Connected-SegNets: A Deep Learning Model for Breast Tumor Segmentation from X-ray Images
Abstract
:Simple Summary
Abstract
1. Introduction
- This study proposes a deep learning model called Connected-SegNets for breast tumor segmentation from X-ray images.
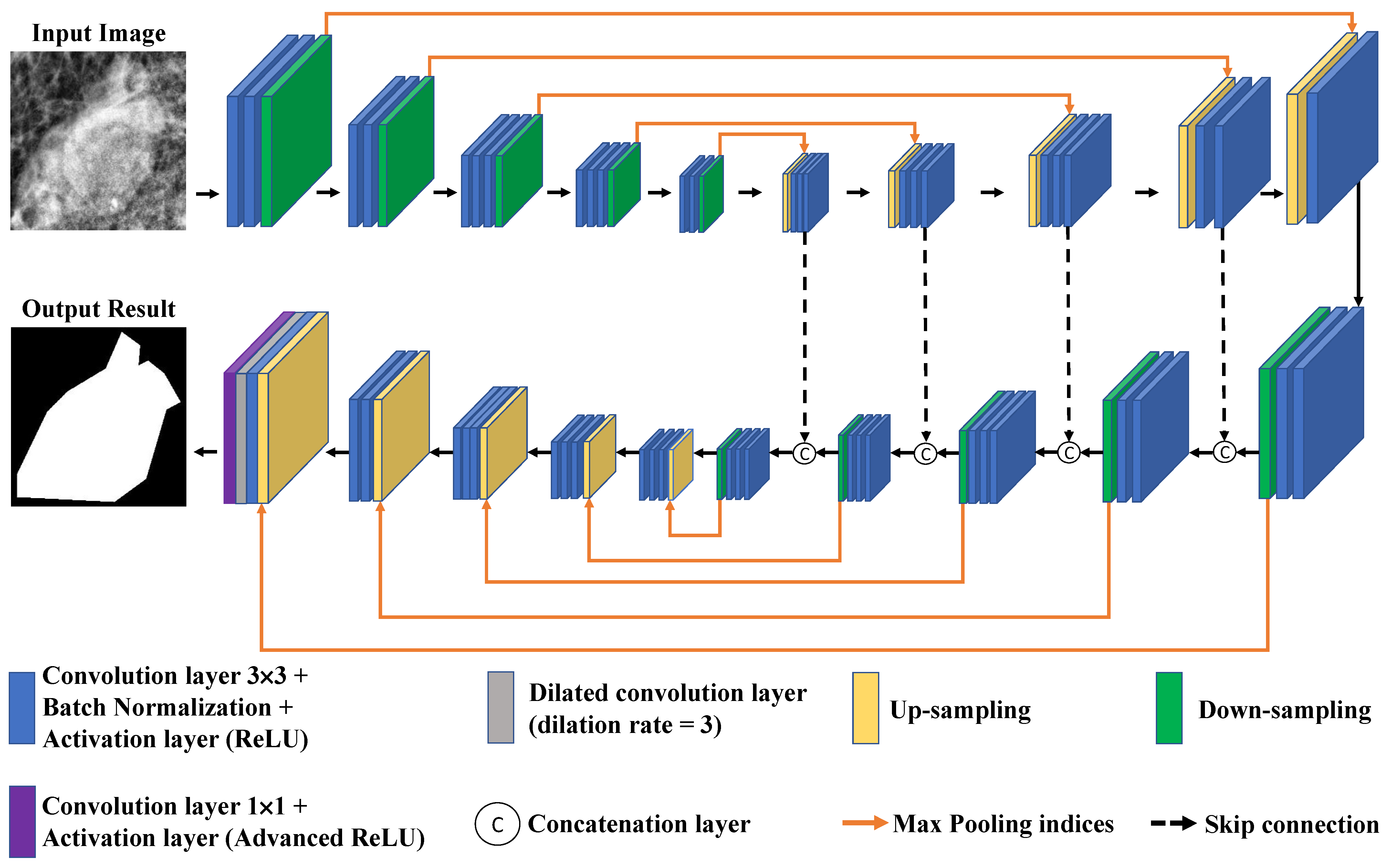
- The proposed model, Connected-SegNets, is designed using skip connections, which helps to recover the spatial information lost during the pooling operations.
- The original SegNet cross-entropy loss function has been replaced by the IoU loss function to overcome any noisy features and enhance the detection of the false negative and false positive cases.
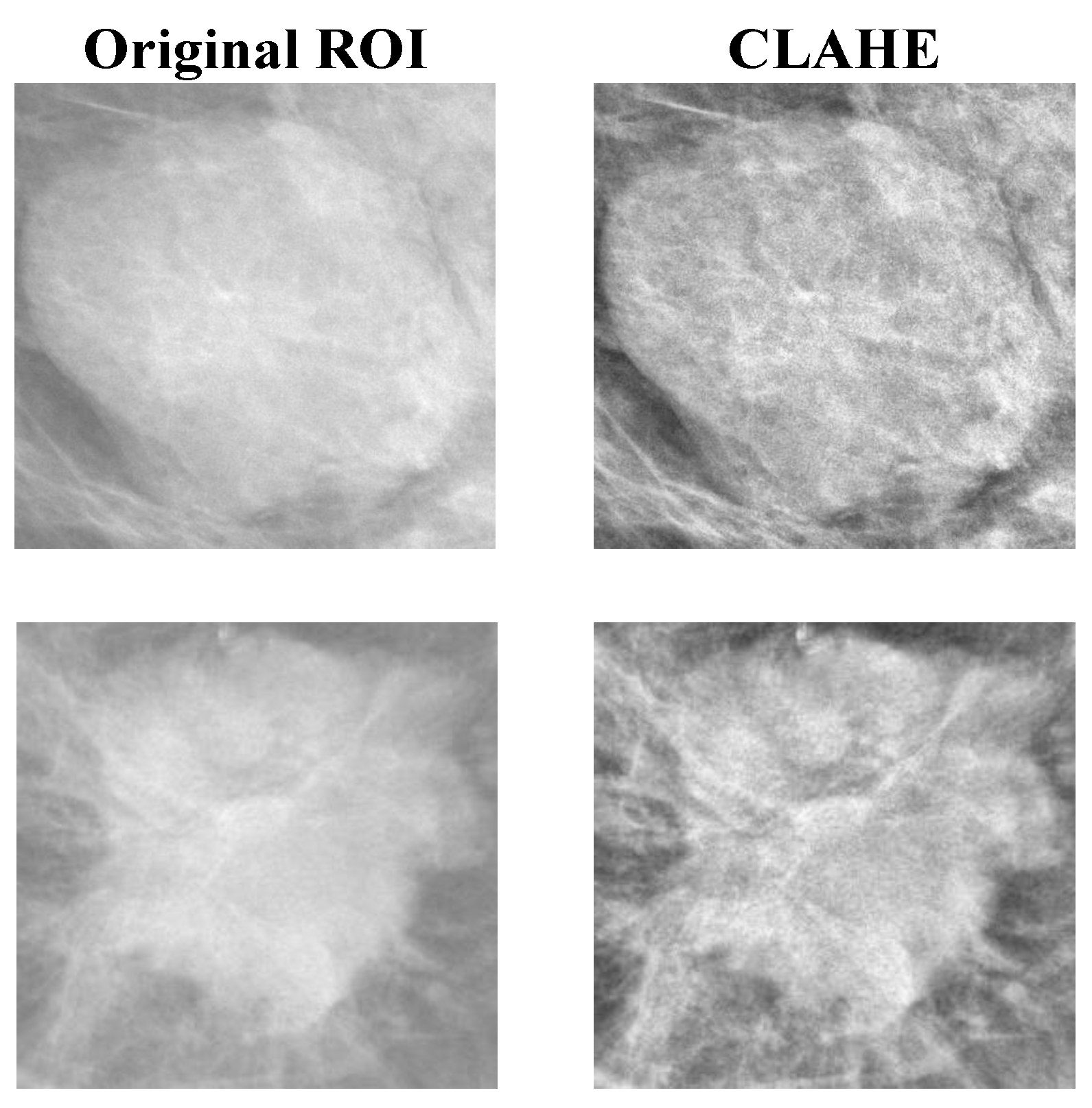
- The histogram equalization method of the contrast limit adapt histogram equalization (CLAHE) is applied to all datasets to enhance the compressed areas and smooth the pixel distribution.
- Image augmentation methods including rotation and flipping have been used to increase the number of training data and to reduce the impact of overfitting.
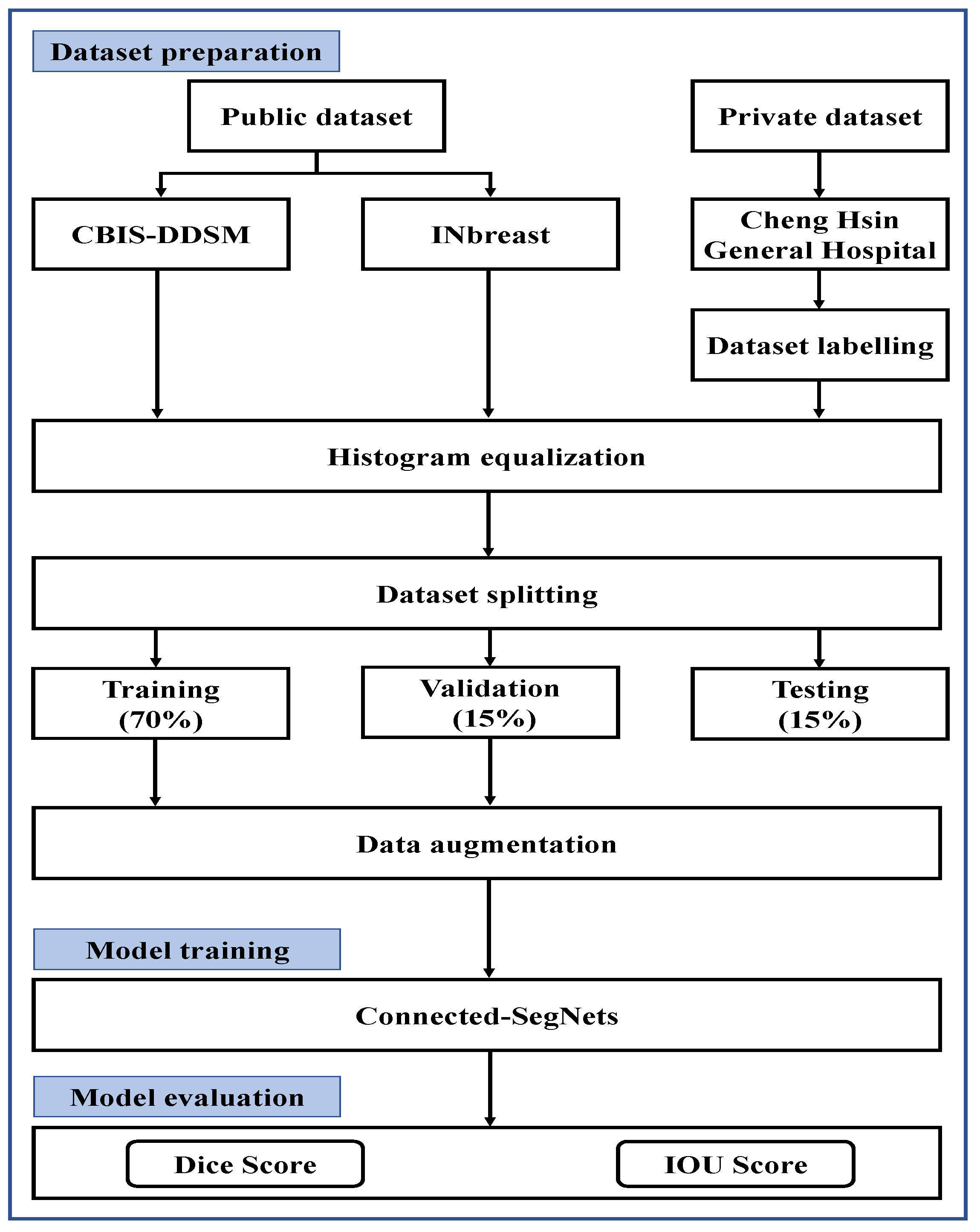
2. Materials and Methods
2.1. Datasets
2.1.1. INbreast Dataset
2.1.2. CBIS-DDSM Dataset
2.1.3. Private Dataset
2.2. Data Preprocessing
2.2.1. Histogram Equalization
2.2.2. Image Augmentation
2.3. Proposed Model
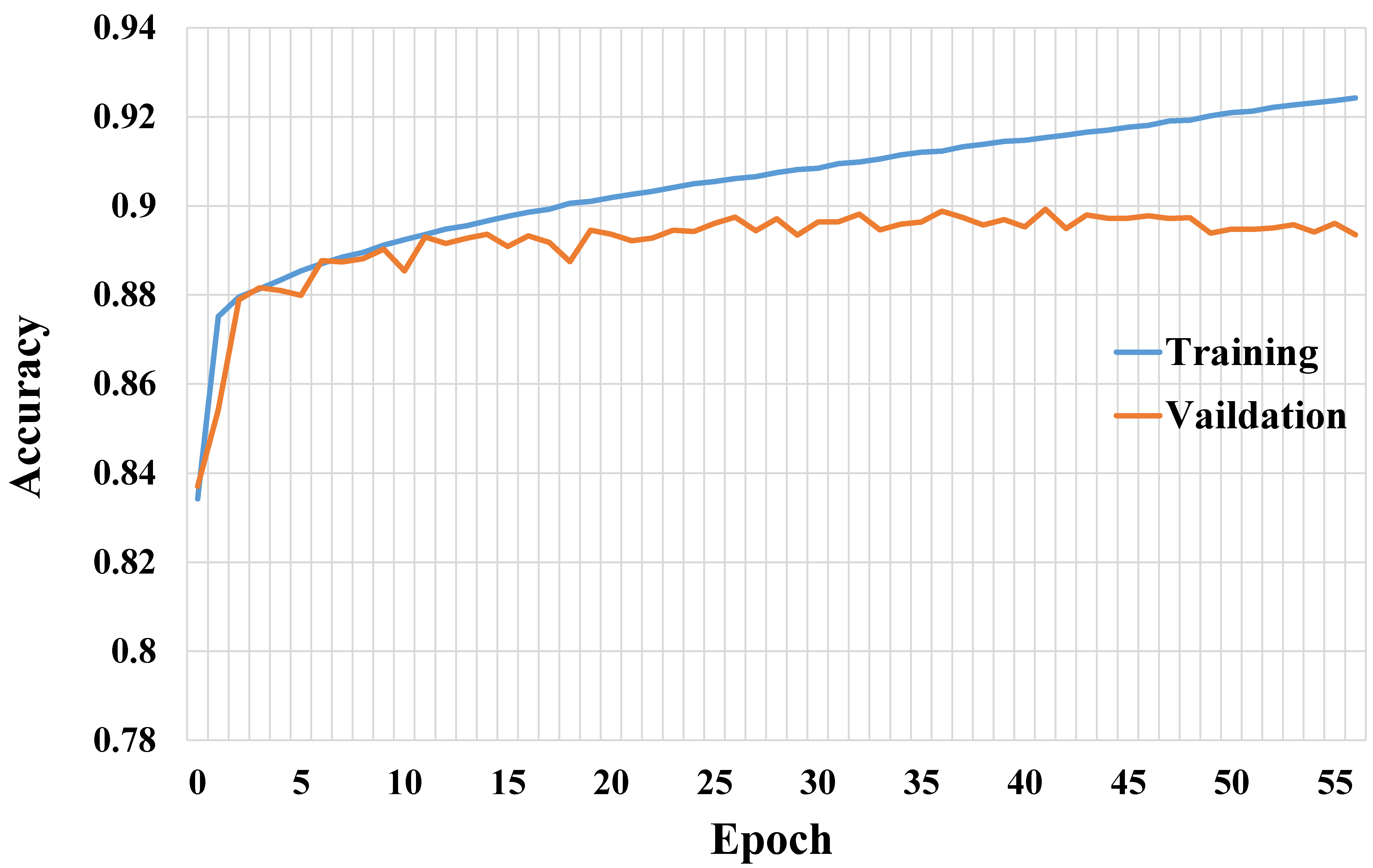
2.4. Experimental Environment and Parameter Settings
2.5. Evaluation Metrics
3. Results
3.1. Results on INbreast Dataset
3.2. Results on CBIS-DDSM Dataset
3.3. Results on Private Dataset
| Algorithm 1 Validation Loss Tracking for Early Stop | |
| Input: LatestValLoss, ActStepSetting | |
| Output: BestValLossScore | |
| 1: | |
| 2: | if then |
| 3: | if then |
| 4: | |
| 5: | |
| 6: | else |
| 7: | |
| 8: | end if |
| 9: | else |
| 10: | |
| 11: | end if |
| 12: | return |
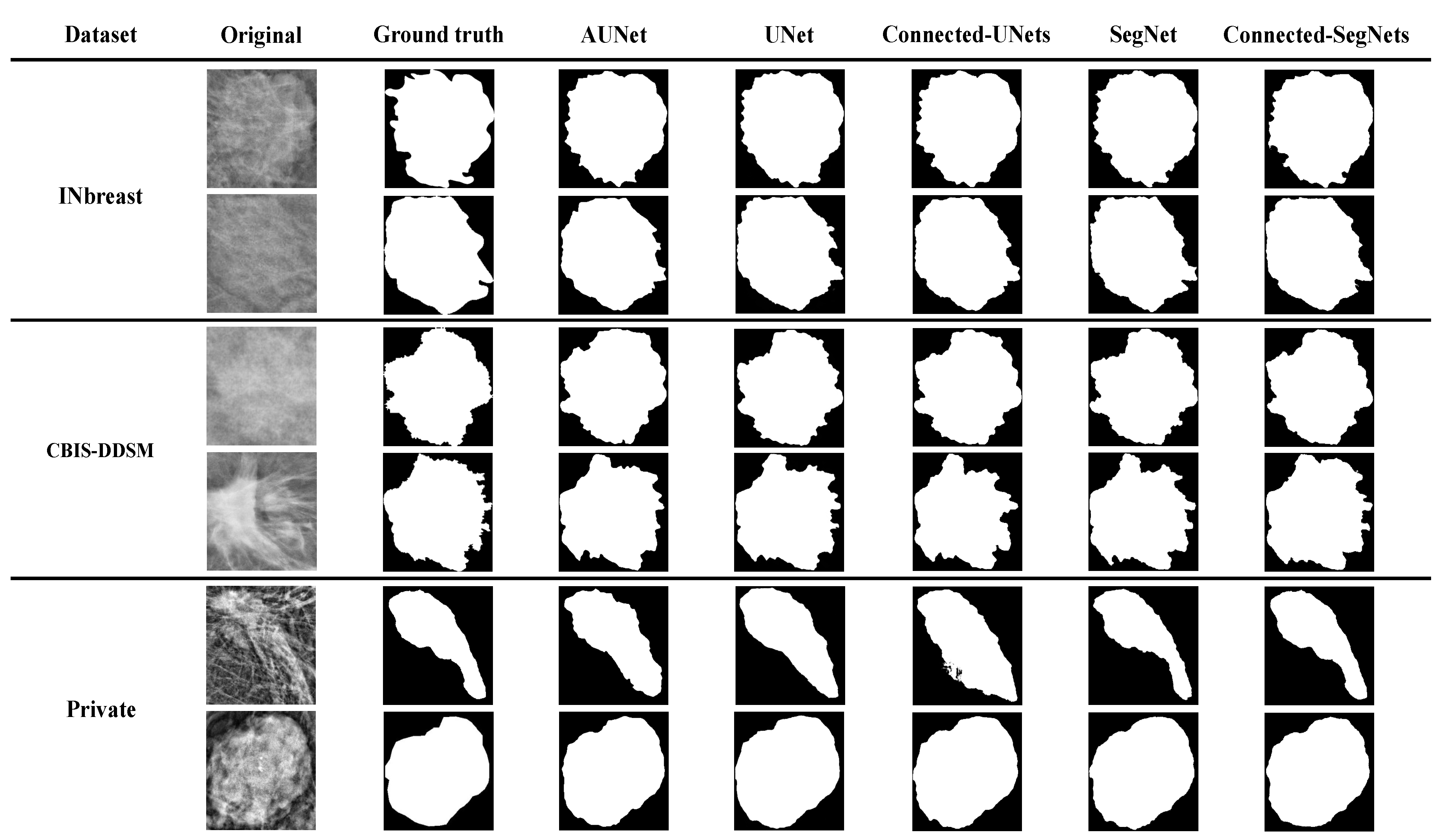
3.4. Comparison of Segmentation Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer statistics, 2022. CA Cancer J. Clin. 2022, 72, 7–33. [Google Scholar] [CrossRef] [PubMed]
- Zou, R.; Loke, S.Y.; Tan, V.K.M.; Quek, S.T.; Jagmohan, P.; Tang, Y.C.; Madhukumar, P.; Tan, B.K.T.; Yong, W.S.; Sim, Y.; et al. Development of a microRNA panel for classification of abnormal mammograms for breast cancer. Cancers 2021, 13, 2130. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Guan, X.; Fan, Z.; Ching, L.M.; Li, Y.; Wang, X.; Cao, W.M.; Liu, D.X. Non-invasive biomarkers for early detection of breast cancer. Cancers 2020, 12, 2767. [Google Scholar] [CrossRef] [PubMed]
- Almalki, Y.E.; Soomro, T.A.; Irfan, M.; Alduraibi, S.K.; Ali, A. Computerized Analysis of Mammogram Images for Early Detection of Breast Cancer. Healthcare 2022, 10, 801. [Google Scholar] [CrossRef] [PubMed]
- Shi, P.; Zhong, J.; Rampun, A.; Wang, H. A hierarchical pipeline for breast boundary segmentation and calcification detection in mammograms. Comput. Biol. Med. 2018, 96, 178–188. [Google Scholar] [CrossRef]
- Waks, A.G.; Winer, E.P. Breast cancer treatment: A review. JAMA 2019, 321, 288–300. [Google Scholar] [CrossRef] [PubMed]
- Salgado, R.; Denkert, C.; Demaria, S.; Sirtaine, N.; Klauschen, F.; Pruneri, G.; Wienert, S.; Van den Eynden, G.; Baehner, F.L.; Pénault-Llorca, F.; et al. The evaluation of tumor-infiltrating lymphocytes (TILs) in breast cancer: Recommendations by an International TILs Working Group 2014. Ann. Oncol. 2015, 26, 259–271. [Google Scholar] [CrossRef]
- Tariq, M.; Iqbal, S.; Ayesha, H.; Abbas, I.; Ahmad, K.T.; Niazi, M.F.K. Medical image based breast cancer diagnosis: State of the art and future directions. Expert Syst. Appl. 2021, 167, 114095. [Google Scholar] [CrossRef]
- Petrillo, A.; Fusco, R.; Di Bernardo, E.; Petrosino, T.; Barretta, M.L.; Porto, A.; Granata, V.; Di Bonito, M.; Fanizzi, A.; Massafra, R.; et al. Prediction of Breast Cancer Histological Outcome by Radiomics and Artificial Intelligence Analysis in Contrast-Enhanced Mammography. Cancers 2022, 14, 2132. [Google Scholar] [CrossRef]
- Ahmed, L.; Iqbal, M.M.; Aldabbas, H.; Khalid, S.; Saleem, Y.; Saeed, S. Images data practices for semantic segmentation of breast cancer using deep neural network. J. Ambient. Intell. Humaniz. Comput. 2020, 1–17. [Google Scholar] [CrossRef]
- Le, E.; Wang, Y.; Huang, Y.; Hickman, S.; Gilbert, F. Artificial intelligence in breast imaging. Clin. Radiol. 2019, 74, 357–366. [Google Scholar] [CrossRef] [PubMed]
- Bi, W.L.; Hosny, A.; Schabath, M.B.; Giger, M.L.; Birkbak, N.J.; Mehrtash, A.; Allison, T.; Arnaout, O.; Abbosh, C.; Dunn, I.F.; et al. Artificial intelligence in cancer imaging: Clinical challenges and applications. CA Cancer J. Clin. 2019, 69, 127–157. [Google Scholar] [CrossRef] [PubMed]
- Shah, S.M.; Khan, R.A.; Arif, S.; Sajid, U. Artificial intelligence for breast cancer analysis: Trends & directions. Comput. Biol. Med. 2022, 142, 105221. [Google Scholar] [PubMed]
- Ketabi, H.; Ekhlasi, A.; Ahmadi, H. A computer-aided approach for automatic detection of breast masses in digital mammogram via spectral clustering and support vector machine. Phys. Eng. Sci. Med. 2021, 44, 277–290. [Google Scholar] [CrossRef]
- Hosny, A.; Parmar, C.; Quackenbush, J.; Schwartz, L.H.; Aerts, H.J. Artificial intelligence in radiology. Nat. Rev. Cancer 2018, 18, 500–510. [Google Scholar] [CrossRef]
- Vobugari, N.; Raja, V.; Sethi, U.; Gandhi, K.; Raja, K.; Surani, S.R. Advancements in Oncology with Artificial Intelligence—A Review Article. Cancers 2022, 14, 1349. [Google Scholar] [CrossRef]
- Alkhaleefah, M.; Wu, C.C. A hybrid CNN and RBF-based SVM approach for breast cancer classification in mammograms. In Proceedings of the 2018 IEEE International Conference on Systems, Man, and Cybernetics (SMC), Miyazaki, Japan, 7–10 October 2018; pp. 894–899. [Google Scholar]
- Kallenberg, M.; Petersen, K.; Nielsen, M.; Ng, A.Y.; Diao, P.; Igel, C.; Vachon, C.M.; Holland, K.; Winkel, R.R.; Karssemeijer, N.; et al. Unsupervised deep learning applied to breast density segmentation and mammographic risk scoring. IEEE Trans. Med. Imaging 2016, 35, 1322–1331. [Google Scholar] [CrossRef]
- Ravitha Rajalakshmi, N.; Vidhyapriya, R.; Elango, N.; Ramesh, N. Deeply supervised u-net for mass segmentation in digital mammograms. Int. J. Imaging Syst. Technol. 2021, 31, 59–71. [Google Scholar]
- Sun, H.; Li, C.; Liu, B.; Liu, Z.; Wang, M.; Zheng, H.; Feng, D.D.; Wang, S. AUNet: Attention-guided dense-upsampling networks for breast mass segmentation in whole mammograms. Phys. Med. Biol. 2020, 65, 055005. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In Medical Image Computing and Computer-Assisted Intervention–MICCAI 2015; Springer International Publishing: Cham, Switzerland, 2015; pp. 234–241. [Google Scholar]
- Baccouche, A.; Garcia-Zapirain, B.; Castillo Olea, C.; Elmaghraby, A.S. Connected-UNets: A deep learning architecture for breast mass segmentation. NPJ Breast Cancer 2021, 7, 1–12. [Google Scholar] [CrossRef]
- Badrinarayanan, V.; Kendall, A.; Cipolla, R. Segnet: A deep convolutional encoder-decoder architecture for image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 2481–2495. [Google Scholar] [CrossRef] [PubMed]
- Moreira, I.C.; Amaral, I.; Domingues, I.; Cardoso, A.; Cardoso, M.J.; Cardoso, J.S. Inbreast: Toward a full-field digital mammographic database. Acad. Radiol. 2012, 19, 236–248. [Google Scholar] [CrossRef] [PubMed]
- Huang, M.L.; Lin, T.Y. Dataset of breast mammography images with masses. Data Brief 2020, 31, 105928. [Google Scholar] [CrossRef]
- Heath, M.; Bowyer, K.; Kopans, D.; Kegelmeyer, P.; Moore, R.; Chang, K.; Munishkumaran, S. Current status of the digital database for screening mammography. In Digital Mammography; Springer: Cham, Switzerland, 1998; pp. 457–460. [Google Scholar]
- Lee, R.S.; Gimenez, F.; Hoogi, A.; Miyake, K.K.; Gorovoy, M.; Rubin, D.L. A curated mammography data set for use in computer-aided detection and diagnosis research. Sci. Data 2017, 4, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Dutta, A.; Zisserman, A. The VIA Annotation Software for Images, Audio and Video. In Proceedings of the 27th ACM International Conference on Multimedia, Nice, France, 21–25 October 2019; ACM: New York, NY, USA, 2019. MM’19. [Google Scholar] [CrossRef]
- Al-Masni, M.A.; Al-Antari, M.A.; Park, J.M.; Gi, G.; Kim, T.Y.; Rivera, P.; Valarezo, E.; Choi, M.T.; Han, S.M.; Kim, T.S. Simultaneous detection and classification of breast masses in digital mammograms via a deep learning YOLO-based CAD system. Comput. Methods Programs Biomed. 2018, 157, 85–94. [Google Scholar] [CrossRef]
- Hai, J.; Qiao, K.; Chen, J.; Tan, H.; Xu, J.; Zeng, L.; Shi, D.; Yan, B. Fully convolutional densenet with multiscale context for automated breast tumor segmentation. J. Healthc. Eng. 2019, 2019, 8415485. [Google Scholar] [CrossRef] [PubMed]
- Dhal, K.G.; Das, A.; Ray, S.; Gálvez, J.; Das, S. Histogram equalization variants as optimization problems: A review. Arch. Comput. Methods Eng. 2021, 28, 1471–1496. [Google Scholar] [CrossRef]
- Huang, Z.; Zhang, Y.; Li, Q.; Zhang, T.; Sang, N. Spatially adaptive denoising for X-ray cardiovascular angiogram images. Biomed. Signal Process. Control. 2018, 40, 131–139. [Google Scholar] [CrossRef]
- Huang, Z.; Li, X.; Wang, N.; Ma, L.; Hong, H. Simultaneous denoising and enhancement for X-ray angiograms by employing spatial-frequency filter. Optik 2020, 208, 164287. [Google Scholar] [CrossRef]
- Huang, Z.; Zhang, Y.; Li, Q.; Li, X.; Zhang, T.; Sang, N.; Hong, H. Joint analysis and weighted synthesis sparsity priors for simultaneous denoising and destriping optical remote sensing images. IEEE Trans. Geosci. Remote Sens. 2020, 58, 6958–6982. [Google Scholar] [CrossRef]
- Rao, B.S. Dynamic histogram equalization for contrast enhancement for digital images. Appl. Soft Comput. 2020, 89, 106114. [Google Scholar] [CrossRef]
- Alkhaleefah, M.; Ma, S.C.; Chang, Y.L.; Huang, B.; Chittem, P.K.; Achhannagari, V.P. Double-shot transfer learning for breast cancer classification from X-ray images. Appl. Sci. 2020, 10, 3999. [Google Scholar] [CrossRef]
- Elasal, N.; Swart, D.M.; Miller, N. Frame augmentation for imbalanced object detection datasets. J. Comput. Vis. Imaging Syst. 2018, 4, 3. [Google Scholar]
- Shorten, C.; Khoshgoftaar, T.M. A survey on image data augmentation for deep learning. J. Big Data 2019, 6, 1–48. [Google Scholar] [CrossRef]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar]
- Al-Antari, M.A.; Al-Masni, M.A.; Choi, M.T.; Han, S.M.; Kim, T.S. A fully integrated computer-aided diagnosis system for digital X-ray mammograms via deep learning detection, segmentation, and classification. Int. J. Med. Infor. 2018, 117, 44–54. [Google Scholar] [CrossRef]


| Dataset | Raw ROIs | Training Samples | Testing Samples |
|---|---|---|---|
| INbreast dataset | 107 | 90 | 17 |
| CBIS-DDSM dataset | 838 | 728 | 110 |
| Private dataset | 196 | 148 | 48 |
| Total | 1141 | 966 | 175 |
| Dataset | Raw Images | Augmented Images | Training | Validation |
|---|---|---|---|---|
| INbreast Dataset | 90 | 720 | 576 | 144 |
| CBIS-DDSM dataset | 728 | 5824 | 4659 | 1165 |
| Private dataset | 148 | 1184 | 947 | 237 |
| Total | 966 | 7728 | 6182 | 1546 |
| SegNet1 | |||||
|---|---|---|---|---|---|
| No. | Layer Name | Output | Filter Size | No. of Filters | No. of Layers |
| 1 | Input | 256 × 256 × 1 | 1 | ||
| 2 | Conv1 | 256 × 256 × 64 | 3 × 3 | 64 | 2 |
| 3 | Maxpool 1 | 128 × 128 × 64 | 1 | ||
| 4 | Conv2 | 128 × 128 × 128 | 3 × 3 | 128 | 2 |
| 5 | Maxpool 1 | 64 × 64 × 128 | 1 | ||
| 6 | Conv3 | 64 × 64 × 256 | 3 × 3 | 256 | 3 |
| 7 | Maxpool 1 | 32 × 32 × 256 | 1 | ||
| 8 | Conv4 | 32 × 32 × 512 | 3 × 3 | 512 | 3 |
| 9 | Maxpool 1 | 16 × 16 × 512 | 1 | ||
| 10 | Conv5 | 16 × 16 × 512 | 3 × 3 | 512 | 3 |
| 11 | Maxpool 1 | 8× 8 × 512 | 1 | ||
| 12 | Upsampling 2 | 16 × 16 × 512 | 1 | ||
| 13 | Conv6 | 16 × 16 × 512 | 3 × 3 | 512 | 3 |
| 14 | Upsampling 2 | 32 × 32 × 512 | 1 | ||
| 15 | Conv7 | 32 × 32 × 512 | 3 × 3 | 512 | 2 |
| 16 | Conv8 | 32 × 32 × 256 | 3 × 3 | 256 | 1 |
| 17 | Upsampling 2 | 64 × 64 × 256 | 1 | ||
| 18 | Conv9 | 64 × 64 × 256 | 3 × 3 | 256 | 2 |
| 19 | Conv10 | 64 × 64 × 128 | 3 × 3 | 128 | 1 |
| 20 | Upsampling 2 | 128 × 128 × 128 | 1 | ||
| 21 | Conv11 | 128 × 128 × 128 | 3 × 3 | 128 | 2 |
| 22 | Conv12 | 128 × 128 × 64 | 3 × 3 | 64 | 1 |
| 23 | Upsampling 2 | 256 × 256 × 64 | 1 | ||
| 24 | Conv13 | 256 × 256 × 64 | 3 × 3 | 64 | 1 |
| 25 | Conv13 | ||||
| 26 | Conv14 | 64 | 2 | ||
| 27 | Maxpool 1 | 1 | |||
| 28 | Concatenate | 1 | |||
| 29 | Conv15 | 128 | 2 | ||
| 30 | Maxpool 1 | 1 | |||
| 31 | Concatenate | 1 | |||
| 32 | Conv16 | 256 | 3 | ||
| 33 | Maxpool 1 | 1 | |||
| 34 | Concatenate | 1 | |||
| 35 | Conv17 | 3 × 3 | 512 | 3 | |
| 36 | Maxpool 1 | 1 | |||
| 37 | Concatenate | 1 | |||
| 38 | Conv18 | 3 × 3 | 512 | 3 | |
| 39 | Maxpool 1 | 1 | |||
| 40 | Upsampling 2 | 1 | |||
| 41 | Conv19 | 512 | 3 | ||
| 42 | Upsampling 2 | 1 | |||
| 43 | Conv20 | 512 | 2 | ||
| 44 | Conv21 | 256 | 1 | ||
| 45 | Upsampling 2 | 1 | |||
| 46 | Conv22 | 256 | 2 | ||
| 47 | Conv23 | 128 | 1 | ||
| 48 | Upsampling 2 | 1 | |||
| 49 | Conv24 | 128 | 2 | ||
| 50 | Conv25 | 64 | 1 | ||
| 51 | Upsampling 2 | 1 | |||
| 52 | Conv26 | 64 | 1 | ||
| 53 | Conv27 | 3 × 3 (D 3 = 3) | 64 | 1 | |
| 54 | Output | 1 | 1 | ||
| Connected-SegNets | |||
|---|---|---|---|
| Ground Truth | |||
| Tumor | Non-Tumor | ||
| Prediction | Tumor | 96% (TP) | 4% (FN) |
| Non-Tumor | 12% (FP) | 88% (TN) | |
| Connected-SegNets | |||
|---|---|---|---|
| Ground Truth | |||
| Tumor | Non-Tumor | ||
| Prediction | Tumor | 93% (TP) | 7% (FN) |
| Non-Tumor | 13% (FP) | 87% (TN) | |
| Connected-SegNets | |||
|---|---|---|---|
| Ground Truth | |||
| Tumor | Non-Tumor | ||
| Prediction | Tumor | 92% (TP) | 8% (FN) |
| Non-Tumor | 11% (FP) | 89% (TN) | |
| Model | INbreast Dataset | CBIS-DDSM Dataset | Private Dataset | |||
|---|---|---|---|---|---|---|
| Dice Score (%) | IoU Score (%) | Dice Score (%) | IoU Score (%) | Dice Score (%) | IoU Score (%) | |
| DS U-Net [19] | 79.00 | 83.40 | 82.70 | 85.70 | NA | NA |
| AUNet [20] | 90.12 | 86.51 | 89.03 | 82.65 | 89.44 | 80.87 |
| UNet [21] | 92.14 | 88.23 | 90.47 | 84.79 | 89.11 | 80.21 |
| Connected-UNets [22] | 94.45 | 89.72 | 90.66 | 85.81 | 90.41 | 81.33 |
| SegNet [23] | 92.01 | 88.77 | 90.52 | 85.30 | 88.49 | 81.97 |
| Connected-SegNets | 96.34 | 91.21 | 92.86 | 87.34 | 92.25 | 83.71 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alkhaleefah, M.; Tan, T.-H.; Chang, C.-H.; Wang, T.-C.; Ma, S.-C.; Chang, L.; Chang, Y.-L. Connected-SegNets: A Deep Learning Model for Breast Tumor Segmentation from X-ray Images. Cancers 2022, 14, 4030. https://doi.org/10.3390/cancers14164030
Alkhaleefah M, Tan T-H, Chang C-H, Wang T-C, Ma S-C, Chang L, Chang Y-L. Connected-SegNets: A Deep Learning Model for Breast Tumor Segmentation from X-ray Images. Cancers. 2022; 14(16):4030. https://doi.org/10.3390/cancers14164030
Chicago/Turabian StyleAlkhaleefah, Mohammad, Tan-Hsu Tan, Chuan-Hsun Chang, Tzu-Chuan Wang, Shang-Chih Ma, Lena Chang, and Yang-Lang Chang. 2022. "Connected-SegNets: A Deep Learning Model for Breast Tumor Segmentation from X-ray Images" Cancers 14, no. 16: 4030. https://doi.org/10.3390/cancers14164030
APA StyleAlkhaleefah, M., Tan, T.-H., Chang, C.-H., Wang, T.-C., Ma, S.-C., Chang, L., & Chang, Y.-L. (2022). Connected-SegNets: A Deep Learning Model for Breast Tumor Segmentation from X-ray Images. Cancers, 14(16), 4030. https://doi.org/10.3390/cancers14164030










