Chronic Wasting Disease: State of the Science
Abstract
1. Introduction
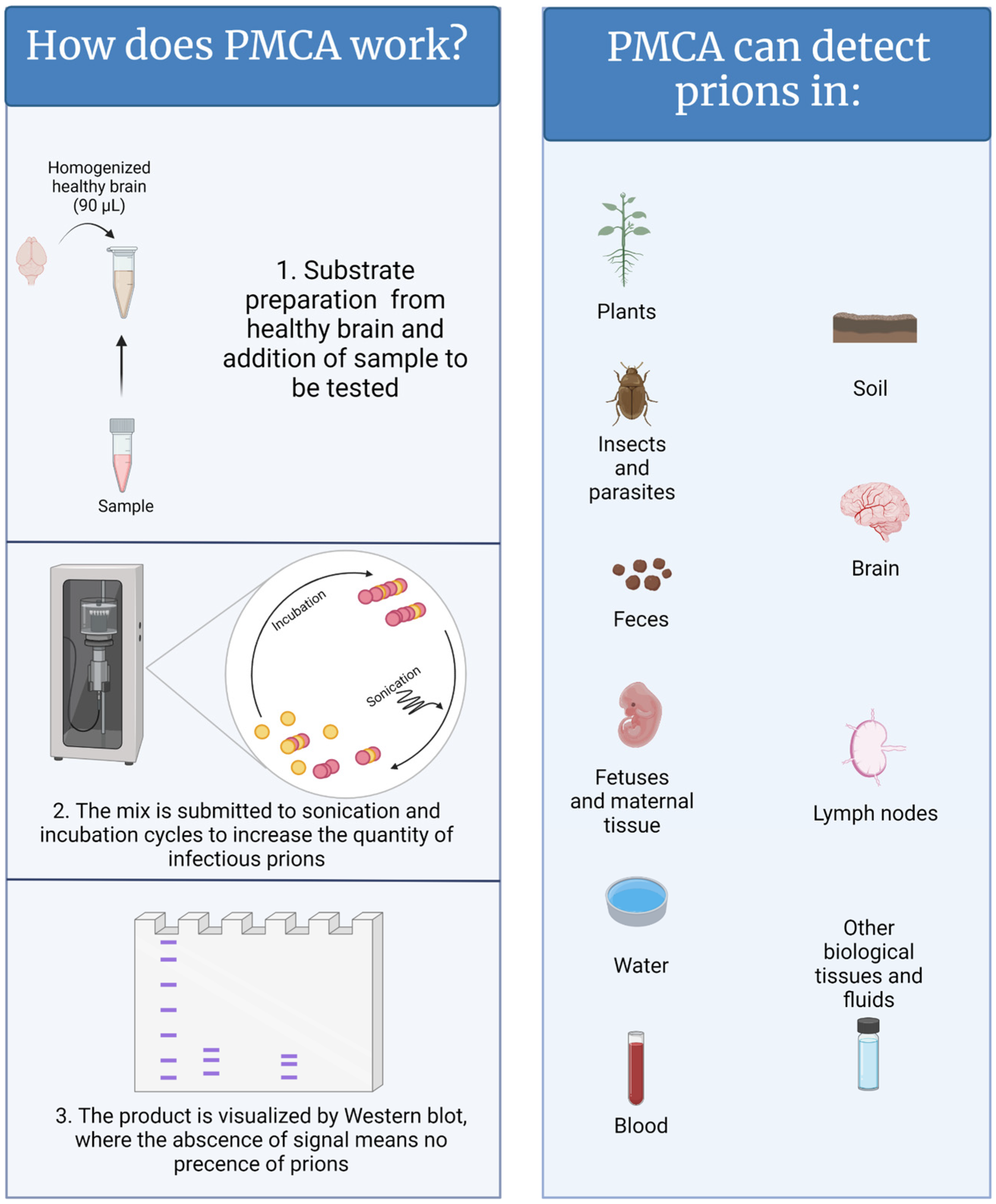
2. Protein Misfolding Cyclic Amplification
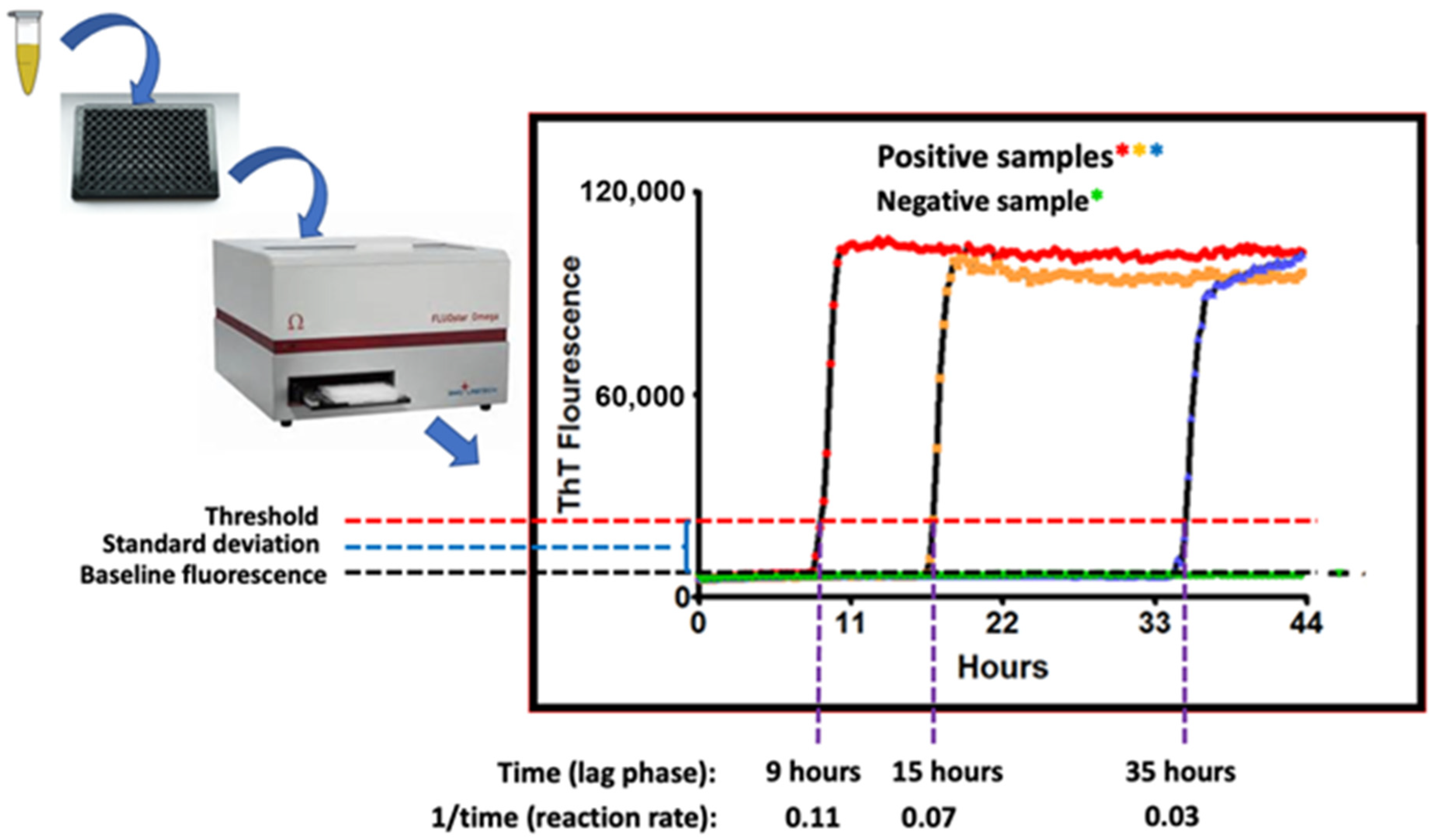
3. Real-Time Quaking-Induced Conversion (RT-QuIC)
4. The Process for CWD Diagnostic Assay Approval by the USDA
5. CWD in the Environment
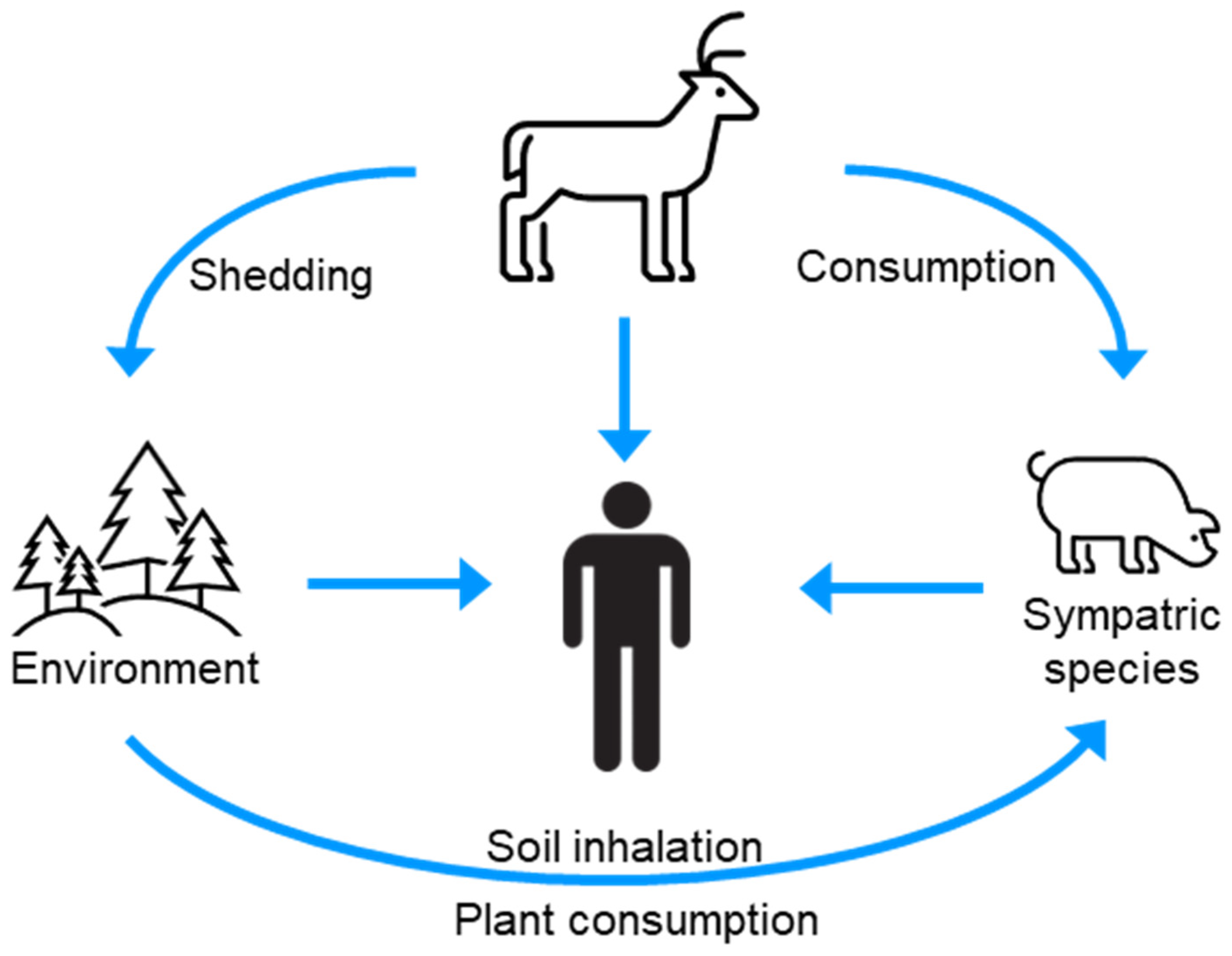
6. Interspecies Transmission of CWD
7. Summary
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Williams, E.S.; Young, S. Chronic wasting disease of captive mule deer: A spongiform encephalopathy. J. Wildl. Dis. 1980, 16, 89–98. [Google Scholar] [CrossRef]
- Deleault, N.R.; Harris, B.T.; Rees, J.R.; Supattapone, S. Formation of native prions from minimal components in vitro. Proc. Natl. Acad. Sci. USA 2007, 104, 9741–9746. [Google Scholar] [CrossRef]
- Prusiner, S.B. Novel proteinaceous infectious particles cause scrapie. Science 1982, 216, 136–144. [Google Scholar] [CrossRef]
- Kraus, A.; Hoyt, F.; Schwartz, C.L.; Hansen, B.; Artikis, E.; Hughson, A.G.; Raymond, G.J.; Race, B.; Baron, G.S.; Caughey, B. High-resolution structure and strain comparison of infectious mammalian prions. Mol. Cell 2021, 81, 4540–4551.e4546. [Google Scholar] [CrossRef]
- Kocisko, D.A.; Come, J.H.; Priola, S.A.; Chesebro, B.; Raymond, G.J.; Lansbury, P.T.; Caughey, B. Cell-free formation of protease-resistant prion protein. Nature 1994, 370, 471–474. [Google Scholar] [CrossRef] [PubMed]
- Oesch, B.; Westaway, D.; Walchli, M.; McKinley, M.P.; Kent, S.B.; Aebersold, R.; Barry, R.A.; Tempst, P.; Teplow, D.B.; Hood, L.E.; et al. A cellular gene encodes scrapie PrP 27-30 protein. Cell 1985, 40, 735–746. [Google Scholar] [CrossRef] [PubMed]
- Caughey, B.; Raymond, G.J. The scrapie-associated form of PrP is made from a cell surface precursor that is both protease- and phospholipase-sensitive. J. Biol. Chem. 1991, 266, 18217–18223. [Google Scholar] [CrossRef] [PubMed]
- Otero, A.; Velasquez, C.D.; Aiken, J.; McKenzie, D. Chronic wasting disease: A cervid prion infection looming to spillover. Vet. Res. 2021, 52, 115. [Google Scholar] [CrossRef]
- Brandel, J.P.; Culeux, A.; Grznarova, K.; Levavasseur, E.; Lamy, P.; Privat, N.; Welaratne, A.; Denouel, A.; Laplanche, J.L.; Haik, S. Amplification techniques and diagnosis of prion diseases. Rev. Neurol. 2019, 175, 458–463. [Google Scholar] [CrossRef]
- Haley, N.J.; Richt, J.A. Evolution of diagnostic tests for chronic wasting eisease, a naturally occurring prion disease of cervids. Pathogens 2017, 6, 35. [Google Scholar] [CrossRef] [PubMed]
- Morales, R.; Duran-Aniotz, C.; Diaz-Espinoza, R.; Camacho, M.V.; Soto, C. Protein misfolding cyclic amplification of infectious prions. Nat. Protoc. 2012, 7, 1397–1409. [Google Scholar] [CrossRef] [PubMed]
- Schmitz, M.; Cramm, M.; Llorens, F.; Muller-Cramm, D.; Collins, S.; Atarashi, R.; Satoh, K.; Orru, C.D.; Groveman, B.R.; Zafar, S.; et al. The real-time quaking-induced conversion assay for detection of human prion disease and study of other protein misfolding diseases. Nat. Protoc. 2016, 11, 2233–2242. [Google Scholar] [CrossRef]
- Castilla, J.; Saa, P.; Hetz, C.; Soto, C. In vitro generation of infectious scrapie prions. Cell 2005, 121, 195–206. [Google Scholar] [CrossRef]
- Saborio, G.P.; Permanne, B.; Soto, C. Sensitive detection of pathological prion protein by cyclic amplification of protein misfolding. Nature 2001, 411, 810–813. [Google Scholar] [CrossRef] [PubMed]
- Saa, P.; Castilla, J.; Soto, C. Ultra-efficient replication of infectious prions by automated protein misfolding cyclic amplification. J. Biol. Chem. 2006, 281, 35245–35252. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.; Morales, R.; Barria, M.A.; Soto, C. Estimating prion concentration in fluids and tissues by quantitative PMCA. Nat. Methods 2010, 7, 519–520. [Google Scholar] [CrossRef]
- McNulty, E.; Nalls, A.V.; Mellentine, S.; Hughes, E.; Pulscher, L.; Hoover, E.A.; Mathiason, C.K. Comparison of conventional, amplification and bio-assay detection methods for a chronic wasting disease inoculum pool. PLoS ONE 2019, 14, e0216621. [Google Scholar] [CrossRef]
- Davenport, K.A.; Hoover, C.E.; Denkers, N.D.; Mathiason, C.K.; Hoover, E.A. Modified protein misfolding cyclic amplification overcomes real-time quaking-induced conversion assay inhibitors in deer saliva to detect chronic wasting disease prions. J. Clin. Microbiol. 2018, 56, e00947-18. [Google Scholar] [CrossRef]
- Kramm, C.; Soto, P.; Nichols, T.A.; Morales, R. Chronic wasting disease (CWD) prion detection in blood from pre-symptomatic white-tailed deer harboring PRNP polymorphic variants. Sci. Rep. 2020, 10, 19763. [Google Scholar] [CrossRef]
- Nalls, A.V.; McNulty, E.E.; Mayfield, A.; Crum, J.M.; Keel, M.K.; Hoover, E.A.; Ruder, M.G.; Mathiason, C.K. Detection of chronic wasting disease prions in fetal tissues of free-ranging white-tailed deer. Viruses 2021, 13, 2430. [Google Scholar] [CrossRef]
- Bravo-Risi, F.; Soto, P.; Eckland, T.; Dittmar, R.; Ramirez, S.; Catumbela, C.S.G.; Soto, C.; Lockwood, M.; Nichols, T.; Morales, R. Detection of CWD prions in naturally infected white-tailed deer fetuses and gestational tissues by PMCA. Sci. Rep. 2021, 11, 18385. [Google Scholar] [CrossRef] [PubMed]
- Pritzkow, S.; Morales, R.; Lyon, A.; Concha-Marambio, L.; Urayama, A.; Soto, C. Efficient prion disease transmission through common environmental materials. J. Biol. Chem. 2018, 293, 3363–3373. [Google Scholar] [CrossRef] [PubMed]
- Plummer, I.H.; Johnson, C.J.; Chesney, A.R.; Pedersen, J.A.; Samuel, M.D. Mineral licks as environmental reservoirs of chronic wasting disease prions. PLoS ONE 2018, 13, e0196745. [Google Scholar] [CrossRef] [PubMed]
- Haley, N.J.; Mathiason, C.K.; Zabel, M.D.; Telling, G.C.; Hoover, E.A. Detection of sub-clinical CWD infection in conventional test-negative deer long after oral exposure to urine and feces from CWD+ deer. PLoS ONE 2009, 4, e7990. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.J.; Aiken, J.M.; McKenzie, D.; Samuel, M.D.; Pedersen, J.A. Highly efficient amplification of chronic wasting disease agent by protein misfolding cyclic amplification with beads (PMCAb). PLoS ONE 2012, 7, e35383. [Google Scholar] [CrossRef] [PubMed]
- Pritzkow, S.; Morales, R.; Moda, F.; Khan, U.; Telling, G.C.; Hoover, E.; Soto, C. Grass plants bind, retain, uptake, and transport infectious prions. Cell Rep. 2015, 11, 1168–1175. [Google Scholar] [CrossRef]
- Saunders, S.E.; Shikiya, R.A.; Langenfeld, K.; Bartelt-Hunt, S.L.; Bartz, J.C. Replication efficiency of soil-bound prions varies with soil type. J. Virol. 2011, 85, 5476–5482. [Google Scholar] [CrossRef]
- Saunders, S.E.; Bartz, J.C.; Vercauteren, K.C.; Bartelt-Hunt, S.L. An enzymatic treatment of soil-bound prions effectively inhibits replication. Appl. Environ. Microbiol. 2011, 77, 4313–4317. [Google Scholar] [CrossRef]
- Kramm, C.; Gomez-Gutierrez, R.; Soto, C.; Telling, G.; Nichols, T.; Morales, R. In Vitro detection of Chronic Wasting Disease (CWD) prions in semen and reproductive tissues of white-tailed deer bucks (Odocoileus virginianus). PLoS ONE 2019, 14, e0226560. [Google Scholar] [CrossRef]
- Nalls, A.V.; McNulty, E.; Powers, J.; Seelig, D.M.; Hoover, C.; Haley, N.J.; Hayes-Klug, J.; Anderson, K.; Stewart, P.; Goldmann, W.; et al. Mother to offspring transmission of chronic wasting disease in reeves’ muntjac deer. PLoS ONE 2013, 8, e71844. [Google Scholar] [CrossRef]
- Barria, M.A.; Telling, G.C.; Gambetti, P.; Mastrianni, J.A.; Soto, C. Generation of a new form of human PrP(Sc) in vitro by interspecies transmission from cervid prions. J. Biol. Chem. 2011, 286, 7490–7495. [Google Scholar] [CrossRef] [PubMed]
- Castilla, J.; Gonzalez-Romero, D.; Saa, P.; Morales, R.; De Castro, J.; Soto, C. Crossing the species barrier by PrP(Sc) replication in vitro generates unique infectious prions. Cell 2008, 134, 757–768. [Google Scholar] [CrossRef] [PubMed]
- Green, K.M.; Castilla, J.; Seward, T.S.; Napier, D.L.; Jewell, J.E.; Soto, C.; Telling, G.C. Accelerated high fidelity prion amplification within and across prion species barriers. PLoS Pathog. 2008, 4, e1000139. [Google Scholar] [CrossRef] [PubMed]
- Morales, R. Prion strains in mammals: Different conformations leading to disease. PLoS Pathog. 2017, 13, e1006323. [Google Scholar] [CrossRef] [PubMed]
- Morales, R.; Abid, K.; Soto, C. The prion strain phenomenon: Molecular basis and unprecedented features. Biochim. Biophys. Acta 2007, 1772, 681–691. [Google Scholar] [CrossRef] [PubMed]
- Wilham, J.M.; Orru, C.D.; Bessen, R.A.; Atarashi, R.; Sano, K.; Race, B.; Meade-White, K.D.; Taubner, L.M.; Timmes, A.; Caughey, B. Rapid end-point quantitation of prion seeding activity with sensitivity comparable to bioassays. PLoS Pathog. 2010, 6, e1001217. [Google Scholar] [CrossRef] [PubMed]
- Atarashi, R.; Satoh, K.; Sano, K.; Fuse, T.; Yamaguchi, N.; Ishibashi, D.; Matsubara, T.; Nakagaki, T.; Yamanaka, H.; Shirabe, S.; et al. Ultrasensitive human prion detection in cerebrospinal fluid by real-time quaking-induced conversion. Nat. Med. 2011, 17, 175–178. [Google Scholar] [CrossRef] [PubMed]
- Henderson, D.M.; Davenport, K.A.; Haley, N.J.; Denkers, N.D.; Mathiason, C.K.; Hoover, E.A. Quantitative assessment of prion infectivity in tissues and body fluids by real-time quaking-induced conversion. J. Gen. Virol. 2015, 96, 210–219. [Google Scholar] [CrossRef]
- Orru, C.D.; Wilham, J.M.; Vascellari, S.; Hughson, A.G.; Caughey, B. New generation QuIC assays for prion seeding activity. Prion 2012, 6, 147–152. [Google Scholar] [CrossRef]
- Orru, C.D.; Wilham, J.M.; Raymond, L.D.; Kuhn, F.; Schroeder, B.; Raeber, A.J.; Caughey, B. Prion disease blood test using immunoprecipitation and improved quaking-induced conversion. mBio 2011, 2, e00078-00011. [Google Scholar] [CrossRef]
- Denkers, N.D.; Henderson, D.M.; Mathiason, C.K.; Hoover, E.A. Enhanced prion detection in biological samples by magnetic particle extraction and real-time quaking-induced conversion. J. Gen. Virol. 2016, 97, 2023–2029. [Google Scholar] [CrossRef]
- Elder, A.M.; Henderson, D.M.; Nalls, A.V.; Hoover, E.A.; Kincaid, A.E.; Bartz, J.C.; Mathiason, C.K. Immediate and Ongoing Detection of Prions in the Blood of Hamsters and Deer following Oral, Nasal, or Blood Inoculations. J. Virol. 2015, 89, 7421–7424. [Google Scholar] [CrossRef]
- Elder, A.M.; Henderson, D.M.; Nalls, A.V.; Wilham, J.M.; Caughey, B.W.; Hoover, E.A.; Kincaid, A.E.; Bartz, J.C.; Mathiason, C.K. In vitro detection of prionemia in TSE-infected cervids and hamsters. PLoS ONE 2013, 8, e80203. [Google Scholar] [CrossRef]
- Haley, N.J.; Van de Motter, A.; Carver, S.; Henderson, D.; Davenport, K.; Seelig, D.M.; Mathiason, C.; Hoover, E. Prion-seeding activity in cerebrospinal fluid of deer with chronic wasting disease. PLoS ONE 2013, 8, e81488. [Google Scholar] [CrossRef] [PubMed]
- Henderson, D.M.; Denkers, N.D.; Hoover, C.E.; Garbino, N.; Mathiason, C.K.; Hoover, E.A. Longitudinal detection of prion shedding in saliva and urine by chronic wasting disease-infected deer by real-time quaking-induced conversion. J. Virol. 2015, 89, 9338–9347. [Google Scholar] [CrossRef] [PubMed]
- Tennant, J.M.; Li, M.; Henderson, D.M.; Tyer, M.L.; Denkers, N.D.; Haley, N.J.; Mathiason, C.K.; Hoover, E.A. Shedding and stability of CWD prion seeding activity in cervid feces. PLoS ONE 2020, 15, e0227094. [Google Scholar] [CrossRef] [PubMed]
- Cooper, S.K.; Hoover, C.E.; Henderson, D.M.; Haley, N.J.; Mathiason, C.K.; Hoover, E.A. Detection of CWD in cervids by RT-QuIC assay of third eyelids. PLoS ONE 2019, 14, e0221654. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, N.C.; Charco, J.M.; Plagenz, J.; Orru, C.D.; Denkers, N.D.; Metrick, M.A., 2nd; Hughson, A.G.; Griffin, K.A.; Race, B.; Hoover, E.A.; et al. Detection of chronic wasting disease in mule and white-tailed deer by RT-QuIC analysis of outer ear. Sci. Rep. 2021, 11, 7702. [Google Scholar] [CrossRef]
- Henderson, D.M.; Tennant, J.M.; Haley, N.J.; Denkers, N.D.; Mathiason, C.K.; Hoover, E.A. Detection of chronic wasting disease prion seeding activity in deer and elk feces by real-time quaking-induced conversion. J. Gen. Virol. 2017, 98, 1953–1962. [Google Scholar] [CrossRef]
- Henderson, D.M.; Denkers, N.D.; Hoover, C.E.; McNulty, E.E.; Cooper, S.K.; Bracchi, L.A.; Mathiason, C.K.; Hoover, E.A. Progression of chronic wasting disease in white-tailed deer analyzed by serial biopsy RT-QuIC and immunohistochemistry. PLoS ONE 2020, 15, e0228327. [Google Scholar] [CrossRef]
- Denkers, N.D.; Hoover, C.E.; Davenport, K.A.; Henderson, D.M.; McNulty, E.E.; Nalls, A.V.; Mathiason, C.K.; Hoover, E.A. Very low oral exposure to prions of brain or saliva origin can transmit chronic wasting disease. PLoS ONE 2020, 15, e0237410. [Google Scholar] [CrossRef]
- Orru, C.D.; Groveman, B.R.; Hughson, A.G.; Zanusso, G.; Coulthart, M.B.; Caughey, B. Rapid and sensitive RT-QuIC detection of human Creutzfeldt-Jakob disease using cerebrospinal fluid. mBio 2015, 6, e02451-14. [Google Scholar] [CrossRef] [PubMed]
- Vascellari, S.; Orru, C.D.; Caughey, B. Real-Time Quaking- Induced Conversion Assays for Prion Diseases, Synucleinopathies, and Tauopathies. Front. Aging Neurosci. 2022, 14, 853050. [Google Scholar] [CrossRef] [PubMed]
- Fairfoul, G.; McGuire, L.I.; Pal, S.; Ironside, J.W.; Neumann, J.; Christie, S.; Joachim, C.; Esiri, M.; Evetts, S.G.; Rolinski, M.; et al. Alpha-synuclein RT-QuIC in the CSF of patients with alpha-synucleinopathies. Ann. Clin. Transl. Neurol. 2016, 3, 812–818. [Google Scholar] [CrossRef] [PubMed]
- Groveman, B.R.; Orru, C.D.; Hughson, A.G.; Raymond, L.D.; Zanusso, G.; Ghetti, B.; Campbell, K.J.; Safar, J.; Galasko, D.; Caughey, B. Rapid and ultra-sensitive quantitation of disease-associated alpha-synuclein seeds in brain and cerebrospinal fluid by alphaSyn RT-QuIC. Acta Neuropathol. Commun. 2018, 6, 7. [Google Scholar] [CrossRef] [PubMed]
- Russo, M.J.; Orru, C.D.; Concha-Marambio, L.; Giaisi, S.; Groveman, B.R.; Farris, C.M.; Holguin, B.; Hughson, A.G.; LaFontant, D.E.; Caspell-Garcia, C.; et al. High diagnostic performance of independent alpha-synuclein seed amplification assays for detection of early Parkinson’s disease. Acta Neuropathol. Commun. 2021, 9, 179. [Google Scholar] [CrossRef] [PubMed]
- Saijo, E.; Ghetti, B.; Zanusso, G.; Oblak, A.; Furman, J.L.; Diamond, M.I.; Kraus, A.; Caughey, B. Ultrasensitive and selective detection of 3-repeat tau seeding activity in Pick disease brain and cerebrospinal fluid. Acta Neuropathol. 2017, 133, 751–765. [Google Scholar] [CrossRef]
- Kraus, A.; Saijo, E.; Metrick, M.A., 2nd; Newell, K.; Sigurdson, C.J.; Zanusso, G.; Ghetti, B.; Caughey, B. Seeding selectivity and ultrasensitive detection of tau aggregate conformers of Alzheimer disease. Acta Neuropathol. 2019, 137, 585–598. [Google Scholar] [CrossRef]
- Metrick, M.A., 2nd; Ferreira, N.D.C.; Saijo, E.; Kraus, A.; Newell, K.; Zanusso, G.; Vendruscolo, M.; Ghetti, B.; Caughey, B. A single ultrasensitive assay for detection and discrimination of tau aggregates of Alzheimer and Pick diseases. Acta Neuropathol. Commun. 2020, 8, 22. [Google Scholar] [CrossRef]
- Mathiason, C.K.; Powers, J.G.; Dahmes, S.J.; Osborn, D.A.; Miller, K.V.; Warren, R.J.; Mason, G.L.; Hays, S.A.; Hayes-Klug, J.; Seelig, D.M.; et al. Infectious prions in the saliva and blood of deer with chronic wasting disease. Science 2006, 314, 133–136. [Google Scholar] [CrossRef]
- Tamguney, G.; Miller, M.W.; Wolfe, L.L.; Sirochman, T.M.; Glidden, D.V.; Palmer, C.; Lemus, A.; DeArmond, S.J.; Prusiner, S.B. Asymptomatic deer excrete infectious prions in faeces. Nature 2009, 461, 529–532. [Google Scholar] [CrossRef]
- Angers, R.C.; Seward, T.S.; Napier, D.; Green, M.; Hoover, E.; Spraker, T.; O’Rourke, K.; Balachandran, A.; Telling, G.C. Chronic wasting disease prions in elk antler velvet. Emerg. Infect. Dis. 2009, 15, 696–703. [Google Scholar] [CrossRef]
- Nalls, A.V.; McNulty, E.; Hoover, C.E.; Pulscher, L.A.; Hoover, E.A.; Mathiason, C.K. Infectious prions in the pregnancy microenvironment of chronic wasting disease-infected Reeves’ muntjac deer. J. Virol. 2017, 91, e00501-17. [Google Scholar] [CrossRef]
- Johnson, C.J.; Pedersen, J.A.; Chappell, R.J.; McKenzie, D.; Aiken, J.M. Oral transmissibility of prion disease is enhanced by binding to soil particles. PLoS Pathog. 2007, 3, e93. [Google Scholar] [CrossRef]
- Russell Greig, J. Observations on the Transmission of the Disease by Mediate Contact. Vet. J. 1940, 96, 203–206. [Google Scholar] [CrossRef]
- Brown, P.; Gajdusek, D.C. Survival of scrapie virus after 3 years’ interment. Lancet 1991, 337, 269–270. [Google Scholar] [CrossRef] [PubMed]
- Somerville, R.A.; Fernie, K.; Smith, A.; Bishop, K.; Maddison, B.C.; Gough, K.C.; Hunter, N. BSE infectivity survives burial for five years with only limited spread. Arch. Virol. 2019, 164, 1135–1145. [Google Scholar] [CrossRef] [PubMed]
- Georgsson, G.; Sigurdarson, S.; Brown, P. Infectious agent of sheep scrapie may persist in the environment for at least 16 years. J. Gen. Virol. 2006, 87, 3737–3740. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.W.; Williams, E.S.; Hobbs, N.T.; Wolfe, L.L. Environmental sources of prion transmission in mule deer. Emerg. Infect. Dis. 2004, 10, 1003–1006. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.J.; Phillips, K.E.; Schramm, P.T.; McKenzie, D.; Aiken, J.M.; Pedersen, J.A. Prions adhere to soil minerals and remain infectious. PLoS Pathog. 2006, 2, e32. [Google Scholar] [CrossRef] [PubMed]
- Giachin, G.; Narkiewicz, J.; Scaini, D.; Ngoc, A.T.; Margon, A.; Sequi, P.; Leita, L.; Legname, G. Prion protein interaction with soil humic substances: Environmental implications. PLoS ONE 2014, 9, e100016. [Google Scholar] [CrossRef] [PubMed]
- Saunders, S.E.; Bartelt-Hunt, S.L.; Bartz, J.C. Resistance of soil-bound prions to rumen digestion. PLoS ONE 2012, 7, e44051. [Google Scholar] [CrossRef] [PubMed]
- David Walter, W.; Walsh, D.P.; Farnsworth, M.L.; Winkelman, D.L.; Miller, M.W. Soil clay content underlies prion infection odds. Nat. Commun. 2011, 2, 200. [Google Scholar] [CrossRef] [PubMed]
- Kuznetsova, A.; McKenzie, D.; Ytrehus, B.; Utaaker, K.S.; Aiken, J.M. Movement of chronic wasting disease prions in prairie, boreal and alpine soils. Pathogens 2023, 12, 269. [Google Scholar] [CrossRef] [PubMed]
- Jacobson, K.H.; Lee, S.; McKenzie, D.; Benson, C.H.; Pedersen, J.A. Transport of the pathogenic prion protein through landfill materials. Environ. Sci. Technol. 2009, 43, 2022–2028. [Google Scholar] [CrossRef][Green Version]
- Yuan, Q.; Telling, G.; Bartelt-Hunt, S.L.; Bartz, J.C. Dehydration of Prions on environmentally relevant surfaces protects them from inactivation by freezing and thawing. J. Virol. 2018, 92, e02191-17. [Google Scholar] [CrossRef]
- Carlson, C.M.; Thomas, S.; Keating, M.W.; Soto, P.; Gibbs, N.M.; Chang, H.; Wiepz, J.K.; Austin, A.G.; Schneider, J.R.; Morales, R.; et al. Plants as vectors for environmental prion transmission. iScience 2023, 26, 108428. [Google Scholar] [CrossRef]
- Nichols, T.A.; Pulford, B.; Wyckoff, A.C.; Meyerett, C.; Michel, B.; Gertig, K.; Hoover, E.A.; Jewell, J.E.; Telling, G.C.; Zabel, M.D. Detection of protease-resistant cervid prion protein in water from a CWD-endemic area. Prion 2009, 3, 171–183. [Google Scholar] [CrossRef]
- Kuznetsova, A.; McKenzie, D.; Cullingham, C.; Aiken, J.M. Long-term incubation PrP(CWD) with soils affects prion recovery but not Infectivity. Pathogens 2020, 9, 311. [Google Scholar] [CrossRef] [PubMed]
- Sohn, H.J.; Park, K.J.; Roh, I.S.; Kim, H.J.; Park, H.C.; Kang, H.E. Sodium hydroxide treatment effectively inhibits PrP(CWD) replication in farm soil. Prion 2019, 13, 137–140. [Google Scholar] [CrossRef] [PubMed]
- Kuznetsova, A.; Cullingham, C.; McKenzie, D.; Aiken, J.M. Soil humic acids degrade CWD prions and reduce infectivity. PLoS Pathog. 2018, 14, e1007414. [Google Scholar] [CrossRef]
- Brown, P.; Rau, E.H.; Lemieux, P.; Johnson, B.K.; Bacote, A.E.; Gajdusek, D.C. Infectivity studies of both ash and air emissions from simulated incineration of scrapie-contaminated tissues. Environ. Sci. Technol. 2004, 38, 6155–6160. [Google Scholar] [CrossRef]
- Bruce, M.E.; Will, R.G.; Ironside, J.W.; McConnell, I.; Drummond, D.; Suttie, A.; McCardle, L.; Chree, A.; Hope, J.; Birkett, C.; et al. Transmissions to mice indicate that ‘new variant’ CJD is caused by the BSE agent. Nature 1997, 389, 498–501. [Google Scholar] [CrossRef] [PubMed]
- Dagleish, M.P.; Martin, S.; Steele, P.; Finlayson, J.; Siso, S.; Hamilton, S.; Chianini, F.; Reid, H.W.; Gonzalez, L.; Jeffrey, M. Experimental transmission of bovine spongiform encephalopathy to European red deer (Cervus elaphus elaphus). BMC Vet. Res. 2008, 4, 17. [Google Scholar] [CrossRef] [PubMed]
- Lasmezas, C.I.; Deslys, J.P.; Demaimay, R.; Adjou, K.T.; Lamoury, F.; Dormont, D.; Robain, O.; Ironside, J.; Hauw, J.J. BSE transmission to macaques. Nature 1996, 381, 743–744. [Google Scholar] [CrossRef] [PubMed]
- Kirkwood, J.K.; Wells, G.A.; Wilesmith, J.W.; Cunningham, A.A.; Jackson, S.I. Spongiform encephalopathy in an arabian oryx (Oryx leucoryx) and a greater kudu (Tragelaphus strepsiceros). Vet. Rec. 1990, 127, 418–420. [Google Scholar] [PubMed]
- Wells, G.A. Pathogenesis of BSE. Vet. Res. Commun. 2003, 27 (Suppl. 1), 25–28. [Google Scholar] [CrossRef] [PubMed]
- Pearson, G.R.; Wyatt, J.M.; Gruffydd-Jones, T.J.; Hope, J.; Chong, A.; Higgins, R.J.; Scott, A.C.; Wells, G.A. Feline spongiform encephalopathy: Fibril and PrP studies. Vet. Rec. 1992, 131, 307–310. [Google Scholar] [CrossRef]
- Wyatt, J.M.; Pearson, G.R.; Smerdon, T.N.; Gruffydd-Jones, T.J.; Wells, G.A.; Wilesmith, J.W. Naturally occurring scrapie-like spongiform encephalopathy in five domestic cats. Vet. Rec. 1991, 129, 233–236. [Google Scholar] [CrossRef] [PubMed]
- Jaumain, E.; Quadrio, I.; Herzog, L.; Reine, F.; Rezaei, H.; Andreoletti, O.; Laude, H.; Perret-Liaudet, A.; Haik, S.; Beringue, V. Absence of evidence for a causal link between bovine spongiform encephalopathy strain variant L-BSE and known forms of sporadic Creutzfeldt-Jakob disease in human PrP transgenic mice. J. Virol. 2016, 90, 10867–10874. [Google Scholar] [CrossRef]
- Osterholm, M.T.; Anderson, C.J.; Zabel, M.D.; Scheftel, J.M.; Moore, K.A.; Appleby, B.S. Chronic wasting disease in cervids: Implications for prion transmission to humans and other animal species. mBio 2019, 10, e01091-19. [Google Scholar] [CrossRef]
- Lasmezas, C.I.; Fournier, J.G.; Nouvel, V.; Boe, H.; Marce, D.; Lamoury, F.; Kopp, N.; Hauw, J.J.; Ironside, J.; Bruce, M.; et al. Adaptation of the bovine spongiform encephalopathy agent to primates and comparison with Creutzfeldt—Jakob disease: Implications for human health. Proc. Natl. Acad. Sci. USA 2001, 98, 4142–4147. [Google Scholar] [CrossRef] [PubMed]
- Mikol, J.; Delmotte, J.; Jouy, D.; Vaysset, E.; Bastian, C.; Deslys, J.P.; Comoy, E. Direct neural transmission of vCJD/BSE in macaque after finger incision. Acta Neuropathol. 2021, 141, 119–122. [Google Scholar] [CrossRef] [PubMed]
- Comoy, E.E.; Mikol, J.; Luccantoni-Freire, S.; Correia, E.; Lescoutra-Etchegaray, N.; Durand, V.; Dehen, C.; Andreoletti, O.; Casalone, C.; Richt, J.A.; et al. Transmission of scrapie prions to primate after an extended silent incubation period. Sci. Rep. 2015, 5, 11573. [Google Scholar] [CrossRef] [PubMed]
- Race, B.; Meade-White, K.D.; Miller, M.W.; Barbian, K.D.; Rubenstein, R.; LaFauci, G.; Cervenakova, L.; Favara, C.; Gardner, D.; Long, D.; et al. Susceptibilities of nonhuman primates to chronic wasting disease. Emerg. Infect. Dis. 2009, 15, 1366–1376. [Google Scholar] [CrossRef] [PubMed]
- Race, B.; Meade-White, K.D.; Phillips, K.; Striebel, J.; Race, R.; Chesebro, B. Chronic wasting disease agents in nonhuman primates. Emerg. Infect. Dis. 2014, 20, 833–837. [Google Scholar] [CrossRef] [PubMed]
- Race, B.; Williams, K.; Orru, C.D.; Hughson, A.G.; Lubke, L.; Chesebro, B. Lack of transmission of chronic wasting disease to Cynomolgus macaques. J. Virol. 2018, 92, e00550-18. [Google Scholar] [CrossRef] [PubMed]
- Race, B.; Williams, K.; Chesebro, B. Transmission studies of chronic wasting disease to transgenic mice overexpressing human prion protein using the RT-QuIC assay. Vet. Res. 2019, 50, 6. [Google Scholar] [CrossRef] [PubMed]
- Wilson, R.; Plinston, C.; Hunter, N.; Casalone, C.; Corona, C.; Tagliavini, F.; Suardi, S.; Ruggerone, M.; Moda, F.; Graziano, S.; et al. Chronic wasting disease and atypical forms of bovine spongiform encephalopathy and scrapie are not transmissible to mice expressing wild-type levels of human prion protein. J. Gen. Virol. 2012, 93, 1624–1629. [Google Scholar] [CrossRef]
- Sandberg, M.K.; Al-Doujaily, H.; Sigurdson, C.J.; Glatzel, M.; O’Malley, C.; Powell, C.; Asante, E.A.; Linehan, J.M.; Brandner, S.; Wadsworth, J.D.; et al. Chronic wasting disease prions are not transmissible to transgenic mice overexpressing human prion protein. J. Gen. Virol. 2010, 91, 2651–2657. [Google Scholar] [CrossRef]
- Tamguney, G.; Giles, K.; Bouzamondo-Bernstein, E.; Bosque, P.J.; Miller, M.W.; Safar, J.; DeArmond, S.J.; Prusiner, S.B. Transmission of elk and deer prions to transgenic mice. J. Virol. 2006, 80, 9104–9114. [Google Scholar] [CrossRef] [PubMed]
- Kong, Q.; Huang, S.; Zou, W.; Vanegas, D.; Wang, M.; Wu, D.; Yuan, J.; Zheng, M.; Bai, H.; Deng, H.; et al. Chronic wasting disease of elk: Transmissibility to humans examined by transgenic mouse models. J. Neurosci. 2005, 25, 7944–7949. [Google Scholar] [CrossRef] [PubMed]
- Kurt, T.D.; Jiang, L.; Fernandez-Borges, N.; Bett, C.; Liu, J.; Yang, T.; Spraker, T.R.; Castilla, J.; Eisenberg, D.; Kong, Q.; et al. Human prion protein sequence elements impede cross-species chronic wasting disease transmission. J. Clin. Investig. 2015, 125, 1485–1496. [Google Scholar] [CrossRef] [PubMed]
- Hannaoui, S.; Zemlyankina, I.; Chang, S.C.; Arifin, M.I.; Beringue, V.; McKenzie, D.; Schatzl, H.M.; Gilch, S. Transmission of cervid prions to humanized mice demonstrates the zoonotic potential of CWD. Acta Neuropathol. 2022, 144, 767–784. [Google Scholar] [CrossRef] [PubMed]
- Wolfe, L.L.; Fox, K.A.; Griffin, K.A.; Miller, M.W. Mountain Lions (Puma concolor) resist long-term dietary exposure to chronic wasting disease. J. Wildl. Dis. 2022, 58, 40–49. [Google Scholar] [CrossRef]
- Rhyan, J.C.; Miller, M.W.; Spraker, T.R.; McCollum, M.; Nol, P.; Wolfe, L.L.; Davis, T.R.; Creekmore, L.; O’Rourke, K.I. Failure of fallow deer (Dama dama) to develop chronic wasting disease when exposed to a contaminated environment and infected mule deer (Odocoileus hemionus). J. Wildl. Dis. 2011, 47, 739–744. [Google Scholar] [CrossRef]
- Williams, E.S.; O’Toole, D.; Miller, M.W.; Kreeger, T.J.; Jewell, J.E. Cattle (Bos taurus) resist chronic wasting disease following oral inoculation challenge or ten years’ natural exposure in contaminated environments. J. Wildl. Dis. 2018, 54, 460–470. [Google Scholar] [CrossRef]
- Mathiason, C.K.; Nalls, A.V.; Seelig, D.M.; Kraft, S.L.; Carnes, K.; Anderson, K.R.; Hayes-Klug, J.; Hoover, E.A. Susceptibility of domestic cats to chronic wasting disease. J. Virol. 2013, 87, 1947–1956. [Google Scholar] [CrossRef] [PubMed]
- Hamir, A.N.; Kunkle, R.A.; Cutlip, R.C.; Miller, J.M.; O’Rourke, K.I.; Williams, E.S.; Miller, M.W.; Stack, M.J.; Chaplin, M.J.; Richt, J.A. Experimental transmission of chronic wasting disease agent from mule deer to cattle by the intracerebral route. J. Vet. Diagn. Investig. 2005, 17, 276–281. [Google Scholar] [CrossRef]
- Hamir, A.N.; Kunkle, R.A.; Miller, J.M.; Greenlee, J.J.; Richt, J.A. Experimental second passage of chronic wasting disease (CWD (mule deer)) agent to cattle. J. Comp. Pathol. 2006, 134, 63–69. [Google Scholar] [CrossRef]
- Hamir, A.N.; Kunkle, R.A.; Cutlip, R.C.; Miller, J.M.; Williams, E.S.; Richt, J.A. Transmission of chronic wasting disease of mule deer to Suffolk sheep following intracerebral inoculation. J. Vet. Diagn. Investig. 2006, 18, 558–565. [Google Scholar] [CrossRef] [PubMed]
- Hamir, A.N.; Miller, J.M.; Kunkle, R.A.; Hall, S.M.; Richt, J.A. Susceptibility of cattle to first-passage intracerebral inoculation with chronic wasting disease agent from white-tailed deer. Vet. Pathol. 2007, 44, 487–493. [Google Scholar] [CrossRef] [PubMed]
- Bartz, J.C.; Marsh, R.F.; McKenzie, D.I.; Aiken, J.M. The host range of chronic wasting disease is altered on passage in ferrets. Virology 1998, 251, 297–301. [Google Scholar] [CrossRef]
- Moore, S.J.; West Greenlee, M.H.; Kondru, N.; Manne, S.; Smith, J.D.; Kunkle, R.A.; Kanthasamy, A.; Greenlee, J.J. Experimental transmission of the chronic wasting disease zgent to swine after oral or intracranial inoculation. J. Virol. 2017, 91, e00926-17. [Google Scholar] [CrossRef] [PubMed]
- Cassmann, E.D.; Greenlee, J.J. Pathogenesis, detection, and control of scrapie in sheep. Am. J. Vet. Res. 2020, 81, 600–614. [Google Scholar] [CrossRef] [PubMed]
- Saa, P.; Sferrazza, G.F.; Ottenberg, G.; Oelschlegel, A.M.; Dorsey, K.; Lasmezas, C.I. Strain-specific role of RNAs in prion replication. J. Virol. 2012, 86, 10494–10504. [Google Scholar] [CrossRef]
- Crowell, J.; Hughson, A.; Caughey, B.; Bessen, R.A. Host Determinants of Prion Strain Diversity Independent of Prion Protein Genotype. J. Virol. 2015, 89, 10427–10441. [Google Scholar] [CrossRef]
- Scott, M.; Foster, D.; Mirenda, C.; Serban, D.; Coufal, F.; Walchli, M.; Torchia, M.; Groth, D.; Carlson, G.; DeArmond, S.J.; et al. Transgenic mice expressing hamster prion protein produce species-specific scrapie infectivity and amyloid plaques. Cell 1989, 59, 847–857. [Google Scholar] [CrossRef]
- Dickinson, A.G.; Fraser, H.; Outram, G.W. Scrapie incubation time can exceed natural lifespan. Nature 1975, 256, 732–733. [Google Scholar] [CrossRef] [PubMed]
- Collinge, J.; Clarke, A.R. A general model of prion strains and their pathogenicity. Science 2007, 318, 930–936. [Google Scholar] [CrossRef] [PubMed]
- Marin-Moreno, A.; Huor, A.; Espinosa, J.C.; Douet, J.Y.; Aguilar-Calvo, P.; Aron, N.; Piquer, J.; Lugan, S.; Lorenzo, P.; Tillier, C.; et al. Radical Change in Zoonotic Abilities of Atypical BSE Prion Strains as Evidenced by Crossing of Sheep Species Barrier in Transgenic Mice. Emerg. Infect. Dis. 2020, 26, 1130–1139. [Google Scholar] [CrossRef]
- Herbst, A.; Velasquez, C.D.; Triscott, E.; Aiken, J.M.; McKenzie, D. Chronic Wasting Disease Prion Strain Emergence and Host Range Expansion. Emerg. Infect. Dis. 2017, 23, 1598–1600. [Google Scholar] [CrossRef]
- Duque Velasquez, C.; Kim, C.; Haldiman, T.; Kim, C.; Herbst, A.; Aiken, J.; Safar, J.G.; McKenzie, D. Chronic wasting disease (CWD) prion strains evolve via adaptive diversification of conformers in hosts expressing prion protein polymorphisms. J. Biol. Chem. 2020, 295, 4985–5001. [Google Scholar] [CrossRef]
- Hannaoui, S.; Triscott, E.; Duque Velasquez, C.; Chang, S.C.; Arifin, M.I.; Zemlyankina, I.; Tang, X.; Bollinger, T.; Wille, H.; McKenzie, D.; et al. New and distinct chronic wasting disease strains associated with cervid polymorphism at codon 116 of the Prnp gene. PLoS Pathog. 2021, 17, e1009795. [Google Scholar] [CrossRef]
- Johnson, C.J.; Herbst, A.; Duque-Velasquez, C.; Vanderloo, J.P.; Bochsler, P.; Chappell, R.; McKenzie, D. Prion protein polymorphisms affect chronic wasting disease progression. PLoS ONE 2011, 6, e17450. [Google Scholar] [CrossRef]
- Bian, J.; Christiansen, J.R.; Moreno, J.A.; Kane, S.J.; Khaychuk, V.; Gallegos, J.; Kim, S.; Telling, G.C. Primary structural differences at residue 226 of deer and elk PrP dictate selection of distinct CWD prion strains in gene-targeted mice. Proc. Natl. Acad. Sci. USA 2019, 116, 12478–12487. [Google Scholar] [CrossRef] [PubMed]
- Perrott, M.R.; Sigurdson, C.J.; Mason, G.L.; Hoover, E.A. Evidence for distinct chronic wasting disease (CWD) strains in experimental CWD in ferrets. J. Gen. Virol. 2012, 93, 212–221. [Google Scholar] [CrossRef] [PubMed]
- Nonno, R.; Di Bari, M.A.; Pirisinu, L.; D’Agostino, C.; Vanni, I.; Chiappini, B.; Marcon, S.; Riccardi, G.; Tran, L.; Vikoren, T.; et al. Studies in bank voles reveal strain differences between chronic wasting disease prions from Norway and North America. Proc. Natl. Acad. Sci. USA 2020, 117, 31417–31426. [Google Scholar] [CrossRef]
- Bian, J.; Kim, S.; Kane, S.J.; Crowell, J.; Sun, J.L.; Christiansen, J.; Saijo, E.; Moreno, J.A.; DiLisio, J.; Burnett, E.; et al. Adaptive selection of a prion strain conformer corresponding to established North American CWD during propagation of novel emergent Norwegian strains in mice expressing elk or deer prion protein. PLoS Pathog. 2021, 17, e1009748. [Google Scholar] [CrossRef] [PubMed]
- Otero, A.; Duque Velasquez, C.; Johnson, C.; Herbst, A.; Bolea, R.; Badiola, J.J.; Aiken, J.; McKenzie, D. Prion protein polymorphisms associated with reduced CWD susceptibility limit peripheral PrP(CWD) deposition in orally infected white-tailed deer. BMC Vet. Res. 2019, 15, 50. [Google Scholar] [CrossRef]
- Nichols, T.A.; Nicholson, E.M.; Liu, Y.; Tao, W.; Spraker, T.R.; Lavelle, M.; Fischer, J.; Kong, Q.; VerCauteren, K.C. Detection of two dissimilar chronic wasting disease isolates in two captive Rocky Mountain elk (Cervus canadensis) herds. Prion 2021, 15, 207–215. [Google Scholar] [CrossRef]
- Moore, J.; Tatum, T.; Hwang, S.; Vrentas, C.; West Greenlee, M.H.; Kong, Q.; Nicholson, E.; Greenlee, J. Novel strain of the chronic wasting disease agent isolated from experimentally inoculated elk with LL132 prion protein. Sci. Rep. 2020, 10, 3148. [Google Scholar] [CrossRef] [PubMed]
- Holec, S.A.M.; Yuan, Q.; Bartz, J.C. Alteration of prion strain emergence by nonhost factors. mSphere 2019, 4, e00630-19. [Google Scholar] [CrossRef]
- Yuan, Q.; Eckland, T.; Telling, G.; Bartz, J.; Bartelt-Hunt, S. Mitigation of prion infectivity and conversion capacity by a simulated natural process—repeated cycles of drying and wetting. PLoS Pathog. 2015, 11, e1004638. [Google Scholar] [CrossRef]
- Kincaid, A.E.; Bartz, J.C. The nasal cavity is a route for prion infection in hamsters. J. Virol. 2007, 81, 4482–4491. [Google Scholar] [CrossRef]
- Jacquemot, C.; Cuche, C.; Dormont, D.; Lazarini, F. High incidence of scrapie induced by repeated injections of subinfectious prion doses. J. Virol. 2005, 79, 8904–8908. [Google Scholar] [CrossRef][Green Version]
- Espinosa, J.C.; Andreoletti, O.; Castilla, J.; Herva, M.E.; Morales, M.; Alamillo, E.; San-Segundo, F.D.; Lacroux, C.; Lugan, S.; Salguero, F.J.; et al. Sheep-passaged bovine spongiform encephalopathy agent exhibits altered pathobiological properties in bovine-PrP transgenic mice. J. Virol. 2007, 81, 835–843. [Google Scholar] [CrossRef]
- Raymond, G.J.; Bossers, A.; Raymond, L.D.; O’Rourke, K.I.; McHolland, L.E.; Bryant, P.K., 3rd; Miller, M.W.; Williams, E.S.; Smits, M.; Caughey, B. Evidence of a molecular barrier limiting susceptibility of humans, cattle and sheep to chronic wasting disease. EMBO J. 2000, 19, 4425–4430. [Google Scholar] [CrossRef] [PubMed]
- Kocisko, D.A.; Priola, S.A.; Raymond, G.J.; Chesebro, B.; Lansbury, P.T., Jr.; Caughey, B. Species specificity in the cell-free conversion of prion protein to protease-resistant forms: A model for the scrapie species barrier. Proc. Natl. Acad. Sci. USA 1995, 92, 3923–3927. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Montalban, N.; Makarava, N.; Ostapchenko, V.G.; Savtchenk, R.; Alexeeva, I.; Rohwer, R.G.; Baskakov, I.V. Highly efficient protein misfolding cyclic amplification. PLoS Pathog. 2011, 7, e1001277. [Google Scholar] [CrossRef]
- Torres, J.M.; Espinosa, J.C.; Aguilar-Calvo, P.; Herva, M.E.; Relano-Gines, A.; Villa-Diaz, A.; Morales, M.; Parra, B.; Alamillo, E.; Brun, A.; et al. Elements modulating the prion species barrier and its passage consequences. PLoS ONE 2014, 9, e89722. [Google Scholar] [CrossRef] [PubMed]
- Harrathi, C.; Fernandez-Borges, N.; Erana, H.; Elezgarai, S.R.; Venegas, V.; Charco, J.M.; Castilla, J. Insights into the bidirectional properties of the sheep-deer prion transmission barrier. Mol. Neurobiol. 2019, 56, 5287–5303. [Google Scholar] [CrossRef] [PubMed]
- Priem, J.; Langeveld, J.P.; van Keulen, L.J.; van Zijderveld, F.G.; Andreoletti, O.; Bossers, A. Enhanced virulence of sheep-passaged bovine spongiform encephalopathy agent is revealed by decreased polymorphism barriers in prion protein conversion studies. J. Virol. 2014, 88, 2903–2912. [Google Scholar] [CrossRef]
- Wang, Z.; Qin, K.; Camacho, M.V.; Cali, I.; Yuan, J.; Shen, P.; Greenlee, J.; Kong, Q.; Mastrianni, J.A.; Zou, W.Q. Generation of human chronic wasting disease in transgenic mice. Acta Neuropathol. Commun. 2021, 9, 158. [Google Scholar] [CrossRef]
- Chianini, F.; Fernandez-Borges, N.; Vidal, E.; Gibbard, L.; Pintado, B.; de Castro, J.; Priola, S.A.; Hamilton, S.; Eaton, S.L.; Finlayson, J.; et al. Rabbits are not resistant to prion infection. Proc. Natl. Acad. Sci. USA 2012, 109, 5080–5085. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bartz, J.C.; Benavente, R.; Caughey, B.; Christensen, S.; Herbst, A.; Hoover, E.A.; Mathiason, C.K.; McKenzie, D.; Morales, R.; Schwabenlander, M.D.; et al. Chronic Wasting Disease: State of the Science. Pathogens 2024, 13, 138. https://doi.org/10.3390/pathogens13020138
Bartz JC, Benavente R, Caughey B, Christensen S, Herbst A, Hoover EA, Mathiason CK, McKenzie D, Morales R, Schwabenlander MD, et al. Chronic Wasting Disease: State of the Science. Pathogens. 2024; 13(2):138. https://doi.org/10.3390/pathogens13020138
Chicago/Turabian StyleBartz, Jason C., Rebeca Benavente, Byron Caughey, Sonja Christensen, Allen Herbst, Edward A. Hoover, Candace K. Mathiason, Debbie McKenzie, Rodrigo Morales, Marc D. Schwabenlander, and et al. 2024. "Chronic Wasting Disease: State of the Science" Pathogens 13, no. 2: 138. https://doi.org/10.3390/pathogens13020138
APA StyleBartz, J. C., Benavente, R., Caughey, B., Christensen, S., Herbst, A., Hoover, E. A., Mathiason, C. K., McKenzie, D., Morales, R., Schwabenlander, M. D., Walsh, D. P., & the NC1209: North American Interdisciplinary Chronic Wasting Disease Research Consortium Members. (2024). Chronic Wasting Disease: State of the Science. Pathogens, 13(2), 138. https://doi.org/10.3390/pathogens13020138






