Simple Summary
A new species of aphid, causing the formation of large galls on a sumac species endemic to Yunnan, China, is named and described. Molecular and morphological diagnostic criteria are provided, and digital specimen data made available.
Abstract
A new species of gall-forming aphid from China, Qiao jinshaensis gen. et sp. nov., is described from Rhus wilsonii Hemsl. Morphological identification and molecular analyses both support the establishment of a new genus. A diagnosis combining morphological and molecular characters from alate viviparae is provided and specimen metadata are published in an open-access and machine-readable format.
Keywords:
Anacardiaceae; aphid; Eriosomatinae; gall; Qiao gen. nov.; Qiao jinshaensis sp. nov.; Sternorrhyncha; sumac 1. Introduction
In China, Rhus-feeding aphids (Hemiptera: Aphididae: Eriosomatinae: Fordini) are of economic importance for the tannins extracted from the galls they produce [1]. There are currently 13 aphid species, in six genera, known to colonize only six species of Rhus (Anacardiaceae) [2,3]. Different species can make distinctly shaped galls on the same plant host. Phylogenetic analyses support the monophyly of most genera, with the exceptional placement of Nurudea ibofushi Matsumura, 1917 and Meitanaphis flavogallis Tang, 1978, which are repeatedly placed within Schlechtendalia Lichtenstein, 1883 and Kaburagia Takagi, 1937 respectively [4,5,6,7].
Aphid galls were collected for the first time on Rhus wilsonii Hemsl. in Yunnan, China. Their shapes were similar to those formed by some Nurudea Matsumura, 1917 and Floraphis Tsai & Tang, 1946 species, but the morphological identification of the aphids did not correspond to any currently known species. Morphological and molecular analyses were conducted but were not conclusive in assigning the species to any existing genus, thus supporting the establishment of a new one. We here formally establish Qiao jinshaensis gen. et sp. nov. Morphological and molecular diagnostic criteria are used following the protocol in Hébert et al. [8]. Additionally, the specimen collection data are published in a machine-readable format, making them readily accessible for future research.
2. Materials and Methods
2.1. Field Sampling
Fresh galls were collected in Yunnan, China (28.014° N, 103.638° E) on the host plant, Rhus wilsonii, and taken to the lab (see below for full collection data). Aphids were extracted from the galls and immersed in 70% ethanol for slide preparation and 100% ethanol for DNA extraction.
2.2. Molecular Analysis
Total genomic DNA was extracted from 20–30 individual aphids collected from a gall using Ezup animal genome DNA extraction kit at Sangon Biotech Co., Ltd., (Shanghai, China), according to the manufacturer’s protocol. The standard primer pair EF2 and EF3 [9] were used to amplify the elongation factor 1-alpha (EF1α) gene and HCO2198 and LCO1490 [10] were used to amplify a region of the mitochondrial cytochrome c oxidase subunit 1 (COI) gene. Polymerase chain reaction (PCR) amplification was carried out using standard protocols [11] with annealing temperatures of 56 °C for the EF1α locus and 55 °C for the COI locus. To ensure the accuracy of the sequence, the EF1α target fragment was incised and purified using a gel extraction kit (EZ-10 Column DNA Purification Kit, Bio Basic Inc., Markham, ON, Canada) according to the manufacturer instructions. Purified PCR products were sequenced in both directions on an ABI-3730XL gene sequencer at Sangon Biotech Co., Ltd. (Shanghai, China). The two sequences have been deposited in GenBank (accession numbers OP536221 and OP548621).
Introns in the EF1α sequence were removed in Geneious Prime 2022 software (Biomatters Ltd., Auckland, New Zealand) for analysis. Simple genetic p-distances were calculated with a Clustal Omega multiple alignment [12], as implemented in Geneious, with EF1α and COI sequences from other Rhus-gall aphids (see Table 1 for accession numbers). Simple (single nucleotide) and compound (multiple nucleotides) molecular diagnostic characters were retrieved manually for the COI sequence. Nucleotide positions are in reference to the COI alignment with the Acyrthosiphon pisum (Harris, 1776) mitochondrial genome (accession FJ411411).

Table 1.
Species compared in this study with their GenBank accession numbers for COI and EF1α genes.
The tannin content of the new galls was measured by ultraviolet spectrometry [13,14].
2.3. Morphological Examination
Fourteen aphids were cleared and slide-mounted in Canada balsam [15]. The identification key of Blackman and Eastop [2], as well as the original published descriptions of other Rhus-gall aphids [16,17,18,19], were used to assess the identity of the new species. Measurements and pictures were taken under brightfield conditions with a Zeiss M2 AxioImager microscope, an AxioCam HRc camera, and Zen 2012 Software, version 1.1.1.0 (Carl Zeiss AG, Oberkochen, Germany). All measurements are in micrometers (μm).
Morphological abbreviations are as follows: BL—body length (measured from the frontal margin of the head to the end of the cauda); BW—basal width; ANT I, II, III, IV, V, VI—antennal segments I, II, III, IV, V, VI or their length; PT—processus terminalis of the last antennal segment or its length; BASE—base of the last antennal segment or its length; URS—ultimate rostral segments (IV + V) or their length; LMF—length of the metafemur; LMT—length of the metatibia; WMT—width of the metatibia (measured in the middle); LMTS—length of the longest setae of the metatibia; HT I—first segment of the hind tarsus or its length (measured on the ventral side); HT II—second segment of the hind tarsus or its length.
3. Results
3.1. Molecular Analysis
The COI genetic p-distances between the new species and other species of Rhus-feeding Fordini ranged from 9.4% to 12.4%, much higher than the mean 7.3% infrageneric divergence found among aphids in general [20], and within the 9.7–11.3% range of intergeneric divergence we measured between Rhus-feeding Fordini (Table 1). Additionally, the average sequence divergence between the new species and other Rhus-feeding genera was 10.6%, greater than the 9.5% divergence between the other genera excluding the new species. The EF1α sequence divergence between the new species and other Rhus-feeding Fordini ranged from 5.1% to 7.7%. The average EF1α sequence divergence between the new species and Rhus-feeding genera was 6.1%, much higher than that between just the other genera (excluding the new species), which was 3.8%. These data confirm that the new species is genetically distinct, at the generic level, from all other Rhus-galling aphids for which DNA sequence data are available.
3.2. Taxonomic Treatment
The genus and species are established together, in accordance with Article 13.4 of the International Code of Zoological Nomenclature (4th Edition) [21].
Qiao gen. nov.
urn:lsid:zoobank.org:act:F2600ADA-F5A0-4478-BECC-F108676F11AE
Qiao jinshaensis sp. nov.
urn:lsid:zoobank.org:act:DFF64645-6FD5-4F86-869C-9CFADD376D66
3.2.1. Types
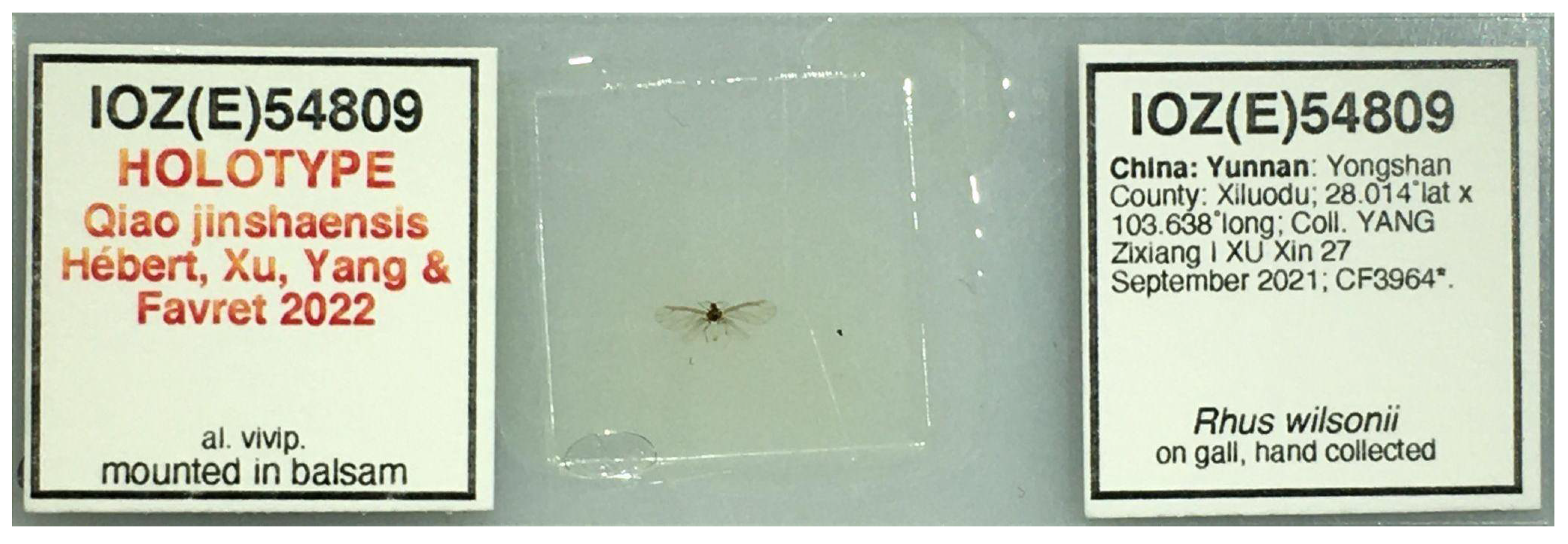
The type species of the genus Qiao gen nov. is Q. jinshaensis sp. nov. The holotype of the Q. jinshaensis is deposited at the Institute of Zoology of the Chinese Academy of Sciences (IOZ(E)) with the catalog number IOZ(E)54809 (Figure 1): alate viviparous female: China: Yunnan Province: Yongshan County: Xiluodu Town; 28.014° N, 103.638° E, 1005 m elev.; coll. 2021-09-27; YANG Zixiang and XU Xin leg.; in gall of Rhus wilsonii Hemsl. Paratypes are deposited at the IOZ(E) and the University of Montreal Ouellet-Robert Entomological Collection (QMOR): 13 alate viviparous females, same data as holotype, IOZ(E)54810-54811, QMOR66201-QMOR66204, QMOR66206-QMOR66212. These same specimen collection data are provided in machine-readable format by Hébert et al. [22].

Figure 1.
Holotype slide of Qiao jinshaensis sp. nov. (IOZ(E)54809).
3.2.2. Diagnosis
The new genus can be distinguished from other Rhus-galling aphids by several characteristics. It has six antennal segments compared to five in Nurudea and Schlechtendalia. It differs from Meitanaphis Tsai & Tang, 1946, by having a short and oblique stigma on the forewing and three (rarely two) hamuli on the hindwing costa (stigma long and sickle-shaped and two (sometimes one) hamuli in Meitanaphis). The numerous ring-like sensoria of Floraphis and Melaphis Walsh, 1867, differentiate them from our new genus, which has large, oblong sensoria. Lastly, Kaburagia has only one large oval sensorium on ANT III-VI, whereas our species has more variation in shapes and numbers of sensoria, with the larger ones on ANT IV-VI enclosing small, sclerotized islands. Thirteen simple and four compound nucleotide characters were unique to our new species when the COI sequence was compared with the other available aphid species causing galls on Rhus (Table 2).

Table 2.
Simple and compound diagnostic characters of Qiao jinshaensis sp. nov. for COI locus when compared with other Rhus-gall aphids.
3.2.3. Description
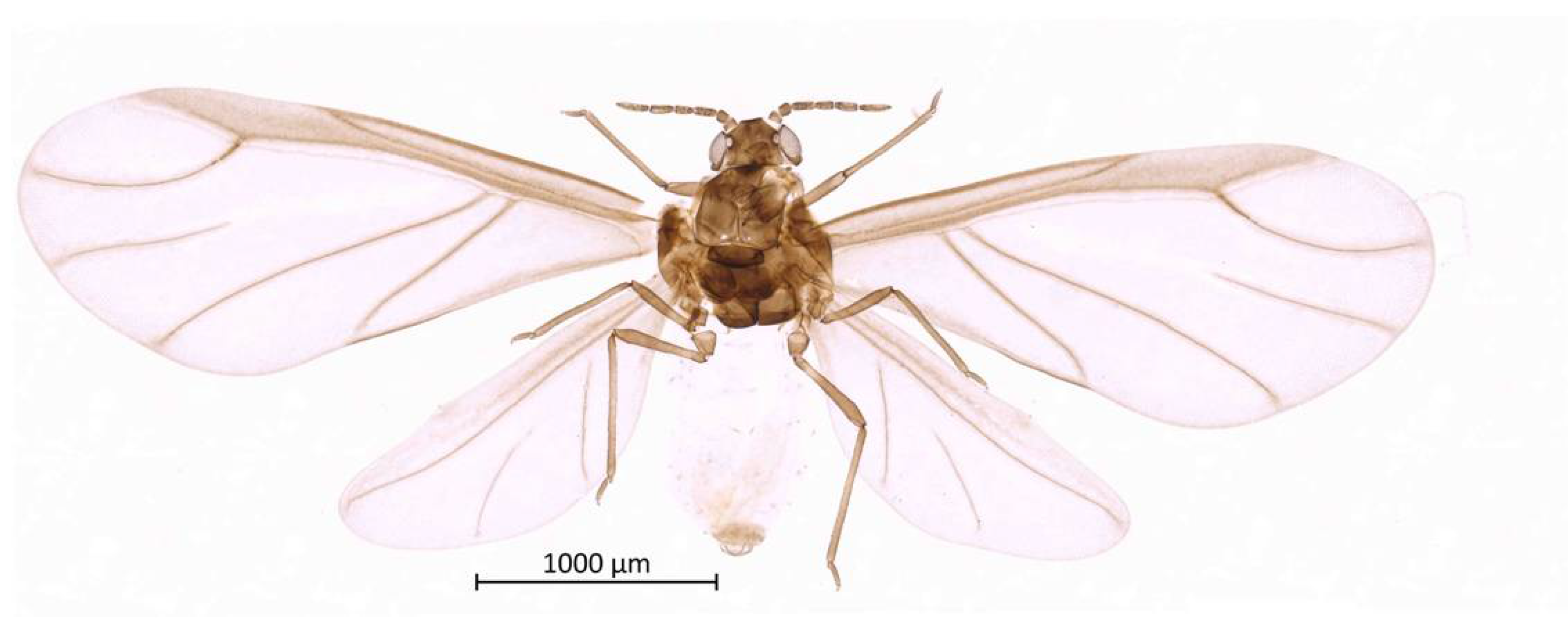
Color. In prepared specimens: Head, antennae, thorax and legs medium-brown (Figure 2). Genital plate, anal plate, and cauda light brown. Abdomen very light, almost transparent.

Figure 2.
Habitus of alate vivipara holotype of Qiao jinshaensis sp. nov. (IOZ(E)54809).
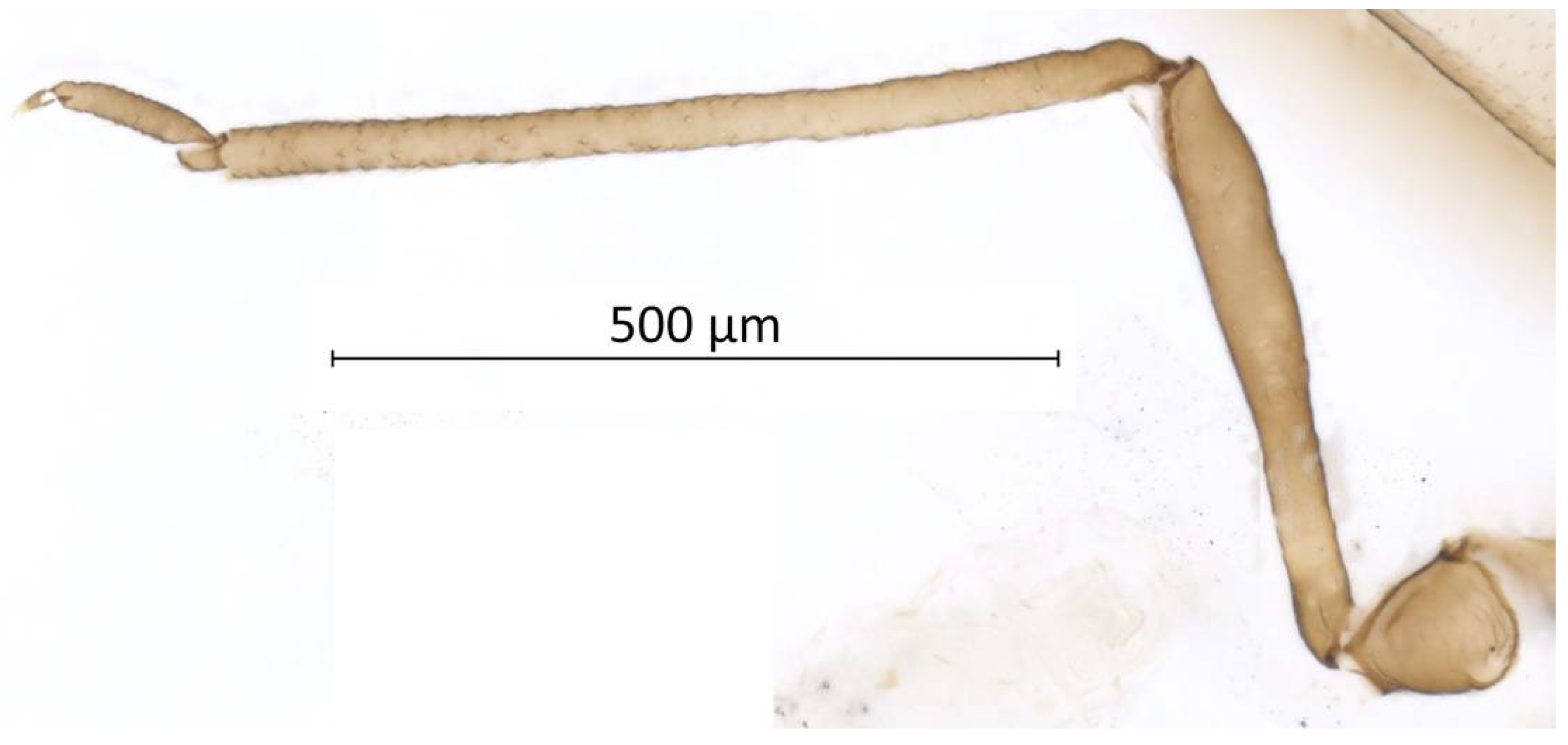
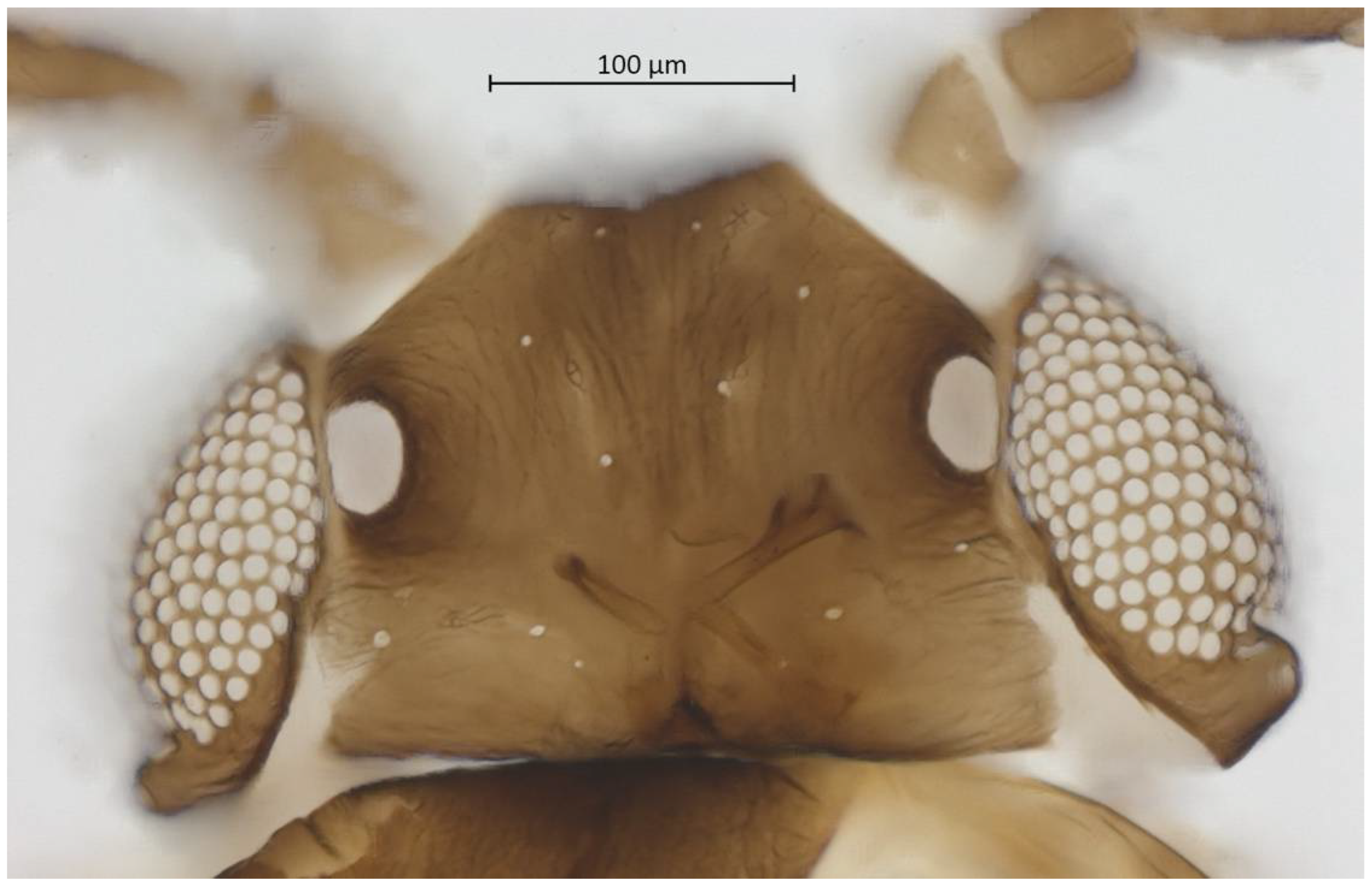
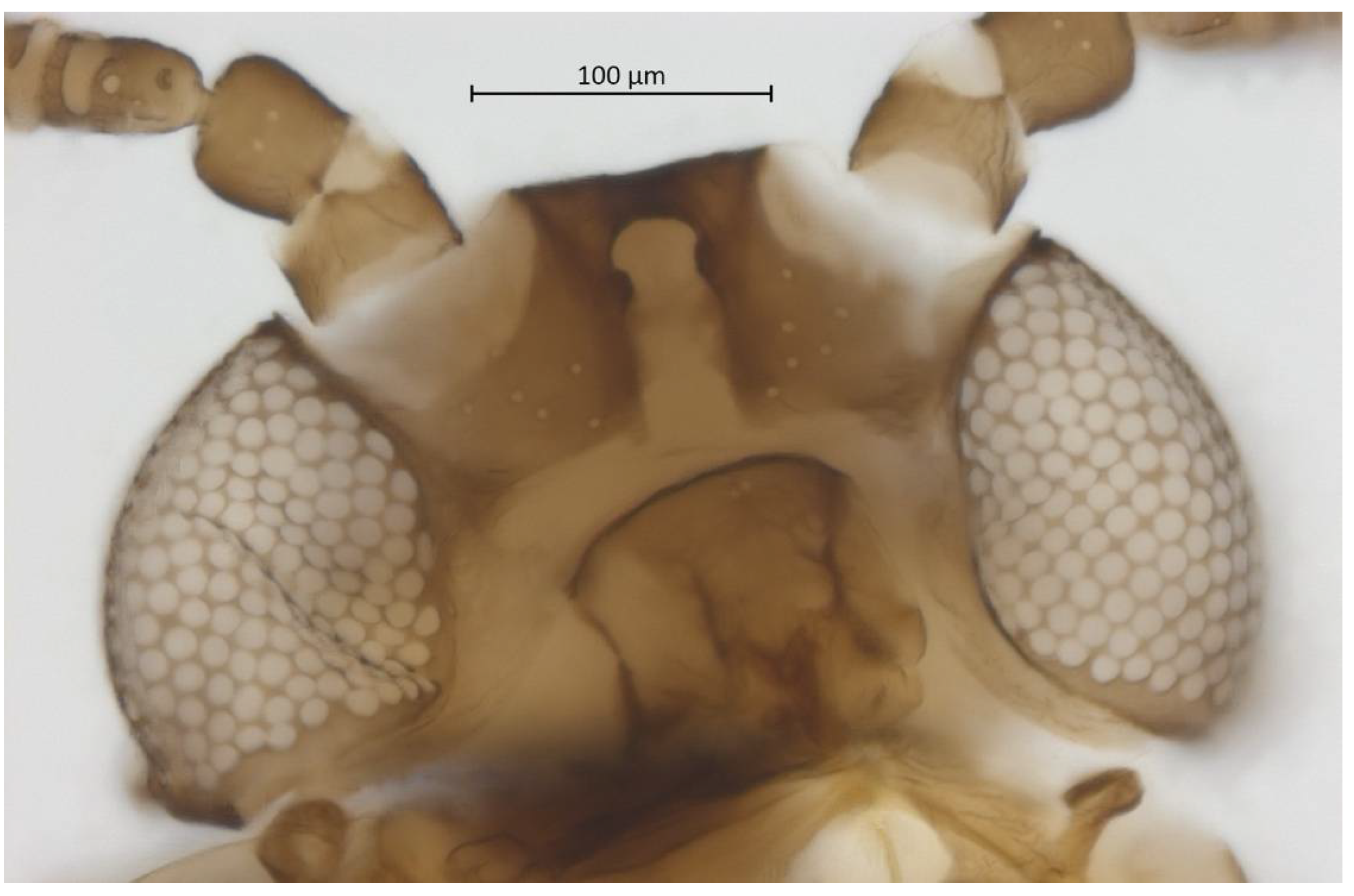
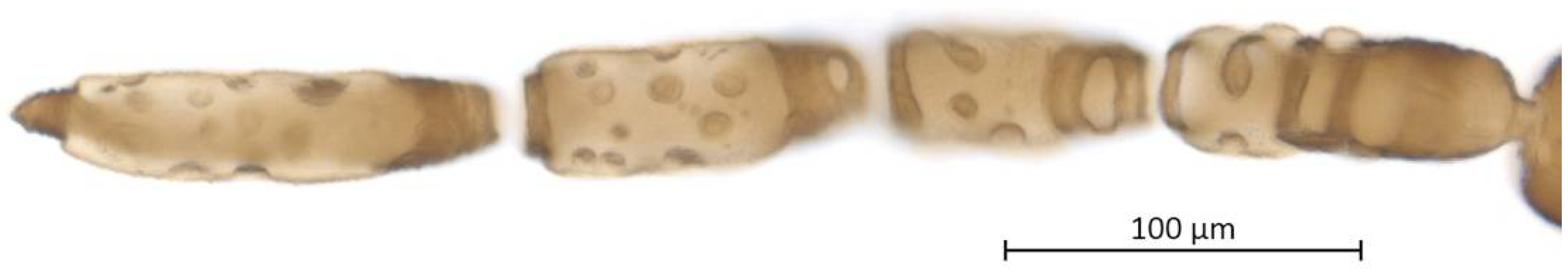
Head. Triommatidium distinct (Figure 3). Pale, linear ventral depression from the middle ocellus to the clypeus (Figure 4). Front margin of the head with two setae, which are 1.25–1.96 × the BW of ANT III. There are 5–6 pairs of dorsal setae and 11–13 ventral setae. There are two to four wax glands. Non-sclerotic region at the insertion of the antennae. Rostrum reaching past the front coxae, with URS 0.55–0.69 × HT II and bearing two accessory setae. Antennal tubercles undeveloped. Antennae are six-segmented (Figure 5). ANT 0.24–0.32 × BL. ANT III and V subequal in length; ANT VI longest. Short PT, 0.09–0.16 × BASE with four apical setae. Sensoria irregular in shape and numbers. ANT IV-V-VI each with one large sensorium occupying most of the length of the segment, enclosing small, scattered islands of cuticle and sometimes accompanied by a smaller round or oval secondary sensorium. Secondary sensoria (between one and four) of ANT III with more shape variation, either as small or large transverse bands, with or without sunken grooves, round or oval. They can also vary between the two antennae of the same specimen.
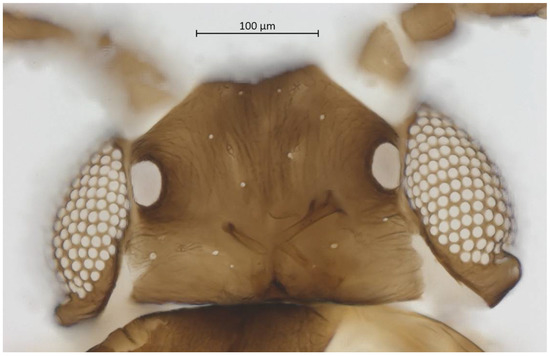
Figure 3.
Dorsal head of alate vivipara holotype of Qiao jinshaensis sp. nov. (IOZ(E)54809).
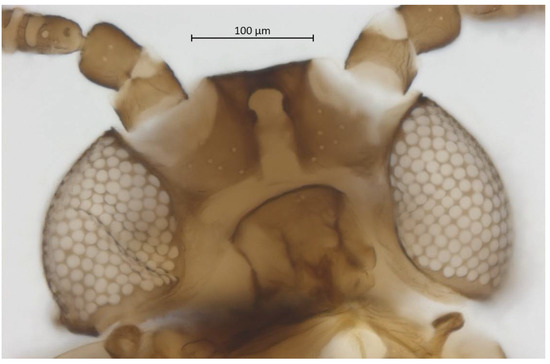
Figure 4.
Ventral head of alate vivipara holotype of Qiao jinshaensis sp. nov. (IOZ(E)54809).
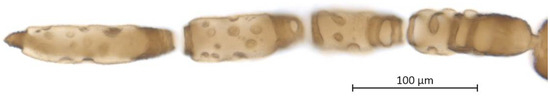
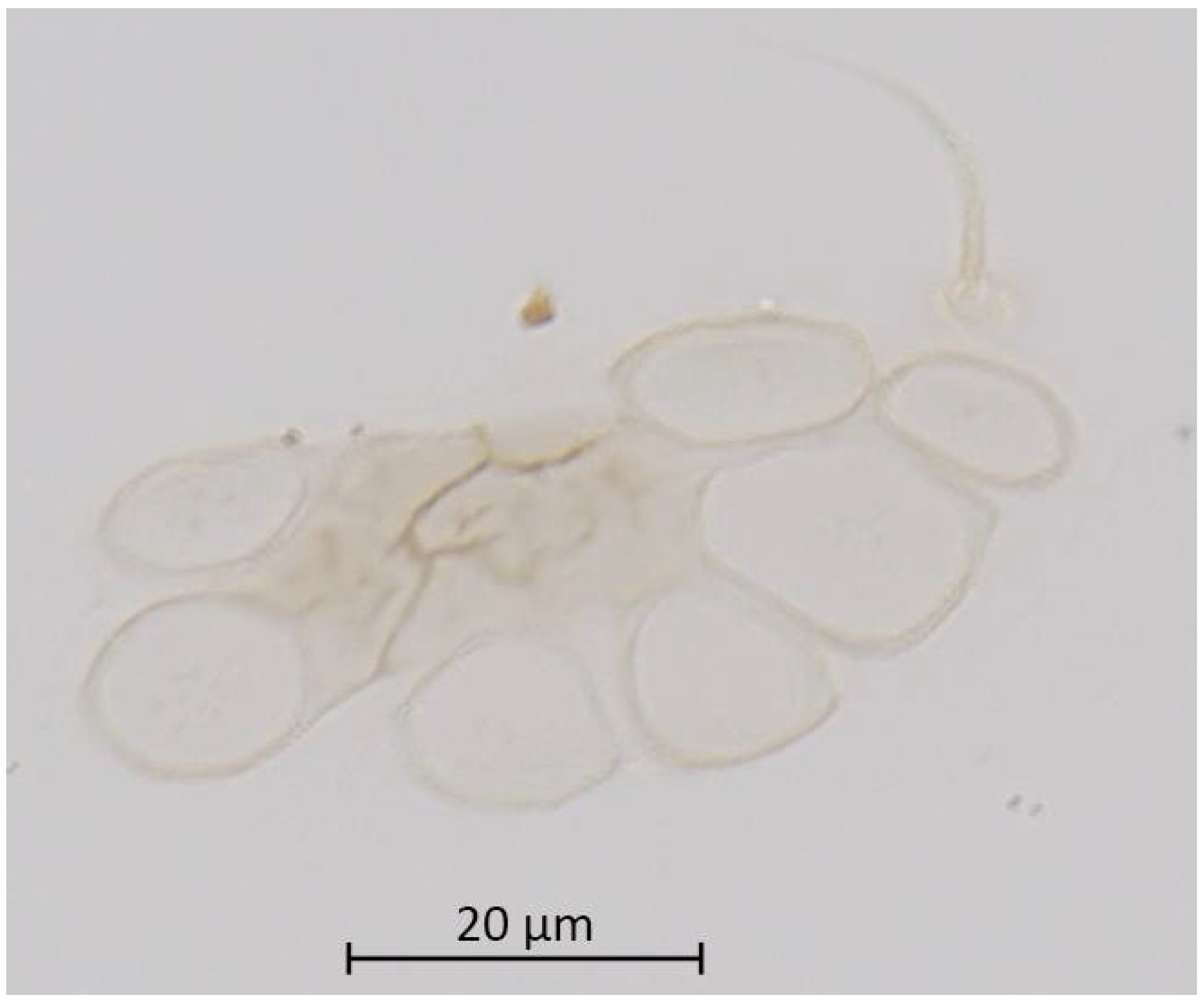
Figure 5.
Antenna (ANT III-VI) and sensoria of Qiao jinshaensis sp. nov. (QMOR66201).
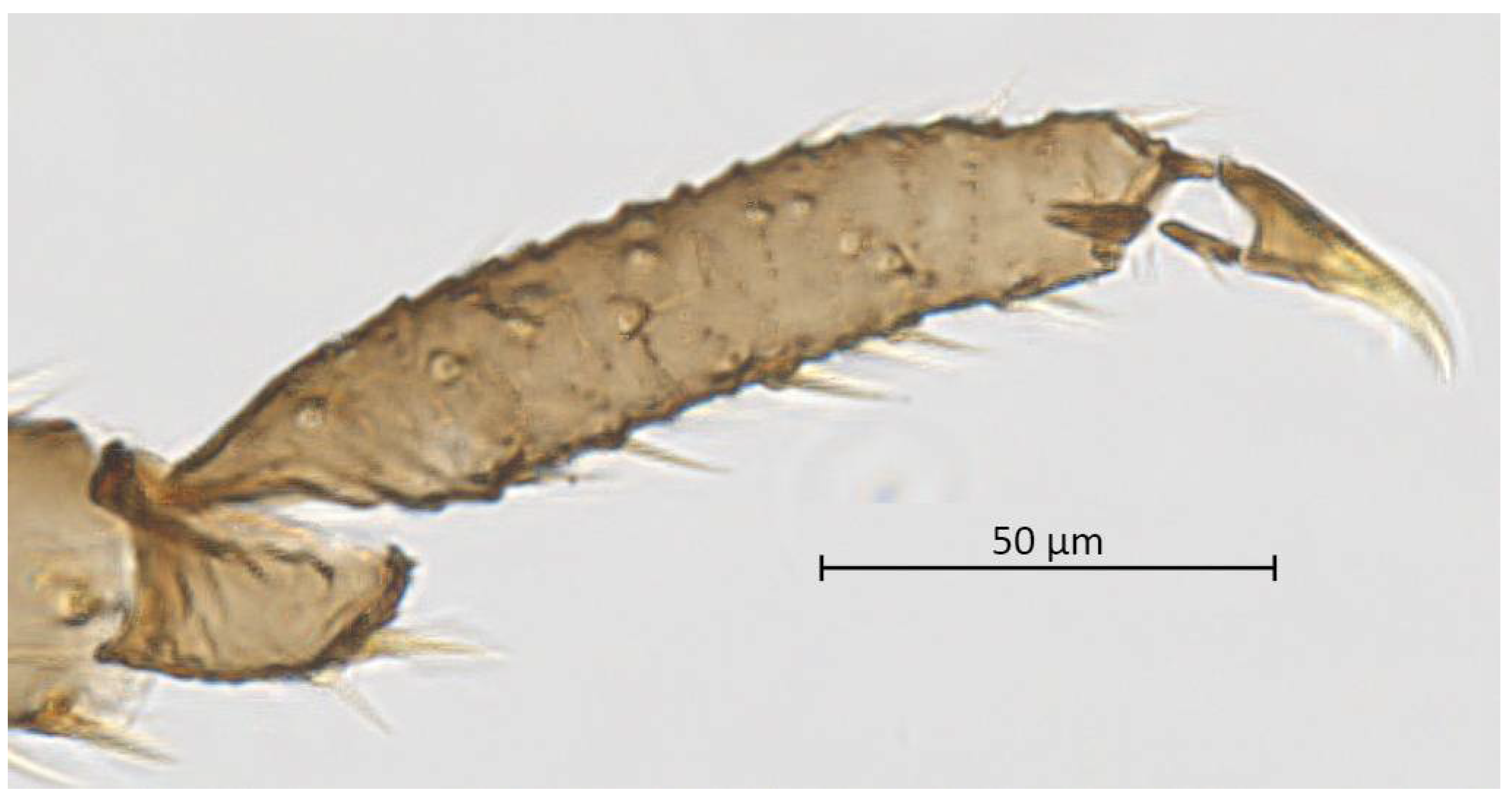
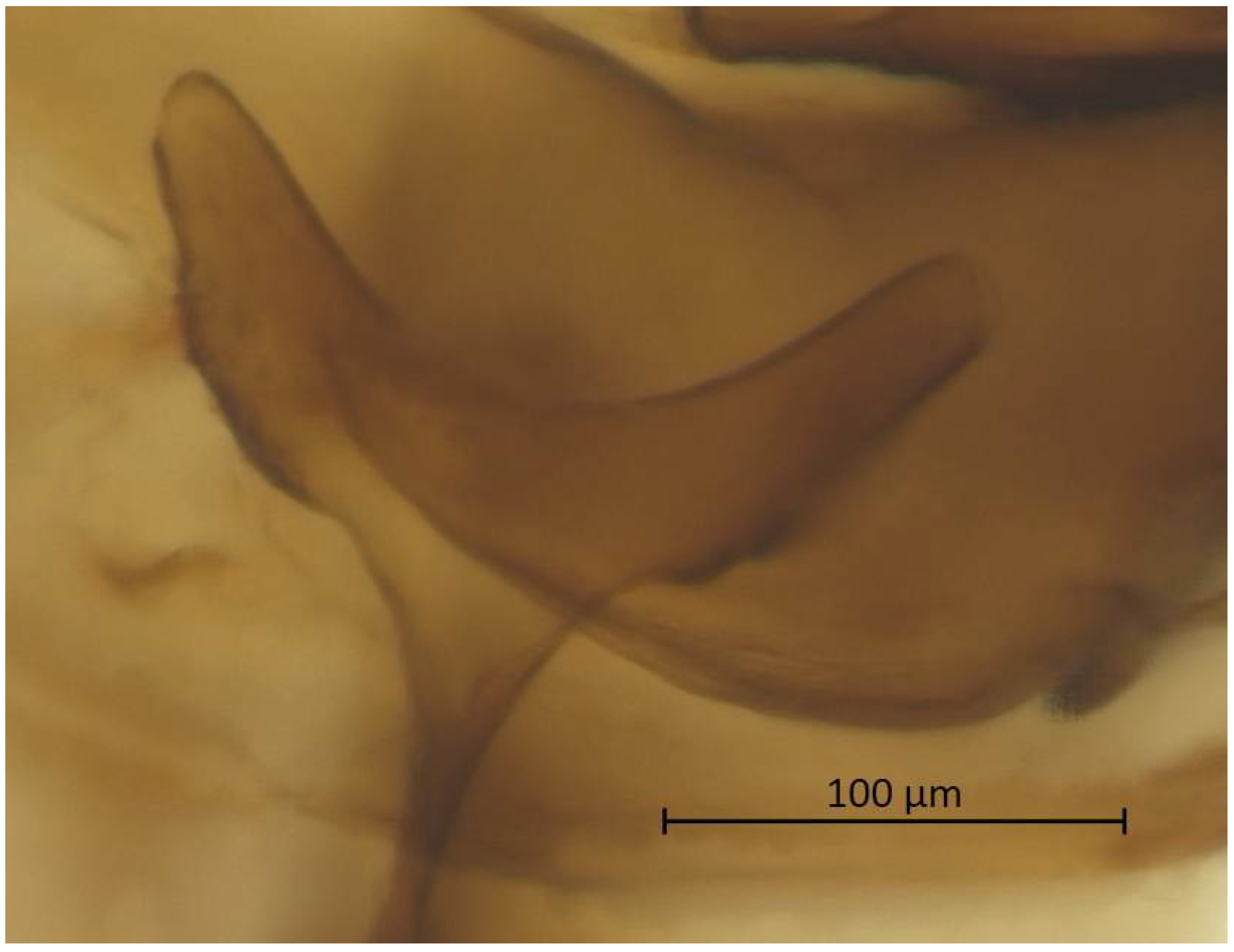
Thorax. Wax glands present. Mesosternal furca ‘Y’-shaped with a well-developed stem (Figure 6). Trochanter and femur fused (Figure 7). LMF 3.86–4.69 × ANT III. Legs slender with short and pointed setae. LMT 0.28-0.37 × BL and LMTS 0.47–0.60 × WMT. HT I triangular, 0.25–0.36 × HT II (Figure 8). First tarsal chaetotaxy: 3-4-4, with distal margin forming small spines. Forewing with two cubital veins well separated at the base, media simple with basal half obsolete, stigma short and oblique, veins fading at their apical extremity (Figure 2). Hindwing with 3 (rarely 2) hamuli on the costal margin and two oblique veins, the outer one shorter and strongly curved inward.
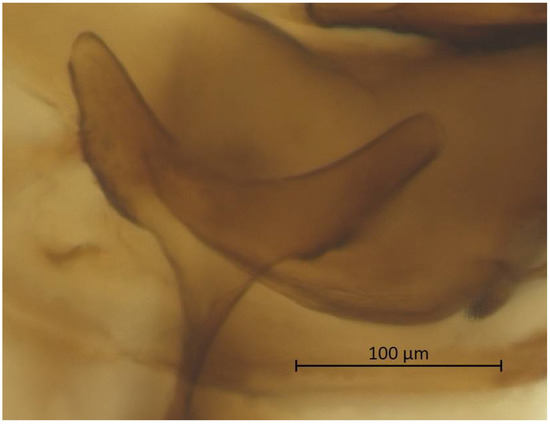

Figure 6.
Mesosternal furca of alate vivipara of Qiao jinshaensis sp. nov. (QMOR66202).

Figure 7.
Hind leg of alate vivipara of Qiao jinshaensis sp. nov. (QMOR66209).

Figure 8.
Hind tarsus of alate vivipara of Qiao jinshaensis sp. nov. (QMOR66209).
Abdomen. Marginal and spinal wax glands (Figure 9). Cauda semicircular, length 0.35–0.57 × its BW, bearing three to four setae. Genital plate bearing 24–33 setae, mostly distributed on the apical margin. Spiracles inconspicuous. Siphunculi absent. Measurements presented in Table 3.
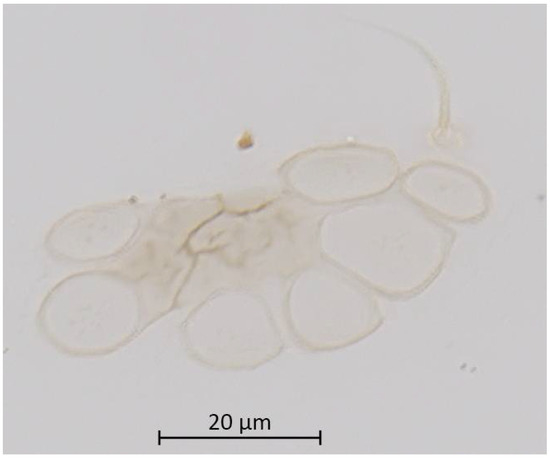
Figure 9.
Dorsal abdominal wax gland and single seta of alate vivipara of Qiao jinshaensis sp. nov. (IOZ(E)54810).

Table 3.
Measurements (in μm) of alate viviparae (n = 14) of Qiao jinshaensis sp. nov.
3.2.4. Etymology
The genus Qiao is named for QIAO Gexia, eminent Chinese aphidologist at the Institute of Zoology of the Chinese Academy of Sciences. The name is feminine in grammatical gender, thus the adjective jinshaensis is also feminine; the masculine and neuter forms are jinshaensis and jinshaense, respectively. The specific epithet is derived from the aphid’s type locality in Yongshan County, along the Jinsha River, tributary of the Yangtze River. The aphid’s host plant, Rhus wilsonii, is mainly distributed at a few narrow areas along the Jinsha River.
3.2.5. Biology
The galls are yellow-green or reddish in color and inflorescence-like in shape (Figure 10). They possess multiple cavities and are positioned on the apical part of the stem. The gall measures approximately 110 by 90 mm and resembles the one formed by Nurudea shiraii (Matsumura, 1917). The autumn migrants of the aphid appear in late September. The tannin content of the gall was 52.3%, higher than the 35.2% of galls formed by N. shiraii [23].
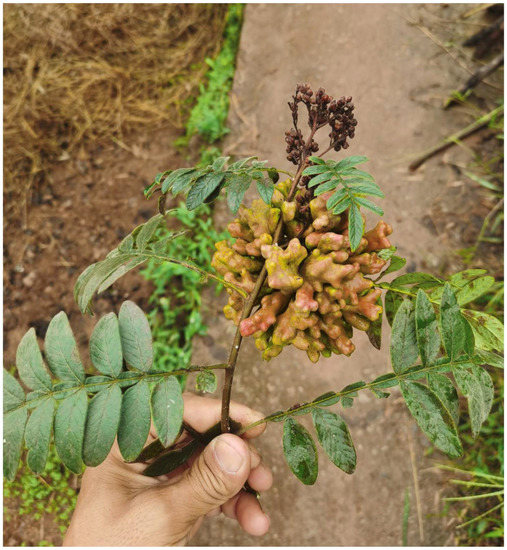

Figure 10.
Galls formed by Qiao jinshaensis sp. nov. on Rhus wilsonii.
3.2.6. Identification Key to Aphid Species on Rhus spp.
The following couplets should be inserted into the key to aphid species feeding on Rhus spp. as currently presented at Aphids on the World’s Plants [2].
- 6
- ANT III-VI each with a single large oblong rhinarium occupying most of its surface area ……Kaburagia rhusicola
- -
- All or most of ANT III-VI with more than one single rhinarium ……6A
- 6A
- Rhinaria on ANT III-VI varying in size, shape and number (some round or in transverse bands, others large sheets enclosing small, scattered islands of cuticle) ……Qiao jinshaensis sp. nov.
- -
- ANT III-VI with only separate, transversely elongate rhinaria ……7
4. Conclusions
Based on gall morphology alone, Qiao jinshaensis gen. et sp. nov. is most similar to species of Nurudea (see Figure 3 in Ren et al., 2018 [24]). However, it is quite distinct genetically and morphologically, suggesting the genus diverged early in the diversification of the Rhus-galling aphids. Despite this apparent divergence, these aphids, sometimes collectively considered the subtribe Melaphidina [5], are not especially speciose (there are now 17 species in 7 genera, including some species without a known host [25]). The genus Rhus dates to approximately 50 mya and is closely related to Pistacia [26], which is also known to harbor several genetically distinct but small Fordini genera. Price (2005) suggested that there may have been an early diversification among the Pistacia-galling aphids, now evident at the generic level, but with reduced recent speciation [27]. This hypothesis is also applicable to the Rhus-galling aphids. So far, most studies have focused on Rhus- or Pistacia-feeding groups separately (e.g., [7,28]), but given the evolutionary proximity of both the aphids and their hosts, a phylogenetic evaluation of the entire tribe is needed.
Author Contributions
Conceptualization, Z.Y. and C.F.; methodology, C.H., X.X. and Z.Y.; validation, C.H., Z.Y. and C.F.; resources, Z.Y. and C.F.; data curation, C.H. and C.F.; writing–original draft preparation, C.H. and C.F.; writing–review and editing, C.H., Z.Y. and C.F.; funding acquisition, Z.Y. and C.F. All authors have read and agreed to the published version of the manuscript.
Funding
The research and APC were funded by the National Natural Science Foundation of China (Grant/Award Number 31872305 to Yang) and the Natural Sciences and Engineering Research Council of Canada (Discovery Grant RGPIN 2017-06287 to Favret and Undergraduate Student Research Award to Hébert).
Data Availability Statement
DNA sequence data are available at GenBank; specimen collection data are available from Hébert et al. [22]; specimens are available at the Institute of Zoology of the Chinese Academy of Sciences and the Ouellet-Robert Entomological Collection of the University of Montreal.
Acknowledgments
The authors thank Anton Bystrov, Malek Kalboussi, and Achille Villeneuve (University of Montreal) for help in the molecular lab, and two anonymous reviewers for their comments on an earlier version of the manuscript.
Conflicts of Interest
The authors declare no conflict of interest. The funders had no role in the design of the study; nor in the collection, analyses, and interpretation of data; nor in the writing of the manuscript; nor in the decision to publish the results.
References
- Zhang, C.-X.; Tang, X.-D.; Cheng, J.-A. The Utilization and Industrialization of Insect Resources in China. Entomol. Res. 2008, 38, S38–S47. [Google Scholar] [CrossRef]
- Blackman, R.L.; Eastop, V.F. Aphids on the World’s Plants . Available online: http://www.aphidsonworldsplants.info/ (accessed on 22 November 2022).
- Holman, J. Host Plant Catalog of Aphids, Palearctic Region; Springer Science: Dordrecht, The Netherlands, 2009; p. 1216. [Google Scholar]
- Yang, Z.; Chen, X.; Havill, N.P.; Feng, Y.; Chen, H. Phylogeny of Rhus Gall Aphids (Hemiptera: Pemphigidae) Based on Combined Molecular Analysis of Nuclear EF1α and Mitochondrial COII Genes. Entomol. Sci. 2010, 13, 351–357. [Google Scholar] [CrossRef]
- Ren, Z.; Zhong, Y.; Kurosu, U.; Aoki, S.; Ma, E.; von Dohlen, C.D.; Wen, J. Historical Biogeography of Eastern Asian–Eastern North American Disjunct Melaphidina Aphids (Hemiptera: Aphididae: Eriosomatinae) on Rhus Hosts (Anacardiaceae). Mol. Phylogenet. Evol. 2013, 69, 1146–1158. [Google Scholar] [CrossRef] [PubMed]
- Ren, Z.; Harris, A.J.; Dikow, R.B.; Ma, E.; Zhong, Y.; Wen, J. Another Look at the Phylogenetic Relationships and Intercontinental Biogeography of Eastern Asian—North American Rhus Gall Aphids (Hemiptera: Aphididae: Eriosomatinae): Evidence from Mitogenome Sequences via Genome Skimming. Mol. Phylogenet. Evol. 2017, 117, 102–110. [Google Scholar] [CrossRef] [PubMed]
- Ren, Z.; von Dohlen, C.D.; Harris, A.J.; Dikow, R.B.; Su, X.; Wen, J. Congruent Phylogenetic Relationships of Melaphidina Aphids (Aphididae: Eriosomatinae: Fordini) According to Nuclear and Mitochondrial DNA Data with Taxonomic Implications on Generic Limits. PLoS ONE 2019, 14, e0213181. [Google Scholar] [CrossRef]
- Hébert, C.; Sano, M.; Favret, C. Morphological and Molecular Diagnosis and Specimen Data Publication of a New Species of Cinara (Hemiptera: Aphididae) from Japan. Entomol. Sci. 2022, submitted.
- von Dohlen, C.D.; Rowe, C.A.; Heie, O.E. A Test of Morphological Hypotheses for Tribal and Subtribal Relationships of Aphidinae (Insecta: Hemiptera: Aphididae) Using DNA Sequences. Mol. Phylogenet. Evol. 2006, 38, 316–329. [Google Scholar] [CrossRef]
- Folmer, O.; Black, M.; Wr, H.; Lutz, R.; Vrijenhoek, R. DNA Primers for Amplification of Mitochondrial Cytochrome C Oxidase Subunit I from Diverse Metazoan Invertebrates. Mol. Mar. Biol. Biotechnol. 1994, 3, 294–299. [Google Scholar]
- Théry, T.; Kanturski, M.; Favret, C. Molecular Phylogenetic Analysis and Species Delimitation in the Pine Needle-Feeding Aphid Genus Essigella (Hemiptera, Sternorrhyncha, Aphididae). Insect Syst. Divers. 2018, 2, 15. [Google Scholar] [CrossRef]
- Sievers, F.; Barton, G.J.; Higgins, D.G. Multiple Sequence Alignment. In Bioinformatics, 4th ed.; Baxevanis, A.D., Bader, G.D., Wishart, D.S., Eds.; Wiley: Hoboken, NJ, USA, 2020; pp. 227–250. [Google Scholar]
- Lv, X.; Yang, Z.-X.; Li, Y.; Liu, P. Comparison of Determination Methods in Tannin Content of Chinese Gallnuts. Southwest China J. Agric. Sci. 2012, 25, 1080–1084. [Google Scholar]
- Huang, H.; Lv, Y.; Sun, X.; Fu, S.; Lou, X.; Liu, Z. Rapid Determination of Tannin in Danshen and Guanxinning Injections Using UV Spectrophotometry for Quality Control. J. Innov. Opt. Health Sci. 2018, 11, 1850034. [Google Scholar] [CrossRef]
- Favret, C. A New Non-Destructive DNA Extraction and Specimen Clearing Technique for Aphids (Hemiptera). Proc. Entomol. Soc. Wash. 2005, 107, 469–470. [Google Scholar]
- Tsai, P.; Tang, C. The Classification of the Chinese Gall Aphids with Descriptions of Three New Genera and Six New Species from Meitan, Kweichow. Trans. R. Entomol. Soc. Lond. 1946, 97, 405–418. [Google Scholar] [CrossRef]
- Takagi, G. Studies in the Artificial Multiplication of the Sumach Gall-Aphid I Especially Schlechtendalia Chinensis Bell. Bull. For. Exp. Stn. 1937, 26, 20–26. [Google Scholar]
- Matsumura, S. Synopsis of the Pemphigidae of Japan. In A Collection of Essays for Mr. Yasushi Nawa, Written in Commemoration of his Sixtieth Birthday, October 8, 1917; Nagano, K., Ed.; Gifu, Japan, 1917; pp. 39–94. [Google Scholar]
- Xiang, H. Studies of Chinese gall-nut aphids on Rhus potaninii Maxim. Entomotaxonomia 1980, 2, 303–313. [Google Scholar]
- Foottit, R.G.; Maw, H.E.L.; Von Dohlen, C.D.; Hebert, P.D.N. Species Identification of Aphids (Insecta: Hemiptera: Aphididae) through DNA Barcodes. Mol. Ecol. Resour. 2008, 8, 1189–1201. [Google Scholar] [CrossRef]
- International Commission on Zoological Nomenclature (ICZN). International Code of Zoological Nomenclature, 4th ed.International Trust for Zoological Nomenclature: London, UK, 1999. [Google Scholar]
- Hébert, C.; Xu, X.; Yang, Z.; Favret, C. Qiao jinshaensis. (Insecta: Hemiptera: Aphididae), a gall-inducing aphid on Rhus wilsonii Hemsl. (Anacardiacaeae) from Yunnan, China. Collection 2022, 1. Available online: https://specimenpub.org/collection/collection_1/ (accessed on 22 November 2022).
- Chang, X.; Cai, D. Experimental Research on the Preparation of Industrial Tannic Acid by a Chinese Gall Beihua. Guangxi For. Sci. Technol. 1993, 22, 64–67. [Google Scholar]
- Ren, Z.; Su, X.; von Dohlen, C.D.; Wen, J. Nurudea zhengii Ren, a New Species of the Rhus Gall Aphids (Aphididae: Eriosomatinae: Fordini) from Eastern China. Pakistan J. Zool. 2018, 50, 2087–2092. [Google Scholar] [CrossRef]
- Favret, C. Aphid Species File. Available online: http://Aphid.SpeciesFile.org (accessed on 22 November 2022).
- Yi, T.; Miller, A.J.; Wen, J. Phylogenetic and Biogeographic Diversification of Rhus (Anacardiaceae) in the Northern Hemisphere. Mol. Phylogenet. Evol. 2004, 33, 861–879. [Google Scholar] [CrossRef]
- Price, P.W. Adaptive radiation of gall-inducing insects. Basic Appl. Ecol. 2005, 6, 413–421. [Google Scholar] [CrossRef]
- Inbar, M.; Wink, M.; Wool, D. The Evolution of Host Plant Manipulation by Insects: Molecular and Ecological Evidence From Gall-Forming Aphids on Pistacia. Mol. Phylogenet. Evol. 2004, 32, 504–511. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).