A Double-Density Clustering Method Based on “Nearest to First in” Strategy
Abstract
1. Introduction
2. Core Objects and Initial Clustering
2.1. Abbreviations
2.2. Density of Each Point
2.3. Core Object
2.4. Initial Clustering
| Algorithm 1: Initial clustering. |
| Input: data set X, k, , . |
| Output: label array L. |
| Step 1. Initialize each element of L to 0, and set to 0; |
| Step 2. for each in X: |
| Step 3. Compute by (1); |
| Step 4. Calculate and by (2) and (3) respectively; |
| Step 5. Calculate the threshold by (4) |
| Step 6. for each in X: |
| Step 7. if and : |
| Step 8. |
| Step 9. Initialize an empty queue Q |
| Step 10. for each in X: |
| Step 11. if : continue; |
| Step 12. , ; |
| Step 13. while Q is not empty: |
| Step 14. , ; |
| Step 15. for each y in : |
| Step 16. if : ; |
| Step 17. return L. |
3. Nearest-to-First-in Strategy
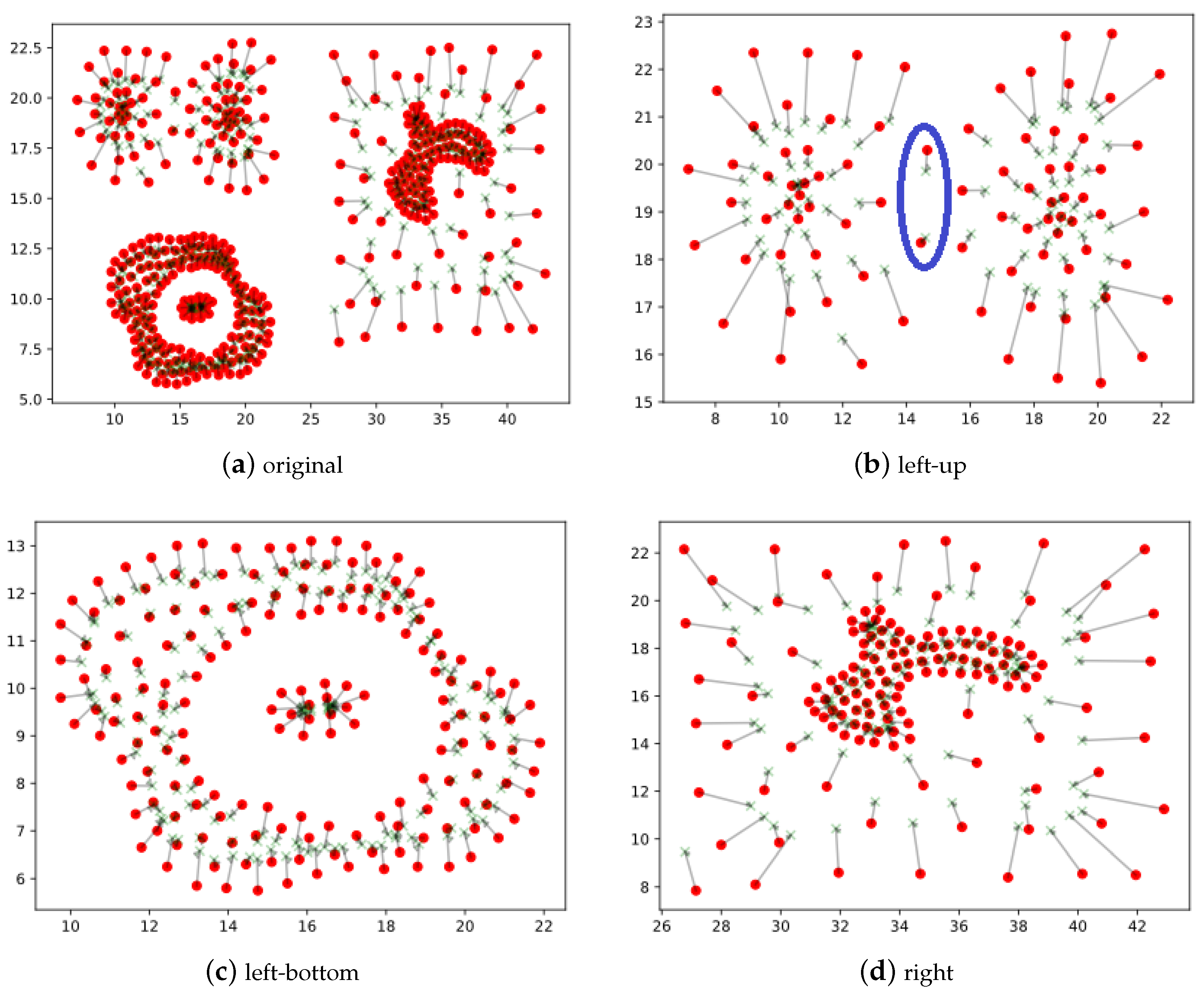
3.1. Basic Idea of the Strategy
3.2. Main Procedures of Nearest-to-First-in Strategy
| Algorithm 2: NTFI. |
| Input: . |
| Output: label array L. |
| Step 1. Put all unclassified point () into an array and create two array D and , each has the same size as . The value of each element in is set to −1 and the value of each element in D is set to ∞; |
| Step 2. while is not empty: |
| Step 3. ; |
| Step 4. for each x in : |
| Step 5. for each y in : |
| Step 6. if and : |
| Step 7. ; |
| Step 8. if : ; |
| Step 9. if : break; |
| Step 10. for each x in : |
| Step 11. if : |
| Step 12. ; |
| Step 13. Remove x from ; |
| Step 14. Return L. |
4. Experiments and Results Analysis
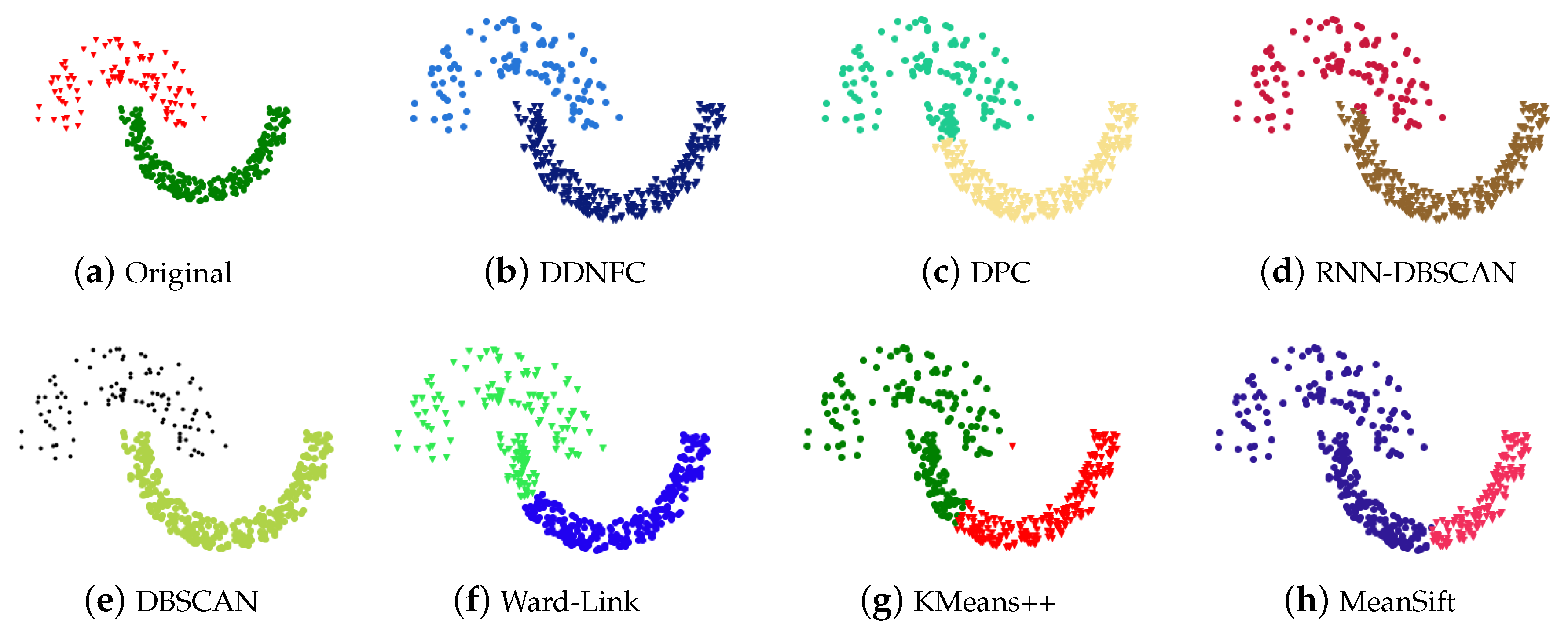
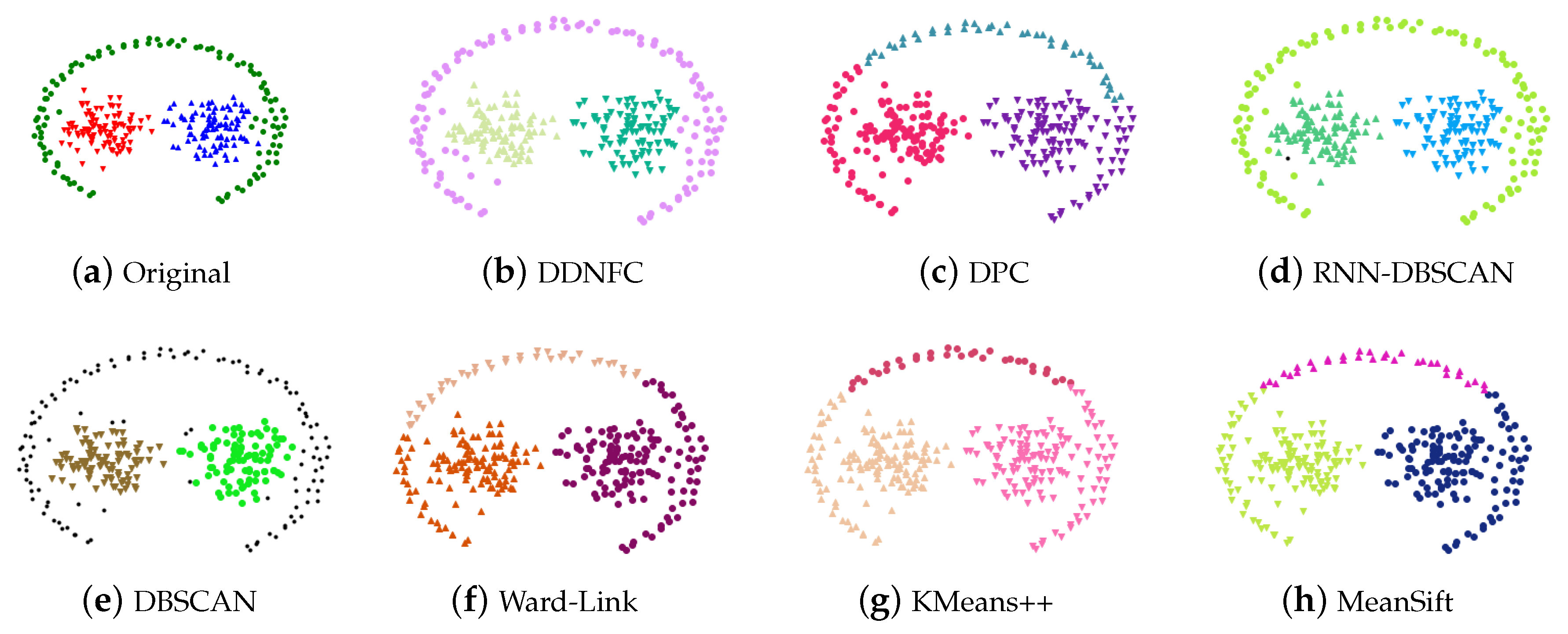
4.1. Experiments on Artificial Data Sets and Results Analysis
4.2. Experiments on Real-World Data Sets and Results Analysis
- Data that have null values, uncertain values, or duplicates were removed.
- Most of data sets have a class attribute. Table 3 only gives the number of none class attributes.
- We conserved from the third to ninth features of the data in Echocardiogram because some of its data has missing values.
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Appendix A. Figures of Experiment Results





References
- Aggarwal, C.C.; Reddy, C.K. Data Clustering Algorithms and Applications, 1st ed.; CRC Press: Boca Raton, FL, USA, 2013; pp. 65–210. [Google Scholar]
- Arthur, D.; Vassilvitskii, S. K-means++: The advantages of careful seeding. In Proceedings of the Annual ACM-SIAM Symposium on Discrete Algorithms, New Orleans, LA, USA, 7–9 January 2007; pp. 1027–1035. [Google Scholar]
- Murtagh, F.; Legendre, P. Ward’s Hierarchical Agglomerative Clustering Method: Which Algorithms Implement Ward’s Criterion? J. Classif. 2014, 31, 274–295. [Google Scholar] [CrossRef]
- Wu, K.L.; Yang, M.S. Mean shift-based clustering. Pattern Recognit. 2007, 40, 3035–3052. [Google Scholar] [CrossRef]
- Jiang, Z.; Liu, X.; Sun, M. A Density Peak Clustering Algorithm Based on the K-Nearest Shannon Entropy and Tissue-Like P System. Math. Probl. Eng. 2019, 2019, 1–13. [Google Scholar] [CrossRef]
- Halim, Z.; Khattak, J.H. Density-based clustering of big probabilistic graphs. Evol. Syst. 2019, 10, 333–350. [Google Scholar] [CrossRef]
- Wu, C.; Lee, J.; Isokawa, T.; Yao, J.; Xia, Y. Efficient Clustering Method Based on Density Peaks with Symmetric Neighborhood Relationship. IEEE Access 2019, 7, 60684–60696. [Google Scholar] [CrossRef]
- Tan, H.; Gao, Y.; Ma, Z. Regularized constraint subspace based method for image set classification. Pattern Recognit. 2018, 76, 434–448. [Google Scholar] [CrossRef]
- Chen, Y.; Tang, S.; Zhou, L.; Wang, C.; Du, J.; Wang, T.; Pei, S. Decentralized Clustering by Finding Loose and Distributed Density Cores. Inf. Sci. 2018, 433, 510–526. [Google Scholar] [CrossRef]
- Wang, Z.; Yu, Z.; Philip Chen, C.L.; You, J.; Gu, T.; Wong, H.S.; Zhang, J. Clustering by Local Gravitation. IEEE Trans. Cybern. 2018, 48, 1383–1396. [Google Scholar] [CrossRef]
- Oktar, Y.; Turkan, M. A review of sparsity-based clustering methods. Signal Process. 2018, 148, 20–30. [Google Scholar] [CrossRef]
- Chen, J.; Zheng, H.; Lin, X.; Wu, Y.; Su, M. A novel image segmentation method based on fast density clustering algorithm. Eng. Appl. Artif. Intell. 2018, 73, 92–110. [Google Scholar] [CrossRef]
- Zhou, R.; Zhang, Y.; Feng, S.; Luktarhan, N. A novel hierarchical clustering algorithm based on density peaks for complex datasets. Complexity 2018, 2018, 1–8. [Google Scholar] [CrossRef]
- Zhang, X.; Liu, H.; Zhang, X. Novel density-based and hierarchical density-based clustering algorithms for uncertain data. Neural Netw. 2017, 93, 240–255. [Google Scholar] [CrossRef] [PubMed]
- Wu, B.; Wilamowski, B.M. A fast density and grid based clustering method for data with arbitrary shapes and noise. IEEE Trans. Ind. Inform. 2017, 13, 1620–1628. [Google Scholar] [CrossRef]
- Lv, Y.; Ma, T.; Tang, M.; Cao, J.; Tian, Y.; Al-Dhelaan, A.; Al-Rodhaan, M. An efficient and scalable density-based clustering algorithm for datasets with complex structures. Neurocomputing 2016, 171, 9–22. [Google Scholar] [CrossRef]
- Ester, M.; Kriegel, H.P.; Sander, J.; Xu, X. A Density-Based Algorithm for Discovering Clusters in Large Spatial Databases with Noise. In Proceedings of the 2nd International Conference on Knowledge Discovery and Data Mining, Portland, OR, USA, 2–4 August 1996; pp. 226–231. [Google Scholar]
- Rodriguez, A.; Laio, A. Clustering by fast search and find of density peaks. Science 2014, 344, 1492–1496. [Google Scholar] [CrossRef] [PubMed]
- Vadapalli, S.; Valluri, S.R.; Karlapalem, K. A simple yet effective data clustering algorithm. In Proceedings of the IEEE International Conference on Data Mining, Hong Kong, China, 18–22 December 2006; pp. 1108–1112. [Google Scholar] [CrossRef]
- Fränti, P.; Sieranoja, S. K-means properties on six clustering benchmark datasets. Appl. Intell. 2018, 48, 4743–4759. [Google Scholar] [CrossRef]
- Bryant, A.; Cios, K. RNN-DBSCAN: A density-based clustering algorithm using reverse nearest neighbor density estimates. IEEE Trans. Knowl. Data Eng. 2018, 30, 1109–1121. [Google Scholar] [CrossRef]
- Romano, S.; Vinh, N.X.; Bailey, J.; Verspoor, K. Adjusting for chance clustering comparison measures. J. Mach. Learn. Res. 2016, 17, 1–32. [Google Scholar]
- Xie, J.; Xiong, Z.Y.; Dai, Q.Z.; Wang, X.X.; Zhang, Y.F. A new internal index based on density core for clustering validation. Inf. Sci. 2020, 506, 346–365. [Google Scholar] [CrossRef]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Von Luxburg, U. A tutorial on spectral clustering. Stat. Comput. 2007, 17, 395–416. [Google Scholar] [CrossRef]
- Karypis, G.; Han, E.H.; Kumar, V. Chameleon: Hierarchical clustering using dynamic modeling. Computer 1999, 32, 68–75. [Google Scholar] [CrossRef]
- Dua, D.; Graff, C. UCI Machine Learning Repository. Available online: https://archive.ics.uci.edu/ml/index.php (accessed on 1 October 2017).





| Data Set | Size | Dimension | Cluster Number |
|---|---|---|---|
| Compound | 399 | 2 | 6 |
| Pathbased | 300 | 2 | 3 |
| Flame | 240 | 2 | 2 |
| Aggregation | 788 | 2 | 7 |
| Jain | 373 | 2 | 3 |
| R15 | 600 | 2 | 15 |
| Unbalance | 6500 | 2 | 8 |
| A3 | 7500 | 2 | 50 |
| S1 | 5000 | 2 | 15 |
| t5.8k | 8000 | 2 | 6 |
| t7.10k | 10,000 | 2 | 9 |
| t8.8k | 8000 | 2 | 8 |
| Spiral | 312 | 2 | 3 |
| Algorithm | Par | c/C | F1 | AMI | ARI | Par | c/C | F1 | AMI | ARI | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Compound | Pathbased | ||||||||||
| DDNFC | 9 | 6/6 | 1.00 | 0.99 | 1.00 | 9 | 3/3 | 0.98 | 0.92 | 0.94 | |
| DPC | 1.0 | 6/6 | 0.78 | 0.82 | 0.62 | 1.5 | 3/3 | 0.74 | 0.55 | 0.50 | |
| RNN-DBSCAN | 8 | 6/6 | 0.89 | 0.86 | 0.86 | 6 | 4/3 | 0.98 | 0.93 | 0.95 | |
| DBSCAN | 0.2/5 | 6/6 | 0.94 | 0.89 | 0.93 | 0.3/11 | 3/3 | 0.96 | 0.84 | 0.88 | |
| Ward-Link | 6 | 6/6 | 0.72 | 0.69 | 0.55 | 3 | 3/3 | 0.72 | 0.53 | 0.48 | |
| KMeans++ | 6 | 6/6 | 0.70 | 0.68 | 0.54 | 3 | 3/3 | 0.70 | 0.51 | 0.46 | |
| MeanShift | - | 4/6 | 0.83 | 0.74 | 0.78 | - | 3/3 | 0.70 | 0.51 | 0.46 | |
| Jain | Flame | ||||||||||
| DDNFC | 18 | 2/2 | 1.00 | 1.00 | 1.00 | 17 | 2/2 | 1.00 | 1.00 | 1.00 | |
| DPC | 1.0 | 2/2 | 0.93 | 0.61 | 0.71 | 4.5 | 2/2 | 1.00 | 0.96 | 0.98 | |
| RNN-DBSCAN | 15 | 2/2 | 0.99 | 0.92 | 0.97 | 8 | 2/2 | 0.99 | 0.93 | 0.97 | |
| DBSCAN | 0.25/14 | 2/2 | 1.00 | 1.00 | 1.00 | 0.45/11 | 3/2 | 0.99 | 0.91 | 0.97 | |
| Ward-Link | 2 | 2/2 | 0.87 | 0.47 | 0.51 | 2 | 2/2 | 0.72 | 0.33 | 0.19 | |
| KMeans++ | 2 | 2/2 | 0.80 | 0.34 | 0.32 | 2 | 2/2 | 0.84 | 0.39 | 0.45 | |
| MeanShift | - | 2/2 | 0.62 | 0.21 | 0.02 | - | 2/2 | 0.86 | 0.43 | 0.50 | |
| cluto-t8.8k | cluto-t7.10k | ||||||||||
| DDNFC | 24 | 9/9 | 0.94 | 0.92 | 0.95 | 35 | 10/10 | 0.90 | 0.87 | 0.89 | |
| DPC | 1.0 | 9/9 | 0.58 | 0.60 | 0.48 | 1.0 | 10/10 | 0.55 | 0.57 | 0.37 | |
| RNN-DBSCAN | 22 | 9/9 | 0.94 | 0.91 | 0.94 | 35 | 10/10 | 0.90 | 0.87 | 0.90 | |
| DBSCAN | 0.1/23 | 10/9 | 0.92 | 0.90 | 0.91 | 0.1/8 | 10/10 | 0.44 | 0.28 | 0.20 | |
| Ward-Link | 9 | 9/9 | 0.49 | 0.53 | 0.33 | 10 | 10/10 | 0.55 | 0.61 | 0.41 | |
| KMeans++ | 9 | 9/9 | 0.52 | 0.55 | 0.36 | 10 | 10/10 | 0.47 | 0.54 | 0.34 | |
| MeanShift | - | 7/9 | 0.51 | 0.54 | 0.36 | - | 6/10 | 0.54 | 0.58 | 0.44 | |
| Aggregation | cluto-t5.8k | ||||||||||
| DDNFC | 6 | 7/7 | 0.99 | 0.99 | 0.99 | 34 | 7/7 | 0.82 | 0.79 | 0.78 | |
| DPC | 2.0 | 7/7 | 1.00 | 1.00 | 1.00 | 2.0 | 7/7 | 0.79 | 0.79 | 0.74 | |
| RNN-DBSCAN | 5 | 7/7 | 1.00 | 0.99 | 0.99 | 39 | 7/7 | 0.82 | 0.80 | 0.79 | |
| DBSCAN | 0.3/23 | 7/7 | 0.94 | 0.91 | 0.91 | 0.15/11 | 7/7 | 0.28 | 0.05 | 0.01 | |
| Ward-Link | 7 | 7/7 | 0.90 | 0.88 | 0.81 | 7 | 7/7 | 0.78 | 0.79 | 0.74 | |
| KMeans++ | 7 | 7/7 | 0.85 | 0.83 | 0.76 | 7 | 7/7 | 0.78 | 0.79 | 0.74 | |
| MeanShift | - | 6/7 | 0.90 | 0.84 | 0.84 | - | 6/7 | 0.81 | 0.79 | 0.78 | |
| Unbalance | R15 | ||||||||||
| DDNFC | 63 | 8/8 | 1.00 | 1.00 | 1.00 | 24 | 15/15 | 0.99 | 0.99 | 0.99 | |
| DPC | 1.0 | 8/8 | 1.00 | 1.00 | 1.00 | 1.0 | 15/15 | 1.00 | 0.99 | 0.99 | |
| RNN-DBSCAN | 25 | 8/8 | 1.00 | 1.00 | 1.00 | 12 | 15/15 | 1.00 | 0.99 | 0.99 | |
| DBSCAN | 0.45/11 | 8/8 | 0.79 | 0.70 | 0.61 | 0.1/17 | 15/15 | 0.76 | 0.62 | 0.27 | |
| Ward-Link | 8 | 8/8 | 1.00 | 1.00 | 1.00 | 15 | 15/15 | 0.99 | 0.99 | 0.98 | |
| KMeans++ | 8 | 8/8 | 1.00 | 1.00 | 1.00 | 15 | 15/15 | 1.00 | 0.99 | 0.99 | |
| MeanShift | - | 8/8 | 1.00 | 1.00 | 1.00 | - | 8/15 | 0.58 | 0.58 | 0.27 | |
| S1 | A3 | ||||||||||
| DDNFC | 29 | 15/15 | 0.99 | 0.99 | 0.99 | 36 | 50/50 | 0.97 | 0.97 | 0.94 | |
| DPC | 2.5 | 15/15 | 0.99 | 0.99 | 0.99 | 1.0 | 50/50 | 0.95 | 0.95 | 0.91 | |
| RNN-DBSCAN | 20 | 16/15 | 0.99 | 0.99 | 0.99 | 39 | 49/50 | 0.97 | 0.97 | 0.95 | |
| DBSCAN | 0.1/8 | 15/15 | 0.94 | 0.93 | 0.89 | 0.1/29 | 37/50 | 0.78 | 0.86 | 0.67 | |
| Ward-Link | 15 | 15/15 | 0.99 | 0.98 | 0.98 | 50 | 50/50 | 0.97 | 0.97 | 0.94 | |
| KMeans++ | 15 | 15/15 | 0.99 | 0.99 | 0.99 | 50 | 50/50 | 0.96 | 0.97 | 0.94 | |
| MeanShift | - | 9/15 | 0.64 | 0.70 | 0.56 | - | - | - | - | - | |
| Data Set | Size | Dimension | Cluster Number |
|---|---|---|---|
| Ionosphere | 351 | 34 | 2 |
| Page-blocks | 5437 | 10 | 5 |
| Haberman | 306 | 3 | 2 |
| SPECT-train | 80 | 22 | 2 |
| Segmentation | 2100 | 19 | 7 |
| Chess | 3196 | 36 | 2 |
| Breast-cancer-Wisconsin | 699 | 10 | 2 |
| Pendigits-train | 7494 | 16 | 10 |
| Sonar | 208 | 60 | 2 |
| Echocardiogram | 106 | 7 | 2 |
| Contraceptive-Method-Choice | 1473 | 9 | 3 |
| Wilt-train | 4339 | 5 | 2 |
| Algorithm | Par | c/C | F1 | AMI | ARI | Par | c/C | F1 | AMI | ARI | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Ionosphere | Page-blocks | ||||||||||
| DDNFC | 22 | 2/2 | 0.89 | 0.47 | 0.60 | 25 | 5/5 | 0.90 | 0.26 | 0.45 | |
| DPC | 4.5 | 2/2 | 0.79 | 0.22 | 0.32 | 1.5 | 5/5 | 0.86 | 0.04 | 0.10 | |
| RNN-DBSCAN | 6 | 2/2 | 0.69 | 0.01 | 0.01 | 19 | 5/5 | 0.90 | 0.26 | 0.48 | |
| DBSCAN | 0.45/8 | 2/2 | 0.65 | 0.09 | −0.05 | 0.1/5 | 7/5 | 0.86 | 0.04 | 0.07 | |
| Ward-Link | 2 | 2/2 | 0.72 | 0.13 | 0.19 | 3 | 5/5 | 0.80 | 0.05 | 0.02 | |
| K-Means++ | 2 | 2/2 | 0.72 | 0.13 | 0.18 | 5 | 5/5 | 0.82 | 0.05 | 0.02 | |
| MeanShift | - | 1/2 | - | - | - | - | 1/5 | - | - | - | |
| Haberman | SPECT-train | ||||||||||
| DDNFC | 10 | 2/2 | 0.73 | 0.01 | 0.05 | 4 | 2/2 | 0.70 | 0.14 | 0.21 | |
| DPC | 1.5 | 2/2 | 0.61 | −0.00 | 0.01 | 1.0 | 2/2 | 0.66 | −0.01 | −0.00 | |
| RNN-DBSCAN | 10 | 2/2 | 0.73 | −0.00 | 0.01 | 19 | 2/2 | 0.64 | 0.10 | 0.04 | |
| DBSCAN | 0.1/5 | 1/2 | - | - | - | 0.1/5 | 2/2 | 0.62 | 0.08 | 0.06 | |
| Ward-Link | 2 | 2/2 | 0.61 | 0.00 | −0.02 | 2 | 2/2 | 0.61 | 0.03 | 0.04 | |
| K-Means++ | 2 | 2/2 | 0.55 | −0.00 | −0.00 | 2 | 2/2 | 0.70 | 0.17 | 0.17 | |
| MeanShift | - | 2/2 | 0.73 | −0.00 | 0.01 | - | 1/2 | - | - | - | |
| Segmentation | Chess | ||||||||||
| DDNFC | 17 | 7/7 | 0.67 | 0.59 | 0.42 | 15 | 2/2 | 0.67 | 0.00 | 0.00 | |
| DPC | 1.0 | 7/7 | 0.57 | 0.52 | 0.35 | 2.0 | 2/2 | 0.61 | 0.00 | 0.01 | |
| RNN-DBSCAN | 13 | 6/7 | 0.60 | 0.56 | 0.38 | 9 | 2/2 | 0.67 | 0.00 | 0.00 | |
| DBSCAN | 0.1/5 | 1/7 | - | - | - | 0.1/5 | 1/2 | - | - | - | |
| Ward-Link | 7 | 7/7 | 0.57 | 0.46 | 0.32 | 2 | 2/2 | 0.59 | 0.01 | 0.00 | |
| K-Means++ | 7 | 7/7 | 0.54 | 0.48 | 0.33 | 2 | 2/2 | 0.51 | 0.00 | 0.00 | |
| MeanShift | - | 7/7 | 0.38 | 0.20 | 0.10 | - | 1/2 | - | - | - | |
| Breast-cancer-Wisconsin | Pendigits-train | ||||||||||
| DDNFC | 27 | 2/2 | 0.94 | 0.65 | 0.77 | 60 | 10/10 | 0.73 | 0.70 | 0.55 | |
| DPC | 3.0 | 2/2 | 0.59 | 0.20 | 0.02 | 1.0 | 10/10 | 0.73 | 0.73 | 0.61 | |
| RNN-DBSCAN | 39 | 2/2 | 0.69 | 0.00 | 0.00 | 32 | 10/10 | 0.68 | 0.63 | 0.41 | |
| DBSCAN | 0.1/26 | 2/2 | 0.68 | 0.03 | −0.03 | 0.1/5 | 1/10 | - | - | - | |
| Ward-Link | 2 | 2/2 | 0.97 | 0.79 | 0.87 | 10 | 10/10 | 0.75 | 0.72 | 0.59 | |
| K-Means++ | 2 | 2/2 | 0.96 | 0.74 | 0.85 | 10 | 10/10 | 0.71 | 0.67 | 0.54 | |
| MeanShift | - | 4/2 | 0.95 | 0.68 | 0.84 | - | 3/10 | 0.34 | 0.23 | 0.17 | |
| Sonar | Echocardiogram | ||||||||||
| DDNFC | 11 | 2/2 | 0.66 | 0.02 | 0.01 | 7 | 2/2 | 0.71 | 0.03 | 0.05 | |
| DPC | 1.0 | 2/2 | 0.57 | −0.00 | −0.00 | 1.0 | 2/2 | 0.70 | 0.01 | 0.06 | |
| RNN-DBSCAN | 7 | 2/2 | 0.67 | −0.00 | −0.00 | 4 | 2/2 | 0.70 | −0.00 | 0.04 | |
| DBSCAN | 0.35/5 | 3/2 | 0.65 | 0.05 | −0.01 | 0.1/5 | 1/2 | - | - | - | |
| Ward-Link | 2 | 2/2 | 0.58 | −0.00 | −0.00 | 2 | 2/2 | 0.72 | 0.10 | 0.16 | |
| K-Means++ | 2 | 2/2 | 0.54 | 0.00 | 0.00 | 2 | 2/2 | 0.73 | 0.11 | 0.19 | |
| MeanShift | - | - | - | - | - | - | 2/2 | 0.71 | 0.00 | 0.04 | |
| Contraceptive-Method-Choice | Wilt-train | ||||||||||
| DDNFC | 9 | 3/3 | 0.52 | 0.00 | −0.00 | 12 | 2/2 | 0.68 | 0.00 | -0.00 | |
| DPC | 1.0 | 3/3 | 0.43 | 0.02 | 0.02 | 1.0 | 2/2 | 0.68 | −0.00 | −0.00 | |
| RNN-DBSCAN | 19 | 2/3 | 0.52 | −0.00 | −0.00 | 29 | 2/2 | 0.68 | −0.00 | −0.00 | |
| DBSCAN | 0.1/5 | 1/3 | - | - | - | 0.1/5 | 1/2 | - | - | - | |
| Ward-Link | 3 | 3/3 | 0.39 | 0.03 | 0.02 | 2 | 2/2 | 0.55 | 0.01 | -0.00 | |
| K-Means++ | 3 | 3/3 | 0.41 | 0.03 | 0.03 | 2 | 2/2 | 0.55 | 0.00 | 0.00 | |
| MeanShift | - | 2/3 | 0.46 | 0.01 | 0.01 | - | 3/2 | 0.68 | 0.01 | −0.01 | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Y.; Liu, D.; Yu, F.; Ma, Z. A Double-Density Clustering Method Based on “Nearest to First in” Strategy. Symmetry 2020, 12, 747. https://doi.org/10.3390/sym12050747
Liu Y, Liu D, Yu F, Ma Z. A Double-Density Clustering Method Based on “Nearest to First in” Strategy. Symmetry. 2020; 12(5):747. https://doi.org/10.3390/sym12050747
Chicago/Turabian StyleLiu, Yaohui, Dong Liu, Fang Yu, and Zhengming Ma. 2020. "A Double-Density Clustering Method Based on “Nearest to First in” Strategy" Symmetry 12, no. 5: 747. https://doi.org/10.3390/sym12050747
APA StyleLiu, Y., Liu, D., Yu, F., & Ma, Z. (2020). A Double-Density Clustering Method Based on “Nearest to First in” Strategy. Symmetry, 12(5), 747. https://doi.org/10.3390/sym12050747





