Abstract
Currently, advances in healthcare technologies are transforming medical diagnostics, particularly for data-driven disease detection. Acute lymphoblastic leukemia is a common and life-threatening blood cancer, especially prevalent in children. Although existing artificial intelligence-based diagnostic models have demonstrated promising accuracy, their black-box nature limits transparency, explainability, and clinical trust. Moreover, most explainable artificial intelligence approaches for leukemia diagnosis rely primarily on qualitative visual explanations and lack a unified quantitative mechanism to measure trust. This study presents a novel, trust-centered explainable deep-learning framework for automated acute lymphoblastic leukemia detection using 153 publicly available microscopic blood smear images and multiple transfer-learning models. Among the evaluated architectures, the fine-tuned EfficientNetB4 achieved high diagnostic performance, attaining an accuracy of 98.31%, and was selected as the base model of the proposed framework. Beyond diagnostic accuracy, this work introduces a novel unified interpretability score that quantitatively assesses model trustworthiness by integrating diagnostic performance, explanation robustness (evaluated using deletion and stability metrics), and clinician trust feedback into a single reliability measure. This quantitative trust formulation represents a distinct advancement over existing studies based on explainable artificial intelligence, which typically rely on isolated or qualitative explainability assessments. The framework further enhances transparency through visual and textual explanations generated using standard post hoc explainable methods and advanced fusion heatmaps, while human-centric evaluation is conducted using two clinician-based trust scales. Overall, this study provides a unified and transparent framework that jointly evaluates accuracy, explainability, and clinician trust, representing a step towards the clinical adoption of trustworthy artificial intelligence driven leukemia diagnostic systems.
1. Introduction
White blood cell cancer is one of the most common diseases that affects a large population worldwide. One of the most common types of white blood cell cancers is leukemia, which may hamper the ability of one’s body to produce healthy blood cells. Acute Lymphoblastic Leukemia (ALL), a common type of leukemia, arises from B- and T–cell precursors in the lymphoid tissues. B- and T-lymphocytes are special types of White Blood Cells (WBCs) [1]. These are defense cells that play a crucial role in protecting the body against various infections and diseases. ALL is a frequently occurring blood cancer that can affect any age group, but is most common in young children. The primary malfunction is the abnormal proliferation of WBCs, which destroy healthy cells and disrupt normal bone marrow function, leading to the formation of many immature cells. Rapid proliferation of abnormal/atypical lymphocyte cells reduces the number of healthy new blood cells, thereby increasing the risk of morbidity and mortality among patients. ALL can ultimately be life-threatening if not detected and cured at an early stage.
Conventional diagnostic approaches to ALL are labor-intensive and rely on microscopic examination of blood samples to detect leukemic features. However, recent methods have successfully applied several Deep-Learning (DL) architectures to automate this process and improve the accuracy. Although these models are efficient, they are black-boxes, meaning the end users cannot easily understand the reasoning behind the model’s decisions.
This research used several sophisticated DL architectures to implement automated ALL detection. In particular, we used pre-trained models VGG19, InceptionV3, DenseNet121, and EfficientNetB4 to classify microscopic blood images. With its deeper architecture, VGG19 is a popular choice for feature extraction in medical imaging [2], and InceptionV3 improves performance using an inception module to capture multiscale features [3]. The architecture of DenseNet121 is highly effective due to its densely connected layers, which allow the network to efficiently learn complex features from input images [4]. The EfficientNetB4, meanwhile, has been widely adopted for medical classification tasks due to its efficient parameter usage and excellent scalability on large datasets [5]. Although these models achieve high accuracy, they are considered black-box models and, therefore, difficult to interpret for deliberate reasoning.
In the last decade, the significant (medical) black-box nature of such DL models sparked extensive research on eXplainable Artificial Intelligence (XAI) to make these models more interpretable [6,7]. XAI was a concept launched by the Defense Advanced Research Projects Agency (DARPA) to provide solutions for black-box models based on Artificial Intelligence (AI) and understand how these systems reach their conclusions. If an AI-based system is to be accepted in fields where human lives are on the line, it is crucial to know how and why the system made a particular decision. The European Union (EU) had gone so far as to make providing a rationale behind AI-driven decisions a legal mandate, again underscoring the need for explainability in AI-based systems [8].
Explaianability is an essential phenomenon for AI-driven healthcare, shaped by XAI, which makes the decision-making process understandable by elucidating the rationale behind the predictions [9].
In addition, the adoption of AI-driven solutions in healthcare mainly depends on the fairness of the model. The main goal of implementing XAI in medical AI systems is to build trust between clinicians and patients. This is achieved by offering clear explanations for the model’s predictions. This kind of approach can really help with clinical decision-making in practice. However, existing XAI-based leukemia studies largely emphasize diagnostic accuracy or qualitative visual explanations, without providing a unified quantitative mechanism to evaluate clinical trust.This research aims to bridge the gap between the transparency of the AI system and its clinical usability in diagnosing leukemia. The focus of the proposed study is to develop a comprehensive framework that not only provides diagnostic accuracy but also instills trust in the explanations generated by multiple XAI methods, providing insight into the classification rationale for ALL images. Three distinctive XAI techniques, including Local Interpretable Model-agnostic Explanations (LIME) [10], Gradient-Weighted Class Activation Mapping (Grad-CAM) [11], and Guided Gradient-Weighted Class Activation Mapping (Guided Grad-CAM) [12], are explored and implemented in the proposed framework. In addition, these XAI methods have been fused using four distinct fusion strategies to generate combined heatmaps, providing a consolidated and deeper insight into the rationale behind the models’ predictions.
The research focuses on implementing a DL framework that can accurately detect ALL and integrate explainability to get transparent and understandable explanations for its predictions. This work may help clinicians to make informed decisions and to improve their trust in AI-based healthcare tools. Furthermore, the explainability provided by the proposed framework has been validated with the evaluation of Stability and Deletion metrics. Collectively, these metrics assess the reliability and robustness of the model’s explanations under noisy conditions, a common challenge for medical systems. The novelty of the framework lies in quantifying the system’s Interpretability Score, which represents its trust and reliability. The Interpretability Score unifies the underlying DL model’s objective evaluation, the evaluation of the explanations generated by the system, and the feedback from medical professionals into a single real clinical trust score.
This quantitative trust formulation represents a distinct contribution beyond existing XAI approaches, which typically assess performance and explainability in isolation. The explanations provided by our model are offered to patients in the form of patient reports. It will provide clinicians with preliminary reasoning behind the predictions, thereby ensuring trust, fairness, and transparency in AI-based diagnostic tools. These reports provide clinicians with clear and interpretable reasoning behind model predictions, thereby supporting transparent and informed clinical decision-making.
The remainder of the paper is organized as follows: Section 2 reviews the existing literature on the diagnosis of ALL and the use of XAI. In Section 3, the proposed methodology is discussed in detail. Section 4 presents the details of the implementation, whereas Section 5 presents the results of the experiment. Section 6 provides a comprehensive discussion of the results along with practical and ethical considerations of this research. The paper concludes in Section 7 with a brief discussion on possible research dimensions for the future.
1.1. Problem Statement
Current AI systems for cancer detection face many challenges. They tend to be opaque, making them “black-box models”. The lack of clarity in these AI setups prevents their use in real medical practice. Doctors and other professionals usually need to understand how decisions are made. This is especially true when it comes to diagnostic tools. Models that work like “black boxes” provide little insight into their own processes, limiting their use in clinical practice [13].
1.2. Research Objectives
In this study, a transfer-learning approach is evaluated to enhance the performance of the automated detection system for ALL microscopic images. The learned features from the pretrained models (trained on image datasets containing millions of images) can be fine-tuned for the specific task of ALL detection. The problem of a relatively limited image dataset will be addressed.
The objectives of this research are listed below.
- Develop a viable AI-based model for the accurate and diagnostic classification of leukemia cells using DL architectures.
- Integration of different XAI techniques in such a way as to improve interpretability and transparency in AI-based predictions.
- Evaluating the performance of the proposed model using some standard Machine-Learning (ML) metrics that include accuracy, F1-score, and AUC-ROC.
- Evaluate the trust and explainability of the underlying DL model by introducing Interpretability Score metrics.
- The trustworthiness and usability of the explanations are also assessed using annotations and feedback provided by medical professionals and other end users.
- Ethical considerations in AI-driven leukemia detection are addressed in a way that ensures fairness and reliability in medical applications.
1.3. Scope of the Study
The scope of the proposed research is described below.
- In this study, a comparison of four state-of-the-art DL architectures, DenseNet121 [14], EfficientNetB4 [15], InceptionV3 [16], VGG19 [2], has been presented. This achieves better accuracy and efficiency. The problem of limited medical image data has been addressed by using pre-trained models, thereby improving the model’s generalizability.
- Some existing XAI methodologies, such as LIME, Grad-CAM, and Guided Grad-CAM, along with four distinct but advanced heatmaps, provide explanations for predicted classification in a consolidated and robust manner.
- The performance evaluation of this framework comprises two levels of validation: human-centric and objective assessment.
2. Literature Review
2.1. Leukemia Detection Using DL and XAI
In medical diagnostics, much attention has been paid to detecting leukemia, particularly ALL. Image-based ALL detection is complex, and early detection is more valuable. DL models, particularly Convolutional Neural Network (CNN) variants, have recently shown significant promise in automating leukemia detection from microscopic blood images. However, because of their black-box nature, medical professionals tend not to prefer them in clinical practice, as they do not trust the predictions of these models. To address this challenge, many research studies have been presented that combine XAI techniques with DL models to improve explainability and trust in these models.
2.2. Previous Approaches to Leukemia Detection Using AI
Many studies have used CNN-based models to detect leukemia with promising diagnostic performance. Thiriveedhi et al. presented a CNN model called ALL-Net that implemented the CAM XAI approach to introduce explainability into the system. Their proposed model achieved 93.4% accuracy for diagnosing ALL [17]. Moreover, Pervez et al. implemented a Hierarchical-Federated-Learning (HFL) approach that integrates multiple XAI techniques. Their proposed model achieved an accuracy of 96.2% [18]. In another study, CNN-based ensemble models were used to classify ALL images, achieving 96.2% accuracy [19]. The study also focused on the usability of the ensemble models to achieve improved generalization and reduction of bias. Bohmrah and Kaur implemented hybrid DL models for disease detection. Multiple datasets were used in the study, yielding varying accuracies. The authors also focused on the need to incorporate explainability to enhance trust and fairness in the underlying models [20]. Furthermore, Shen et al. designed fine-tuned pre-trained DL models based on the concept of domain adaptation; they implemented ResNet and VGG19 to detect ALL cells from healthy cells [21]. The proposed model achieved 94.8% accuracy, demonstrating that these models can perform better on medical image classification tasks, especially in cases with limited datasets. In another study, Salman et al. explored multimodal DL by integrating clinical data with microscopic image analysis, reporting an accuracy of 92.7%. The underlying model presented robustness and accuracy for leukemia diagnosis [22].
A significant research gap in these studies is the lack of trust in the underlying ML models. These models achieve diagnostic accuracy but often lack explainability, a crucial factor for clinical adoption. This lack of explainability and interpretability is directly related to trust and fairness, posing a challenge because clinicians require justification and reasoning behind AI decisions, especially in life-threatening diseases like ALL.
2.3. XAI Techniques in Leukemia Detection
XAI has gained a fair amount of traction, especially in recent years, in the domain of ALL, to address the black-box nature of the DL models and enhance trust and interpretability in clinical practice. A custom-built CNN, an XAI-enhanced version called ALL-Net, has been shown to detect ALL with high accuracy from blood smear images, with visual explanations provided by heatmaps and feature importance. Such findings further substantiate the model’s transferability and clinical relevance, with performance accuracy of 98.38% in multicenter datasets [23]. Similarly, hybrid frameworks that combine HFL with XAI are also presented, aiming to develop privacy-preserving methods for ALL classification and introducing features such as saliency maps and occlusion sensitivity to generate interpretable predictions on histopathological cell images. Such systems support the research areas mentioned above to advance transparent AI in oncology, where regulatory issues such as GDPR, as well as clinician uptake, are paramount [18]. The enhanced developments comprise decision-support systems that integrate DL with XAI for segmentation and classification of ALL using LIME and Grad-CAM-based techniques to explain model decisions on microscopic blood samples, thus enhancing diagnostics in resource-poor settings [24]. Another interesting approach is to use an attention-based DL model, DDRNet, which imposes hard attention mechanisms to attend to relevant cellular features in blood smears, thereby providing strong justification and reducing the risk of overfitting to noisy datasets [25].
A deep CNN ensemble model Skin-NeT, has been presented by Alshehri et al. for skin cancer diagnosis. Their framework used LIME to enhance explainability, achieving an accuracy of 85% [26]. DCEN-WCNet, Moreover, in their work, Chaddad et al. implemented adversarial models and used Projected Gradient Descent (PGD) for the model analysis and transparency. Their reported accuracy is 87.9% [27]. Although the trade-off between model accuracy and explainability in the context of medical diagnosis has not been fully explored in the literature, it has been identified as a major research gap in the field [28].
2.4. Emerging Trends in XAI for Medical Diagnostics
Recently, there has been a paradigm shift in the domain of DL, focusing on enhanced accuracy with better explainability of the underlying models. A 2.5D CNN model has been implemented by Song et al., which distinguishes between lymphoma and tuberculous lymphadenitis, achieving classification accuracy of 90.3%. They also incorporated activation mapping to build transparency and explainability [29]. Gouveia has explored the significance of incorporating explainability towards building trust, particularly for ALL detection. Their work presents a detailed analysis, suggesting an utmost need for designing ethical and reasonable AI-based systems [30].
The above-mentioned research emphasizes integrating explainability and transparency in the black-box DL models to enhance trust and to resolve data bias and inequities, ensuring fairness. The importance of XAI in healthcare applications, in general and specifically for leukemia detection, is growing rapidly, impacting DL-based classification models, Federated-Learning (FL) frameworks, and adversarial explainability techniques. Hence, integrating XAI approaches will help to bridge the gap between AI model performance and trustworthiness in medical diagnostics. Moreover, several qualitative assessment criteria have been presented, but there is a lack of unified quantitative evaluation criteria to measure the trust and reliability of the system. Table 1 provides a comprehensive comparison of the existing studies on the diagnosis of ALL.

Table 1.
Summary of Existing Studies on XAI-Based Leukemia Diagnostic Models.
2.5. Research Gap and Advanced Contributions
Most previous studies focus on single techniques or narrow datasets. Our work addresses this gap by combining multiple pretrained DL models (such as VGG19, InceptionV3, DenseNet121, and EfficientNetB4). The underlying DL architecture employs a dynamic and scalable preprocessing pipeline to apply feature engineering, including cropping and resizing, noise reduction, and data normalization to standardize the pixel intensities for the input data, handling the missing data, managing class imbalances by data augmentation using custom generators, during training, to improve the performance of the model over complex datatypes.
In addition, incorporating XAI methods improves model interpretation by introducing interactive adaptive visualizations. Although existing studies have explored XAI techniques for leukemia detection, there remains a gap in integrating multiple XAI techniques with state-of-the-art DL models to achieve comprehensive interpretability and performance.
The framework incorporates three gradient- and perturbation-based XAI methods (LIME, Grad-CAM, and Guided Grad-CAM) to provide a more multidimensional interpretation of model decisions. This allows clinicians to obtain comprehensive information about the image regions that influence the model’s predictions. In addition, it can increase confidence in the key identified features of the model and potentially uncover biases and inconsistencies specific to the method used.
Furthermore, all of the hybrid heatmap approaches that we propose offer more precise visual explanations than using any of these methods individually, and this aspect of combined interpretability has not been fully explored in the context of leukemia detection. This addresses a key challenge in interpreting AI-based medical image analysis: the trade-off between local fidelity and high-level feature relevance, offered by individual XAI methods. In the proposed model, for a given leukemic cell image, hybrid heatmaps fuse the strengths of individual XAI methods, such as LIME’s local fidelity and Grad-CAM’s coarse localization, and provide fine-grained pixel-level details via Guided Grad-CAM.
In addition, a strong correlation has been observed among visual explanations and expert’s annotations and feedback provided by the medical professionals. The framework also generates a patient report in a consolidated format, with visual and textual explanations.
A quantitative evaluation of XAI approaches has also been presented in the research. However, there was a need for real-time feedback and evaluation mechanisms from domain experts and end users, which consistently hindered explainability and interpretability. This has been addressed by presenting a comprehensive, multidimensional evaluation pipeline that assesses three aspects of model performance: robustness, efficiency, and trust.
The study also presents a unique Interpretability Score to evaluate the system’s trustworthiness and reliability. The Interpretability Score consolidates all three dimensions of clinical trust —accuracy, explainability, and clinical feedback—into a single, easy-to-read score ranging from 0 to 1. The closer to 1, the more you can rely on the AI-based model in the daily clinical environments.
To sum it all up, the novelty of the proposed framework lies in the advanced integration of all the above-mentioned contributions in a synergistic manner. To our knowledge, no existing study has implemented a framework to calculate and evaluate the system’s reliability by integrating a fine-tuned DL model with multiple established XAI methods, further exploring the insights and relevant strengths and weaknesses.
We believe that this integration in our work constitutes a multifaceted solution for transparent AI in healthcare, whose effectiveness is validated through direct comparison with significant regions identified by experts to evaluate both the quantitative and qualitative performance of the system, in terms of spatial alignment, trust, reliability, and the relevance of the explanations.
3. Proposed Methodology
This research focuses on bridging the gap between the trust in AI systems for clinical usability and advanced AI diagnostic methods. Our model provides a unified yet vital evaluation framework, i.e., accuracy, interpretability, and trust evaluation, along with objective and human-centered evaluation mechanisms.
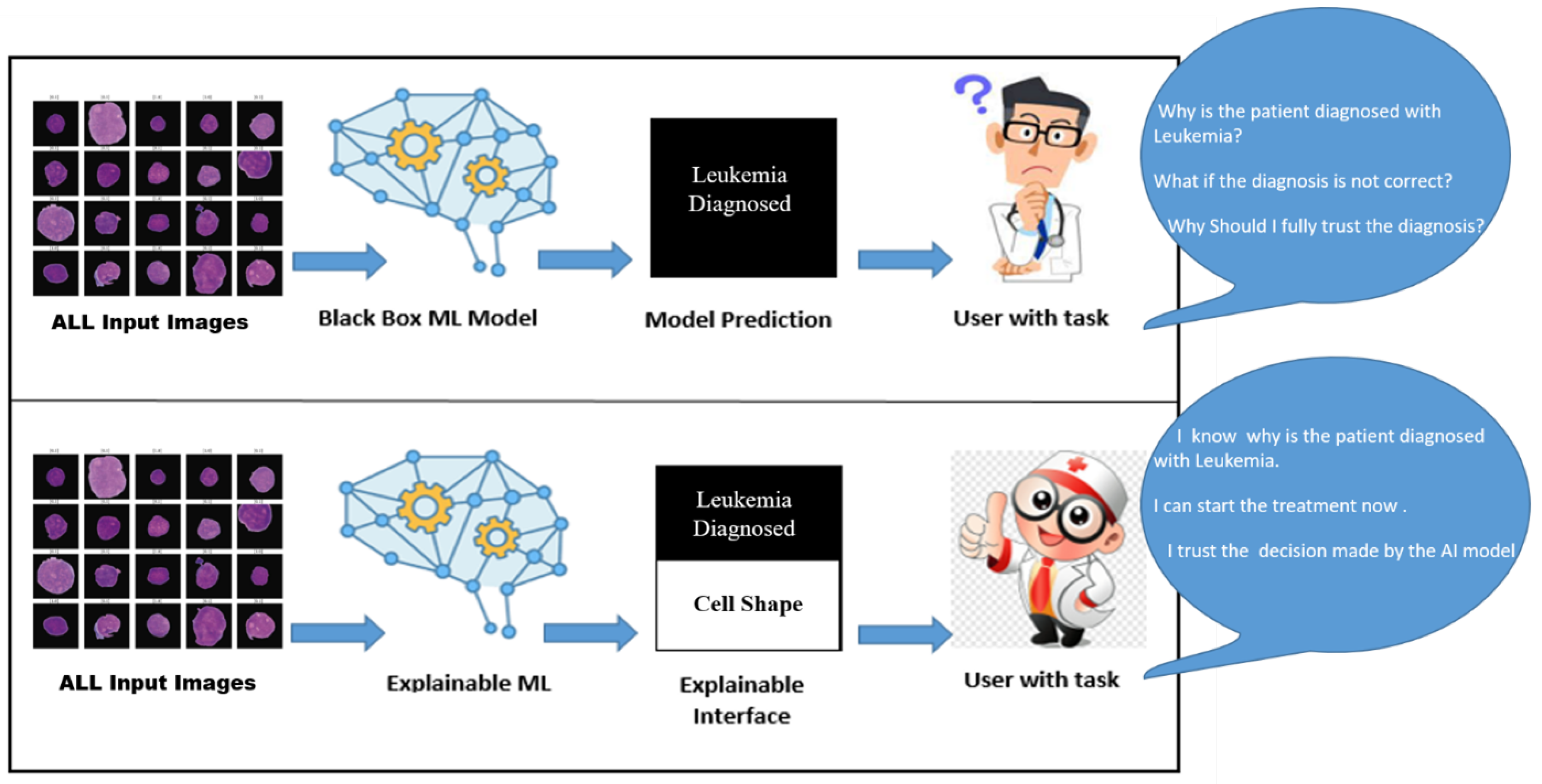
Figure 1 shows a comparative analysis of an AI-based ALL diagnostic with/without XAI. In the upper half, the AI black-box model takes input data (ALL microscopic images) and, after complex, unknown processing, produces results in ALL predictions. This result might be accurate but is vague and unclear to end users. In this case, the end user could be experts in the medical domain (oncologists, medical specialists, histopathologists), patients, pharmaceutical manufacturers, or the scientific community. They may have unanswered questions about the AI model’s trustworthiness, including verification, fairness, debugging, and model monitoring. The most important aspect is the fairness and the removal of bias in the AI model’s decisions. However, the lower half of Figure 1 shows an XAI-based model for ALL with transparent and understandable diagnosis predictions/recommendations. The XAI will help generate explanations for end users and stakeholders to answer questions like ’Why was this prediction made? Why don’t you diagnose the disease? How can we improve the results? What are the reasons for success/failure?’ and so on. This kind of understanding consequently leads to gaining trust in the model’s fairness and transparency.
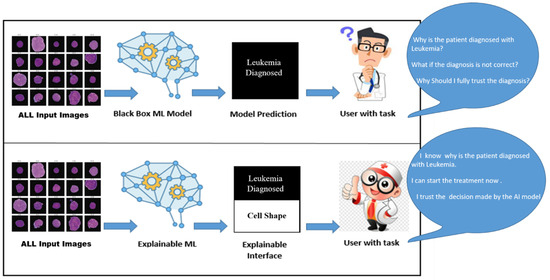
Figure 1.
Role of XAI in Medical Diagnostic of ALL.
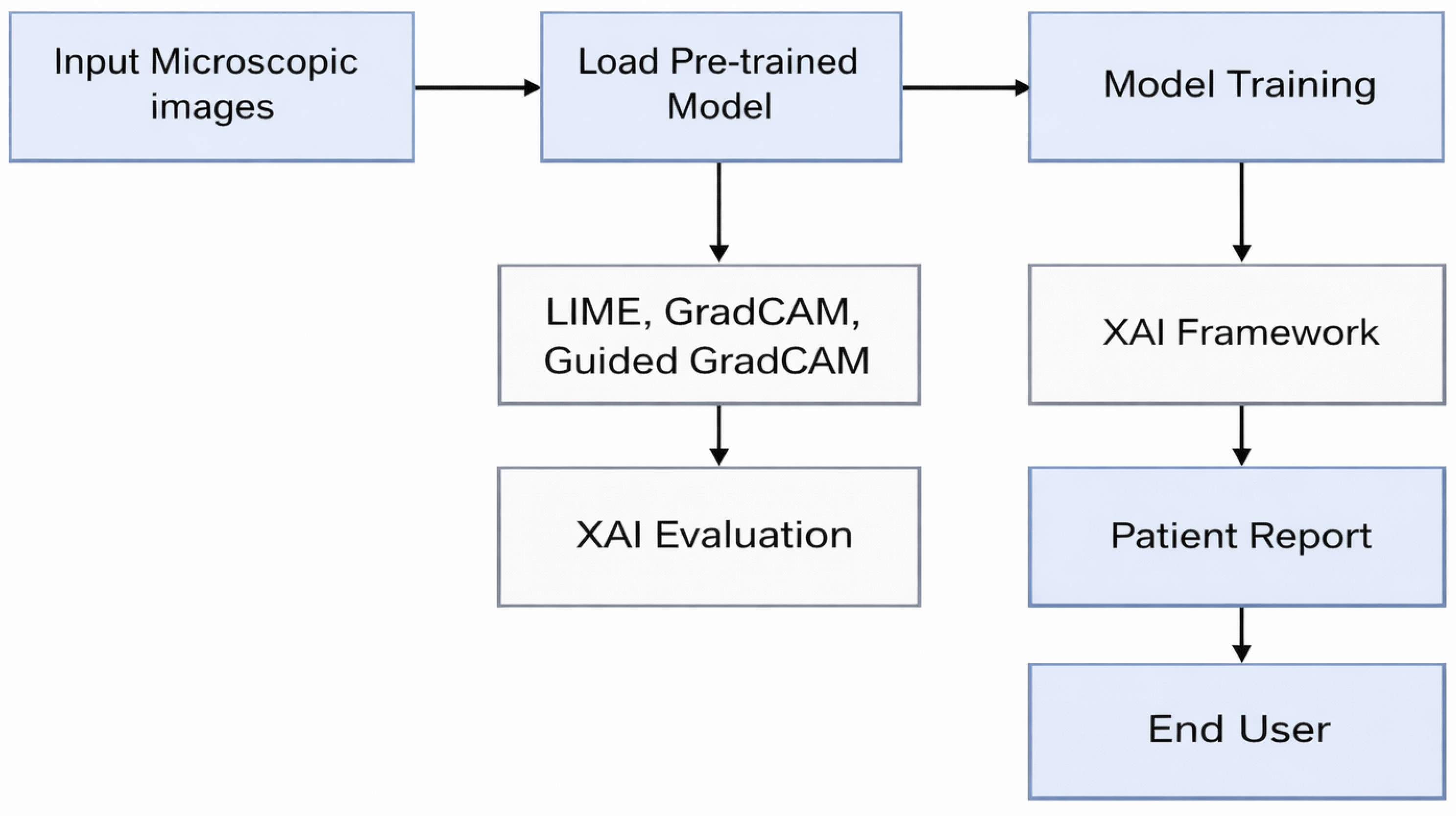
In this research, we propose an advanced framework for ALL detection that combines state-of-the-art DL methods with XAI techniques to achieve high diagnostic accuracy while providing transparency and trust evaluation of AI-assisted predictions. Figure 2 presents the flow of the proposed model.
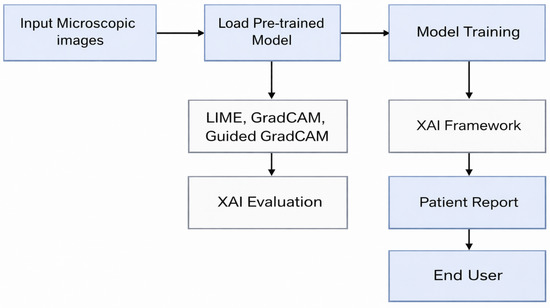
Figure 2.
Block Diagram of the Proposed Framework.
We initially built the framework using a transfer-learning approach employing multiple pretrained DL models: EfficientNetB4, VGG19, DenseNet121, and InceptionV3. These models are adapted for the ALL detection task by applying custom modifications to their architecture, specifically by replacing the thickest, final fully connected layers and fine-tuning them for task-specific output.
Additional performance metrics are reported to evaluate performance, including precision, Kappa, MCC, recall, F1 score, and AUC (Area Under the Curve). These metrics are essential for assessing the model’s performance in correctly identifying leukemia cells while avoiding false positives and negatives. This is a crucial challenge in designing systems with imbalanced datasets, a common issue in the medical domain. Furthermore, the model is fine-tuned to identify optimal hyperparameter values, such as learning rate, momentum, and dropout rate, thereby improving model training and achieving higher accuracy and better generalization, which is vital for improving the performance of medical image classification models.
3.1. XAI Methods
The proposed framework incorporates explainability and transparency by integrating three XAI methods to generate heatmaps that highlight the features of the input image that influenced the model’s prediction/decision. This integration provides clinicians with transparent, understandable, and clinically aligned explanations of the rationale behind each prediction, effectively addressing a significant challenge in medical AI: trust and transparency. The three most popular post hoc XAI methods from the literature have been selected for implementation in the proposed research to provide explanations. LIME [10], Grad-CAM [11], and Guided Grad-CAM [12].
3.1.1. LIME
LIME provides feature-level details/explanations for the underlying model. LIME can help analyze a model’s feature importance because it focuses on explaining individual predictions rather than the model as a whole. This local technique lets users learn how different aspects of an input sample influence the model’s output for a given instance. LIME provides instance-level details about the model, revealing its behavior locally.
3.1.2. Grad-CAM
Grad-CAM is an XAI method that provides explanations only for image-based data sets. Its working principle generates heat maps of gradient layers for a CNN, which contribute to the output prediction and highlight the areas and pixels most important to the classification process.
3.1.3. Guided Grad-CAM
Guided Grad-CAM combines Grad-CAM and Guided Backpropagation by point-wise multiplication of both, as the former results in a coarse-grained activation map and the latter provides high-resolution maps. The feature maps of the last convolutional layers are used to compute the Guided Grad-CAM. It identifies trivial pixels in an image by setting negative gradients to zero, yielding fine-grained activation maps.
3.2. XAI Fusion Approaches
We also employed four distinct heatmap fusion methods to generate the explanations, providing a more comprehensive and robust understanding of the model’s decision-making process. Common areas in the combined heatmap indicate regions of high importance shared by multiple methods, while uncommon areas offer unique insights from individual techniques. This explainability process yields better model interpretation, serving two primary purposes. First, it addresses any ethical concerns arising from the black-box nature of AI-based models in healthcare. Second, it serves as an initial tool to build better trust with clinicians and promote clinical adoption of the model. We also compare the performance of these combined/fused heatmaps using evaluation metrics and qualitative analysis.
The motivation for implementing XAI fusion approaches is based on ensemble theory with the aim of improved robustness, interpretability, and reduced variance. The need of exploring fusion strategies to merge multiple complementary explanations has been focused upon in existing literature. Recent research has also explored the strengths of fusing different XAI-based techniques [32,33,34]. Moreover, much focus has been placed on exploring XAI fusion to achieve uniform evaluation frameworks to enhance trust in model interpretability. A brief overview of the fusion strategies mentioned above is given below.
3.2.1. Weighted Average Fusion
This approach assigns weights to the explanations generated by each XAI method and averages the pixel values to produce the final output. In this approach, all the XAI methods are given equal importance.
3.2.2. Max Fusion
The maximum value against each pixel for individual heatmaps is selected to generate the final heatmap. This approach focuses on identifying any area that may be marked as important by at least one method.
3.2.3. Simple Average Fusion
Each visual explanation generated by a particular XAI method is assigned a weight, and all pixels in the heatmaps are averaged to create the final map. In this approach, each explanation method in the combined visualization is treated equally.
3.2.4. Consensus Fusion Approach
This approach focuses on generating consensus-based maps from regions consistently identified as necessary by maximum XAI techniques. Identifies important regions by highlighting pixels with scores above a threshold.The consensus heatmap highlights features of strong agreement, improving confidence by reducing the noise from individual methods.
3.3. XAI Evaluation
Two evaluation metrics have also been introduced in the proposed work: Stability and Deletion to evaluate the XAI methods used in our analysis.
- Stability Metric: The Stability measures the reliability and consistency of the underlying XAI method under noisy conditions by altering the input data. This metric measures the change in the visualizations by adding perturbations to the input. Stability analyzes the robustness and reliability of the explanations generated by the XAI method.
- Deletion Metric: The Deletion metric identifies critical features in an input by evaluating the impact of deleting specific pixels. Consequently, it measures the faithfulness of an XAI method, which is crucial in ensuring the trust and reliability of the model in clinical applications.
3.4. Interpretability Score
This study introduces a novel reliability measure, the Interpretability Score, to evaluate trust in the underlying AI-based diagnostic model. This contribution addresses the crucial gap between the model’s diagnostic accuracy and reliability. The Interpretability Score quantifies the essential components required for a trustworthy system, such as accuracy, clinician ratings, and explanation quality.
The values of all components are normalized to the range 0 to 1. Moreover, each value is assigned a weight to emphasize model transparency further. The Interpretability Score can be mathematically defined as follows:
In Equation (1), denotes denotes normalized Diagnostic Accuracy of the model, which is the AUC value. represents the normalized Clinical Trust Score, which is calculated from the feedback provided by the clinicians. This value represents the real-world trust score evaluated by the domain experts. stands for Explanation Quality Score, calculated by normalizing the explanation quality evaluation. whereas, , , are the respective weights assigned to each score.The values of , , and are set to 0.4, 0.3, and 0.3, respectively. The final Explainability Value ranges from 0 to 1.
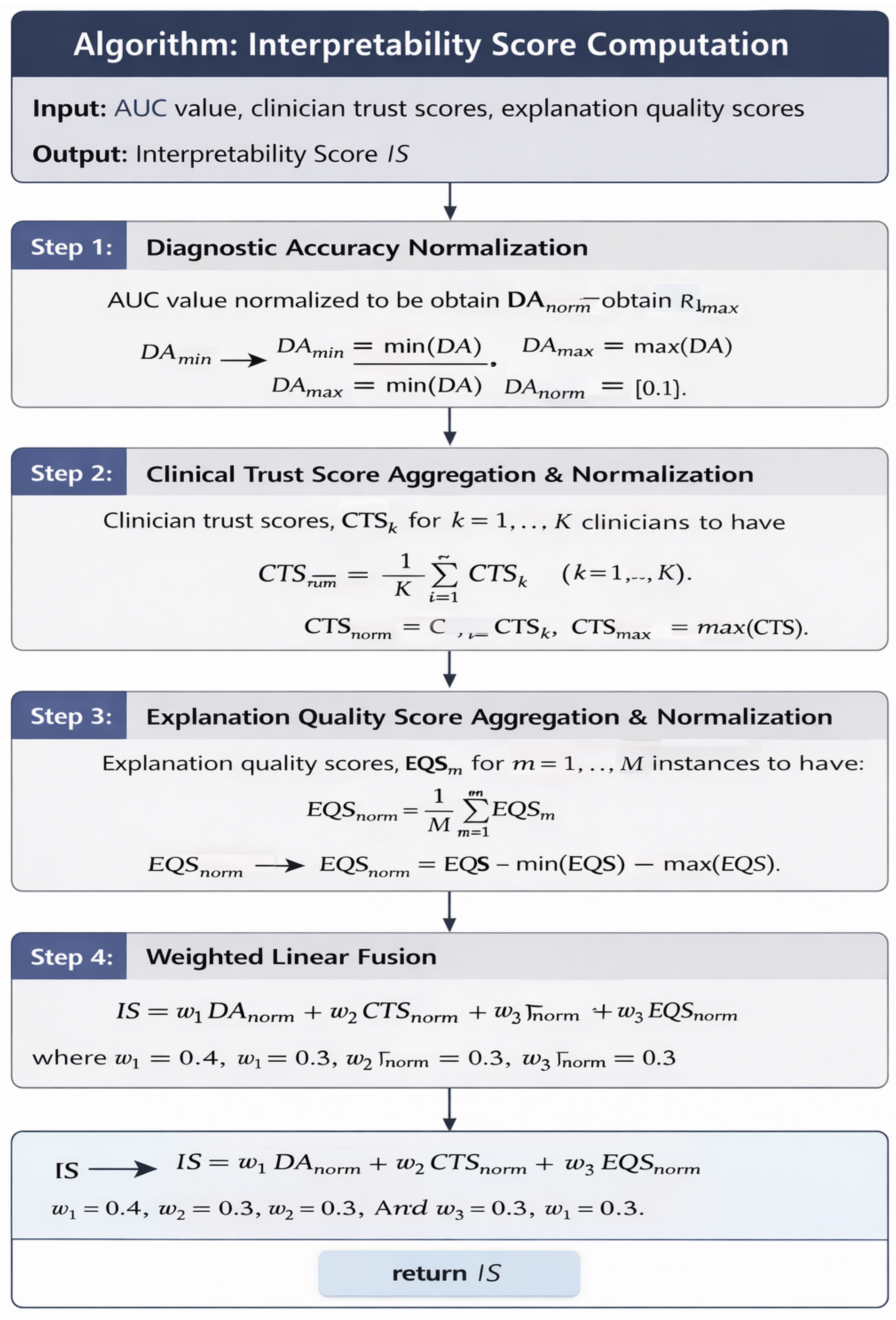
The algorithm presented in Figure 3 describes the step wise computational details along with the mathematical formulation of the proposed Interpretability Score.
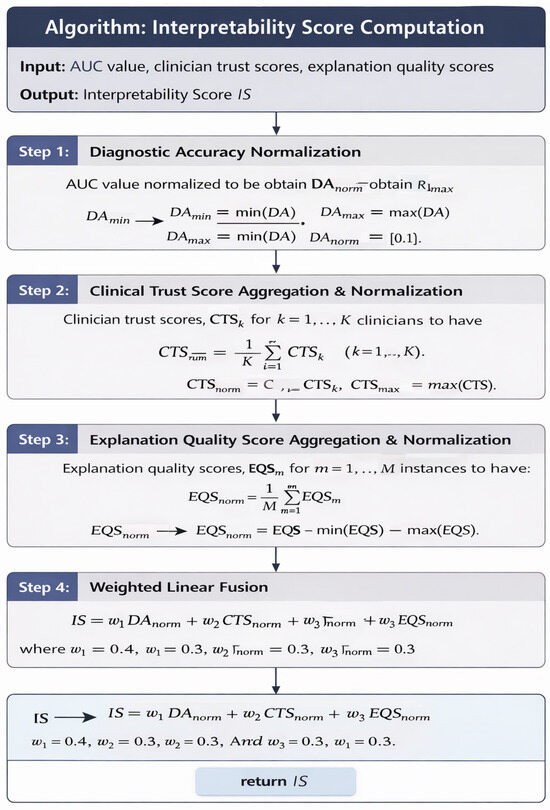
Figure 3.
Algorithm for the Interpretability Score Computation.
3.5. Patient Report
The framework also generates a patient report that summarizes the explanation (both visual and textual) of the predicted decision. This report provides details that lead to a diagnostic conclusion. It also deepens understanding of the model’s predictions within the clinical context. Hence, the proposed framework comprehensively addresses ALL detection in an interpretable and understandable manner.
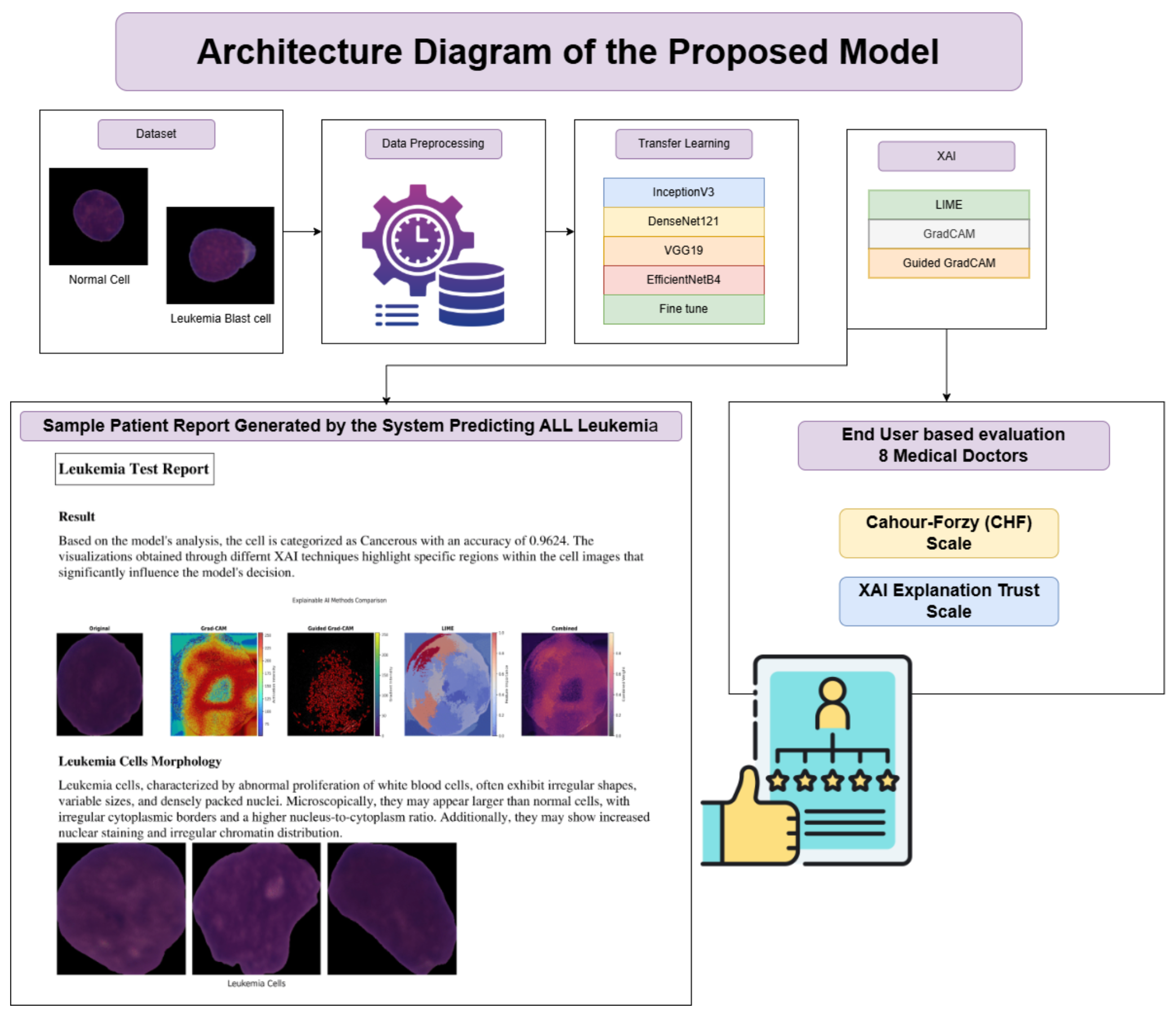
Figure 4 presents the architecture of the proposed model.
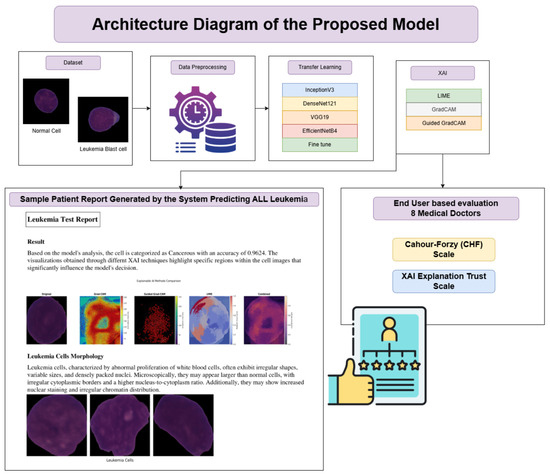
Figure 4.
Architecture Diagram of the Proposed Framework.
4. Implementation Details
4.1. Data Collection
The proposed framework has been trained on a publicly available ALL image dataset from Kaggle, which provides segmented images of lymphocytic cells with staining noise, scaled to a consistent size of 224 × 224 pixels with three color channels (RGB). These images are also padded with black to ensure uniformity in size and enhance focus. However, according to sources, the original blood smear images have a higher resolution of 2560 × 1920.
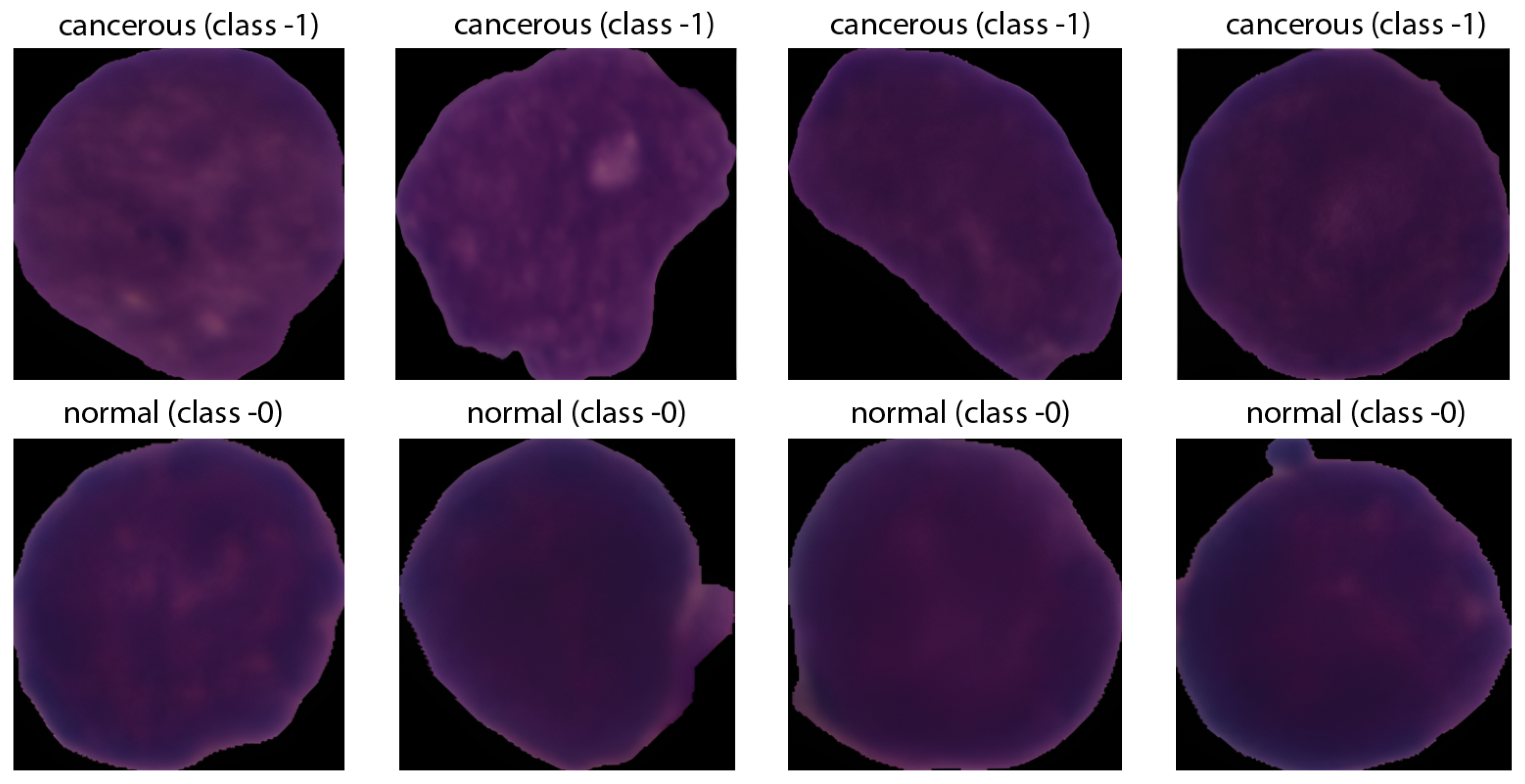
The ground-truth labels were derived from expert oncology annotations. The dataset comprises 15,135 images of 118 patients, classified into two categories: normal cells and leukemia blasts, as shown in Figure 5.
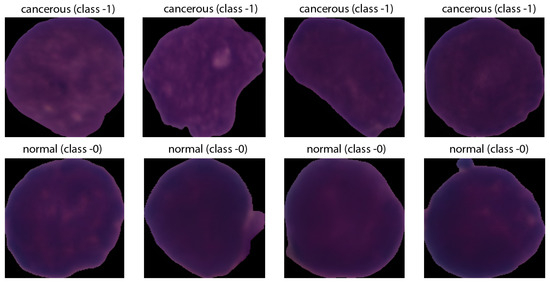
Figure 5.
Sample Leukemic Cell Images.
4.2. Data Preprocessing
Identifying immature leukemic blasts from normal cells is challenging due to their morphological similarities. Preprocessing included noise reduction via Gaussian blurring and image augmentation techniques such as rotation, flipping, and zooming to enhance the model’s robustness against overfitting and its ability to generalize from the training data to unseen test data. Data augmentation is mathematically modeled in the following manner:
- Rotation:
- Width and Height Shifts:
- Horizontal and Vertical Flip:
- Horizontal flip:
- Vertical flip:where h and w are the height and width of the image, respectively.
4.3. Test Environment
The framework was developed in Python version 3.11 using the TensorFlow DL programming library, and Google Collaboratory was used to conduct the experiments. The A100 NVIDIA GPU (premium GPU) with 80 GB of memory was used for the experimental hardware to enable fast, efficient processing.
4.4. ML Model Selection
The research uses four transfer-learning-pretrained architectures (DenseNet121, EfficientNetB4, InceptionV3, VGG19) that are well known for their performance on complex image classification tasks, whereas the dataset split ratio was 70:15:15.
4.5. Parameter Settings
The base models are fine-tuned on the leukemia image dataset to adapt the learned features to the specific appearances and phenotypes of the cells. In addition, the hyperparameter range is adopted from popular methodologies in medical image classification tasks. The hyperparameters and their respective values are summarized in Table 2. However, consistent performance was achieved along with stable convergence for a batch size of 100 epochs, making it the final choice for the model training. In order to prevent over-fitting and to achieve better convergence stability early stopping with a patience value of 15 epochs was selected. All models are trained with similar hyperparameter values to ensure a fair comparison and consistency to measure their performance.

Table 2.
Hyper Parameters for Training the Proposed Model.
5. Experimentation Results
The experimentation was divided into two phases. The first phase covered training the base transfer-learning models, and the second phase fine-tuned them.
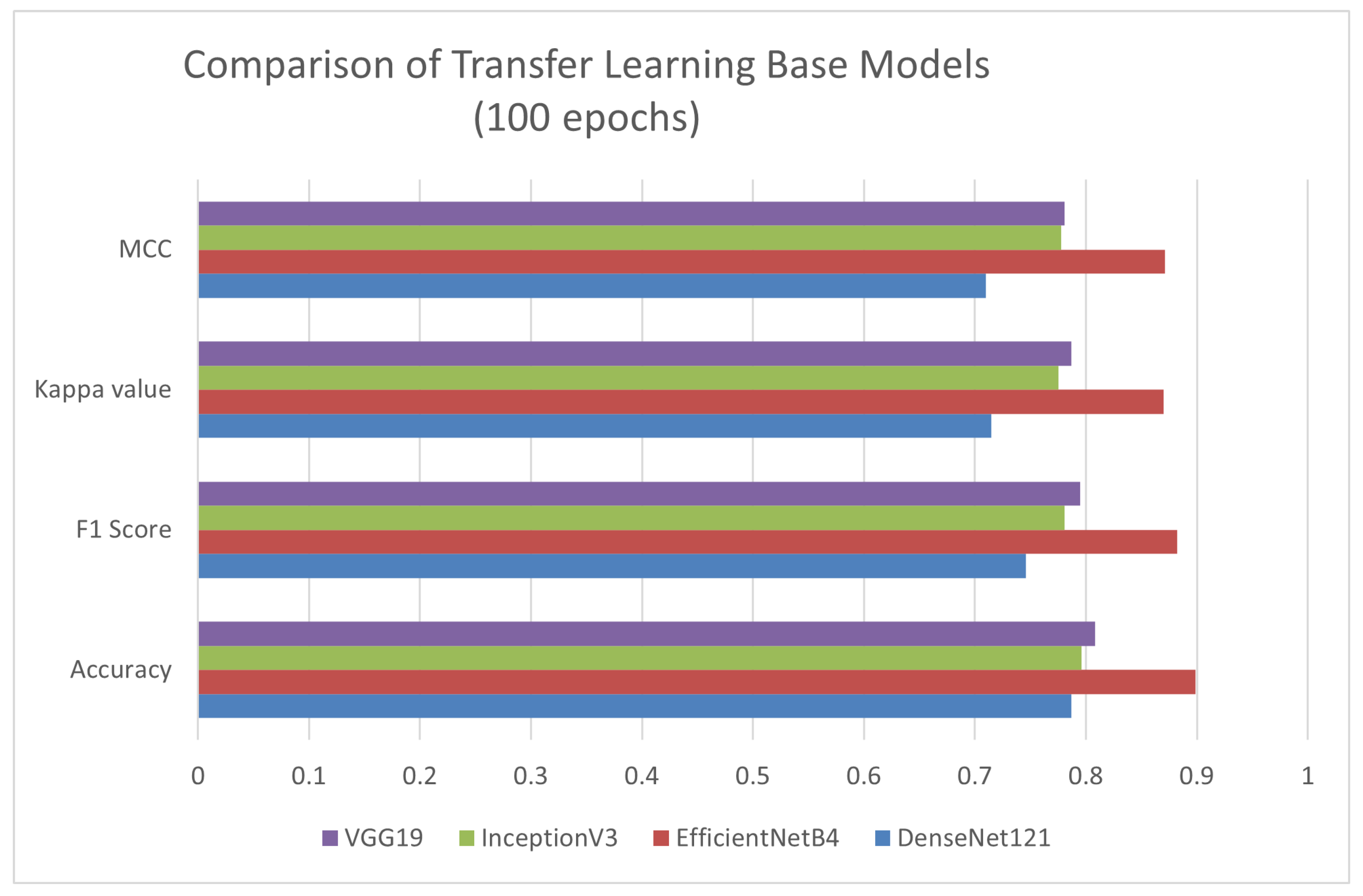
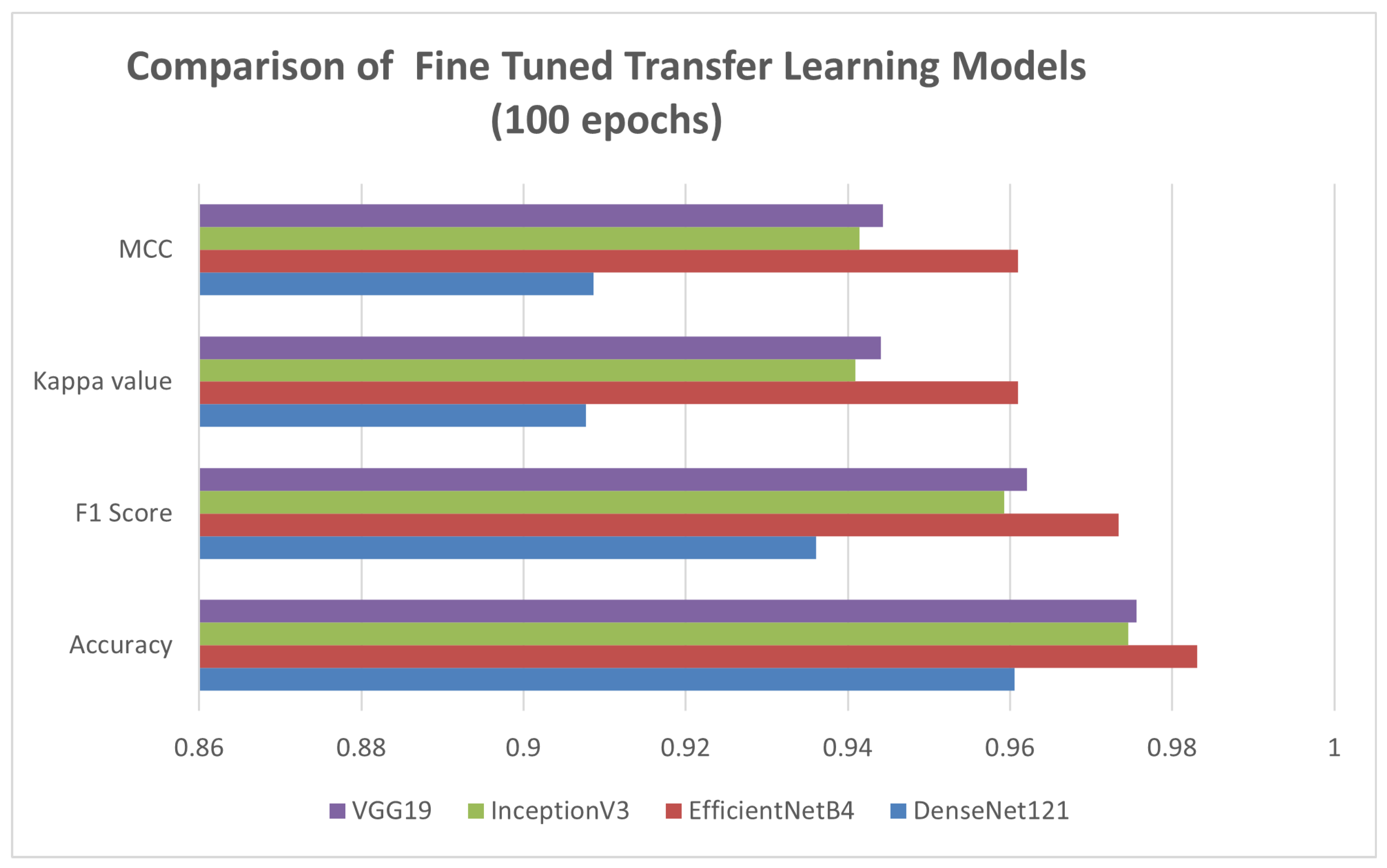
Figure 6 evaluates the performance of four different transfer-learning architectures (VGG19, DenseNet121, EfficientNetB4, InceptionV3) across four metrics: Accuracy, F1 Score, Kappa, and MCC (Matthews Correlation Coefficient). EfficientNetB4 performed distinctly across all metrics. Therefore, it is selected as the base model for further experimentation. Figure 7 presents the comparative performance evaluation of the models discussed above (after fine-tuning). However, a specific performance breakdown is presented below.
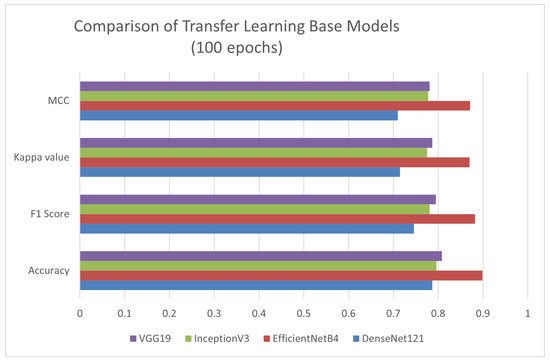
Figure 6.
Performance Evaluation of Different Transfer-Learning Models.
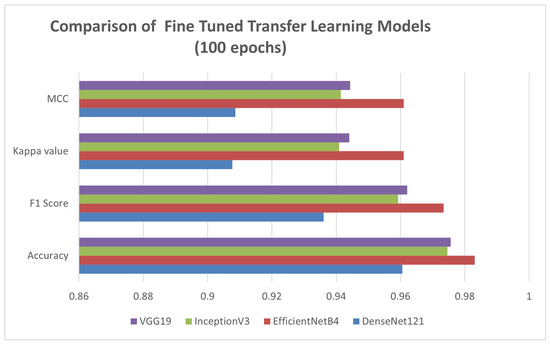
Figure 7.
Performance Evaluation of Different Transfer-Learning Models Fine-Tuned.
- Accuracy: EfficientNetB4 demonstrated the best diagnostic performance by achieving 98.31% accuracy.
- MCC: EfficientNetB4 outperformed in MCC, also suggesting a strong correlation between the predicted and actual classes in all categories.
- Kappa: EfficientNetB4 achieved a high Cohen’s Kappa score, indicating strong agreement between predicted and true class labels beyond chance.
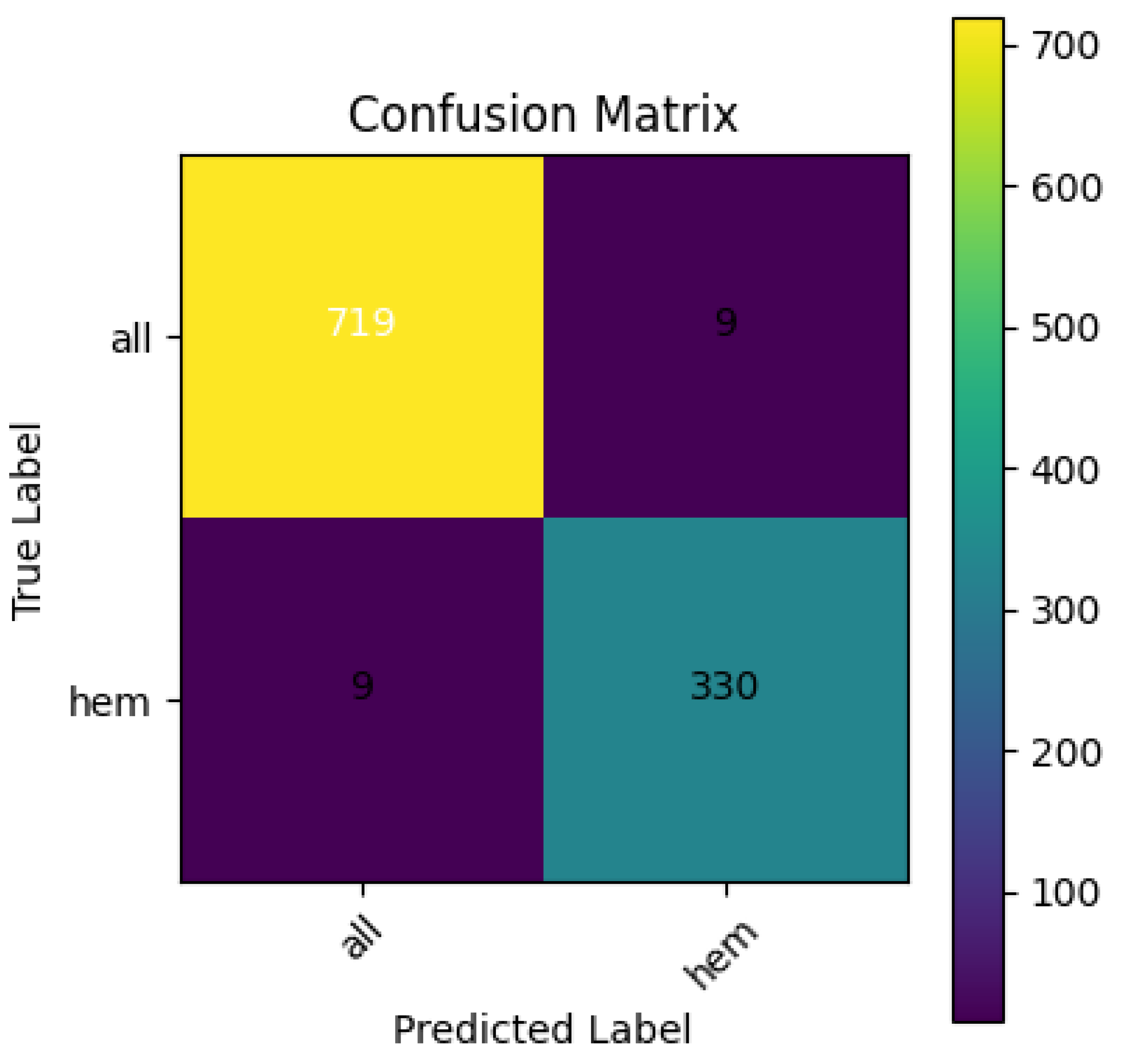
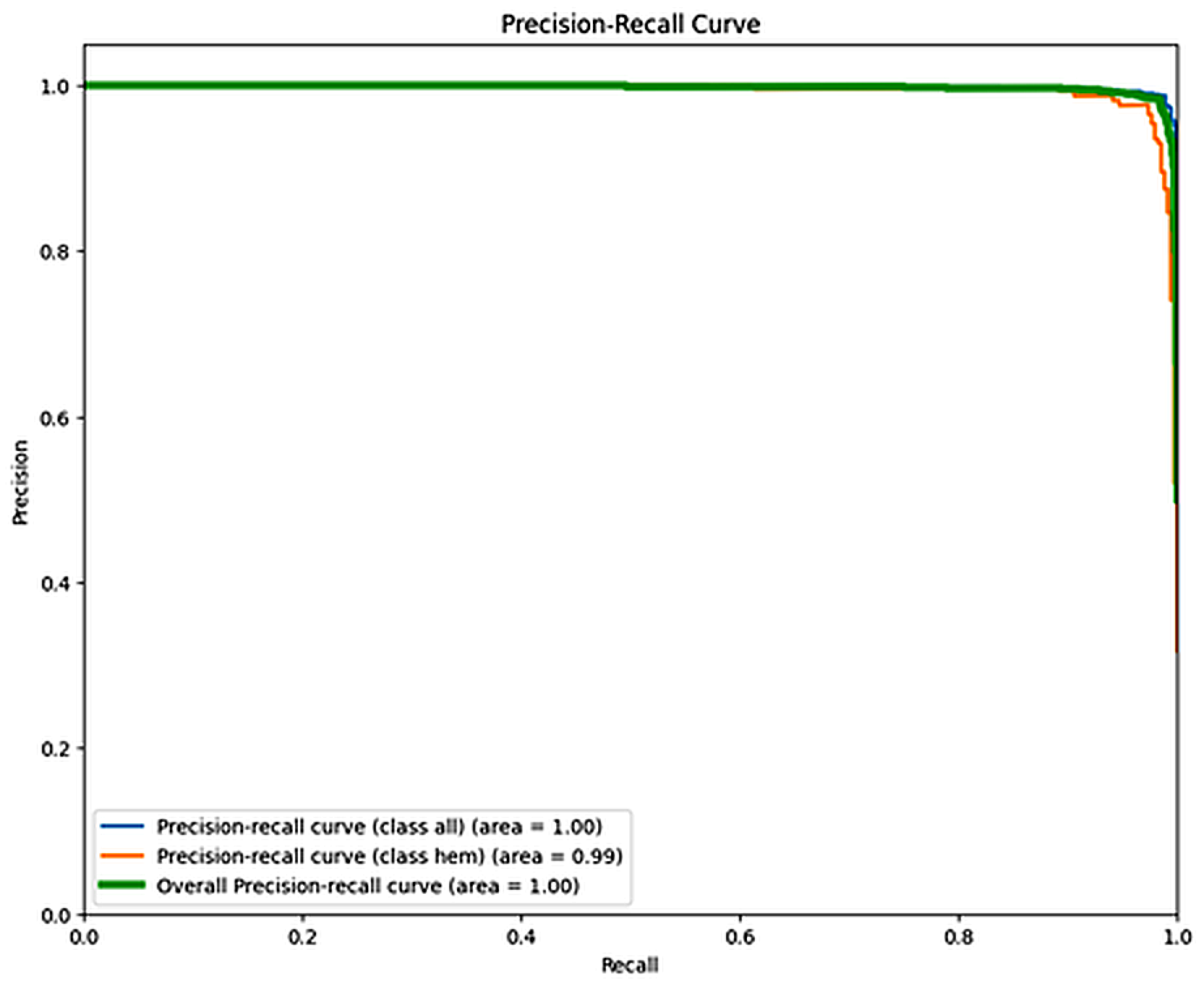
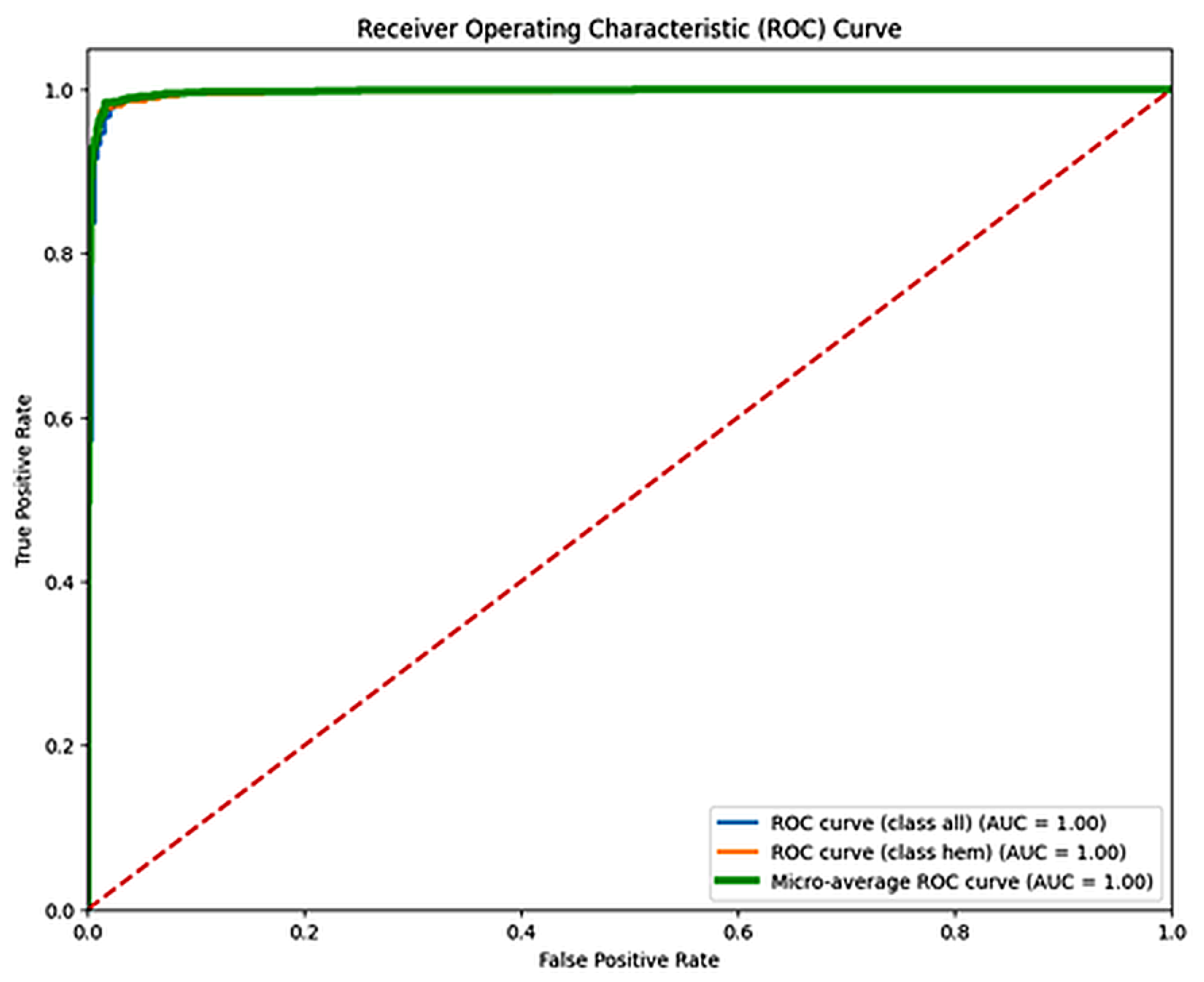
Figure 8 shows the confusion matrix for the EfficientNetB4 model, with an accuracy of 98.31% and an F1 score of 0.9734. Figure 9 presents the Precision–Recall Curve of the model. The area under the curve (AUC) for the proposed model is 1.00 for the ALL class and 0.99 for the hem class, with an overall AUC of 1.00. However, Figure 10 presents the ROC curve. These curves are essential for evaluating the model’s diagnostic accuracy, especially in the medical domain where reliable, accurate decision-making is critical.
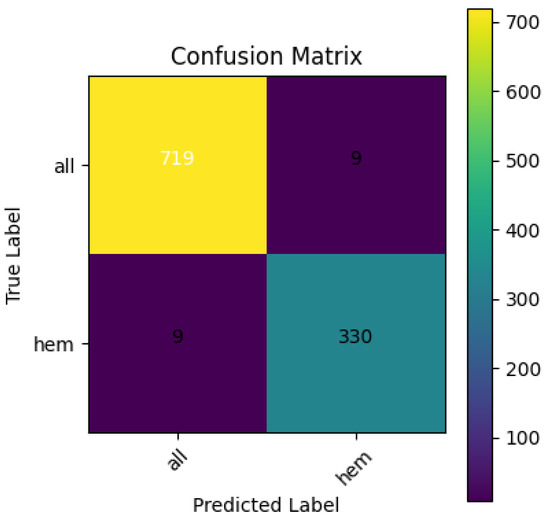
Figure 8.
Confusion Matrix for EfficientNetB4.
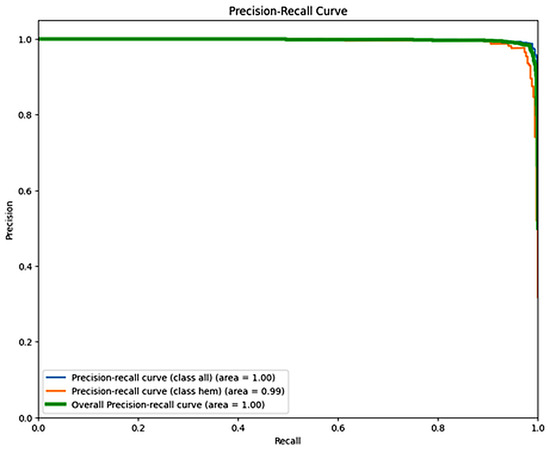
Figure 9.
Recall Curve for EfficientNetB4.
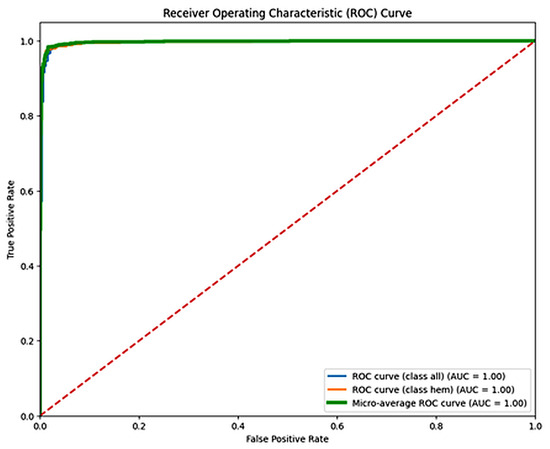
Figure 10.
ROC Curve for EfficientNetB4.The dashed diagonal line represents the performance of a random classifier (AUC = 0.5), serving as a baseline for comparison.
5.1. XAI-Based Results
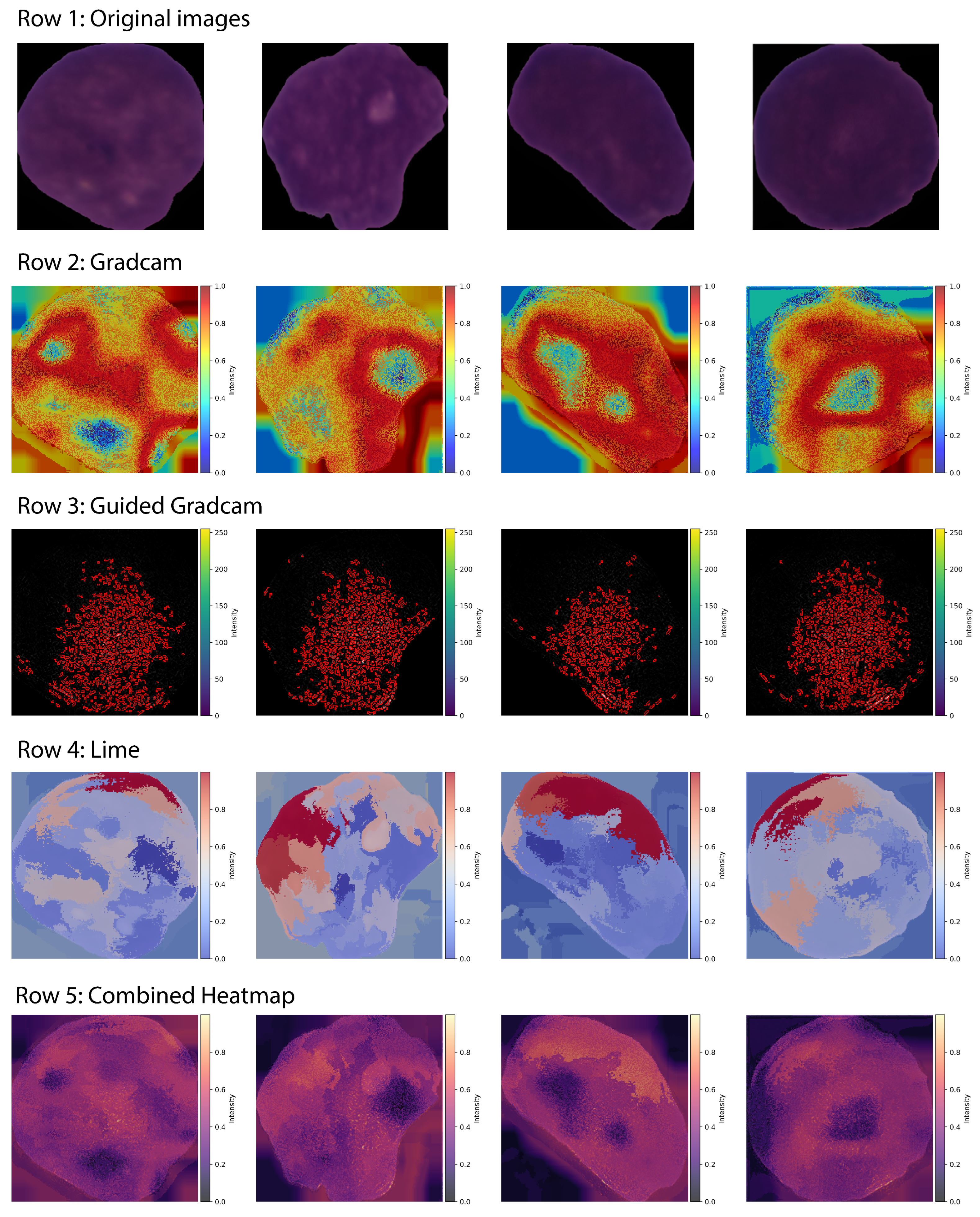
Figure 11 illustrates sample visual explanations generated by XAI methods (LIME, Grad-CAM, and Guided Grad-CAM) for the proposed framework.
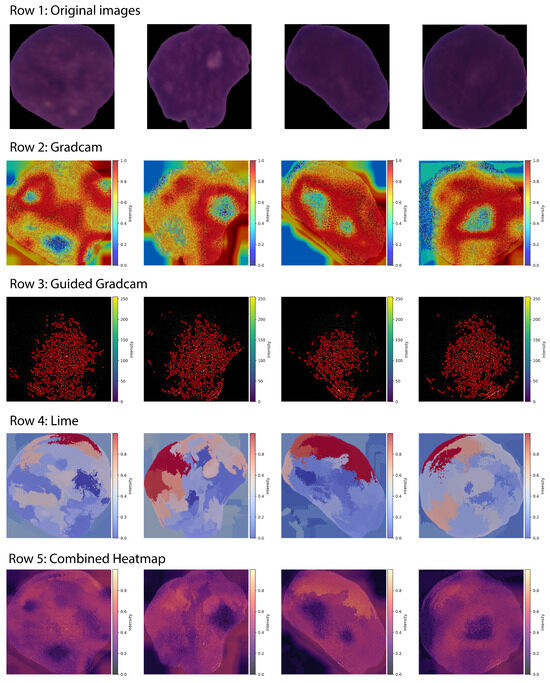
Figure 11.
XAI Visualizations Generated by LIME, Grad-CAM, Guided Grad-CAM.
Moreover, by combining the above-discussed explanations, the model also generates advanced fusion heatmaps that balance the trade off between feature importance and explanation diversity, thereby incorporating the most important rationale behind the classification decision.
Let:
- = normalized Grad-CAM heatmap
- = normalized Guided Grad-CAM heatmap (averaged over channels if 3D)
- = normalized LIME heatmap
Given below are the mathematical equations for calculating simple average fusion Equation (6), weighted average fusion Equation (7), max fusion Equation (8), and consensus fusion Equation (9), and Equation (10), all representing advanced combined heatmaps.
Let:
Then:
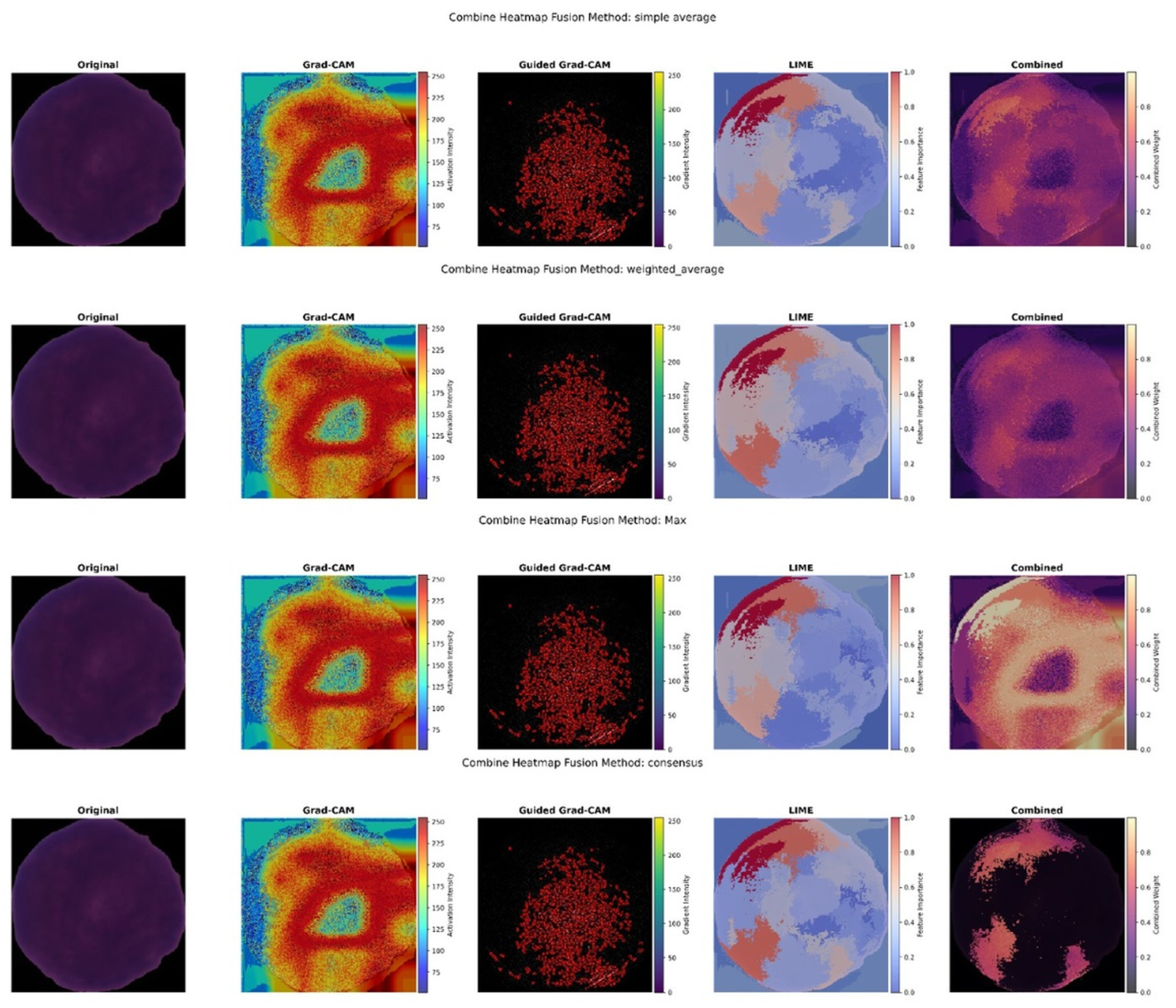
Figure 12 shows the combined XAI visuals achieved by applying different fusion approaches.
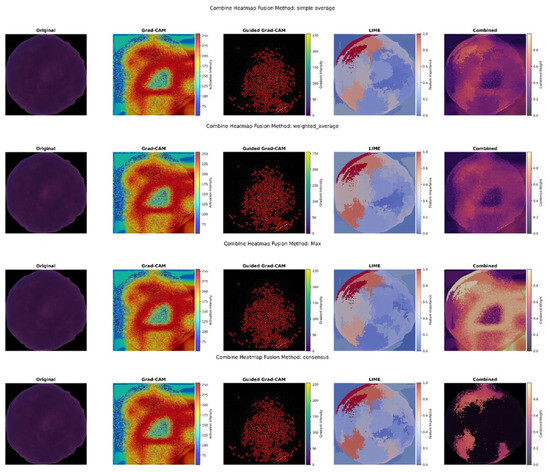
Figure 12.
Combined Hybrid XAI Visuals Generated by Different Fusion Approaches.
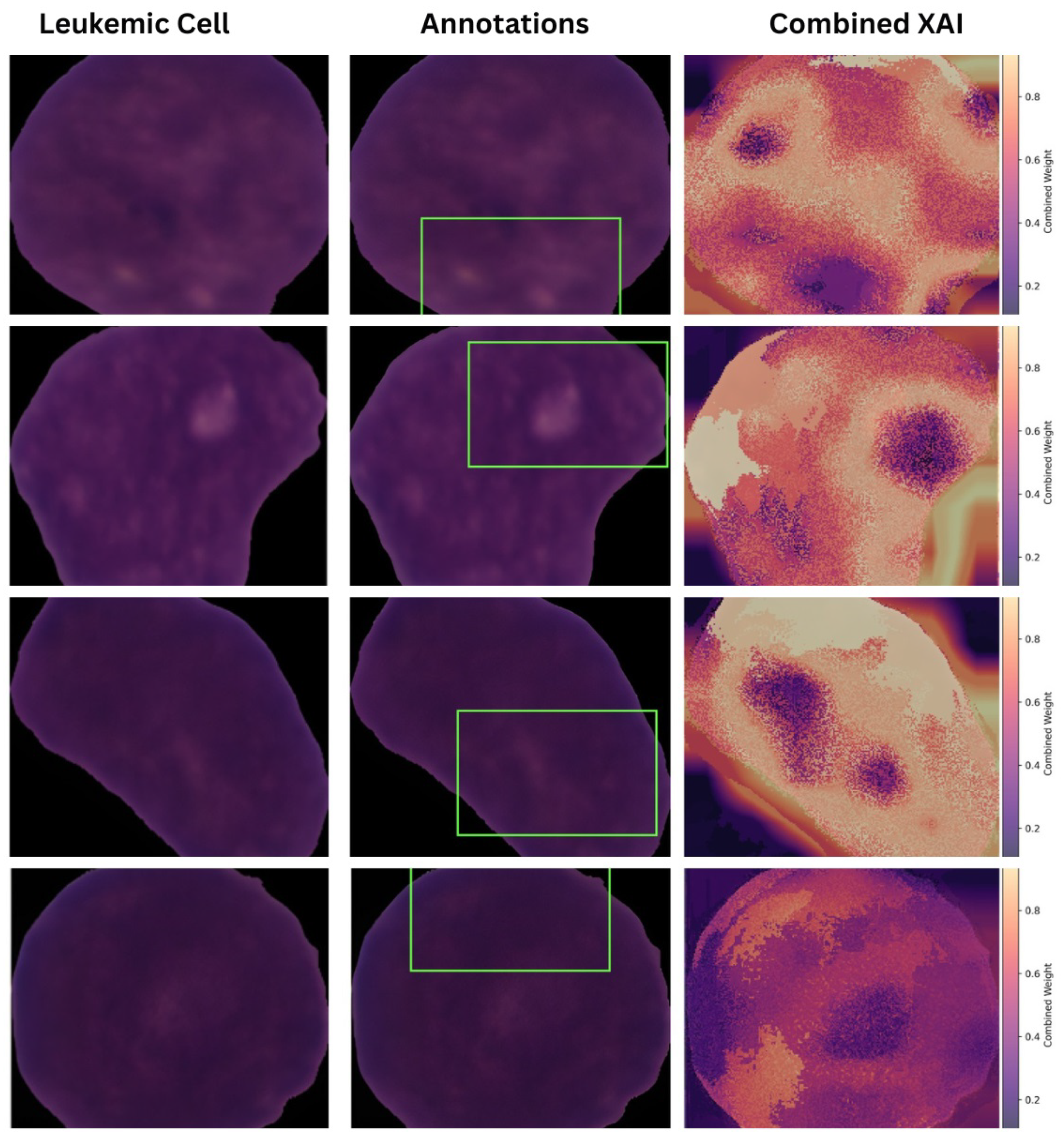
All fusion heatmaps were analyzed and compared with the experts’ annotations. Furthermore, visuals derived from the max fusion strategy were selected as the best in terms of interpretability and the explanations they provided. Figure 13 presents the annotations with the fusion heat maps generated by the max approach for leukemic cell images.
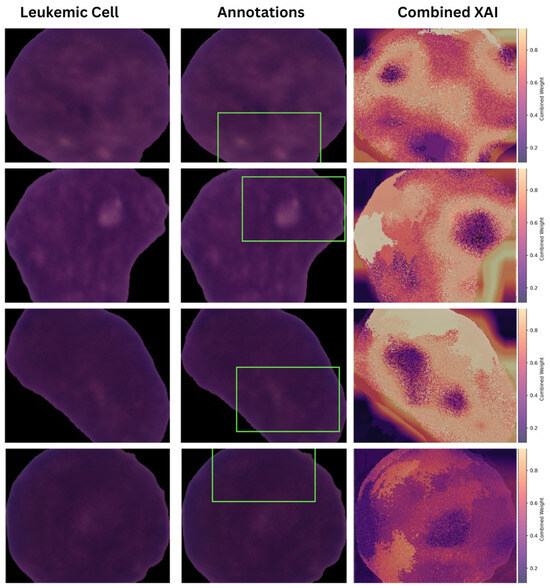
Figure 13.
Comparative Analysis of Expert Annotations with Combined Heatmaps (Max Fusion).
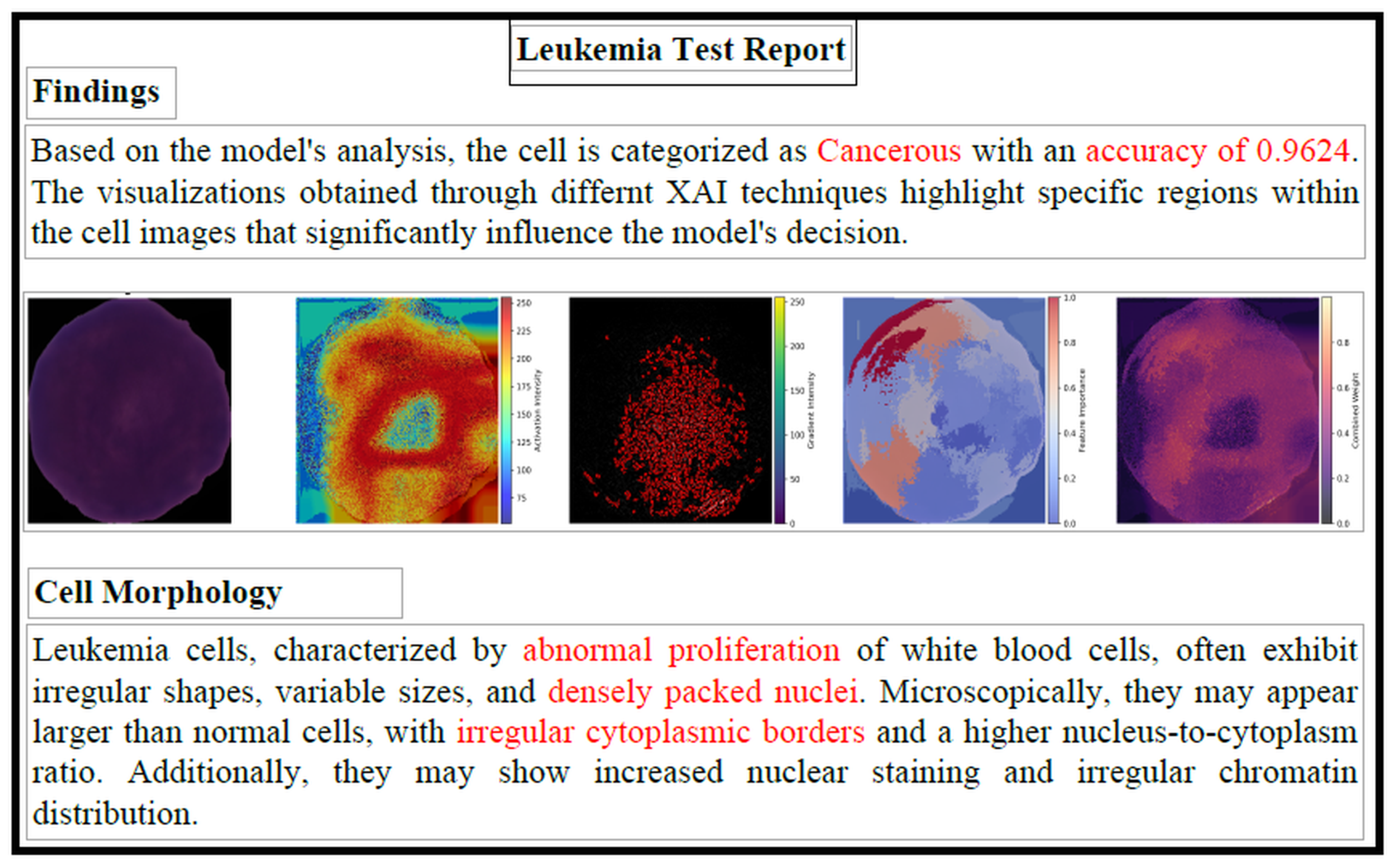
The diagnostic report comprehensively explains the decision-making process of the proposed model. Using fused heat maps derived from LIME, Grad-CAM, and Guided Grad-CAM, the critical regions in microscopic images that influence the classification decisions of the underlying DL model are highlighted. Textual explanations are provided by analyzing the highlighted areas alongside the model’s diagnostic decisions. For example, if a specific morphological feature is highlighted in the combined heat map, a textual explanation of the feature (under the supervision of experts) is provided, along with its correlation to the diagnosis decision. A logical flow of information is ensured by presenting the predicted diagnosis, the visual explanation, and the textual explanation in a structured manner. The clinical performance and relevance of the proposed explanation system are supported by the close alignment between the highlighted regions in the fused heat maps and those identified by experienced pathologists. Figure 14 shows a sample diagnostic report generated by the framework in PDF format.
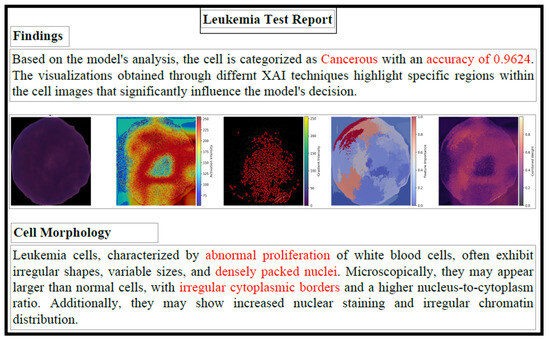
Figure 14.
Sample Leukemia Test Report. The red-highlighted text emphasizes key diagnostic outputs, including the model’s predicted class, confidence level, and abnormal cellular morphology features used to support the classification.
5.2. Quantitative and Interpretability Evaluation of the XAI Approaches
The effectiveness of XAI-generated explanations was evaluated using the following criteria: faithfulness, consistency, and reliability. The evaluation of the implemented XAI techniques included two distinct metrics:
- Stability
- Deletion
For a particular XAI approach, let N be the total perturbations required, be the original selected input image, and be the selected perturbation. The Stability metrics is derived as below (Equation (11)),
whereas M are the total regions considered to be removed, and is the image after removing the jth region from the original image. The mathematical formulation of Deletion metrics is given below (Equation (12)).
This study introduces an Interpretability Score, a single, consolidated metric ranging from 0 to 1 that is directly proportional to the clinical trust and reliability of the AI model. The Interpretability Score combines three normalized dimensions of trust in medical systems: diagnostic accuracy, domain expert feedback ratings, and explainability. Each dimension reflects a distinct yet vital aspect of the reliability of the AI-based diagnostics.
For the proposed framework, Interpretability Scores are calculated for each XAI method. The reported DA of 0.9831 and CTS of 0.76 are used. However, the EQS obtained jointly for the Deletion and Stability metric varied for every method. According to the results, both LIME and Grad-CAM obtained an Interpretability Score of 0.771, followed by 0.707 for Guided Grad-CAM, validating the model’s trustworthiness and reliability.
Table 3 presents the quantitative analysis of the employed XAI methods (LIME, Grad-CAM, Guided Grad-CAM, average fusion, weighted average fusion, max fusion, and consensus fusion), against the Stability and Deletion metrics, where a lower score indicates better performance. Max fusion approach demonstrated best Stability score whereas LIME showed the best values for the Deletion metrics.

Table 3.
Stability and Deletion Metrics for XAI Methods.
5.3. Assessment of Proposed Model Using Experts
The qualitative assessment of the model has been conducted through a human-centric approach; it included evaluations by end users, specifically clinical experts. The CTS value contributing 30% towards computing the Interpretability Score is derived from qualitative evaluation performed by these medical practitioners. A group of eight medical doctors, mainly pathologists, was selected to evaluate histopathological images and the corresponding output reports generated by the proposed framework.
The survey is based on two of the most frequently used scales for XAI systems: the Cahour-Forzy (CHF) Scale (bipolar) [35] and the XAI Explanation Trust Scale (Likert) [36]. The personal details of the survey participants have also been recorded, including their age, specialty, years of experience, and prior knowledge of AI-based systems. Before the survey, each participant is also introduced to ’What is AI?’ to give a clear understanding of the system.
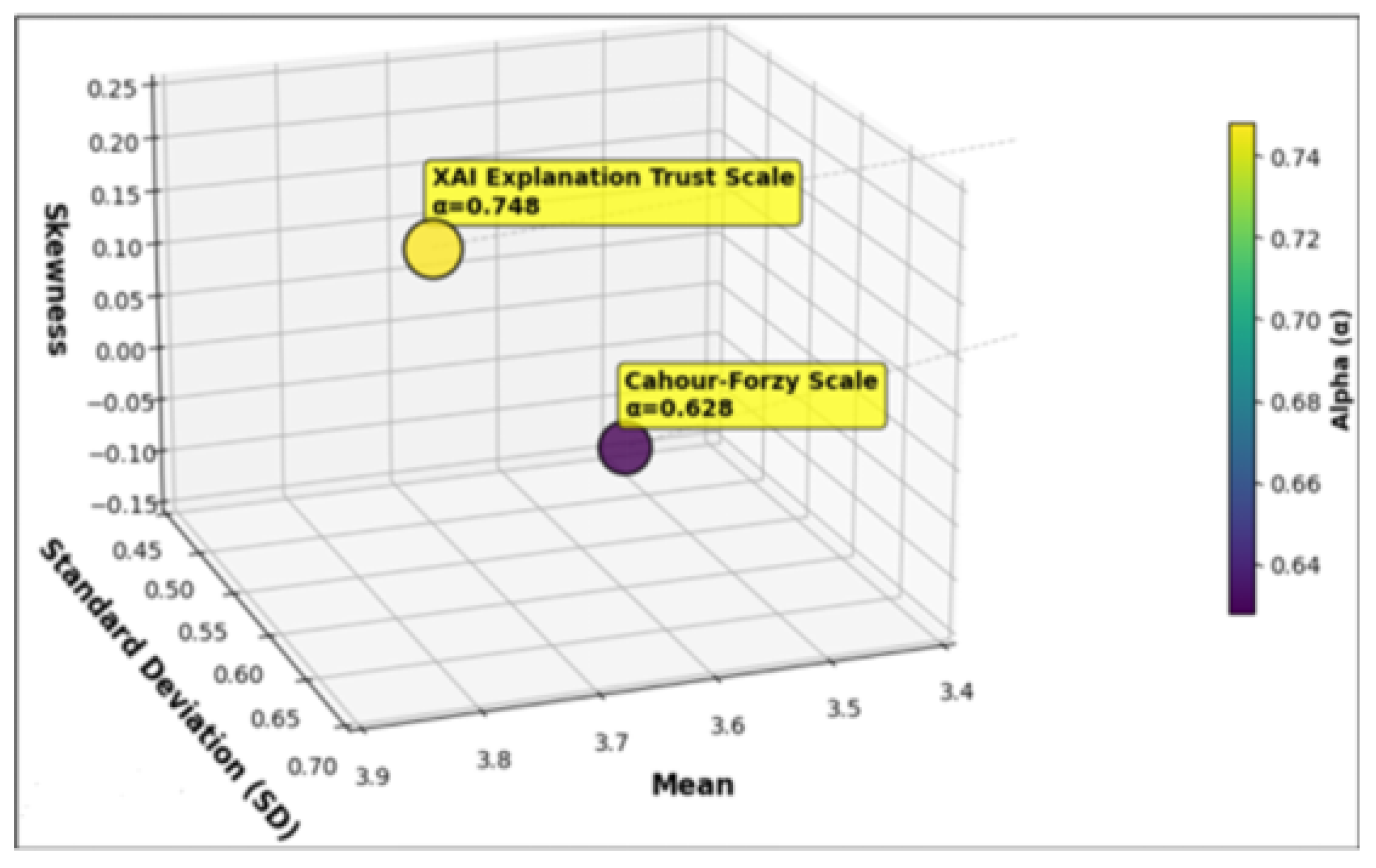
Mean and Standard Deviation (SD) to evaluate the Normal distribution for the selected scales are calculated for the scores obtained by them. The mean value for the XAI Explanation Trust scale is 3.8, and the SD is 0.648. For the CHF scale, the recorded mean value is 3.525 with 0.505 SD. Furthermore, to measure the asymmetry in the distribution, the Skewness test is also conducted.
The Skewness test confirmed that the dataset is normally distributed. In addition, to evaluate the internal consistency of the items for both scales, Cronbach’s alpha () is also calculated (a higher value indicates greater consistency and homogeneity among the items). The Cronbach’s alpha for the XAI Explanation Trust Scale is recorded to be , and for the CHF scale it is , showing that items of both the scales are consistent and satisfactory. Figure 15 presents these results.
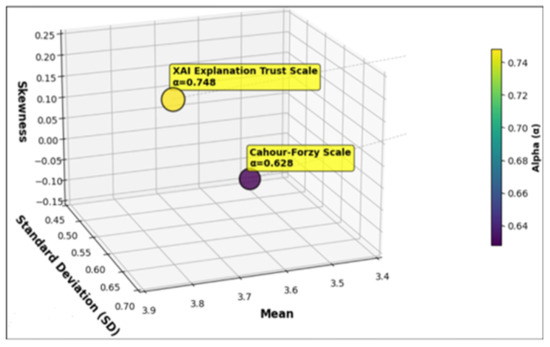
Figure 15.
Visualization of Mean, Standard Deviation, and Skewness for the XAI Explanation Trust and CHF Scales.
6. Discussion
The performance evaluation of the proposed framework showed excellent diagnostic accuracy with a high number of True Positives (TP) and True Negatives (TN). At the same time, this remarkable performance is further validated by only a few cases reported as False Positives (FP) and False Negatives (FN). Among the four transfer-learning models, EfficientNetB4 showed the best performance for classifying leukemia cells from normal cells, as evidenced by high ROC and Precision-Recall scores, indicating the diagnostic strength and robustness of the DL model.
The integration of XAI techniques has proven essential for improving the transparency and understandability of the model’s predictions. The combination of LIME, Grad-CAM, and Guided Grad-CAM provides both post hoc and local explainability, generating understandable explanations of the most important features and image regions that contributed significantly to the diagnosis of ALL. Amongst all employed XAI methods, LIME demonstrated better explainability, in terms of the Deletion metrics and max fusion for the Stability metrics. However, both Grad-CAM and Guided Grad-CAM and the rest of fusion strategies have also shown promising results. Further analysis of these explanations could be beneficial in addressing challenges like misclassified cases and data bias.
The visual explanations generated by the fusion approaches provide detailed insight into the model’s classification decision. These visual explanations were compared with the expert oncologists’ annotations. Furthermore, the visual maps generated by max fusion heatmaps were found to be closely aligned with expert annotations. It validates that the underlying DL model successfully captures relevant regions and anomalies in the microscopic images, which may support clinicians when making diagnostic decisions.
The novelty of the proposed framework lies in introducing a unique Interpretability Score metric to evaluate trust in the system. The primary focus of previous studies has been on achieving diagnostic accuracy or implementing distinctive metrics of explainability. However, this study presents a consolidated quantitative metric to evaluate the reliability of the underlying DL model, derived by combining diagnostic accuracy, clinicians’ trust ratings, and evaluations of XAI-based explanations. With an Interpretability Score of 0.771, the proposed explainable ALL diagnosis framework demonstrates strong performance in terms of explainability and reliability. It also indicates potential of alignment with clinical needs, ethical standards, and regulatory expectations for trustworthy AI in healthcare.
In addition, clinicians’ feedback has also shown encouraging results, demonstrating that such explanation-based frameworks have a potential to be integrated into the healthcare domain, in the future.
6.1. Comparison with Existing Methods
The proposed method outperformed traditional diagnostic procedures, which are prone to human error and difficult to scale. Unlike previous AI studies that did not prioritize trust and explainability evaluation, our research equally values accuracy, trust, and interpretability to make AI more practical for medical use.
Compared with recent studies that use DL models such as CNNs, ensembles, and transfer learning for ALL classification, the proposed framework achieves 98.3% accuracy. The deep CNN model proposed by Oybek et al. achieved 93.7% accuracy for ALL detection [37], and Rajpurkar et al. reported 94.6% accuracy via implementing DL models for medical image classification [38]. The proposed approach achieved improved diagnostic accuracy and higher precision and recall.
To our knowledge, no previous study has presented a comprehensive XAI-based trustworthy framework combining perturbation-based and gradient-based post hoc explanation methods (LIME, Grad-CAM, and Grad-CAM), multi-point fusion heatmaps, and calculating the Interpretability Score for the model explainability, along with extensive feedback from the domain experts in terms of both accuracy and explainability on the local datasets. Furthermore, the framework provides visual and textual explanations to present transparency. Shortly, this work not only achieves diagnostic accuracy but also supports trust and usability for ALL detection. This comprehensive and extensive amalgam offers a deeper understanding of model behavior from a human-centric perspective, making it an initial step towards trustworthy solutions for clinical adoption. Table 4 provides a detailed comparative analysis of the presented framework with the current research.

Table 4.
Comparison of Interpretability Score with Existing Work.
6.2. Ethical and Clinical Considerations
There is a strong need to develop strategies to integrate AI into clinical practice, which can be achieved by bridging the gap between technological innovations and their clinical adoption. Given below is an overview of the practical and ethical implications of deploying the proposed framework in clinical settings.
- Although AI-based models achieve diagnostic accuracy, clinicians remain the real decision-makers. Especially for decision-making and treatment planning in life-threatening diseases, the final decision-maker should always be medical professionals, and AI-based systems will assist and facilitate this labor-intensive process.
- There is a shift toward AI-based modeling to achieve ethical, trustworthy systems. Much focus has been put on ethical compliance while using AI-based systems, especially in the medical domain. National and international regulatory bodies, such as the GDPR, HIPAA, and the WHO, place restrictions on the use of patient data without their consent. Furthermore, they require obtaining patient consent whenever the diagnosis and treatment plan are generated by AI-based models.
- The potential of AI systems to work with existing healthcare imbalances cannot be overlooked. These methodologies are accessible in different regions and socioeconomic groups. Therefore, it is essential to avoid increasing health inequities.
- The design and implementation of ethical AI for medical diagnostics must integrate with human-centered principles. The proposed framework is based on trust and fairness, while minimizing potential data and training biases [43].
6.3. Industrial Benefits and Practical Relevance of the Proposed Framework
Recent studies across multiple AI domains have demonstrated that advanced DL architectures, multi-branch model structures, and multi-scale feature aggregation are effective for complex pattern recognition tasks in noisy, imbalanced, and real-world environments. Research in areas such as underwater acoustic communication, bioacoustic signal classification, UAV navigation, image processing, biomedical image analysis, reinforcement learning, and hybrid NLP–ML systems consistently emphasizes the value of robust preprocessing, informed feature engineering, and adaptive learning strategies to improve model reliability.
At the same time, the evolution of clinically calibrated AI frameworks has underscored the need for trustworthiness, reproducibility, and interpretability in medical decision-support systems. These developments highlight that predictive performance alone is insufficient; transparent and clinically meaningful explanations are essential for safe and responsible deployment.
Together, advances in deep neural network design, imbalance-handling techniques (such as focal loss, SMOTE), optimization strategies, and XAI establish a solid methodological basis for developing a trust-centered, multi-model fusion framework for ALL detection with integrated interpretability assessment [44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61].
The proposed framework takes a more practical approach building on current practices by incorporating multiple explanation methods and introducing quantitative checks on the model’s interpretability. The goal is to support interpretability research and model validation, allowing for a more structured and reliable analysis of model explanations.
7. Conclusions
ALL is known to be one of the main types of blood cancer, especially in children, as it contributes more than a third of childhood cancers. Accurate detection and timely treatment of ALL can benefit both patients and the global healthcare sector. Therefore, to address all the above concerns, the training of an ML model in combination with XAI is chosen to diagnose ALL.
The available training dataset is small, making it unsuitable for DL modeling; hence, the need for data diversity in ML training naturally leads to the use of transfer-learning approaches. This research has effectively explored the application of various transfer-learning-based approaches to train base models, specifically focusing on the performance of VGG19, DenseNet121, EfficientNetB4, and InceptionV3 to achieve higher accuracy and to conduct a comparative analysis of the underlying ML models. Consequently, EfficientNetB4 demonstrated superior performance, outperforming the other models in terms of accuracy, and was selected as the base model. Furthermore, explainability has been incorporated into the framework through various XAI approaches. Real-time feedback on trust and reliability from end users and domain experts has also been conducted in the study. This work may serve as a foundation for future efforts by the scientific community and domain experts to facilitate the widespread deployment of AI-based trustworthy diagnosis systems in clinical settings.
Thus, this quantitative amalgam of human-focused and algorithmic aspects has established a milestone for evaluating AI-based medical systems and has also strengthened the scientific impact of this study. However, the findings of the research may serve as a foundation step towards designing systems for real-world clinical settings. This research can serve as preliminary evidence for developing verification tools for medical professionals’ diagnostic decisions.
Emerging Research Directions
The core aim of the research has been to explore the potential applications of XAI for classifying ALL histopathological images, towards achieving transparent, understandable, and justifiable AI. This framework can be successfully adopted in resource-constrained countries such as Pakistan, which have a limited and underdeveloped healthcare infrastructure, need to meet the healthcare needs of the population, especially when treating critical and life-threatening diseases. Early diagnosis can prompt treatment initiation, thereby improving mortality rates in such countries. To our knowledge, no such work has been presented that focuses on integrating and quantifying XAI (along with the end-user evaluation paradigm) to build trust in predicting ALL at an early stage. This work lays the foundation for reliable, transparent solutions in translational cancer research, particularly for histopathological images. Therefore, our research work can be further extended in the following domains and directions.
- Advanced DL explanation models can be designed using diverse multimodal and multisource datasets, such as patient clinical data, genomic expression data, and image-based data. This diverse data source can be extremely beneficial in advanced diagnostics by revolutionizing patient care planning, tracking disease spread, developing post-care treatment plans, advancing translational oncology, and enabling advanced drug discovery.
- The above-suggested specific solutions apply only to the field of cancer and oncology, and will be beneficial in the field of translational cancer research. Additionally, domain experts, especially clinicians, can participate in research to design domain-specific solutions, as the heterogeneous nature of oncology data is enormous and multidimensional and could result in the loss of significant patterns and information if mishandled [62].
- Moreover, achieving accuracy alone is not a comprehensive measure to evaluate the efficiency and efficacy of the underlying ML model. Fewer data variations and more detailed annotations are another hurdle to translational cancer research, which is essential for better treatment (especially for rare diseases). Bridging the gap between generalized and personalized treatment measures, such as medication and care plans, is a limitation to the prognosis of critical and complex treatments, including different types of cancer [63].
- In addition to this, an infrastructure is also required that involves the active participation of all stakeholders in the healthcare domain, including researchers, pharmaceuticals, and medical practitioners, to obtain an abundance of diverse multimodal biomedical datasets, clinical statistical data, complex and rare phenotypes, and multisource datasets to improve performance toward achieving goals in the healthcare domain. FL-based paradigms supported by XAI offer an ideal approach to achieving this diversity. This domain has not yet been fully explored, but the scientific research community and healthcare specialists can join hands to realize the full potential/benefits for humanity.
- In the domain of XAI, there is a lack of a standard, uniform evaluation framework to measure the levels of explanation performance. Multiple taxonomies have been proposed in the literature to evaluate an XAI-based framework; however, there is no uniform, standardized guideline for assessing XAI performance. Additionally, there is a lack of XAI-based methods that provide multi-modal explanations for a given input data. For example, in oncology, there is a need to design a standard XAI-based framework that operates on a patient’s multimodal dataset. Raising patients’ awareness of AI’s benefits, such as improved diagnostic accuracy and support for clinical decision-making, may help address issues in healthcare. To clear hindrances and ensure a smooth path, it is vital to implement effective training systems that equip clinicians with the skills to take on technical tasks, including system operation, interpreting XAI results, and troubleshooting.
Author Contributions
Methodology, conceptualization, results, and writing—original draft, K.P.; validation and supervision, S.I.S.; writing—review, editing, and project management, M.B.; conceptualization, visualization, and funding acquisition, W.S.; verification and proofreading, A.R.; analysis and results verification, M.J. All authors have read and agreed to the published version of the manuscript.
Funding
The authors received no specific funding for this work.
Data Availability Statement
The original data presented in the study are openly available in https://www.kaggle.com/datasets/andrewmvd/leukemia-classification (accessed on 5 March 2025).
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Adenola, V. Artificial Intelligence Based Access Management System. Ph.D. Thesis, East Carolina University, Greenville, NC, USA, 2023. [Google Scholar]
- Wang, C.; Chen, D.; Hao, L.; Liu, X.; Zeng, Y.; Chen, J.; Zhang, G. Pulmonary image classification based on inception-v3 transfer learning model. IEEE Access 2019, 7, 146533–146541. [Google Scholar] [CrossRef]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- Szegedy, C.; Vanhoucke, V.; Ioffe, S.; Shlens, J.; Wojna, Z. Rethinking the inception architecture for computer vision. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 2818–2826. [Google Scholar]
- Huang, G.; Liu, Z.; Van Der Maaten, L.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2018. [Google Scholar]
- Arrieta, A.B.; Díaz-Rodríguez, N.; Del Ser, J.; Bennetot, A.; Tabik, S.; Barbado, A.; García, S.; Gil-López, S.; Molina, D.; Benjamins, R.; et al. Explainable Artificial Intelligence (XAI): Concepts, taxonomies, opportunities and challenges toward responsible AI. Inf. Fusion 2020, 58, 82–115. [Google Scholar] [CrossRef]
- Paul, S.; Kantarjian, H.; Jabbour, E.J. Adult acute lymphoblastic leukemia. Mayo Clin. Proc. 2016, 91, 1645–1666. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Chaw, J.K.; Ang, M.C.; Daud, M.M.; Liu, L. A Diabetes Prediction Model with Visualized Explainable Artificial Intelligence (XAI) Technology. In Proceedings of the International Visual Informatics Conference, Bangi, Malaysia, 15 November 2023; Springer: Berlin/Heidelberg, Germany, 2023; pp. 648–661. [Google Scholar]
- Tan, M.; Le, Q. Efficientnet: Rethinking model scaling for convolutional neural networks. In Proceedings of the International Conference on Machine Learning (ICML 2019), Long Beach, CA, USA, 9–15 June 2019; pp. 6105–6114. [Google Scholar]
- Ribeiro, M.T.; Singh, S.; Guestrin, C. “Why should i trust you?” Explaining the predictions of any classifier. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 1135–1144. [Google Scholar]
- Joo, H.T.; Kim, K.J. Visualization of deep reinforcement learning using grad-CAM: How AI plays atari games? In Proceedings of the 2019 IEEE Conference on Games (CoG), London, UK, 20–23 August 2019; pp. 1–2. [Google Scholar]
- Selvaraju, R.R.; Das, A.; Vedantam, R.; Cogswell, M.; Parikh, D.; Batra, D. Grad-CAM: Why did you say that? arXiv 2016, arXiv:1611.07450. [Google Scholar]
- Rai, A. Explainable AI: From black box to glass box. J. Acad. Mark. Sci. 2020, 48, 137–141. [Google Scholar]
- Wadden, J.J. Defining the undefinable: The black box problem in healthcare artificial intelligence. J. Med. Ethics 2022, 48, 764–768. [Google Scholar] [CrossRef] [PubMed]
- Chhabra, M.; Kumar, R. A smart healthcare system based on classifier DenseNet 121 model to detect multiple diseases. In Mobile Radio Communications and 5G Networks: Proceedings of Second MRCN 2021; Springer: Singapore, 2022; pp. 297–312. [Google Scholar]
- Preetha, R.; Priyadarsini, M.J.P.; Nisha, J. Automated Brain Tumor Detection from Magnetic Resonance Images Using Fine-Tuned EfficientNet-B4 Convolutional Neural Network. IEEE Access 2024, 12, 112181–112195. [Google Scholar] [CrossRef]
- Dey, N.; Zhang, Y.D.; Rajinikanth, V.; Pugalenthi, R.; Raja, N.S.M. Customized VGG19 architecture for pneumonia detection in chest X-rays. Pattern Recognit. Lett. 2021, 143, 67–74. [Google Scholar] [CrossRef]
- Pervez, K.; Sohail, S.I.; Parwez, F.; Zia, M.A. Towards trustworthy AI-driven leukemia diagnosis: A hybrid Hierarchical Federated Learning and explainable AI framework. Inform. Med. Unlocked 2025, 53, 101618. [Google Scholar] [CrossRef]
- Das, P.K.; Diya, V.; Meher, S.; Panda, R.; Abraham, A. A systematic review on recent advancements in deep and machine learning based detection and classification of acute lymphoblastic leukemia. IEEE Access 2022, 10, 81741–81763. [Google Scholar] [CrossRef]
- Saeed, A.; Shoukat, S.; Shehzad, K.; Ahmad, I.; Eshmawi, A.; Amin, A.H.; Tag-Eldin, E. A deep learning-based approach for the diagnosis of acute lymphoblastic leukemia. Electronics 2022, 11, 3168. [Google Scholar] [CrossRef]
- Shen, Y.; Yu, J.; Zhou, J.; Hu, G. Twenty-Five Years of Evolution and Hurdles in Electronic Health Records and Interoperability in Medical Research: Comprehensive Review. J. Med. Internet Res. 2025, 27, e59024. [Google Scholar] [CrossRef] [PubMed]
- Salman, M.; Das, P.K.; Mohanty, S.K. A Systematic Review on Recent Advancements in Deep Learning and Mathematical Modeling for Efficient Detection of Glioblastoma. IEEE Trans. Instrum. Meas. 2024, 73, 2533134. [Google Scholar] [CrossRef]
- Thiriveedhi, A.; Ghanta, S.; Biswas, S.; Pradhan, A.K. ALL-Net: Integrating CNN and explainable-AI for enhanced diagnosis and interpretation of acute lymphoblastic leukemia. PeerJ Comput. Sci. 2025, 11, e2600. [Google Scholar] [CrossRef] [PubMed]
- Genovese, A.; Piuri, V.; Scotti, F. A decision support system for acute lymphoblastic leukemia detection based on explainable artificial intelligence. Image Vis. Comput. 2024, 151, 105298. [Google Scholar] [CrossRef]
- Jawahar, M.; Anbarasi, L.J.; Narayanan, S.; Gandomi, A.H. An attention-based deep learning for acute lymphoblastic leukemia classification. Sci. Rep. 2024, 14, 17447. [Google Scholar] [CrossRef]
- Alshehri, A. Skin-NeT: Skin Cancer Diagnosis Using VGG and ResNet-Based Ensemble Learning Approaches. Trait. Signal 2024, 41, 1689–1705. [Google Scholar] [CrossRef]
- Kinger, S.; Kulkarni, V. A review of explainable AI in medical imaging: Implications and applications. Int. J. Comput. Appl. 2024, 46, 983–997. [Google Scholar] [CrossRef]
- Diallo, R.; Edalo, C.; Awe, O.O. Machine Learning Evaluation of Imbalanced Health Data: A Comparative Analysis of Balanced Accuracy, MCC, and F1 Score. In Practical Statistical Learning and Data Science Methods: Case Studies from LISA 2020 Global Network, USA; Springer: Cham, Switzerland, 2024; pp. 283–312. [Google Scholar]
- Rai, H.M.; Yoo, J.; Razaque, A. Comparative analysis of machine learning and deep learning models for improved cancer detection: A comprehensive review of recent advancements in diagnostic techniques. Expert Syst. Appl. 2024, 255, 124838. [Google Scholar] [CrossRef]
- Vimbi, V.; Shaffi, N.; Mahmud, M. Interpreting artificial intelligence models: A systematic review on the application of LIME and SHAP in Alzheimer’s disease detection. Brain Inform. 2024, 11, 10. [Google Scholar]
- Xing, F.; Xie, Y.; Su, H.; Liu, F.; Yang, L. Deep learning in microscopy image analysis: A survey. IEEE Trans. Neural Netw. Learn. Syst. 2017, 29, 4550–4568. [Google Scholar]
- Bello, M.; Amador, R.; García, M.-M.; Bello, R.; Cordón, O.; Herrera, F. Meta-Explainers: A Unified Ensemble Approach for Multifaceted XAI. Int. J. Intell. Syst. 2025, 2025, 4841666. [Google Scholar]
- Akgündoğdu, A.; Çelikbaş, Ş. Explainable Deep Learning Framework for Brain Tumor Detection: Integrating LIME, Grad-CAM, and SHAP for Enhanced Accuracy. Med. Eng. Phys. 2025, 144, 104405. [Google Scholar] [CrossRef]
- Saarela, M.; Podgorelec, V. Recent Applications of Explainable AI (XAI): A Systematic Literature Review. Appl. Sci. 2024, 14, 8884. [Google Scholar] [CrossRef]
- Cahour, B.; Forzy, J.F. Does projection into use improve trust and exploration? An example with a cruise control system. Saf. Sci. 2009, 47, 1260–1270. [Google Scholar] [CrossRef]
- Landis, J.R.; Koch, G.G. The measurement of observer agreement for categorical data. Biometrics 1977, 33, 159–174. [Google Scholar] [CrossRef] [PubMed]
- Oybek Kizi, R.F.; Theodore Armand, T.P.; Kim, H.C. A Review of Deep Learning Techniques for Leukemia Cancer Classification Based on Blood Smear Images. Appl. Biosci. 2025, 4, 9. [Google Scholar] [CrossRef]
- Rajpurkar, P.; Irvin, J.; Ball, R.L.; Zhu, K.; Yang, B.; Mehta, H.; Duan, T.; Ding, D.; Bagul, A.; Langlotz, C.P.; et al. Deep learning for chest radiograph diagnosis: A retrospective comparison of the CheXNeXt algorithm to practicing radiologists. PLoS Med. 2018, 15, e1002686. [Google Scholar]
- Makem, M.; Tamas, L.; Busoniu, L. A Reliable Approach for Identifying Acute Lymphoblastic Leukaemia in Microscopic Imaging. Front. Artif. Intell. 2025, 8, 1620252. [Google Scholar]
- Hasanaath, A.A.; Mohammed, A.S.; Latif, G.; Abdelhamid, S.E.; Alghazo, J.; Hussain, A.A. Acute lymphoblastic leukemia detection using ensemble features from multiple deep CNN models. Electron. Res. Arch. 2024, 32, 2407–2423. [Google Scholar] [CrossRef]
- Ahmed, F. Transfer learning with efficientnet for accurate leukemia cell classification. arXiv 2025, arXiv:2508.06535. [Google Scholar]
- Singh, S.; Avasthi, D.; Singh, N.; Singh, A.; Prakash, S. Leveraging Pre-Trained CNNs and Ensemble Methods for Improved Diagnosis of Acute Lymphoblastic Leukemia. In Proceedings of the 2024 International Conference on Decision Aid Sciences and Applications (DASA), Manama, Bahrain, 11–12 December 2024; pp. 1–6. [Google Scholar]
- Wabro, A.; Herrmann, M.; Winkler, E.C. When time is of the essence: Ethical reconsideration of XAI in time-sensitive environments. J. Med. Ethics 2025, 51, 516–520. [Google Scholar]
- Javeed, M.U.; Aslam, S.M.; Sadiqa, H.A.; Raza, A.; Iqbal, M.M.; Akram, M. Phishing Website URL Detection Using a Hybrid Machine Learning Approach. J. Comput. Biomed. Inform. 2025, 9. [Google Scholar]
- Zuberi, H.H.; Liu, S.; Bilal, M.; Alharbi, A.; Jaffar, A.; Mohsan, S.A.H.; Miyajan, A.; Khan, M.A. Deep-Neural-Network-Based Receiver Design for Downlink Non-Orthogonal Multiple-Access Underwater Acoustic Communication. J. Mar. Sci. Eng. 2023, 11, 2184. [Google Scholar] [CrossRef]
- Ma, X.; Raza, W.; Wu, Z.; Bilal, M.; Zhou, Z.; Ali, A. A nonlinear distortion removal based on deep neural network for underwater acoustic OFDM communication with the mitigation of peak to average power ratio. Appl. Sci. 2020, 10, 4986. [Google Scholar] [CrossRef]
- Raza, A.; Zongxin, S.; Qiao, G.; Javed, M.; Bilal, M.; Zuberi, H.H.; Mohsin, M. Automated classification of humpback whale calls in four regions using convolutional neural networks and multi scale deep feature aggregation (MSDFA). Measurement 2025, 255, 118038. [Google Scholar] [CrossRef]
- Liu, S.; Zuberi, H.H.; Arfeen, Z.; Zhang, X.; Bilal, M.; Sun, Z. Spectral Efficient Neural Network-Based M-ary Chirp Spread Spectrum Receivers for Underwater Acoustic Communication. Arab. J. Sci. Eng. 2024, 49, 16593–16609. [Google Scholar] [CrossRef]
- Farid, G.; Bilal, M.; Zhang, L.; Alharbi, A.A.; Ahmed, I.; Azhar, M. An Improved Deep Q-Learning Approach for Navigation of an Autonomous UAV Agent in 3D Obstacle-Cluttered Environment. Drones 2025, 9, 518. [Google Scholar] [CrossRef]
- Adil, M.; Liu, S.; Mazhar, S.; Jan, M.; Khan, A.; Bilal, M. A Fully Connected Neural Network Driven UWA Channel Estimation for Reliable Communication. In 2023 International Conference on Frontiers of Information Technology (FIT); IEEE: Piscataway, NJ, USA, 2023; Volume 12, pp. 310–315. [Google Scholar] [CrossRef]
- Xuezhi, X.; Ali, S.; Farid, G. Image Processing Applications in Visual Tracking Using Various Techniques with Embedment of Particle Filter- A Critical Review. J. Flow Vis. Image Process. 2017, 23. [Google Scholar] [CrossRef]
- Ali, S.; Bilal, M.; Alharbi, A.; Amin, R. A Novel Deep Reinforcement Learning Based Extended Fractal Radial Basis Function Network for State-of-Charge Estimation. IET Power Electron. 2025, 18, e70101. [Google Scholar] [CrossRef]
- Raza, A.; Qiao, G.; Li, L.; Javed, M.; Rakha, M.; Bilal, M.; Sankoh, A.; Javed, S. A Deep Learning Framework for Imbalanced Bioacoustic Call-Type Classification Using Focal Loss and SMOTE. In 2025 International Conference on Frontiers of Information Technology (FIT); IEEE: Piscataway, NJ, USA, 2026; Volume 12, pp. 1–6. [Google Scholar] [CrossRef]
- Sankoh, A.P.; Raza, A.; Parwez, K.; Shishah, W.; Alharbi, A.; Javed, M.; Bilal, M. Automated Facial Pain Assessment Using Dual-Attention CNN with Clinically Calibrated High-Reliability and Reproducibility Framework. Biomimetics 2026, 11, 51. [Google Scholar] [CrossRef]
- Raza, A.; Javed, M.; Aslam, S.M.; Javeed, M.U.; Samual, P.; Raheem, S.; Ejaz, G.; Arshad, M.A. AI-powered sentiment analysis for social media opinion mining: A hybrid NLP and machine learning approach. Int. J. Adv. Comput. Emerg. Technol. 2025, 1, 1–15. [Google Scholar] [CrossRef]
- Sankoh, A.P.; Raza, A.; Parwez, K.; Shishah, W.; Alharbi, A.; Javed, M.; Bilal, M. Automated Facial Pain Assessment Using Dual-Attention CNN with Medical-Grade Calibration and Reproducibility Framework. Preprints 2025, 2025112089. [Google Scholar]
- Javeed, M.U.; Aslam, S.M.; Aslam, S.; Iqbal, M.; Farhan, M.; Javed, M.; Raza, A. Unveiling ambivalence in reviews: Using Sentence-BERT and K-NN for airline recommendations. Tech. J. 2025, 30, 51–59. [Google Scholar]
- Raza, A.; Javeed, M.U.; Arshad, M.A.; Akram, W.; Aslam, S.M.; Ejaz, G.; Shoukat, A.; Shaukat, A. Intelligent image gallery system using deep learning for automated fruit and vegetable classification. Int. J. Adv. Comput. Emerg. Technol. 2025, 1, 1–15. [Google Scholar]
- Raza, A.; Javeed, M.U.; Javed, M.; Aslam, S.M.; Saleem, R.; Irshad, W.; Maqbool, H.; Ejaz, G. A hybrid machine learning framework for personalized news recommendation. Int. J. Adv. Comput. Emerg. Technol. 2025, 1, 49–62. [Google Scholar]
- Nauman, M.; Ashraf, S.; Javeed, M.U.; Aslam, S.M.; Farooq, U.; Raza, A. Deep transfer learning for COVID-19 screening: Benchmarking ResNet50, VGG16, and GoogleNet on chest X-ray images. Int. J. Adv. Comput. Emerg. Technol. 2025, 1, 69–83. [Google Scholar]
- Raza, A.; Javed, M.; Fayad, A.; Khan, A.Y. Advanced deep learning-based predictive modelling for analyzing trends and performance metrics in stock market. J. Account. Financ. Emerg. Econ. 2023, 9, 277–294. [Google Scholar]
- Hossain, M.I.; Zamzmi, G.; Mouton, P.R.; Salekin, M.S.; Sun, Y.; Goldgof, D. Explainable AI for medical data: Current methods, limitations, and future directions. ACM Comput. Surv. 2025, 57, 148. [Google Scholar] [CrossRef]
- Basiouni, A.; Abdelqader, K.; Shaalan, K. Unlocking the Future: Systematic Review of the Progress and Challenges in Explainable Artificial Intelligence (XAI). SSRN Electron. J. 2024. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.