Unsupervised Event Graph Representation and Similarity Learning on Biomedical Literature
Abstract
:1. Introduction
- In-depth literature analysis. We offer newcomers in the field a global perspective on the problem with insightful discussions and an extensive reference list, also providing systematic taxonomies.
- Deep Divergence Event Graph Kernels. A novel method of event graph similarity learning centered on constructing general event embeddings with deep graph kernels, considering both structure and semantics.
- Experimental results. We conduct extensive experiments to demonstrate the effectiveness of DDEGK in real-world scenarios. We show that our solution successfully recognizes fine- and coarse-grained similarities between biomedical events. Precisely, when used as features, the event representations learned by DDEGK achieve new state-of-the-art or competitive results on different extrinsic evaluation tasks, comprising sentence similarity, event classification, and clustering. To shed light on the performance of different embedding techniques, we compare with a rich set of baselines on nine datasets having distinct biological views.
2. Related Work
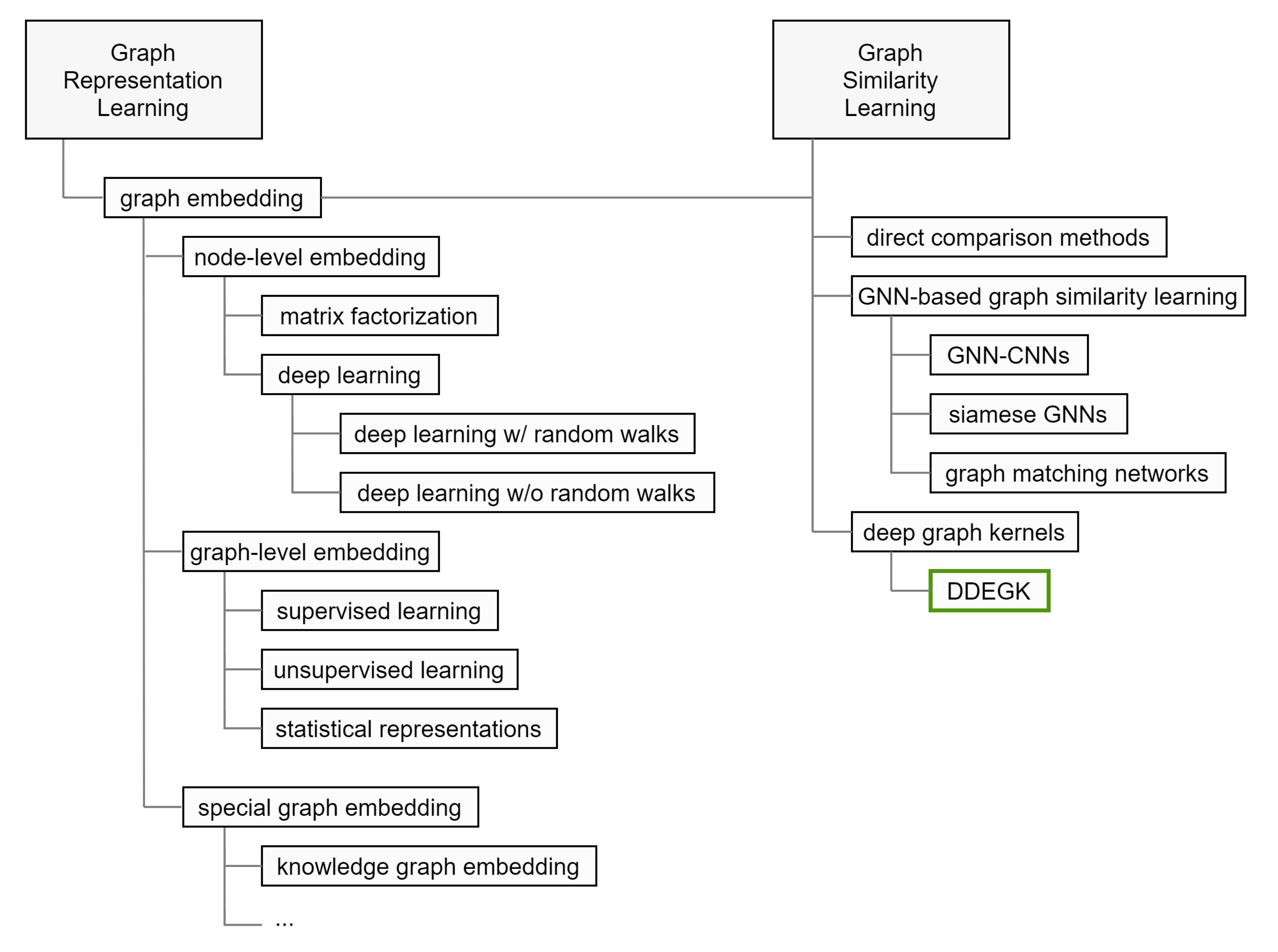
2.1. Individual Graph Embedding
2.1.1. Node Embedding
2.1.2. Whole-Graph Embedding
2.1.3. Knowledge Graph Embedding
2.2. Graph Similarity Computation with Cross-Graph Feature Interaction
2.2.1. Direct Comparison Methods
2.2.2. GNN-Based Graph Similarity Learning
2.2.3. Graph Kernels
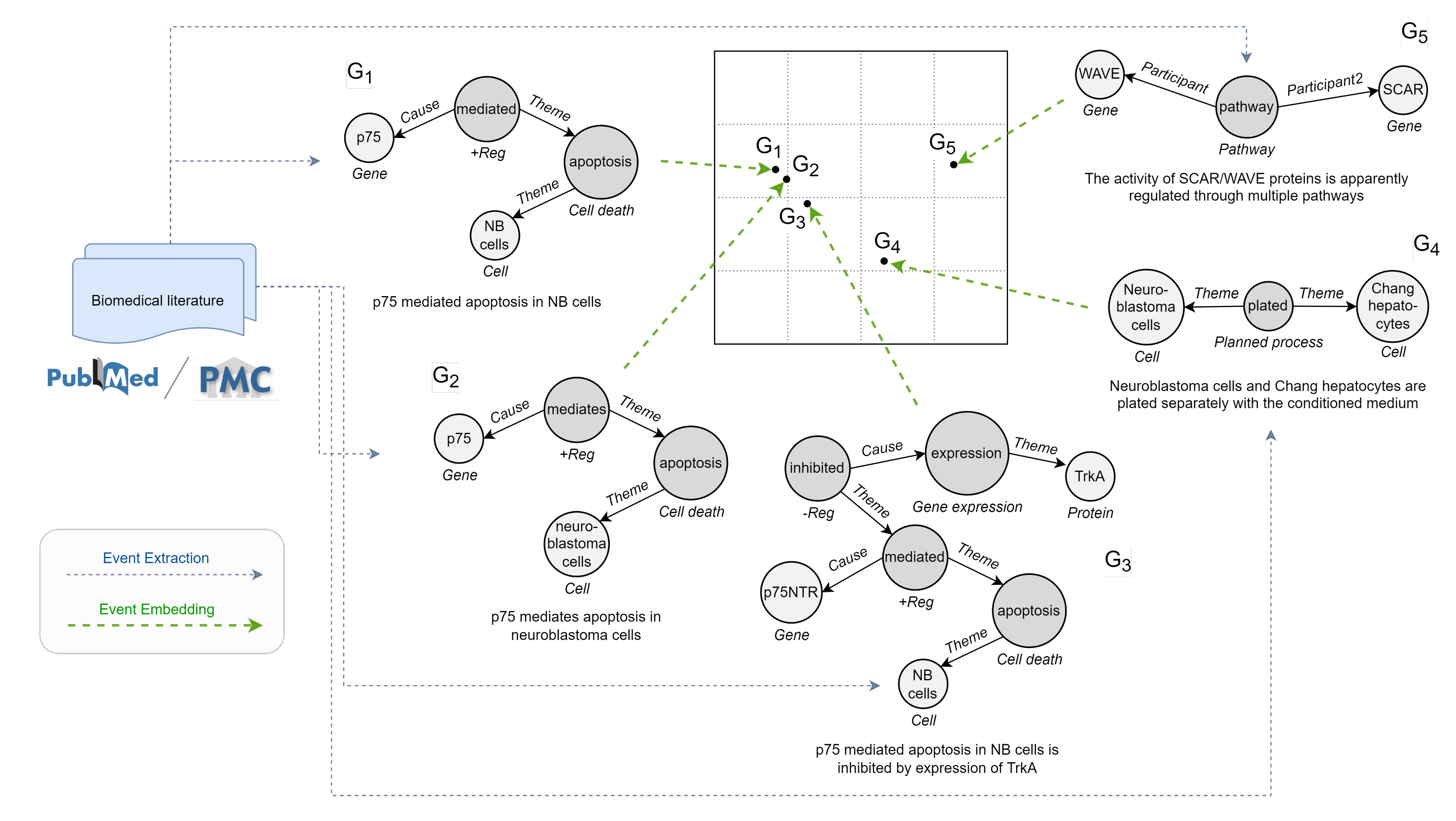
2.3. Event Embedding
3. Notation and Preliminaries
4. Materials and Methods
4.1. Datasets
- BioNLP-ST 2009 (ST09) [4]. Dataset taken from the first BioNLP-ST challenge, consisting of a sub-portion of the GENIA event corpus. It includes 13,623 events (total between train, validation, and test sets) mentioned in 1210 MEDLINE abstracts on human blood cells and transcription factors.
- Genia Event 2011 (GE11) [94]. Extended version of ST09, also including ≈4500 events collected from 14 PMC full-text articles.
- Epigenetics and Post-translational Modifications (EPI11) [95]. Dataset on epigenetic change and common protein post-translational modifications. It contains 3714 events extracted from 1200 abstracts.
- Infectious Diseases (ID11) [96]. Dataset on two-component regulatory systems; 4150 events recognized in 30 full papers.
- Multi-Level Event Extraction (MLEE) [3]. Dataset on blood vessel development from the subcellular to the whole organism; 6667 events from 262 abstracts.
- Genia Event 2013 (GE13) [97]. Updated version of GE11, with 9364 events extracted exclusively from 30 full papers.
- Cancer Genetics (CG13) [98]. Dataset on cancer biology, with 17,248 events from 600 abstracts.
- Pathway Curation (PC13) [99]. Dataset on reactions, pathways, and curation; 12,125 events from 525 abstracts.
- Gene Regulation Ontology (GRO13) [100]. Dataset on human gene regulation and transcription; 5241 events from 300 abstracts.
4.1.1. Data Preprocessing and Sampling
4.2. Deep Divergence Event Graph Kernels
| Dataset | # Graphs | # Nodes | # Edges | # Labels | ||||
|---|---|---|---|---|---|---|---|---|
| Min | Mean | Max | Mean | Graph | Node | Edge | ||
| ST09 | 1007 | 2 | 4 | 14 | 3 | 9 | 11 | 3 |
| GE11 | 1001 | 2 | 4 | 14 | 3 | 9 | 11 | 2 |
| EPI11 | 1002 | 2 | 3 | 6 | 2 | 10 | 11 | 1 |
| ID11 | 1001 | 2 | 3 | 14 | 2 | 9 | 16 | 3 |
| MLEE | 1012 | 2 | 4 | 15 | 3 | 15 | 39 | 8 |
| GE13 | 1011 | 2 | 3 | 13 | 2 | 8 | 13 | 2 |
| CG13 | 1033 | 2 | 3 | 13 | 2 | 23 | 50 | 8 |
| PC13 | 1020 | 2 | 4 | 18 | 3 | 15 | 25 | 9 |
| GRO13 | 1006 | 2 | 3 | 5 | 2 | 18 | 140 | 4 |
| BIO_ALL | 1216 | 2 | 3 | 14 | 3 | 53 | 101 | 12 |
4.2.1. Problem Definition
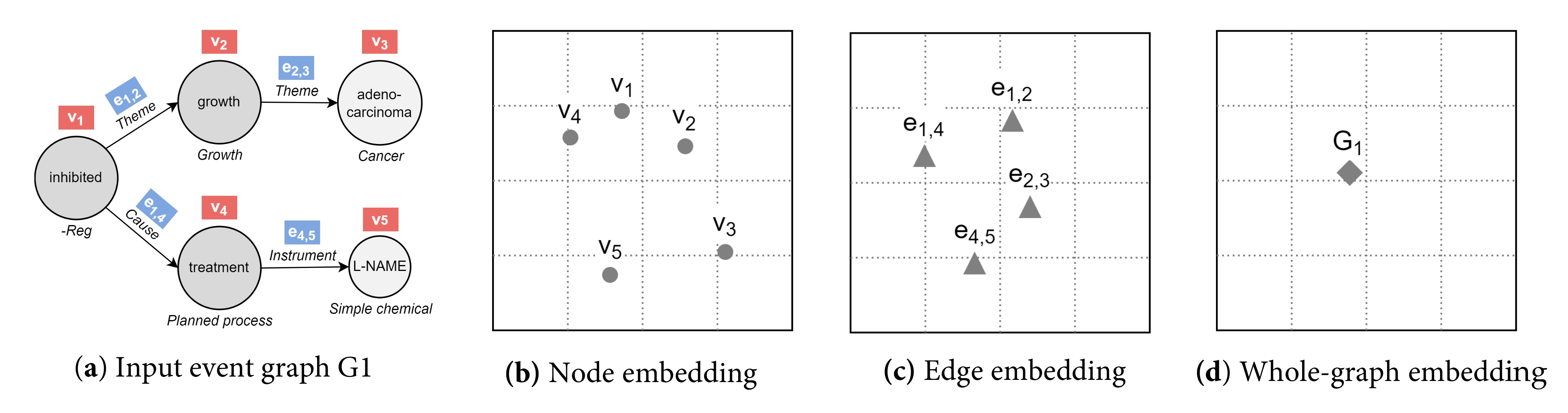
4.2.2. Event Graph Representation Alignment
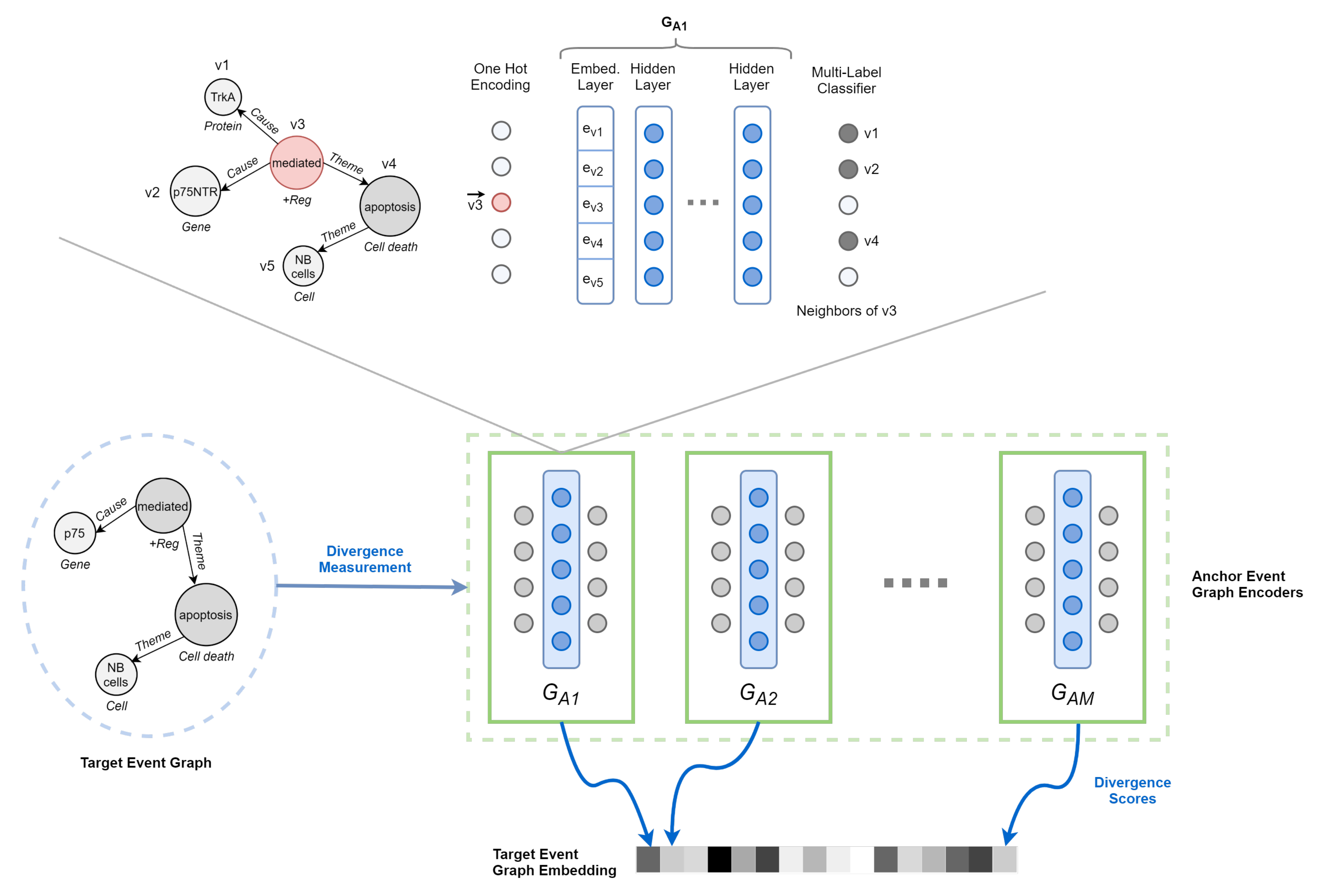
4.2.3. Event Graph Divergence and Embedding
4.2.4. Training
4.2.5. Scalability
4.3. Hardware and Software Setup
5. Experiments
5.1. Event Graph Classification
5.1.1. Baseline Methods
- Node embedding flat pooling. Each event is represented as the unsupervised aggregation of its constituent node vectors. We experiment with multiple unweighted flat pooling strategies, namely, mean, sum, and max. As for node representations, we examine (i) contextualized word embeddings from large-scale language models pre-trained on scientific and biomedical texts, (ii) node2vec [26]. The first point is realized by applying SciBERT [106] and BioBERT [108] (768 embedding size) on trigger and entity text spans: a common approach in the event-GRL area [88]. It condenses the semantic gist of an event based on the involved entities; it totally ignores structure and argument roles. In contrast, node2vec is a baseline for sequential methods which efficiently trade off between different proximity levels. The default walk length is 80, the number of walks per node is 10, return and in-out hyper-parameters are 1, and embedding size is 128.
- Node embedding + CNN. We use node2vec-PCA [40] (d = 2, i.e., one channel), which composes graph matrices from node2vec and then applies a CNN for supervised classification.
- Whole-graph embedding. We use graph2vec [51] to generate unsupervised structure-aware graph-level representations for our biomedical events. We work on labeled graphs, with labels denoting numerical identifiers for event types and entities. Default embedding size is 128, and the number of epochs is 100.
- GNN + supervised pooling. We use DGCNN [45], an end-to-end graph classification model made by GCNs with a sort pooling layer to derive permutation invariant graph embeddings. 1D-CNN then extracts features along with a fully-connected layer. Default k (normalized graph size) is 35.
5.1.2. Hyperparameters Search
5.1.3. Results
5.2. Between-Graph Clustering
5.2.1. Results
5.2.2. Influence of Embedding Dimensions
| Method | Event Type Clustering | Dataset Clustering | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ST09 | GE11 | EPI11 | ID11 | MLEE | GE13 | CG13 | PC13 | GRO13 | BIO_ALL | AVG | ||
| SciBERT (AVG) | −0.00915 | −0.00940 | 0.04082 | 0.01347 | −0.01139 | −0.00784 | 0.00087 | −0.01251 | −0.00343 | −0.01974 | −0.00183 | −0.01123 |
| 0.03638 | 0.06641 | 0.06644 | 0.09005 | 0.05810 | 0.04079 | 0.06644 | 0.04022 | 0.07669 | 0.04075 | 0.05823 | 0.05373 | |
| node2vec (AVG) | −0.48715 | −0.48165 | −0.58339 | −0.58004 | −0.46614 | −0.57659 | −0.51334 | −0.49366 | −0.54964 | −0.58171 | −0.53133 | −0.27637 |
| −0.02987 | −0.02539 | −0.00234 | −0.02405 | −0.05510 | −0.03748 | −0.04139 | −0.03689 | −0.03148 | −0.03830 | −0.03223 | 0.00068 | |
| graph2vec | −0.25687 | −0.20579 | −0.14446 | −0.27879 | −0.35015 | −0.28666 | −0.39072 | −0.36219 | −0.41060 | −0.45318 | −0.31394 | −0.16632 |
| 0.03789 | 0.04679 | 0.09058 | 0.14508 | 0.03394 | 0.07483 | 0.02883 | 0.02123 | 0.01443 | 0.02412 | 0.05177 | 0.02761 | |
| DDEGK (ours) | ||||||||||||
| w/random anchors | 0.21371 | 0.17281 | 0.27985 | 0.09267 | 0.08572 | 0.22813 | −0.05122 | −0.04587 | 0.05914 | −0.32140 | 0.07108 | 0.00686 |
| 0.33177 | 0.34138 | 0.54129 | 0.44621 | 0.16441 | 0.51622 | 0.17242 | 0.13066 | 0.39070 | 0.14349 | 0.31786 | 0.14349 | |
| w/random anchors per type | 0.23428 | 0.24598 | 0.28723 | 0.15141 | 0.10136 | 0.31270 | 0.08123 | 0.01809 | 0.08047 | −0.29036 | 0.12224 | 0.00007 |
| 0.35625 | 0.38622 | 0.39765 | 0.29661 | 0.15441 | 0.43387 | 0.15185 | 0.09296 | 0.23732 | 0.08320 | 0.25903 | 0.28101 | |
5.3. Visualization
5.4. Cross-Graph Attention
5.5. Semantic Textual Similarity
6. Discussion
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A


References
- Landhuis, E. Scientific literature: Information overload. Nature 2016, 535, 457–458. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Frisoni, G.; Moro, G.; Carbonaro, A. A Survey on Event Extraction for Natural Language Understanding: Riding the Biomedical Literature Wave. IEEE Access 2021, 9, 160721–160757. [Google Scholar] [CrossRef]
- Pyysalo, S.; Ohta, T.; Miwa, M.; Cho, H.; Tsujii, J.; Ananiadou, S. Event extraction across multiple levels of biological organization. Bioinformatics 2012, 28, 575–581. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Ohta, T.; Pyysalo, S.; Kano, Y.; Tsujii, J. Overview of BioNLP’09 Shared Task on Event Extraction. In Proceedings of the BioNLP 2009 Workshop Companion Volume for Shared Task, BioNLP@HLT-NAACL 2009–Shared Task, Boulder, CO, USA, 5 June 2009; Association for Computational Linguistics: Stroudsburg, PA, USA, 2009; pp. 1–9. [Google Scholar]
- Kim, J.; Pyysalo, S.; Ohta, T.; Bossy, R.; Nguyen, N.L.T.; Tsujii, J. Overview of BioNLP Shared Task 2011. In Proceedings of the BioNLP Shared Task 2011 Workshop, Portland, OR, USA, 24 June 2011; Tsujii, J., Kim, J., Pyysalo, S., Eds.; Association for Computational Linguistics: Stroudsburg, PA, USA, 2011; pp. 1–6. [Google Scholar]
- Nédellec, C.; Bossy, R.; Kim, J.; Kim, J.; Ohta, T.; Pyysalo, S.; Zweigenbaum, P. Overview of BioNLP Shared Task 2013. In Proceedings of the BioNLP Shared Task 2013 Workshop, Sofia, Bulgaria, 9 August 2013; Nédellec, C., Bossy, R., Kim, J., Kim, J., Ohta, T., Pyysalo, S., Zweigenbaum, P., Eds.; Association for Computational Linguistics: Stroudsburg, PA, USA, 2013; pp. 1–7. [Google Scholar]
- Henry, S.; McInnes, B.T. Literature Based Discovery: Models, methods, and trends. J. Biomed. Inf. 2017, 74, 20–32. [Google Scholar] [CrossRef]
- Björne, J.; Ginter, F.; Pyysalo, S.; Tsujii, J.; Salakoski, T. Complex event extraction at PubMed scale. Bioinformatics 2010, 26, 382–390. [Google Scholar] [CrossRef] [Green Version]
- Miwa, M.; Ohta, T.; Rak, R.; Rowley, A.; Kell, D.B.; Pyysalo, S.; Ananiadou, S. A method for integrating and ranking the evidence for biochemical pathways by mining reactions from text. Bioinformatics 2013, 29, 44–52. [Google Scholar] [CrossRef] [Green Version]
- Zhang, T.; Chen, M.; Bui, A.A.T. Diagnostic Prediction with Sequence-of-sets Representation Learning for Clinical Events. In Proceedings of the 18th International Conference on Artificial Intelligence in Medicine AIME 2020, Minneapolis, MN, USA, 25–28 August 2020; Michalowski, M., Moskovitch, R., Eds.; Lecture Notes in Comput. Science. Springer: Berlin/Heidelberg, Germany, 2020; Volume 12299, pp. 348–358. [Google Scholar] [CrossRef]
- Berant, J.; Srikumar, V.; Chen, P.; Linden, A.V.; Harding, B.; Huang, B.; Clark, P.; Manning, C.D. Modeling Biological Processes for Reading Comprehension. In Proceedings of the 2014 Conference on Empirical Methods in Natural Language Processing (EMNLP) 2014, Doha, Qatar, 25–29 October 2014; A Meeting of SIGDAT, a Special Interest Group of the ACL. Moschitti, A., Pang, B., Daelemans, W., Eds.; Association for Computational Linguistics: Stroudsburg, PA, USA, 2014. [Google Scholar]
- Bronstein, M.M.; Bruna, J.; LeCun, Y.; Szlam, A.; Vandergheynst, P. Geometric Deep Learning: Going beyond Euclidean data. IEEE Signal Process. Mag. 2017, 34, 18–42. [Google Scholar] [CrossRef] [Green Version]
- Wu, Z.; Pan, S.; Chen, F.; Long, G.; Zhang, C.; Yu, P.S. A Comprehensive Survey on Graph Neural Networks. IEEE Trans. Neural Netw. Learn. Syst. 2021, 32, 4–24. [Google Scholar] [CrossRef] [Green Version]
- Blumenthal, D.B.; Gamper, J. On the exact computation of the graph edit distance. Pattern Recognit. Lett. 2020, 134, 46–57. [Google Scholar] [CrossRef]
- Ma, G.; Ahmed, N.K.; Willke, T.L.; Yu, P.S. Deep graph similarity learning: A survey. Data Min. Knowl. Discov. 2021, 35, 688–725. [Google Scholar] [CrossRef]
- Chen, F.; Wang, Y.C.; Wang, B.; Kuo, C.C.J. Graph representation learning: A survey. APSIPA Trans. Signal Inf. Process. 2020, 9, E15. [Google Scholar] [CrossRef]
- Li, Y.; Gu, C.; Dullien, T.; Vinyals, O.; Kohli, P. Graph Matching Networks for Learning the Similarity of Graph Structured Objects. In Proceedings of the 36th International Conference on Machine Learning, ICML 2019, Long Beach, CA, USA, 9–15 June 2019; Volume 97, pp. 3835–3845. [Google Scholar]
- Sousa, R.T.; Silva, S.; Pesquita, C. Supervised biomedical semantic similarity. bioRxiv 2021. [Google Scholar] [CrossRef]
- Al-Rfou, R.; Perozzi, B.; Zelle, D. DDGK: Learning Graph Representations for Deep Divergence Graph Kernels. In Proceedings of the World Wide Web Conference (WWW 2019), San Francisco, CA, USA, 13–17 May 2019; Liu, L., White, R.W., Mantrach, A., Silvestri, F., McAuley, J.J., Baeza-Yates, R., Zia, L., Eds.; ACM: Baltimore, MD, USA, 2019; pp. 37–48. [Google Scholar] [CrossRef] [Green Version]
- Cao, S.; Lu, W.; Xu, Q. GraRep: Learning Graph Representations with Global Structural Information. In Proceedings of the 24th ACM International on Conference on Information and Knowledge Management, Melbourne, Australia, 18–23 October 2015; pp. 891–900. [Google Scholar]
- Ou, M.; Cui, P.; Pei, J.; Zhang, Z.; Zhu, W. Asymmetric Transitivity Preserving Graph Embedding. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 1105–1114. [Google Scholar]
- Qiu, J.; Dong, Y.; Ma, H.; Li, J.; Wang, K.; Tang, J. Network Embedding as Matrix Factorization: Unifying DeepWalk, LINE, PTE, and node2vec. In Proceedings of the Eleventh ACM International Conference on Web Search and Data Mining, Los Angeles, CA, USA, 5–9 February 2018; pp. 459–467. [Google Scholar]
- Perozzi, B.; Al-Rfou, R.; Skiena, S. DeepWalk: Online learning of social representations. In Proceedings of the 20th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, New York, NY, USA, 24–27 August 2014; pp. 701–710. [Google Scholar]
- Makarov, I.; Kiselev, D.; Nikitinsky, N.; Subelj, L. Survey on graph embeddings and their applications to machine learning problems on graphs. PeerJ Comput. Sci. 2021, 7, e357. [Google Scholar] [CrossRef]
- Tang, J.; Qu, M.; Wang, M.; Zhang, M.; Yan, J.; Mei, Q. LINE: Large-scale Information Network Embedding. In Proceedings of the 24th International Conference on World Wide Web, Florence, Italy, 18–22 May 2015; pp. 1067–1077. [Google Scholar]
- Grover, A.; Leskovec, J. node2vec: Scalable Feature Learning for Networks. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 855–864. [Google Scholar]
- Dong, Y.; Chawla, N.V.; Swami, A. metapath2vec: Scalable Representation Learning for Heterogeneous Networks. In Proceedings of the 23rd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, Halifax, NS, Canada, 13–17 August 2017; pp. 135–144. [Google Scholar]
- Chen, H.; Perozzi, B.; Hu, Y.; Skiena, S. HARP: Hierarchical Representation Learning for Networks. In Proceedings of the AAAI Conference on Artificial Intelligence, New Orleans, LA, USA, 2–7 February 2018; pp. 2127–2134. [Google Scholar]
- Abu-El-Haija, S.; Perozzi, B.; Al-Rfou, R.; Alemi, A.A. Watch Your Step: Learning Node Embeddings via Graph Attention. In Proceedings of the 2018 Conference on Neural Information Processing Systems (NeurIPS), Montreal, QC, Canada, 3–8 December 2018; pp. 9198–9208. [Google Scholar]
- Rozemberczki, B.; Davies, R.; Sarkar, R.; Sutton, C. GEMSEC: Graph embedding with self clustering. In Proceedings of the 2019 IEEE/ACM International Conference on Advances in Social Networks Analysis and Mining, Vancouver, BC, Canada, 27–30 August 2019; pp. 65–72. [Google Scholar]
- Yang, C.; Liu, Z.; Zhao, D.; Sun, M.; Chang, E.Y. Network Representation Learning with Rich Text Information. In Proceedings of the Twenty-Fourth International Joint Conference on Artificial Intelligence, Buenos Aires, Argentina, 25–31 July 2015; AAAI Press: Menlo Park, CA, USA, 2015; pp. 2111–2117. [Google Scholar]
- Ahn, S.; Kim, M.H. Variational Graph Normalized Auto-Encoders. In Proceedings of the 30th ACM International Conference on Information & Knowledge Management, Online, 1–5 November 2021. [Google Scholar]
- Kipf, T.N.; Welling, M. Semi-Supervised Classification with Graph Convolutional Networks. arXiv 2016, arXiv:1609.02907. [Google Scholar]
- Velickovic, P.; Cucurull, G.; Casanova, A.; Romero, A.; Liò, P.; Bengio, Y. Graph Attention Networks. arXiv 2017, arXiv:1710.10903. [Google Scholar]
- Hamilton, W.L.; Ying, Z.; Leskovec, J. Inductive Representation Learning on Large Graphs. In Proceedings of the Annual Conference on Neural Information Processing Systems (NIPS), Long Beach, CA, USA, 4–9 December 2017; pp. 1024–1034. [Google Scholar]
- Cai, H.; Zheng, V.W.; Chang, K.C. A Comprehensive Survey of Graph Embedding: Problems, Techniques, and Applications. IEEE Trans. Knowl. Data Eng. 2018, 30, 1616–1637. [Google Scholar] [CrossRef] [Green Version]
- Chami, I.; Abu-El-Haija, S.; Perozzi, B.; Ré, C.; Murphy, K. Machine Learning on Graphs: A Model and Comprehensive Taxonomy. arXiv 2020, arXiv:2005.03675. [Google Scholar]
- Nikolentzos, G.; Meladianos, P.; Vazirgiannis, M. Matching Node Embeddings for Graph Similarity. In Proceedings of the Thirty-First AAAI Conference on Artificial Intelligence, San Francisco, CA, USA, 4–9 February 2017; AAAI Press: Menlo Park, CA, USA, 2017; pp. 2429–2435. [Google Scholar]
- Rubner, Y.; Tomasi, C.; Guibas, L.J. The Earth Mover’s Distance as a Metric for Image Retrieval. Int. J. Comput. Vis. 2000, 40, 99–121. [Google Scholar] [CrossRef]
- Tixier, A.J.; Nikolentzos, G.; Meladianos, P.; Vazirgiannis, M. Graph Classification with 2D Convolutional Neural Networks. In Lecture Notes in Computer Science, Proceedings of the International Conference on Artificial Neural Networks ICANN, Munich, Germany, 17–19 September 2019; Springer: Berlin/Heidelberg, Germany, 2019; Volume 11731, pp. 578–593. [Google Scholar]
- Battaglia, P.W.; Hamrick, J.B.; Bapst, V.; Sanchez-Gonzalez, A.; Zambaldi, V.F.; Malinowski, M.; Tacchetti, A.; Raposo, D.; Santoro, A.; Faulkner, R.; et al. Relational inductive biases, deep learning, and graph networks. arXiv 2018, arXiv:1806.01261. [Google Scholar]
- Atamna, A.; Sokolovska, N.; Crivello, J.C. SPI-GCN: A simple permutation-invariant graph convolutional network. Available online: https://hal.archives-ouvertes.fr/hal-02093451/ (accessed on 11 December 2021).
- Zhang, J. Graph Neural Distance Metric Learning with Graph-Bert. arXiv 2020, arXiv:2002.03427. [Google Scholar]
- Ying, Z.; You, J.; Morris, C.; Ren, X.; Hamilton, W.L.; Leskovec, J. Hierarchical Graph Representation Learning with Differentiable Pooling. In Proceedings of the 2018 Conference on Neural Information Processing Systems (NeurIPS), Montreal, QC, Canada, 3–8 December 2018; pp. 4805–4815. [Google Scholar]
- Zhang, M.; Cui, Z.; Neumann, M.; Chen, Y. An End-to-End Deep Learning Architecture for Graph Classification. In Proceedings of the Thirty-Second AAAI Conference on Artificial Intelligence, New Orleans, LA, USA, 2–7 February 2018; AAAI Press: Menlo Park, CA, USA, 2018; pp. 4438–4445. [Google Scholar]
- Gao, H.; Ji, S. Graph U-Nets. In Proceedings of the International Conference on Machine Learning, Long Beach, CA, USA, 10–15 June 2019; Volume 97, pp. 2083–2092. [Google Scholar]
- Lee, J.; Lee, I.; Kang, J. Self-Attention Graph Pooling. In Proceedings of the International Conference on Machine Learning, Long Beach, CA, USA, 10–15 June 2019; Volume 97, pp. 3734–3743. [Google Scholar]
- Ahmadi, A.H.K.; Hassani, K.; Moradi, P.; Lee, L.; Morris, Q. Memory-Based Graph Networks. In Proceedings of the International Conference on Learning Representations (ICLR), Addis Ababa, Ethiopia, 26–30 April 2020. [Google Scholar]
- Niepert, M.; Ahmed, M.; Kutzkov, K. Learning Convolutional Neural Networks for Graphs. In Proceedings of the International Conference on Machine Learning, New York, NY, USA, 19–24 June 2016; Volume 48, pp. 2014–2023. [Google Scholar]
- Adhikari, B.; Zhang, Y.; Ramakrishnan, N.; Prakash, B.A. Distributed Representations of Subgraphs. In Proceedings of the 2017 IEEE International Conference on Data Mining Workshops (ICDMW), New Orleans, LA, USA, 18–21 November 2017; pp. 111–117. [Google Scholar]
- Narayanan, A.; Chandramohan, M.; Venkatesan, R.; Chen, L.; Liu, Y.; Jaiswal, S. graph2vec: Learning Distributed Representations of Graphs. arXiv 2017, arXiv:1707.05005. [Google Scholar]
- Liu, S.; Demirel, M.F.; Liang, Y. N-Gram Graph: Simple Unsupervised Representation for Graphs, with Applications to Molecules. In Proceedings of the 2019 Conference on Neural Information Processing Systems (NeurIPS), Vancouver, BC, Canada, 8–14 December 2019; pp. 8464–8476. [Google Scholar]
- Bai, Y.; Ding, H.; Bian, S.; Chen, T.; Sun, Y.; Wang, W. SimGNN: A Neural Network Approach to Fast Graph Similarity Computation. In Proceedings of the Twelfth ACM International Conference on Web Search and Data Mining, Melbourne, Australia, 11–15 February 2019; pp. 384–392. [Google Scholar]
- Domeniconi, G.; Moro, G.; Pasolini, R.; Sartori, C. A Comparison of Term Weighting Schemes for Text Classification and Sentiment Analysis with a Supervised Variant of tf.idf. In Proceedings of the International Conference on Data Management Technologies and Applications, Colmar, France, 20–22 July 2015; pp. 39–58. [Google Scholar] [CrossRef]
- Papadimitriou, P.; Dasdan, A.; Garcia-Molina, H. Web graph similarity for anomaly detection. J. Internet Serv. Appl. 2010, 1, 19–30. [Google Scholar] [CrossRef] [Green Version]
- Faloutsos, C.; Koutra, D.; Vogelstein, J.T. DELTACON: A Principled Massive-Graph Similarity Function. In Proceedings of the 2013 SIAM International Conference on Data Mining, Austin, TX, USA, 2–4 May 2013; pp. 162–170. [Google Scholar]
- Ji, S.; Pan, S.; Cambria, E.; Marttinen, P.; Yu, P.S. A Survey on Knowledge Graphs: Representation, Acquisition and Applications. IEEE Trans. Neural Netw. Learn. Syst. 2021, 1–21. [Google Scholar] [CrossRef]
- Cai, L.; Wang, W.Y. KBGAN: Adversarial Learning for Knowledge Graph Embeddings. In Proceedings of the 2018 Conference of the North American Chapter of the Association for Computational Linguistics, NAACL-HLT 2018, New Orleans, LA, USA, 2–4 June 2018; pp. 1470–1480. [Google Scholar]
- Bordes, A.; Usunier, N.; García-Durán, A.; Weston, J.; Yakhnenko, O. Translating Embeddings for Modeling Multi-relational Data. In Proceedings of the 26th International Conference on Neural Information Processing Systems (NIPS), Lake Tahoe, NA, USA, 5–10 December 2013; pp. 2787–2795. [Google Scholar]
- Yang, B.; Yih, W.; He, X.; Gao, J.; Deng, L. Embedding Entities and Relations for Learning and Inference in Knowledge Bases. In 3rd International Conference on Learning Representations (ICLR), San Diego, CA, USA, 7–9 May 2015. [Google Scholar]
- Nickel, M.; Rosasco, L.; Poggio, T.A. Holographic Embeddings of Knowledge Graphs. In Proceedings of the Thirtieth AAAI Conference on Artificial Intelligence (AAAI), Phoenix, AZ, USA, 12–17 February 2016; AAAI Press: Palo Alto, CA, USA, 2016; pp. 1955–1961. [Google Scholar]
- Trouillon, T.; Welbl, J.; Riedel, S.; Gaussier, É.; Bouchard, G. Complex Embeddings for Simple Link Prediction. In Proceedings of the International Conference on Machine Learning, New York, NY, USA, 19–24 June 2016; Volume 48, pp. 2071–2080. [Google Scholar]
- Socher, R.; Chen, D.; Manning, C.D.; Ng, A.Y. Reasoning With Neural Tensor Networks for Knowledge Base Completion. In Proceedings of the Annual Conference on Neural Information Processing Systems (NIPS), Lake Tahoe, NA, USA, 5–10 December 2013; pp. 926–934. [Google Scholar]
- Ristoski, P.; Paulheim, H. RDF2Vec: RDF Graph Embeddings for Data Mining. In Proceedings of the International Semantic Web Conference (ISWC), Kobe, Japan, 17–21 October 2016; Volume 9981, pp. 498–514. [Google Scholar]
- Dettmers, T.; Minervini, P.; Stenetorp, P.; Riedel, S. Convolutional 2D Knowledge Graph Embeddings. In Proceedings of the AAAI Conference on Artificial Intelligence (AAAI), New Orleans, LA, USA, 2–7 February 2018; AAAI Press: Palo Alto, CA, USA, 2018; pp. 1811–1818. [Google Scholar]
- Schlichtkrull, M.S.; Kipf, T.N.; Bloem, P.; van den Berg, R.; Titov, I.; Welling, M. Modeling Relational Data with Graph Convolutional Networks. In Lecture Notes in Computer Science, Proceedings of the 2018 Extended Semantic Web Conference (ESWC), Crete, Greece, 3–7 June 2018; Springer: Heidelberg, Germany, 2018; Volume 10843, pp. 593–607. [Google Scholar]
- Xie, R.; Liu, Z.; Jia, J.; Luan, H.; Sun, M. Representation Learning of Knowledge Graphs with Entity Descriptions. In Proceedings of the AAAI Conference on Artificial Intelligence (AAAI), Phoenix, AZ, USA, 12–17 February 2016; AAAI Press: Palo Alto, CA, USA, 2016; pp. 2659–2665. [Google Scholar]
- Wang, Q.; Huang, P.; Wang, H.; Dai, S.; Jiang, W.; Liu, J.; Lyu, Y.; Zhu, Y.; Wu, H. CoKE: Contextualized Knowledge Graph Embedding. arXiv 2019, arXiv:1911.02168. [Google Scholar]
- Hu, L.; Zhang, M.; Li, S.; Shi, J.; Shi, C.; Yang, C.; Liu, Z. Text-Graph Enhanced Knowledge Graph Representation Learning. Front. Artif. Intell. 2021, 4, 697856. [Google Scholar] [CrossRef]
- Grohe, M. word2vec, node2vec, graph2vec, X2vec: Towards a Theory of Vector Embeddings of Structured Data. In Proceedings of the 39th ACM SIGMOD-SIGACT-SIGAI Symposium on Principles of Database Systems, Portland, OR, USA, 14–19 June 2020; pp. 1–16. [Google Scholar]
- Wu, L.; Yen, I.E.; Xu, F.; Ravikumar, P.; Witbrock, M. D2KE: From Distance to Kernel and Embedding. arXiv 2018, arXiv:1802.04956. [Google Scholar]
- Bunke, H.; Allermann, G. Inexact graph matching for structural pattern recognition. Pattern Recognit. Lett. 1983, 1, 245–253. [Google Scholar] [CrossRef]
- Bunke, H.; Shearer, K. A graph distance metric based on the maximal common subgraph. Pattern Recognit. Lett. 1998, 19, 255–259. [Google Scholar] [CrossRef]
- Liang, Y.; Zhao, P. Similarity Search in Graph Databases: A Multi-Layered Indexing Approach. In Proceedings of the 2017 IEEE 33rd International Conference on Data Engineering (ICDE), San Diego, CA, USA, 19–22 April 2017; pp. 783–794. [Google Scholar]
- Daller, É.; Bougleux, S.; Gaüzère, B.; Brun, L. Approximate Graph Edit Distance by Several Local Searches in Parallel. In Proceedings of the 7th International Conference on Pattern Recognition Applications and Methods, Funchal, Portugal, 16–18 January 2018; pp. 149–158. [Google Scholar]
- Bai, Y.; Ding, H.; Sun, Y.; Wang, W. Convolutional Set Matching for Graph Similarity. arXiv 2018, arXiv:1810.10866. [Google Scholar]
- Ktena, S.I.; Parisot, S.; Ferrante, E.; Rajchl, M.; Lee, M.C.H.; Glocker, B.; Rueckert, D. Metric learning with spectral graph convolutions on brain connectivity networks. NeuroImage 2018, 169, 431–442. [Google Scholar] [CrossRef]
- Ma, G.; Ahmed, N.K.; Willke, T.L.; Sengupta, D.; Cole, M.W.; Turk-Browne, N.B.; Yu, P.S. Deep Graph Similarity Learning for Brain Data Analysis. In Proceedings of the 28th ACM International Conference on Information and Knowledge Management, Beijing, China, 3–7 November 2019; pp. 2743–2751. [Google Scholar]
- Wang, S.; Chen, Z.; Yu, X.; Li, D.; Ni, J.; Tang, L.; Gui, J.; Li, Z.; Chen, H.; Yu, P.S. Heterogeneous Graph Matching Networks for Unknown Malware Detection. In Proceedings of the 2019 International Joint Conference on Artificial Intelligence (IJCAI), Macao, China, 10–16 August 2019; pp. 3762–3770. [Google Scholar]
- Bai, Y.; Ding, H.; Qiao, Y.; Marinovic, A.; Gu, K.; Chen, T.; Sun, Y.; Wang, W. Unsupervised inductive graph-level representation learning via graph-graph proximity. arXiv 2019, arXiv:1904.01098. [Google Scholar]
- Borgwardt, K.M.; Kriegel, H. Shortest-Path Kernels on Graphs. In Proceedings of the 5th IEEE International Conference on Data Mining (ICDM 2005), Houston, TX, USA, 27–30 November 2005; pp. 74–81. [Google Scholar]
- Shervashidze, N.; Vishwanathan, S.V.N.; Petri, T.; Mehlhorn, K.; Borgwardt, K.M. Efficient graphlet kernels for large graph comparison. In Proceedings of the 12th International Conference on Artificial Intelligence and Statistics (AISTATS), Clearwater Beach, FL, USA, 16–18 April 2009; Volume 5, pp. 488–495. [Google Scholar]
- Shervashidze, N.; Schweitzer, P.; van Leeuwen, E.J.; Mehlhorn, K.; Borgwardt, K.M. Weisfeiler-Lehman Graph Kernels. J. Mach. Learn. Res. 2011, 12, 2539–2561. [Google Scholar]
- Kondor, R.; Pan, H. The Multiscale Laplacian Graph Kernel. In Proceedings of the 2016 Conference on Neural Information Processing Systems (NIPS), Barcelona, Spain, 5–10 December 2016; pp. 2982–2990. [Google Scholar]
- Yanardag, P.; Vishwanathan, S.V.N. Deep Graph Kernels. In Proceedings of the 21th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, Sydney, Australi, 10–13 August 2015; pp. 1365–1374. [Google Scholar]
- Ding, X.; Zhang, Y.; Liu, T.; Duan, J. Deep Learning for Event-Driven Stock Prediction. In Proceedings of the Twenty-Fourth International Joint Conference on Artificial Intelligence, Buenos Aires, Argentina, 25–31 July 2015; AAAI Press: Palo Alto, CA, USA, 2015; pp. 2327–2333. [Google Scholar]
- Ding, X.; Zhang, Y.; Liu, T.; Duan, J. Knowledge-Driven Event Embedding for Stock Prediction. In Proceedings of the coling 2016, the 26th International Conference on Computational Linguistics: Technical Papers, Osaka, Japan, 11–16 December 2016; pp. 2133–2142. [Google Scholar]
- Weber, N.; Balasubramanian, N.; Chambers, N. Event Representations with Tensor-Based Compositions. In Proceedings of the Thirty-Second AAAI Conference on Artificial Intelligence (AAAI), New Orleans, LA, USA, 2–7 February 2018; AAAI Press: Palo Alto, CA, USA, 2018; pp. 4946–4953. [Google Scholar]
- Ding, X.; Liao, K.; Liu, T.; Li, Z.; Duan, J. Event Representation Learning Enhanced with External Commonsense Knowledge. In Proceedings of the 9th International Joint Conference on Natural Language Processing, EMNLP-IJCNLP 2019, Hong Kong, China, 3–7 November 2019; pp. 4893–4902. [Google Scholar]
- Kavumba, P.; Inoue, N.; Inui, K. Exploring Supervised Learning of Hierarchical Event Embedding with Poincaré Embeddings. In Proceedings of the 25th Annual Meeting of the Association for Natural Language Processing (ANLP), Kyoto, Japan, 12–25 March 2019; pp. 217–220. [Google Scholar]
- Trieu, H.; Tran, T.T.; Nguyen, A.D.; Nguyen, A.; Miwa, M.; Ananiadou, S. DeepEventMine: End-to-end neural nested event extraction from biomedical texts. Bioinformatics 2020, 36, 4910–4917. [Google Scholar] [CrossRef] [PubMed]
- Gui, H.; Liu, J.; Tao, F.; Jiang, M.; Norick, B.; Kaplan, L.M.; Han, J. Embedding Learning with Events in Heterogeneous Information Networks. IEEE Trans. Knowl. Data Eng. 2017, 29, 2428–2441. [Google Scholar] [CrossRef]
- Kriege, N.M.; Johansson, F.D.; Morris, C. A survey on graph kernels. Appl. Netw. Sci. 2020, 5, 6. [Google Scholar] [CrossRef] [Green Version]
- Kim, J.; Nguyen, N.L.T.; Wang, Y.; Tsujii, J.; Takagi, T.; Yonezawa, A. The Genia Event and Protein Coreference tasks of the BioNLP Shared Task 2011. BMC Bioinf. 2012, 13, S1. [Google Scholar] [CrossRef] [Green Version]
- Ohta, T.; Pyysalo, S.; Tsujii, J. Overview of the Epigenetics and Post-translational Modifications (EPI) task of BioNLP Shared Task 2011. In Proceedings of the BioNLP Shared Task 2011 Workshop, Portland, OR, USA, 24 June 2011; Tsujii, J., Kim, J., Pyysalo, S., Eds.; Association for Computational Linguistics: Stroudsburg, PA, USA, 2011; pp. 16–25. [Google Scholar]
- Pyysalo, S.; Ohta, T.; Rak, R.; Sullivan, D.E.; Mao, C.; Wang, C.; Sobral, B.W.S.; Tsujii, J.; Ananiadou, S. Overview of the Infectious Diseases (ID) task of BioNLP Shared Task 2011. In Proceedings of the BioNLP Shared Task 2011 Workshop, Portland, OR, USA, 24 June 2011; Tsujii, J., Kim, J., Pyysalo, S., Eds.; Association for Computational Linguistics: Stroudsburg, PA, USA, 2011; pp. 26–35. [Google Scholar]
- Kim, J.; Wang, Y.; Yamamoto, Y. The Genia Event Extraction Shared Task, 2013 Edition—Overview. In Proceedings of the BioNLP Shared Task 2013 Workshop, Sofia, Bulgaria, 9 August 2013; Nédellec, C., Bossy, R., Kim, J., Kim, J., Ohta, T., Pyysalo, S., Zweigenbaum, P., Eds.; Association for Computational Linguistics: Stroudsburg, PA, USA, 2013; pp. 8–15. [Google Scholar]
- Pyysalo, S.; Ohta, T.; Ananiadou, S. Overview of the Cancer Genetics (CG) task of BioNLP Shared Task 2013. In Proceedings of the BioNLP Shared Task 2013 Workshop, Sofia, Bulgaria, 9 August 2013; Nédellec, C., Bossy, R., Kim, J., Kim, J., Ohta, T., Pyysalo, S., Zweigenbaum, P., Eds.; Association for Computational Linguistics: Stroudsburg, PA, USA, 2013; pp. 58–66. [Google Scholar]
- Ohta, T.; Pyysalo, S.; Rak, R.; Rowley, A.; Chun, H.; Jung, S.; Choi, S.; Ananiadou, S.; Tsujii, J. Overview of the Pathway Curation (PC) task of BioNLP Shared Task 2013. In Proceedings of the BioNLP Shared Task 2013 Workshop, Sofia, Bulgaria, 9 August 2013; Nédellec, C., Bossy, R., Kim, J., Kim, J., Ohta, T., Pyysalo, S., Zweigenbaum, P., Eds.; Association for Computational Linguistics: Stroudsburg, PA, USA, 2013; pp. 67–75. [Google Scholar]
- Kim, J.; Han, X.; Lee, V.; Rebholz-Schuhmann, D. GRO Task: Populating the Gene Regulation Ontology with events and relations. In Proceedings of the BioNLP Shared Task 2013 Workshop, Sofia, Bulgaria, 9 August 2013; Nédellec, C., Bossy, R., Kim, J., Kim, J., Ohta, T., Pyysalo, S., Zweigenbaum, P., Eds.; Association for Computational Linguistics: Stroudsburg, PA, USA, 2013; pp. 50–57. [Google Scholar]
- Dobson, P.D.; Doig, A.J. Distinguishing enzyme structures from non-enzymes without alignments. J. Mol. Biol. 2003, 330, 771–783. [Google Scholar] [CrossRef] [Green Version]
- Toivonen, H.; Srinivasan, A.; King, R.D.; Kramer, S.; Helma, C. Statistical Evaluation of the Predictive Toxicology Challenge 2000–2001. Bioinformatics 2003, 19, 1183–1193. [Google Scholar] [CrossRef] [Green Version]
- Borgwardt, K.M.; Ong, C.S.; Schönauer, S.; Vishwanathan, S.V.N.; Smola, A.J.; Kriegel, H. Protein function prediction via graph kernels. Bioinformatics 2005, 21, 47–56. [Google Scholar] [CrossRef] [Green Version]
- Debnath, A.K.; Lopez de Compadre, R.L.; Debnath, G.; Shusterman, A.J.; Hansch, C. Structure-activity relationship of mutagenic aromatic and heteroaromatic nitro compounds. correlation with molecular orbital energies and hydrophobicity. J. Med. Chem. 1991, 34, 786–797. [Google Scholar] [CrossRef]
- Freitas, S.; Dong, Y.; Neil, J.; Chau, D.H. A Large-Scale Database for Graph Representation Learning. arXiv 2020, arXiv:2011.07682. [Google Scholar]
- Beltagy, I.; Lo, K.; Cohan, A. SciBERT: A Pretrained Language Model for Scientific Text. In Proceedings of the 9th International Joint Conference on Natural Language Processing, EMNLP-IJCNLP 2019, Hong Kong, China, 3–7 November 2019; Association for Computational Linguistics: Stroudsburg, PA, USA, 2019; pp. 3613–3618. [Google Scholar]
- Gionis, A.; Indyk, P.; Motwani, R. Similarity Search in High Dimensions via Hashing. In Proceedings of the 16th International Conference on Very Large Data Bases (VLDB), Queensland, Australia, 13–16 August 1999; pp. 518–529. [Google Scholar]
- Lee, J.; Yoon, W.; Kim, S.; Kim, D.; Kim, S.; So, C.H.; Kang, J. BioBERT: A pre-trained biomedical language representation model for biomedical text mining. Bioinformatics 2020, 36, 1234–1240. [Google Scholar] [CrossRef]
- Blei, D.M.; Ng, A.Y.; Jordan, M.I. Latent Dirichlet Allocation. In Proceedings of the Advances in Neural Information Processing Systems (NIPS), Vancouver, BC, Canada, 3–8 December 2001; MIT Press: Cambridge, MA, USA, 2001; pp. 601–608. [Google Scholar]
- Miwa, M.; Pyysalo, S.; Ohta, T.; Ananiadou, S. Wide coverage biomedical event extraction using multiple partially overlapping corpora. BMC Bioinform. 2013, 14, 175. [Google Scholar] [CrossRef] [Green Version]
- Schaeffer, S.E. Graph clustering. Comput. Sci. Rev. 2007, 1, 27–64. [Google Scholar] [CrossRef]
- Sgall, P.; Hajicová, E.; Hajicová, E.; Panevová, J.; Panevova, J. The Meaning of the Sentence in Its Semantic and Pragmatic Aspects; Springer Science & Business Media: New York, NY, USA, 1986. [Google Scholar]
- Sogancioglu, G.; Öztürk, H.; Özgür, A. BIOSSES: A semantic sentence similarity estimation system for the biomedical domain. Bioinformatics 2017, 33, i49–i58. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Afzal, N.; Fu, S.; Wang, L.; Shen, F.; Rastegar-Mojarad, M.; Liu, H. MedSTS: A resource for clinical semantic textual similarity. Lang. Resour. Eval. 2020, 54, 57–72. [Google Scholar] [CrossRef] [Green Version]
- Lithgow-Serrano, O.; Gama-Castro, S.; Ishida-Gutiérrez, C.; Mejía-Almonte, C.; Tierrafría, V.H.; Martínez-Luna, S.; Santos-Zavaleta, A.; Velázquez-Ramírez, D.A.; Collado-Vides, J. Similarity corpus on microbial transcriptional regulation. J. Biomed. Semant. 2019, 10, 8. [Google Scholar] [CrossRef] [Green Version]
- Reimers, N.; Gurevych, I. Sentence-BERT: Sentence Embeddings using Siamese BERT-Networks. In Proceedings of the 9th International Joint Conference on Natural Language Processing, EMNLP-IJCNLP 2019, Hong Kong, China, 3–7 November 2019; Association for Computational Linguistics: Stroudsburg, PA, USA, 2019; pp. 3980–3990. [Google Scholar]
- Domeniconi, G.; Moro, G.; Pagliarani, A.; Pasolini, R. On Deep Learning in Cross-Domain Sentiment Classification. In Proceedings of the 9th International Joint Conference on Knowledge Discovery, Knowledge Engineering and Knowledge Management IC3K, Funchal, Portugal, 1–3 November 2017; pp. 50–60. [Google Scholar] [CrossRef]
- Moro, G.; Pagliarani, A.; Pasolini, R.; Sartori, C. Cross-domain & In-domain Sentiment Analysis with Memory-based Deep Neural Networks. In Proceedings of the 10th International Joint Conference on Knowledge Discovery, Knowledge Engineering and Knowledge Management, Seville, Spain, 18–20 September 2018; pp. 127–138. [Google Scholar] [CrossRef]
- Lewis, P.S.H.; Perez, E.; Piktus, A.; Petroni, F.; Karpukhin, V.; Goyal, N.; Küttler, H.; Lewis, M.; Yih, W.; Rocktäschel, T.; et al. Retrieval-Augmented Generation for Knowledge-Intensive NLP Tasks. arXiv 2005, arXiv:2005.11401. [Google Scholar]
- Moro, G.; Valgimigli, L. Efficient Self-Supervised Metric Information Retrieval: A Bibliography Based Method Applied to COVID Literature. Sensors 2021, 21, 6430. [Google Scholar] [CrossRef]
- Li, S.; Jia, K.; Wen, Y.; Liu, T.; Tao, D. Orthogonal Deep Neural Networks. IEEE Trans. Pattern Anal. Mach. Intell. 2021, 43, 1352–1368. [Google Scholar] [CrossRef] [Green Version]
- Frisoni, G.; Moro, G.; Carbonaro, A. Learning Interpretable and Statistically Significant Knowledge from Unlabeled Corpora of Social Text Messages: A Novel Methodology of Descriptive Text Mining. In Proceedings of the 9th International Conference on Data Science, Technologies and Applications (DATA), Online, 7–9 July 2020; pp. 121–132. Available online: https://www.scitepress.org/Papers/2020/98920/98920.pdf (accessed on 11 December 2021).
- Frisoni, G.; Moro, G. Phenomena Explanation from Text: Unsupervised Learning of Interpretable and Statistically Significant Knowledge. In Proceedings of the International Conference on Data Management Technologies and Applications, Prague, Czech Republic, 26–28 July 2020; Springer: Berlin/Heidelberg, Germany, 2020; Volume 1446, pp. 293–318. [Google Scholar] [CrossRef]
- Frisoni, G.; Moro, G.; Carbonaro, A. Unsupervised Descriptive Text Mining for Knowledge Graph Learning. In Proceedings of the 12th International Conference on Knowledge Discovery and Information Retrieval KDIR, Budapest, Hungary, 2–4 November 2020; Volume 1, pp. 316–324. [Google Scholar]
- Frisoni, G.; Moro, G.; Carbonaro, A. Towards Rare Disease Knowledge Graph Learning from Social Posts of Patients. In Proceedings of the International Research & Innovation Forum, Athens, Greece, 15–17 April 2020; Springer: Berlin/Heidelberg, Germany; pp. 577–589. [Google Scholar] [CrossRef]
- Domeniconi, G.; Semertzidis, K.; López, V.; Daly, E.M.; Kotoulas, S.; Moro, G. A Novel Method for Unsupervised and Supervised Conversational Message Thread Detection. In Proceedings of the 5th International Conference on Data Management Technologies and Applications, Lisbon, Portugal, 24–26 July 2016; pp. 43–54. [Google Scholar] [CrossRef]
- Domeniconi, G.; Moro, G.; Pasolini, R.; Sartori, C. Iterative Refining of Category Profiles for Nearest Centroid Cross-Domain Text Classification. In Proceedings of the 6th International Joint Conference, Rome, Italy, 21–24 October 2014; pp. 50–67. [Google Scholar] [CrossRef]
- Moro, G.; Ragazzi, L. Semantic Self-segmentation for Abstractive Summarization of Long Legal Documents in Low-resource Regimes. In Proceedings of the Thirty-Six AAAI Conference on Artificial Intelligence, Vancouver, BC, Canada, 22 February–1 March 2022; pp. 1–9. [Google Scholar]
- Riccucci, S.; Carbonaro, A.; Casadei, G. Knowledge Acquisition in Intelligent Tutoring System: A Data Mining Approach. In Proceedings of the 6th Mexican International Conference on Artificial Intelligence, Aguascalientes, Mexico, 4–10 November 2007; pp. 1195–1205. [Google Scholar]
- Riccucci, S.; Carbonaro, A.; Casadei, G. An Architecture for Knowledge Management in Intelligent Tutoring System. In Proceedings of the Cognition and Exploratory Learning in Digital Age, CELDA 2005, Porto, Portugal, 14–16 December 2005; pp. 473–476. [Google Scholar]
- Andronico, A.; Carbonaro, A.; Colazzo, L.; Molinari, A. Personalisation services for learning management systems in mobile settings. Int. J. Contin. Eng. Educ. Life-Long Learn. 2004, 14, 353–369. [Google Scholar] [CrossRef]
- Domeniconi, G.; Moro, G.; Pagliarani, A.; Pasini, K.; Pasolini, R. Job Recommendation from Semantic Similarity of LinkedIn Users’ Skills. In Proceedings of the 5th International Conference on Pattern Recognition Applications and Methods, Rome, Italy, 24–26 February 2016; pp. 270–277. [Google Scholar] [CrossRef] [Green Version]




| Hyperparameter | Values |
|---|---|
| Node embedding | 2, 4, 8, 16, 32 |
| Encoder layers | 1, 2, 3, 4 |
| Learning rate | , , , , 1 |
| Encoding epochs | 100, 300, 600 |
| Scoring epochs | 100, 300, 600 |
| , , preserving | {7, 7, 7}, {8, 8, 4} |
| loss coefficients | {10, 6, 4}, {15, 0, 5} |
| Method | Unsupervised | Event Type Classification | Dataset Classification | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ST09 | GE11 | EPI11 | ID11 | MLEE | GE13 | CG13 | PC13 | GRO13 | BIO_ALL | AVG | ||||
| SciBERT | AVG | ✓ | 72.16 | 73.24 | 89.55 | 85.45 | 72.18 | 82.53 | 69.50 | 72.59 | 78.08 | 58.72 | 75.40 | 53.11 |
| SUM | ✓ | 56.32 | 58.91 | 94.34 | 76.97 | 58.35 | 81.02 | 52.68 | 53.51 | 65.35 | 42.63 | 64.01 | 47.68 | |
| MAX | ✓ | 64.54 | 53.95 | 90.47 | 75.95 | 56.16 | 74.35 | 63.91 | 60.39 | 72.16 | 51.11 | 66.30 | 49.22 | |
| BioBERT | AVG | ✓ | 71.43 | 70.11 | 90.60 | 85.40 | 71.91 | 79.81 | 66.77 | 70.40 | 76.72 | 54.59 | 73.77 | 52.61 |
| SUM | ✓ | 61.49 | 58.73 | 93.32 | 73.22 | 56.03 | 76.94 | 56.36 | 56.53 | 59.60 | 43.64 | 63.59 | 50.17 | |
| MAX | ✓ | 56.38 | 53.95 | 90.47 | 75.95 | 56.16 | 74.35 | 57.26 | 56.86 | 76.72 | 45.72 | 64.38 | 51.82 | |
| node2vec | AVG | ✓ | 19.65 | 21.03 | 23.81 | 14.18 | 18.58 | 19.24 | 7.51 | 12.60 | 9.91 | 11.38 | 15.79 | 14.05 |
| SUM | ✓ | 25.90 | 23.43 | 23.71 | 22.05 | 15.85 | 19.37 | 13.62 | 15.19 | 8.89 | 8.68 | 17.67 | 14.31 | |
| MAX | ✓ | 28.13 | 22.06 | 24.31 | 29.26 | 16.45 | 22.80 | 13.00 | 15.42 | 14.91 | 11.21 | 19.76 | 15.20 | |
| node2vec-PCA | ✓ | 26.71 | 24.17 | 32.04 | 31.06 | 22.53 | 34.43 | 17.90 | 24.51 | 23.16 | 16.54 | 25.31 | 18.26 | |
| graph2vec | ✓ | 54.12 | 58.60 | 57.78 | 62.25 | 41.34 | 63.47 | 44.59 | 40.51 | 33.81 | 40.48 | 49.70 | 43.06 | |
| DGCNN | 89.19 | 89.04 | 90.40 | 88.70 | 93.19 | 86.65 | 95.73 | 93.35 | 94.23 | 89.55 | 91.00 | 89.55 | ||
| DDEGK (ours) | ||||||||||||||
| w/random anchors | ✓ | 99.00 | 97.99 | 100 | 100 | 98.01 | 98.02 | 92.89 | 99.67 | 100 | 92.28 | 97.86 | 63.39 | |
| w/random anchors per type | ✓ | 99.01 | 97.99 | 100 | 100 | 98.02 | 97.70 | 90.32 | 99.67 | 100 | 90.87 | 97.36 | 62.79 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Frisoni, G.; Moro, G.; Carlassare, G.; Carbonaro, A. Unsupervised Event Graph Representation and Similarity Learning on Biomedical Literature. Sensors 2022, 22, 3. https://doi.org/10.3390/s22010003
Frisoni G, Moro G, Carlassare G, Carbonaro A. Unsupervised Event Graph Representation and Similarity Learning on Biomedical Literature. Sensors. 2022; 22(1):3. https://doi.org/10.3390/s22010003
Chicago/Turabian StyleFrisoni, Giacomo, Gianluca Moro, Giulio Carlassare, and Antonella Carbonaro. 2022. "Unsupervised Event Graph Representation and Similarity Learning on Biomedical Literature" Sensors 22, no. 1: 3. https://doi.org/10.3390/s22010003
APA StyleFrisoni, G., Moro, G., Carlassare, G., & Carbonaro, A. (2022). Unsupervised Event Graph Representation and Similarity Learning on Biomedical Literature. Sensors, 22(1), 3. https://doi.org/10.3390/s22010003









