Impact of Pre- and Post-Processing Steps for Supervised Classification of Colorectal Cancer in Hyperspectral Images
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Patient Data
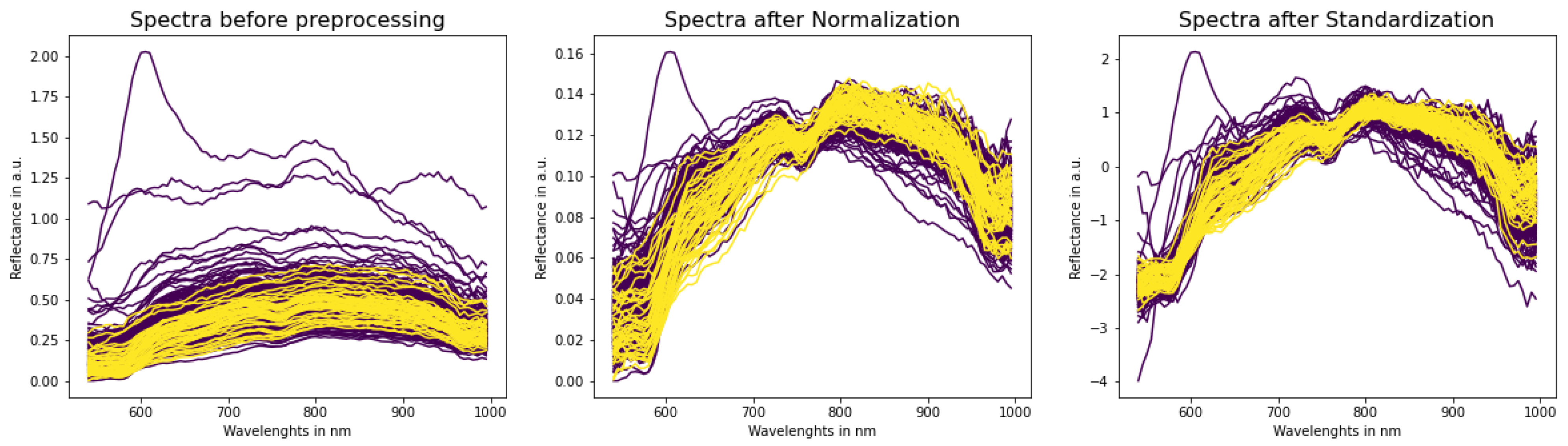
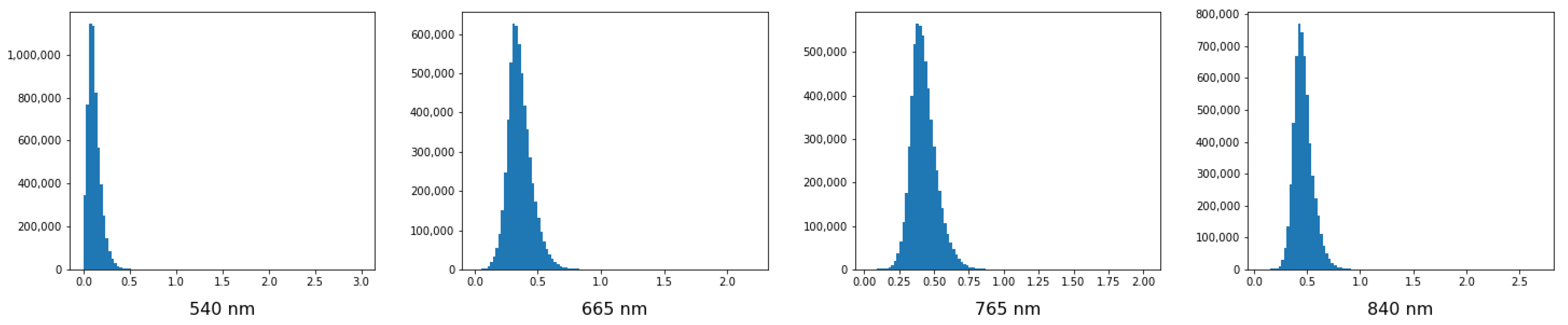
2.2. Pre-Processing
2.3. Supervised Binary Classification
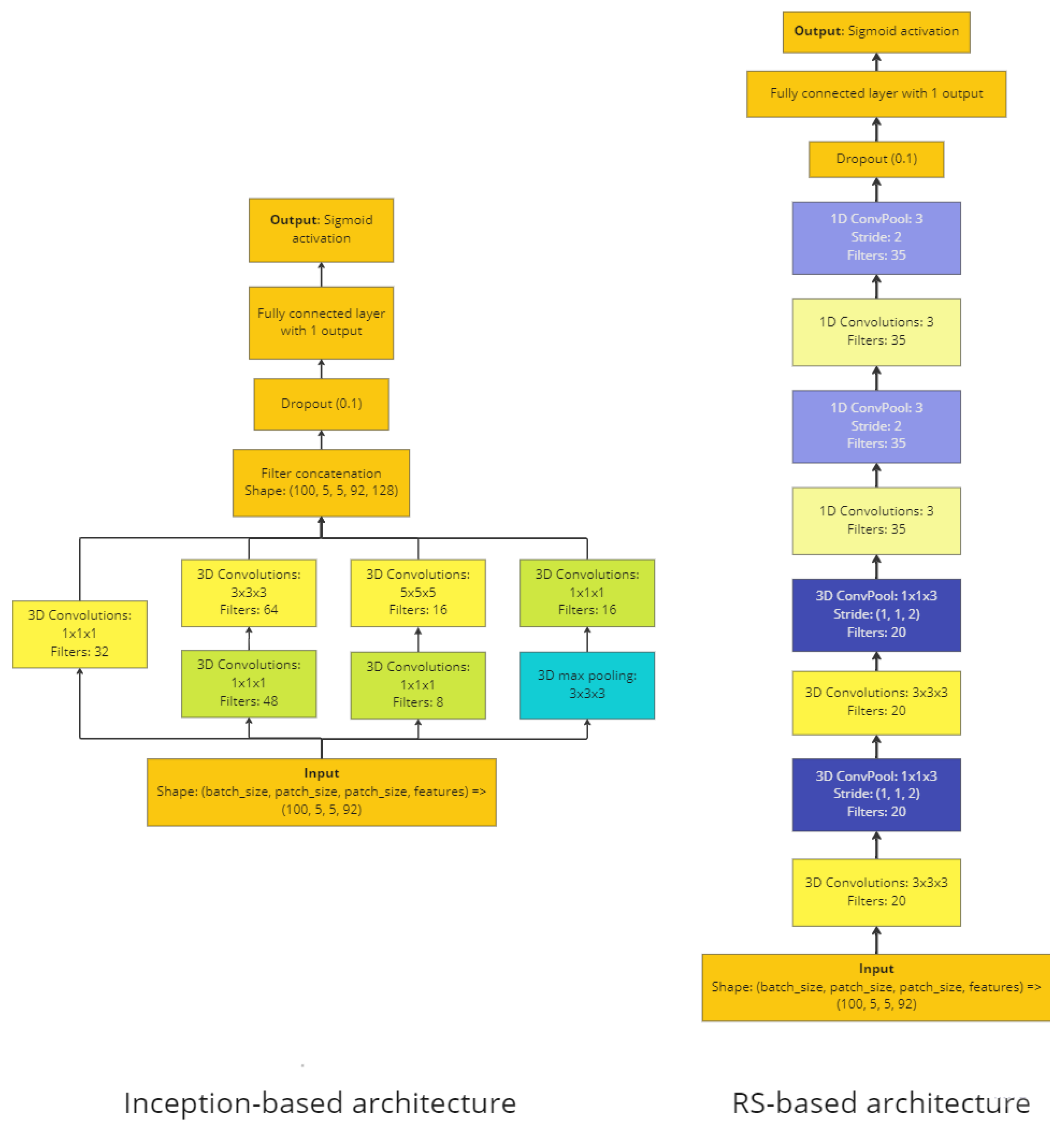
2.3.1. Architecture 1: 3D-CNN with Inception Architecture
2.3.2. Architecture 2: RS-Based 3D-CNN
2.3.3. Training Parameters
2.3.4. Thresholding
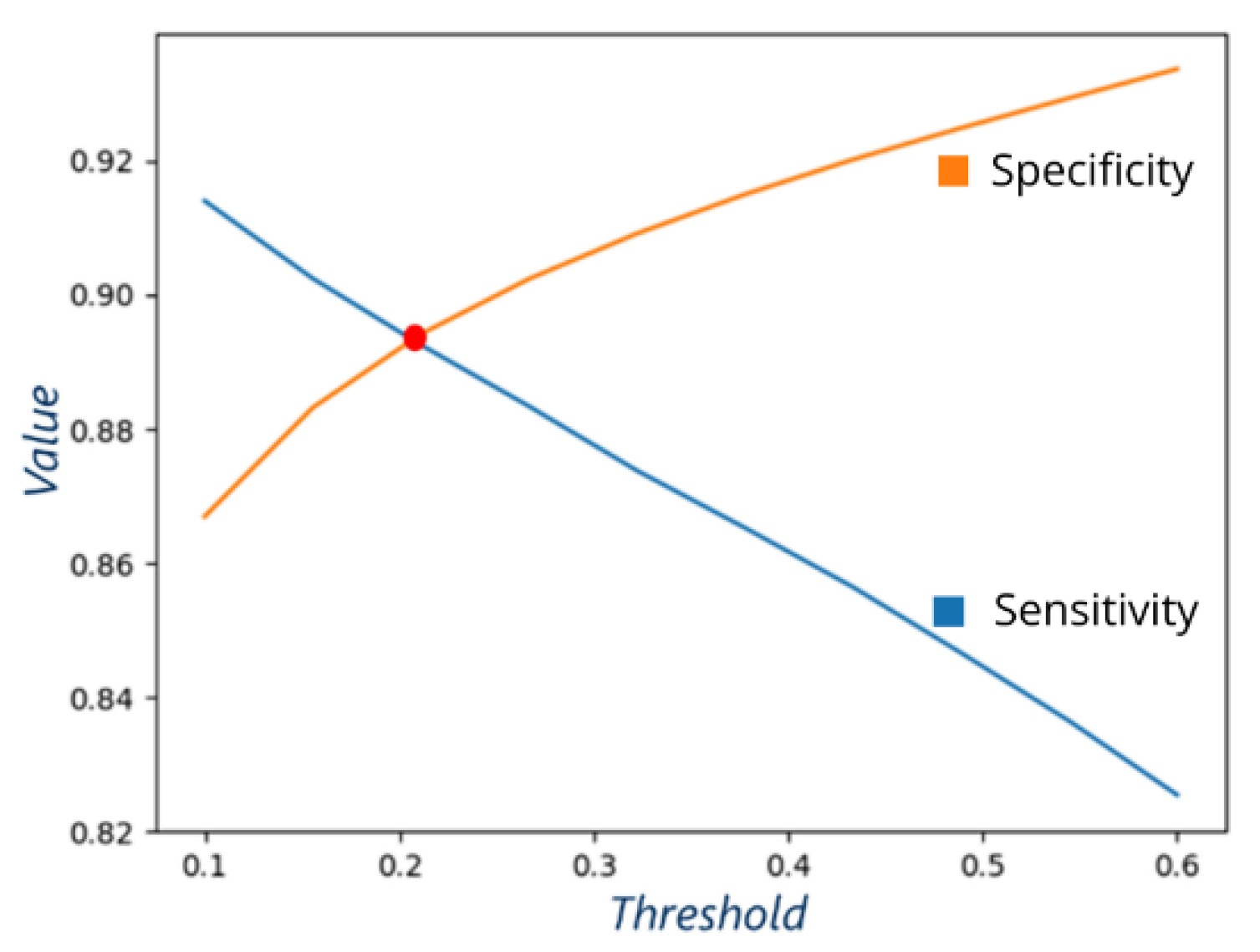
2.3.5. Choosing the Best Threshold
- A threshold where sensitivity and specificity have similar values allows for easier comparison between different models: both sensitivity and specificity are either better or worse.
- This approach allows balancing between false negatives and false positives.
2.4. Post-Processing
2.4.1. Definitions
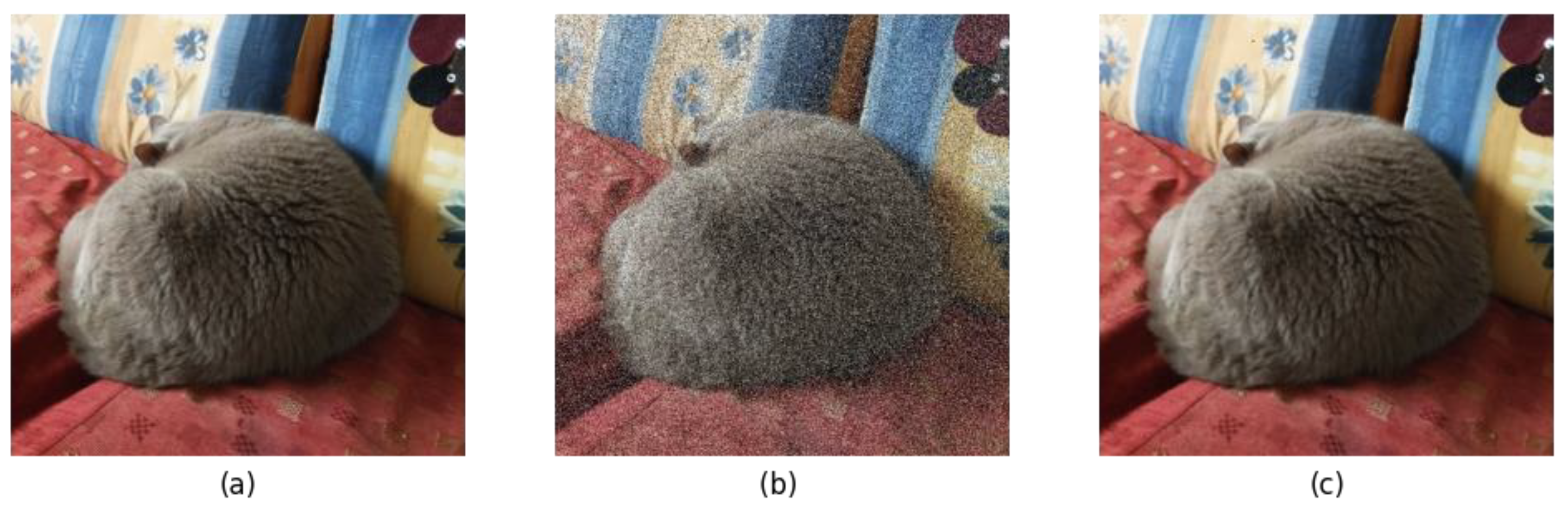
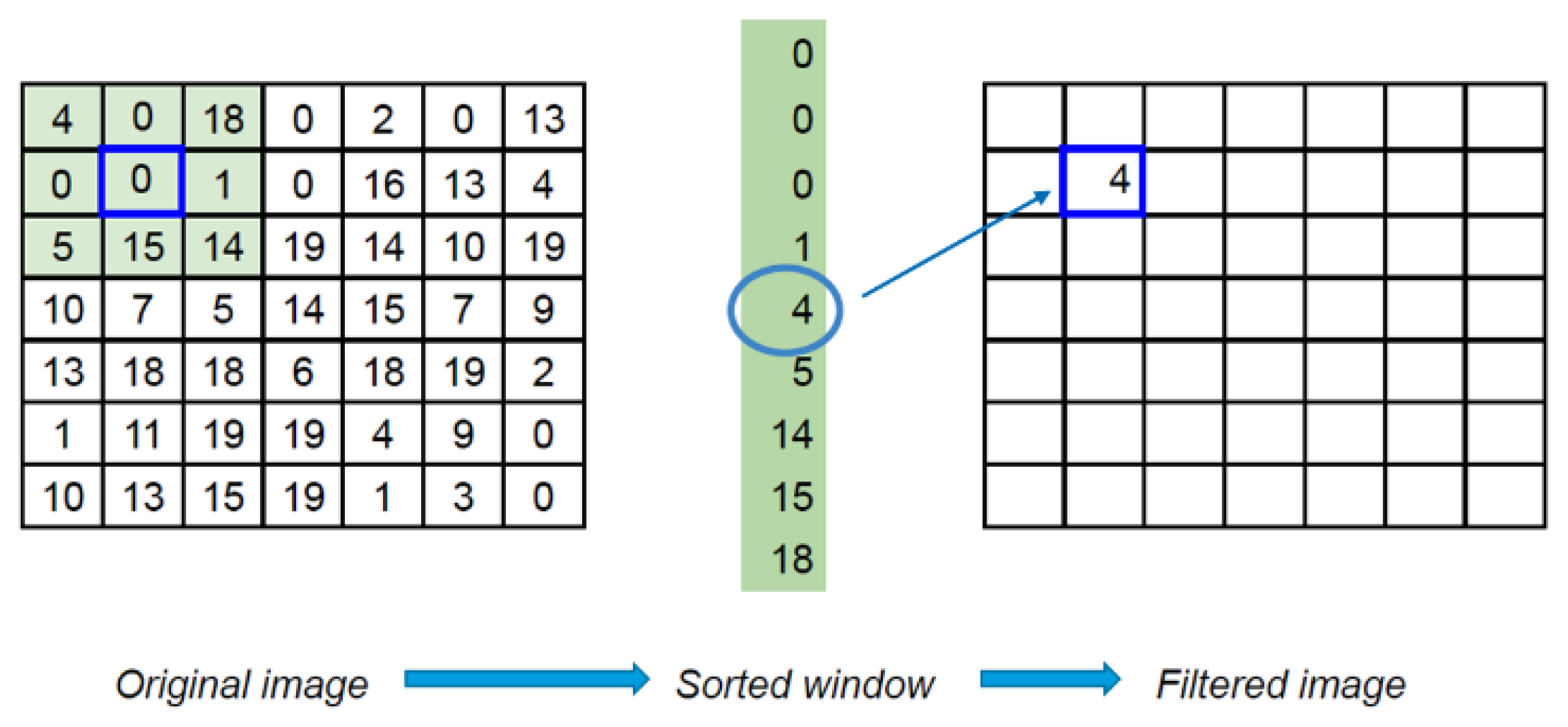
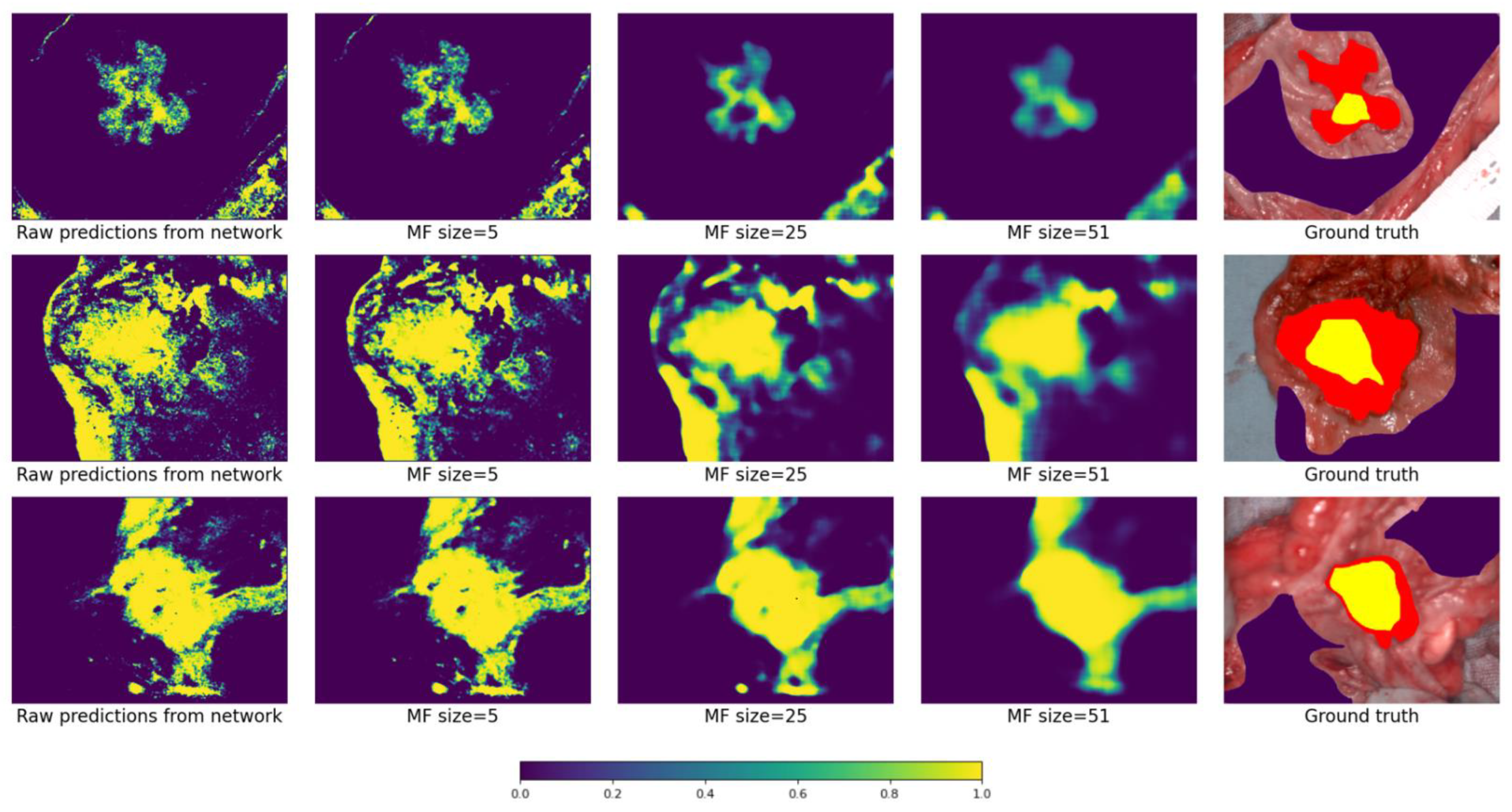
2.4.2. Using Median Filter (MF)
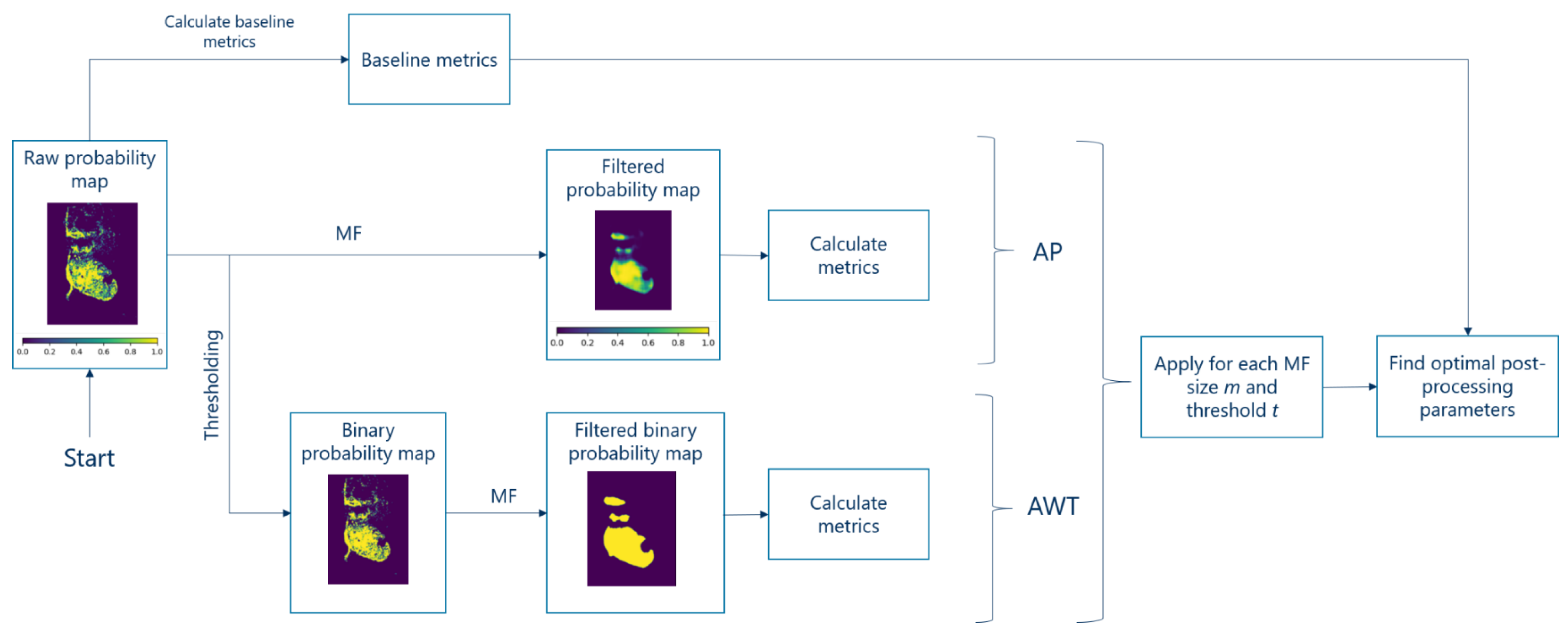
2.4.3. Post-Processing Methods: The Plain Algorithm (AP) and the Algorithm with Threshold (AWT)
2.4.4. Finding Optimal Parameters of Post-Processing and Estimation of the Performance Improvement
| Algorithm 1. Finding optimal parameters of post-processing. | |
| Inputs: | |
| sensbaseline, specbaseline | (Section 2.3.5) |
| MF_sizes_to_test | |
| thresholds_to_test | |
| Initialize: | |
| improvementsi,j ← 0, where i = 1, …, length(MF_sizes_to_test) | |
| j = 1, …, length(thresholds_to_test) | |
| Result: optimal MF size mopt, optimal threshold size topt | |
| for each m in MF_sizes_to_test do | |
| for each t in thresholds_to_test do | |
| apply MF size m and threshold t to prediction maps | |
| calculate sensm,t and specm,t | |
| improvementspec← specm,t – specbaseline | (3) |
| improvementsens ← sensm,t – sensbaseline | (4) |
| if improvementsens > 0 and improvementspec > 0 then | |
| improvementm,t ← improvementspec + improvementsens | (5) |
| end if | |
| end for | |
| end for | |
| mopt, topt ← argmax(improvements) | (6) |
2.5. Evaluation
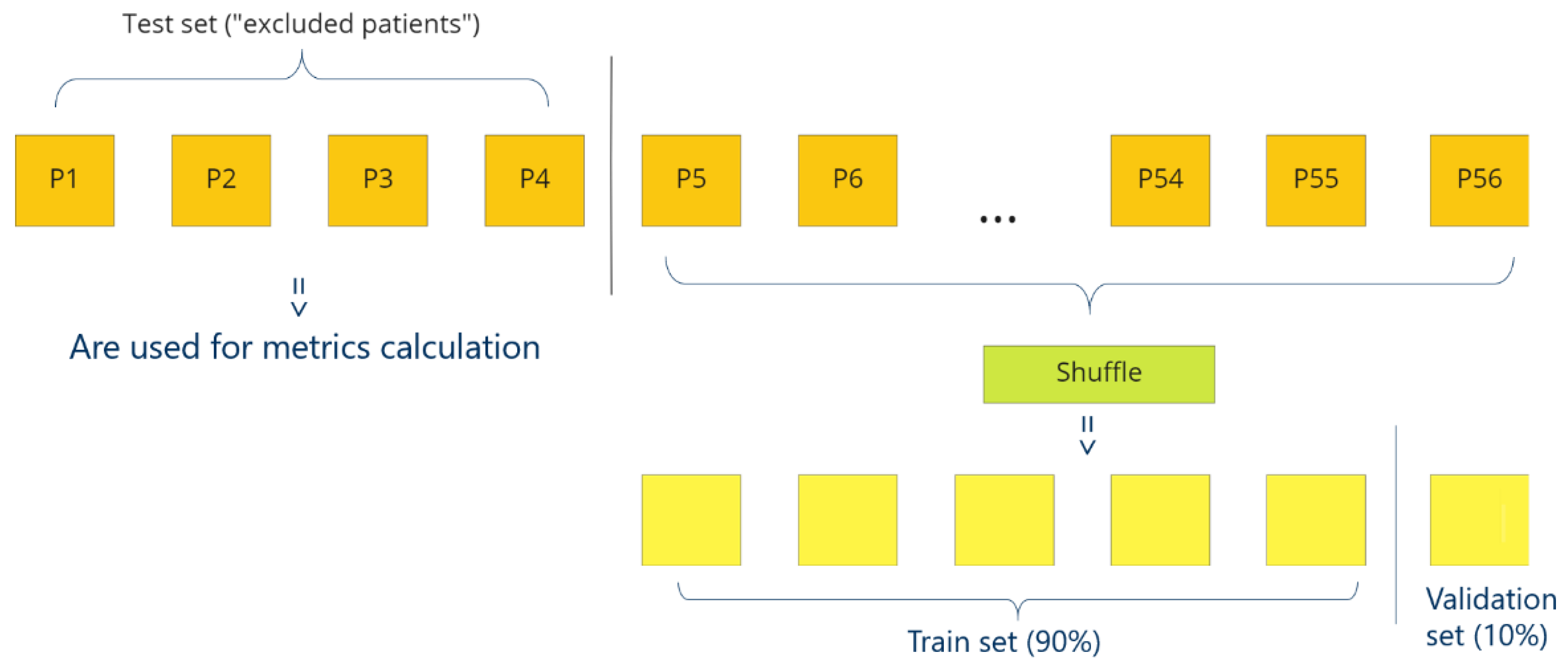
2.5.1. Leave-K-Out-Cross-Validation (LKOCV)
2.5.2. Metrics
2.6. Models
- “Architecture_” +
- “Pre-processing_” +
- “NumberOfExcludedPatients_” +
- “T(rue)/F(alse)” if every third sample was used.
- “Inc” for inception-based models;
- “RS” for RS-based models;
- “Norm” for Normalization pre-processing;
- “Stan” for Standardization pre-processing.
3. Results
3.1. Baseline Results (before Any Post-Processing)
- Inception-based models perform better than RS-based ones. The best three models in Table 6 are inception-based.
- Inception-based networks work better with Normalization (model Inc_Norm_4_T clearly better than model Inc_Stan_1_T and other inception models even despite the fact that model Inc_Stan_1_T uses the most complete set of data) and RS-based models with Standardization (models RS_Stan_4_T and RS_Stan_4_T + SW are better than model RS_Norm_4_T).
- 3.
- Moreover, we noticed that the thresholds are very low (below 0.05) for models that use Standardization as pre-processing, unlike those that use Normalization, for which raw thresholds show regular values (Table 6).
- 4.
- Excluding one patient gives better results than excluding four patients, which is the expected behavior, because in the case of one patient there is more training data.
3.2. Post-Processing Results (AWP and AP)
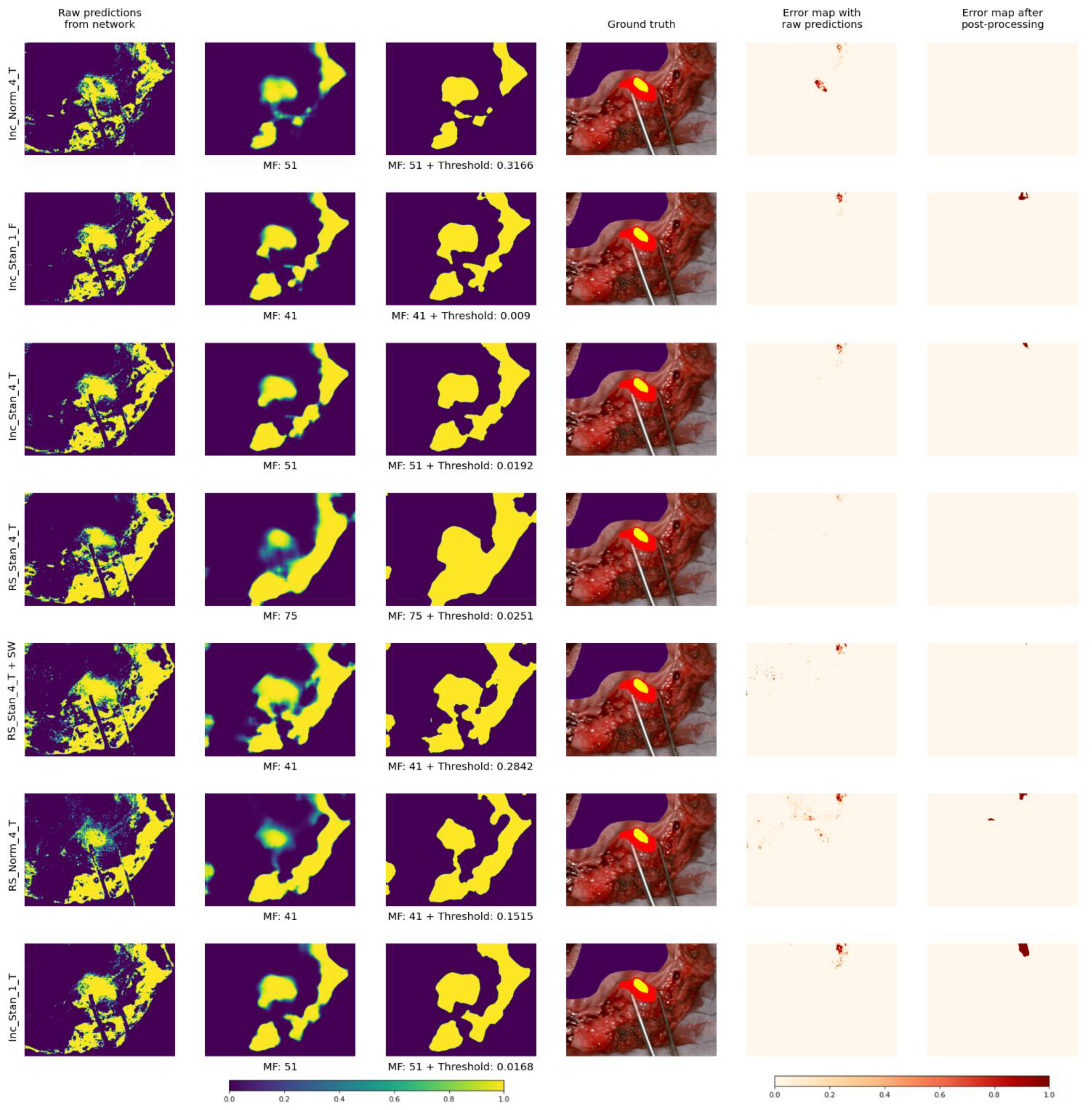
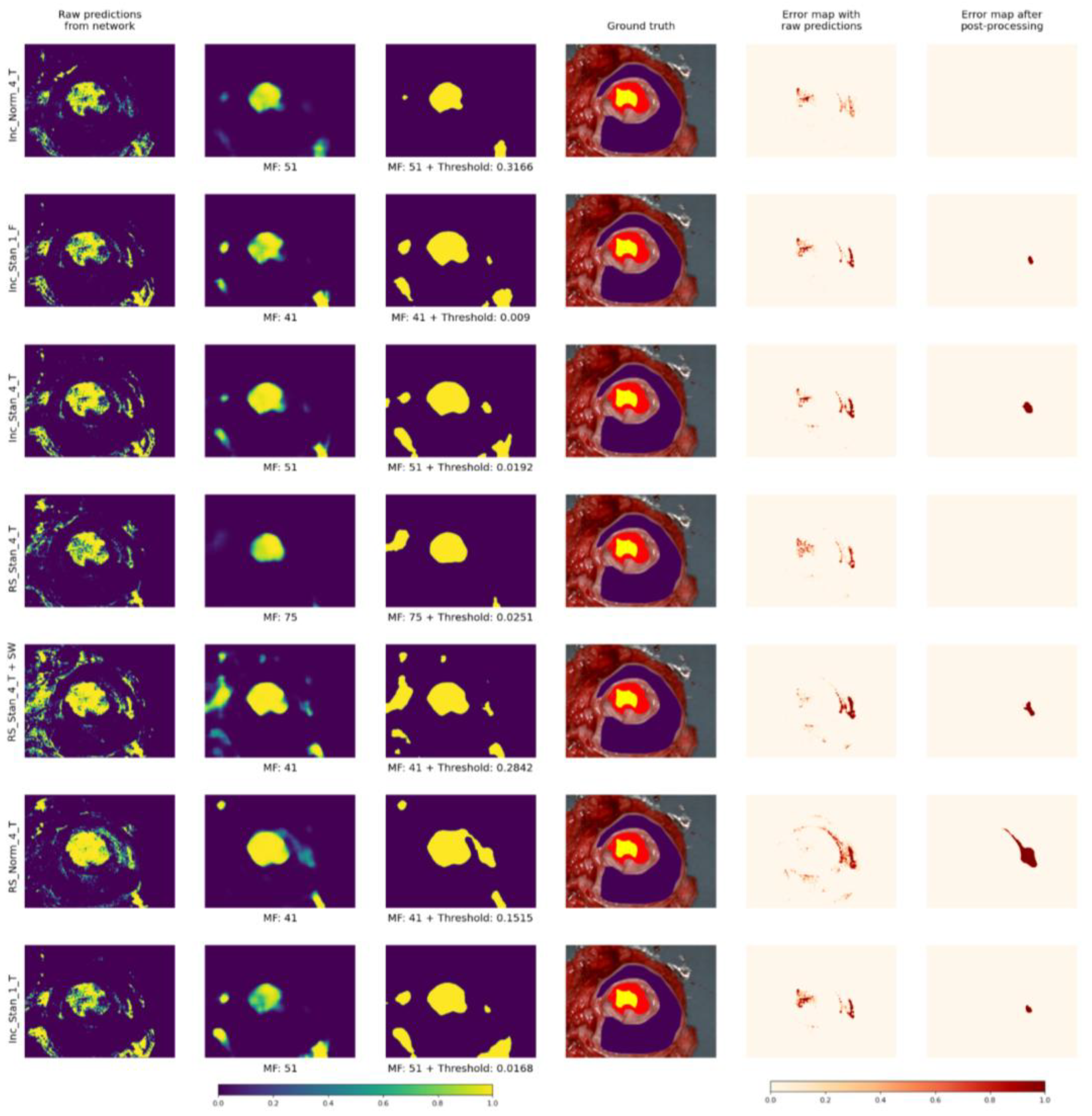
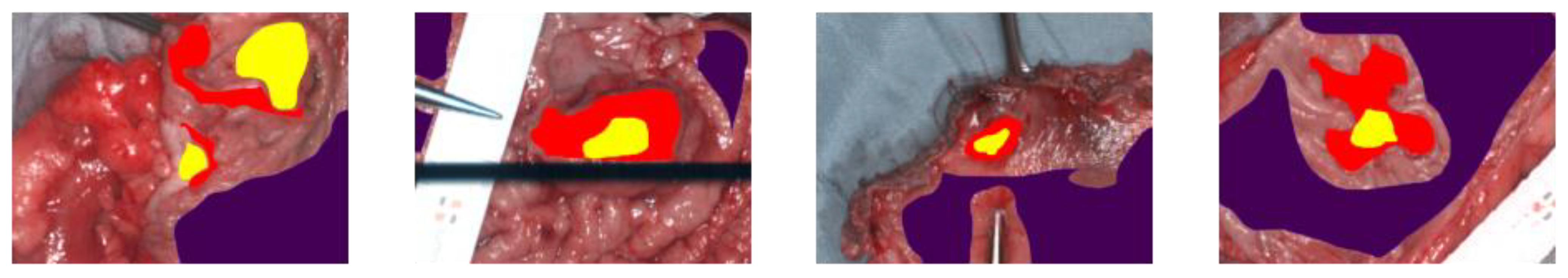
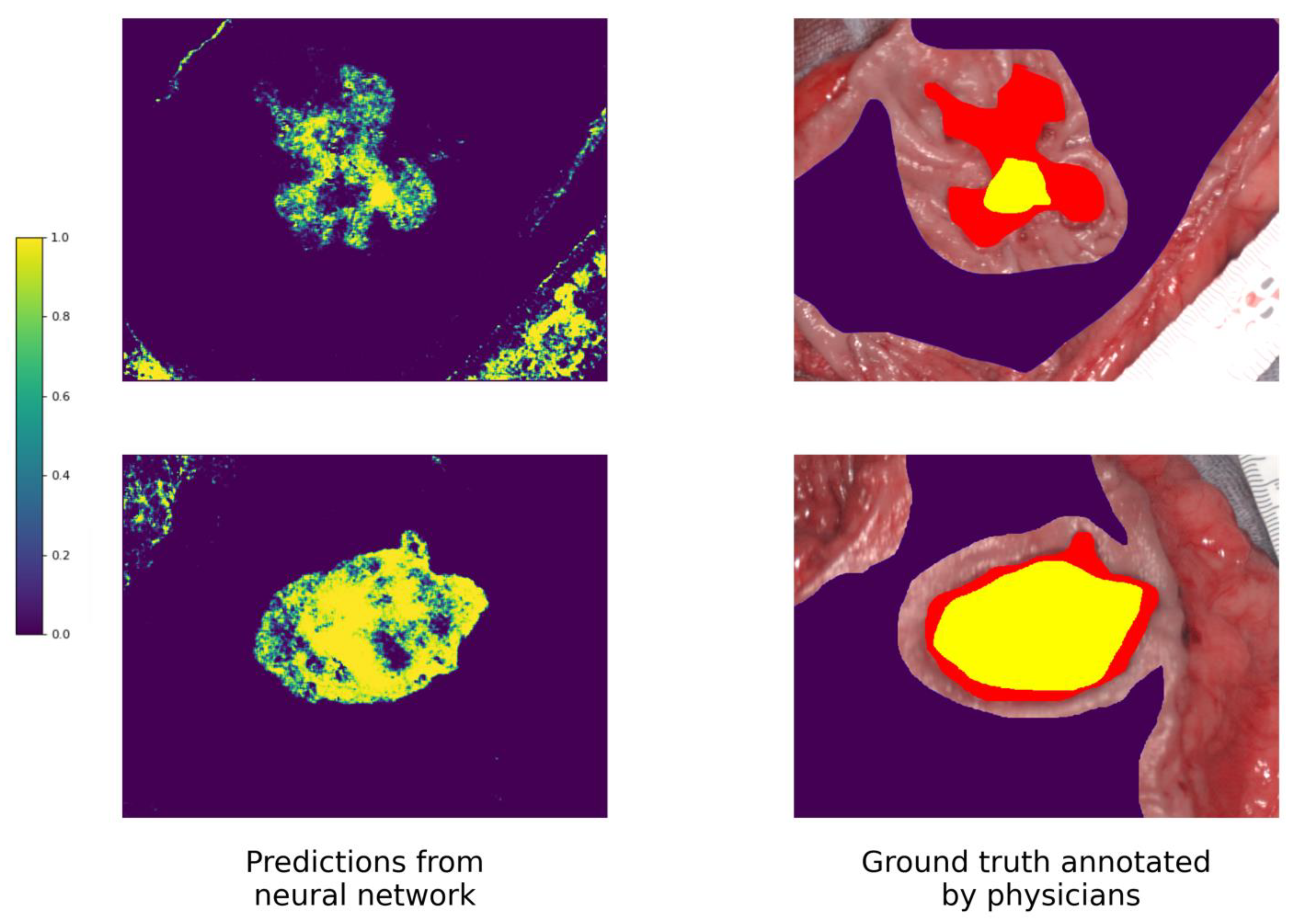
3.3. Visual Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A. Key Wavelengths Detection Algorithm

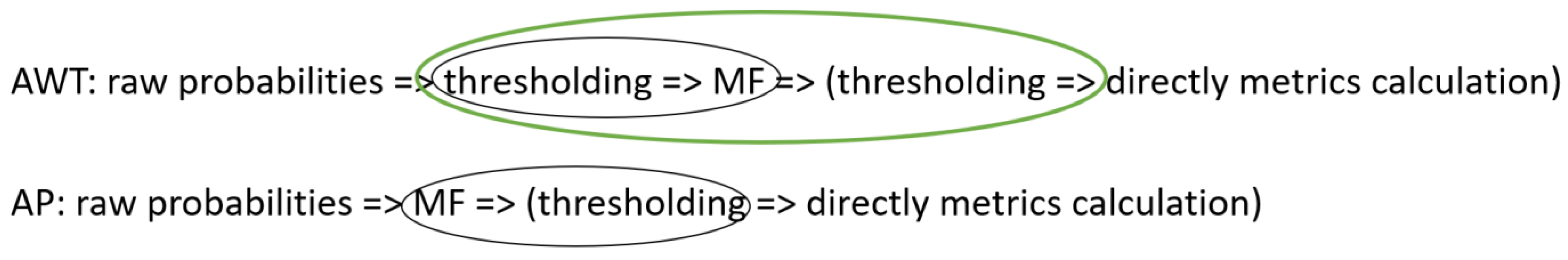
Appendix B. Proof Why AP and AWT Give the Same Results
Appendix B.1. MF and Thresholding Are Commutative: Simple Example

Appendix B.2. Proof That MF and Thresholding Are Commutative
- a > threshold and b > threshold. Then a′ = A, b′ = A.A = A => a′ >= b′. Satisfies the theorem.
- a > threshold and b < threshold. Then a′ = A, b′ = BA > B => a′ >= b′. Satisfies the theorem.
- a < threshold and b > threshold. Not possible, because in this case a < threshold < b => a < b, which contradicts the given conditions.
- a < threshold and b < threshold. Then a′ = B, b′ = B.B = B => a′ >= b′. Satisfies the theorem.
- a = b. Then a′ = b′. Satisfies the theorem.
Appendix B.3. Detailed Description of AWT and AP
References
- Global Cancer Observatory. Available online: https://gco.iarc.fr (accessed on 29 March 2023).
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef]
- Winawer, S.J. The history of colorectal cancer screening: A personal perspective. Dig. Dis. Sci. 2015, 60, 596–608. [Google Scholar] [CrossRef] [PubMed]
- Barberio, M.; Benedicenti, S.; Pizzicannella, M.; Felli, E.; Collins, T.; Jansen-Winkeln, B.; Marescaux, J.; Viola, M.G.; Diana, M. Intraoperative Guidance Using Hyperspectral Imaging: A Review for Surgeons. Diagnostics 2021, 11, 2066. [Google Scholar] [CrossRef]
- Glover, B.; Teare, J.; Patel, N. The Status of Advanced Imaging Techniques for Optical Biopsy of Colonic Polyps. Clin. Transl. Gastroenterol. 2020, 11, e00130. [Google Scholar] [CrossRef] [PubMed]
- Pfahl, A.; Köhler, H.; Thomaßen, M.T.; Maktabi, M.; Bloße, A.M.; Mehdorn, M.; Lyros, O.; Moulla, Y.; Niebisch, S.; Jansen-Winkeln, B.; et al. Video: Clinical evaluation of a laparoscopic hyperspectral imaging system. Surg. Endosc. 2022, 10, 7794–7799. [Google Scholar] [CrossRef]
- Ortega, S.; Halicek, M.; Fabelo, H.; Callico, G.M.; Fei, B. Hyperspectral and multispectral imaging in digital and computational pathology: A systematic review [Invited]. Biomed Opt. Express. 2020, 11, 3195–3233. [Google Scholar] [CrossRef] [PubMed]
- Sucher, R.; Wagner, T.; Köhler, H.; Sucher, E.; Quice, H.; Recknagel, S.; Lederer, A.; Hau, H.M.; Rademacher, S.; Schneeberger, S.; et al. Hyperspectral Imaging (HSI) of Human Kidney Allografts. Ann. Surg. 2022, 276, e48–e55. [Google Scholar] [CrossRef]
- Hren, R.; Sersa, G.; Simoncic, U.; Milanic, M. Imaging perfusion changes in oncological clinical applications by hyperspectral imaging: A literature review. Radiol. Oncol. 2022, 56, 420–429. [Google Scholar] [CrossRef]
- Clancy, T.; Jones, G.; Maier-Hein, L.; Elson, D.S.; Stoyanov, D. Surgical spectral imaging. Med. Image Anal. 2020, 63, 101699. [Google Scholar] [CrossRef]
- Chalopin, C.; Nickel, F.; Pfahl, A.; Köhler, H.; Maktabi, M.; Thieme, R.; Sucher, R.; Jansen-Winkeln, B.; Studier-Fischer, A.; Seidlitz, S.; et al. Künstliche Intelligenz und hyperspektrale Bildgebung zur bildgestützten Assistenz in der minimal-invasiven Chirurgie [Artificial intelligence and hyperspectral imaging for image-guided assistance in minimally invasive surgery]. Chirurgie 2022, 93, 940–947. (In German) [Google Scholar] [CrossRef]
- Liu, L.; Qi, M.; Li, Y.; Liu, Y.; Liu, X.; Zhang, Z.; Qu, J. Staging of Skin Cancer Based on Hyperspectral Microscopic Imaging and Machine Learning. Biosensors 2022, 12, 790. [Google Scholar] [CrossRef] [PubMed]
- Aggarwal, S.L.P.; Papay, F.A. Applications of multispectral and hyperspectral imaging in dermatology. Exp. Dermatol. 2022, 31, 1128–1135. [Google Scholar] [CrossRef]
- Li, Y.; Xie Johansen, T.H.; Møllersen, K.; Ortega, S.; Fabelo, H.; Garcia, A.; Callico, G.M.; Godtliebsen, F. Recent Advances in Hyperspectral Imaging for Melanoma Detection. WIREs Comput. Stat. 2020, 12, e1465. [Google Scholar]
- Li, Y.; Xie, X.; Yang, X.; Guo, L.; Liu, Z.; Zhao, X.; Luo, Y.; Jia, W.; Huang, F.; Zhu, S.; et al. Diagnosis of early gastric cancer based on fluorescence hyperspectral imaging technology combined with partial-least-square discriminant analysis and support vector machine. J. Biophotonics 2019, 12, e201800324. [Google Scholar] [CrossRef] [PubMed]
- Jayanthi, J.L.; Nisha, G.U.; Manju, S.; Philip, E.K.; Jeemon, P.; Baiju, K.V.; Beena, V.T.; Subhash, N. Diffuse reflectance spectroscopy: Diagnostic accuracy of a non-invasive screening technique for early detection of malignant changes in the oral cavity. BMJ Open 2011, 1, e000071. [Google Scholar] [CrossRef]
- Aboughaleb, I.H.; Aref, M.H.; El-Sharkawy, Y.H. Hyperspectral imaging for diagnosis and detection of ex-vivo breast cancer. Photodiagnosis Photodyn. Ther. 2020, 31, 101922. [Google Scholar] [CrossRef]
- Martinez, B.; Leon, R.; Fabelo, H.; Ortega, S.; Piñeiro, J.F.; Szolna, A.; Hernandez, M.; Espino, C.; O’Shanahan, A.J.; Carrera, D.; et al. Most Relevant Spectral Bands Identification for Brain Cancer Detection Using Hyperspectral Imaging. Sensors 2019, 19, 5481. [Google Scholar] [CrossRef] [PubMed]
- Leon, R.; Fabelo, H.; Ortega, S.; Piñeiro, J.F.; Szolna, A.; Hernandez, M.; Espino, C.; O’Shanahan, A.J.; Carrera, D.; Bisshopp, S.; et al. VNIR-NIR hyperspectral imaging fusion targeting intraoperative brain cancer detection. Sci. Rep. 2021, 11, 19696. [Google Scholar] [CrossRef]
- Fabelo, H.; Halicek, M.; Ortega, S.; Shahedi, M.; Szolna, A.; Piñeiro, J.F.; Sosa, C.; O’Shanahan, A.J.; Bisshopp, S.; Espino, C.; et al. Deep Learning-Based Framework for In Vivo Identification of Glioblastoma Tumor Using Hyperspectral Images of Human Brain. Sensors 2019, 19, 920. [Google Scholar] [CrossRef]
- Fabelo, H.; Ortega, S.; Lazcano, R.; Madroñal, D.; Callicó, G.M.; Juárez, E.; Salvador, R.; Bulters, D.; Bulstrode, H.; Szolna, A.; et al. An Intraoperative Visualization System Using Hyperspectral Imaging to Aid in Brain Tumor Delineation. Sensors 2018, 18, 430. [Google Scholar] [CrossRef]
- Giannoni, L.; Lange, F.; Tachtsidis, I. Hyperspectral Imaging Solutions for Brain Tissue Metabolic and Hemodynamic Monitoring: Past, Current and Future Developments. J. Opt. 2018, 20, 044009. [Google Scholar] [CrossRef] [PubMed]
- Eggert, D.; Bengs, M.; Westermann, S.; Gessert, N.; Gerstner, A.O.H.; Mueller, N.A.; Bewarder, J.; Schlaefer, A.; Betz, C.; Laffers, W. In Vivo Detection of Head and Neck Tumors by Hyperspectral Imaging Combined with Deep Learning Methods. J. Biophotonics 2022, 15, e202100167. [Google Scholar] [CrossRef] [PubMed]
- Jansen-Winkeln, B.; Barberio, M.; Chalopin, C.; Schierle, K.; Diana, M.; Köhler, H.; Gockel, I.; Maktabi, M. Feedforward Artificial Neural Network-Based Colorectal Cancer Detection Using Hyperspectral Imaging: A Step towards Automatic Optical Biopsy. Cancers 2021, 13, 967. [Google Scholar] [CrossRef]
- Collins, T.; Maktabi, M.; Barberio, M.; Bencteux, V.; Jansen-Winkeln, B.; Chalopin, C.; Marescaux, J.; Hostettler, A.; Diana, M.; Gockel, I. Automatic Recognition of Colon and Esophagogastric Cancer with Machine Learning and Hyperspectral Imaging. Diagnostics 2021, 11, 1810. [Google Scholar] [CrossRef]
- Martinez-Vega, B.; Tkachenko, M.; Matkabi, M.; Ortega, S.; Fabelo, H.; Balea-Fernandez, F.; La Salvia, M.; Torti, E.; Leporati, F.; Callico, G.M.; et al. Evaluation of Preprocessing Methods on Independent Medical Hyperspectral Databases to Improve Analysis. Sensors 2022, 22, 8917. [Google Scholar] [CrossRef] [PubMed]
- Halicek, M.; Fabelo, H.; Ortega, S.; Little, J.V.; Wang, X.; Chen, A.Y.; Callico, G.M.; Myers, L.; Sumer, B.D.; Fei, B. Hyperspectral imaging for head and neck cancer detection: Specular glare and variance of the tumor margin in surgical specimens. J. Med. Imaging 2019, 6, 035004. [Google Scholar] [CrossRef]
- Halicek, M.; Dormer, J.D.; Little, J.V.; Chen, A.Y.; Fei, B. Tumor detection of the thyroid and salivary glands using hyperspectral imaging and deep learning. Biomed. Opt. Express. 2020, 11, 1383–1400. [Google Scholar] [CrossRef]
- Fabelo, H.; Halicek, M.; Ortega, S.; Szolna, A.; Morera, J.; Sarmiento, R.; Callico, G.M.; Fei, B. Surgical Aid Visualization System for Glioblastoma Tumor Identification based on Deep Learning and In-Vivo Hyperspectral Images of Human Patients. Proc. SPIE Int. Soc. Opt. Eng. 2019, 10951, 1095110. [Google Scholar] [CrossRef] [PubMed]
- Rajendran, T.; Prajoona Valsalan Amutharaj, J.; Jenifer, M.; Rinesh, S.; Charlyn Pushpa Latha, G.; Anitha, T. Hyperspectral Image Classification Model Using Squeeze and Excitation Network with Deep Learning. Comput. Intell. Neurosci. 2022, 4, 9430779. [Google Scholar] [CrossRef]
- Hong, S.M.; Baek, S.S.; Yun, D.; Kwon, Y.H.; Duan, H.; Pyo, J.; Cho, K.H. Monitoring the vertical distribution of HABs using hyperspectral imagery and deep learning models. Sci. Total Environ. 2021, 10, 148592. [Google Scholar] [CrossRef]
- Li, C.; Li, Z.; Liu, X.; Li, S. The Influence of Image Degradation on Hyperspectral Image Classification. Remote Sens. 2022, 14, 5199. [Google Scholar] [CrossRef]
- Kang, X.; Li, S.; Benediktsson, J.A. Spectral–Spatial Hyperspectral Image Classification with Edge-Preserving Filtering. IEEE Trans. Geosci. Remote Sens. 2014, 52, 2666–2677. [Google Scholar] [CrossRef]
- Tarabalka, Y.; Fauvel, M.; Chanussot, J.; Benediktsson, J.A. SVM- and MRF-Based Method for Accurate Classification of Hyperspectral Images. IEEE Geosci. Remote Sens. Lett. 2010, 7, 736–740. [Google Scholar] [CrossRef]
- Szegedy, C.; Liu, W.; Jia, Y.; Sermanet, P.; Reed, S.; Anguelov, D.; Erhan, D.; Vanhoucke, V.; Rabinovich, A. Going deeper with convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Boston, MA, USA, 7–12 June 2015; pp. 1–9. [Google Scholar] [CrossRef]
- Ben Hamida, A.; Benoit, A.; Lambert, P.; Ben Amar, C. 3-D Deep Learning Approach for Remote Sensing Image Classification. IEEE Trans. Geosci. Remote Sens. 2018, 56, 4420–4434. [Google Scholar] [CrossRef]
- Seidlitz, S.; Sellner, J.; Odenthal, J.; Özdemir, B.; Studier-Fischer, A.; Knödler, S.; Ayala, L.; Adler, T.J.; Kenngott, H.G.; Tizabi, M.; et al. Robust deep learning-based semantic organ segmentation in hyperspectral images. Med. Image Anal. 2022, 80, 102488. [Google Scholar] [CrossRef] [PubMed]
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization. In Proceedings of the IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017; pp. 618–626. [Google Scholar]





| Standardization (Z-Score) | Scaling to Unit Length (Normalization) |
|---|---|
| —standard deviation | ||x||—Euclidian length of x |
| x—original spectrum x′—scaled spectrum | |
| Training Parameter Name | Value |
|---|---|
| Epochs | 40 |
| Batch size | 100 |
| Loss | Binary cross entropy |
| Optimizer | Adam with β1 = 0.9 and β2 = 0.99 |
| Learning rate | 0.0001 |
| Dropout | 0.1 |
| Activation | ReLU, except the last layer, where Sigmoid |
| Number of wavelengths | 92 (we exclude the first 8 values because they are very noisy) |
| Number of parameters in models | 393,633 (inception) and 27,156 (RS) |
| Shape of samples | [5, 5] |
| Abbreviation | Description | Meaning |
|---|---|---|
| TP | True Positives | Cancerous detected as cancerous |
| TN | True Negatives | Non-malignant tissue detected as non-malignant tissue |
| FP | False Positives | Non-malignant tissue detected as cancerous |
| FN | False Negatives | Cancerous detected as non-malignant tissue |
| Metric | Formula/Description |
|---|---|
| Sensitivity (also known as recall) | |
| Specificity | |
| F1-score (also known as Sørensen–Dice coefficient) | , where |
| AUC | Area Under the Receiver Operating Characteristic Curve (ROC AUC) |
| MCC (Matthew correlation coefficient) |
| Name | Pre-Processing | Architecture | How Many Patients Are Excluded for Cross-Validation | Every Third Sample | |||
|---|---|---|---|---|---|---|---|
| Normalization | Standardization | Inception | RS | 1 Patient | 4 Patients | ||
| Inc_Norm_4_T | x | x | x | x | |||
| Inc_Stan_1_F | x | x | x | ||||
| Inc_Stan_4_T | x | x | x | x | |||
| RS_Stan_4_T | x | x | x | x | |||
| RS_Stan_4_T + SW 1 | x | x | x | x | |||
| RS_Norm_4_T | x | x | x | x | |||
| Inc_Stan_1_T | x | x | x | x | |||
| Name | Threshold | Accuracy | Sensitivity | Specificity | F1-n 1 | F1-c 1 | AUC | MCC |
|---|---|---|---|---|---|---|---|---|
| Inc_Norm_4_T | 0.211 | 90.2 ± 15 | 89.1 ± 19 | 89.5 ± 17 | 92.8 ± 14 | 65.9 ± 29 | 96.8 ± 10 | 64.3 ± 30 |
| Inc_Stan_1_F | 0.0189 | 89.2 ± 14 | 88.5 ± 21 | 88.2 ± 15 | 92.5 ± 11 | 60.8 ± 29 | 95.7 ± 13 | 59.7 ± 28 |
| Inc_Stan_4_T | 0.0456 | 87.6 ± 16 | 88 ± 21 | 87.1 ± 18 | 91.5 ± 14 | 59 ± 29 | 95.6 ± 13 | 57.8 ± 30 |
| RS_Stan_4_T | 0.0367 | 89 ± 12 | 87.1 ± 20 | 88.3 ± 14 | 92.7 ± 9 | 61.4 ± 28 | 95.5 ± 9 | 60.3 ± 27 |
| RS_Stan_4_T + SW | 0.45 | 87.1 ± 18 | 87 ± 21 | 86.1 ± 20 | 90.5 ± 15 | 59.1 ± 29 | 95.2 ± 10 | 57.4 ± 29 |
| RS_Norm_4_T | 0.1556 | 88.1 ± 16 | 86.8 ± 22 | 87.5 ± 18 | 91.6 ± 13 | 62.6 ± 29 | 94.9 ± 10 | 61.4 ± 28 |
| Inc_Stan_1_T | 0.0456 | 88.6 ± 16 | 86.1 ± 23 | 88 ± 17 | 92 ± 13 | 60.3 ± 30 | 95.5 ± 14 | 59.1 ± 29 |
| Name | MF Size | Threshold | Accuracy | Sensitivity | Specificity | F1-n 1 | F1-c 2 | AUC | MCC |
|---|---|---|---|---|---|---|---|---|---|
| Inc_Norm_4_T | 51 | 0.3166 | 94.2 ± 14 *** | 91.6 ± 22 * | 93.6 ± 16 *** | 95.3 ± 14 * | 78.4 ± 29 *** | 98.4 ± 9 *** | 76.6 ± 32 *** |
| Inc_Stan_1_F | 41 | 0.009 | 91 ± 16 ** | 92.6 ± 21 *** | 89.8 ± 18 ** | 93.2 ± 14 | 70.5 ± 31 *** | 97.2 ± 15 *** | 69.1 ± 32 *** |
| Inc_Stan_4_T | 51 | 0.0192 | 89.2 ± 20 * | 92.7 ± 21 *** | 88.4 ± 22 | 91.5 ± 20 | 68.8 ± 32 *** | 97.2 ± 15 ** | 66.5 ± 38 *** |
| RS_Stan_4_T | 75 | 0.0251 | 92.8 ± 12 *** | 87.8 ± 29 | 92.1 ± 14 *** | 95 ± 9 *** | 68.7 ± 34 * | 96.7 ± 14 | 68.1 ± 33 * |
| RS_Stan_4_T + SW | 41 | 0.2842 | 88.1 ± 21 | 90.5 ± 23 | 86.7 ± 24 | 90.1 ± 21 | 67.3 ± 33 ** | 96.9 ± 10 *** | 65.4 ± 35 ** |
| RS_Norm_4_T | 41 | 0.1515 | 90.9 ± 17 *** | 90.5 ± 25 * | 90.1 ± 19 *** | 93.1 ± 14 *** | 71.9 ± 30 *** | 97.6 ± 7 ** | 71.5 ± 30 *** |
| Inc_Stan_1_T | 51 | 0.0168 | 89.9 ± 19 * | 91.4 ± 23 *** | 88.7 ± 21 | 91.9 ± 18 | 69.1 ± 31 *** | 97.1 ± 15 *** | 67.6 ± 34 *** |
| Name | Improvement Sensitivity (Algorithm 1. Equation (4)) | Improvement Specificity (Algorithm 1. Equation (3)) | Improvement (Algorithm 1. Equation (5)) |
|---|---|---|---|
| Inc_Norm_4_T | 2.34 | 4.292 | 6.638 |
| Inc_Stan_1_F | 4.20 | 1.486 | 5.689 |
| Inc_Stan_4_T | 5.12 | 0.796 | 5.925 |
| RS_Stan_4_T | 0.02 | 4.378 | 4.399 |
| RS_Stan_4_T + SW | 3.87 | 0.155 | 4.032 |
| RS_Norm_4_T | 3.33 | 2.905 | 6.241 |
| Inc_Stan_1_T | 4.42 | 1.641 | 6.065 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tkachenko, M.; Chalopin, C.; Jansen-Winkeln, B.; Neumuth, T.; Gockel, I.; Maktabi, M. Impact of Pre- and Post-Processing Steps for Supervised Classification of Colorectal Cancer in Hyperspectral Images. Cancers 2023, 15, 2157. https://doi.org/10.3390/cancers15072157
Tkachenko M, Chalopin C, Jansen-Winkeln B, Neumuth T, Gockel I, Maktabi M. Impact of Pre- and Post-Processing Steps for Supervised Classification of Colorectal Cancer in Hyperspectral Images. Cancers. 2023; 15(7):2157. https://doi.org/10.3390/cancers15072157
Chicago/Turabian StyleTkachenko, Mariia, Claire Chalopin, Boris Jansen-Winkeln, Thomas Neumuth, Ines Gockel, and Marianne Maktabi. 2023. "Impact of Pre- and Post-Processing Steps for Supervised Classification of Colorectal Cancer in Hyperspectral Images" Cancers 15, no. 7: 2157. https://doi.org/10.3390/cancers15072157
APA StyleTkachenko, M., Chalopin, C., Jansen-Winkeln, B., Neumuth, T., Gockel, I., & Maktabi, M. (2023). Impact of Pre- and Post-Processing Steps for Supervised Classification of Colorectal Cancer in Hyperspectral Images. Cancers, 15(7), 2157. https://doi.org/10.3390/cancers15072157







