Performance and Phenotypic Stability of Norway Spruce Provenances, Families, and Clones Growing under Diverse Climatic Conditions in Four Nordic Countries
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials, Trials, and Measurements
2.2. Measures of Phenotypic Stability
2.3. Calculations and Statistical Analyses
2.3.1. Provenance Data
2.3.2. Family Data
2.3.3. Clonal Data
3. Results
3.1. Mortality and Stem Defects
3.2. Height
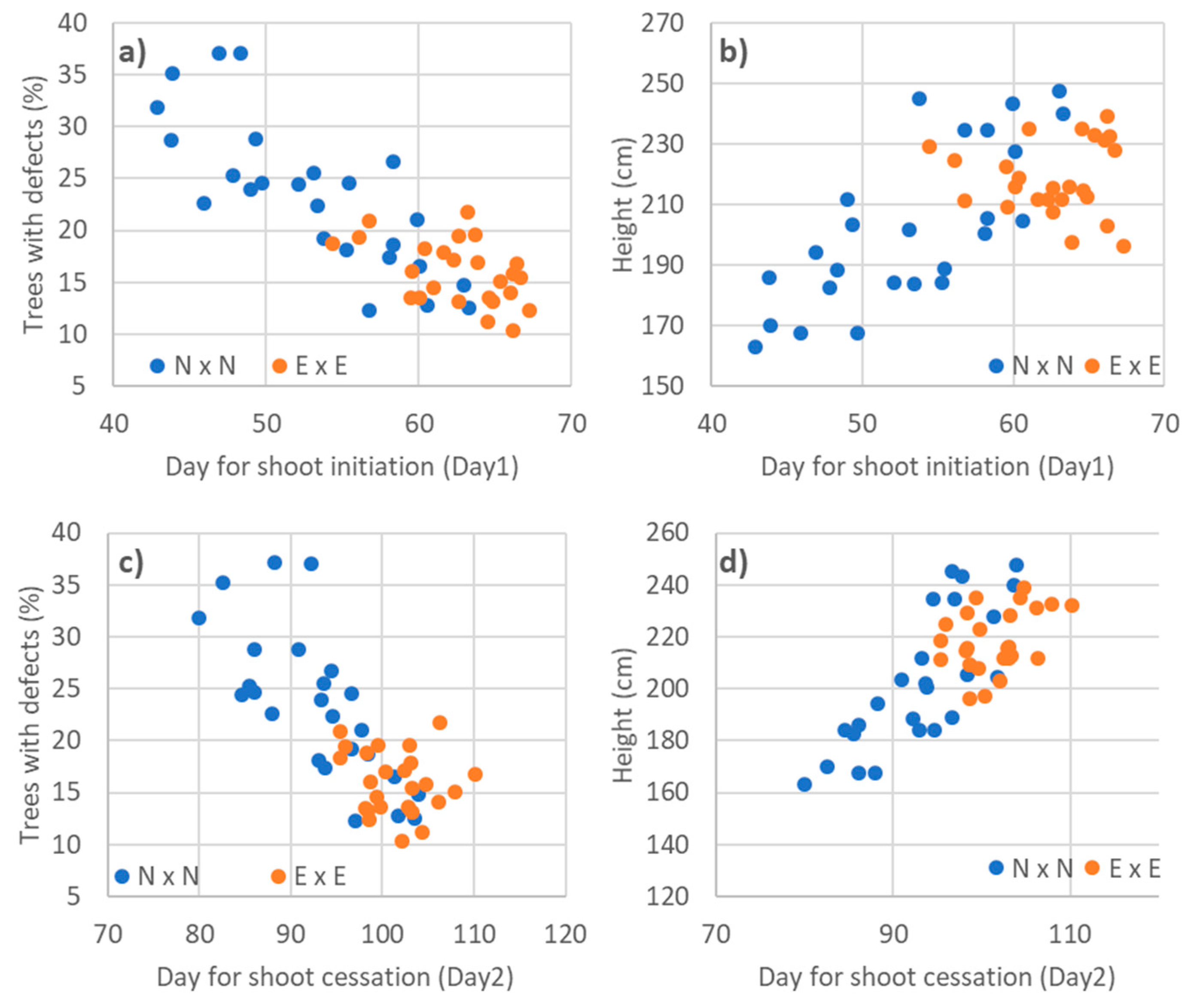
3.3. Relationships to Phenology Traits
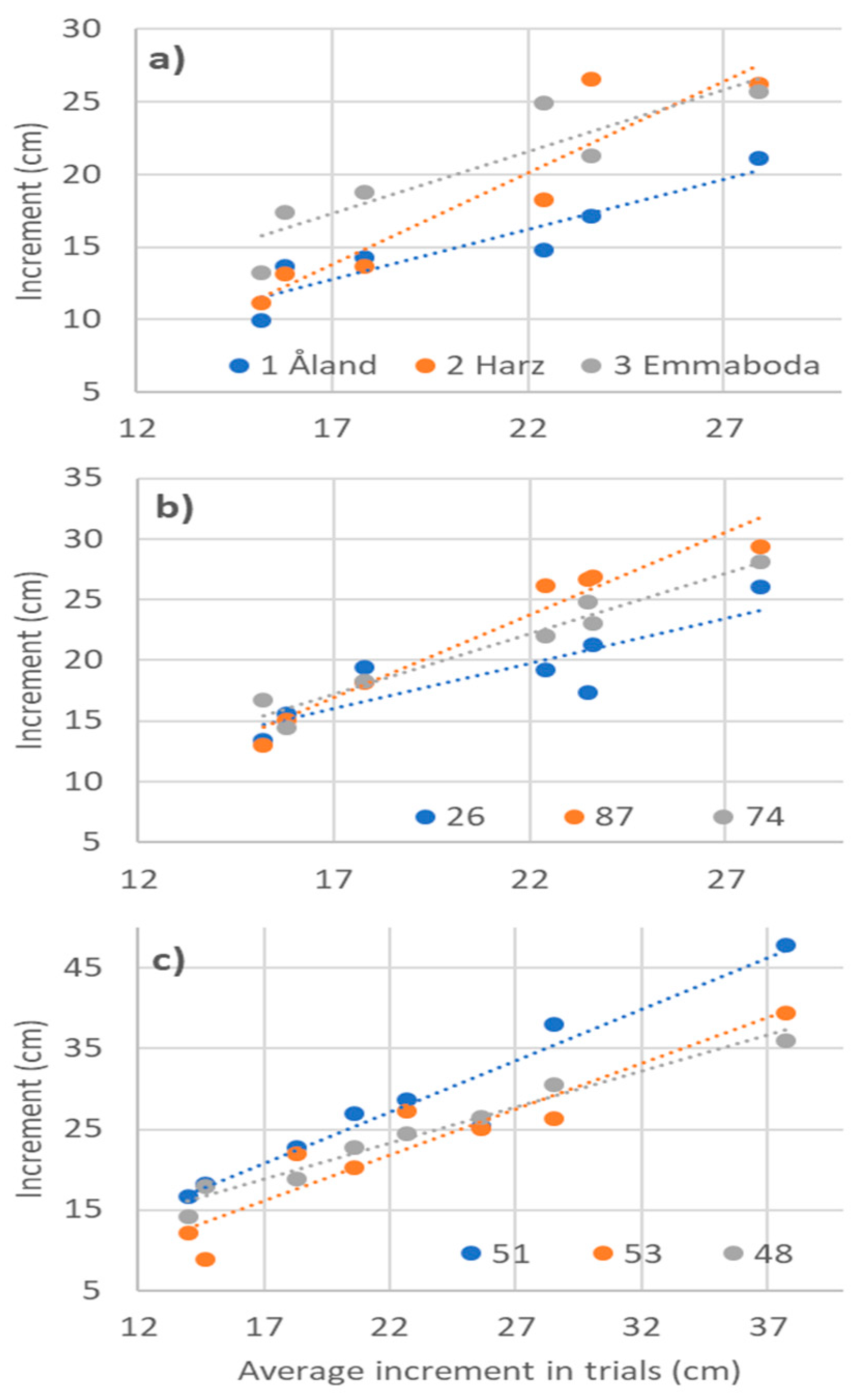
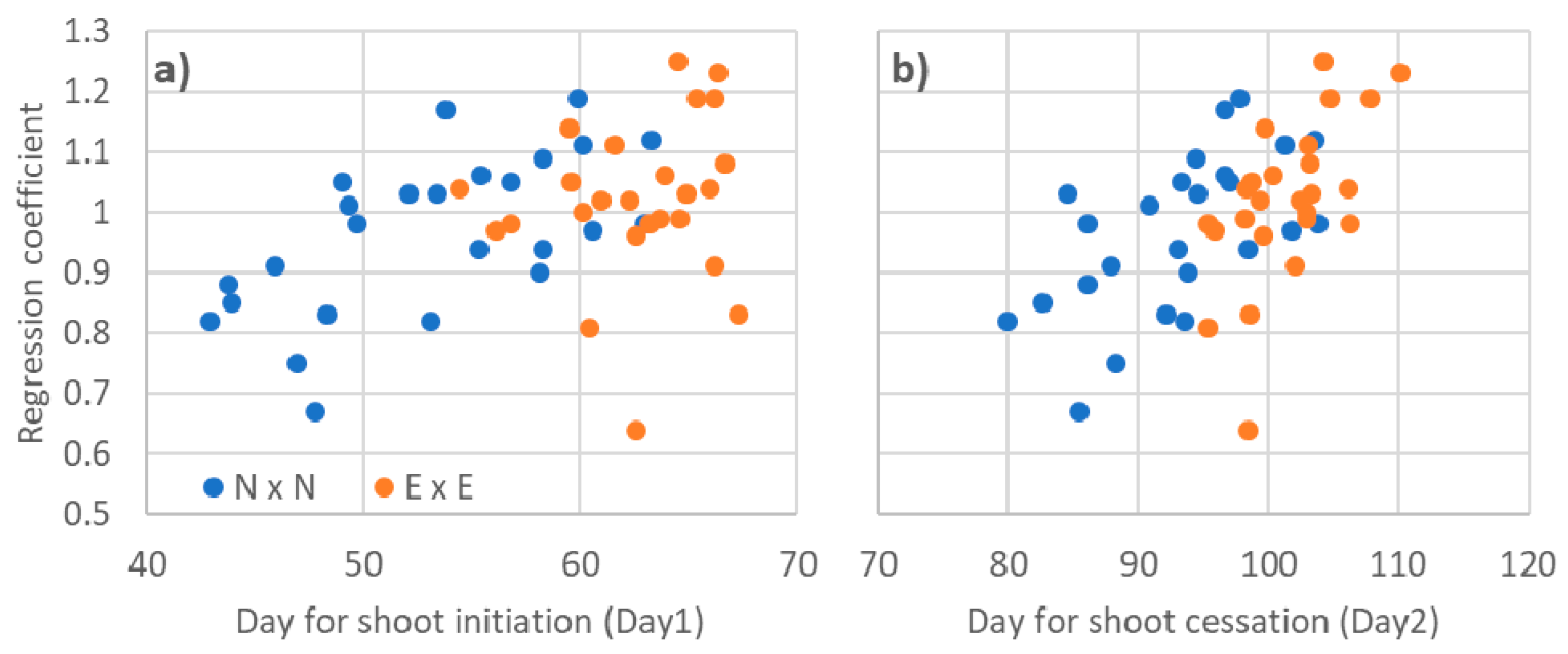
3.4. Phenotypic Stability Parameters
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Becker, H.; Leon, J. Stability analysis in plant breeding. Plant Breed. 1988, 101, 1–23. [Google Scholar] [CrossRef]
- Berlin, M.; Jansson, G.; Högberg, K.A. Genotype by environment interaction in the southern Swedish breeding population of Picea abies using new climatic indices. Scand. J. For. Res. 2015, 30, 112–121. [Google Scholar] [CrossRef]
- Persson, B.; Persson, A. Survival and Quality of Norway Spruce (Picea abies (L.) Karst.) Provenances at the Three Swedish Sites of the IUFRO 1964/68 Picea abies Provenance Experiment; 1992 Report 29; Dept for Yield Research, SLU: Garpenberg, Sweden, 1992.
- Skrøppa, T. A critical evaluation of methods available to estimate the genotype × environment interactions. Studia For. Suec. 1984, 166, 3–14. [Google Scholar]
- Hannerz, T.; Sonesson, J.; Ekberg, I. Genetic correlations between growth and growth rhythm observed in short-tern test and performance in long-term field trials of Norway spruce. Can. J. For. Res. 1999, 29, 768–778. [Google Scholar] [CrossRef]
- Benzer, B.; Foster, G.S.; Hellberg, A.R.; Podzorski, A.C. Genotype × environment interaction in Norway spruce involving three levels of genetic control: Seed source, clone mixture and clone. Can. J. For. Res. 1988, 18, 1172–1181. [Google Scholar] [CrossRef]
- Karlsson, B.; Wellendorf, H.; Roulund, H.; Werner, M. Genotype x trial interaction and stability across sites in 11 combined provenance and clone experiments with Picea abies in Denmark and Sweden. Can. J. For. Res. 2001, 31, 1826–1836. [Google Scholar] [CrossRef]
- Chen, Z.Q.; Karlsson, B.; Wu, H.X. Patterns of additive genotype-by-environment interaction in tree height of Norway spruce in southern and central Sweden. Tree Genet. Genomes 2017, 13–25. [Google Scholar] [CrossRef]
- Berlin, M.; Jansson, G.; Högberg, K.A.; Helmersson, A. Analysis of non-additive genetic effects in Norway spruce. Tree Genet. Genomes 2019, 15, 42. [Google Scholar] [CrossRef]
- Skrøppa, T.; Steffenrem, A. Selection in a provenance trial of Norway spruce. (Picea abies L. Karst) produced a land race with desirable properties. Scand. J. For. Res. 2016, 31, 439–449. [Google Scholar] [CrossRef]
- Langlet, O. Mellaneuropeiska granprovenienser i svenske skogsbruk. [Central European provenances in Swedish forestry]. Kungl. Skogs och Lantbr. Tidskrift. 1960, 99, 259–329. [Google Scholar]
- Solvin, T.M.; Steffenrem, A. Modelling the epigenetic response of increased temperature during reproduction on Norway spruce phenology. Scand. J. For. Res. 2019, 34, 83–93. [Google Scholar] [CrossRef]
- Bärring, L.; Berlin, M.; Andersson Gull, B. Tailored climate indices for climate-proofing operational forestry applications in Sweden and Finland. Int. J. Climatol. 2017, 123–142. [Google Scholar] [CrossRef]
- Krutzsch, P. Norway Spruce Development of Buds; IUFRO S2.02.11; The Royal College of Forestry: Stockholm, Sweden, 1973. [Google Scholar]
- Lynch, M.; Walsh, B. Genetics and Analysis of Quantitative Traits; Sinauer Associates, Inc.: Sunderland, MA, USA, 1998. [Google Scholar]
- Tveite, B. Bonitetskurver for gran. [Site-index curves for Norway spruce]; Meddelelser fra Norsk Institutt for Skogforskning: Ås, Norway, 1977. [Google Scholar]
- SAS Institute. SAS/STAT User’s Guide; Version 9; SAS Institute: Cary, CA, USA, 2003. [Google Scholar]
- Mullin, T.J.; Morgenstern, E.K.; Park, Y.S.; Fowler, D.P. Genetic parameters from a clonally replicated test of black spruce (Picea mariana). Can. J. For. Res. 1992, 22, 24–36. [Google Scholar] [CrossRef]
- Russel, J.; Costa e Silva, J.; Baltunis, B.S. Multisite genetic parameter estimates from a Callitropsis kookiness diallel study with clonally replicated progeny. Can. J. For. Res. 2015, 45, 689–697. [Google Scholar] [CrossRef]
- Fottland, H.; Skrøppa, T. The IUFRO 1964/68 provenance experiment with Norway spruce (Picea abies) in Norway: Variation in mortality and height growth. Meld. Skogforsk 1989, 43, 1–30. [Google Scholar]
- Falconer, D.S.; Mackey, T. Introduction to Quantitative Genetics, 4th ed.; Pearson Education Ltd.: Harlow, UK, 1996. [Google Scholar]
- Shelbourne, C.J.A. Genotype-environment interaction: Its study and its implications in forest tree improvement. Proc. IUFRO Genet. SABRAO Joint Seem Tokyo 1972, B-1, 1–28. [Google Scholar]
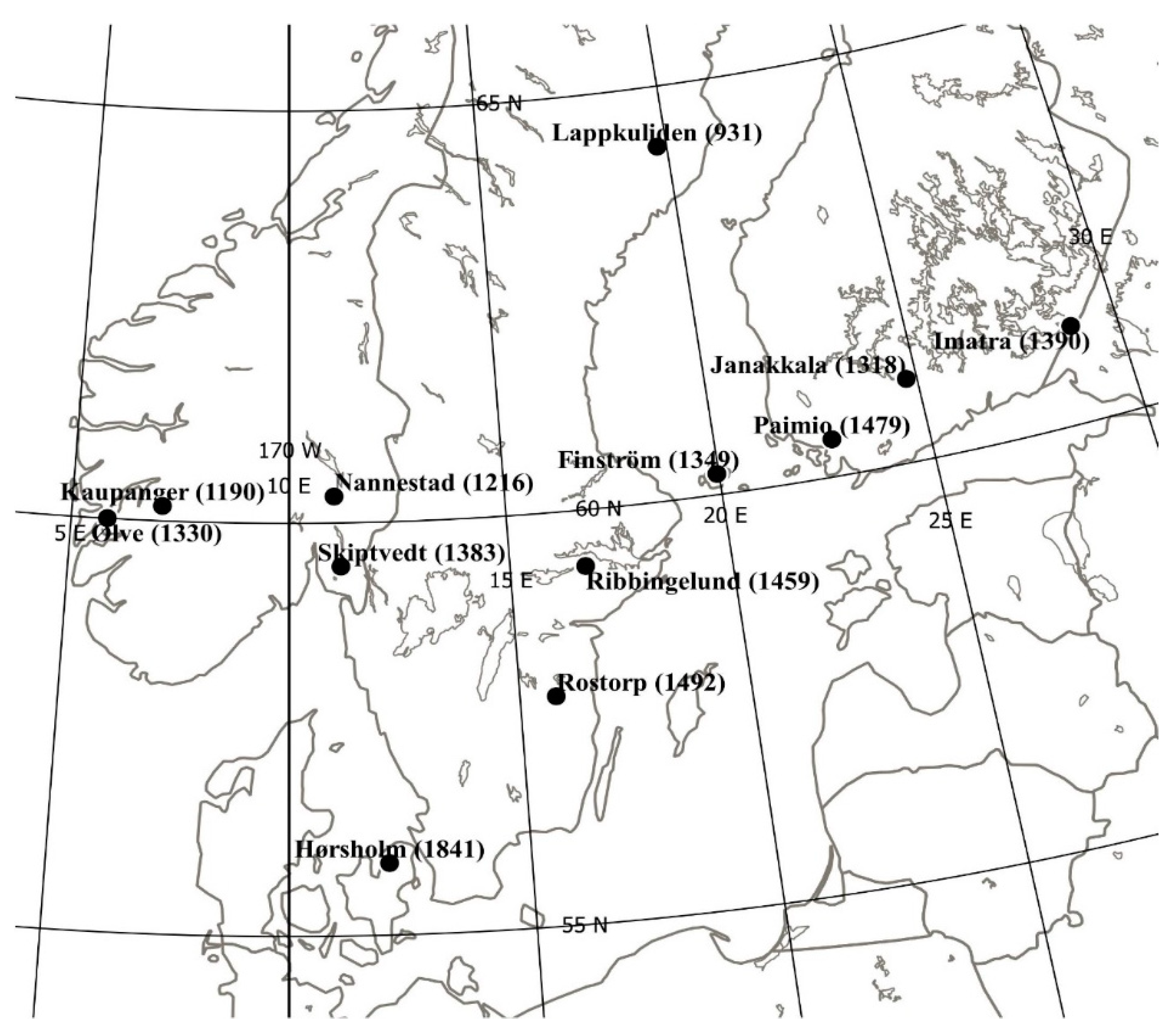
| Trial and Location | Latitude °N | Longitude °E | Altitude M | Mortality % | Height cm | Stem Defects % |
|---|---|---|---|---|---|---|
| 1 Ølve | 60°00′ | 5°50′ | 50 | 10.2 | 260 ** | 8.9 |
| 2 Skiptvedt | 59°28′ | 11°11′ | 100 | 24.8 | 224 | 15.5 |
| 3 Rostorp | 57°47′ | 15°47′ | 165 | 66.0 | 235 | 18.1 |
| 4 Lappkuliden | 64°16′ | 19°37′ | 190 | 52.8 | 174 ** | 19.3 |
| 5 Paimio | 60°27′ | 22°44′ | 40 | 4.1 | 279 | 21.7 |
| 6 Janakkala | 61°00′ | 24°45′ | 130 | 27.6 | 178 | 16.0 |
| 7 Imatra | 61°10′ | 28°53′ | 80 | 42.9 | 152 | 26.9 |
| 11 Kaupanger | 61°11′ | 7°05′ | 80 | 8.0 | 183 | 10.0 |
| 12 Nannestad | 60°19′ | 11°05′ | 200 | 22.1 | 256 | 14.5 |
| 13 Ribbingelund | 59°19′ | 16°42′ | 50 | 31.9 | 206 | 11.5 |
| 14 Lappkuliden | 64°1′ | 19°37′ | 190 | 33.2 | 147 | 37.0 |
| 15 Hørsholm | 55°52′ | 12°04′ | 50 | 14.7 | 377 | 31.9 |
| 16 Finström | 60°15′ | 19°57′ | 5 | 24.3 | 227 | 27.4 |
| 17 Paimio | 60°27′ | 22°44′ | 40 | 4.5 | 257 * | 36.1 |
| 18 Imatra | 61°10′ | 28°53′ | 80 | 28.9 | 126 * | 39.0 |
| Famale Parents | Male Parents | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Norwegian (N) | Eastern European (E) | ||||||||||
| 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | ||
| Norwegian (N) | 1 | X | X | X | X | X | X | X | X | X | X |
| 2 | X | X | X | X | X | X | X | X | X | X | |
| 3 | X | X | X | X | X | X | X | X | X | X | |
| 4 | X | X | X | X | X | X | X | X | X | X | |
| 5 | X | X | X | X | X | X | X | X | X | X | |
| Eastern European (E) | 6 | X | X | X | X | X | X | X | X | X | X |
| 7 | X | X | X | X | X | X | X | X | X | X | |
| 8 | X | X | X | X | X | X | X | X | X | X | |
| 9 | X | X | X | X | X | X | X | X | X | X | |
| 10 | X | X | X | X | X | X | X | X | X | X | |
| Source/Cross | N × N | N × E | E × N | E × E |
|---|---|---|---|---|
| Mean Height (cm) | 205 | 221 | 209 | 219 |
| F | 215.5 (0.0002) | 81.3 (0.0002) | 0.0 | 0.0 |
| M | 476.8 (<0.0001) | 59.5 (0.007) | 423.3 (<0.0001) | 105.2 (0.001) |
| F × M | 38.5 (0.07) | 73.5 (0.0012) | 22.3 (0.20) | 26.0 (0.05) |
| F × S | 0.0 | 69.5 (0.0009) | 88.8 (<0.0001) | 66.3 (<0.0001) |
| M × S | 71.9 (0.01) | 114.7 (<0.0001) | 117.5 (<0.0001) | 47.7 (0.004) |
| F × M × S | 101.2 (0.002) | 2.7 (0.54) | 0.0 | 0.0 |
| Error | 5926.4 | 6559.1 | 6066.1 | 5861.8 |
| h2 | 0.20 (0.15) | 0.03 (0.04) |
| Source | N × N | N × E | E × N | E × E |
|---|---|---|---|---|
| Mean height (cm) | 169 | 178 | 194 | 187 |
| Family | 768.3 (<0.0001) | 651.2 (<0.0001) | 191.8 (0.05) | 269.6 (0.003) |
| Family × site | 164.3 (<0.0001) | 84.5 (0.01) | 279.0 (<0.0001) | 200.5 (<0.0001) |
| Clone (family) | 503.5 (<0.0001) | 462.3 (<0.0001) | 755.1 (<0.0001) | 373.1 (<0.0001) |
| Clone (family) × site | 141.7 (0.01) | 292.9 (<0.0001) | 174.1 (0.04) | 131.6 (0.04) |
| Error | 3291.8 | 3451.2 | 4221.4 | 3333.1 |
| H2 | 0.26 (0.20) | 0.15 (0.11) |
| Male Female | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | Mean |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 1.05 | 0.94 | 1.03 | 0.88 | 0.98 | 1.00 | 0.93 | 0.86 | 1.14 | 0.75 | 0.96 |
| 2 | 1.09 | 1.19 | 0.94 | 0.67 | 1.03 | 1.00 | 1.15 | 1.22 | 0.89 | 0.92 | 1.01 |
| 3 | 0.75 | 0.82 | 0.91 | 0.85 | 0.82 | 0.74 | 0.86 | 0.98 | 1.03 | 0.79 | 0.86 |
| 4 | 1.17 | 1.11 | 1.06 | 0.83 | 1.01 | 0.94 | 1.02 | 1.20 | 1.19 | 1.04 | 1.06 |
| 5 | 0.98 | 1.12 | 0.97 | 1.05 | 0.90 | 0.69 | 1.05 | 1.15 | 0.87 | 1.04 | 0.98 |
| 6 | 0.93 | 0.77 | 0.99 | 1.01 | 0.79 | 0.81 | 0.64 | 1.19 | 0.97 | 0.96 | 0.91 |
| 7 | 1.07 | 1.05 | 0.85 | 0.97 | 0.99 | 0.98 | 1.14 | 1.25 | 1.04 | 1.05 | 1.04 |
| 8 | 1.20 | 1.06 | 0.88 | 1.07 | 0.98 | 0.98 | 1.02 | 1.23 | 1.00 | 1.06 | 1.05 |
| 9 | 1.44 | 1.36 | 1.01 | 1.20 | 1.01 | 1.08 | 0.99 | 1.19 | 1.11 | 1.03 | 1.14 |
| 10 | 1.23 | 1.07 | 0.96 | 0.89 | 0.78 | 0.83 | 0.99 | 1.04 | 1.02 | 0.91 | 0.97 |
| Mean | 1.09 | 1.05 | 0.96 | 0.94 | 0.93 | 0.90 | 0.98 | 1.13 | 1.02 | 0.96 | 1.00 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Skrøppa, T.; Steffenrem, A. Performance and Phenotypic Stability of Norway Spruce Provenances, Families, and Clones Growing under Diverse Climatic Conditions in Four Nordic Countries. Forests 2021, 12, 230. https://doi.org/10.3390/f12020230
Skrøppa T, Steffenrem A. Performance and Phenotypic Stability of Norway Spruce Provenances, Families, and Clones Growing under Diverse Climatic Conditions in Four Nordic Countries. Forests. 2021; 12(2):230. https://doi.org/10.3390/f12020230
Chicago/Turabian StyleSkrøppa, Tore, and Arne Steffenrem. 2021. "Performance and Phenotypic Stability of Norway Spruce Provenances, Families, and Clones Growing under Diverse Climatic Conditions in Four Nordic Countries" Forests 12, no. 2: 230. https://doi.org/10.3390/f12020230
APA StyleSkrøppa, T., & Steffenrem, A. (2021). Performance and Phenotypic Stability of Norway Spruce Provenances, Families, and Clones Growing under Diverse Climatic Conditions in Four Nordic Countries. Forests, 12(2), 230. https://doi.org/10.3390/f12020230






