HEMIGEN: Human Embryo Image Generator Based on Generative Adversarial Networks
Abstract
1. Introduction
2. Materials and Methods
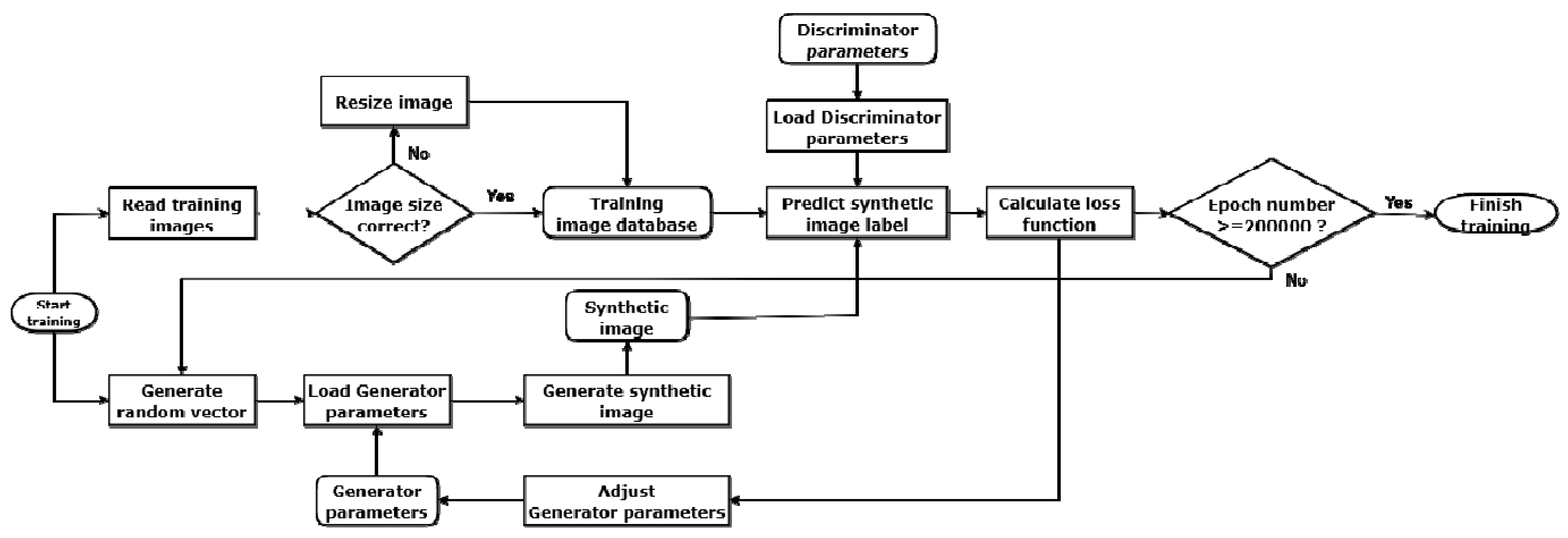
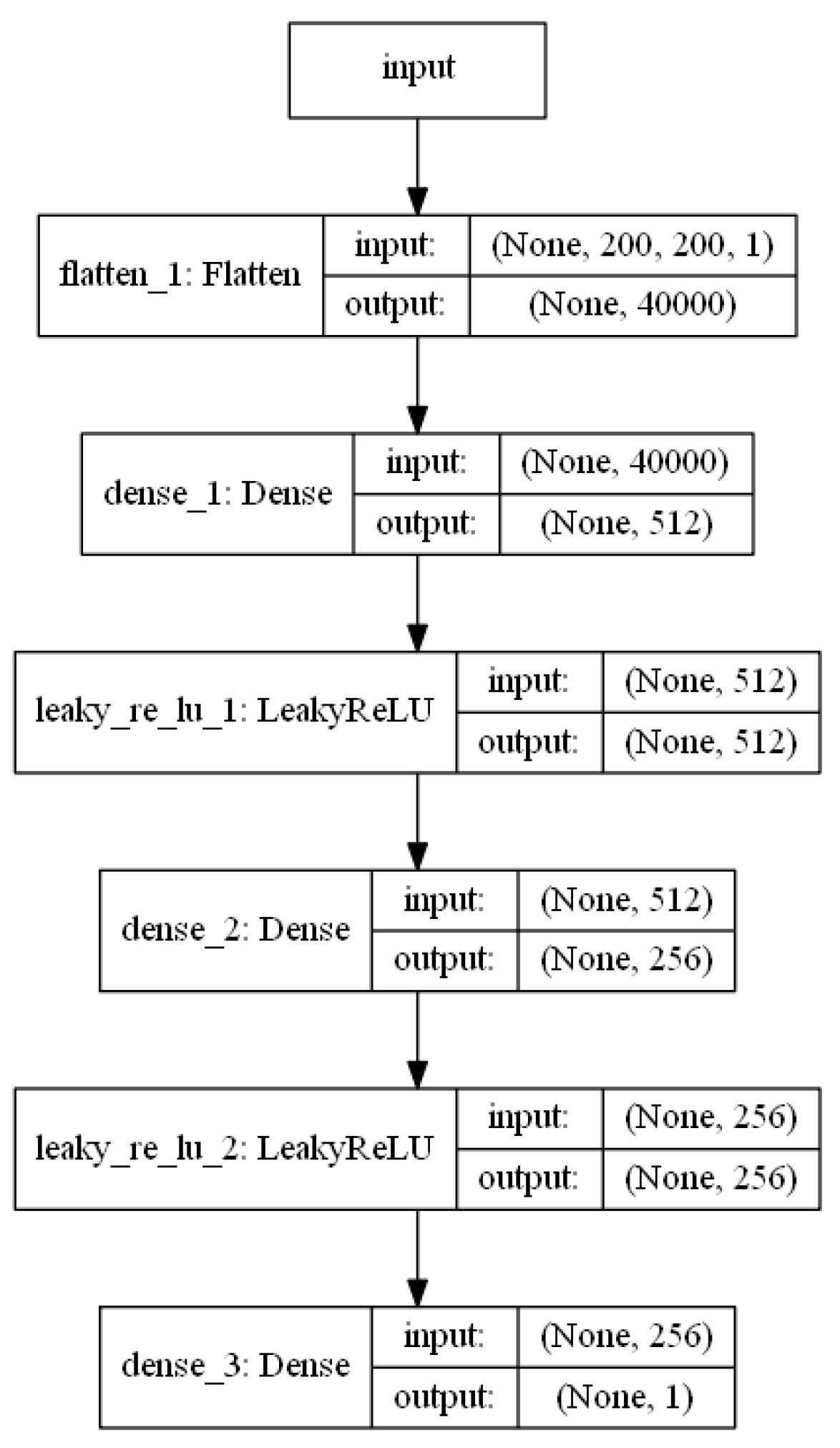
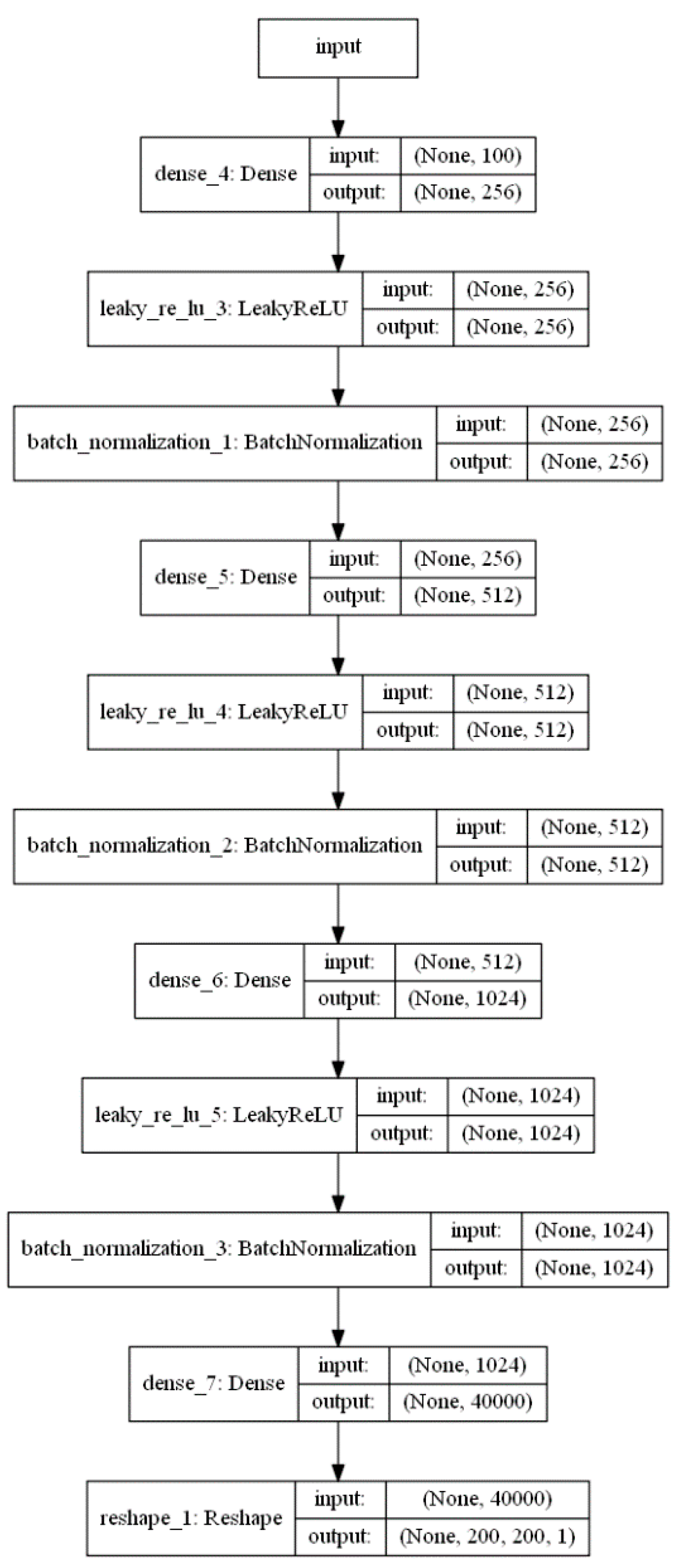
2.1. Architecture of the Generative Adversarial Network (GAN)
2.2. Training
2.3. Evaluation of Image Quality
2.4. Ethics Declaration
3. Experiment and Results
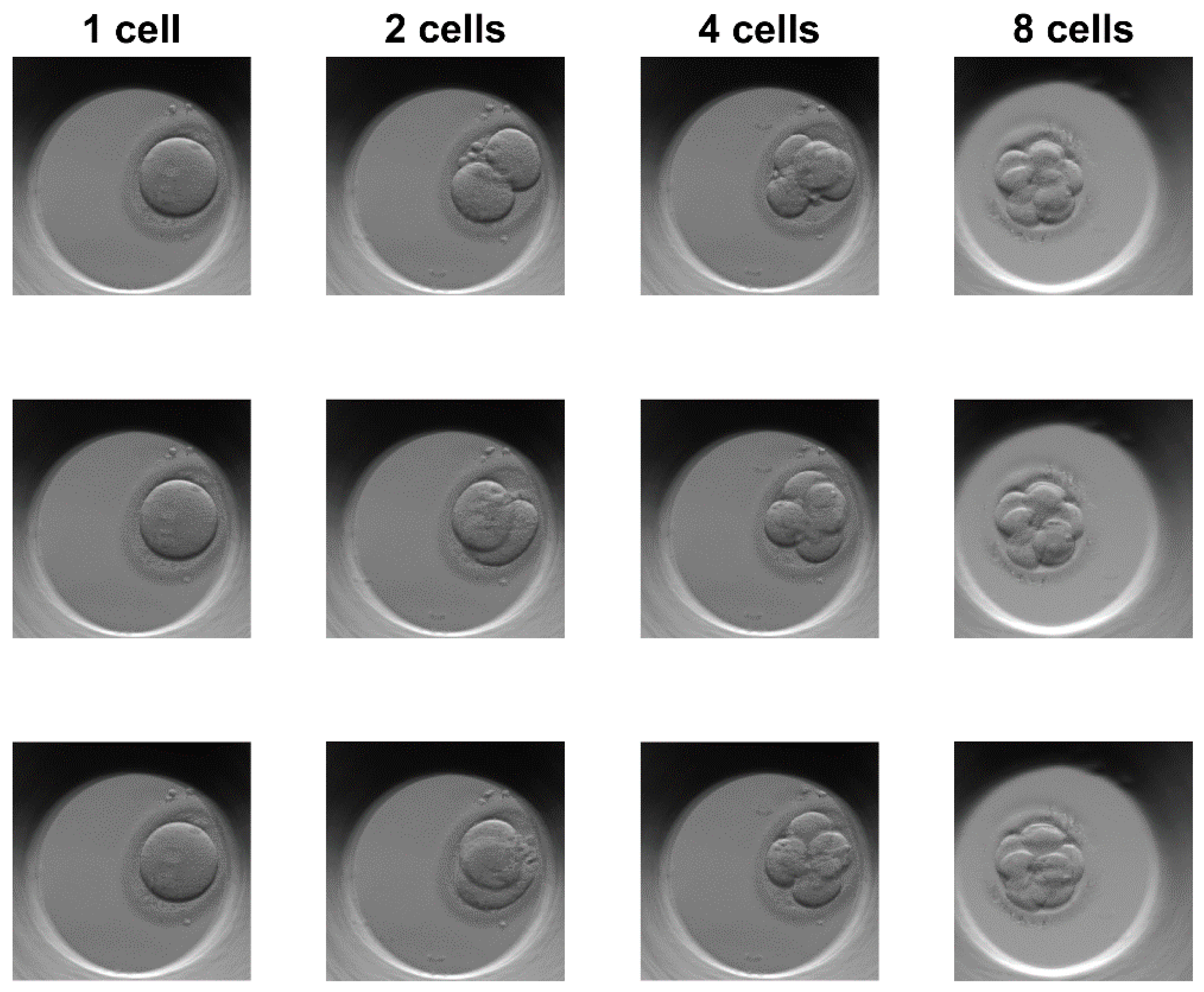
3.1. Dataset and Equipment
3.2. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Russakovsky, O.; Deng, J.; Su, H.; Krause, J.; Satheesh, S.; Ma, S.; Huang, Z.; Karpathy, A.; Khosla, A.; Bernstein, M.; et al. ImageNet Large Scale Visual Recognition Challenge. Int. J. Comput. Vis. 2015, 115, 211. [Google Scholar] [CrossRef]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. Commun. ACM 2017, 60, 84–90. [Google Scholar] [CrossRef]
- Donahue, J.; Jia, Y.; Vinyals, O.; Hoffman, J.; Zhang, N.; Tzeng, E.; Darrell, T. DeCAF: A Deep Convolutional Activation Feature for Generic Visual Recognition. In Proceedings of the 31st International Conference on Machine Learning, Beijing, China, 21–26 June 2014; Volume 32, pp. 647–655. [Google Scholar]
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition. In Proceedings of the International Conference on Learning Representations ICLR, San Diego, CA, USA, 7–9 May 2015; pp. 1–14. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Jersey, NJ, USA, 26 June–1 July 2016; IEEE: Piscataway, NJ, USA. [Google Scholar] [CrossRef]
- Long, J.; Shelhamer, E.; Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the 2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Boston, MA, USA, 7–12 June 2015; pp. 3431–3440. [Google Scholar] [CrossRef]
- Girshick, R.B.; Donahue, J.; Darrell, T.; Malik, J. Rich feature hierarchies for accurate object detection and semantic segmentation. In Proceedings of the 2014 IEEE Conference on Computer Vision and Pattern Recognition (CVPR ’14), Washington, DC, USA, 23–28 June 2014; pp. 580–587. [Google Scholar] [CrossRef]
- Esteva, A.; Kuprel, B.; Novoa, R.A.; Ko, J.; Swetter, S.M.; Blau, H.M.; Thrun, S. Dermatologist-level classification of skin cancer with deep neural networks. Nature 2017, 542, 115–118. [Google Scholar] [CrossRef] [PubMed]
- Fakoor, R.; Ladhak, F.; Nazi, A.; Huber, M. Using deep learning to enhance cancer diagnosis and classification. In ICML Workshop on the Role of Machine Learning in Transforming Healthcare Proceedings of the International Conference on Machine Learning, Atlanta, AG, USA, 16–21 June 2013; ACM: New York, NY, USA, 2013; pp. 1–7. [Google Scholar]
- Ciresan, D.C.; Giusti, A.; Gambardella, L.M.; Schmidhuber, J. Mitosis Detection in Breast Cancer Histology Images with Deep Neural Networks. In Proceedings of the Medical Image Computing and Computer-Assisted Intervention, MICCAI 2013, Nagoya, Japan, 22–26 September 2013; pp. 411–418. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In Proceedings of the Medical Image Computing and Computer-Assisted Intervention, MICCAI 2015, Munich, Germany, 5–9 October 2015; Lecture Notes in Computer Science. Springer: Cham, Switzerland, 2015; Volume 9351, pp. 234–241. [Google Scholar]
- Dong, B.; Shao, L.; Da Costa, M.; Bandmann, O.; Frangi, A.F. Deep learning for automatic cell detection in wide-field microscopy zebrafish images. In Proceedings of the 2015 IEEE 12th International Symposium on Biomedical Imaging (ISBI), Brooklyn, NY, USA, 16–19 April 2015; pp. 772–776. [Google Scholar]
- Kheradmand, S.; Singh, A.; Saeedi, P.; Au, J.; Havelock, J. Inner cell mass segmentation in human HMC embryo images using fully convolutional network. In Proceedings of the 2017 IEEE International Conference on Image Processing (ICIP), Beijing, China, 17–20 September 2017; pp. 1752–1756. [Google Scholar]
- Manna, C.; Nanni, L.; Lumini, A.; Pappalardo, S. Artificial intelligence techniques for embryo and oocyte classification. Reprod. Biomed. Online 2013, 26, 42–49. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Moussavi, F.; Lorenzen, P. Automated Embryo Stage Classification in Time-Lapse Microscopy Video of Early Human Embryo Development. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, MICCAI (2013), Nagoya, Japan, 22–26 September 2013; pp. 460–467. [Google Scholar] [CrossRef]
- Khan, A.; Gould, S.; Salzmann, M. Automated monitoring of human embryonic cells up to the 5-cell stage in time-lapse microscopy images. In Proceedings of the 2015 IEEE 12th International Symposium on Biomedical Imaging (ISBI), New York, NY, USA, 16–19 April 2015; pp. 389–393. [Google Scholar]
- Khan, A.; Gould, S.; Salzmann, M. A Linear Chain Markov Model for Detection and Localization of Cells in Early Stage Embryo Development. In Proceedings of the 2015 IEEE Winter Conference on Applications of Computer Vision, Waikoloa, HI, USA, 6–9 January 2015; pp. 526–533. [Google Scholar]
- Nandy, K.; Kim, J.; McCullough, D.P.; McAuliffe, M.; Meaburn, K.J.; Yamaguchi, T.P.; Gudla, P.R.; Lockett, S.J. Segmentation and Quantitative Analysis of Individual Cells in Developmental Tissues. In Mouse Molecular Embryology; Methods in Molecular Biology (Methods and Protocols); Lewandoski, M., Ed.; Humana Press: Boston, MA, USA, 2014; Volume 1092, pp. 235–253. [Google Scholar]
- Baggett, D.; Nakaya, M.-A.; McAuliffe, M.; Yamaguchi, T.P.; Lockett, S. Whole cell segmentation in solid tissue sections. Cytometry 2005, 67A, 137–143. [Google Scholar] [CrossRef] [PubMed]
- Dirvanauskas, D.; Maskeliunas, R.; Raudonis, V.; Damasevicius, R. Embryo development stage prediction algorithm for automated time lapse incubators. Comput. Methods Programs Biomed. 2019, 177, 161–174. [Google Scholar] [CrossRef] [PubMed]
- Osokin, A.; Chessel, A.; Salas, R.E.C.; Vaggi, F. GANs for Biological Image Synthesis. In Proceedings of the 2017 IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017; IEEE: Piscataway, NJ, USA. [Google Scholar] [CrossRef]
- Karras, T.; Aila, T.; Laine, S.; Lehtinen, J. Progressive growing of GANs for improved quality, stability, and variation. arXiv 2017, arXiv:1710.10196. [Google Scholar]
- Han, C.; Rundo, L.; Araki, R.; Furukawa, Y.; Mauri, G.; Nakayama, H.; Hayashi, H. Infinite Brain MR Images: PGGAN-based Data Augmentation for Tumor Detection. arXiv 2019, arXiv:1903.12564. [Google Scholar]
- Khan, S.H.; Hayat, M.; Barnes, N. Adversarial Training of Variational Auto-encoders for High Fidelity Image Generation. In Proceedings of the 2018 IEEE Winter Conference on Applications of Computer Vision (WACV), Waikoloa Village, HI, USA, 12–15 March 2018; IEEE: Piscataway, NJ, USA, 2018. [Google Scholar] [CrossRef]
- Yi, X.; Walia, E.; Babyn, P. Generative adversarial network in medical imaging: A review. arXiv 2018, arXiv:1809.07294. [Google Scholar]
- Bengio, Y.; Thibodeau-Laufer, É.; Alain, G.; Yosinski, J. Deep Generative Stochastic Networks Trainable by Backprop. In Proceedings of the 31st International Conference on International Conference on Machine Learning, Beijing, China, 21–26 June 2014; Volume 32, pp. II-226–II-234. [Google Scholar]
- Gregor, K.; Danihelka, I.; Graves, A.; Jimenez Rezende, D.; Wierstra, D. DRAW: A Recurrent Neural Network for Image Generation. In Proceedings of the 32nd International Conference on Machine Learning, Lille, France, 6–11 July 2015; Volume 37, pp. 1462–1471. [Google Scholar]
- Dosovitskiy, A.; Springenberg, J.T.; Brox, T. Learning to generate chairs with convolutional neural networks. In Proceedings of the 2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Boston, MA, USA, 7–12 June 2015; pp. 1538–1546. [Google Scholar]
- Denton, E.; Chintala, S.; Szlam, A.; Fergus, R. Deep Generative Image Models using a Laplacian Pyramid of Adversarial Networks. In Proceedings of the 28th International Conference on Neural Information Processing Systems-Volume 1 (NIPS’15), Montreal, QC, Canada, 7–12 December 2015; MIT Press: Cambridge, MA, USA, 2015; Volume 1, pp. 1486–1494. [Google Scholar]
- Radford, A.; Metz, L.; Chintala, S. Unsupervised Representation Learning with Deep Convolutional Generative Adversarial Networks. arXiv 2015, arXiv:1511.06434. [Google Scholar]
- Reed, S.; Akata, Z.; Yan, X.; Logeswaran, L.; Schiele, B.; Lee, H. Generative Adversarial Text to Image Synthesis. In Proceedings of the 33rd International Conference on Machine Learning, New York, NY, USA, 20–22 June 2016; Volume 48, pp. 1060–1069. [Google Scholar]
- Zhang, H.; Xu, T.; Li, H.; Zhang, S.; Wang, X.; Huang, X.; Metaxas, D. StackGAN: Text to Photo-realistic Image Synthesis with Stacked Generative Adversarial Networks. In Proceedings of the 2017 IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017; pp. 5908–5916. [Google Scholar]
- Tulyakov, S.; Liu, M.-Y.; Yang, X.; Kautz, J. MoCoGAN: Decomposing Motion and Content for Video Generation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Salt Lake, UT, USA, 18–22 June 2018; pp. 1526–1535. [Google Scholar]
- Vondrick, C.; Pirsiavash, H.; Torralba, A. Generating Videos with Scene Dynamics. In Proceedings of the 30th International Conference on Neural Information Processing Systems (NIPS’16), Long Beach, CA, USA, 4–9 December 2017; pp. 613–621. [Google Scholar]
- Saito, M.; Matsumoto, E.; Saito, S. Temporal Generative Adversarial Nets with Singular Value Clipping. In Proceedings of the 2017 IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017; pp. 2849–2858. [Google Scholar] [CrossRef]
- Wu, J.; Zhang, C.; Xue, T.; Freeman, W.T.; Tenenbaum, J.B. Learning a Probabilistic Latent Space of Object Shapes via 3D Generative-Adversarial Modeling. In Proceedings of the 30th International Conference on Neural Information Processing Systems (NIPS’16), Boston, MA, USA, 15–19 September 2016; pp. 82–90. [Google Scholar]
- Li, C.; Guo, Y.; Liu, Q.; Liu, X. DR-Net: A Novel Generative Adversarial Network for Single Image Deraining. Secur. Commun. Netw. 2018, 2018, 7350324. [Google Scholar] [CrossRef]
- Zhu, D.; Dai, L.; Luo, Y.; Zhang, G.; Shao, X.; Itti, L.; Lu, J. Multi-Scale Adversarial Feature Learning for Saliency Detection. Symmetry 2018, 10, 457. [Google Scholar] [CrossRef]
- Ma, Y.; Liu, K.; Guan, Z.; Xu, X.; Qian, X.; Bao, H. Background Augmentation Generative Adversarial Networks (BAGANs): Effective Data Generation Based on GAN-Augmented 3D Synthesizing. Symmetry 2018, 10, 734. [Google Scholar] [CrossRef]
- Han, C.; Hayashi, H.; Rundo, L.; Araki, R.; Shimoda, W.; Muramatsu, S.; Nakayama, H. GAN-based synthetic brain MR image generation. In Proceedings of the 2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018), Washington, DC, USA, 4–7 April 2018; IEEE: Piscataway, NJ, USA, 2018. [Google Scholar] [CrossRef]
- Goodfellow, I.J.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative adversarial nets. In Proceedings of the 27th International Conference on Neural Information Processing Systems-Volume 2 (NIPS’14), Montreal, QC, Canada, 8–13 December 2014; MIT Press: Cambridge, MA, USA, 2014; pp. 2672–2680. [Google Scholar]
- Maas, A.L.; Hannun, A.Y.; Ng, A.Y. Rectifier nonlinearities improve neural network acoustic model. In Proceedings of the International Conference on Machine ICML 2013, Atlanta, GA, USA, 16–21 June 2013; Volume 30, p. 3. [Google Scholar]
- Ioffe, S.; Szegedy, C. Batch Normalization: Accelerating Deep Network Training by Reducing Internal Covariate Shift. In Proceedings of the 32nd International Conference on International Conference on Machine Learning-Volume 37 (ICML’15), Lille, France, 7–9 July 2015; Volume 37, pp. 448–456. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A Method for Stochastic Optimization. In Proceedings of the 3rd International Conference on Learning Representations, Indianapolis, IN, USA, 24–28 March 2014. [Google Scholar]
- Theis, L.; van den Oord, A.; Bethge, M. A note on the evaluation of generative models. In Proceedings of the International Conference on Learning Representations (ICLR), San Juan, Puerto Rico, 19 November 2015. [Google Scholar]
- Im, D.J.; Kim, C.D.; Jiang, H.; Memisevic, R. Generating Images with Recurrent Adversarial Networks. arXiv 2016, arXiv:1602.05110. [Google Scholar]
- Geman, D.; Geman, S.; Hallonquist, N.; Younes, L. Visual Turing test for computer vision systems. In Proceedings of the National Academy of Sciences, Washington, DC, USA, 22–29 December 2015; p. 201422953. [Google Scholar] [CrossRef]
- Salimans, T.; Goodfellow, I.; Zaremba, W.; Cheung, V.; Radford, A.; Chen, X. Improved techniques for training GANs. In Proceedings of the 30th International Conference on Neural Information Processing Systems (NIPS’16), Barcelona, Spain, 5–10 December 2016; Curran Associates Inc.: Red Hook, NY, USA, 2016; pp. 2234–2242. [Google Scholar]
- Tazehkandi, A.A. Computer Vision with OpenCV 3 and Qt5; Packt Publishing: Birmingham, UK, 2018. [Google Scholar]
- Haralick, R.M.; Shanmugam, K.; Dinstein, I. Textural Features for Image Classification. IEEE Trans. Syst. Man Cybern. 1973, SMC-3, 610–621. [Google Scholar] [CrossRef]
- Kingma, D.P.; Rezende, D.J.; Mohamed, S.; Welling, M. Semi-supervised learning with deep generative models. In Proceedings of the 27th International Conference on Neural Information Processing Systems-Volume 2 (NIPS’14), Montreal, QC, Canada, 8–13 December 2014; MIT Press: Cambridge, MA, USA, 2014; pp. 3581–3589. [Google Scholar]
- Maaløe, L.; Sønderby, C.K.; Sønderby, S.K.; Winther, O. Auxiliary deep generative models. In Proceedings of the 33rd International Conference on Machine Learning, New York, NY, USA, 19–24 June 2016; Volume 48, pp. 1445–1453. [Google Scholar]
- Dumoulin, V.; Belghazi, I.; Poole, B.; Lamb, M.A.; Mastropietro, O.; Courville, A. Adversarially Learned Inference. In Proceedings of the International Conference on Learning Representations (ICLR), Toulon, France, 24–26 April 2017. [Google Scholar]
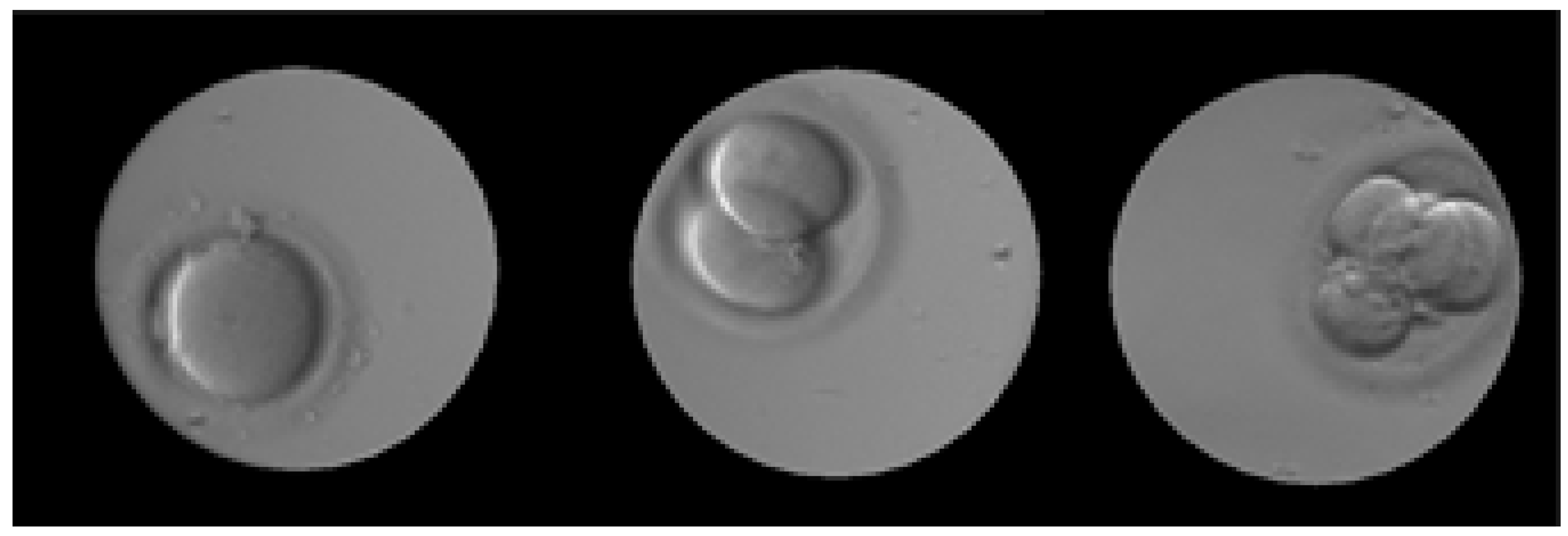

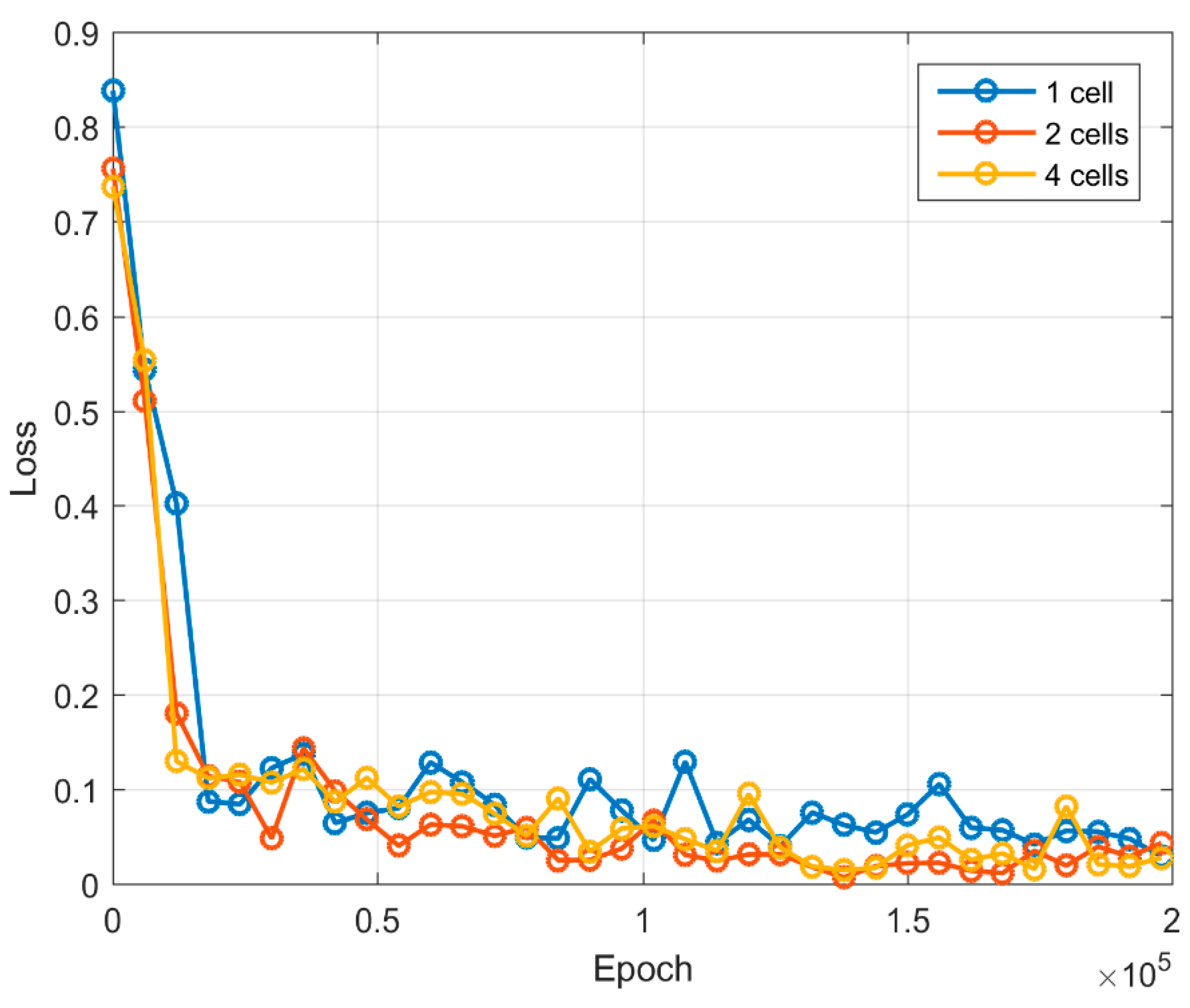
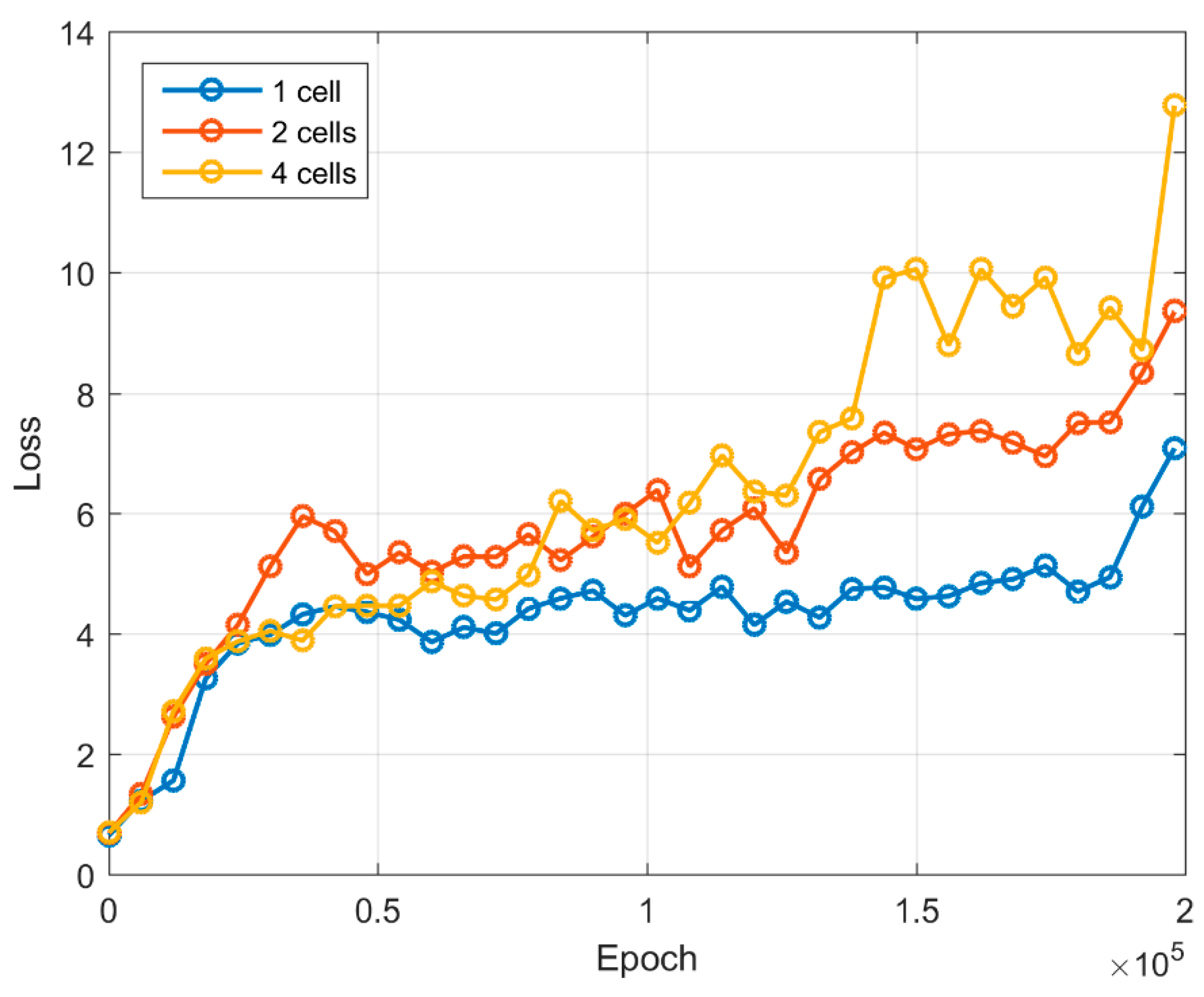
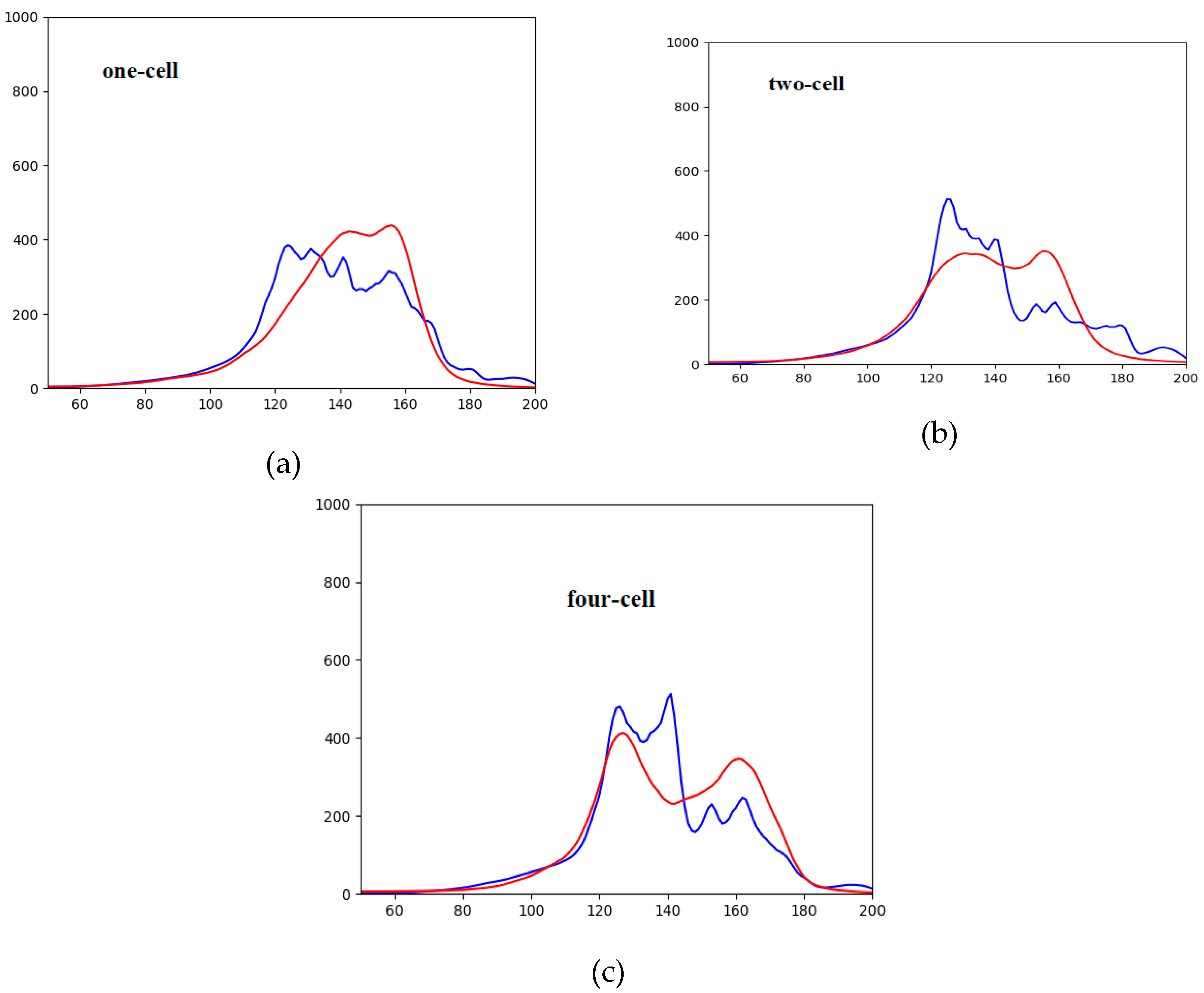
| Epoch | 0 | 25,000 | 50,000 | 75,000 | 100,000 | 125,000 | 150,000 | 175,000 | 200,000 |
|---|---|---|---|---|---|---|---|---|---|
| one-cell |  |  |  |  |  |  |  |  |  |
| two-cells |  |  |  |  |  |  |  |  |  |
| four-cells |  |  |  |  |  |  |  |  |  |
| Class | Test Images (Generated) | Good Images | Accuracy by Expert Selection |
|---|---|---|---|
| one-cell | 500 | 481 | 96.2% |
| two-cells | 500 | 434 | 86.8% |
| four-cells | 500 | 400 | 80.00% |
| One-cell | Two-cells | Four-cells | |
|---|---|---|---|
| Correlation | 0.995 | 0.990 | 0.986 |
| Chi-square | 0.236 | 0.455 | 0.442 |
| Intersection | 1.883 | 1.849 | 1.873 |
| Bhattacharyya | 0.147 | 0.210 | 0.208 |
| One-cell | Two-cells | Four-cells | |
|---|---|---|---|
| Explained variance of PC1 (original) | 99.90% | 99.94% | 99.86% |
| Explained variance of PC1 (synthetic) | 98.95% | 99.60% | 99.73% |
| Correlation between values of PC1 (original) and PC1 (synthetic) | 1.0000 | 0.9997 | 0.9999 |
| Model | Images Analyzed | Overall Misclassification Rate |
|---|---|---|
| DGN [51] | Digits 0 to 9 and combination (based on SVHN, and NORB sets) (realistic images) | 36.02% |
| Skip Deep Generative Model [52] | Digits 0 to 9 and combination (based on SVHN, and NORB sets) (realistic images) | 16.61% |
| GAN (feature matching) [48] | LSVRC2012 dataset with 1,000 categories (non-realistic images) | 8.11% |
| ALI [53] | 25 types of shapes (non-realistic images) | 7.42% |
| WGAN [40] | 6 classes of brain MR images (realistic images) | 50% |
| WGAN-GP [21] | 8 classes of florescent microscopy images (realistic images) | 6.9% |
| HEMIGEN (our approach) | Once cell, Two cells, Four cells (realistic images) | 12.3% |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dirvanauskas, D.; Maskeliūnas, R.; Raudonis, V.; Damaševičius, R.; Scherer, R. HEMIGEN: Human Embryo Image Generator Based on Generative Adversarial Networks. Sensors 2019, 19, 3578. https://doi.org/10.3390/s19163578
Dirvanauskas D, Maskeliūnas R, Raudonis V, Damaševičius R, Scherer R. HEMIGEN: Human Embryo Image Generator Based on Generative Adversarial Networks. Sensors. 2019; 19(16):3578. https://doi.org/10.3390/s19163578
Chicago/Turabian StyleDirvanauskas, Darius, Rytis Maskeliūnas, Vidas Raudonis, Robertas Damaševičius, and Rafal Scherer. 2019. "HEMIGEN: Human Embryo Image Generator Based on Generative Adversarial Networks" Sensors 19, no. 16: 3578. https://doi.org/10.3390/s19163578
APA StyleDirvanauskas, D., Maskeliūnas, R., Raudonis, V., Damaševičius, R., & Scherer, R. (2019). HEMIGEN: Human Embryo Image Generator Based on Generative Adversarial Networks. Sensors, 19(16), 3578. https://doi.org/10.3390/s19163578





