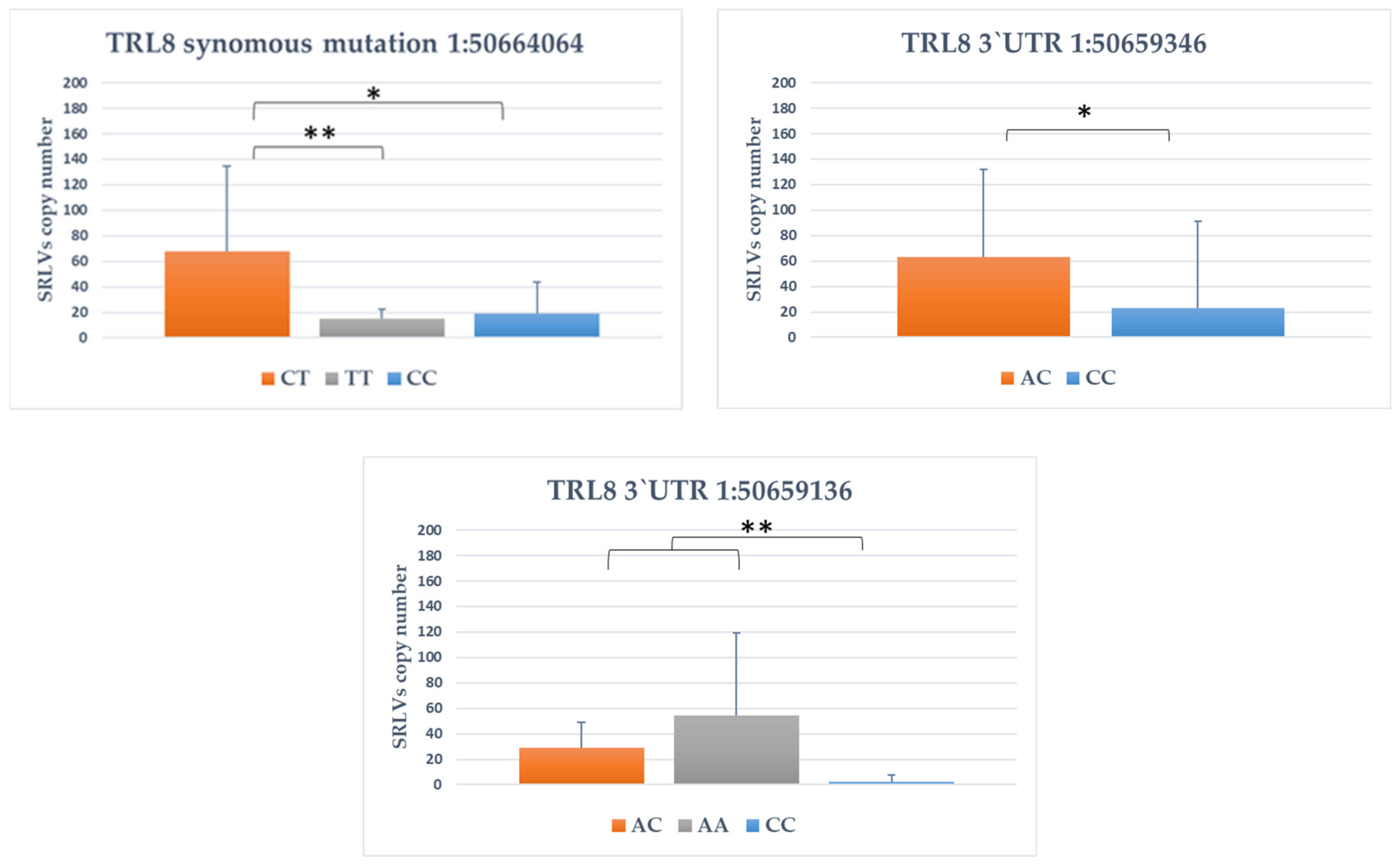
3.4. Association between SNPs and Provirus Copy Number
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Polledo, L.; González, J.; Benavides, J.; Morales, S.; Martínez-Fernández, B.; Delgado, L.; Reina, R.; Glaria, I.; Pérez, V.; Ferreras, M.C.; et al. Patterns of lesion and local host cellular immune response in natural cases of ovine maedi-visna. J. Comp. Pathol. 2012, 147, 1–10. [Google Scholar] [CrossRef]
- Minguijón, E.; Reina, R.; Pérez, M.; Polledo, L.; Villoria, M.; Ramírez, H.; Leginagoikoa, I.; Badiola, J.J.; García-Marín, J.F.; De Andrés, D.; et al. Small ruminant lentivirus infections and diseases. Vet. Microbiol. 2015, 181, 75–89. [Google Scholar] [CrossRef] [PubMed]
- Kalogianni, A.I.; Bossis, I.; Ekateriniadou, L.V.; Gelasakis, A.I. Etiology, Epizootiology and Control of Maedi-Visna in Dairy Sheep: A Review. Animals 2020, 10, 616. [Google Scholar] [CrossRef] [PubMed]
- Reina, R.; Berriatua, E.; Luján, L.; Juste, R.; Sánchez, A.; De Andrés, D.; Amorena, B. Prevention strategies against small ruminant lentiviruses: An update. Vet. J. 2009, 182, 31–37. [Google Scholar] [CrossRef] [PubMed]
- Pérez, M.; Biescas, E.; De Andrés, X.; Leginagoikoa, I.; Salazar, E.; Berriatua, E.; Reina, R.; Bolea, R.; De Andrés, D.; Juste, R.A.; et al. Visna/maedi virus serology in sheep: Survey, risk factors and implementation of a successful control programme in Aragón (Spain). Vet. J. 2010, 186, 221–225. [Google Scholar] [CrossRef] [PubMed]
- Olech, M.; Osiński, Z.; Kuźmak, J. Bayesian estimation of seroprevalence of small ruminant lentiviruses in sheep from Poland. Prev. Vet. Med. 2017, 147, 66–78. [Google Scholar] [CrossRef] [PubMed]
- Kaba, J.; Czopowicz, M.; Ganter, M.; Nowicki, M.; Witkowski, L.; Nowicka, D.; Szaluś-Jordanow, O. Risk factors associated with seropositivity to small ruminant lentiviruses in goat herds. Res. Vet. Sci. 2013, 94, 225–227. [Google Scholar] [CrossRef]
- Ravazzolo, A.P.; Nenci, C.; Vogt, H.R.; Waldvogel, A.; Obexer-Ruff, G.; Peterhans, E.; Bertoni, G. Viral load, organ distribution, histopathological lesions, and cytokine mRNA expression in goats infected with a molecular clone of the caprine arthritis encephalitis virus. Virology 2006, 350, 116–127. [Google Scholar] [CrossRef] [PubMed]
- Herrmann-Hoesing, L.M.; Noh, S.M.; White, S.N.; Snekvik, K.R.; Truscott, T.; Knowles, D.P. Peripheral ovine progressive pneumonia provirus levels correlate with and predict histological tissue lesion severity in naturally infected sheep. Clin. Vaccine Immunol. 2009, 16, 551–557. [Google Scholar] [CrossRef][Green Version]
- Crespo, H.; Bertolotti, L.; Proffiti, M.; Cascio, P.; Cerruti, F.; Acutis, P.L.; De Andrés, D.; Reina, R.; Rosati, S. Low proviral small ruminant lentivirus load as biomarker of natural restriction in goats. Vet. Microbiol. 2016, 192, 152–162. [Google Scholar] [CrossRef]
- Stonos, N.; Wootton, S.K.; Karrow, N. Immunogenetics of small ruminant lentiviral infections. Viruses 2014, 6, 3311–3333. [Google Scholar] [CrossRef] [PubMed]
- Cardinaux, L.; Zahno, M.L.; Deubelbeiss, M.; Zanoni, R.; Vogt, H.R.; Bertoni, G. Virological and phylogenetic characterization of attenuated small ruminant lentivirus isolates eluding efficient serological detection. Vet. Microbiol. 2013, 162, 572–581. [Google Scholar] [CrossRef]
- Larruskain, A.; Bernales, I.; Luján, L.; De Andrés, D.; Amorena, B.; Jugo, B.M. Expression analysis of 13 ovine immune response candidate genes in Visna/Maedi disease progression. Comp. Immunol. Microbiol. Infect. Dis. 2013, 36, 405–413. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Lucia, E.; Barquero, N.; Domenech, A. Maedi-Visna virus: Current perspectives. Vet. Med. 2018, 21, 11–21. [Google Scholar] [CrossRef] [PubMed]
- Skevaki, C.; Pararas, M.; Kostelidou, K.; Tsakris, A.; Routsias, J.G. Single nucleotide polymorphisms of Toll-like receptors and susceptibility to infectious diseases. Clin. Exp. Immunol. 2015, 180, 165–177. [Google Scholar] [CrossRef]
- Lester, S.N.; Li, K. Toll-like receptors in antiviral innate immunity. J. Mol. Biol. 2014, 426, 1246–1264. [Google Scholar] [CrossRef]
- Heil, F.; Hemmi, H.; Hochrein, H.; Ampenberger, F.; Kirschning, C.; Akira, S.; Lipford, G.; Wagner, H.; Bauer, S. Species-specific recognition of single-stranded RNA via toll-like receptor 7 and 8. Science 2004, 303, 1526–1529. [Google Scholar] [CrossRef]
- Akira, S.; Takeda, K. Toll-like receptor signalling. Nat. Rev. Immunol. 2004, 4, 499–511. [Google Scholar] [CrossRef]
- De Pablo-Maiso, L.; Doménech, A.; Echeverría, I.; Gómez-Arrebola, C.; De Andrés, D.; Rosati, S.; Gómez-Lucia, E.; Reina, R. Prospects in Innate Immune Responses as Potential Control Strategies against Non-Primate Lentiviruses. Viruses 2018, 10, 435. [Google Scholar] [CrossRef]
- Jarczak, J.; Kaba, J.; Reczyńska, D.; Bagnicka, E. Impaired Expression of Cytokines as a Result of Viral Infections with an Emphasis on Small Ruminant Lentivirus Infection in Goats. Viruses 2016, 8, 186. [Google Scholar] [CrossRef]
- Blacklaws, B.A. Small ruminant lentiviruses: Immunopathogenesis of visna-maedi and caprine arthritis and encephalitis virus. Comp. Immunol. Microbiol. Infect. Dis. 2012, 35, 259–269. [Google Scholar] [CrossRef]
- Fang, X.; Liu, X.; Meng, C.; Fu, Y.; Wang, X.; Li, B.; Tu, F.; Zhao, F.; Ren, S. Breed-linked polymorphisms of porcine toll-like receptor 2 (TLR2) and TLR4 and the primary investigation on their relationship with prevention against Mycoplasma pneumoniae and bacterial LPS challenge. Immunogenetics 2013, 65, 829–834. [Google Scholar] [CrossRef]
- Kloch, A.; Wenzel, M.A.; Laetsch, D.R.; Michalski, O.; Bajer, A.; Behnke, J.M.; Welc-Falęciak, R.; Piertney, S.B. Signatures of balancing selection in toll-like receptor (TLRs) genes–novel insights from a free-living rodent. Sci. Rep. 2018, 8, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Novák, K. Functional polymorphisms in Toll-like receptor genes for innate immunity in farm animals. Vet. Immunol. Immunopathol. 2014, 157, 1–11. [Google Scholar] [CrossRef]
- Beutler, B.; Jiang, Z.; Georgel, P.; Crozat, K.; Croker, B.; Rutschmann, S.; Du, X.; Hoebe, K. Genetic analysis of host resistance: Toll-like receptor signaling and immunity at large. Annu. Rev. Immunol. 2006, 24, 353–389. [Google Scholar] [CrossRef]
- Dutta, S.K.; Tripathi, A. Association of toll-like receptor polymorphisms with susceptibility to chikungunya virus infection. Virology 2017, 511, 207–213. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.H.; Eng, H.L.; Lin, K.H.; Chang, C.H.; Hsieh, C.A.; Lin, Y.L.; Lin, T.M. TLR7 and TLR8 gene variations and susceptibility to hepatitis C virus infection. PLoS ONE 2011, 6, e26235. [Google Scholar] [CrossRef]
- Zhang, T.; Zhu, J.; Su, B.; Cao, L.; Li, Z.; Wei, H.; Huang, X.; Zheng, K.; Li, A.; Chen, N.; et al. Effects of TLR7 Polymorphisms on the Susceptibility and Progression of HIV-1 Infection in Chinese MSM Population. Front. Immunol. 2020, 11, 589010. [Google Scholar] [CrossRef]
- Arslan, S.; Engin, A.; Özbilüm, N.; Bakır, M. Toll-like receptor 7 Gln11Leu, c.4-151A/G, and +1817G/T polymorphisms in Crimean Congo hemorrhagic fever. J. Med. Virol. 2015, 87, 1090–1095. [Google Scholar] [CrossRef] [PubMed]
- Mikula, I.; Bhide, M.; Pastorekova, S.; Mikula, I. Characterization of ovine TLR7 and TLR8 protein coding regions, detection of mutations and Maedi Visna virus infection. Vet. Immunol. Immunopathol. 2010, 138, 51–59. [Google Scholar] [CrossRef]
- Mikula, I., Jr.; Mikula, I., Sr. Characterization of ovine Toll-like receptor 9 protein coding region, comparative analysis, detection of mutations and maedi visna infection. Dev. Comp. Immunol. 2011, 35, 182–192. [Google Scholar] [CrossRef]
- De Sena Brandine, G.; Smith, A.D. Falco: High-speed FastQC emulation for quality control of sequencing data. F1000Research 2019, 8, 1874. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Ghosh, S.; Chan, C.K. Analysis of RNA-Seq Data Using TopHat and Cufflinks. Methods Mol. Biol. 2016, 1374, 339–361. [Google Scholar]
- Sánchez-Sevilla, J.F.; Vallarino, J.G.; Osorio, S.; Bombarely, A.; Posé, D.; Merchante, C.; Botella, M.A.; Amaya, I.; Valpuesta, V. Gene expression atlas of fruit ripening and transcriptome assembly from RNA-seq data in octoploid strawberry (Fragaria × ananassa). Sci. Rep. 2017, 7, 13737. [Google Scholar] [CrossRef]
- Danecek, P.; Auton, A.; Abecasis, G.; Albers, C.A.; Banks, E.; DePristo, M.A.; Handsaker, R.E.; Lunter, G.; Marth, G.T.; Sherry, S.T.; et al. 1000 Genomes Project Analysis Group. The variant call format and VCFtools. Bioinformatics 2011, 27, 2156–2158. [Google Scholar] [CrossRef] [PubMed]
- McLaren, W.; Gil, L.; Hunt, S.E.; Riat, H.S.; Ritchie, G.R.; Thormann, A.; Flicek, P.; Cunningham, F. The Ensembl Variant Effect Predictor. Genome Biol. 2016, 17, 122. [Google Scholar] [CrossRef]
- Olech, M.; Valas, S.; Kuźmak, J. Epidemiological survey in single-species flocks from Poland reveals expanded genetic and antigenic diversity of small ruminant lentiviruses. PLoS ONE 2018, 13, e0193892. [Google Scholar] [CrossRef]
- Olech, M.; Kuźmak, J. Molecular Characterization of Small Ruminant Lentiviruses of Subtype A5 Detected in Naturally Infected but Clinically Healthy Goats of Carpathian Breed. Pathogens 2020, 9, 992. [Google Scholar] [CrossRef] [PubMed]
- Gabriel, S.B.; Schaffner, S.F.; Nguyen, H.; Moore, J.M.; Roy, J.; Blumenstiel, B.; Higgins, J.; DeFelice, M.; Lochner, A.; Faggart, M.; et al. The structure of haplotype blocks in the human genome. Science 2002, 296, 2225–2229. [Google Scholar] [CrossRef] [PubMed]
- Sauna, Z.E.; Kimchi-Sarfaty, C. Understanding the contribution of synonymous mutations to human disease. Nat. Rev. Genet. 2011, 12, 683–691. [Google Scholar] [CrossRef]
- Fåhraeus, R.; Marin, M.; Olivares-Illana, V. Whisper mutations: Cryptic messages within the genetic code. Oncogene 2016, 35, 3753–3759. [Google Scholar] [CrossRef]
- Hunt, R.C.; Simhadri, V.L.; Iandoli, M.; Sauna, Z.E.; Kimchi-Sarfaty, C. Exposing synonymous mutations. Trends Genet. 2014, 30, 308–321. [Google Scholar] [CrossRef]
- Fernández-Calero, T.; Cabrera-Cabrera, F.; Ehrlich, R.; Marín, M. Silent Polymorphisms: Can the tRNA Population Explain Changes in Protein Properties? Life 2016, 6, 9. [Google Scholar] [CrossRef]
- Diederichs, S.; Bartsch, L.; Berkmann, J.C.; Fröse, K.; Heitmann, J.; Hoppe, C.; Iggena, D.; Jazmati, D.; Karschnia, P.; Linsenmeier, M.; et al. The dark matter of the cancer genome: Aberrations in regulatory elements, untranslated regions, splice sites, non-coding RNA and synonymous mutations. EMBO Mol. Med. 2016, 8, 442–457. [Google Scholar] [CrossRef]
- Carmody, S.R.; Wente, S.R. mRNA nuclear export at a glance. J. Cell Sci. 2009, 122, 1933–1937. [Google Scholar] [CrossRef]
- Pickering, B.M.; Willis, A.E. The implications of structured 5′ untranslated regions on translation and disease. Semin. Cell Dev. Biol. 2005, 16, 39–47. [Google Scholar] [CrossRef]
- Georges, M.; Clop, A.; Marcq, F.; Takeda, H.; Pirottin, D.; Hiard, S.; Tordoir, X.; Caiment, F.; Meish, F.; Bibé, B.; et al. Polymorphic microRNA-target interactions: A novel source of phenotypic variation. Cold Spring Harb. Symp. Quant. Biol. 2006, 71, 343–350. [Google Scholar] [CrossRef][Green Version]
- Conne, B.; Stutz, A.; Vassalli, J.D. The 3′ untranslated region of messenger RNA: A molecular ‘hotspot’ for pathology? Nat. Med. 2000, 6, 637–641. [Google Scholar] [CrossRef]
- Herrmann-Hoesing, L.M.; White, S.N.; Mousel, M.R.; Lewis, G.S.; Knowles, D.P. Ovine progressive pneumonia provirus levels associate with breed and Ovar-DRB1. Immunogenetics 2008, 60, 749–758. [Google Scholar] [CrossRef]
- White, S.N.; Mousel, M.R.; Reynolds, J.O.; Lewis, G.S.; Herrmann-Hoesing, L.M. Common promoter deletion is associated with 3.9-fold differential transcription of ovine CCR5 and reduced proviral level of ovine progressive pneumonia virus. Anim. Genet. 2009, 40, 583–589. [Google Scholar] [CrossRef]
- Alshanbari, F.A.; Mousel, M.R.; Reynolds, J.O.; Herrmann-Hoesing, L.M.; Highland, M.A.; Lewis, G.S.; White, S.N. Mutations in Ovis aries TMEM154 are associated with lower small ruminant lentivirus proviral concentration in one sheep flock. Anim. Genet. 2014, 45, 565–571. [Google Scholar] [CrossRef] [PubMed]
- Said, E.A.; Al-Yafei, F.; Zadjali, F.; Hasson, S.S.; Al-Balushi, M.S.; Al-Mahruqi, S.; Koh, C.Y.; Al-Naamani, K.; Al-Busaidi, J.Z.; Idris, M.A.; et al. Association of single-nucleotide polymorphisms in TLR7 (Gln11Leu) and TLR9 (1635A/G) with a higher CD4T cell count during HIV infection. Immunol. Lett. 2014, 160, 58–64. [Google Scholar] [CrossRef]
- Anokhin, V.V.; Bakhteeva, L.B.; Khasanova, G.R.; Khaiboullina, S.F.; Martynova, E.V.; Tillett, R.L.; Schlauch, K.A.; Lombardi, V.C.; Rizvanov, A.A. Previously Unidentified Single Nucleotide Polymorphisms in HIV/AIDS Cases Associate with Clinical Parameters and Disease Progression. Biomed. Res. Int. 2016, 2016, 2742648. [Google Scholar] [CrossRef]
- Browne, E.P. The Role of Toll-Like Receptors in Retroviral Infection. Microorganisms 2020, 8, 1787. [Google Scholar] [CrossRef]
- Oh, D.Y.; Taube, S.; Hamouda, O.; Kücherer, C.; Poggensee, G.; Jessen, H.; Eckert, J.K.; Neumann, K.; Storek, A.; Pouliot, M.; et al. A functional toll-like receptor 8 variant is associated with HIV disease restriction. J. Infect. Dis. 2008, 198, 701–709. [Google Scholar] [CrossRef]
- Larruskain, A.; Jugo, B.M. Retroviral infections in sheep and goats: Small ruminant lentiviruses and host interaction. Viruses 2013, 5, 2043–2061. [Google Scholar] [CrossRef]
- Murphy, B.; Hillman, C.; Castillo, D.; Vapniarsky, N.; Rowe, J. The presence or absence of the gamma-activated site determines IFN gamma-mediated transcriptional activation in CAEV promoters cloned from the mammary gland and joint synovium of a single CAEV-infected goat. Virus Res. 2012, 163, 537–545. [Google Scholar] [CrossRef]
- Willie, B.; Hall, N.B.; Stein, C.M.; Jurevic, R.J.; Weinberg, A.; Mehlotra, R.K.; Zimmerman, P.A. Association of Toll-like receptor polymorphisms with HIV status in North Americans. Genes Immun. 2014, 15, 569–577. [Google Scholar] [CrossRef]
- Peterhans, E.; Greenland, T.; Badiola, J.; Harkiss, G.; Bertoni, G.; Amorena, B.; Eliaszewicz, M.; Juste, R.A.; Krassnig, R.; Lafont, J.-P.; et al. Routes of transmission and consequences of small ruminant lentiviruses (SRLVs) infection and eradication schemes. Vet. Res. 2004, 35, 257–274. [Google Scholar] [CrossRef] [PubMed]

| Reference Sequence | Sequences of Primers (5′-3′) | Orientation | Size | Identified SNP Based on RNA-Seq Data |
|---|---|---|---|---|
| TLR7 ENSCHIG00000021012 | GACATGAGGTTGCCCTGATTG | F | 338 bp | LWLT01000021.1_50718645_C/T |
| TATGATTTGGGTCCCTTCCCC | R | |||
| TCTCTGAGTTTCTTACCTTTGGGA | F | 285 bp | LWLT01000021.1_50703293_T/C | |
| GCTGAGGTCCAGATGTCGC | R | |||
| CTTGATCGGCTCACCCATGC | F | 297 bp | LWLT01000021.1_50702074_T/C | |
| AAGTGCAGTTCTTGGTGACTG | R | |||
| ACCTGCTTAAATATGGCTCTGG | F | 418 bp | LWLT01000021.1_50701297_C/T | |
| TGGATGAGGCAACTTTTCTTTGG | R | |||
| TLR8 ENSCHIG00000016625 | GGGCATTTCTCAACCTCAAA | F | 316 bp | LWLT01000021.1_50666071_A/G |
| GTTAGCGAGCTTGGCAGACT | R | |||
| AATGCGCACTATTTCCGAAT | F | 301 bp | LWLT01000021.1_50664682_A/G | |
| TCCTGGGCAAGTTAAGGAAG | R | |||
| GCAGCCTGATACACCTCGAT | F | 424 bp | LWLT01000021.1_50664208_C/T LWLT01000021.1_50664064_C/T | |
| TGGGATGTGGACAGAGACCT | R | |||
| CTGATGTTGCTGGGAGTCCT | F | 576 bp | LWLT01000021.1_50659559_C/T LWLT01000021.1_50659202_T/C | |
| TAGCTGAGAGGGGAATTGCC | R |
| Location | Mutant Allele | Variant Type | Gene Reference | Transcript Reference | cDNA Position | Codons | |
|---|---|---|---|---|---|---|---|
| TLR7 | |||||||
| LWLT01000021.1_50701297_C/T | LWLT01000021.1:50701297-50701297 | T | 3_prime_UTR_variant | ENSCHIG00000021012 | ENSCHIT00000031299.1 | 4304 | - |
| LWLT01000021.1_50702074_T/C | LWLT01000021.1:50702074-50702074 | C | 3_prime_UTR_variant | ENSCHIG00000021012 | ENSCHIT00000031299.1 | 3527 | - |
| LWLT01000021.1_50703293_T/C | LWLT01000021.1:50703293-50703293 | C | synonymous_variant | ENSCHIG00000021012 | ENSCHIT00000031309.1 | 2088 | ggA/ggG |
| LWLT01000021.1_50718645_C/T | LWLT01000021.1:50718645-50718645 | T | 5_prime_UTR_variant | ENSCHIG00000021012 | ENSCHIT00000031299.1 | 23 | - |
| TLR8 | |||||||
| LWLT01000021.1_50659202_T/C | LWLT01000021.1:50659202-50659202 | C | 3_prime_UTR_variant | ENSCHIG00000016625 | ENSCHIT00000024133.1 | 4708 | - |
| LWLT01000021.1_50659559_C/T | LWLT01000021.1:50659559-50659559 | T | 3_prime_UTR_variant | ENSCHIG00000016625 | ENSCHIT00000024133.1 | 4351 | - |
| LWLT01000021.1_50664064_C/T | LWLT01000021.1:50664064-50664064 | T | synonymous_variant | ENSCHIG00000016625 | ENSCHIT00000024133.1 | 2460 | gcG/gcA |
| LWLT01000021.1_50664208_G/A | LWLT01000021.1:50664208-50664208 | A | synonymous_variant | ENSCHIG00000016625 | ENSCHIT00000024133.1 | 2316 | ttC/ttT |
| LWLT01000021.1_50664682_A/G | LWLT01000021.1:50664682-50664682 | G | synonymous_variant | ENSCHIG00000016625 | ENSCHIT00000024133.1 | 1842 | ctT/ctC |
| LWLT01000021.1_50666071_A/G | LWLT01000021.1:50666071-50666071 | G | synonymous_variant | ENSCHIG00000016625 | ENSCHIT00000024133.1 | 453 | aaT/aaC |
| Location | Mutant Allele | Variant Type | Gene Reference | Transcript Reference | cDNA Position | |
|---|---|---|---|---|---|---|
| TLR7 | ||||||
| LWLT01000021.1_50718760_G/A | LWLT01000021.1:50718760-1: 50718760 | G | Promoter | ENSCHIG00000021012 | ENSCHIT00000031299.1 | - |
| LWLT01000021.1_0718466_C/T | LWLT01000021.1:50717866-50717866 | C | Intron 1 | ENSCHIG00000021012 | ENSCHIT00000031299.1 | - |
| TLR8 | ||||||
| LWLT01000021.1_50659346_C/A | LWLT01000021.1: 50659346-50659346 | C | 3_prime_UTR_variant | ENSCHIG00000016625 | ENSCHIT00000024133.1 | 4564 |
| LWLT01000021.1_50659136_A/C | LWLT01000021.1: 50659136-50659136 | A | 3_prime_UTR_variant | ENSCHIG00000016625 | ENSCHIT00000024133.1 | 4774 |
| SNP | Genotypes | Alleles | HWE p Value | |||
|---|---|---|---|---|---|---|
| CC | CT | TT | C | T | ||
| SV T/C 1:50703293 | 11 (31%) | 16 (46%) | 5 (23%) | 38 (0.54) | 26 (0.46) | ns |
| CC | CT | TT | C | T | ||
| 3′UTR C/T 1:50701297 | 5 (23%) | 16 (46%) | 11 (31%) | 38 (0.54) | 26 (0.46) | ns |
| CC | CT | TT | C | T | ||
| 5′UTR C/T 1:50718645 | 5 (16%) | 17 (53%) | 10 (31%) | 27 (0.42) | 37 (0.58) | ns |
| CC | CT | TT | C | T | ||
| INTRON1 C/T 1:50718466 | 29 (91%) | 2 (6%) | 1 (3%) | 58 (0.94) | 6 (0.06) | 0.00001 |
| Haplotype | Frequency | S.E. | |
|---|---|---|---|
| TLR7 | |||
| 1 | T/T/C/T/A/C | 51% | 0.003 |
| 2 | C/T/T/C/A/C | 29% | 0.004 |
| 3 | C/T/T/C/A/T | 10% | 0.004 |
| 4 | T/T/T/C/A/C | 4% | 0.002 |
| 5 | T/T/C/C/A/C | 4% | 0.0003 |
| 6 | T/T/C/T/G/C | 1.20% | 0.000 |
| 7 | T/T/T/C/A/T | <1% | 0.002 |
| 8 | T/T/C/T/A/T | <1% | 0.003 |
| TLR8 | |||
| 1 | C/C/C/G/A/A/C/A | 29.70% | 0.010 |
| 2 | C/T/T/A/A/G/A/A | 19.90% | 0.003 |
| 3 | C/C/C/G/A/A/C/C | 14.80% | 0.008 |
| 4 | C/T/C/G/A/A/C/A | 13.40% | 0.007 |
| 5 | C/T/C/G/A/A/C/C | 13.30% | 0.008 |
| 6 | C/T/T/A/A/G/C/A | 5.70% | 0.001 |
| 7 | C/T/C/G/G/A/C/A | 2.70% | 0.003 |
| 8 | C/C/T/A/A/G/A/A | <1% | 0.002 |
| 9 | C/T/C/G/G/A/C/C | <1% | 0.003 |
| 10 | C/T/T/A/A/G/A/C | <1% | 0.002 |
| SNP | Genotypes | Alleles | HWE p Value | |||
|---|---|---|---|---|---|---|
| AA | GA | GG | A | G | ||
| SV A/G 1:50666071 | 18 (56%) | 11 (34%) | 3 (10%) | 47 (0.73) | 17 (0.27) | ns |
| CC | CT | TT | G | A | ||
| SV G/A 1:506642208 | 18 (56%) | 11 (34%) | 3 (10%) | 47 (0.73) | 17 (0.27) | ns |
| CC | CT | TT | C | T | ||
| SV C/T 1:50664064 | 18 (56%) | 11 (34%) | 3 (10%) | 47 (0.73) | 17 (0.27) | ns |
| AA | AC | CC | A | C | ||
| 3′UTR A/C 1:50659136 | 17 (55%) | 10 (32%) | 4 (13%) | 42 (0.71) | 18 (0.29) | ns |
| CC | AC | AA | C | A | ||
| 3′UTR C/A 1:50659346 | 17 (55%) | 13 (42%) | 1 (3%) | 47 (0.76) | 15 (0.24) | ns |
| CC | TC | TT | C | T | ||
| 3″UTR C/T 1:50659559 | 15 (48%) | 13 (42%) | 3 (10%) | 43 (0.69) | 19 (0.31) | ns |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Olech, M.; Ropka-Molik, K.; Szmatoła, T.; Piórkowska, K.; Kuźmak, J. Single Nucleotide Polymorphisms in Genes Encoding Toll-Like Receptors 7 and 8 and Their Association with Proviral Load of SRLVs in Goats of Polish Carpathian Breed. Animals 2021, 11, 1908. https://doi.org/10.3390/ani11071908
Olech M, Ropka-Molik K, Szmatoła T, Piórkowska K, Kuźmak J. Single Nucleotide Polymorphisms in Genes Encoding Toll-Like Receptors 7 and 8 and Their Association with Proviral Load of SRLVs in Goats of Polish Carpathian Breed. Animals. 2021; 11(7):1908. https://doi.org/10.3390/ani11071908
Chicago/Turabian StyleOlech, Monika, Katarzyna Ropka-Molik, Tomasz Szmatoła, Katarzyna Piórkowska, and Jacek Kuźmak. 2021. "Single Nucleotide Polymorphisms in Genes Encoding Toll-Like Receptors 7 and 8 and Their Association with Proviral Load of SRLVs in Goats of Polish Carpathian Breed" Animals 11, no. 7: 1908. https://doi.org/10.3390/ani11071908
APA StyleOlech, M., Ropka-Molik, K., Szmatoła, T., Piórkowska, K., & Kuźmak, J. (2021). Single Nucleotide Polymorphisms in Genes Encoding Toll-Like Receptors 7 and 8 and Their Association with Proviral Load of SRLVs in Goats of Polish Carpathian Breed. Animals, 11(7), 1908. https://doi.org/10.3390/ani11071908






