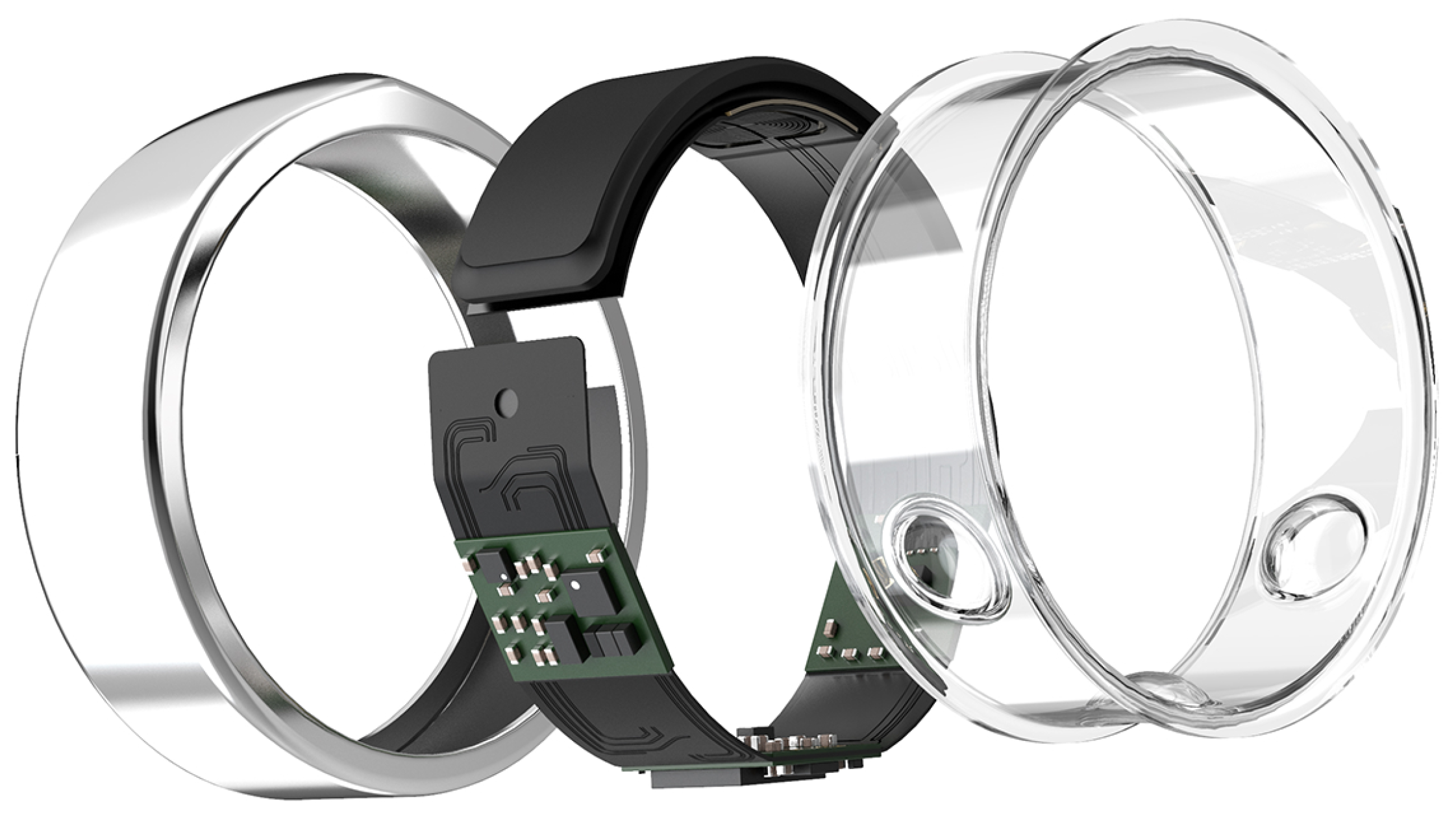
The Promise of Sleep: A Multi-Sensor Approach for Accurate Sleep Stage Detection Using the Oura Ring
Abstract
:1. Introduction
2. Materials and Methods
2.1. Datasets and Data Acquisition Protocols
2.2. Participants
2.2.1. Dataset 1: Singapore
2.2.2. Dataset 2: Finland
2.2.3. Dataset 3: USA
2.3. Features Extraction
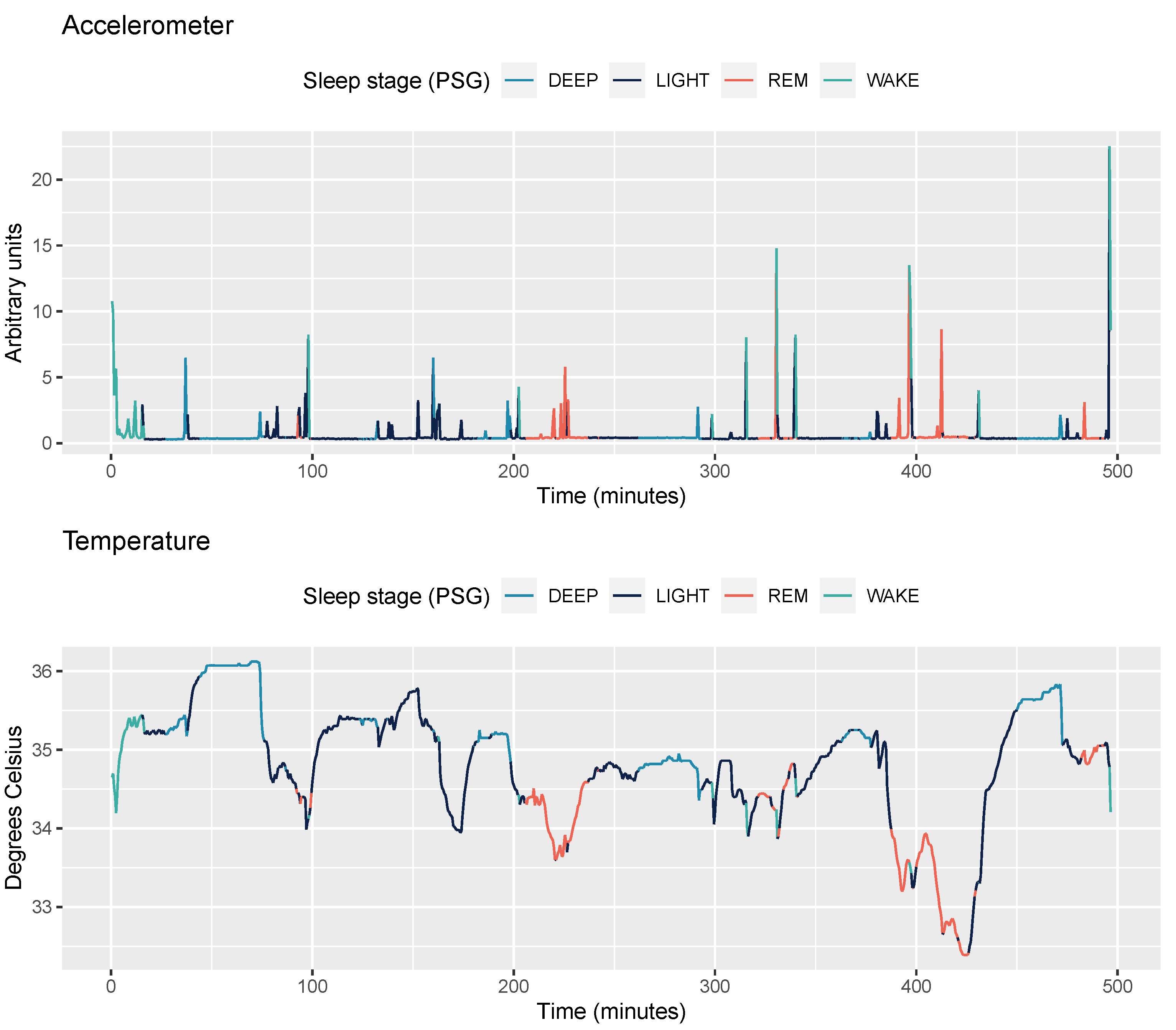
2.3.1. Accelerometer Data
2.3.2. Temperature Data
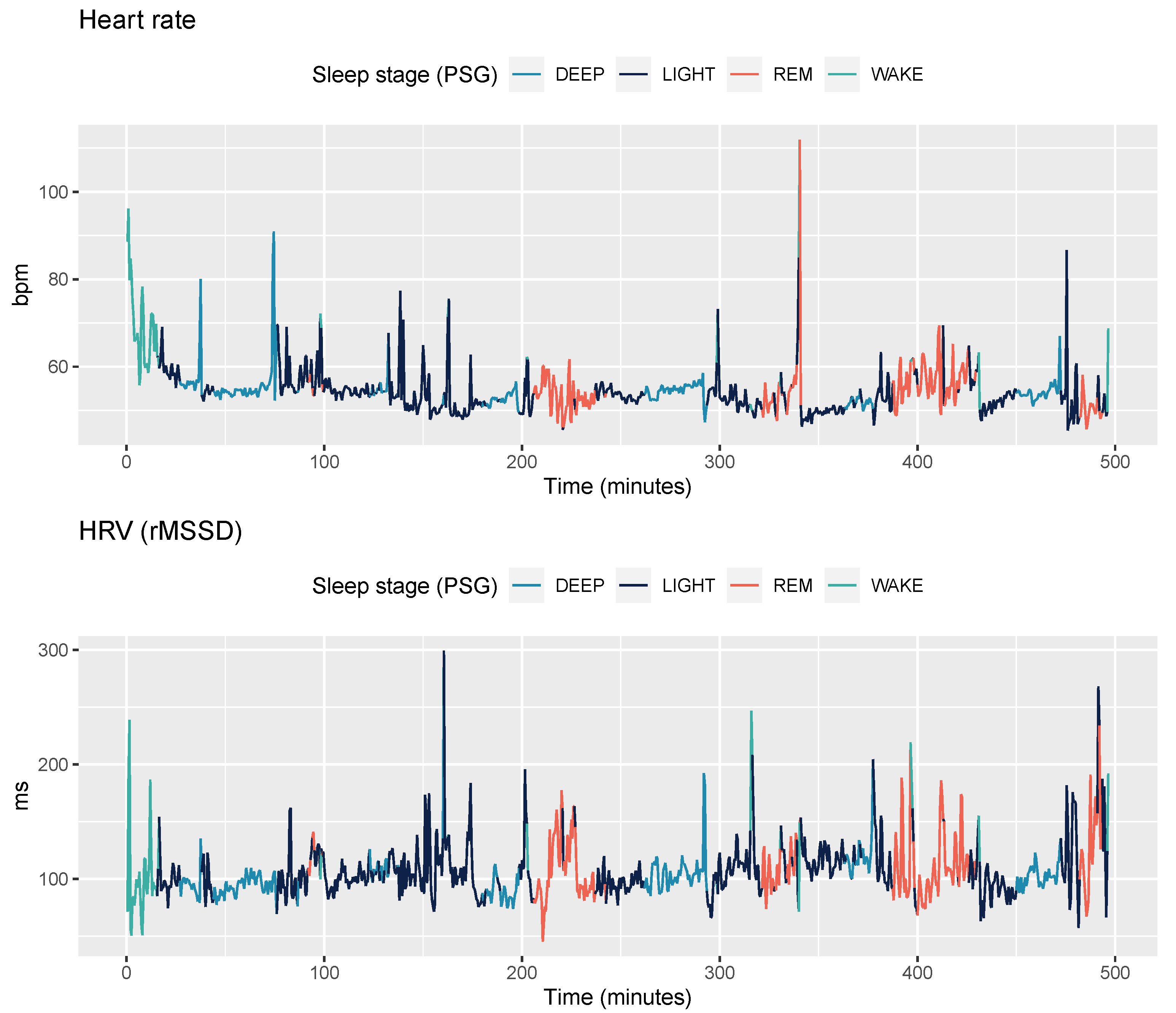
2.3.3. PPG data
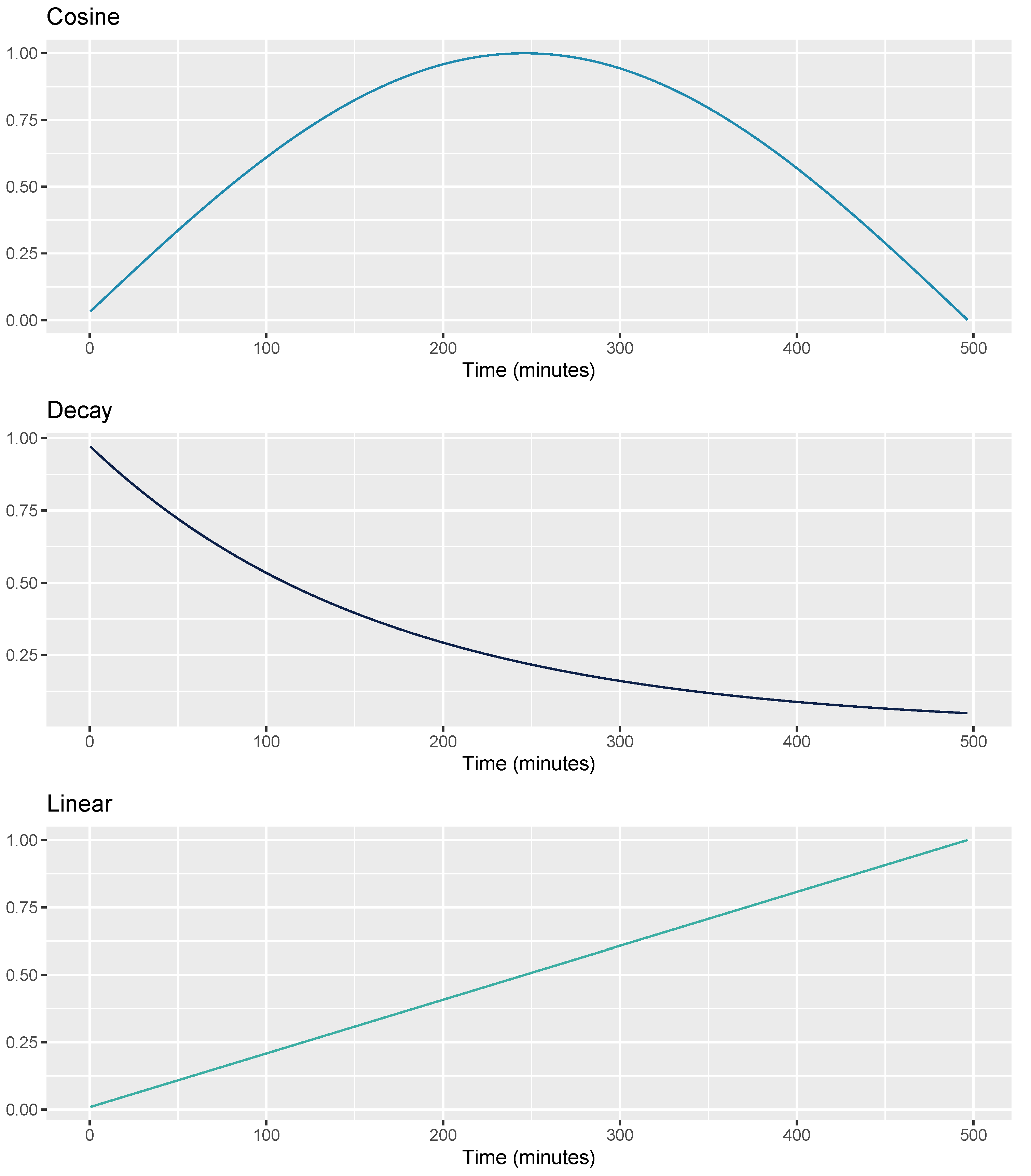
2.3.4. Sensor-Independent Circadian Features
2.4. Feature Normalization
2.5. Machine Learning
2.6. Validation Strategy and Evaluation Metrics
3. Results
3.1. 2-Stage Classification
3.2. 4-Stage Classification
4. Discussion
4.1. Physiological Considerations
4.2. Generalizability
4.3. Limitations
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Grandner, M.A. Sleep, health, and society. Sleep Med. Clin. 2017, 12, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Spiegel, K.; Tasali, E.; Leproult, R.; Van Cauter, E. Effects of poor and short sleep on glucose metabolism and obesity risk. Nat. Rev. Endocrinol. 2009, 5, 253. [Google Scholar] [CrossRef]
- Peppard, P.E.; Young, T.; Palta, M.; Dempsey, J.; Skatrud, J. Longitudinal study of moderate weight change and sleep-disordered breathing. JAMA 2000, 284, 3015–3021. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krause, A.J.; Simon, E.B.; Mander, B.A.; Greer, S.M.; Saletin, J.M.; Goldstein-Piekarski, A.N.; Walker, M.P. The sleep-deprived human brain. Nat. Rev. Neurosci. 2017, 18, 404. [Google Scholar] [CrossRef]
- Freeman, D.; Sheaves, B.; Goodwin, G.M.; Yu, L.M.; Nickless, A.; Harrison, P.J.; Emsley, R.; Luik, A.I.; Foster, R.G.; Wadekar, V.; et al. The effects of improving sleep on mental health (OASIS): A randomised controlled trial with mediation analysis. Lancet Psychiatry 2017, 4, 749–758. [Google Scholar] [CrossRef] [Green Version]
- De Zambotti, M.; Cellini, N.; Goldstone, A.; Colrain, I.M.; Baker, F.C. Wearable sleep technology in clinical and research settings. Med. Sci. Sport. Exerc. 2019, 51, 1538. [Google Scholar] [CrossRef]
- Depner, C.M.; Cheng, P.C.; Devine, J.K.; Khosla, S.; de Zambotti, M.; Robillard, R.; Vakulin, A.; Drummond, S.P. Wearable technologies for developing sleep and circadian biomarkers: A summary of workshop discussions. Sleep 2020, 43, zsz254. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shelgikar, A.V.; Anderson, P.F.; Stephens, M.R. Sleep tracking, wearable technology, and opportunities for research and clinical care. Chest 2016, 150, 732–743. [Google Scholar] [CrossRef] [PubMed]
- Berryhill, S.; Morton, C.J.; Dean, A.; Berryhill, A.; Provencio-Dean, N.; Patel, S.I.; Estep, L.; Combs, D.; Mashaqi, S.; Gerald, L.B.; et al. Effect of wearables on sleep in healthy individuals: A randomized crossover trial and validation study. J. Clin. Sleep Med. 2020, 16, 775–783. [Google Scholar] [CrossRef] [PubMed]
- Ameen, M.S.; Cheung, L.M.; Hauser, T.; Hahn, M.A.; Schabus, M. About the accuracy and problems of consumer devices in the assessment of sleep. Sensors 2019, 19, 4160. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- de Zambotti, M.; Rosas, L.; Colrain, I.M.; Baker, F.C. The sleep of the ring: Comparison of the ŌURA sleep tracker against polysomnography. Behav. Sleep Med. 2019, 17, 124–136. [Google Scholar] [CrossRef]
- Kuula, L.; Gradisar, M.; Martinmäki, K.; Richardson, C.; Bonnar, D.; Bartel, K.; Lang, C.; Leinonen, L.; Pesonen, A. Using big data to explore worldwide trends in objective sleep in the transition to adulthood. Sleep Med. 2019, 62, 69–76. [Google Scholar] [CrossRef]
- Khosla, S.; Deak, M.C.; Gault, D.; Goldstein, C.A.; Hwang, D.; Kwon, Y.; O’Hearn, D.; Schutte-Rodin, S.; Yurcheshen, M.; Rosen, I.M.; et al. Consumer sleep technology: An American Academy of Sleep Medicine position statement. J. Clin. Sleep Med. 2018, 14, 877–880. [Google Scholar] [CrossRef] [Green Version]
- Landolt, H.P.; Dijk, D.J.; Gaus, S.E.; Borbély, A.A. Caffeine reduces low-frequency delta activity in the human sleep EEG. Neuropsychopharmacology 1995, 12, 229–238. [Google Scholar] [CrossRef]
- Mourtazaev, M.; Kemp, B.; Zwinderman, A.; Kamphuisen, H. Age and gender affect different characteristics of slow waves in the sleep EEG. Sleep 1995, 18, 557–564. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rosenberg, R.S.; Van Hout, S. The American Academy of Sleep Medicine inter-scorer reliability program: Sleep stage scoring. J. Clin. Sleep Med. 2013, 9, 81–87. [Google Scholar] [CrossRef] [Green Version]
- Van De Water, A.T.; Holmes, A.; Hurley, D.A. Objective measurements of sleep for non-laboratory settings as alternatives to polysomnography—A systematic review. J. Sleep Res. 2011, 20, 183–200. [Google Scholar] [CrossRef] [PubMed]
- Scott, H.; Lack, L.; Lovato, N. A systematic review of the accuracy of sleep wearable devices for estimating sleep onset. Sleep Med. Rev. 2020, 49, 101227. [Google Scholar] [CrossRef]
- Sundararajan, K.; Georgievska, S.; Te Lindert, B.H.; Gehrman, P.R.; Ramautar, J.; Mazzotti, D.R.; Sabia, S.; Weedon, M.N.; van Someren, E.J.; Ridder, L.; et al. Sleep classification from wrist-worn accelerometer data using random forests. Sci. Rep. 2021, 11, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Fonseca, P.; Weysen, T.; Goelema, M.S.; Møst, E.I.; Radha, M.; Lunsingh Scheurleer, C.; van den Heuvel, L.; Aarts, R.M. Validation of photoplethysmography-based sleep staging compared with polysomnography in healthy middle-aged adults. Sleep 2017, 40, zsx097. [Google Scholar] [CrossRef]
- de Zambotti, M.; Goldstone, A.; Claudatos, S.; Colrain, I.M.; Baker, F.C. A validation study of Fitbit Charge 2™ compared with polysomnography in adults. Chronobiol. Int. 2018, 35, 465–476. [Google Scholar] [CrossRef] [PubMed]
- Beattie, Z.; Pantelopoulos, A.; Ghoreyshi, A.; Oyang, Y.; Statan, A.; Heneghan, C. 0068 estimation of sleep stages using cardiac and accelerometer data from a wrist-worn device. Sleep 2017, 40, A26. [Google Scholar] [CrossRef]
- Kortelainen, J.M.; Mendez, M.O.; Bianchi, A.M.; Matteucci, M.; Cerutti, S. Sleep staging based on signals acquired through bed sensor. IEEE Trans. Inf. Technol. Biomed. 2010, 14, 776–785. [Google Scholar] [CrossRef]
- Hedner, J.; White, D.P.; Malhotra, A.; Herscovici, S.; Pittman, S.D.; Zou, D.; Grote, L.; Pillar, G. Sleep staging based on autonomic signals: A multi-center validation study. J. Clin. Sleep Med. 2011. [Google Scholar] [CrossRef] [Green Version]
- Long, X. On the analysis and classification of sleep stages from cardiorespiratory activity. In Sleep Wake; Technische Universiteit Eindhoven: Eindhoven, The Netherlands, 2015; pp. 1–232. [Google Scholar]
- Walch, O.; Huang, Y.; Forger, D.; Goldstein, C. Sleep stage prediction with raw acceleration and photoplethysmography heart rate data derived from a consumer wearable device. Sleep 2019, 42, zsz180. [Google Scholar] [CrossRef] [PubMed]
- Danzig, R.; Wang, M.; Shah, A.; Trotti, L.M. The wrist is not the brain: Estimation of sleep by clinical and consumer wearable actigraphy devices is impacted by multiple patient-and device-specific factors. J. Sleep Res. 2020, 29, e12926. [Google Scholar] [CrossRef] [PubMed]
- Lee, I.; Park, N.; Lee, H.; Hwang, C.; Kim, J.H.; Park, S. Systematic Review on Human Skin-Compatible Wearable Photoplethysmography Sensors. Appl. Sci. 2021, 11, 2313. [Google Scholar] [CrossRef]
- Longmore, S.K.; Lui, G.Y.; Naik, G.; Breen, P.P.; Jalaludin, B.; Gargiulo, G.D. A comparison of reflective photoplethysmography for detection of heart rate, blood oxygen saturation, and respiration rate at various anatomical locations. Sensors 2019, 19, 1874. [Google Scholar] [CrossRef] [Green Version]
- Menghini, L.; Cellini, N.; Goldstone, A.; Baker, F.C.; de Zambotti, M. A standardized framework for testing the performance of sleep-tracking technology: Step-by-step guidelines and open-source code. Sleep 2021, 44, zsaa170. [Google Scholar] [CrossRef]
- Patanaik, A.; Ong, J.L.; Gooley, J.J.; Ancoli-Israel, S.; Chee, M.W. An end-to-end framework for real-time automatic sleep stage classification. Sleep 2018, 41, zsy041. [Google Scholar] [CrossRef]
- Chee, N.I.; Ghorbani, S.; Golkashani, H.A.; Leong, R.L.; Ong, J.L.; Chee, M.W. Multi-night validation of a sleep tracking ring in adolescents compared with a research actigraph and polysomnography. Nat. Sci. Sleep 2021, 13, 177. [Google Scholar] [CrossRef]
- te Lindert, B.H.; Van Someren, E.J. Sleep estimates using microelectromechanical systems (MEMS). Sleep 2013, 36, 781–789. [Google Scholar] [CrossRef] [Green Version]
- Vähä-Ypyä, H.; Vasankari, T.; Husu, P.; Mänttäri, A.; Vuorimaa, T.; Suni, J.; Sievänen, H. Validation of cut-points for evaluating the intensity of physical activity with accelerometry-based mean amplitude deviation (MAD). PLoS ONE 2015, 10, e0134813. [Google Scholar] [CrossRef] [Green Version]
- Van Hees, V.T.; Sabia, S.; Anderson, K.N.; Denton, S.J.; Oliver, J.; Catt, M.; Abell, J.G.; Kivimäki, M.; Trenell, M.I.; Singh-Manoux, A. A novel, open access method to assess sleep duration using a wrist-worn accelerometer. PLoS ONE 2015, 10, e0142533. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kinnunen, H.; Rantanen, A.; Kenttä, T.; Koskimäki, H. Feasible assessment of recovery and cardiovascular health: Accuracy of nocturnal HR and HRV assessed via ring PPG in comparison to medical grade ECG. Physiol. Meas. 2020, 41, 04NT01. [Google Scholar] [CrossRef]
- Xiao, M.; Yan, H.; Song, J.; Yang, Y.; Yang, X. Sleep stages classification based on heart rate variability and random forest. Biomed. Signal Process. Control 2013, 8, 624–633. [Google Scholar] [CrossRef]
- Borbély, A.A. A two process model of sleep regulation. Hum. Neurobiol. 1982, 1, 195–204. [Google Scholar] [PubMed]
- Borbély, A.A.; Daan, S.; Wirz-Justice, A.; Deboer, T. The two-process model of sleep regulation: A reappraisal. J. Sleep Res. 2016, 25, 131–143. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Mayer, C.; Cheng, P.; Siddula, A.; Burgess, H.J.; Drake, C.; Goldstein, C.; Walch, O.; Forger, D.B. Predicting circadian phase across populations: A comparison of mathematical models and wearable devices. Sleep 2019. [Google Scholar] [CrossRef]
- Ke, G.; Meng, Q.; Finley, T.; Wang, T.; Chen, W.; Ma, W.; Ye, Q.; Liu, T.Y. Lightgbm: A highly efficient gradient boosting decision tree. Adv. Neural Inf. Process. Syst. 2017, 30, 3146–3154. [Google Scholar]
- Vinayak, R.K.; Gilad-Bachrach, R. Dart: Dropouts meet multiple additive regression trees. Artif. Intell. Stat. 2015, 38, 489–497. [Google Scholar]
- Imtiaz, S.A. A Systematic Review of Sensing Technologies for Wearable Sleep Staging. Sensors 2021, 21, 1562. [Google Scholar] [CrossRef]
- Supratak, A.; Dong, H.; Wu, C.; Guo, Y. DeepSleepNet: A model for automatic sleep stage scoring based on raw single-channel EEG. IEEE Trans. Neural Syst. Rehabil. Eng. 2017, 25, 1998–2008. [Google Scholar] [CrossRef] [Green Version]
- Haghayegh, S.; Khoshnevis, S.; Smolensky, M.H.; Diller, K.R.; Castriotta, R.J. Accuracy of wristband Fitbit models in assessing sleep: Systematic review and meta-analysis. J. Med. Internet Res. 2019, 21, e16273. [Google Scholar] [CrossRef] [PubMed]
- Stone, J.D.; Rentz, L.E.; Forsey, J.; Ramadan, J.; Markwald, R.R.; Finomore, V.S. Evaluations of Commercial Sleep Technologies for Objective Monitoring During Routine Sleeping Conditions. Nat. Sci. Sleep 2020, 12, 821. [Google Scholar] [CrossRef] [PubMed]
- Gradisar, M.; Lack, L. Relationships between the circadian rhythms of finger temperature, core temperature, sleep latency, and subjective sleepiness. J. Biol. Rhythm. 2004, 19, 157–163. [Google Scholar] [CrossRef] [PubMed]
- Harding, E.C.; Franks, N.P.; Wisden, W. The temperature dependence of sleep. Front. Neurosci. 2019, 13, 336. [Google Scholar] [CrossRef]
- Campbell, S.S.; Broughton, R.J. Rapid decline in body temperature before sleep: Fluffing the physiological pillow? Chronobiol. Int. 1994, 11, 126–131. [Google Scholar] [CrossRef]
- Harding, E.C.; Franks, N.P.; Wisden, W. Sleep and thermoregulation. Curr. Opin. Physiol. 2020, 15, 7–13. [Google Scholar] [CrossRef]
- Altevogt, B.M.; Colten, H.R. Sleep Disorders and Sleep Deprivation: An Unmet Public Health Problem; The National Academies Press, Institute of Medicine: Washington, DC, USA, 2006. [Google Scholar]
- Stein, P.K.; Pu, Y. Heart rate variability, sleep and sleep disorders. Sleep Med. Rev. 2012, 16, 47–66. [Google Scholar] [CrossRef]
- Carskadon, M.A.; Dement, W.C. Normal human sleep: An overview. Princ. Pract. Sleep Med. 2005, 4, 13–23. [Google Scholar]
- Buysse, D.J. Sleep health: Can we define it? Does it matter? Sleep 2014, 37, 9–17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tilley, A.J.; Wilkinson, R.T. The effects of a restricted sleep regime on the composition of sleep and on performance. Psychophysiology 1984, 21, 406–412. [Google Scholar] [CrossRef] [PubMed]
- Stone, J.D.; Ulman, H.K.; Tran, K.; Thompson, A.G.; Halter, M.D.; Ramadan, J.H.; Stephenson, M.; Finomore, V.S., Jr.; Galster, S.M.; Rezai, A.R.; et al. Assessing the Accuracy of Popular Commercial Technologies That Measure Resting Heart Rate and Heart Rate Variability. Front. Sport. Act. Living 2021, 3, 37. [Google Scholar] [CrossRef]
- Parak, J.; Tarniceriu, A.; Renevey, P.; Bertschi, M.; Delgado-Gonzalo, R.; Korhonen, I. Evaluation of the beat-to-beat detection accuracy of PulseOn wearable optical heart rate monitor. In Proceedings of the 2015 37th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Milan, Italy, 25–29 August 2015; pp. 8099–8102. [Google Scholar]
- Koskimäki, H.; Kinnunen, H.; Rönkä, S.; Smarr, B. Following the heart: What does variation of resting heart rate tell about us as individuals and as a population. In Adjunct Proceedings of the 2019 ACM International Joint Conference on Pervasive and Ubiquitous Computing and Proceedings of the 2019 ACM International Symposium on Wearable Computers; Association for Computing Machinery: New York, NY, USA, 2019; pp. 1178–1181. [Google Scholar]
- Phan, H.; Andreotti, F.; Cooray, N.; Chén, O.Y.; De Vos, M. SeqSleepNet: End-to-end hierarchical recurrent neural network for sequence-to-sequence automatic sleep staging. IEEE Trans. Neural Syst. Rehabil. Eng. 2019, 27, 400–410. [Google Scholar] [CrossRef] [Green Version]
Short Biography of Authors
 | Marco Altini, born in 1984 in Ravenna, Italy, received his Master’s degree in Computer Science Engineering from the University of Bologna, Doctor of Philosophy in Electrical Engineering from Eindhoven University of Technology and Master’s degree in Human Movement Sciences and High Performance Coaching from Vrije Universiteit Amsterdam. He is the founder of HRV4Training and has developed and validated several tools to acquire and analyze physiological data using mobile phones. He is an advisor at Oura and guest lecturer in Data in Sport and Health at Vrije Universiteit Amsterdam. Marco has published more than 50 peer-reviewed papers at the intersection between physiology, technology, health and performance. |
 | Hannu Kinnunen, born in 1972 in Pattijoki, Finland, received his Master degree in Biophysics and Doctor of Science (Technology) degree in Electrical Engineering from University of Oulu, Oulu, Finland in 1997 and 2020, respectively. He has worked in the industry with pioneering wearable companies Polar Electro (1996–2014) and Oura Health (2014–2021) in various specialist and leadership roles. In addition to algorithms embedded in consumer products and research tools, his work has resulted in numerous scientific publications and patents. His past research interests included feasible methodology for metabolic estimations utilizing accelerometers and heart rate sensors, and lately his work has concentrated on wearable multi-sensor approach for the quantification of human health behavior, sleep, and recovery. |

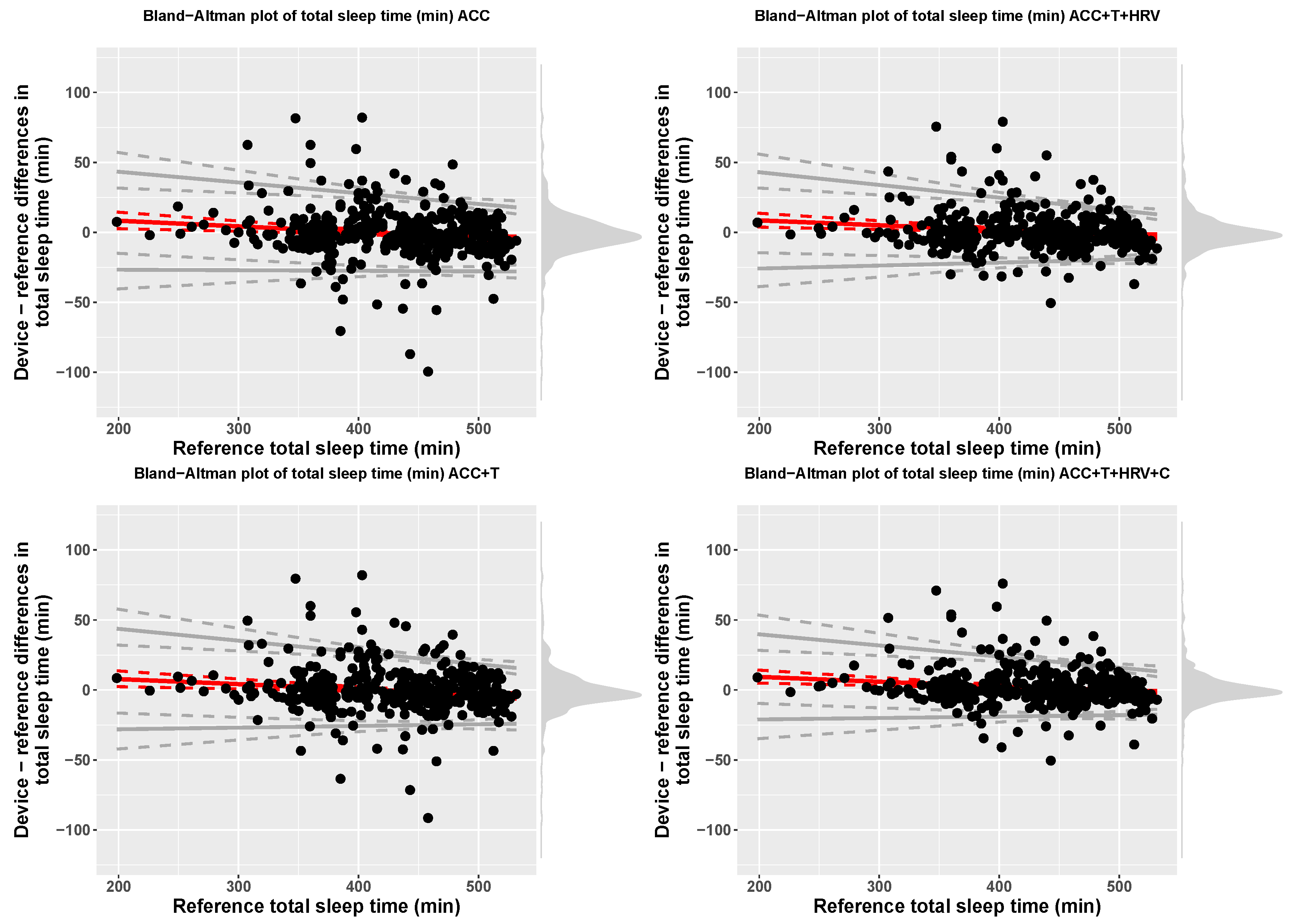
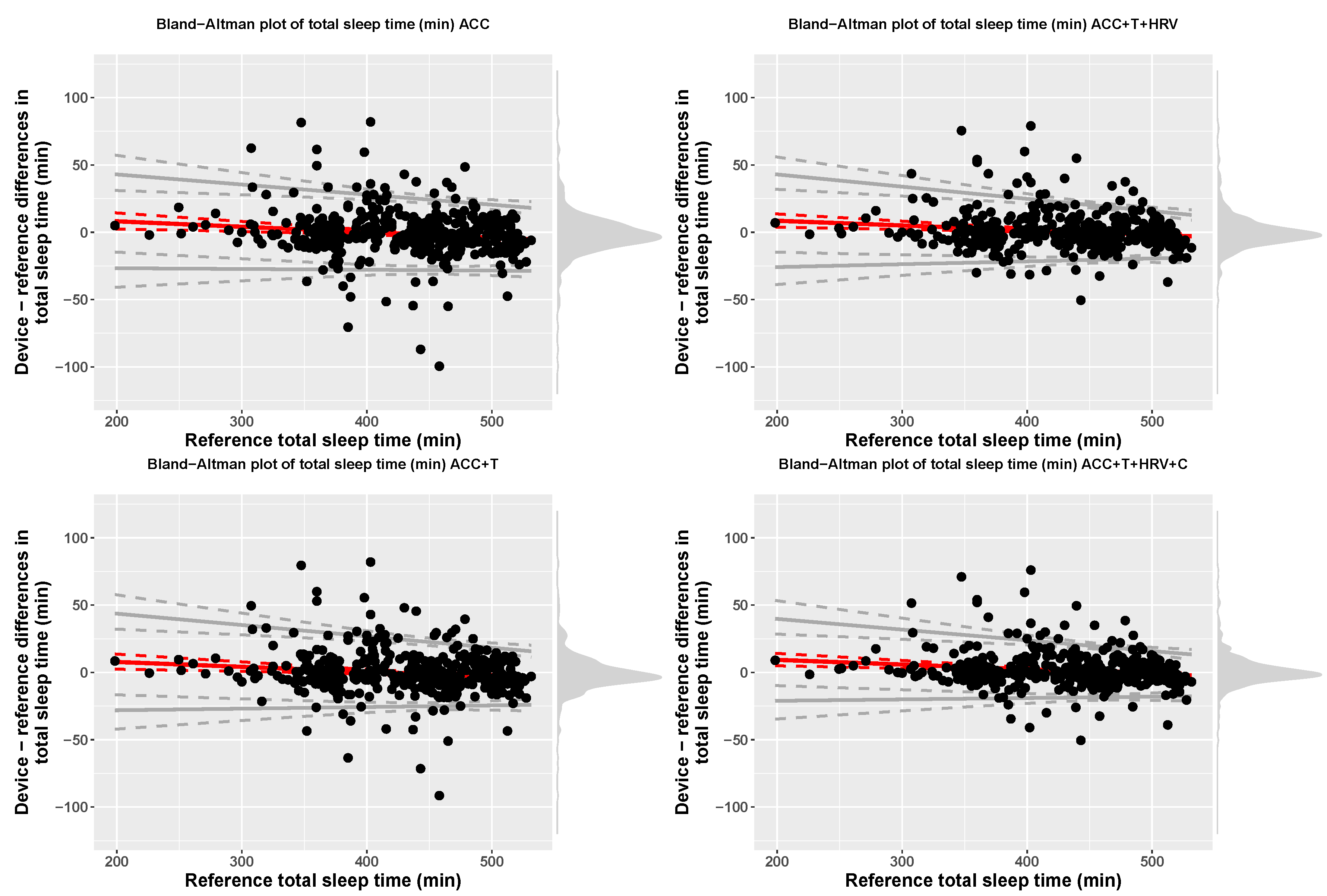
| Model | Device | Reference | Bias | LOA.Lower | LOA.Upper |
|---|---|---|---|---|---|
| ACC | 429.67 (61.05) | 430.66 (61.12) | 16.38 + −0.04 x ref | bias − 2.46(17.19 + −0.01 x ref) | bias + 2.46(17.19 + −0.01 x ref) |
| ACC+T | 430.1 (61) | 430.66 (61.12) | 14.98 + −0.04 x ref | bias − 2.46(18.51 + −0.02 x ref) | bias + 2.46(18.51 + −0.02 x ref) |
| ACC+T+HRV | 431.2 (60.48) | 430.66 (61.12) | 15.49 + −0.03 x ref | bias − 2.46(18.54 + −0.02 x ref) | bias + 2.46(18.54 + −0.02 x ref) |
| ACC+T+HRV+C | 432 (60.34) | 430.66 (61.12) | 16.27 + −0.03 x ref | bias − 2.46(16.02 + −0.02 x ref) | bias + 2.46(16.02 + −0.02 x ref) |
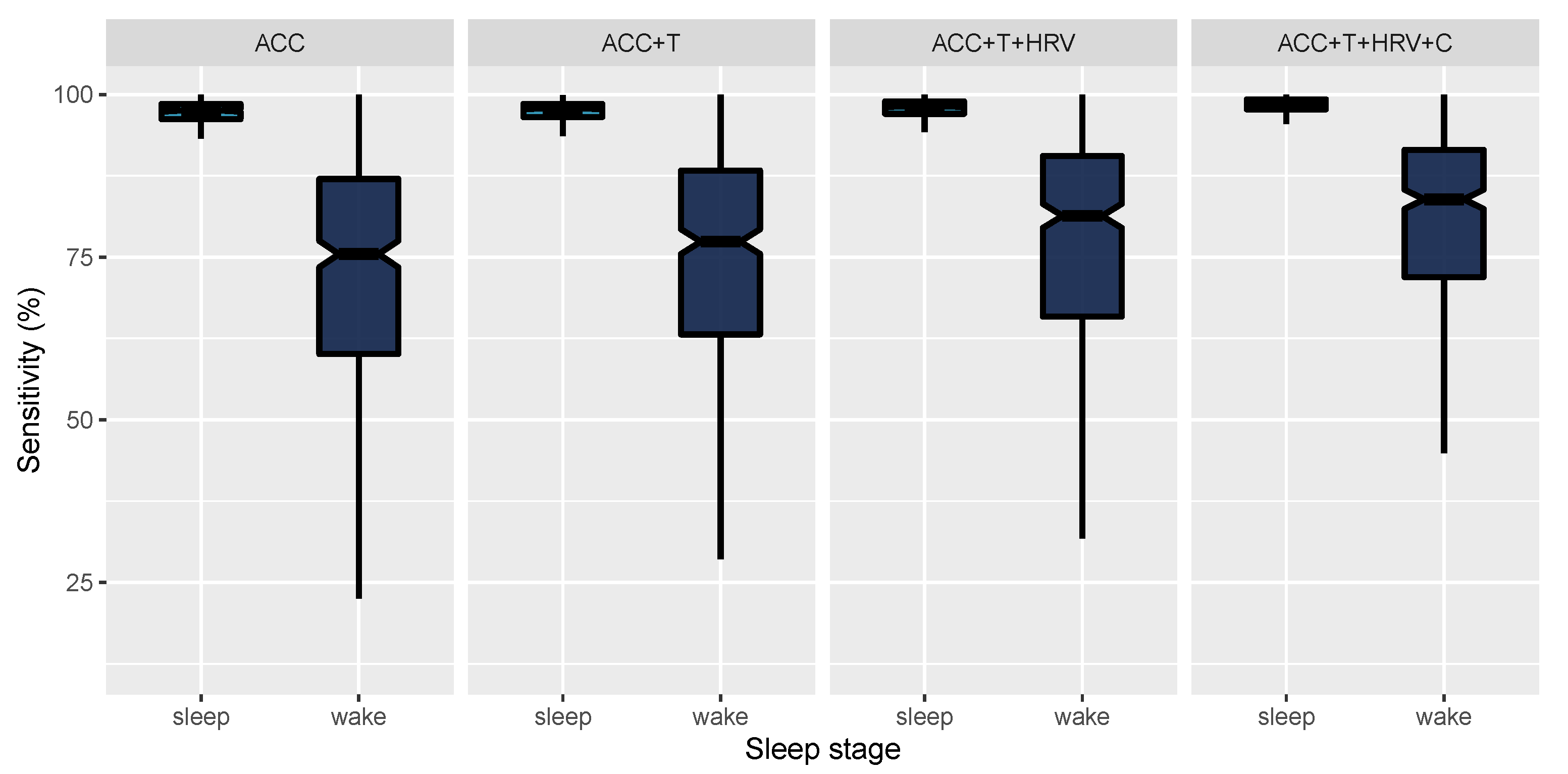
| Model | Sensitivity | Specificity |
|---|---|---|
| ACC | 72.08 (18.44) [70.35, 73.86] | 96.82 (3.04) [96.54, 97.11] |
| ACC+T | 73.71 (17.9) [72.06, 75.42] | 97.05 (2.79) [96.8, 97.31] |
| ACC+T+HRV | 77.18 (16.77) [75.62, 78.76] | 97.61 (2.11) [97.42, 97.81] |
| ACC+T+HRV+C | 80.74 (14.12) [79.44, 82.07] | 98.15 (1.87) [97.98, 98.33] |
| Model | Measure | Device | Reference | Bias | LOA.Lower | LOA.Upper |
|---|---|---|---|---|---|---|
| ACC | TST (min) | 429.49 (61.05) | 430.66 (61.12) | 16.23 + −0.04 x ref | bias − 2.46(16.96 + −0.01 x ref) | bias + 2.46(16.96 + −0.01 x ref) |
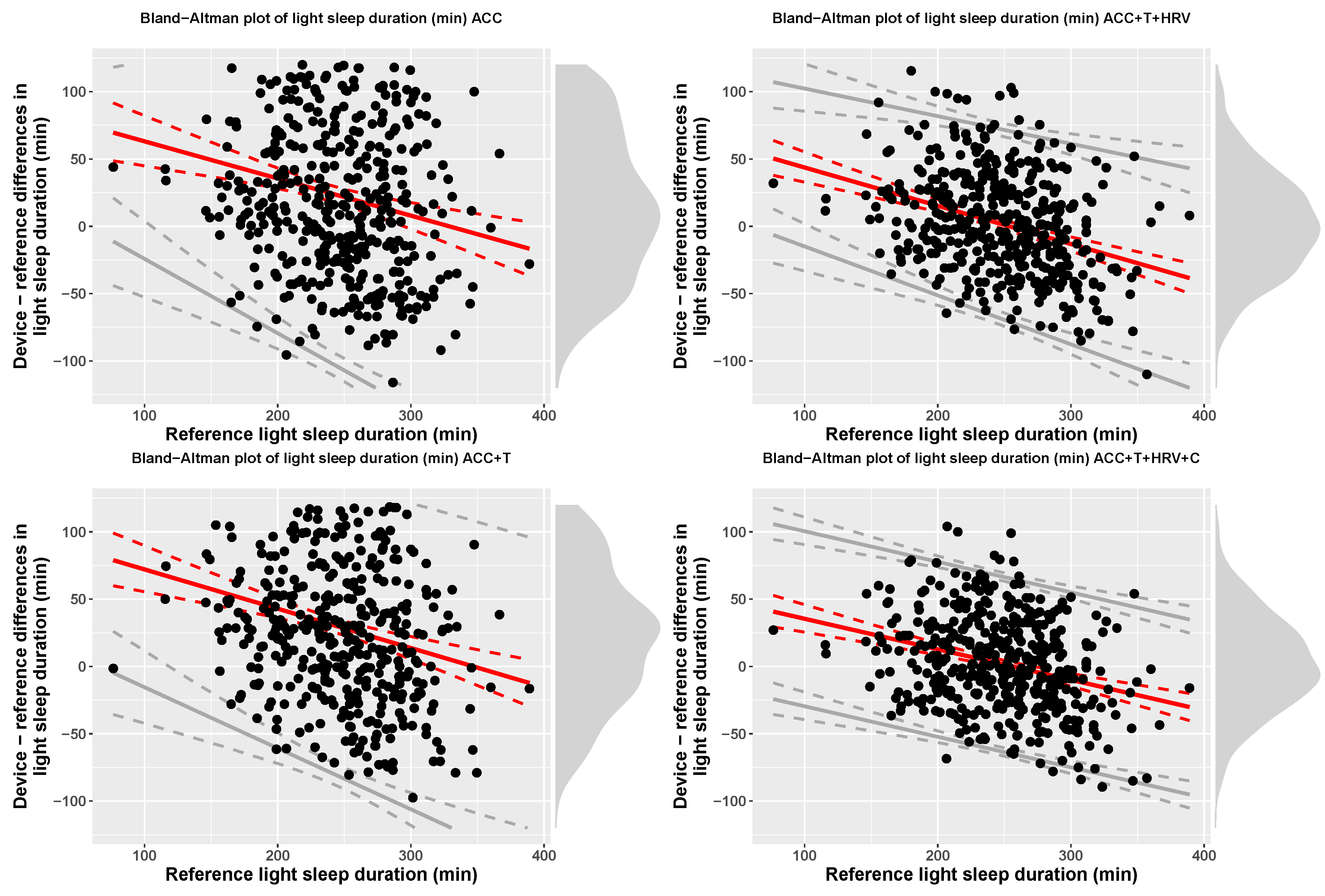
| ACC | Light (min) | 269.78 (71.24) | 247.31 (45.3) | 90.71 + −0.28 x ref | bias − 2.46(24.26 + 0.11 x ref) | bias + 2.46(24.26 + 0.11 x ref) |
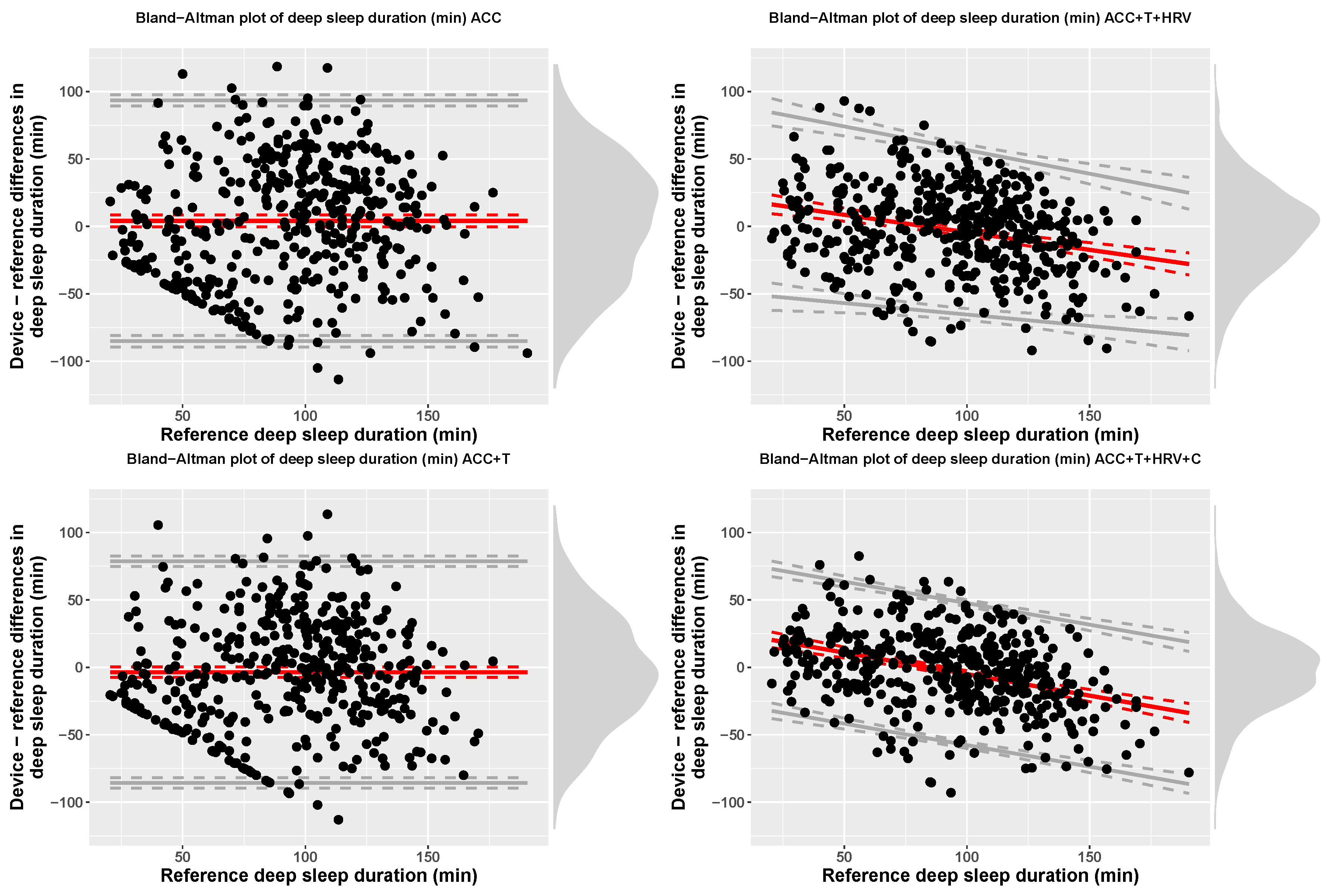
| ACC | Deep (min) | 97.45 (59.24) | 93.34 (34.19) | 4.11 (45.55) | −85.16 | 93.38 |
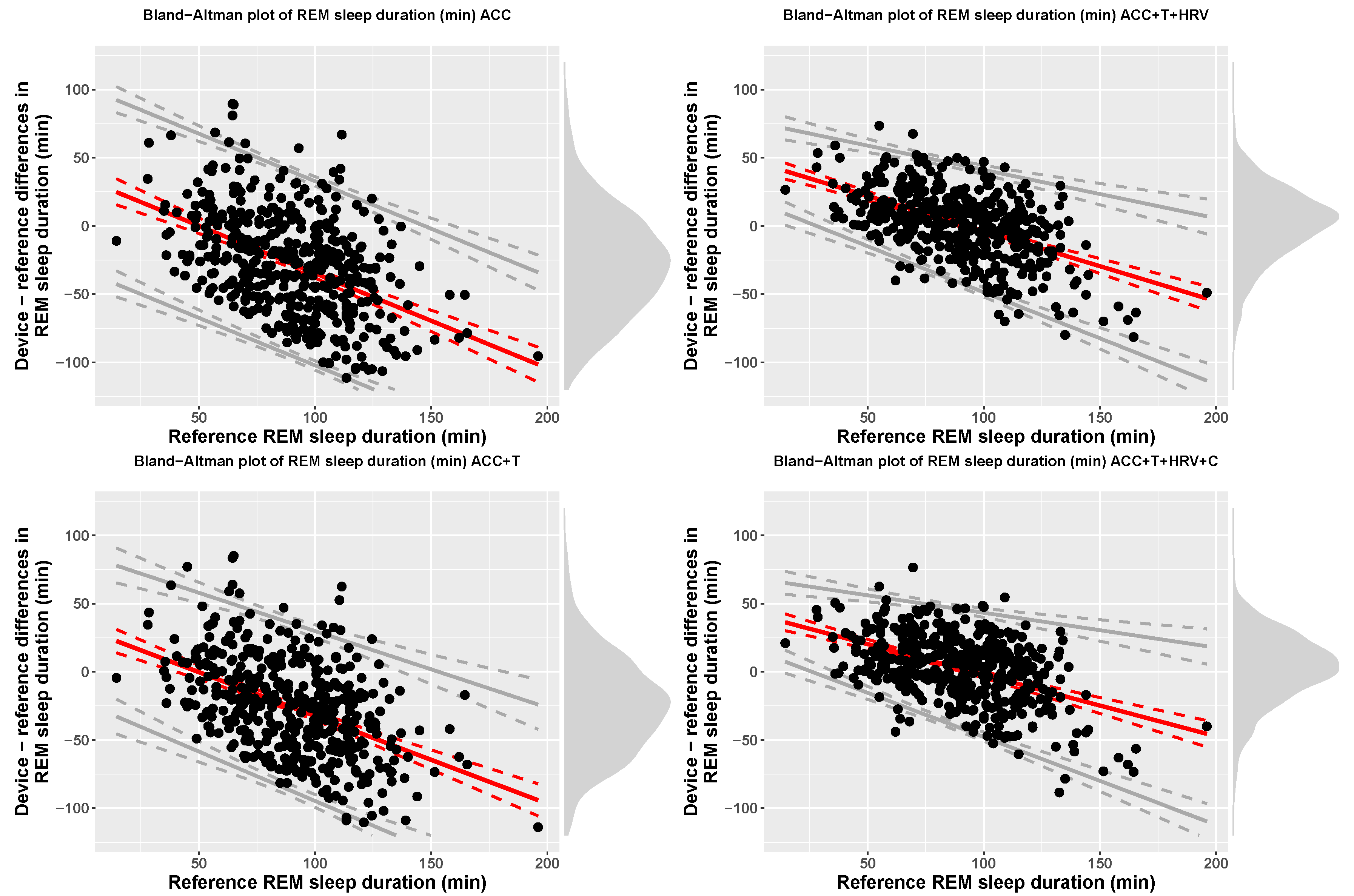
| ACC | REM (min) | 62.26 (35.36) | 90.01 (26.24) | 34.95 + −0.7 x ref | bias − 67.53 | bias + 67.53 |
| ACC+T | TST (min) | 430.1 (61) | 430.66 (61.12) | 14.98 + −0.04 x ref | bias − 2.46(18.51 + −0.02 x ref) | bias + 2.46(18.51 + −0.02 x ref) |
| ACC+T | Light (min) | 276.39 (64.67) | 247.31 (45.3) | 101.27 + −0.29 x ref | bias − 2.46(29.01 + 0.07 x ref) | bias + 2.46(29.01 + 0.07 x ref) |
| ACC+T | Deep (min) | 89.72 (53.45) | 93.34 (34.19) | −3.62 (41.93) | −85.8 | 78.56 |
| ACC+T | REM (min) | 63.98 (32.59) | 90.01 (26.24) | 31.86 + −0.64 x ref | bias − 61.18 | bias + 61.18 |
| ACC+T+HRV | TST (min) | 431.2 (60.48) | 430.66 (61.12) | 15.49 + −0.03 x ref | bias − 2.46(18.54 + −0.02 x ref) | bias + 2.46(18.54 + −0.02 x ref) |
| ACC+T+HRV | Light (min) | 249.12 (48.06) | 247.31 (45.3) | 72.1 + −0.28 x ref | bias − 2.46(20.61 + 0.03 x ref) | bias + 2.46(20.61 + 0.03 x ref) |
| ACC+T+HRV | Deep (min) | 90.69 (40.12) | 93.34 (34.19) | 21.61 + −0.26 x ref | bias − 2.46(28.47 + −0.04 x ref) | bias + 2.46(28.47 + −0.04 x ref) |
| ACC+T+HRV | REM (min) | 91.39 (25.24) | 90.01 (26.24) | 47.76 + −0.52 x ref | bias − 2.46(11.74 + 0.07 x ref) | bias + 2.46(11.74 + 0.07 x ref) |
| ACC+T+HRV+C | TST (min) | 432 (60.34) | 430.66 (61.12) | 16.27 + −0.03 x ref | bias − 2.46(16.02 + −0.02 x ref) | bias + 2.46(16.02 + −0.02 x ref) |
| ACC+T+HRV+C | Light (min) | 249.31 (48.24) | 247.31 (45.3) | 58.1 + −0.23 x ref | bias − 65 | bias + 65 |
| ACC+T+HRV+C | Deep (min) | 90.43 (35.54) | 93.34 (34.19) | 26.85 + −0.32 x ref | bias − 52.63 | bias + 52.63 |
| ACC+T+HRV+C | REM (min) | 92.26 (26.19) | 90.01 (26.24) | 42.83 + −0.45 x ref | bias − 2.46(10.58 + 0.08 x ref) | bias + 2.46(10.58 + 0.08 x ref) |
| Model | Stage | Accuracy | Sensitivity | Specificity |
|---|---|---|---|---|
| ACC | Wake | 94.53 (3.79) [94.18, 94.88] | 72.07 (17.61) [70.44, 73.74] | 96.85 (2.94) [96.58, 97.14] |
| ACC | Light | 61.27 (5.72) [60.74, 61.8] | 67.74 (14.22) [66.45, 69.03] | 53.11 (16.2) [51.63, 54.62] |
| ACC | Deep | 80.44 (6.01) [79.87, 81] | 48.14 (28.42) [45.51, 50.77] | 87.15 (7.88) [86.42, 87.9] |
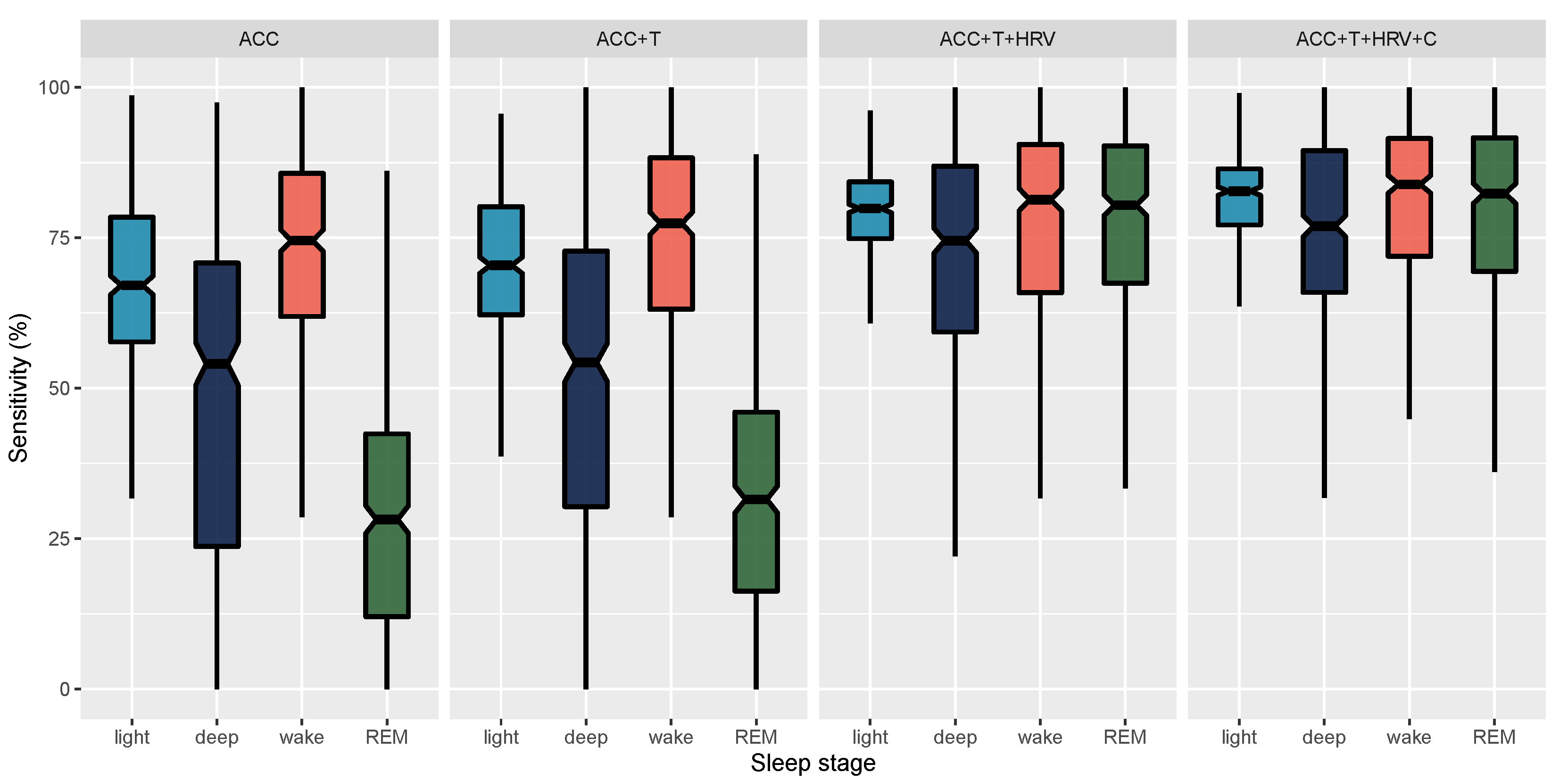
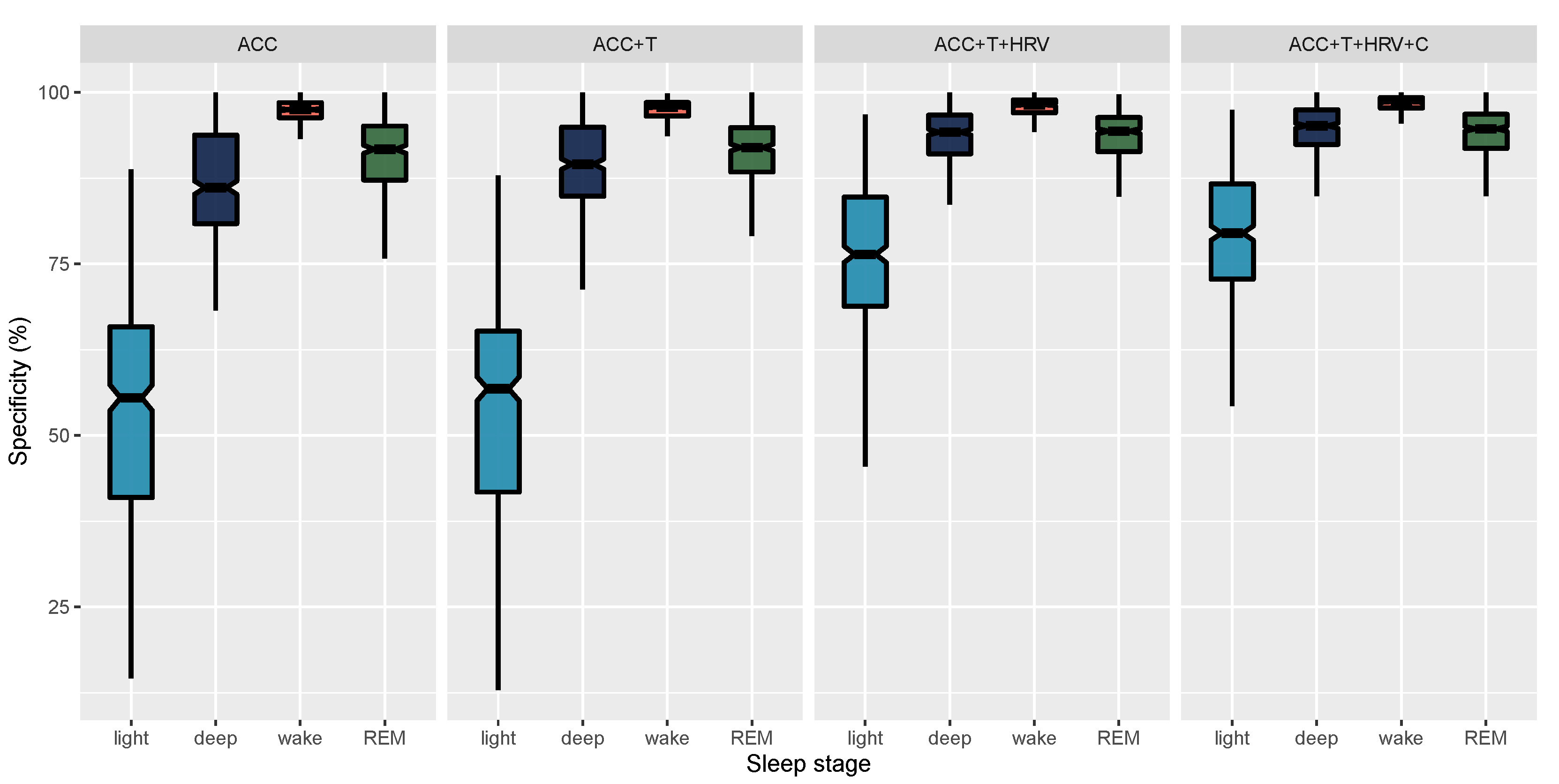
| ACC | REM | 78.98 (5.2) [78.5, 79.47] | 28.99 (19.37) [27.19, 30.81] | 90.72 (5.89) [90.17, 91.27] |
| ACC+T | Wake | 94.79 (3.93) [94.43, 95.17] | 73.71 (17.9) [72.07, 75.41] | 97.05 (2.79) [96.79, 97.32] |
| ACC+T | Light | 63.09 (6.8) [62.46, 63.72] | 70.86 (12) [69.76, 71.98] | 53.77 (16.32) [52.27, 55.31] |
| ACC+T | Deep | 82.69 (5.91) [82.14, 83.25] | 50.06 (28.88) [47.33, 52.71] | 89.67 (6.66) [89.07, 90.29] |
| ACC+T | REM | 79.9 (5.35) [79.4, 80.4] | 32.38 (19.33) [30.55, 34.18] | 91.04 (5.44) [90.55, 91.57] |
| ACC+T+HRV | Wake | 95.58 (3.5) [95.25, 95.92] | 77.18 (16.77) [75.65, 78.76] | 97.61 (2.11) [97.42, 97.81] |
| ACC+T+HRV | Light | 77.48 (6.14) [76.91, 78.05] | 79.13 (7.38) [78.45, 79.82] | 75.73 (11.75) [74.62, 76.84] |
| ACC+T+HRV | Deep | 89.11 (4.25) [88.72, 89.51] | 69.57 (23.84) [67.39, 71.8] | 93.73 (4.16) [93.35, 94.12] |
| ACC+T+HRV | REM | 90.16 (4.18) [89.78, 90.57] | 75.89 (18.09) [74.23, 77.56] | 93.75 (3.4) [93.44, 94.06] |
| ACC+T+HRV+C | Wake | 96.38 (3.12) [96.1, 96.68] | 80.74 (14.12) [79.43, 82.06] | 98.15 (1.87) [97.98, 98.33] |
| ACC+T+HRV+C | Light | 80.2 (5.53) [79.68, 80.73] | 81.7 (6.97) [81.06, 82.35] | 78.67 (10.63) [77.7, 79.67] |
| ACC+T+HRV+C | Deep | 90.64 (3.75) [90.29, 90.99] | 74.44 (20.28) [72.56, 76.34] | 94.63 (3.77) [94.28, 94.99] |
| ACC+T+HRV+C | REM | 90.87 (4.12) [90.49, 91.26] | 78.08 (17.39) [76.5, 79.71] | 94.12 (3.41) [93.8, 94.44] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Altini, M.; Kinnunen, H. The Promise of Sleep: A Multi-Sensor Approach for Accurate Sleep Stage Detection Using the Oura Ring. Sensors 2021, 21, 4302. https://doi.org/10.3390/s21134302
Altini M, Kinnunen H. The Promise of Sleep: A Multi-Sensor Approach for Accurate Sleep Stage Detection Using the Oura Ring. Sensors. 2021; 21(13):4302. https://doi.org/10.3390/s21134302
Chicago/Turabian StyleAltini, Marco, and Hannu Kinnunen. 2021. "The Promise of Sleep: A Multi-Sensor Approach for Accurate Sleep Stage Detection Using the Oura Ring" Sensors 21, no. 13: 4302. https://doi.org/10.3390/s21134302
APA StyleAltini, M., & Kinnunen, H. (2021). The Promise of Sleep: A Multi-Sensor Approach for Accurate Sleep Stage Detection Using the Oura Ring. Sensors, 21(13), 4302. https://doi.org/10.3390/s21134302







