The Placenta as a Target of Epigenetic Alterations in Women with Gestational Diabetes Mellitus and Potential Implications for the Offspring
Abstract
1. Introduction
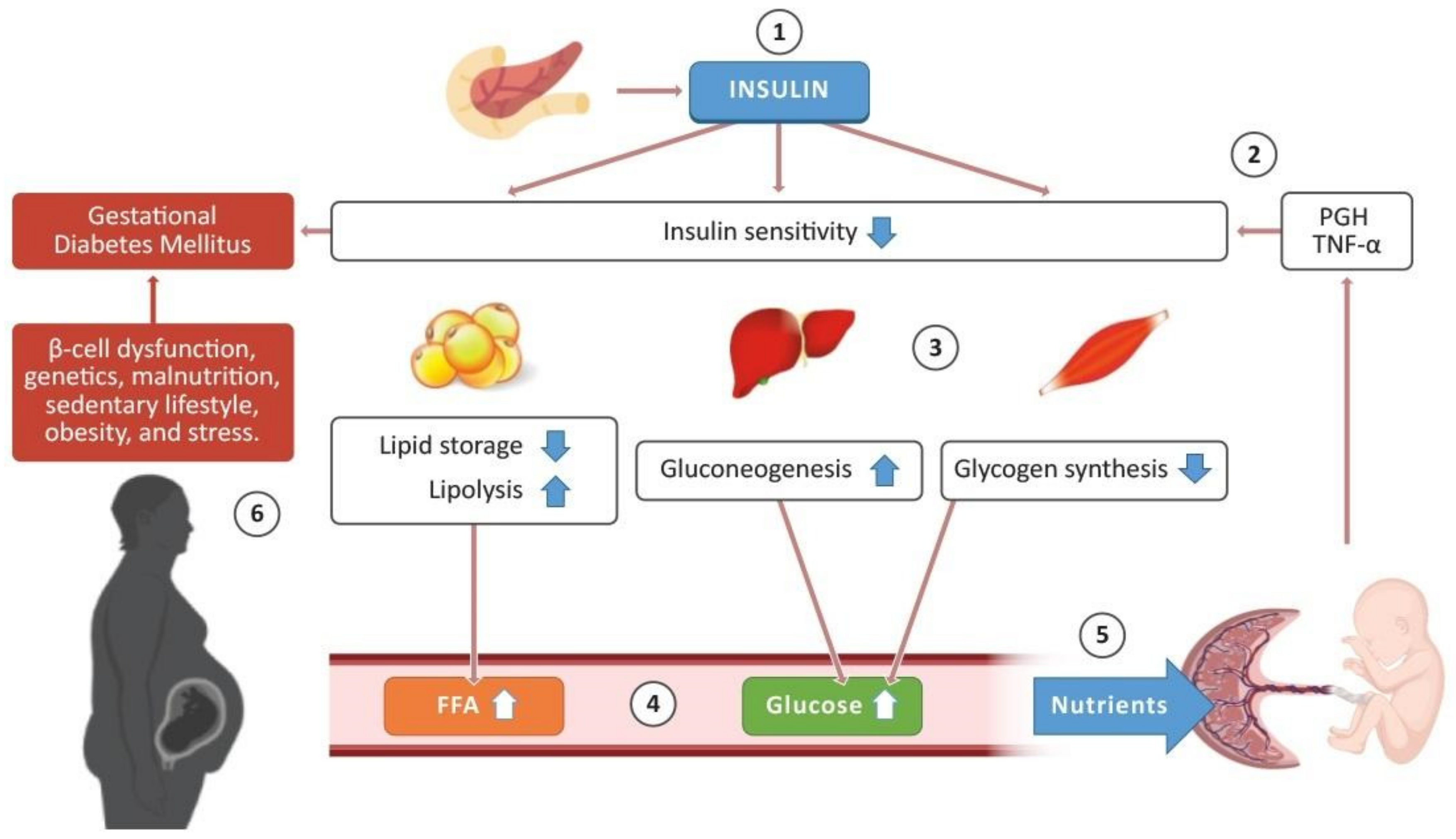
2. Insulin Sensitivity in Pregnancy
The Role of the Placenta
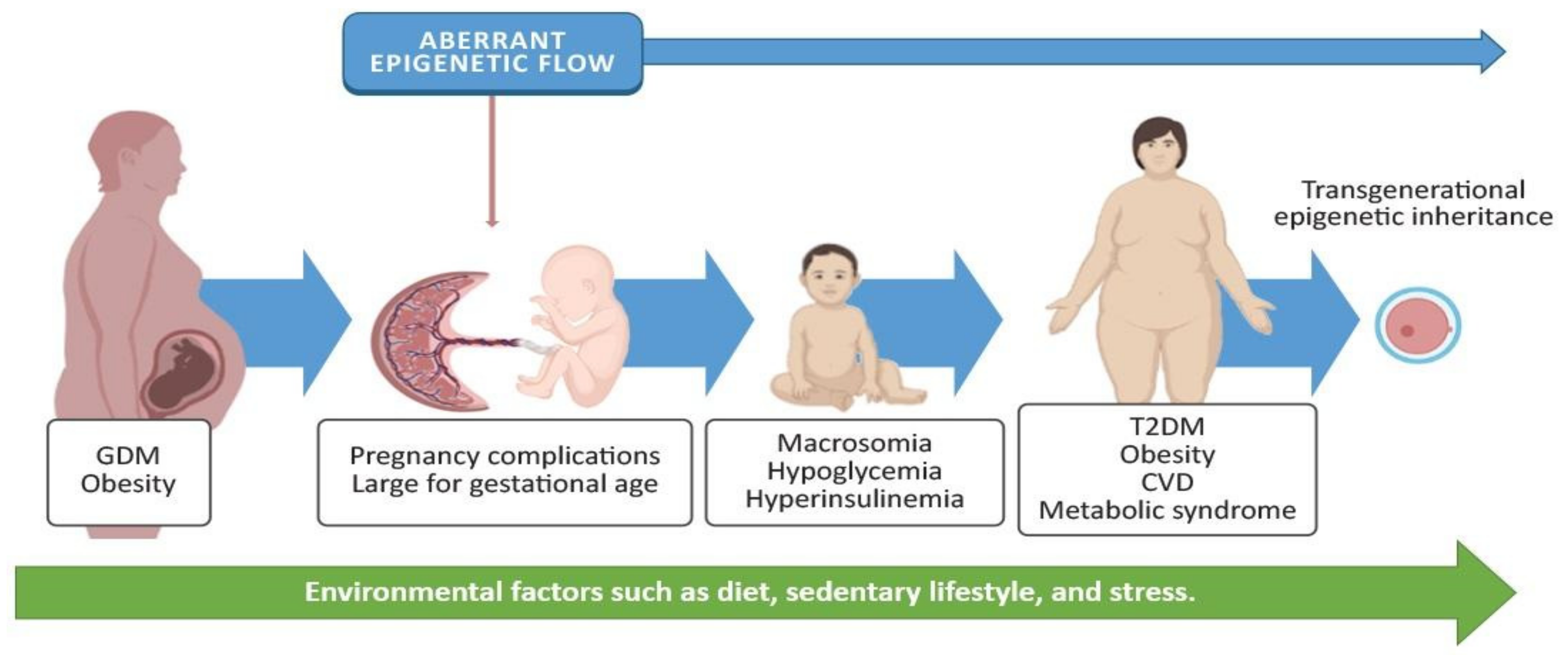
3. Gestational Diabetes Mellitus and Adverse Effects on the Offspring
4. Alterations in the Placenta in Women with GDM
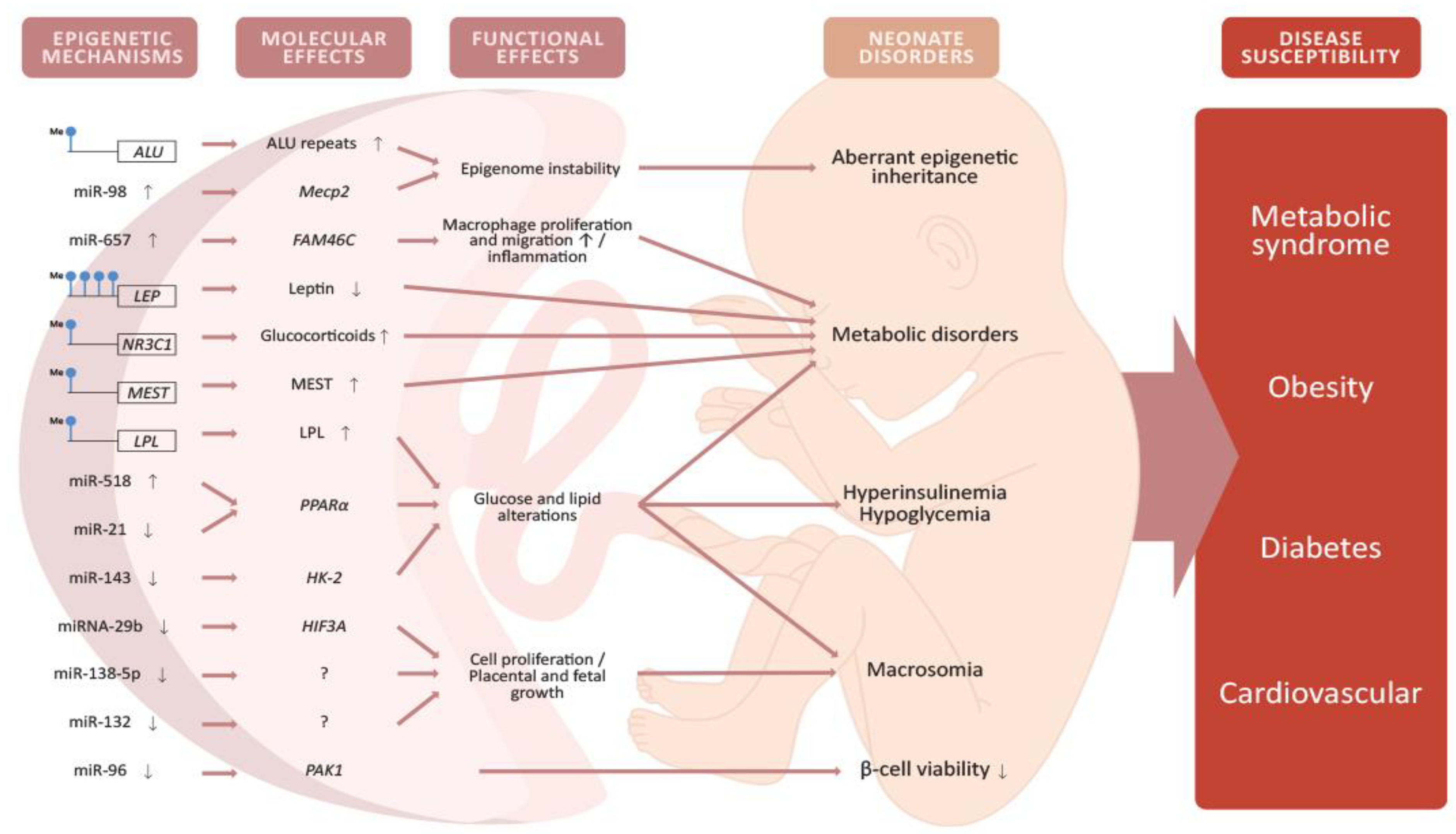
4.1. Epigenetic Modifications in the Placenta Caused by GDM
4.2. Placental DNA Methylation
4.3. miRNA Expression in the Placenta
4.4. Epigenetic Alterations in the Offspring and Potential Implications
5. Concluding Remarks
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- International Diabetes Federation. IDF Diabetes Atlas, 9th ed.; International Diabetes Federation: Brussels, Belgium, 2019; p. 53. [Google Scholar]
- Desai, M.; Beall, M.; Ross, M.G. Developmental Origins of Obesity: Programmed Adipogenesis. Curr. Diabetes Rep. 2013, 13, 27–33. [Google Scholar] [CrossRef] [PubMed]
- Catalano, P.M.; McIntyre, H.D.; Cruickshank, J.K.; McCance, D.R.; Dyer, A.R.; Metzger, B.E.; Lowe, L.P.; Trimble, E.R.; Coustan, D.R.; Hadden, D.R.; et al. The Hyperglycemia and Adverse Pregnancy Outcome Study: Associations of GDM and obesity with pregnancy outcomes. Diabetes Care 2012, 35, 780–786. [Google Scholar] [CrossRef]
- El-Sawy, N.A.; Iqbal, M.S.; AlKushi, A.G. Histomorphological Study of Placenta in Gestational Diabetes Mellitus. Int. J. Morphol. 2018, 36, 687–692. [Google Scholar] [CrossRef]
- Rudge, M.V.C.; Barbosa, A.M.P.; Sobrevia, L.; Gelaleti, R.B.; Hallur, R.L.S.; Marcondes, J.P.C.; Salvadori, D.M.F.; Prudêncio, C.B.; Magalhães, C.G.; Costa, R.; et al. Altered maternal metabolism during mild gestational hyperglycemia as a predictor of adverse perinatal outcomes: A comprehensive analysis. Biochim. Biophys. Acta (BBA) Mol. Basis Dis. 2020, 1866, 165478. [Google Scholar] [CrossRef]
- Plows, J.F.; Stanley, J.L.; Baker, P.N.; Reynolds, C.M.; Vickers, M.H. The Pathophysiology of Gestational Diabetes Mellitus. Int. J. Mol. Sci. 2018, 19, 3342. [Google Scholar] [CrossRef]
- Starikov, R.; Dudley, N.; Reddy, U.M. Stillbirth in the Pregnancy Complicated by Diabetes. Curr. Diabetes Rep. 2015, 15, 11. [Google Scholar] [CrossRef] [PubMed]
- Young, B.C.; Ecker, J.L. Fetal Macrosomia and Shoulder Dystocia in Women with Gestational Diabetes: Risks Amenable to Treatment? Curr. Diabetes Rep. 2013, 13, 12–18. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Arah, O.; Liew, Z.; Cnattingius, S.; Olsen, J.; Sørensen, H.T.; Qin, G.; Li, J. Maternal diabetes during pregnancy and early onset of cardiovascular disease in offspring: Population based cohort study with 40 years of follow-up. BMJ 2019, 367, 6398. [Google Scholar] [CrossRef] [PubMed]
- Lawlor, D.A.; Lichtenstein, P.; Långström, N. Association of Maternal Diabetes Mellitus in Pregnancy with Offspring Adiposity Into Early Adulthood. Circulation 2011, 123, 258–265. [Google Scholar] [CrossRef]
- Dabelea, D.; Hanson, R.L.; Lindsay, R.S.; Pettitt, D.J.; Imperatore, G.; Gabir, M.M.; Roumain, J.; Bennett, P.H.; Knowler, W.C. Intrauterine exposure to diabetes conveys risks for type 2 diabetes and obesity: A study of discordant sibships. Diabetes 2000, 49, 2208–2211. [Google Scholar] [CrossRef]
- Moore, T.R. Fetal exposure to gestational diabetes contributes to subsequent adult metabolic syndrome. Am. J. Obstet. Gynecol. 2010, 202, 643–649. [Google Scholar] [CrossRef]
- Barker, D. The Developmental Origins of Adult Disease. J. Am. Coll. Nutr. 2004, 23, 588S–595S. [Google Scholar] [CrossRef] [PubMed]
- Barker, D.J.P.; Thornburg, K.L.; Osmond, C.; Kajantie, E.; Eriksson, J.G. Beyond birthweight: The maternal and placental origins of chronic disease. J. Dev. Orig. Health Dis. 2010, 1, 360–364. [Google Scholar] [CrossRef]
- Thompson, R.F.; Einstein, F.H. Epigenetic Basis for Fetal Origins of Age-Related Disease. J. Women’s Health 2010, 19, 581–587. [Google Scholar] [CrossRef] [PubMed]
- Stiemsma, L.T.; Michels, K.B. The Role of the Microbiome in the Developmental Origins of Health and Disease. Pediatrics 2018, 141, e20172437. [Google Scholar] [CrossRef]
- Lacal, I.; Ventura, R. Epigenetic Inheritance: Concepts, Mechanisms and Perspectives. Front. Mol. Neurosci. 2018, 11, 292. [Google Scholar] [CrossRef] [PubMed]
- Al Aboud, N.M.; Shahid, H.; Jialal, I. Genetics, Epigenetic Mechanism; StatPearls Publishing: Treasure Island, FL, USA, 2020. [Google Scholar]
- Horsthemke, B. A critical view on transgenerational epigenetic inheritance in humans. Nat. Commun. 2018, 9, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Sailasree, S.P.; Srivastava, S.; Mishra, R.K. The placental gateway of maternal transgenerational epigenetic inheritance. J. Genet. 2017, 96, 465–482. [Google Scholar] [CrossRef]
- Nelissen, E.C.; Van Montfoort, A.P.; Dumoulin, J.C.; Evers, J.L. Epigenetics and the placenta. Hum. Reprod. Updat. 2010, 17, 397–417. [Google Scholar] [CrossRef] [PubMed]
- Januar, V.; Desoye, G.; Novakovic, B.; Cvitic, S.; Saffery, R. Epigenetic regulation of human placental function and pregnancy outcome: Considerations for causal inference. Am. J. Obstet. Gynecol. 2015, 213, S182–S196. [Google Scholar] [CrossRef]
- Lesseur, C.; Chen, J. Adverse Maternal Metabolic Intrauterine Environment and Placental Epigenetics: Implications for Fetal Metabolic Programming. Curr. Environ. Health Rep. 2018, 5, 531–543. [Google Scholar] [CrossRef]
- Filardi, T.; Catanzaro, G.; Mardente, S.; Zicari, A.; Santangelo, C.; Lenzi, A.; Morano, S.; Ferretti, E. Non-Coding RNA: Role in Gestational Diabetes Pathophysiology and Complications. Int. J. Mol. Sci. 2020, 21, 4020. [Google Scholar] [CrossRef] [PubMed]
- The Global Diabetes Community. Available online: https://www.diabetes.co.uk/insulin/insulin-sensitivity.html (accessed on 15 January 2019).
- Parsons, J.; Brelje, T.C.; Sorenson, R.L. Adaptation of islets of Langerhans to pregnancy: Increased islet cell proliferation and insulin secretion correlates with the onset of placental lactogen secretion. Endocrinolology 1992, 130, 1459–1466. [Google Scholar] [CrossRef]
- Freemark, M. Regulation of Maternal Metabolism by Pituitary and Placental Hormones: Roles in Fetal Development and Metabolic Programming. Horm. Res. Paediatr. 2006, 65, 41–49. [Google Scholar] [CrossRef]
- Newbern, D.; Freemark, M. Placental hormones and the control of maternal metabolism and fetal growth. Curr. Opin. Endocrinol. Diabetes Obes. 2011, 18, 409–416. [Google Scholar] [CrossRef]
- Sonagra, A.D. Normal Pregnancy- A State of Insulin Resistance. J. Clin. Diagn. Res. 2014, 8, CC01–CC03. [Google Scholar] [CrossRef]
- Lacroix, M.; Guibourdenche, J.; Frendo, J.; Muller, F.; Evain-Brion, D. Human Placental Growth Hormone—A Review. Placenta 2002, 23, S87–S94. [Google Scholar] [CrossRef]
- Barbour, L.A.; Shao, J.; Qiao, L.; Pulawa, L.K.; Jensen, D.R.; Bartke, A.; Garrity, M.; Draznin, B.; Friedman, J.E. Human placental growth hormone causes severe insulin resistance in transgenic mice. Am. J. Obstet. Gynecol. 2002, 186, 512–517. [Google Scholar] [CrossRef] [PubMed]
- Barbour, L.A.; Shao, J.; Qiao, L.; Leitner, W.; Anderson, M.; Friedman, J.E.; Draznin, B. Human Placental Growth Hormone Increases Expression of the P85 Regulatory Unit of Phosphatidylinositol 3-Kinase and Triggers Severe Insulin Resistance in Skeletal Muscle. Endocrinolology 2004, 145, 1144–1150. [Google Scholar] [CrossRef] [PubMed]
- Kirwan, J.P.; Mouzon, S.H.-D.; Lepercq, J.; Challier, J.-C.; Huston-Presley, L.; Friedman, J.E.; Kalhan, S.C.; Catalano, P.M. TNF Is a Predictor of Insulin Resistance in Human Pregnancy. Diabetes 2002, 51, 2207–2213. [Google Scholar] [CrossRef]
- Wieser, V.; Moschen, A.R.; Tilg, H. Inflammation, Cytokines and Insulin Resistance: A Clinical Perspective. Arch. Immunol. Ther. Exp. 2013, 61, 119–125. [Google Scholar] [CrossRef]
- American Diabetes Association. Classification and Diagnosis of Diabetes: Standards of Medical Care in Diabetes-2019. Diabetes Care 2018, 42 (Suppl. 1), S13–S28. [Google Scholar] [CrossRef]
- World Health Organization. Classification of Diabetes Mellitus; World Health Organization: Geneva, Switzerland, 2019; Licence: CC BY-NC-SA 3.0 IGO. [Google Scholar]
- Peters, R.; Xiang, A.; Kjos, S.; Buchanan, T. Long-term diabetogenic effect of single pregnancy in women with previous gestational diabetes mellitus. Lancet 1996, 347, 227–230. [Google Scholar] [CrossRef]
- Mondestin, M.A.; Ananth, C.V.; Smulian, J.C.; Vintzileos, A.M. Birth weight and fetal death in the United States: The effect of maternal diabetes during pregnancy. Am. J. Obstet. Gynecol. 2002, 187, 922–926. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.-R.; Dye, T.D.; Li, D. Effects of pre-gestational diabetes mellitus and gestational diabetes mellitus on macrosomia and birth defects in Upstate New York. Diabetes Res. Clin. Pract. 2019, 155, 107811. [Google Scholar] [CrossRef] [PubMed]
- De Lapertosa, S.G.; Alvariñas, J.; Elgart, J.F.; Salzberg, S.; Gagliardino, J.J.; EduGest Group. The triad macrosomia, obesity, and hypertriglyceridemia in gestational diabetes. Diabetes Metab. Res. Rev. 2020, 36, e03302. [Google Scholar] [CrossRef] [PubMed]
- Esakoff, T.F.; Cheng, Y.W.; Sparks, T.N.; Caughey, A.B. The association between birthweight 4000 g or greater and perinatal outcomes in patients with and without gestational diabetes mellitus. Am. J. Obstet. Gynecol. 2009, 200, 672.e1–672.e4. [Google Scholar] [CrossRef] [PubMed]
- Senniappan, S.; Shanti, B.; James, C.; Hussain, K. Hyperinsulinaemic hypoglycaemia: Genetic mechanisms, diagnosis and management. J. Inherit. Metab. Dis. 2012, 35, 589–601. [Google Scholar] [CrossRef]
- Xiang, A.H.; Wang, X.; Martinez, M.P.; Walthall, J.C.; Curry, E.S.; Page, K.; Buchanan, T.A.; Coleman, K.J.; Getahun, D. Association of Maternal Diabetes with Autism in Offspring. JAMA 2015, 313, 1425–1434. [Google Scholar] [CrossRef]
- Page, K.A.; Luo, S.; Wang, X.; Chow, T.; Alves, J.; Buchanan, T.A.; Xiang, A.H. Children Exposed to Maternal Obesity or Gestational Diabetes Mellitus During Early Fetal Development Have Hypothalamic Alterations That Predict Future Weight Gain. Diabetes Care 2019, 42, 1473–1480. [Google Scholar] [CrossRef]
- Higueras Sanz, M.T. Malformaciones de la placenta y el cordón umbilical. In Medicina Fetal, 1st ed.; Gratacós, E., Gómez, R., Nicolaides, K., Romero, R., Cabero, L., Eds.; Panamericana: España, Mexico, 2009; p. 822. [Google Scholar]
- Abdelhalim, N.Y.; Shehata, M.H.; Gadallah, H.N.; Sayed, W.M.; Othman, A.A. Morphological and ultrastructural changes in the placenta of the diabetic pregnant Egyptian women. Acta Histochem. 2018, 120, 490–503. [Google Scholar] [CrossRef] [PubMed]
- Carrasco-Wong, I.; Moller, A.; Giachini, F.R.; Lima, V.V.; Toledo, F.; Stojanova, J.; Sobrevia, L.; Martín, S.S. Placental structure in gestational diabetes mellitus. Biochim. Biophys. Acta (BBA) Mol. Basis Dis. 2020, 1866, 165535. [Google Scholar] [CrossRef]
- Daskalakis, G.; Marinopoulos, S.; Krielesi, V.; Papapanagiotou, A.; Papantoniou, N.; Mesogitis, S.; Antsaklis, A. Placental pathology in women with gestational diabetes. Acta Obstet. Gynecol. Scand. 2008, 87, 403–407. [Google Scholar] [CrossRef]
- Magee, T.R.; Ross, M.G.; Wedekind, L.; Desai, M.; Kjos, S.; Belkacemi, L. Gestational diabetes mellitus alters apoptotic and inflammatory gene expression of trophobasts from human term placenta. J. Diabetes Its Complicat. 2014, 28, 448–459. [Google Scholar] [CrossRef]
- Sandovici, I.; Hoelle, K.; Angiolini, E.; Constância, M. Placental adaptations to the maternal–fetal environment: Implications for fetal growth and developmental programming. Reprod. Biomed. Online 2012, 25, 68–89. [Google Scholar] [CrossRef]
- Myatt, L. Placental adaptive responses and fetal programming. J. Physiol. 2006, 572, 25–30. [Google Scholar] [CrossRef] [PubMed]
- Daskalakis, N.P.; Bagot, R.C.; Parker, K.J.; Vinkers, C.H.; de Kloet, E. The three-hit concept of vulnerability and resilience: Toward understanding adaptation to early-life adversity outcome. Psychoneuroendocrinology 2013, 38, 1858–1873. [Google Scholar] [CrossRef]
- Jin, B.; Li, Y.; Robertson, K.D. DNA Methylation: Superior or Subordinate in the Epigenetic Hierarchy? Genes Cancer 2011, 2, 607–617. [Google Scholar] [CrossRef] [PubMed]
- Weaver, I.C.G.; Korgan, A.C.; Lee, K.; Wheeler, R.V.; Hundert, A.S.; Goguen, D. Stress and the Emerging Roles of Chromatin Remodeling in Signal Integration and Stable Transmission of Reversible Phenotypes. Front. Behav. Neurosci. 2017, 11, 41. [Google Scholar] [CrossRef] [PubMed]
- Tessier, D.; Ferraro, Z.; Gruslin, A. Role of leptin in pregnancy: Consequences of maternal obesity. Placenta 2013, 34, 205–211. [Google Scholar] [CrossRef]
- Lesseur, C.; Armstrong, D.A.; Paquette, A.G.; Li, Z.; Padbury, J.F.; Marsit, C.J. Maternal obesity and gestational diabetes are associated with placental leptin DNA methylation. Am. J. Obstet. Gynecol. 2014, 211, 654.e1–654.e9. [Google Scholar] [CrossRef]
- Klok, M.D.; Jakobsdottir, S.; Drent, M.L. The role of leptin and ghrelin in the regulation of food intake and body weight in humans: A review. Obes. Rev. 2006, 8, 21–34. [Google Scholar] [CrossRef]
- Pérez-Pérez, A.; Maymo, J.; Gambino, Y.; Guadix, P.; Dueñas, J.L.; Varone, C.; Sánchez-Margalet, V. Activated Translation Signaling in Placenta from Pregnant Women with Gestational Diabetes Mellitus: Possible Role of Leptin. Horm. Metab. Res. 2013, 45, 436–442. [Google Scholar] [CrossRef]
- Pérez-Pérez, A.; Toro, A.; Vilariño-García, T.; Maymó, J.; Guadix, P.; Dueñas, J.L.; Fernández-Sánchez, M.; Varone, C.; Sánchez-Margalet, V. Leptin action in normal and pathological pregnancies. J. Cell. Mol. Med. 2017, 22, 716–727. [Google Scholar] [CrossRef]
- Gagné-Ouellet, V.; Houde, A.-A.; Guay, S.-P.; Perron, P.; Gaudet, D.; Guérin, R.; Jean-Patrice, B.; Hivert, M.-F.; Brisson, D.; Bouchard, L. Placental lipoprotein lipase DNA methylation alterations are associated with gestational diabetes and body composition at 5 years of age. Epigenetics 2017, 12, 616–625. [Google Scholar] [CrossRef]
- Pirahanchi, Y.; Anoruo, M.D.; Sharma, S. Biochemistry, Lipoprotein Lipase; StatPearls Publishing: Treasure Island, FL, USA, 2020. [Google Scholar]
- El Hajj, N.; Pliushch, G.; Schneider, E.; Dittrich, M.; Müller, T.; Korenkov, M.; Aretz, M.; Zechner, U.; Lehnen, H.; Haaf, T. Metabolic Programming of MEST DNA Methylation by Intrauterine Exposure to Gestational Diabetes Mellitus. Diabetes 2012, 62, 1320–1328. [Google Scholar] [CrossRef] [PubMed]
- Shi, W.; Lefebvre, L.; Yu, Y.; Otto, S.; Krella, A.; Orth, A.; Fundele, R. Loss-of-imprinting ofPeg1 in mouse interspecies hybrids is correlated with altered growth. Genestics 2004, 39, 65–72. [Google Scholar] [CrossRef]
- Paquette, A.G.; Lester, B.M.; Lesseur, C.; Armstrong, D.; Guerin, D.J.; Appleton, A.; Marsit, C.J. Placental epigenetic patterning of glucocorticoid response genes is associated with infant neurodevelopment. Epigenomics 2015, 7, 767–779. [Google Scholar] [CrossRef] [PubMed]
- Oberlander, T.F.; Weinberg, J.; Papsdorf, M.; Grunau, R.; Misri, S.; Devlin, A.M. Prenatal exposure to maternal depression, neonatal methylation of human glucocorticoid receptor gene (NR3C1) and infant cortisol stress responses. Epigenetics 2008, 3, 97–106. [Google Scholar] [CrossRef]
- Pray, L. Transposons: The jumping genes. Nat. Educ. 2008, 1, 204. [Google Scholar]
- Vryer, R.; Saffery, R. What’s in a name? Context-dependent significance of ‘global’ methylation measures in human health and disease. Clin. Epigenetics 2017, 9, 2. [Google Scholar] [CrossRef] [PubMed]
- Nomura, Y.; Lambertini, L.; Rialdi, A.; Lee, M.; Ba, E.Y.M.; Grabie, M.; Bs, I.M.; Huynh, N.; Finik, J.; Davey, M.; et al. Global Methylation in the Placenta and Umbilical Cord Blood from Pregnancies with Maternal Gestational Diabetes, Preeclampsia, and Obesity. Reprod. Sci. 2013, 21, 131–137. [Google Scholar] [CrossRef]
- Reichetzeder, C.; Putra, S.E.D.; Pfab, T.; Slowinski, T.; Neuber, C.; Kleuser, B.; Hocher, B. Increased global placental DNA methylation levels are associated with gestational diabetes. Clin. Epigenetics 2016, 8, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Hivert, M.-F.; Cardenas, A.; Allard, C.; Doyon, M.; Powe, C.E.; Catalano, P.M.; Perron, P.; Bouchard, L. Interplay of Placental DNA Methylation and Maternal Insulin Sensitivity in Pregnancy. Diabetes 2019, 69, 484–492. [Google Scholar] [CrossRef]
- Cai, M.; Kolluru, G.K.; Ahmed, A. Small Molecule, Big Prospects: MicroRNA in Pregnancy and Its Complications. J. Pregnancy 2017, 2017, 6972732. [Google Scholar] [CrossRef]
- Cao, J.-L.; Zhang, L.; Li, J.; Tian, S.; Lv, X.-D.; Wang, X.-Q.; Su, X.; Li, Y.; Hu, Y.; Ma, X.; et al. Up-regulation of miR-98 and unraveling regulatory mechanisms in gestational diabetes mellitus. Sci. Rep. 2016, 6, 32268. [Google Scholar] [CrossRef]
- Li, C.H.; Coffey, E.L.; Dall’Agnese, A.; Hannett, N.M.; Tang, X.; Henninger, J.E.; Platt, J.M.; Oksuz, O.; Zamudio, A.V.; Afeyan, L.K.; et al. MeCP2 links heterochromatin condensates and neurodevelopmental disease. Nature 2020, 586, 440–444. [Google Scholar] [CrossRef] [PubMed]
- Bach, S.; Ryan, N.M.; Guasoni, P.; Corvin, A.P.; El-Nemr, R.A.; Khan, D.; Sanfeliu, A.; Tropea, D. Methyl-CpG-binding protein 2 mediates overlapping mechanisms across brain disorders. Sci. Rep. 2020, 10, 1–13. [Google Scholar] [CrossRef]
- Zhao, C.; Zhang, T.; Shi, Z.; Ding, H.; Ling, X. MicroRNA-518d regulates PPARα protein expression in the placentas of females with gestational diabetes mellitus. Mol. Med. Rep. 2014, 9, 2085–2090. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Cai, Y.; Jian, L.; Cheung, C.W.; Zhang, L.; Xia, Z. Impact of peroxisome proliferator-activated receptor-α on diabetic cardiomyopathy. Cardiovasc. Diabetol. 2021, 20, 1–15. [Google Scholar] [CrossRef]
- Guan, C.-Y.; Tian, S.; Cao, J.-L.; Wang, X.-Q.; Ma, X.; Xia, H.-F. Down-Regulated miR-21 in Gestational Diabetes Mellitus Placenta Induces PPAR-α to Inhibit Cell Proliferation and Infiltration. Diabetes Metab. Syndr. Obes. Targets Ther. 2020, 13, 3009–3034. [Google Scholar] [CrossRef] [PubMed]
- Muralimanoharan, S.; Maloyan, A.; Myatt, L. Mitochondrial function and glucose metabolism in the placenta with gestational diabetes mellitus: Role of miR-143. Clin. Sci. 2016, 130, 931–941. [Google Scholar] [CrossRef]
- Nakajima, K.; Kawashima, I.; Koshiisi, M.; Kumagai, T.; Suzuki, M.; Suzuki, J.; Mitsumori, T.; Kirito, K. Glycolytic enzyme hexokinase II is a putative therapeutic target in B-cell malignant lymphoma. Exp. Hematol. 2019, 78, 46–55.e3. [Google Scholar] [CrossRef]
- Zhou, X.; Xiang, C.; Zheng, X. miR-132 serves as a diagnostic biomarker in gestational diabetes mellitus and its regulatory effect on trophoblast cell viability. Diagn. Pathol. 2019, 14, 1–7. [Google Scholar] [CrossRef]
- Li, L.; Wang, S.; Li, H.; Wan, J.; Zhou, Q.; Zhou, Y.; Zhang, C. microRNA-96 protects pancreatic β-cell function by targeting PAK1 in gestational diabetes mellitus. BioFactors 2018, 44, 539–547. [Google Scholar] [CrossRef]
- Ahn, M.; Yoder, S.M.; Wang, Z.; Oh, E.; Ramalingam, L.; Tunduguru, R.; Thurmond, D.C. The p21-activated kinase (PAK1) is involved in diet-induced beta cell mass expansion and survival in mice and human islets. Diabetologia 2016, 59, 2145–2155. [Google Scholar] [CrossRef] [PubMed]
- Sun, D.-G.; Tian, S.; Zhang, L.; Hu, Y.; Guan, C.-Y.; Ma, X.; Xia, H.-F. The miRNA-29b Is Downregulated in Placenta During Gestational Diabetes Mellitus and May Alter Placenta Development by Regulating Trophoblast Migration and Invasion Through a HIF3A-Dependent Mechanism. Front. Endocrinol. 2020, 11, 169. [Google Scholar] [CrossRef]
- Pfeiffer, S.; Krüger, J.; Maierhofer, A.; Böttcher, Y.; Klöting, N.; El Hajj, N.; Schleinitz, D.; Schön, M.R.; Dietrich, A.; Fasshauer, M.; et al. Hypoxia-inducible factor 3A gene expression and methylation in adipose tissue is related to adipose tissue dysfunction. Sci. Rep. 2016, 6, 27969. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Wang, Z.; Liu, G.; Jin, C.; Zhang, Q.; Man, S.; Wang, Z. miR-657 Promotes Macrophage Polarization toward M1 by Targeting FAM46C in Gestational Diabetes Mellitus. Mediat. Inflamm. 2019, 2019, 4851214. [Google Scholar] [CrossRef]
- Mroczek, S.; Chlebowska, J.; Kuliński, T.M.; Gewartowska, O.; Gruchota, J.; Cysewski, D.; Liudkovska, V.; Borsuk, E.; Nowis, D.; Dziembowski, A. The non-canonical poly(A) polymerase FAM46C acts as an onco-suppressor in multiple myeloma. Nat. Commun. 2017, 8, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Song, L.; Zhou, L.; Wu, J.; Sheng, C.; Chen, H.; Liu, Y.; Gao, S.; Huang, W. A MicroRNA Signature in Gestational Diabetes Mellitus Associated with Risk of Macrosomia. Cell. Physiol. Biochem. 2015, 37, 243–252. [Google Scholar] [CrossRef]
- Marte, B.M. PKB/Akt: Connecting phosphoinositide 3-kinase to cell survival and beyond. Trends Biochem. Sci. 1997, 22, 355–358. [Google Scholar] [CrossRef]
- Nair, S.; Jayabalan, N.; Guanzon, D.; Palma, C.; Scholz-Romero, K.; Elfeky, O.; Zuñiga, F.; Ormazabal, V.; Diaz, E.; Rice, G.E.; et al. Human placental exosomes in gestational diabetes mellitus carry a specific set of miRNAs associated with skeletal muscle insulin sensitivity. Clin. Sci. 2018, 132, 2451–2467. [Google Scholar] [CrossRef] [PubMed]
- Meloche, S.; Pouysségur, J. The ERK1/2 mitogen-activated protein kinase pathway as a master regulator of the G1- to S-phase transition. Oncogene 2007, 26, 3227–3239. [Google Scholar] [CrossRef] [PubMed]
- Ding, R.; Guo, F.; Zhang, Y.; Liu, X.-M.; Xiang, Y.-Q.; Zhang, C.; Liu, Z.-W.; Sheng, J.-Z.; Huang, H.-F.; Zhang, J.-Y.; et al. Integrated Transcriptome Sequencing Analysis Reveals Role of miR-138-5p/TBL1X in Placenta from Gestational Diabetes Mellitus. Cell. Physiol. Biochem. 2018, 51, 630–646. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zhang, T.; Sun, D.; Cheng, G.; Ren, H.; Hong, H.; Chen, L.; Jiao, X.; Du, Y.; Zou, Y.; et al. Diagnostic value of dysregulated microribonucleic acids in the placenta and circulating exosomes in gestational diabetes mellitus. J. Diabetes Investig. 2021. [Google Scholar] [CrossRef]
- Tang, L.; Li, P.; Li, L. Whole transcriptome expression profiles in placenta samples from women with gestational diabetes mellitus. J. Diabetes Investig. 2020, 11, 1307–1317. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Feng, J.; Cheng, F.; Cui, X.; Gao, L.; Chen, Y.; Wang, F.; Zhong, T.; Li, Y.; Liu, L. Circular RNA expression profiles in placental villi from women with gestational diabetes mellitus. Biochem. Biophys. Res. Commun. 2018, 498, 743–750. [Google Scholar] [CrossRef]
- Gu, Y.; Chu, X.; Morgan, J.A.; Lewis, D.F.; Wang, Y. Upregulation of METTL3 expression and m6A RNA methylation in placental trophoblasts in preeclampsia. Placenta 2021, 103, 43–49. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Huang, Q.; Liu, Y.; Garmire, L.X. Single cell transcriptome research in human placenta. Reproduction 2020, 160, R155–R167. [Google Scholar] [CrossRef] [PubMed]
- Howe, C.G.; Cox, B.; Fore, R.; Jungius, J.; Kvist, T.; Lent, S.; Miles, H.E.; Salas, L.A.; Rifas-Shiman, S.; Starling, A.P.; et al. Maternal Gestational Diabetes Mellitus and Newborn DNA Methylation: Findings from the Pregnancy and Childhood Epigenetics Consortium. Diabetes Care 2020, 43, 98–105. [Google Scholar] [CrossRef]
- Kim, E.; Kwak, S.H.; Chung, H.R.; Ohn, J.H.; Bae, J.H.; Choi, S.H.; Park, K.S.; Hong, J.-S.; Sung, J.; Jang, H.C. DNA methylation profiles in sibling pairs discordant for intrauterine exposure to maternal gestational diabetes. Epigenetics 2017, 12, 825–832. [Google Scholar] [CrossRef] [PubMed]
- Hjort, L.; Martino, D.; Grunnet, L.G.; Naeem, H.; Maksimovic, J.; Olsson, A.H.; Zhang, C.; Ling, C.; Olsen, S.F.; Saffery, R.; et al. Gestational diabetes and maternal obesity are associated with epigenome-wide methylation changes in children. JCI Insight 2018, 3, 122572. [Google Scholar] [CrossRef] [PubMed]
- Houshmand-Oeregaard, A.; Schrölkamp, M.; Kelstrup, L.; Hansen, N.S.; Hjort, L.; Thuesen, A.C.B.; Broholm, C.; Mathiesen, E.R.; Clausen, T.D.; Vaag, A.; et al. Increased expression of microRNA-15a and microRNA-15b in skeletal muscle from adult offspring of women with diabetes in pregnancy. Hum. Mol. Genet. 2018, 27, 1763–1771. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lizárraga, D.; García-Gasca, A. The Placenta as a Target of Epigenetic Alterations in Women with Gestational Diabetes Mellitus and Potential Implications for the Offspring. Epigenomes 2021, 5, 13. https://doi.org/10.3390/epigenomes5020013
Lizárraga D, García-Gasca A. The Placenta as a Target of Epigenetic Alterations in Women with Gestational Diabetes Mellitus and Potential Implications for the Offspring. Epigenomes. 2021; 5(2):13. https://doi.org/10.3390/epigenomes5020013
Chicago/Turabian StyleLizárraga, Dennise, and Alejandra García-Gasca. 2021. "The Placenta as a Target of Epigenetic Alterations in Women with Gestational Diabetes Mellitus and Potential Implications for the Offspring" Epigenomes 5, no. 2: 13. https://doi.org/10.3390/epigenomes5020013
APA StyleLizárraga, D., & García-Gasca, A. (2021). The Placenta as a Target of Epigenetic Alterations in Women with Gestational Diabetes Mellitus and Potential Implications for the Offspring. Epigenomes, 5(2), 13. https://doi.org/10.3390/epigenomes5020013






