Future Directions of Marine Myxobacterial Natural Product Discovery Inferred from Metagenomics
Abstract
1. Introduction
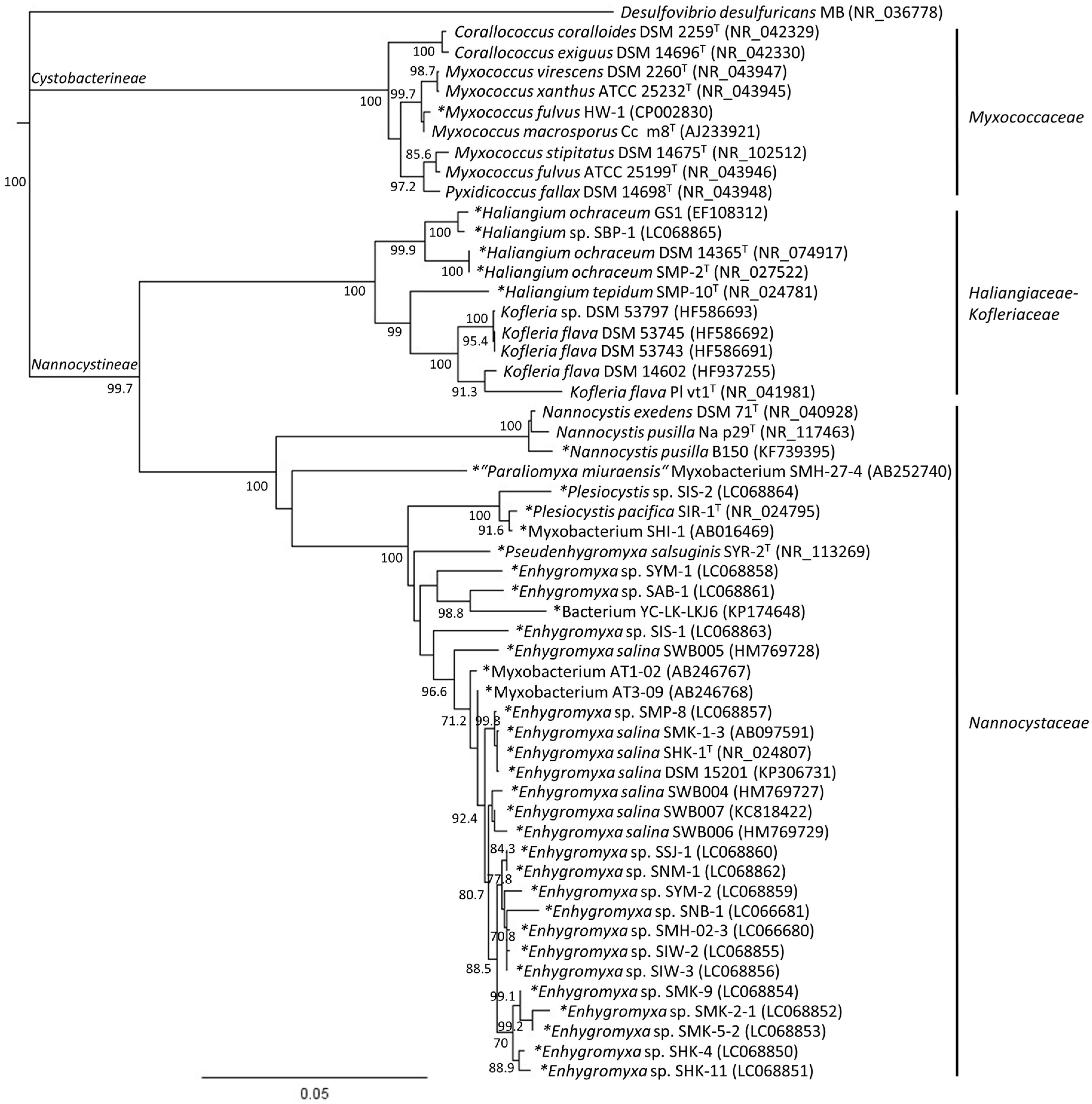
2. Phylogenetic Analysis of Cultured Marine- and Estuarine-Derived Myxobacteria
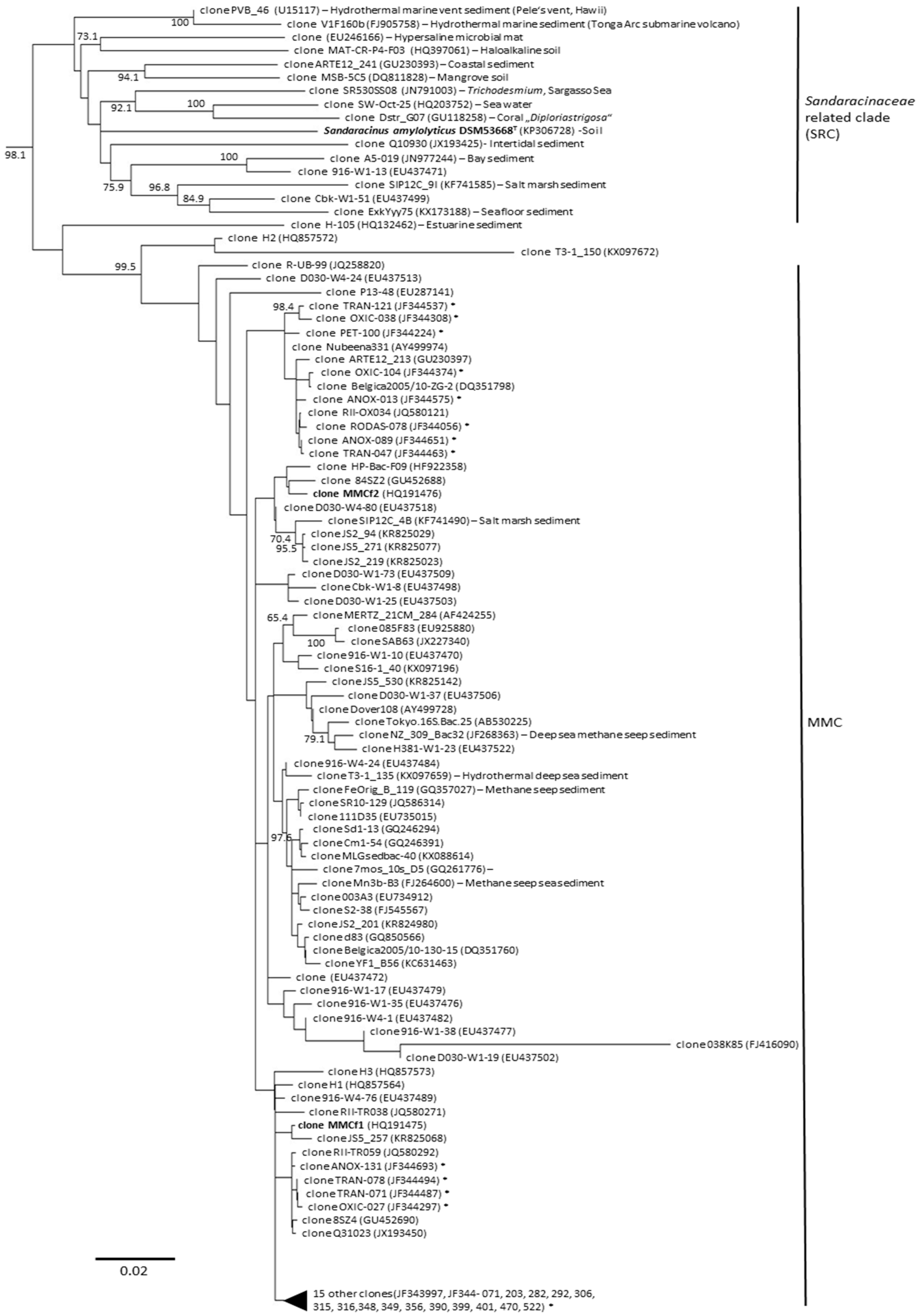
3. Diversity of Myxobacteria in the Suborder Sorangiineae as Inferred from Metagenomic Analyses
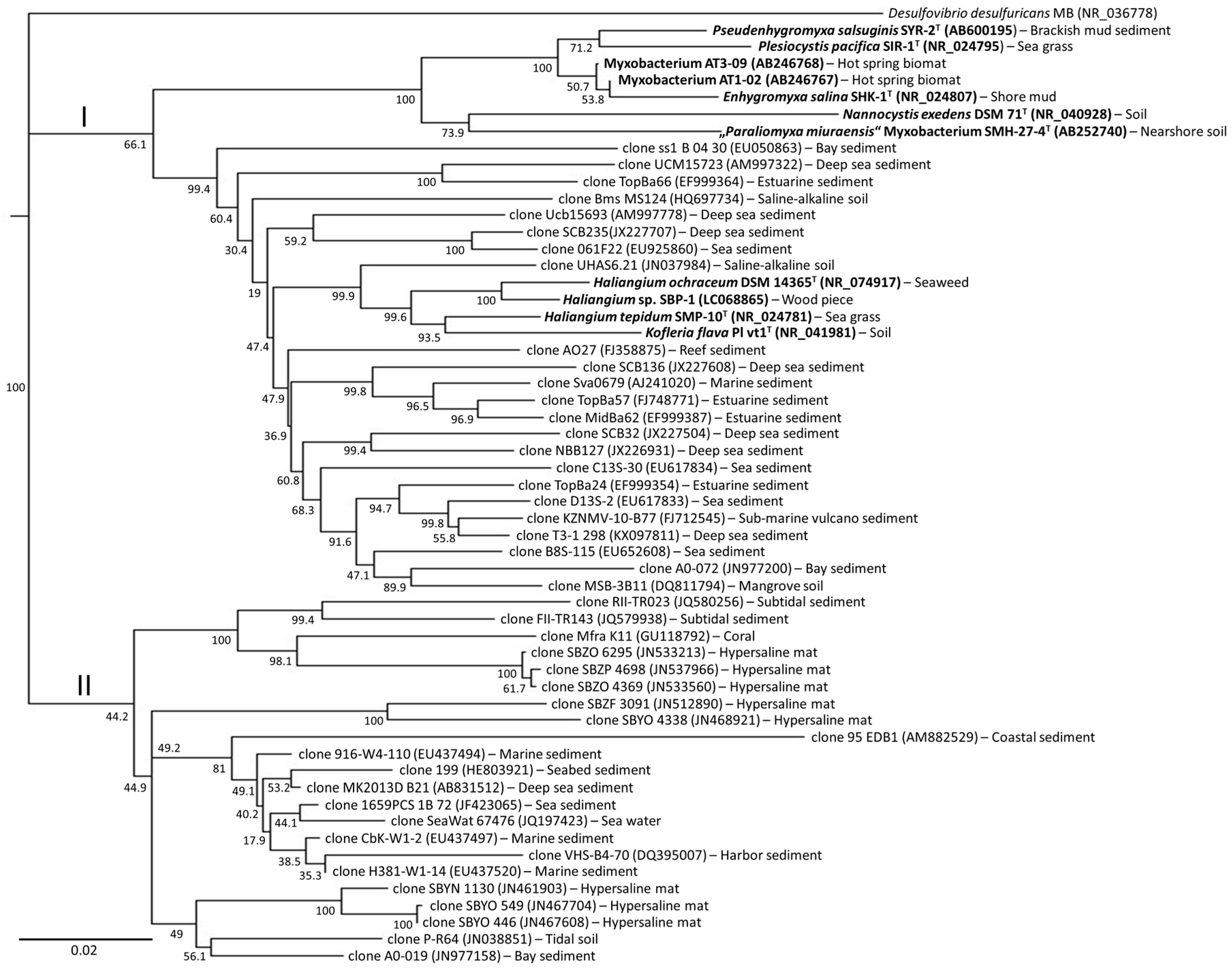
4. Diversity of Myxobacteria in the Nannocystineae Suborder
5. Characteristics of Novel Marine Myxobacteria and Parameters to Consider for Their Cultivation
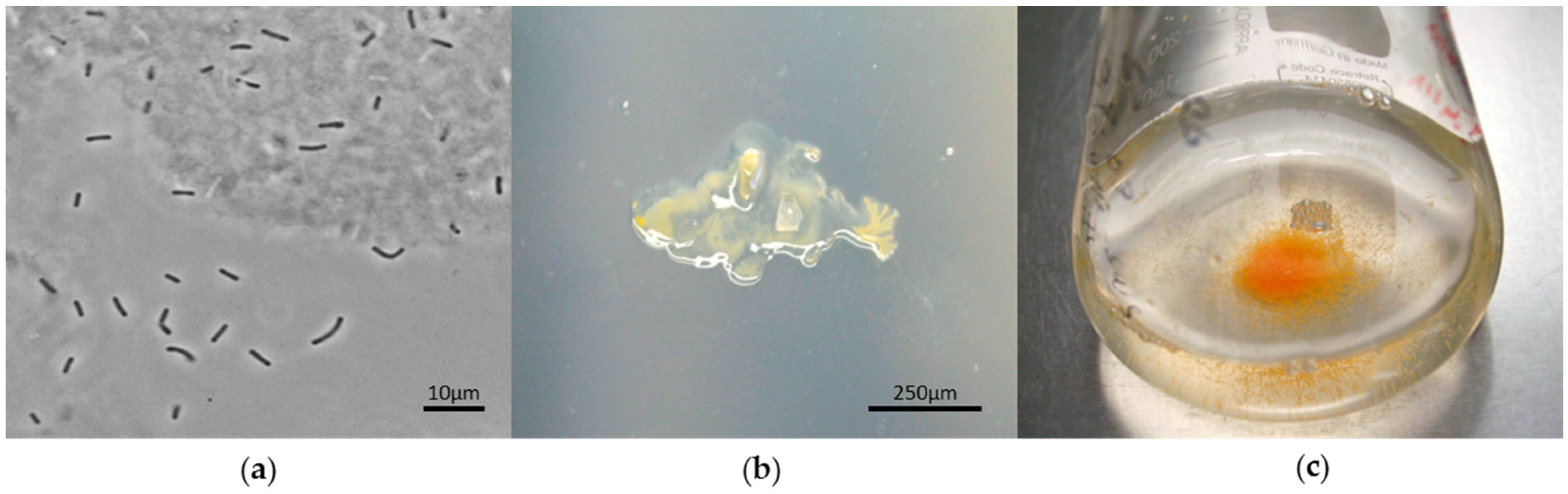
5.1. Structural Features: Rod-Shaped Cells with Blunted Ends
5.2. Reproductive Features: Reduction in Forming Fruiting Bodies
5.3. Chemical Predation and Defense: Lack of Microbial Predation and Cellulose Degradation but Agar Degradation
5.4. Requirement for Salinity: A Need for Salt and Associate Metal Cations
5.5. Temperature Regulation: Growth Possible in Wider Temperature Range
5.6. Life Cycle: Potentially Slow Growth
6. Can Marine Myxobacteria Access Novel Bioactive Secondary Metabolites?
7. Conclusions
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Starr, T.J.; Ordal, E.J. A Study of Marine Myxobacteria; Technical Report for University of Washington Department of Oceanography: Seattle, WA, USA, December 1953. [Google Scholar]
- Brockman, E.R. Fruiting myxobacteria from the South Carolina coast. J. Bacteriol. 1967, 94, 1253–1254. [Google Scholar] [PubMed]
- Rückert, G. Zur Verbreitung von Fruchtkörper-bildenden Myxobakterien in europäischen Strand- und Dünenböden. Zentralbl. Bakteriol. Parasitenkd. Infektionskr. Hyg. 1975, 130, 343–347. [Google Scholar] [CrossRef]
- Iizuka, T.; Jojima, Y.; Fudou, R.; Yamanaka, S. Isolation of myxobacteria from the marine environment. FEMS Microbiol. Lett. 1998, 169, 317–322. [Google Scholar] [CrossRef] [PubMed]
- Iizuka, T.; Jojima, Y.; Fudou, R.; Tokura, M.; Hiraishi, A.; Yamanaka, S. Enhygromyxa salina gen. nov., sp. nov., a slightly halophilic myxobacterium isolated from the coastal areas of Japan. Syst. Appl. Microbiol. 2003, 26, 189–196. [Google Scholar] [CrossRef] [PubMed]
- Iizuka, T.; Jojima, Y.; Fudou, R.; Hiraishi, A.; Ahn, J.W.; Yamanaka, S. Plesiocystis pacifica gen. nov., sp. nov., a marine myxobacterium that contains dihydrogenated menaquinone, isolated from the pacific coasts of Japan. Int. J. Syst. Evol. Microbiol. 2003, 53, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Iizuka, T.; Jojima, Y.; Hayakawa, A.; Fujii, T.; Yamanaka, S.; Fudou, R. Pseudenhygromyxa salsuginis gen. nov., sp. nov., a myxobacterium isolated from an estuarine marsh. Int. J. Syst. Evol. Microbiol. 2013, 63, 1360–1369. [Google Scholar] [CrossRef] [PubMed]
- Fudou, R.; Jojima, Y.; Iizuka, T.; Yamanaka, S. Haliangium ochraceum gen. nov., sp nov and Haliangium tepidum sp nov.: Novel moderately halophilic myxobacteria isolated from coastal saline environments. J. Gen. Appl. Microbiol. 2002, 48, 109–115. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.Z.; Hu, W.; Zhang, Y.Q.; Qiu, Z.; Zhang, Y.; Wu, B.H. A simple method to isolate salt-tolerant myxobacteria from marine samples. J. Microbiol. Methods 2002, 50, 205–209. [Google Scholar] [CrossRef]
- Gemperlein, K.; Zaburannyi, N.; Garcia, R.; La Clair, J.J.; Müller, R. Metabolic and biosynthetic diversity in marine myxobacteria. Mar. Drugs 2018, in press. [Google Scholar]
- Jiang, D.M.; Kato, C.; Zhou, X.W.; Wu, Z.H.; Sato, T.; Li, Y.Z. Phylogeographic separation of marine and soil myxobacteria at high levels of classification. ISME J. 2010, 4, 1520–1530. [Google Scholar] [CrossRef] [PubMed]
- Garcia, R.; Müller, R. The Family Myxococcaceae. In The Prokaryotes: Deltaproteobacteria and Epsilonproteobacteria; Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E., Thompson, F., Eds.; Springer: Berlin/Heidelberg, Germany, 2014; pp. 191–212. [Google Scholar]
- Li, Z.F.; Li, X.; Liu, H.; Liu, X.; Han, K.; Wu, Z.H.; Hu, W.; Li, F.F.; Li, Y.Z. Genome sequence of the halotolerant marine bacterium Myxococcus fulvus HW-1. J. Bacteriol. 2011, 193, 5015–5016. [Google Scholar] [CrossRef] [PubMed]
- Garcia, R.; Gerth, K.; Stadler, M.; Dogma, I.J., Jr.; Müller, R. Expanded phylogeny of myxobacteria and evidence for cultivation of the ‘unculturables’. Mol. Phylogenet. Evol. 2010, 57, 878–887. [Google Scholar] [CrossRef] [PubMed]
- Ojika, M.; Inukai, Y.; Kito, Y.; Hirata, M.; Iizuka, T.; Fudou, R. Miuraenamides: Antimicrobial cyclic depsipeptides isolated from a rare and slightly halophilic myxobacterium. Chem. Asian J. 2008, 3, 126–133. [Google Scholar] [CrossRef] [PubMed]
- Iizuka, T.; Fudou, R.; Jojima, Y.; Ogawa, S.; Yamanaka, S.; Inukai, Y.; Ojika, M. Miuraenamides A and B, novel antimicrobial cyclic depsipeptides from a new slightly halophilic myxobacterium: Taxonomy, production, and biological properties. J. Antibiot. 2006, 59, 385–391. [Google Scholar] [CrossRef] [PubMed]
- Brinkhoff, T.; Fischer, D.; Vollmers, J.; Voget, S.; Beardsley, C.; Thole, S.; Mussmann, M.; Kunze, B.; Wagner-Döbler, I.; Daniel, R.; Simon, M. Biogeography and phylogenetic diversity of a cluster of exclusively marine myxobacteria. ISME J. 2012, 6, 1260–1272. [Google Scholar] [CrossRef] [PubMed]
- López-García, P.; Duperron, S.; Philippot, P.; Foriel, J.; Susini, J.; Moreira, D. Bacterial diversity in hydrothermal sediment and epsilonproteobacterial dominance in experimental microcolonizers at the Mid-Atlantic Ridge. Environ. Microbiol. 2003, 5, 961–976. [Google Scholar] [CrossRef] [PubMed]
- Acosta-González, A.; Rosselló-Móra, R.; Marqués, S. Characterization of the anaerobic microbial community in oil-polluted subtidal sediments: Aromatic biodegradation potential after the Prestige oil spill. Environ. Microbiol. 2013, 15, 77–92. [Google Scholar] [CrossRef] [PubMed]
- Forget, N.L.; Murdock, S.A.; Juniper, S.K. Bacterial diversity in Fe-rich hydrothermal sediments at two South Tonga Arc submarine volcanoes. Geobiology 2010, 8, 417–432. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.H.; Liao, L.; Wang, C.S.; Ma, W.L.; Meng, F.X.; Wu, M.; Xu, X.W. A comparison of microbial communities in deep-sea polymetallic nodules and the surrounding sediments in the Pacific Ocean. Deep Sea Res. Part I 2013, 79, 40–49. [Google Scholar] [CrossRef]
- Schauer, R.; Bienhold, C.; Ramette, A.; Harder, J. Bacterial diversity and biogeography in deep-sea surface sediments of the South Atlantic Ocean. ISME J. 2010, 4, 159–170. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.; Zheng, Y.; Peng, X.; Zhou, H.; Zhang, C.; Xiao, X.; Wang, F. Vertical distribution and diversity of sulfate-reducing prokaryotes in the Pearl River estuarine sediments, Southern China. FEMS Microbiol. Ecol. 2009, 70, 93–106. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Ki, J.S.; Qian, P.Y. Microbial diversity in polluted harbor sediments I: Bacterial community assessment based on four clone libraries of 16S rDNA. Estuar. Coast. Shelf Sci. 2008, 76, 668–681. [Google Scholar] [CrossRef]
- Aoki, M.; Ehara, M.; Saito, Y.; Yoshioka, H.; Miyazaki, M.; Saito, Y.; Miyashita, A.; Kawakami, S.; Yamaguchi, T.; Ohashi, A.; et al. A long-term cultivation of an anaerobic methane-oxidizing microbial community from deep-sea methane-seep sediment using a continuous-flow bioreactor. PLoS ONE 2014, 9, e105356. [Google Scholar] [CrossRef] [PubMed]
- Paissé, S.; Coulon, F.; Goñi-Urriza, M.; Peperzak, L.; McGenity, T.J.; Duran, R. Structure of bacterial communities along a hydrocarbon contamination gradient in a coastal sediment. FEMS Microbiol. Ecol. 2008, 66, 295–305. [Google Scholar] [CrossRef] [PubMed]
- Harris, J.K.; Caporaso, J.G.; Walker, J.J.; Spear, J.R.; Gold, N.J.; Robertson, C.E.; Hugenholtz, P.; Goodrich, J.; McDonald, D.; Knights, D.; et al. Phylogenetic stratigraphy in the Guerrero Negro hypersaline microbial mat. ISME J. 2013, 7, 50–60. [Google Scholar] [CrossRef] [PubMed]
- Sunagawa, S.; Woodley, C.M.; Medina, M. Threatened corals provide underexplored microbial habitats. PLoS ONE 2010, 5, e9554. [Google Scholar] [CrossRef] [PubMed]
- Reichenbach, H. Order VIII. Myxococcales. Tchan, Pochon and Prévot 1948, 398AL. In Bergey’s Manual® of Systematic Bacteriology, 2nd ed.; Garrity, G., Brenner, D.J., Krieg, N.R., Staley, J.T., Eds.; Springer: New York, NY, USA, 2005; Volume 2, Part C; pp. 1059–1144. [Google Scholar]
- Garcia, R.; Müller, R. The Family Haliangiaceae. In The Prokaryotes; Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E., Thompson, F., Eds.; Springer: Berlin/Heidelberg, Germany, 2014; pp. 173–181. [Google Scholar]
- Garcia, R.; Müller, R. The Family Nannocystaceae. In The Prokaryotes; Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E., Thompson, F., Eds.; Springer: Berlin/Heidelberg, Germany, 2014; pp. 213–229. [Google Scholar]
- Mohr, K.I.; Garcia, R.O.; Gerth, K.; Irschik, H.; Müller, R. Sandaracinus amylolyticus gen. nov., sp. nov., a starch-degrading soil myxobacterium, and description of Sandaracinaceae fam. nov. Int. J. Syst. Evol. Microbiol. 2012, 62, 1191–1198. [Google Scholar] [CrossRef] [PubMed]
- Kimura, Y.; Kawasaki, S.; Yoshimoto, H.; Takegawa, K. Glycine betaine biosynthesized from glycine provides an osmolyte for cell growth and spore germination during osmotic stress in Myxococcus xanthus. J. Bacteriol. 2010, 192, 1467–1470. [Google Scholar] [CrossRef] [PubMed]
- Pan, H.W.; Tan, Z.G.; Liu, H.; Li, Z.F.; Zhang, C.Y.; Li, C.Y.; Li, J.; Li, Y.Z. Hdsp, a horizontally transferred gene required for social behavior and halotolerance in salt-tolerant Myxococcus fulvus HW-1. ISME J. 2010, 4, 1282–1289. [Google Scholar] [CrossRef] [PubMed]
- Moghaddam, J.A.; Boehringer, N.; Burdziak, A.; Kunte, H.J.; Galinski, E.A.; Schäberle, T.F. Different strategies of osmoadaptation in the closely related marine myxobacteria Enhygromyxa salina SWB007 and Plesiocystis pacifica SIR-1. Microbiology 2016. [Google Scholar] [CrossRef] [PubMed]
- Felder, S.; Kehraus, S.; Neu, E.; Bierbaum, G.; Schäberle, T.F.; König, G.M. Salimyxins and enhygrolides: Antibiotic, sponge-related metabolites from the obligate marine myxobacterium Enhygromyxa salina. ChemBioChem 2013, 14, 1363–1371. [Google Scholar] [CrossRef] [PubMed]
- Tomura, T.; Nagashima, S.; Yamazaki, S.; Iizuka, T.; Fudou, R.; Ojika, M. An unusual diterpene-enhygromic acid and deoxyenhygrolides from a marine myxobacterium, Enhygromyxa sp. Mar. Drugs 2017, 15, 109. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Tomura, T.; Sato, J.; Iizuka, T.; Fudou, R.; Ojika, M. Isolation and biosynthetic analysis of haliamide, a new PKS-NRPS hybrid metabolite from the marine myxobacterium Haliangium ochraceum. Molecules 2016, 21, 59. [Google Scholar] [CrossRef] [PubMed]
- Fudou, R.; Iizuka, T.; Sato, S.; Ando, T.; Shimba, N.; Yamanaka, S. Haliangicin, a novel antifungal metabolite produced by a marine myxobacterium. 2. Isolation and structural elucidation. J. Antibiot. 2001, 54, 153–156. [Google Scholar] [CrossRef] [PubMed]
- Fudou, R.; Iizuka, T.; Yamanaka, S. Haliangicin, a novel antifungal metabolite produced by a marine myxobacterium. 1. Fermentation and biological characteristics. J. Antibiot. 2001, 54, 149–152. [Google Scholar] [CrossRef] [PubMed]
- Felder, S.; Dreisigacker, S.; Kehraus, S.; Neu, E.; Bierbaum, G.; Wright, P.R.; Menche, D.; Schäberle, T.F.; König, G.M. Salimabromide: Unexpected chemistry from the obligate marine myxobacterium Enhygromxya salina. Chemistry 2013, 19, 9319–9324. [Google Scholar] [CrossRef] [PubMed]
- Davila-Cespedes, A.; Hufendiek, P.; Crusemann, M.; Schaberle, T.F.; Konig, G.M. Marine-derived myxobacteria of the suborder Nannocystineae: An underexplored source of structurally intriguing and biologically active metabolites. Beilstein J. Org. Chem. 2016, 12, 969–984. [Google Scholar] [CrossRef] [PubMed]
- Sumiya, E.; Shimogawa, H.; Sasaki, H.; Tsutsumi, M.; Yoshita, K.; Ojika, M.; Suenaga, K.; Uesugi, M. Cell-morphology profiling of a natural product library identifies bisebromoamide and miuraenamide A as actin filament stabilizers. ACS Chem. Biol. 2011, 6, 425–431. [Google Scholar] [CrossRef] [PubMed]
- Moghaddam, J.A.; Poehlein, A.; Fisch, K.; Alanjary, M.; Daniel, R.; König, G.M.; Schäberle, T.F. Draft Genome Sequences of the Obligatory Marine Myxobacterial Strains Enhygromyxa salina SWB005 and SWB007. Genome Announc. 2018, 6, e00324-18. [Google Scholar] [CrossRef] [PubMed]
- Ivanova, N.; Daum, C.; Lang, E.; Abt, B.; Kopitz, M.; Saunders, E.; Lapidus, A.; Lucas, S.; Glavina Del Rio, T.; Nolan, M.; et al. Complete genome sequence of Haliangium ochraceum type strain (SMP-2). Stand. Genom. Sci. 2010, 2, 96–106. [Google Scholar] [CrossRef] [PubMed]
- Komaki, H.; Fudou, R.; Iizuka, T.; Nakajima, D.; Okazaki, K.; Shibata, D.; Ojika, M.; Harayama, S. PCR detection of type I polyketide synthase genes in myxobacteria. Appl. Environ. Microbiol. 2008, 74, 5571–5574. [Google Scholar] [CrossRef] [PubMed]
- La Clair, J.J.; Loveridge, S.T.; Tenney, K.; O’Neil-Johnson, M.; Chapman, E.; Crews, P. In situ natural product discovery via an artificial marine sponge. PLoS ONE 2014, 9, e100474. [Google Scholar] [CrossRef] [PubMed]

| Bacterial Strains | GenBank Accession Number |
|---|---|
| Bacterium YC-LK-LKJ6 | KP174648 |
| Corallococcus coralloides DSM 2259 T | NR_042329 |
| Corallococcus exiguus DSM 14696 T | NR_042330 |
| Desulfovibrio desulfuricans MB | NR_036778 |
| Enhygromyxa sp. SAB-1 | LC068861 |
| Enhygromyxa salina SHK-1 T | NR_024807 |
| Enhygromyxa sp. SHK-4 | LC068850 |
| Enhygromyxa sp. SHK-11 | LC068851 |
| Enhygromyxa sp. SIS-1 | LC068863 |
| Enhygromyxa sp. SIW-2 | LC068855 |
| Enhygromyxa sp. SIW-3 | LC068856 |
| Enhygromyxa sp. SMH-02-3 | LC066680 |
| Enhygromyxa salina SMK-1-3 | AB097591 |
| Enhygromyxa sp. SMK-2-1 | LC068852 |
| Enhygromyxa sp. SMK-5-2 | LC068853 |
| Enhygromyxa sp. SMK-9 | LC068854 |
| Enhygromyxa sp. SMP-8 | LC068857 |
| Enhygromyxa sp. SNB-1 | LC066681 |
| Enhygromyxa sp. SNM-1 | LC068862 |
| Enhygromyxa sp. SSJ-1 | LC068860 |
| Enhygromyxa sp. SYM-1 | LC068858 |
| Enhygromyxa sp. SYM-2 | LC068859 |
| Enhygromyxa salina DSM 15201 | KP306731 |
| Enhygromyxa salina SWB004 | HM769727 |
| Enhygromyxa salina SWB005 | HM769728 |
| Enhygromyxa salina SWB006 | HM769729 |
| Enhygromyxa salina SWB007 | KC818422 |
| Haliangium sp. SBP-1 | LC068865 |
| Haliangium ochraceum GS1 | EF108312 |
| Haliangium ochraceum DSM 14365 T | NR_074917 |
| Haliangium ochraceum SMP-2 T | NR_027522 |
| Haliangium tepidum SMP-10 T | NR_024781 |
| Kofleria sp. DSM 53797 | HF586693 |
| Kofleria flava DSM 53745 | HF586692 |
| Kofleria flava DSM 53743 | HF586691 |
| Kofleria flava DSM 14602 | HF937255 |
| Kofleria flava Pl vt1 T | NR_041981 |
| Myxobacterium SHI-1 | AB016469 |
| Myxobacterium SMH-27-4 (“Paraliomyxa miuraensis”) * | AB252740 |
| Myxobacterium AT1-02 | AB246767 |
| Myxobacterium AT3-09 | AB246768 |
| Myxococcus fulvus HW-1 | CP002830 |
| Myxococcus fulvus ATCC 25199 T | NR_043946 |
| Myxococcus macrosporus Cc m8 T | AJ233921 |
| Myxococcus stipitatus DSM 14675 T | NR_102512 |
| Myxococcus virescens DSM 2260 T | NR_043947 |
| Myxococcus xanthus ATCC 25232 T | NR_043945 |
| Nannocystis exedens DSM 71 T | NR 040928 |
| Nannocystis pusilla Na p29 T | NR_117463 |
| Nannocystis pusilla B150 | KF739395 |
| Plesiocystis pacifica SIR-1 T | NR_024795 |
| Plesiocystis sp. SIS-2 | LC068864 |
| Pseudenhygromyxa salsuginis SYR-2 T | NR_113269 |
| Pyxidicoccus fallax DSM 14698 T | NR_043948 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Garcia, R.; La Clair, J.J.; Müller, R. Future Directions of Marine Myxobacterial Natural Product Discovery Inferred from Metagenomics. Mar. Drugs 2018, 16, 303. https://doi.org/10.3390/md16090303
Garcia R, La Clair JJ, Müller R. Future Directions of Marine Myxobacterial Natural Product Discovery Inferred from Metagenomics. Marine Drugs. 2018; 16(9):303. https://doi.org/10.3390/md16090303
Chicago/Turabian StyleGarcia, Ronald, James J. La Clair, and Rolf Müller. 2018. "Future Directions of Marine Myxobacterial Natural Product Discovery Inferred from Metagenomics" Marine Drugs 16, no. 9: 303. https://doi.org/10.3390/md16090303
APA StyleGarcia, R., La Clair, J. J., & Müller, R. (2018). Future Directions of Marine Myxobacterial Natural Product Discovery Inferred from Metagenomics. Marine Drugs, 16(9), 303. https://doi.org/10.3390/md16090303





