Clinical and Mutational Spectrum of Xeroderma Pigmentosum in Egypt: Identification of Six Novel Mutations and Implications for Ancestral Origins
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients
2.2. Clinical Evaluation
2.3. Molecular Investigation
3. Results
3.1. Clinical Description
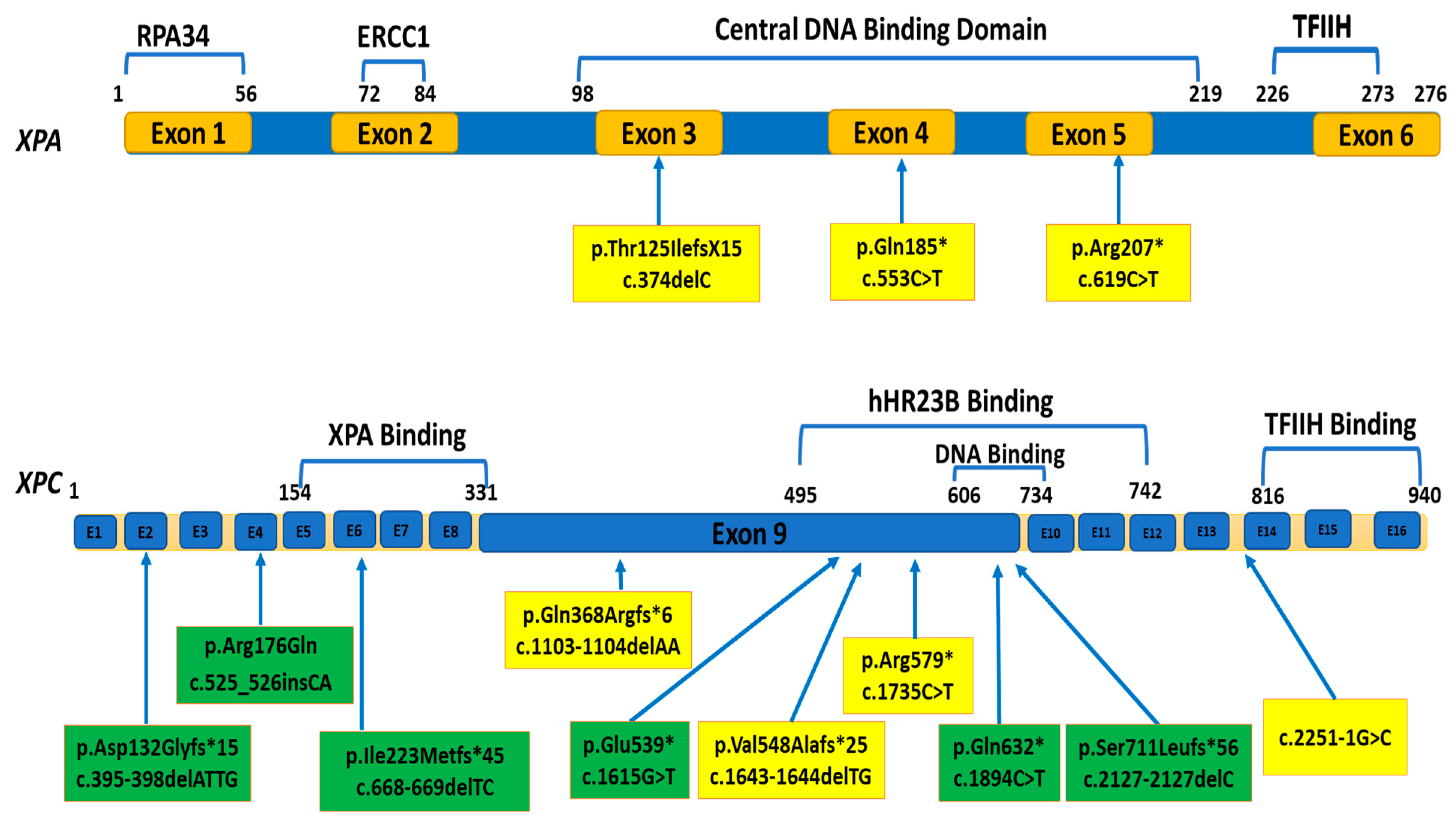
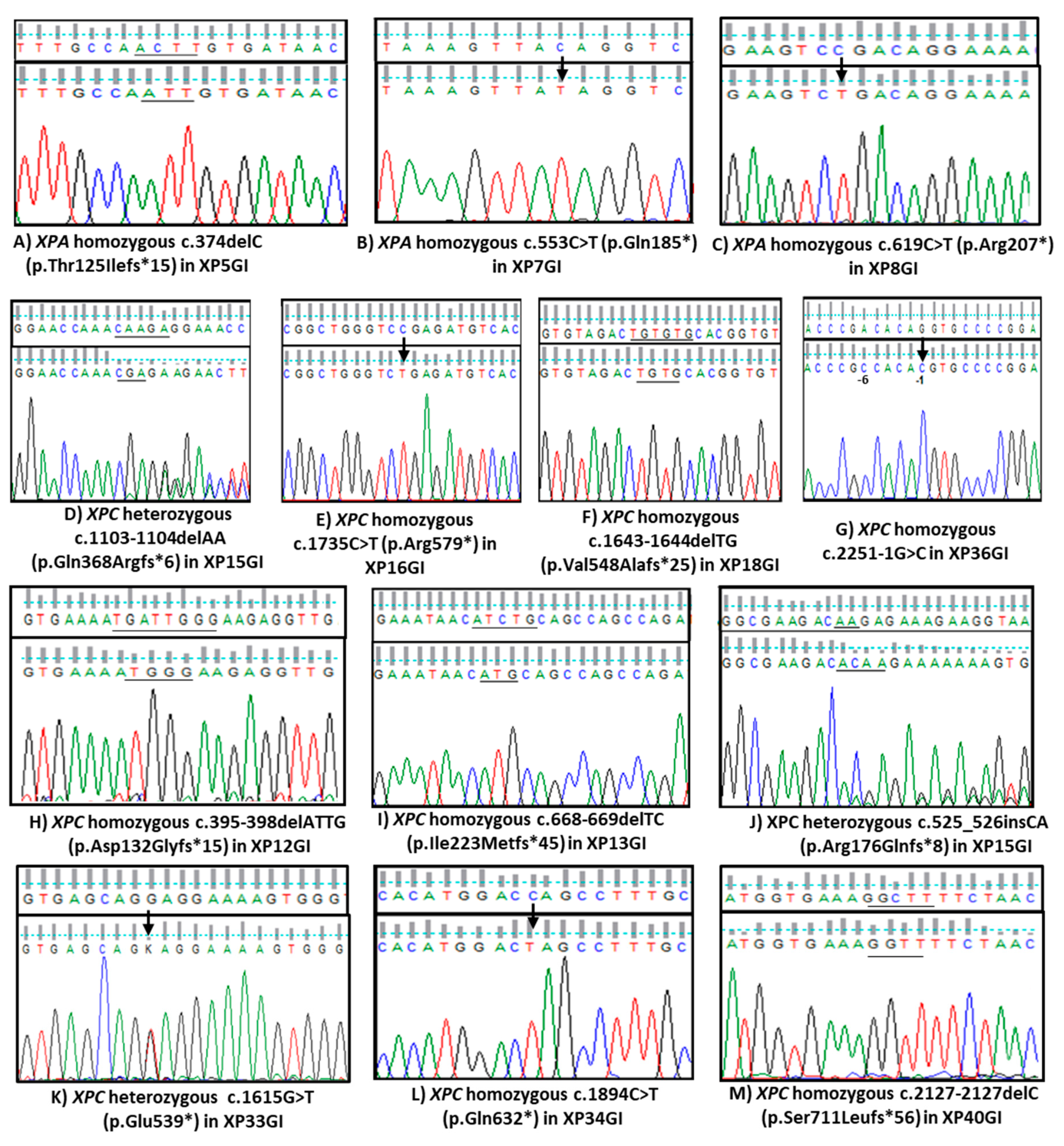
3.2. Molecular Results
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Bradford, P.T.; Goldstein, A.M.; Tamura, D.; Khan, S.G.; Ueda, T.; Boyle, J.; Oh, K.-S.; Imoto, K.; Inui, H.; Moriwaki, S.-I.; et al. Cancer and neurologic degeneration in xeroderma pigmentosum: Long term follow-up characterises the role of DNA repair. J. Med. Genet. 2010, 48, 168–176. [Google Scholar] [CrossRef]
- Sethi, M.; Lehmann, A.; Fawcett, H.; Stefanini, M.; Jaspers, N.; Mullard, K.; Turner, S.; Robson, A.; McGibbon, D.; Sarkany, R.; et al. Patients with xeroderma pigmentosum complementation groups C, E and V do not have abnormal sunburn reactions. Br. J. Dermatol. 2013, 169, 1279–1287. [Google Scholar] [CrossRef]
- Lim, R.; Sethi, M.; Morley, A.M. Ophthalmic manifestations of xeroderma pigmentosum. Ophthalmology 2017, 124, 1652–1661. [Google Scholar] [CrossRef]
- Totonchy, M.B.; Tamura, D.; Pantell, M.S.; Zalewski, C.; Bradford, P.T.; Merchant, S.N.; Nadol, J.; Khan, S.G.; Schiffmann, R.; Pierson, T.M.; et al. Auditory analysis of xeroderma pigmentosum 1971–2012: Hearing function, sun sensitivity and DNA repair predict neurological degeneration. Brain 2013, 136, 194–208. [Google Scholar] [CrossRef]
- Rapin, I.; Lindenbaum, Y.; Dickson, D.W.; Kraemer, K.H.; Robbins, J.H. Cockayne syndrome and xeroderma pigmentosum. Neurology 2000, 55, 1442–1449. [Google Scholar] [CrossRef] [PubMed]
- Anttinen, A.; Koulu, L.; Nikoskelainen, E.; Portin, R.; Kurki, T.; Erkinjuntti, M.; Jaspers, N.G.; Raams, A.; Green, M.H.; Lehmann, A.R.; et al. Neurological symptoms and natural course of xeroderma pigmentosum. Brain 2008, 131, 1979–1989. [Google Scholar] [CrossRef] [PubMed]
- Lindenbaum, Y.; Dickson, D.; Rosenbaum, P.; Kraemer, K.; Robbins, J.; Rapin, I. Xeroderma pigmentosum/cockayne syndrome complex: First neuropathological study and review of eight other cases. Eur. J. Paediatr. Neurol. 2001, 5, 225–242. [Google Scholar] [CrossRef] [PubMed]
- Bologna, S.B.; Harumi Nakajima Teshima, T.; Lourenço, S.V.; Nico, M.M.S. An atrophic, telangiectatic patch at the distal border of the tongue: A mucous membrane manifestation of xeroderma pigmentosum. Pediatr. Dermatol. 2014, 31, e38–e41. [Google Scholar] [CrossRef]
- Brooks, B.P.; Thompson, A.H.; Bishop, R.J.; Clayton, J.A.; Chan, C.-C.; Tsilou, E.T.; Zein, W.M.; Tamura, D.; Khan, S.G.; Ueda, T.; et al. Ocular manifestations of xeroderma pigmentosum. Ophthalmology 2013, 120, 1324–1336. [Google Scholar] [CrossRef] [PubMed]
- Kraemer, K.H.; Lee, M.M.; Scotto, J. Xeroderma Pigmentosum: Cutaneous, Ocular, and Neurologic Abnormalities in 830 Published Cases. Arch. Dermatol. 1987, 123, 241–250. [Google Scholar] [CrossRef]
- Fassihi, H.; Sethi, M.; Fawcett, H.; Wing, J.; Chandler, N.; Mohammed, S.; Craythorne, E.; Morley, A.M.S.; Lim, R.; Turner, S.; et al. Deep phenotyping of 89 xeroderma pigmentosum patients reveals unexpected heterogeneity dependent on the precise molecular defect. Proc. Natl. Acad. Sci. USA 2016, 113, E1236–E1245. [Google Scholar] [CrossRef]
- Tamura, D.; Ono, R.; DiGiovanna, J.J.; Kraemer, K.H. Management of xeroderma pigmentosum. In DNA Repair Disorders; Springer: Singapore, 2019; pp. 203–221. [Google Scholar]
- Rastogi, R.P.; Richa, K.A.; Tyagi, M.B.; Sinha, R.P. Molecular mechanisms of ultraviolet radiation-induced DNA damage and repair. J. Nucleic Acids 2010, 2010, 592980. [Google Scholar] [CrossRef] [PubMed]
- Van Steeg, H.; Kraemer, K.H. Xeroderma pigmentosum and the role of UV-induced DNA damage in skin cancer. Mol. Med. Today 1999, 5, 86–94. [Google Scholar] [CrossRef]
- Kraemer, K.H.; DiGiovanna, J.J. Global contributions to the understanding of DNA repair and skin cancer. J. Investig. Dermatol. 2014, 134, E8–E17. [Google Scholar] [CrossRef] [PubMed]
- Sugasawa, K.; Okuda, Y.; Saijo, M.; Nishi, R.; Matsuda, N.; Chu, G.; Mori, T.; Iwai, S.; Tanaka, K.; Tanaka, K.; et al. UV-Induced ubiquitylation of XPC protein mediated by UV-DDB-Ubiquitin Ligase Complex. Cell 2005, 121, 387–400. [Google Scholar] [CrossRef]
- Volker, M.; Moné, M.J.; Karmakar, P.; Van Hoffen, A.; Schul, W.; Vermeulen, W.; Hoeijmakers, J.H.J.; Van Driel, R.; Van Zeeland, A.A.; Mullenders, L.H.F. Sequential assembly of the nucleotide excision repair factors in vivo. Mol. Cell 2001, 8, 213–224. [Google Scholar] [CrossRef]
- Fadda, E. Role of the XPA protein in the NER pathway: A perspective on the function of structural disorder in macromolecular assembly. Comput. Struct. Biotechnol. J. 2016, 14, 78–85. [Google Scholar] [CrossRef]
- Ogi, T.; Limsirichaikul, S.; Overmeer, R.M.; Volker, M.; Takenaka, K.; Cloney, R.; Nakazawa, Y.; Niimi, A.; Miki, Y.; Jaspers, N.G.; et al. Three DNA polymerases, recruited by different mechanisms, carry out NER repair synthesis in human cells. Mol. Cell 2010, 37, 714–727. [Google Scholar] [CrossRef]
- Stallons, L.J.; McGregor, W.G. Translesion synthesis polymerases in the prevention and promotion of carcinogenesis. J. Nucleic Acids 2010, 2010. [Google Scholar] [CrossRef] [PubMed]
- Masutani, C.; Kusumoto, R.; Yamada, A.; Dohmae, N.; Yokoi, M.; Yuasa, M.; Araki, M.; Iwai, S.; Takio, K.; Hanaoka, F. The XPV (xeroderma pigmentosum variant) gene encodes human DNA polymerase η. Nature 1999, 399, 700–704. [Google Scholar] [CrossRef]
- Rivera-Begeman, A.; McDaniel, L.D.; Schultz, R.A.; Friedberg, E.C. A novel XPC pathogenic variant detected in archival material from a patient diagnosed with Xeroderma Pigmentosum: A case report and review of the genetic variants reported in XPC. DNA Repair 2007, 6, 100–114. [Google Scholar] [CrossRef] [PubMed]
- Kleijer, W.J.; Laugel, V.; Berneburg, M.; Nardo, T.; Fawcett, H.; Gratchev, A.; Jaspers, N.G.; Sarasin, A.; Stefanini, M.; Lehmann, A.R. Incidence of DNA repair deficiency disorders in western Europe: Xeroderma pigmentosum, Cockayne syndrome and trichothiodystrophy. DNA Repair 2008, 7, 744–750. [Google Scholar] [CrossRef] [PubMed]
- Tamura, D.; DiGiovanna, J.J.; Kraemer, K.H. Founder mutations in xeroderma pigmentosum. J. Investig. Dermatol. 2010, 130, 1491–1493. [Google Scholar] [CrossRef]
- Zghal, M.; El-Fekih, N.; Fazaa, B.; Fredj, M.; Zhioua, R.; Mokhtar, I.; Mrabet, A.; Ferjani, M.; Gaigi, S.; Kamoun, M.R. Xeroderma pigmentosum. Cutaneous, ocular, and neurologic abnormalities in 49 Tunisian cases. Tunis. Med. 2005, 83, 760–763. [Google Scholar]
- Jerbi, M.; Ben Rekaya, M.; Naouali, C.; Jones, M.; Messaoud, O.; Tounsi, H.; Nagara, M.; Chargui, M.; Kefi, R.; Boussen, H.; et al. Clinical, genealogical and molecular investigation of the xeroderma pigmentosum type C complementation group in Tunisia. Br. J. Dermatol. 2015, 174, 439–443. [Google Scholar] [CrossRef] [PubMed]
- Hirai, Y.; Kodama, Y.; Moriwaki, S.-I.; Noda, A.; Cullings, H.M.; Macphee, D.G.; Kodama, K.; Mabuchi, K.; Kraemer, K.H.; Land, C.E.; et al. Heterozygous individuals bearing a founder mutation in the XPA DNA repair gene comprise nearly 1% of the Japanese population. Mutat. Res. Mol. Mech. Mutagen. 2006, 601, 171–178. [Google Scholar] [CrossRef]
- Temtamy, S.A.; Aglan, M.S.; Meguid, N.A. Genetic Disorders in Egypt. In Genetic Disorders Among Arab Populations; Teebi, A.S., Ed.; Springer: Berlin/Heidelberg, Germany, 2010; pp. 219–272. [Google Scholar]
- Shawky, R.M.; El-Awady, M.Y.; Elsayed, S.M.; Hamadan, G.E. Consanguineous matings among Egyptian population. Egypt. J. Med. Hum. Genet. 2011, 12, 157–163. [Google Scholar] [CrossRef]
- Hashem, N.; Bootsma, D.; Keijzer, W.; Greene, A.; Coriell, L.; Thomas, G.; Cleaver, J.E. Clinical characteristics, DNA repair, and complementation groups in xeroderma pigmentosum patients from Egypt. Cancer Res. 1980, 40, 13–18. [Google Scholar] [PubMed]
- Cleaver, J.E.; Zelle, B.; Hashem, N.; El-Hefnawi, M.H.; German, J. Xeroderma pigmentosum patients from Egypt: II. Preliminary correlations of epidemiology, clinical symptoms and molecular biology. J. Investig. Dermatol. 1981, 77, 96–101. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ridley, A.J.; Colley, J.; Wynford-Thomas, D.; Jones, C.J. Characterisation of novel mutations in Cockayne syndrome type A and xeroderma pigmentosum group C subjects. J. Hum. Genet. 2005, 50, 151–154. [Google Scholar] [CrossRef]
- Soufir, N.; Ged, C.; Bourillon, A.; Austerlitz, F.; Chemin, C.; Stary, A.; Armier, J.; Pham, D.; Khadir, K.; Roume, J.; et al. A prevalent mutation with founder effect in xeroderma pigmentosum group C from North Africa. J. Investig. Dermatol. 2010, 130, 1537–1542. [Google Scholar] [CrossRef]
- Amr, K.; Messaoud, O.; El Darouti, M.; Abdelhak, S.; El-Kamah, G. Mutational spectrum of Xeroderma pigmentosum group A in Egyptian patients. Gene 2014, 533, 52–56. [Google Scholar] [CrossRef]
- Sarasin, A.; Munier, P.; Cartault, F. How history and geography may explain the distribution in the Comorian archipelago of a novel mutation in DNA repair-deficient xeroderma pigmentosum patients. Genet. Mol. Biol. 2020, 43, e20190046. [Google Scholar] [CrossRef] [PubMed]
- Romdhane, L.; Mezzi, N.; Hamdi, Y.; El-Kamah, G.; Barakat, A.; Abdelhak, S. Consanguinity and inbreeding in health and disease in North African populations. Annu. Rev. Genom. Hum. Genet. 2019, 20, 155–179. [Google Scholar] [CrossRef] [PubMed]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015, 17, 405–423. [Google Scholar] [CrossRef] [PubMed]
- Satokata, I.; Tanaka, K.; Okada, Y. Molecular basis of group A xeroderma pigmentosum: A missense mutation and two deletions located in a zinc finger consensus sequence of the XPAC gene. Qual. Life Res. 1992, 88, 603–607. [Google Scholar] [CrossRef]
- Satokata, I.; Tanaka, K.; Miura, N.; Narita, M.; Mimaki, T.; Satoh, Y.; Kondo, S.; Okada, Y. Three nonsense mutations responsible for group A xeroderma pigmentosum. Mutat. Res. Repair 1992, 273, 193–202. [Google Scholar] [CrossRef]
- Chavanne, F.; Broughton, B.C.; Pietra, D.; Nardo, T.; Browitt, A.; Lehmann, A.R.; Stefanini, M. Mutations in the XPC gene in families with xeroderma pigmentosum and consequences at the cell, protein, and transcript levels. Cancer Res. 2000, 60, 1974–1982. [Google Scholar] [PubMed]
- Li, L.; Bales, E.S.; Peterson, C.A.; Legerski, R.J. Characterization of molecular defects in xeroderma pigmentosum group C. Nat. Genet. 1993, 5, 413–417. [Google Scholar] [CrossRef] [PubMed]
- Cartault, F.; Nava, C.; Malbrunot, A.-C.; Munier, P.; Hebert, J.-C.; N’Guyen, P.; Djeridi, N.; Pariaud, P.; Pariaud, J.; Dupuy, A.; et al. A new XPC gene splicing mutation has lead to the highest worldwide prevalence of xeroderma pigmentosum in black Mahori patients. DNA Repair 2011, 10, 577–585. [Google Scholar] [CrossRef]
- Schaefer, A.; Hofmann, L.; Gratchev, A.; Laspe, P.; Schubert, S.; Schuerer, A.; Ohlenbusch, A.; Tzvetkov, M.; Hallermann, C.; Reichrath, J.; et al. Molecular genetic analysis of 16 XP-C patients from Germany: Environmental factors predominately contribute to phenotype variations. Exp. Dermatol. 2013, 22, 24–29. [Google Scholar] [CrossRef] [PubMed]
- Senhaji, M.A.; Abidi, O.; Nadifi, S.; Benchikhi, H.; Khadir, K.; Ben Rekaya, M.; Eloualid, A.; Messaoud, O.; Abdelhak, S.; Barakat, A. c.1643_1644delTG XPC mutation is more frequent in Moroccan patients with xeroderma pigmentosum. Arch. Dermatol. Res. 2012, 305, 53–57. [Google Scholar] [CrossRef] [PubMed]
- Ben Rekaya, M.; Jerbi, M.; Messaoud, O.; Ben Brick, A.S.; Zghal, M.; Mbarek, C.; Chadli-Debbiche, A.; Jones, M.; Mokni, M.; Boussen, H.; et al. Further evidence of mutational heterogeneity of the XPC gene in Tunisian families: A spectrum of private and ethnic specific mutations. BioMed Res. Int. 2013, 2013. [Google Scholar] [CrossRef] [PubMed]
- Bensenouci, S.; Louhibi, L.; De Verneuil, H.; Mahmoudi, K.; Saïdi-Mehtar, N. Diagnosis of Xeroderma pigmentosum groups A and C by detection of two prevalent mutations in West Algerian population: A rapid genotyping tool for the frequent XPC mutation c.1643_1644delTG. BioMed Res. Int. 2016, 2016, 2180946. [Google Scholar] [CrossRef] [PubMed]
- Mahindra, P.; DiGiovanna, J.J.; Tamura, D.; Brahim, J.S.; Hornyak, T.J.; Stern, J.B.; Lee, C.-C.R.; Khan, S.G.; Brooks, B.P.; Smith, J.A.; et al. Skin cancers, blindness, and anterior tongue mass in African brothers. J. Am. Acad. Dermatol. 2008, 59, 881–886. [Google Scholar] [CrossRef]
- El-Harith, E.A.; Pahl, L.; Al-Nutaifi, K.; Bukhari, I.; Schmidtke, J.; Stuhrmann, M. Diagnosis of xeroderma pigmentosum C by detection of the founder mutation c.1643_1644delTG (p. Val548Ala fsX25) in a Sudanese family. J. Saudi Soc. Dermatol. Dermatol. Surg. 2012, 16, 85–86. [Google Scholar] [CrossRef][Green Version]
- Santiago, K.; Castro, L.; Neto, J.; De Nóbrega, A.; Pinto, C.; Ashton-Prolla, P.; Pinto e Vairo, E.; De Medeiros, P.; Ribeiro, E.; Ribeiro, B.; et al. Comprehensive germline mutation analysis and clinical profile in a large cohort of Brazilian xeroderma pigmentosum patients. J. Eur. Acad. Dermatol. Venereol. 2020, 34, 2392–2401. [Google Scholar] [CrossRef]
- Khan, S.G. The human XPC DNA repair gene: Arrangement, splice site information content and influence of a single nucleotide polymorphism in a splice acceptor site on alternative splicing and function. Nucleic Acids Res. 2002, 30, 3624–3631. [Google Scholar] [CrossRef] [PubMed]
- Kgokolo, M.; Morice-Picard, F.; Rezvani, H.R.; Austerlitz, F.; Cartault, F.; Sarasin, A.; Sathekge, M.; Taieb, A.; Ged, C. Xeroderma pigmentosum in South Africa: Evidence for a prevalent founder effect. Br. J. Dermatol. 2019, 181, 1070–1072. [Google Scholar] [CrossRef]
- Ijaz, A.; Basit, S.; Gul, A.; Batool, L.; Hussain, A.; Afzal, S.; Ramzan, K.; Ahmad, J.; Wali, A. XPC gene mutations in families with xeroderma pigmentosum from Pakistan; prevalent founder effect. Congenit. Anom. 2019, 59, 18–21. [Google Scholar] [CrossRef] [PubMed]
- Coulombe, J.; Orbach, D.; Soufir, N.; Hadj-Rabia, S. Primary gingival squamous cell carcinoma in a xeroderma pigmentosum type C patient. J. Eur. Acad. Dermatol. Venereol. 2016, 30, e157–e158. [Google Scholar] [CrossRef]
- Jacyk, W.K. Xeroderma pigmentosum in black South Africans. Int. J. Dermatol. 1999, 38, 511–514. [Google Scholar] [CrossRef] [PubMed]
- Brenner, M.; Hearing, V.J. The protective role of melanin against UV damage in human skin. Photochem. Photobiol. 2007, 84, 539–549. [Google Scholar] [CrossRef]
- Gozukara, E.M.; Khan, S.G.; Metin, A.; Emmert, S.; Busch, D.B.; Shahlavi, T.; Coleman, D.M.; Miller, M.; Chinsomboon, N.; Stefanini, M.; et al. A stop codon in xeroderma pigmentosum group C families in Turkey and Italy: Molecular genetic evidence for a common ancestor. J. Investig. Dermatol. 2001, 117, 197–204. [Google Scholar] [CrossRef][Green Version]
- De Jesus, B.M.B.; Bjørås, M.; Coin, F.; Egly, J.M. Dissection of the molecular defects caused by pathogenic mutations in the DNA repair factor XPC. Mol. Cell. Biol. 2008, 28, 7225–7235. [Google Scholar] [CrossRef] [PubMed]
- Stenson, P.D.; Mort, M.; Ball, E.V.; Shaw, K.; Phillips, A.D.; Cooper, D.N. The human gene mutation database: Building a comprehensive mutation repository for clinical and molecular genetics, diagnostic testing and personalized genomic medicine. Qual. Life Res. 2014, 133, 1–9. [Google Scholar] [CrossRef]
- Khan, S.G.; Oh, K.-S.; Emmert, S.; Imoto, K.; Tamura, D.; DiGiovanna, J.J.; Shahlavi, T.; Armstrong, N.; Baker, C.C.; Neuburg, M.; et al. XPC initiation codon mutation in xeroderma pigmentosum patients with and without neurological symptoms. DNA Repair 2009, 8, 114–125. [Google Scholar] [CrossRef] [PubMed]
- Khan, S.G.; Oh, K.-S.; Shahlavi, T.; Ueda, T.; Busch, D.B.; Inui, H.; Emmert, S.; Imoto, K.; Muniz-Medina, V.; Baker, C.C.; et al. Reduced XPC DNA repair gene mRNA levels in clinically normal parents of xeroderma pigmentosum patients. Carcinogenesis 2005, 27, 84–94. [Google Scholar] [CrossRef]
- Khan, S.G. Two essential splice lariat branchpoint sequences in one intron in a xeroderma pigmentosum DNA repair gene: Mutations result in reduced XPC mRNA levels that correlate with cancer risk. Hum. Mol. Genet. 2003, 13, 343–352. [Google Scholar] [CrossRef]
- Khan, S.G.; Levy, H.L.; Legerski, R.; Quackenbush, E.; Reardon, J.T.; Emmert, S.; Sancar, A.; Li, L.; Schneider, T.D.; Cleaver, J.E.; et al. Xeroderma pigmentosum group C splice mutation associated with autism and hypoglycinemia. J. Investig. Dermatol. 1998, 111, 791–796. [Google Scholar] [CrossRef] [PubMed]
- Maquat, L.E. Nonsense-mediated mRNA decay in mammals. J. Cell Sci. 2005, 118, 1773–1776. [Google Scholar] [CrossRef]
- Messaoud, O.; Ben Rekaya, M.; Ouragini, H.; Benfadhel, S.; Azaiez, H.; Kefi, R.; Gouider-Khouja, N.; Mokhtar, I.; Amouri, A.; Boubaker, M.S.; et al. Severe phenotypes in two Tunisian families with novel XPA mutations: Evidence for a correlation between mutation location and disease severity. Arch. Dermatol. Res. 2012, 304, 171–176. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, Y.; Endo, Y.; Sugiyama, Y.; Inoue, S.; Iijima, M.; Tomita, Y.; Kuru, S.; Takigawa, M.; Moriwaki, S. XPA gene mutations resulting in subtle truncation of protein in xeroderma pigmentosum group A patients with mild skin symptoms. J. Investig. Dermatol. 2010, 130, 2481–2488. [Google Scholar] [CrossRef]
- States, J.; McDuffie, E.; Myrand, S.; McDowell, M.; Cleaver, J. Distribution of mutations in the human xero-derma pigmentosum group A gene and their relationships to the functional regions of the DNA damage recognition protein. Hum. Mutat. 1998, 12, 103–113. [Google Scholar] [CrossRef]
- Bartels, C.L.; Lambert, M.W. Domains in the XPA protein important in its role as a processivity factor. Biochem. Biophys. Res. Commun. 2007, 356, 219–225. [Google Scholar] [CrossRef] [PubMed]
- Santiago, K.M.; França de Nóbrega, A.; Rocha, R.M.; Rogatto, S.R.; Achatz, M.I. Xeroderma pigmentosum: Low prevalence of germline XPA mutations in a brazilian XP population. Int. J. Mol. Sci. 2015, 16, 8988–8996. [Google Scholar] [CrossRef] [PubMed]
- Alberto, F.; Figueiredo, M.; Zago, M.; Araújo, A.; Dos-Santos, J. The lebanese mutation as an important cause of familial hypercholesterolemia in Brazil. Braz. J. Med. Biol. Res. 1999, 32, 739–745. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Alagoz, M.; Kherad, N. Advance genome editing technologies in the treatment of human diseases: CRISPR therapy (Review). Int. J. Mol. Med. 2020, 46, 521–534. [Google Scholar] [CrossRef] [PubMed]
| Exon | Forward Primer | Amplicon Size (bp) | Annealing Temperature (°C) |
|---|---|---|---|
| Reverse Primer | |||
| XPA 1 | 5′-GGAGTGGGCCAGAGATGG-3′ | 300 | 61 |
| 5′-ATACGCCAGCGGAGTTGAC-3′ | |||
| XPA 2 | 5′-TTGTGGACATCCTTGTGTTG-3′ | 319 | 59 |
| 5′-GGCATTATTTAGCATCACTTTGC-3′ | |||
| XPA 3 | 5′-AGGCATTGCATACATGCTG-3′ | 373 | 58 |
| 5′-AAGCACAGATTTACAGTATTTGGC-3′ | |||
| XPA 4 | 5′-CTGTGTGTGCCCCTAAGTTG-3′ | 342 | 59 |
| 5′-TCTGTAAGCAAAAGCCAAACC-3′ | |||
| XPA 5 | 5′-TGGTACCTTTGGATTTGACAG-3′ | 368 | 59 |
| 5′-TGCCTTGAAGACCAACATACTG-3′ | |||
| XPA 6 | 5′-ACATGGCTGAAAGCTTGATG-3′ | 423 | 59 |
| 5′-CCAGGTGACCTTCACTGAAAC-3′ | |||
| XPC 1 | 5′-TTGTGCTCTTTCCTGCTTCCC-3′ | 372 | 61 |
| 5′- TCTGGACTCCGCCCTGCCTCTG-3′ | |||
| XPC 2 | 5′-ATAGAGCCGTTTTATGCCCC-3′ | 381 | 60 |
| 5′-TGGACCCCAGTGACAAGTAAG-3′ | |||
| XPC 3 | 5′-GTTGATGGAGGAAGTGAGGC-3′ | 346 | 60 |
| 5′-TCTGACTCCAAACAGAATCAAAC-3′ | |||
| XPC 4 | 5′-TGTCTAGGGGTCTCTGTGGG-3′ | 356 | 60 |
| 5′-TGGTCCCCTACAAGTTTCTCC-3′ | |||
| XPC 5 | 5′-AGGAAATAGCTGGCTTGCAG-3′ | 323 | 60 |
| 5′-CCAGAAATAAAGCCTCGGTG-3′ | |||
| XPC 6 | 5′-CATGTCTTGACTTTGGCAGC-3′ | 336 | 60 |
| 5′-TCAGGGAAGGTCTGTGGAAG-3′ | |||
| XPC 7 | 5′-AGTTAGCTAGACGGGCTGGG-3′ | 352 | 60 |
| 5′-AACACACCTGGAATGGCATC-3′ | |||
| XPC 8 | 5′-CTGGCTGTTTCCAGCTTTTC-3′ | 326 | 55 |
| 5′-GCTCGAAAGAACCCACACTC-3′ | |||
| XPC 9a | 5′-CAACCCTGAAGGATAGCTGG-3′ | 662 | 60 |
| 5′-AAGCTTGGGTCCTTACGATG-3′ | |||
| XPC 9b | 5′-GAGAGTGGGAGTGATGAGGC-3′ | 661 | 60 |
| 5′-GCTGGGCATATATAAGGTGCTC-3′ | |||
| XPC 10 | 5′-GTTCATGAAACCTTGGCTCC-3′ | 396 | 59 |
| 5′-CCGAGAATGCTGTCCAGTC-3′ | |||
| XPC 11 | 5′-CTAGCACAGCTTCTCTGGGC-3′ | 331 | 60 |
| 5′-GGGAGGCTCATCATCACTTC-3′ | |||
| XPC 12–13 | 5′-TGGTAGGTGTGTTCTGAGGG-3′ | 629 | 59 |
| 5′-CTGAAAATTGGAGCCACCAG-3′ | |||
| XPC 14 | 5′-AGATGTGGCCCACTGTCTTC-3′ | 310 | 60 |
| 5′-ATGATGTCAGAGAGGGCTGG-3′ | |||
| XPC 15 | 5′-GAGACTTGGTGTGAAGGAGAGG-3′ | 336 | 60 |
| 5′-AAGCTATCCCTGACTTGAGGC-3′ | |||
| XPC 16 | 5′-AACTTGCTGCCTCTTCATGG-3′ | 467 | 60 |
| 5′-TCAGCTTGGCCTCGTCTC-3′ |
| Fam | P | Sex | XP gp | Age of Onset | Age at Examin-Ation | Skin | Ocular | Malignancies | Neurological Symptoms | Others | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Photo-Sensitivity | Xerosis | Freckles/Lentigines | Poikilo-Derma | Others | Photo-Phobia | Conjunc-Tivitis | Keratitis | Others | Skin | Eye | Others | |||||||||||
| Face & Extremities | Trunk | Micro-Cephaly | ID | Cerebellar Hypoplasia | ||||||||||||||||||
| 1 | XP5GI | F | A | 4 m | 7 y | + | - | - | - | - | Actinic keratosis | + | - | - | Conjunctival papules | + BCC, SCC | + SCC | - | + | + | + | Neuropathy |
| 1 | XP6GI | F | A | 4 m | 4 y 3 m | - | + | - | - | - | - | - | - | - | - | - | - | - | + | + | + | - |
| 2 | XP7GI | F | A | 6 m | 2 y 3 m | + | + | + | - | + | - | - | + | - | - | - | - | - | + | + | + | Hyperreflexia, moderate to severe hearing loss |
| 3 | XP8GI | F | A | 6 m | 4 y | + | + | + | - | + | - | + | - | + | Conjunctival papules | - | - | - | + | + | + | |
| 4 | XP9GI | M | A | 5 m | 6 y 7 m | + | + | + | - | - | - | - | - | - | - | - | - | + | + | - | Delayed Milestones | |
| 5 | XP10GI | M | A | 3 m | 3 y | + | - | - | - | - | - | - | + | - | - | - | - | - | + | - | - | |
| 5 | XP11GI | M | A | 3 m | 4 y | + | - | - | - | - | - | - | - | - | - | - | - | - | + | - | - | |
| 6 | XP12GI | M | C | 6 m | 7 y | + | + | + | - | + | - | + | + | + | - | + + BCC, SCC | - | - | - | - | - | |
| 7 | XP13GI | F | C | 2 y | 16 y | + | + | + | + | - | Telangiectasia | + | + | + | Right eye enucleation | ++ SCC | ++ BCC | BCC of the tongue | - | - | - | |
| 7 | XP14GI | M | C | 8 m | 12 y | + | + | + | + | + | - | + | + | + | Left and right eyes enucleation | + + + | + + + BCC, SCC | - | - | - | - | |
| 8 | XP15GI | F | C | 8 m | 6 y | + | + | + | - | + | - | + | + | - | - | - | - | small submand-ibular tumor | - | - | - | |
| 9 | XP16GI | F | C | 8 m | 10 y 5 m | + | + | + | - | + | - | + | + | - | - | ++ BCC, SCC | + | - | - | - | - | |
| 9 | XP17GI | F | C | 2 y | 12 y | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | |
| 10 | XP18GI | M | C | 6 m | 8 y | + | + | + | - | + | - | + | + | + | Corneal opacities | + + BCC on nose &scalp | + | - | - | - | - | |
| 11 | XP19GI | M | C | 2 y | 32 y | + | + | + | - | + | - | + | + | + | - | + + + BCC, SCC, Nose & left eyelid | - | Melanoma | - | - | - | |
| 12 | XP20GI | F | C | 8 m | 10 y 6 m | + | + | + | - | + | - | + | + | + | - | + + BCC on nose & scalp | + | - | - | - | - | |
| 13 | XP21GI | F | C | 8 m | 10 y 6 m | + | + | + | - | + | - | + | + | + | - | + + BCC, SCC | + | - | - | - | - | |
| 14 | XP22GI | F | C | 2 y | 15 y | + | - | + | - | + | Multiple nodules on face and non-healing ulcer | + | + | - | - | + BCC on nose | - | - | - | - | - | |
| 14 | XP23GI | M | C | 8 m | 11 y | + | - | + | - | + | - | + | + | - | - | - | - | - | - | - | - | |
| 14 | XP24GI | M | C | 2 y | 6 y | - | - | +, face only | - | - | - | - | - | - | - | - | - | - | - | - | - | |
| 15 | XP25GI | F | C | 2 y | 10 y | - | - | + | - | + | - | + | + | - | Pterygium | - | - | - | - | - | - | |
| 15 | XP26GI | F | C | 2 y | 11 y | - | - | + | - | + + | - | + | + | - | - | - | - | - | - | - | - | |
| 15 | XP27GI | F | C | 1 y 6 m | 20 y | - | - | + | - | + | - | + | + | - | Nystagmus and severe myopia (−18.5) | - | - | - | - | - | - | |
| 16 | XP28GI | F | C | 2 y 6 m | 20 y | - | - | + | - | + | - | - | + | - | - | - | - | - | - | - | - | |
| 17 | XP29GI | M | C | 2 y | 8 y | + | - | + | + | - | - | - | - | - | - | - | - | - | - | - | - | |
| 17 | XP30GI | M | C | N/A | 2 m | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | |
| 18 | XP31GI | M | C | 9 m | 25 y | + | - | +, extre-mities only | - | + | - | + | + | - | Fibroses of conjunctiva and lower lid | + + + SCC | - | - | - | - | - | Hyperreflexia, patellar and ankle clonus |
| 19 | XP32GI | M | C | 6 m | 4 y | - | - | + | - | + | - | + | - | - | Progressive loss of vision | - | - | - | - | - | - | |
| 20 | XP33GI | M | C | 2 y | 6 y 2 m | + | - | + | + | + | Granuloma | - | - | - | - | + BCC | - | - | - | - | - | |
| 21 | XP34GI | M | C | 11 m | 12 y 4 m | + | + | + | - | - | Actinic keratosis | + | + | + | Corneal opacities | + + + BCC | - | - | - | - | - | |
| 22 | XP35GI | F | C | 1 y | 8 y | + | + | + | - | - | Multiple nodules on face & scalp | + | + | - | - | BCC on nose & right eyelid | - | - | - | - | - | |
| 23 | XP36GI | F | C | 6 m | 9 y | + | + | + | - | + | - | + | + | + | Ectropion of left lower lid | - | + SCC | - | - | - | - | |
| 23 | XP37GI | F | C | 6 m | 5 y 6 m | + | + | + | - | + | Papules on left cheeks | + | + | + | - | - | - | - | - | - | - | |
| 24 | XP38GI | F | C | 6 m | 4 y | - | + | + | - | + | + Skin papules on forehead, nose, and upper lip | + | + | + | - | + + + BCC, SCC | - | - | - | - | - | |
| 25 | XP39GI | F | C | 3 y | 25 y | + | + | + | + | + | - | + | + | + | Corneal opacities | + + + BCC, SCC | - | - | - | - | - | |
| 26 | XP40GI | F | C | 1 y | 1 y 3 m | - | + | + | + | + | - | - | - | - | - | - | - | - | - | - | - | |
| Family | Patient | Gene | Exon (E) or Intron (IVS) | Mutation | Type of mutation | Pathogenicity c | Genotype | Reference d | Number of Heterozygous Carriers/Screened Family Members e | |
|---|---|---|---|---|---|---|---|---|---|---|
| Nucleotide Change a | Protein or mRNA Changes b | |||||||||
| 1 | XP5GI&XP6GI | XPA | E3 | c.374delC | p.Thr125Ilefs*15 | Frameshift/ Small deletion | Pathogenic | Homozygous | [38] | 2/2 |
| 2 | XP7GI | XPA | E4 | c.553C>T | p.Gln185* | Nonsense | Pathogenic | Homozygous | [34] | 4/5 |
| 3–5 | XP8GI-XP11GI | XPA | E5 | c.619C>T | p.Arg207* | Nonsense | Pathogenic | Homozygous | [39] | 6/8 |
| 6 | XP12GI | XPC | E3 | c.395-398delATTG | p.Asp132Glyfs*15 | Frameshift/ Small deletion | Pathogenic | Homozygous | Current study | 2/2 |
| 7 | XP13GI& XP14GI | XPC | E6 | c.668-669delTC | p.Ile223Metfs*45 | Frameshift/ Small deletion | Pathogenic | Homozygous | Current study | 3/3 |
| 8 | XP15GI | XPC | E4 | c.525_526insCA | p.Arg176Glnfs*8 | Frameshift/ Small insertion | Pathogenic | Compound heterozygous | Current study | 1/2 |
| E9 | c.1103-1104delAA | p.Gln368Argfs*6 | Frameshift/ Small deletion | Pathogenic | [40] | 1/2 | ||||
| 9 | XP16GI& XP17GI | XPC | E9 | c.1735C>T | p.Arg579* | Nonsense | Pathogenic | Homozygous | [40] | 4/5 |
| 10–19 | XP18GI- XP32GI | XPC | E9 | c.1643-1644delTG | p.Val548Alafs*25 | Frameshift/ Small deletion | Pathogenic | Homozygous | [41] | 23/24 |
| 20 | XP33GI | XPC | E9 | c.1615G>T | p.Glu539* | Nonsense | Pathogenic | Compound heterozygous | Current study | 1/1 |
| E9 | c.1643-1644delTG | p.Val548Alafs*25 | Frameshift/ Small deletion | Pathogenic | [41] | 1/1 | ||||
| 21–22 | XP34GI& XP35GI | XPC | E10 | c.1894C>T | p.Gln632* | Nonsense | Pathogenic | Homozygous | Current study | 5/5 |
| 23–25 | XP36GI-XP39GI | XPC | IVS12 | c.2251-1G>C | Three abnormally spliced mRNA: exon 13 skipping, intron 12 retention, and 44 bp deletion in exon 13 | Splicing | Pathogenic | Homozygous | [42] | 6/6 |
| 26 | XP40GI | XPC | E12 | c.2127-2127delC | p.Ser711Leufs*56 | Frameshift/Small deletion | Pathogenic | Homozygous | Current study | 2/2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rabie, E.; Amr, K.; Zada, S.; El-Sayed, H.; El Darouti, M.; El-Kamah, G. Clinical and Mutational Spectrum of Xeroderma Pigmentosum in Egypt: Identification of Six Novel Mutations and Implications for Ancestral Origins. Genes 2021, 12, 295. https://doi.org/10.3390/genes12020295
Rabie E, Amr K, Zada S, El-Sayed H, El Darouti M, El-Kamah G. Clinical and Mutational Spectrum of Xeroderma Pigmentosum in Egypt: Identification of Six Novel Mutations and Implications for Ancestral Origins. Genes. 2021; 12(2):295. https://doi.org/10.3390/genes12020295
Chicago/Turabian StyleRabie, Eman, Khalda Amr, Suher Zada, Heba El-Sayed, Mohamad El Darouti, and Ghada El-Kamah. 2021. "Clinical and Mutational Spectrum of Xeroderma Pigmentosum in Egypt: Identification of Six Novel Mutations and Implications for Ancestral Origins" Genes 12, no. 2: 295. https://doi.org/10.3390/genes12020295
APA StyleRabie, E., Amr, K., Zada, S., El-Sayed, H., El Darouti, M., & El-Kamah, G. (2021). Clinical and Mutational Spectrum of Xeroderma Pigmentosum in Egypt: Identification of Six Novel Mutations and Implications for Ancestral Origins. Genes, 12(2), 295. https://doi.org/10.3390/genes12020295






