1. Introduction
Again since the one face, constant in symmetry, appears sometimes fair and sometimes not, can we doubt that beauty is something more than symmetry, that symmetry itself owes its beauty to a remoter principle?
[
1] (Ennead I, Sixt Tractate, p66).
The remote principle envisaged by Plotinus is still a symmetry principle but in a modern definition involving group theory and algebraic geometry. Recently, we wrote a paper about a common algebra possibly ruling the beauty and structure in poems, music and proteins [
2]. We found that free groups govern the structure of such disparate topics where a language emerges from pure randomness. We coined the concept of ‘syntactical freedom’ for qualifying the occurrence of symbols and rules organized according to group theory and aperiodicity [
3,
4]. One favorite decomposition of the secondary structure of proteins is in term of
-helices,
-sheets and coils [
5] but this decomposition and the resulting syntax is model dependent [
2]. According to our view, at the genome scale, the escape to ‘syntactical freedom’ means a lack of harmony and the signature of a potential disease. The secondary structures in the sequence of proteins or viruses are, most of the time, organized according to the rules of free groups and, otherwise they may be a witness of a potential aberrant topology.
Of course, there are other techniques to identify a potential disease in the DNA/RNA structures. The loose of homochirality of DNA may indicate an age-related disease [
6]. Data mining techniques may be employed to identify cancer in gene expression [
7]. The role of miRNAs in the regulation of pathogenesis during infection is investigated in [
8].
Apart from the canonical double helix B-DNA we now know that there exists a diversity of non-canonical coding/decoding sequences organized in structures such as Z-DNA (often encountered in transcription factors [
9,
10]), G-quadruplex (in telomeres) and other types that are single-stranded, two-stranded, or multistranded [
11]. RNA is usually a single-stranded molecule in a short chain of nucleotides, as is the case for a messenger RNA (mRNA) or a (non-coding) microRNA (miRNA).
In our approach, the investigated sequences define a finitely generated group
whose structure of subgroups is close or away from a free group
of rank
r, where
is the number of distinct amino acids in the sequence (or the number of distinct secondary structures considered in the protein chain) [
4]. Recently, we also introduced concepts for representing the groups
over the Lie group
. The
character variety of
and its Groebner basis are topological ingredients, they feature algebraic geometric properties of the group
under question [
12,
13,
14]. A reminder of the theory is given in the next section.
In this paper, we focus upon transcription factors and miRNAs, both serve at properly decoding and regulating the genes and their action, either independently of each other or together by targeting common genes [
15].
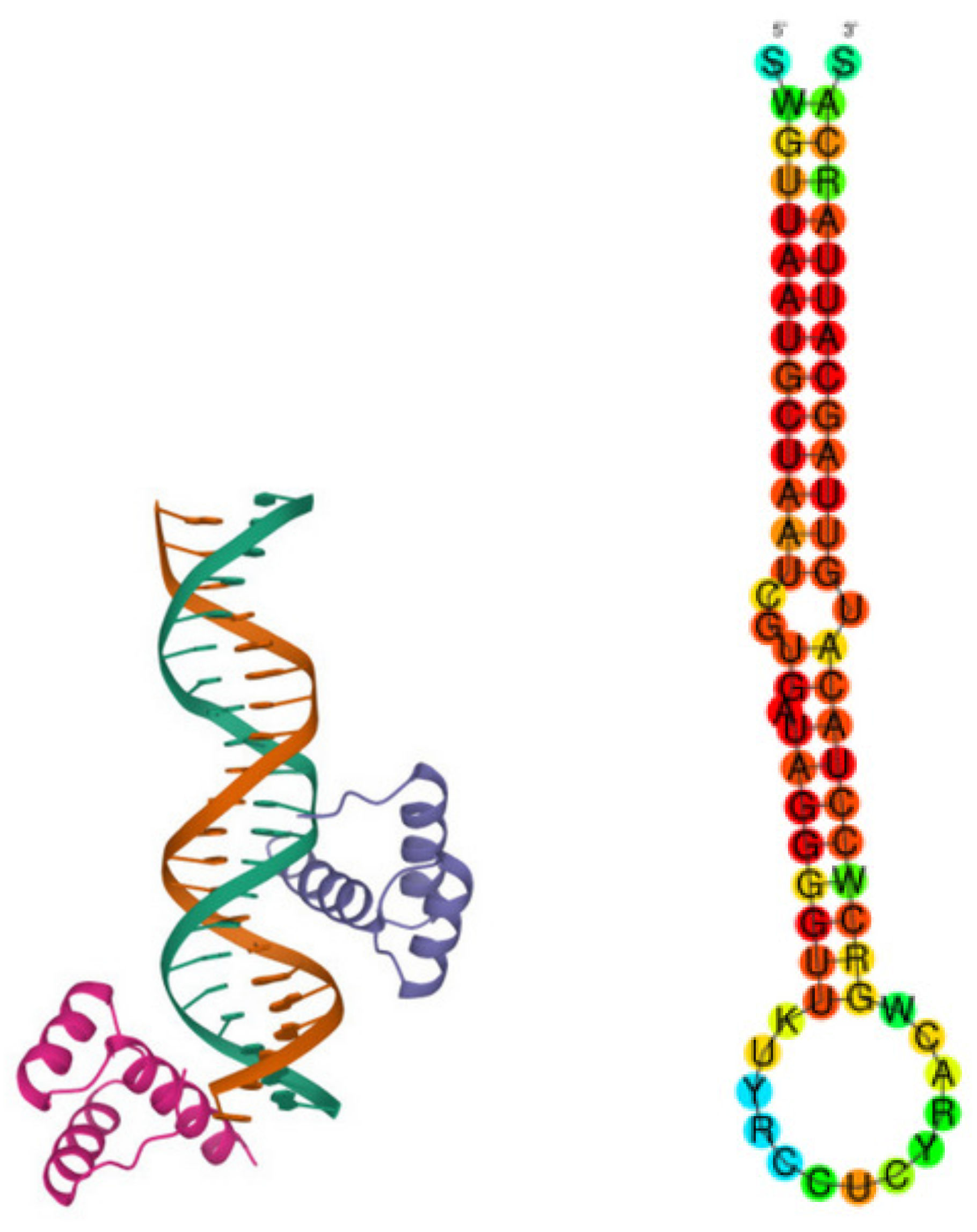
Figure 1 (Left) is a picture of the pluripotent transcription factor Nanog.
Figure 1 (Right) is an example of a pre-miRNA associated to a disease [
16]. Both are investigated in detail in this paper.
In
Section 2, we describe the mathematical methods and the software needed for describing the algebraic surfaces relevant to DNA/RNA sequences. This includes the definition of infinite groups under question, of the free groups
of rank
to 3 corresponding to 2- to 4-base sequences and the calculation of
representation of such groups. A special care is needed to compute a Groebner basis of the character variety.
Section 3 is a discussion about the type of singular surfaces encountered in this research. They play an important role in our view of the discrimination of a potential disease.
In
Section 4, the methods are applied to representative examples of sequences taken from transcription factors and microRNAs whose group is close or away from a free group, and whose Groebner basis of the variety contains simple singularities. Our theory applies to many TFs and miRNAs that are known to be related to an identified disease.
Section 5 summarizes our paper and opens a few perspectives.
2. Theory
2.1. Finitely Generated Groups, Free Groups and Their Conjugacy Classes
Let be the free group on r generators (r is called the rank).
The number of conjugacy classes of
of a given index
d is known and is a signature of the isomorphism, or the closeness, of a group
to
[
4,
17]. The cardinality structure of conjugacy classes of index
d in
will be called the card seq of
. We need the cases from
r = 1 to 3 that correspond to the number of distinct bases in a DNA/RNA sequence. The card seq of
is in
Table 1 for the 3 sequences of interest in the context of DNA/RNA.
The free group of rank 1 may be defined as (with one generator a and no relation) or (with two generators a and b and the relation ). Similarly, the free group of rank 2 is and so on for higher rank r.
Next, given a finitely generated group with a relation (rel) given by a sequence motif, we are interested in the number of conjugacy classes of subgroups of index d (the card seq of ). Often, for a selected DNA/RNA motif taken as the generator of a finitely generated group , the card seq that is obtained is close to that of a free group , with being the number of distinct bases involved in the motif.
The closeness of to can be checked by its signature in the finite range of indices of the card seq.
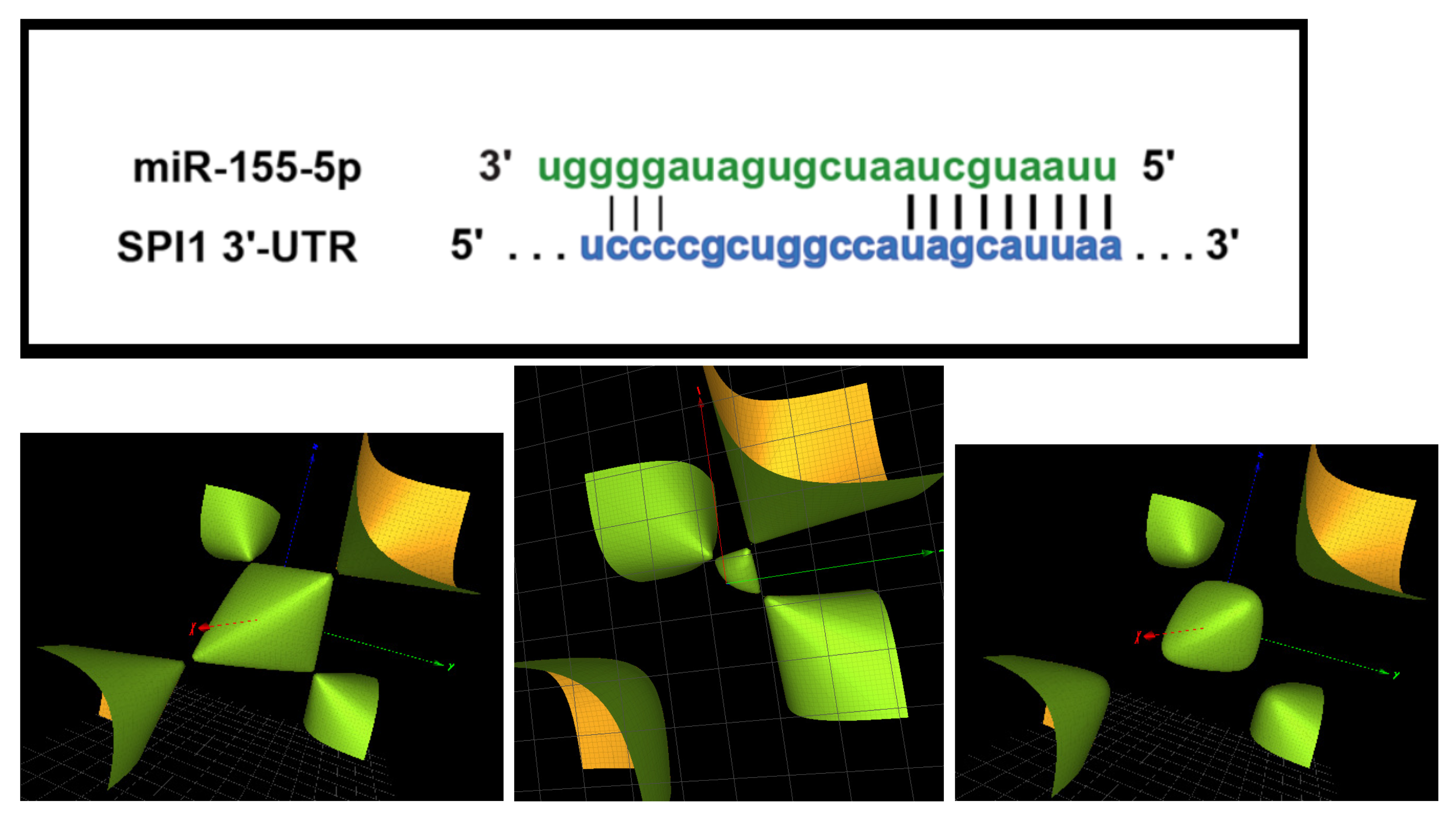
To illustrate our methodology, let us consider the seed UCCUACA of the microRNA sequence mir-155-3p that is investigated in detail in
Section 4.3; see also
Figure 2. The finitely generated group is
. The card seq of
is found to be the series
. The group
is of rank 2 and the card seq corresponds to that of the group we call
in the tables of
Section 4. In this case, the card seq of
is ‘away’ from the card seq of the free group of the same rank
and the two groups
and
are of course ‘away’ to each other. However, if we omit the last nucleotide A of the seed in the generator of
, then the group
obtains the same card seq than
(at least in the finite range of the card seq series that we can check) and, in this sense, both groups
and
become ‘close’ to each other.
2.2. The Character Variety of a Finitely Generated Group and a Groebner Basis
Let be a finitely generated group, we describe the representations of in , the group of matrices with complex entries and determinant 1. The group may be seen simultaneously as a ‘space-time’ (a Lorentz group) and a ‘quantum’ (a spin) group.
Such a group describes representations as degrees of freedom for all quantum fields and is the gauge group for Einstein–Cartan theory. The later contains the Einstein–Hilbert action and Einstein’s field equations [
19]. The so-called Holst action used in loop quantum gravity has quantum gravity states given in terms of
representations [
20].
Representations of
in
are homomorphisms
with character
,
. The notation
means the trace of the matrix
. The set of characters allows to determine an algebraic set by taking the quotient of the set of representations
by the group
, that acts by conjugation on representations [
21,
22].
Such an algebraic set is called the
character variety of
. The set is made of a sequence of multivariate polynomials called a scheme
X. The ideal
is defined by the vanishing of polynomials living in the scheme
X. Below, we use a particular basis
of the polynomial ring
, called a Groebner basis. The Groebner basis
has to follow algorithmic rules (similar to the Euclidean division for univariate polynomials) [
14].
For the effective calculations of the character variety, we make use of a software on Sage [
23]. We also need Magma [
24] for the calculation of a Groebner basis, at least for 3- and 4-base sequences.
2.3. Algebraic Geometry and Topology of DNA/RNA Sequences
2.3.1. Two-Base Sequences
Following [
25], in this section, we are interested by the special case of representations for the once punctured torus
and the relevance of the extended mapping class group
in its action on surfaces of type
,
.
The fundamental group of
, that we denote
, is the free group
on two generators
a and
b. The boundary component of
consists of a single loop around the puncture expressed by the commutator
with
and
. Taking the traces
,
,
, the trace of the commutator is the surface [
21,
25]
According to the Dehn–Nielsen–Baer theorem [
26], for a surface of genus
, we have
where
is the mapping class group. It denotes the group of isotopy classes of orientation-preserving diffeomorphisms of
S.
The extended mapping class group
is the group of isotopy classes of all homeomorphisms of
S (including the orientation-reversing ones). The outer automorphism group of
is denoted
. This leads to the (topological) action of
on the once punctured torus as follows
The automorphism group
acts by composition on the representations
and induces an action of the extended mapping class group
on the character variety by polynomial diffeomorphisms of the surface
defined by [
25]
Within the family
, the Cayley surface
is obtained from the character variety for the fundamental group of the Hopf link
(the link of two unknotted curves). For the Hopf link, the fundamental group is
where the notation
is for the 3-sphere and
means that a small tubular neighborhood around the Hopf link
is removed from
to define the corresponding Hopf link 3-manifold.
The Cayley surface
possesses 4 simple (isolated) singularities. At these points, the surface loses its smoothness. We already showed that it plays a role in the context of Z-DNA conformations of transcription factors ([
13], Tables 2 and 5); see also,
Section 4 below, and notably
Figure 3 (Left).
The surface
lies within the character variety for the fundamental group of the link L6a1 [
27]. We show below that this surface also lies in the generic Groebner basis obtained for 4-base sequences; see
Section 4 below.
2.3.2. Three-Base Sequences
Our main object in this section is the four punctured sphere for which the fundamental group is the free group
of rank 3 whose character variety generalizes the Fricke cubic surface (
2) to the surface
in
. From now,
a and
b are no longer generators of a group but belong to the set of parameters of the surface
V.
We follow the work of references [
21,
25,
28].
Let be the quadruply punctured sphere. The fundamental group for can be expressed in terms of the boundary components , , , as .
A representation
is a quadruple
Let us associate the seven traces
where
a,
b,
c,
d are boundary traces and
x,
y and
z are traces of elements
,
and
representing simple loops on
.
The character variety for
satisfies the equation ([
21], Section 5.2), ([
25], Section 2.1), ([
28], Section 3B), ([
29], Equation (1.9)) or ([
30], Equation (39)).
with
,
,
and
The 4-punctured sphere, whose fundamental group is the free group
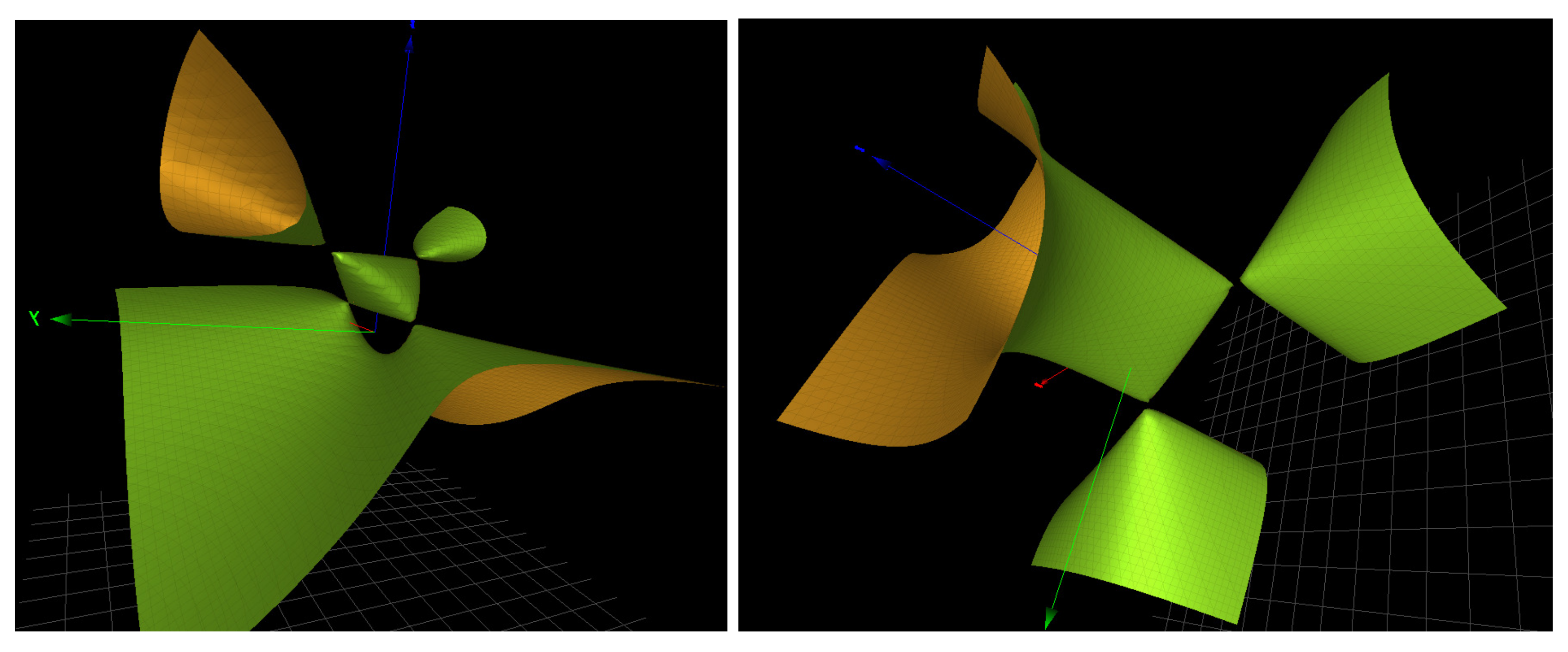
with generator the product of the 4 letters, is a generic topology. It is straightforward to check that the Groebner basis for
contains (among other surfaces and depending on the choice of parameters) a single copy of the generic surfaces
,
and
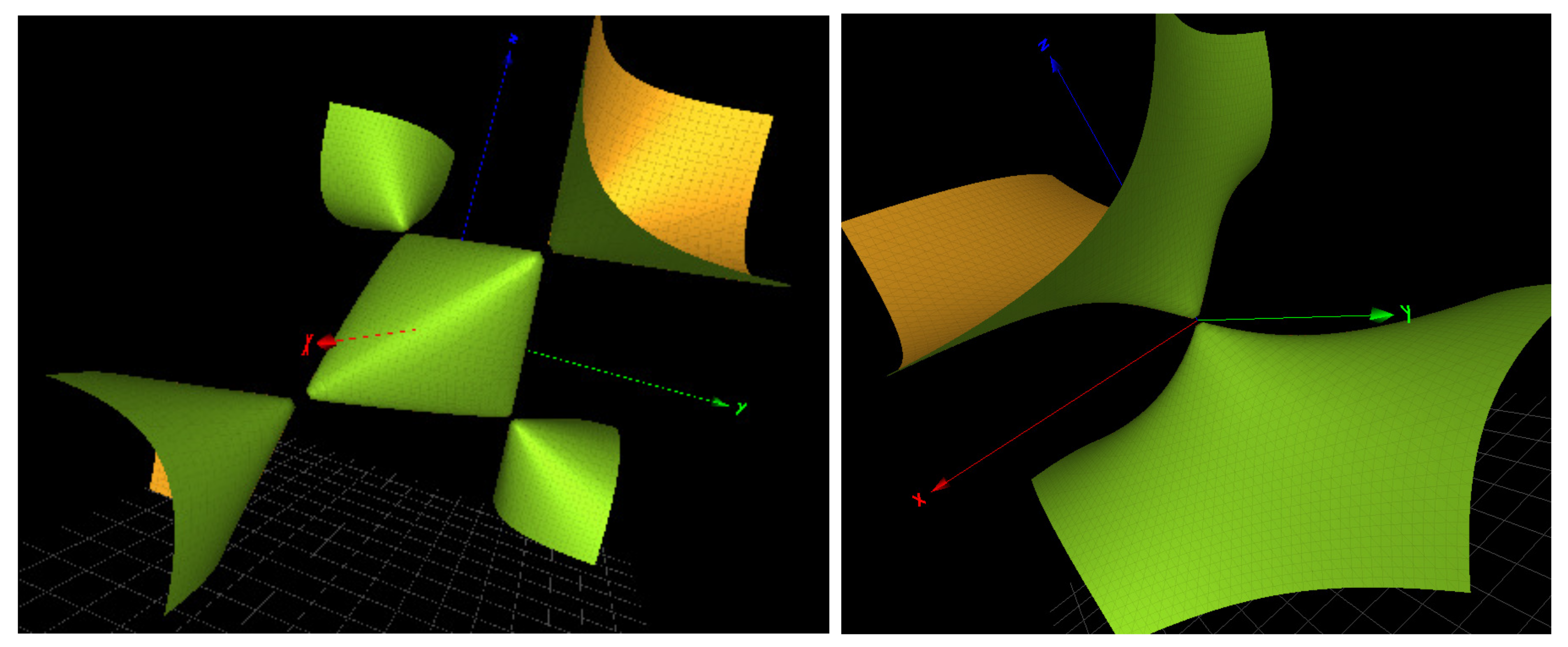
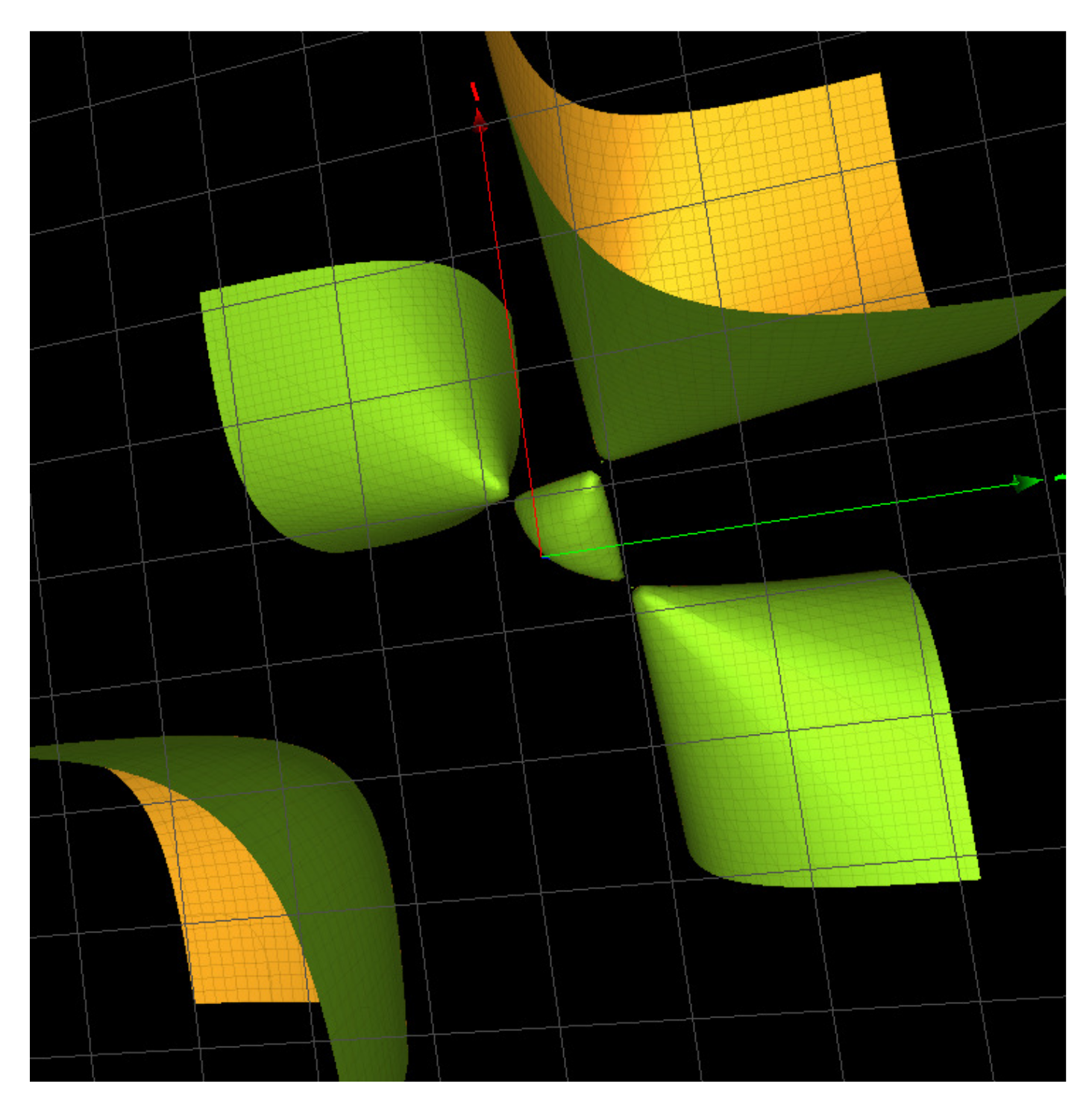
, a surface we also denote
because it contains 3 simple singularities of type
as shown in
Figure 4 and
Figure 5.
There are other surfaces encountered in our study of the Groebner basis for transcription factors and miRNAs when the generated group is close or away from the free group
(for 3-base sequences) or the free group
(for 4 base sequences). These surfaces are described in
Section 4.
2.3.3. Four-Base Sequences
There does not exist a huge difference in the structure of a Groebner basis of the character variety in the case of a 4-base sequence compared to the case of a 3-basis sequence. One difference is that one has to manage a 14-dimensional hypersurface in (instead of a 7-dimensional one as in the previous subsection). In general, after the appropriate choice of the 8 parameters , the Groebner basis contains more than one copy of the generic Groebner basis. Each copy S of a relevant surface may be of the form , , or .
3. Discussion
Given an ordinary projective surface
S in the projective space
over a number field, when
S is birationally equivalent to a rational surface, the software Magma [
24] determines the map to such a rational surface and returns its type within five categories. The returned type of
S is
for the projective plane, a quadric surface (for a degree 2 surface in
), a rational ruled surface, a conic bundle or a degree
p Del Pezzo surface where
.
One important attribute of a projective surface is its degree of singularity. Most surfaces
S of interest below are almost not singular in the sense that they have at worst simple (isolated) singularities. A simple singularity is referred to as an A-D-E singularity [
31]. It has to be of the type
,
,
,
,
,
or
.
The type and the number of simple singularities are denoted in an exponent such as for l singularities of type . All such surfaces are degree 3 del Pezzo surfaces. For the Cayley cubic , the exponent means .
There are additional facilities offered by Magma for studying a singular scheme. As already mentioned, a scheme is any geometric object defined by a set of polynomials in a projective space. An algebraic surface is a scheme. If the scheme has simple singularities, one can calculate the degree and the support of the reduced singular subscheme that are signatures of the scheme. In the examples of this paper, for a surface , the degree is l and the support contains l simple singular points. Otherwise, we add a lower index to to qualify the index and the support of the singular subscheme. The notation means that there are l singularities of type , that the degree of the singular subscheme is m and that the support is the empty set.
Most of the time, the surfaces in the character variety attached to a transcription factor or a microRNA only contain isolated singularities.
This contrasts with some sequences encountered in another context. For instance, a complete turn of A-DNA (PDB; 2D47) defines the dodecamer sequence CCCCCGCGGGGG whose attached character variety contains the surface ([
13], Figure 5).
This surface contains many non isolated singularities (that cannot be resolved by blow ups). Methods to desingularize a surface of such a type are presented in a companion paper [
32].
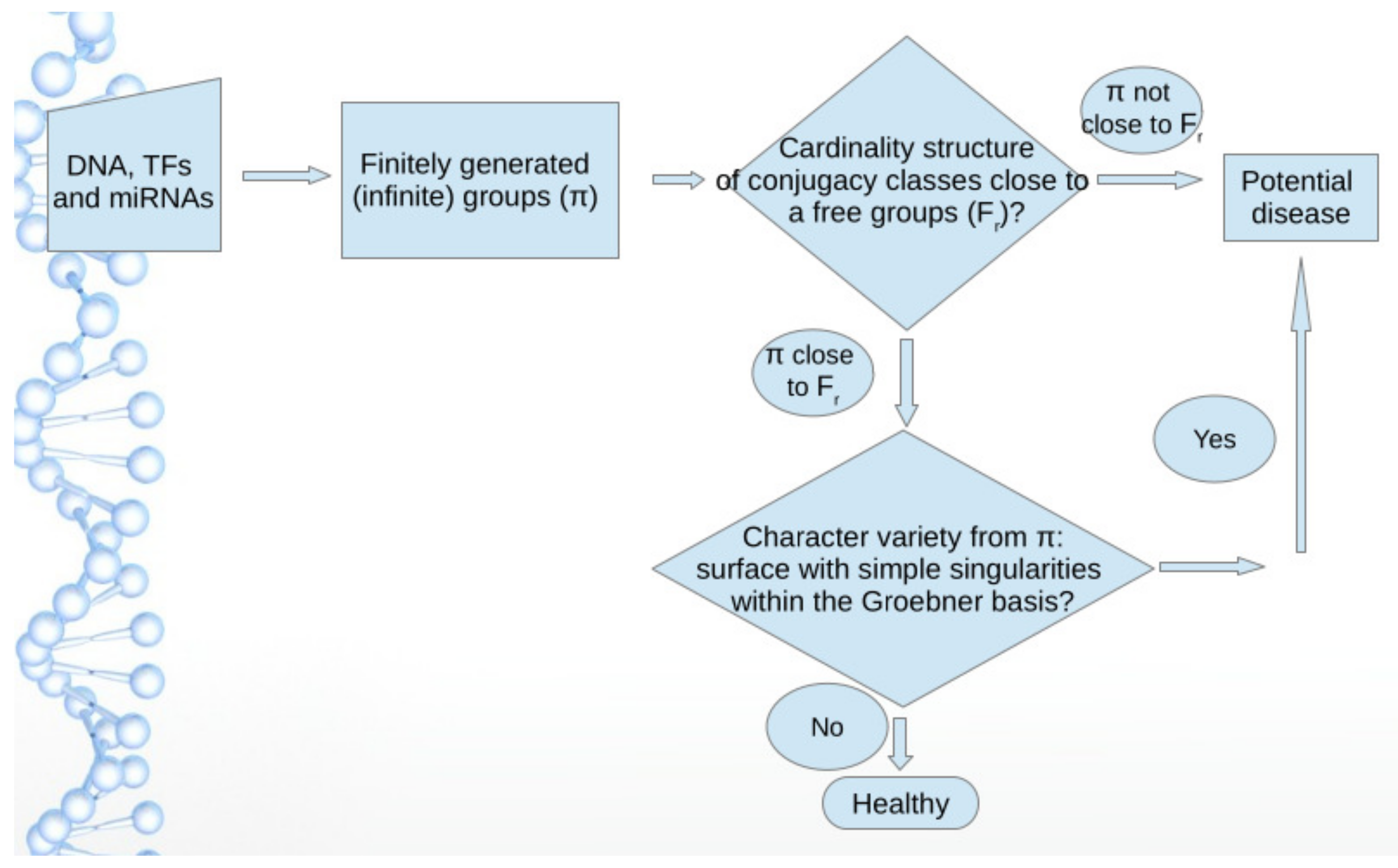
To summarize the important issues below, a noteworthy result of our approach is to recognize that optimal regulation occurs when the group underlying the sequence looks similar to a free group
(
to 3) in the cardinality sequence of its subgroups, a result obtained in our previous papers. A non-free group structure features a potential disease. A second noteworthy result is about the structure of the Groebner basis of the variety. A surface with simple singularities (such as the well known Cayley cubic) within the Groebner basis is a signature of a potential disease even when the generated group looks similar to a free group
in its structure of subgroups. Our methods apply to groups with a generating sequence made of two to four distinct DNA/RNA bases in
. Several human TFs and miRNAs are investigated in detail thanks to our approach. We summarize this discussion in the diagram presented in
Figure 6.
4. Results
In this section, we apply the
representation theory to groups generated by DNA/RNA sequences occurring in transcription factors and microRNAs. Both play a leading role in the decoding of the genome and in genome-scale regulatory networks. Two-letter transcription factors (TFs) whose structure is close or away from the free group
were already investigated in ([
13], Table 2). The occurrence of the Cayley cubic
in the Groebner basis of the character variety was found to be a signature in the former case. In this case, this surface seems to possess a regulatory action that may be lost in the latter case. In ([
13], Table 3), the potential diseases associated with a non-free group structure are mentioned. In the present section, one explores, in
Table 2, the case of a 3-letter sequence of a TF and in
Table 3, the case of 3-letter sequence of a miRNA. The case of a 4-letter sequence of a miRNA is summarized in
Table 4. The role of surfaces with simple singularities in the Groebner basis is emphasized.
4.1. Algebraic Morphology of the Transcription Factor Prdm1
The transcriptional repressor PR domain containing 1 (Prdm1), also known as B-lymphocyte-induced maturation protein-1 (Blimp1), is essential for normal development and immunity [
33]. It is of a zinc finger type. The consensus sequence ACTTTC corresponds to the code MA0508.2 in [
34].
4.1.1. The Character Variety
The ideal for the character variety
for a few values of the parameters is
where
,
are Fricke surfaces [
27] and
is the surface drawn in
Figure 4. The subscript
is for featuring the three singularities of type
.
4.1.2. The Groebner Basis
The singular surfaces found in the Groebner basis of the ideal are not similar to those in the ideal. One of them is obtained at values featuring two simple singularities of type and with a singular subscheme of degree 3 and the singular point of type in its support. The other surface obtained in the Groebner basis at values is a conic bundle of the type whose singular subscheme is non zero dimensional and of degree 1.
These two surfaces are non standard in our context of TFs and miRNAs. The formal desingularization of the surface
is given in [
32].
4.2. Algebraic Morphology of Homeodomains for Nanog and Xvent
The pluripotency in embryonic stem cells and their regulation is characterized by the expression of several transcription factors [
35,
36]. Among them, the transcription factor Nanog is present in the embryonic stages of life of several vertebrate species. Nanog binds to promoter elements of hundreds of target genes as a regulatory element. It has a conserved DNA-binding homeodomain with consensus sequence TAATGG. The closest homolog of Nanog is the (nonmammalia) Xenopus, a Xvent transcription factor with consensus sequence CTAATT [
36]. In this subsection, we investigate the algebraic morphology of both transcription factors Nanog and Xvent thanks to their consensus sequences.
Table 2.
A few (three-base) transcription factors whose group structure is away from a free group or whose Groebner basis of the
character variety contains a (possibly almost) singular surface. The symbol gene is for the identification of the transcription factor in the Jaspar database [
34], motif is for the consensus sequence of the transcription factor, card seq is for the cardinality sequence of conjugacy classes of subgroups of the group whose motif is the generator, simple sing is for the identification of a surface with simple singularities within the Groebner basis and the last column is for a reference paper and the corresponding disease. The group
is the free group of rank two. The card seq for
is
, close to the card seq of the group
. The latter group is found as governing the structure of many transcription factors and is associated to the link found in ([
13],
Figure 2). The card seq for
is
. The surface
(not defined in the text) is part of the character variety for the genes Pitx1, OTX1, etc.
Table 2.
A few (three-base) transcription factors whose group structure is away from a free group or whose Groebner basis of the
character variety contains a (possibly almost) singular surface. The symbol gene is for the identification of the transcription factor in the Jaspar database [
34], motif is for the consensus sequence of the transcription factor, card seq is for the cardinality sequence of conjugacy classes of subgroups of the group whose motif is the generator, simple sing is for the identification of a surface with simple singularities within the Groebner basis and the last column is for a reference paper and the corresponding disease. The group
is the free group of rank two. The card seq for
is
, close to the card seq of the group
. The latter group is found as governing the structure of many transcription factors and is associated to the link found in ([
13],
Figure 2). The card seq for
is
. The surface
(not defined in the text) is part of the character variety for the genes Pitx1, OTX1, etc.
| Gene | Motif | Card Seq | Simple Sing | Ref & Disease |
|---|
| Prdm1 | ACTTTC | | | [34], MA0508.2
lupus, rheumatoid arthritis
MA1549.1
lung adenocarcinoma
MA0076.2
gastric cancer
[MA0712.2, MA0883.1]
medulloblastomas
[37]
drug sensitivity |
| POU6F1 | TAATGAG | | no |
| ELK4 | CTTCCGG | . | no, Fricke |
| OTX2 | GGATTA | | no |
| N-box | TTCCGG | . | no, Fricke |
| Pitx1,OTX1,⋯ | TAATCC | . | | [34], [MA0682.1,MA0711.1]
autism, epilepsy, ⋯ |
| Nanog | TAATGG | . | | [35]
cancer cells |
| Xvent | CTAATT | F2 | | [36] |
The Groebner basis for Xvent
takes the form
where
is the Cayley cubic (with its 4 simple singularities) and
(a surface with a single simple singularity of type
) as shown in
Figure 3 (Right). The forgotten factors are factors for planes or trivial smooth surfaces.
The Groebner basis for Xvent
takes the form
where
(a surface with three simple singularity of type
). The missing term does not contain surfaces with singularities. The character variety
contains the cubic surface
(with two simple singularities of type
) and other factors for planes or trivial smooth surfaces. Both surfaces
and
are pictured in
Figure 7.
Table 2 lists a few selected transcription factors, their card seq and the corresponding singular surfaces, if any. As announced, in the selected transcription factors, there exists a correlation between the lack of ‘syntactical freedom’, or the presence of a surface with isolated singularities in the character variety, with an identified disease.
4.3. Algebraic Morphology of microRNAs
MicroRNAs (miRNAs) are important players of the expression and regulation of genes by targeting specific messenger RNAs (mRNAs) for degradation or translational repression [
38,
39]. The miRNAs are approximately 22 nt (where nt is for nucleotide) long single-stranded RNA molecules. The genes encoding miRNAs have a length much longer than the processed mature miRNA molecule. Many miRNAs reside in introns of their pre-mRNA host genes. They share their regulatory elements, primary transcript and have a similar expression profile. MicroRNAs are transcribed by RNA polymerase II as large RNA precursors called pri-miRNAs. The pre-miRNAs are approximately 70-nucleotides in length and are folded into imperfect stem-loop structures; see
Figure 1 (Right) for an example.
Each miRNA is synthesized as a miRNA duplex comprising two strands (-5p and -3p). However, only one of the two strands is selectively incorporated into the RNA-induced silencing complex to act as a template for the transcript of a complementary mRNA [
40,
41]. For details about the mirRNA sequences, we use the Mir database [
42,
43].
Plant miRNAs usually have near-perfect pairing with their mRNA targets so that gene repression proceeds through cleavage of the target transcripts. In contrast, animal miRNAs are able to recognize their target mRNAs by using as few as 6 to 8 nucleotides (the seed region), which is not enough pairing for leading to cleavage of the target mRNAs. A given miRNA may have hundreds of different mRNA targets, and a given target might be regulated by multiple miRNAs.
Disregulation of miRNAs may lead to a disease such as cancer. A key microRNA known as an oncommir (involved in immunity and cancer) is mir-155.
Specifically the -3p strand is mir-155-3p.
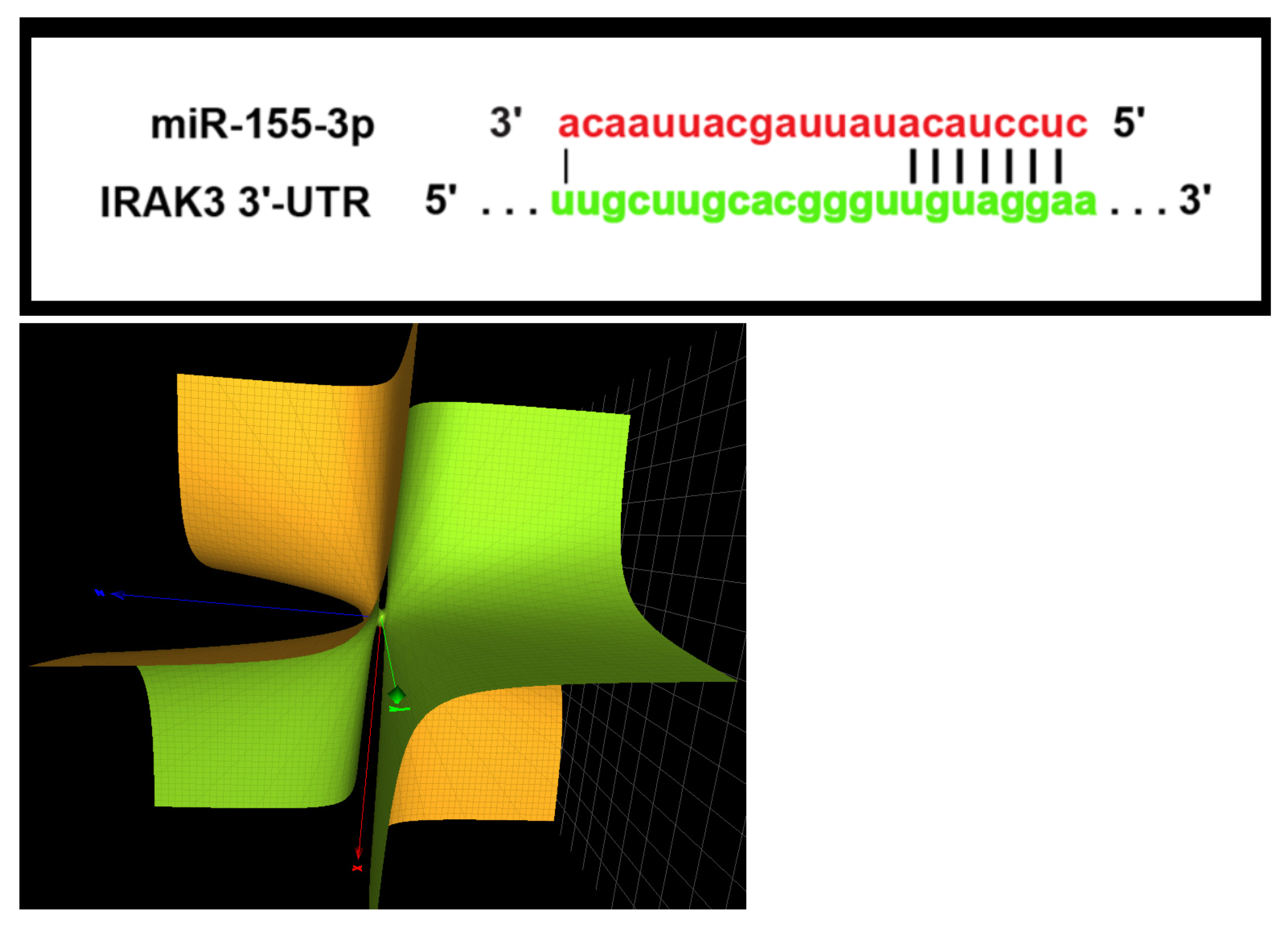
Figure 2 (top) illustrates the complementary base-pairing between miR-155-3p and the human IRAK3 (interleukin-1 receptor-associated kinase 3) mRNA ([
16],
Figure 4) and the relevant seed sequence UCCUAC. The card seq for this sequence is the two-letter free group
and the Groebner basis for the corresponding character variety contains the surface
that has a single simple singularity as shown in
Figure 2 (down). If one retains the full seed sequence is UCCUAC(A) then the card seq passes to that of the free group
to the group
and the singular surface is lost. This is a case where the ‘bandwidth’ of the seed is critical in the (dis)regulation of the miRNA. These results are transcribed in
Table 3.
Table 3.
A few human (prefix ‘hsa’) microRNAs whose group structure is away from a free group or whose Groebner basis of the
character variety contains a singular surface. The symbol mir is for the identification in the Mir database [
43], seed is for the seed of the miRNA, card seq is for the cardinality sequence of conjugacy classes of subgroups of the group whose seed is the generator, sing is the identification of a singular surface within the Groebner basis and the last column is for a reference paper and the corresponding disease [
40]. The card seq for
and
are given in ([
4], Table 5). The card seq for
is
. For hsa-mir-124-1-3p, one encounters the Fricke surface
in the character variety.
Table 3.
A few human (prefix ‘hsa’) microRNAs whose group structure is away from a free group or whose Groebner basis of the
character variety contains a singular surface. The symbol mir is for the identification in the Mir database [
43], seed is for the seed of the miRNA, card seq is for the cardinality sequence of conjugacy classes of subgroups of the group whose seed is the generator, sing is the identification of a singular surface within the Groebner basis and the last column is for a reference paper and the corresponding disease [
40]. The card seq for
and
are given in ([
4], Table 5). The card seq for
is
. For hsa-mir-124-1-3p, one encounters the Fricke surface
in the character variety.
| mir | Seed | Card Seq | Simple Sing | Ref & Disease |
|---|
| hsa-mir-193b-5p | GGGGUU | | no | [40,43]
lung cancer |
| | GGGGUUU | | no |
| hsa-mir-155-3p | UCCUAC | | | [40,41,43]
multiple sclerosis |
| | UCCUACA | | no |
| hsa-mir-193a-5p | GGGUCUU | | | [40,43]
breast cancer |
| hsa-mir-223-5p | GUGUAUU | . | . | . |
| hsa-mir-133-3p | UUGGUC | | | [40,43]
atrial fibrillation |
| | UUGGUCC | | no |
| hsa-mir-124-3p | AAGGCA | | | [43,44] |
| | AAGGCAC | . | no sing | Alzheimer’s disease |
For the case of -5p strand mir-155-3p, the seed sequence UUAAUGCUA contains four distinct letters. This case is similar to generic Groebner bases obtained from four letter seeds. Depending on the choices of parameters
, the Groebner basis contains the Cayley cubic
, the Fricke surface
(that is related to the link L6a1 ([
27],
Figure 2)), the surface
shown in
Figure 4 and other surfaces. In this generic case, the surface is found with (at most) 4 copies where each copy is attached to a distinct puncture of the 4-punctured 4-sphere
.
In
Table 3, this generic case is denoted
(or
for mir-133-5p). These results are transcribed in
Table 4.
Table 4.
The opposite strand of the microRNA considered in
Table 3. The seed sequence is made of 4 distinct bases and the corresponding card seq is the free group
of rank 3. The Groebner basis contains 4 copies of the generic collection of surfaces
,
,
, etc., as shown in
Figure 5, except for the -5p strand of mir-133, where there are only 3 copies of the generic surfaces.
Table 4.
The opposite strand of the microRNA considered in
Table 3. The seed sequence is made of 4 distinct bases and the corresponding card seq is the free group
of rank 3. The Groebner basis contains 4 copies of the generic collection of surfaces
,
,
, etc., as shown in
Figure 5, except for the -5p strand of mir-133, where there are only 3 copies of the generic surfaces.
| mir | Seed | Card Seq | Sing | Ref & Disease |
|---|
| hsa-mir-193b-3p | ACUGGCC | | generic | [40,43] |
| hsa-mir-155-5p | UUAAUGCUA | . | . | [40,41,43] |
| hsa-mir-193a-3p | ACUGGCC | . | . | [40,43] |
| hsa-mir-223-3p | GUCAGUU | . | . | . |
| hsa-mir-124-5p | GUGUUCA | . | . | . |
| hsa-mir-133-5p | GCUGGUA | . | generic | [43,44] |
A small list of huma miRNAs is investigated in
Table 3 and
Table 4 corresponding to 3-letter and 4-letter seeds. the prefix ‘hsa’ is for the human species. Similar to transcription factors in
Table 2, the lack of ‘syntactical freedom’, or the occurrence of a singular surface in the character variety, is symptomatic of a disease.
5. Conclusions
We found, in this work, that a signature of a disease may be given in terms of the group structure of a DNA/RNA sequence and the related character variety representing the group. The DNA motif of a transcription factor, or the seed of a microRNA, defines the generator of a group
. As soon as
is away from a free group
(with
the number of distinct bases in the sequence) or the
character variety
of
contains singular surfaces with isolated singularities, a potential disease is on sight, as shown in
Figure 6.
For example, for mir-155-3p, examined in much detail in
Section 4.3, the seed UCCUAC serves as the generator of the group
whose card seq is that of the free group
and whose Groebner basis of the variety contains the surface
possessing an isolated singularity of type
. The potential disease is multiple sclerosis. Note that a longer seed (with A added at the right hand side) is not appropriate to the detection. A partial list of other potential diseases identified from the structure of a miRNA seed can be seen in
Table 3. The diseases are a lung cancer associated with mir-193b-5p, a breast cancer associated with mir-194a-5p or mir-223-5p, an atrial fibrillation associated with mir-133-3p and an Alzheimer’s disease associated with mir-124-3p.
One would like to be more predictive in identifying the potential disease with peculiar groups or singular surfaces. First of all, most of the time, the surfaces encountered in the context of TFs and miRNAs are degree 3 del Pezzo, in contrast to surfaces obtained from other DNA sequences, as in Equation (
4). However, the degree 3 del Pezzo family is very rich. For instance, the singular surface
(see
Figure 2) is part of the character variety of TF Pitx1 (see
Table 2) and of miRNAs 155-3p and 193a-5p (in
Table 3). Then, the singular surface
is part of the character variety of mirRNA 133-3p. Both surfaces have a simple singular point of type
but distinct singular subschemes (see
Section 3 for the notation).
An exception to the degree 3 del Pezzo rule was found in investigating the character variety for the Prdm1 transcription factor in
Section 4.1. Do these features and other ones to be described later help for the diagnostic of a potential disease? There is room for much work in the future along these lines, as shown in our recent new paper [
32].
In a separate direction, the work of Reference [
45] about biochirality and the CPT theorem may be possibly related to the space-time-spin group
that served as a prism of DNA/RNA structure in our approach. Related ideas about the concept of a time crystal are also worthwhile to be pointed out, e.g., Reference [
46].