The Landscape of Genetic Variation and Disease Risk in Romania: A Single-Center Study of Autosomal Recessive Carrier Frequencies and Molecular Variants
Abstract
1. Introduction
2. Results
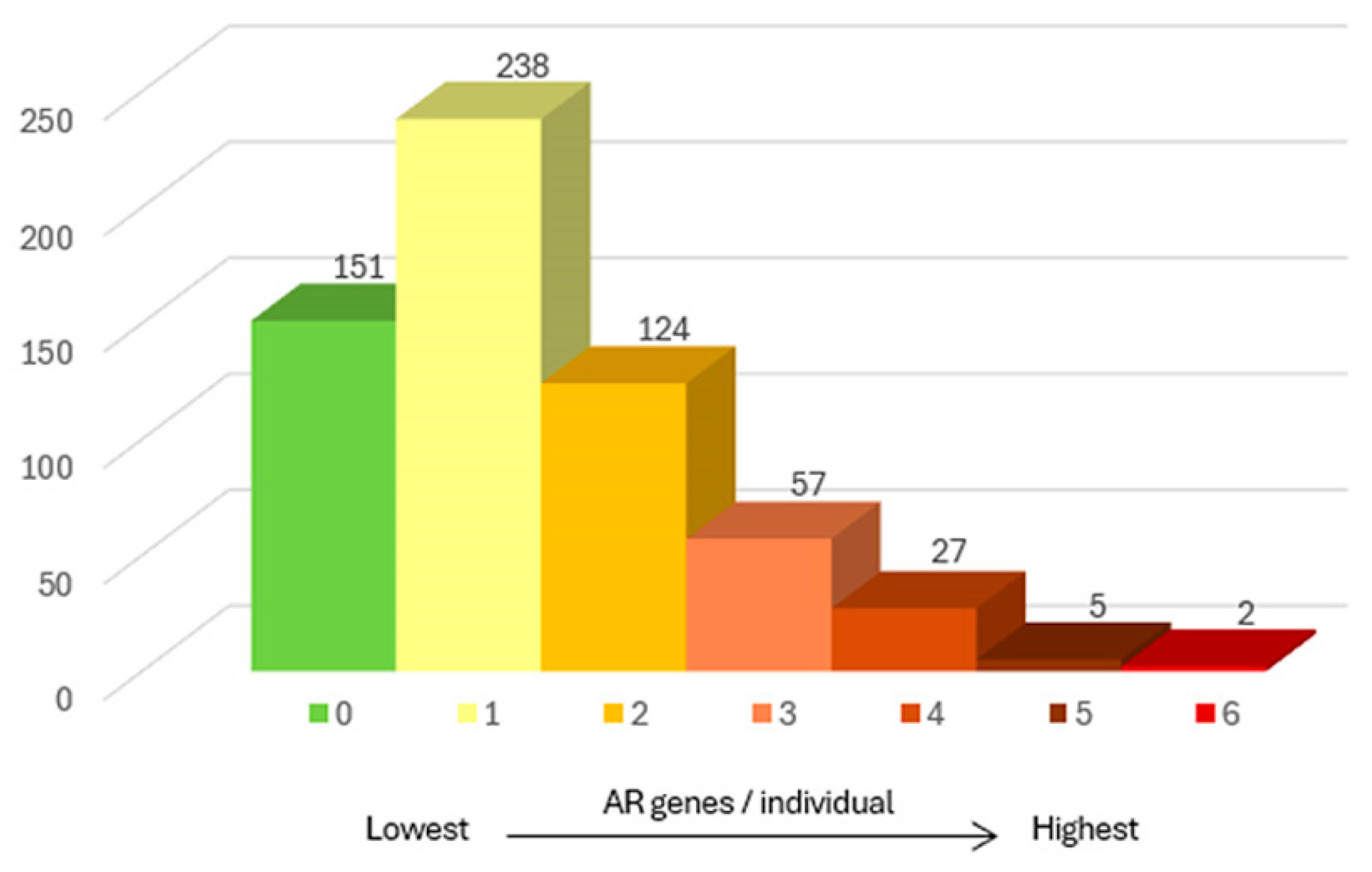
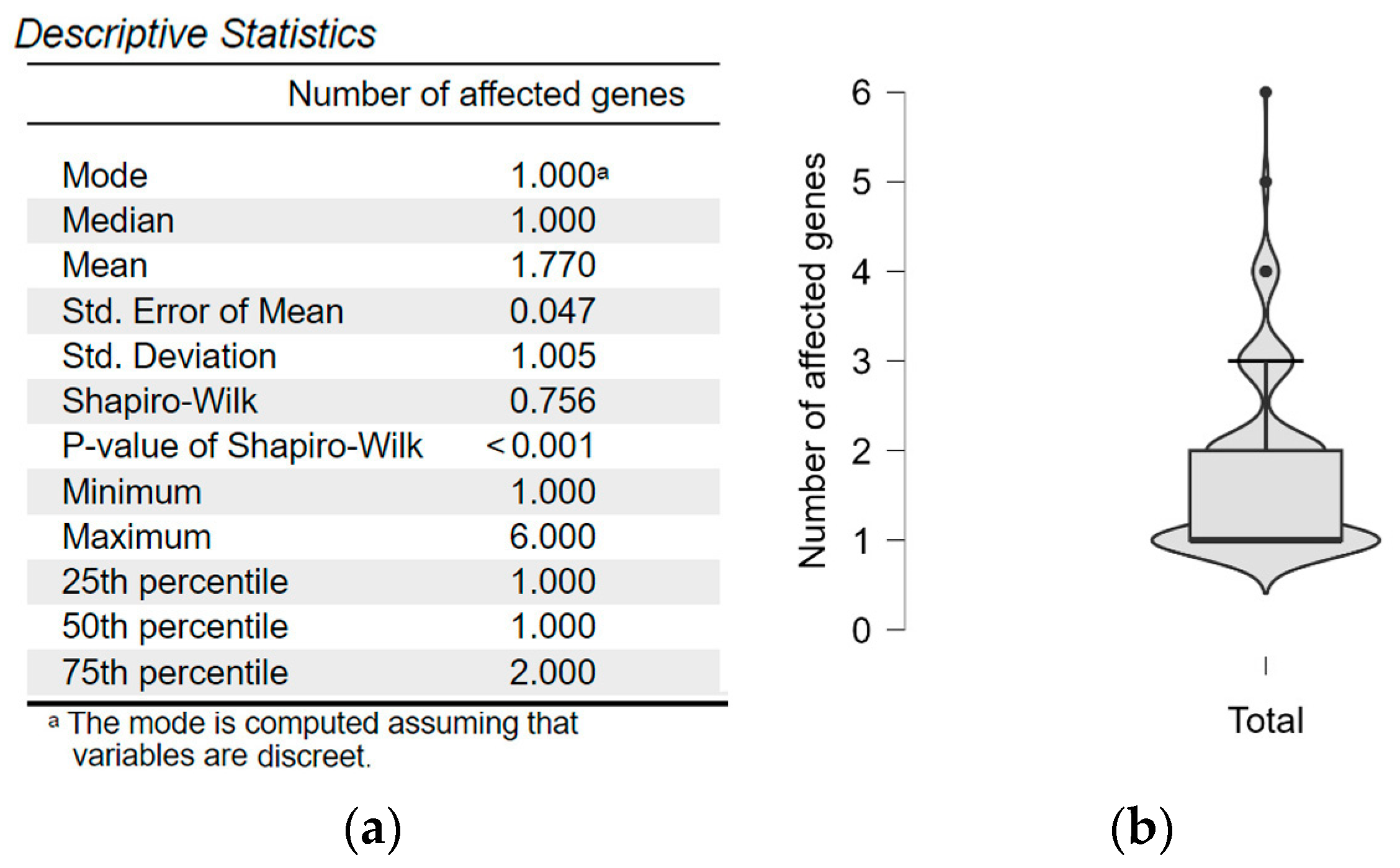
2.1. Gene Frequency
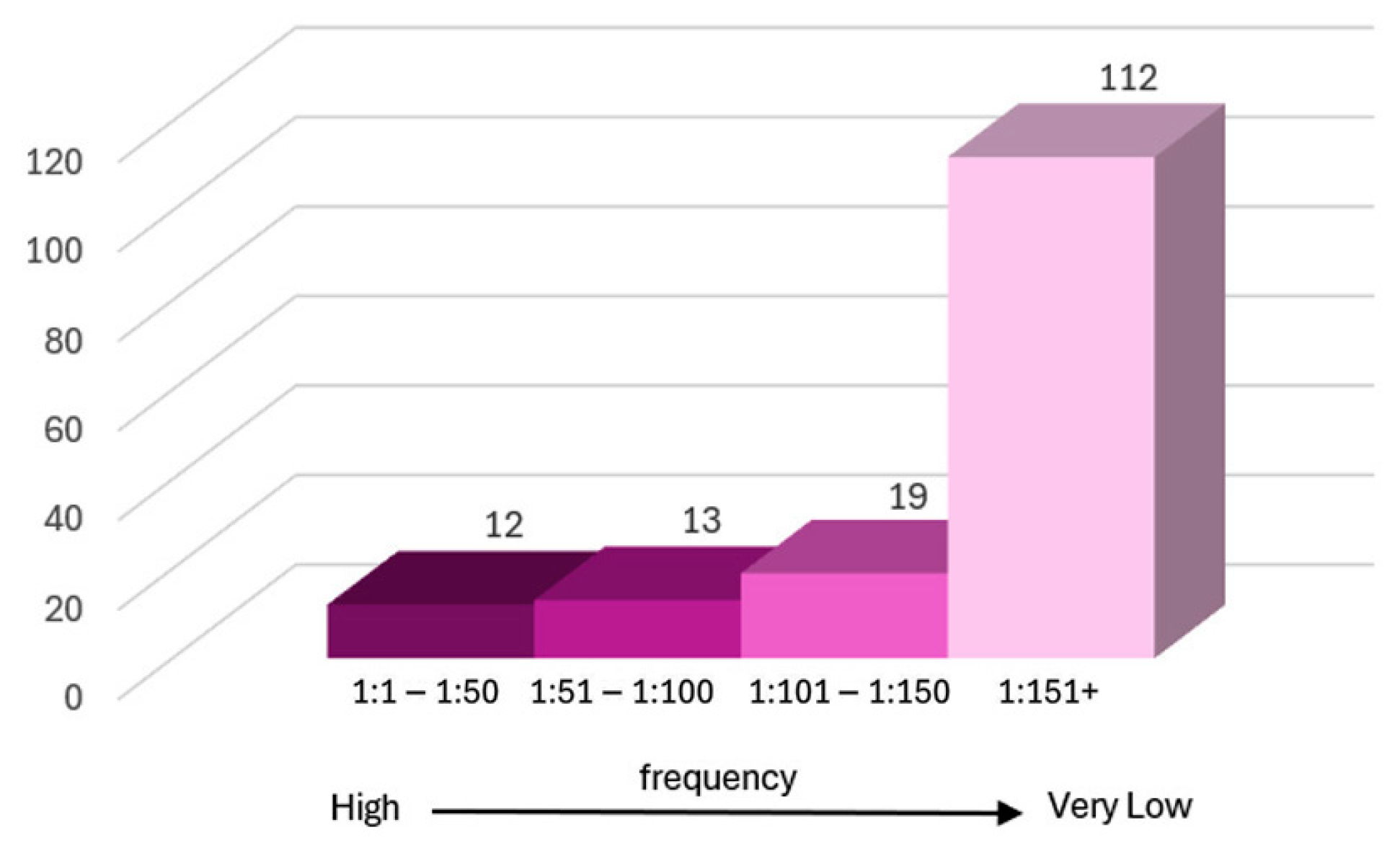
- High frequency (up to 1:50): 12 genes.
- Moderate frequency (between 1:51 and 1:100): 13 genes.
- Low frequency (between 1:101 and 1:150): 19 genes.
- Very low frequency (greater than 1:151): the remaining 112 genes.
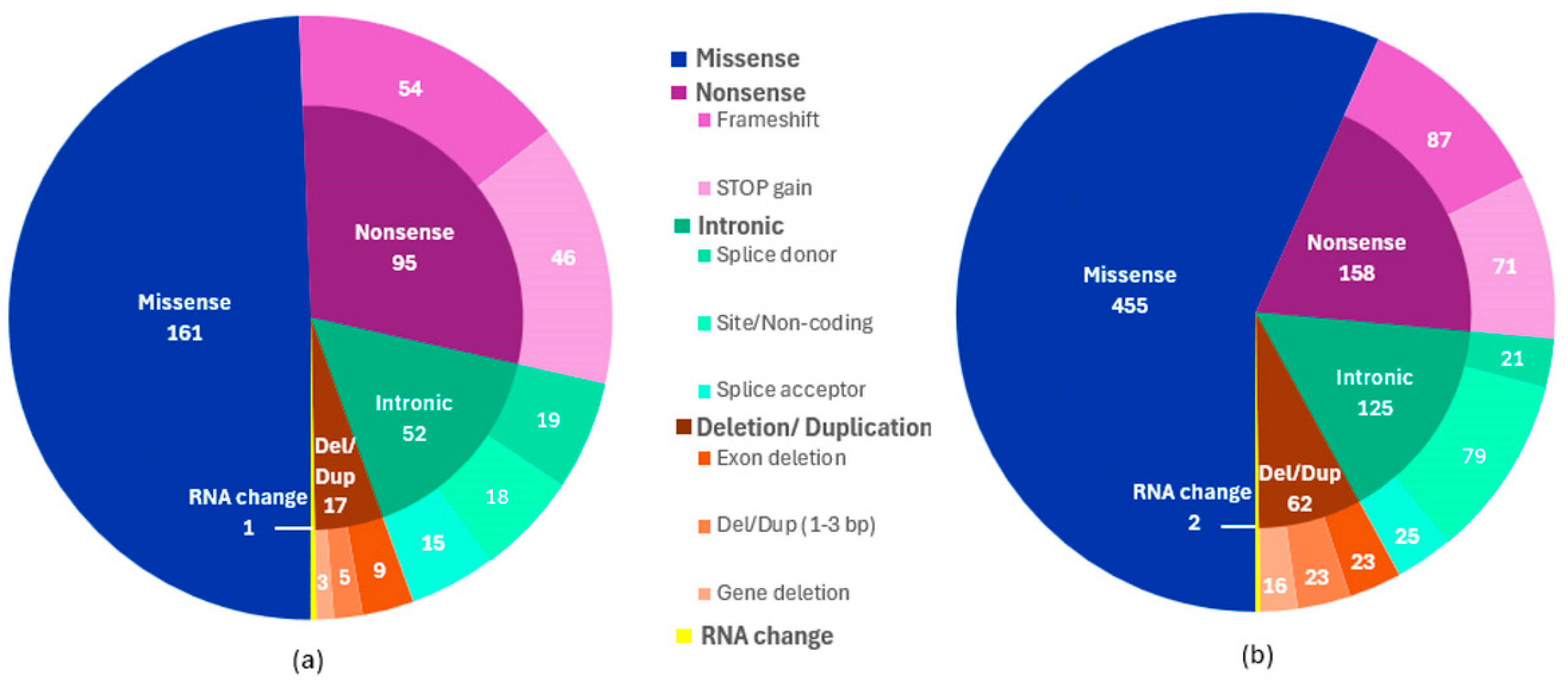
2.2. Type of Mutated Variants Identified
2.3. Gene Rankings
2.4. Allelic Heterogeneity
2.5. Classification of Diseases Risk by Morbidity and Mortality
3. Discussion
4. Materials and Methods
4.1. Selection of Participants and Clinical Data
4.2. Ethical Approval
4.3. Genetic Counseling
4.4. Carrier Screen Test and Panel Genes
4.5. Statistical Analysis of Variant Frequencies
5. Conclusions and Future Perspectives
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ACMG | American College of Medical Genetics and Genomics |
| ACMG-25 | 25-variant CFTR carrier panel (recommended by ACMG) |
| ACOG | American College of Obstetricians and Gynecologists |
| AD | Autosomal dominant |
| AR | Autosomal recessive |
| bp | Base pairs |
| CAVD | Congenital bilateral absence of the vas deferens |
| CF | Cystic fibrosis |
| CFTR | Cystic fibrosis transmembrane conductance regulator |
| CFTR-100 | 100-variant CFTR screening panel |
| DNA | Deoxyribonucleic acid |
| Def | Deficiency |
| gnomAD | Genome Aggregation Database |
| GRCh37 | Genome Reference Consortium Human build 37 |
| IQR | Interquartile range |
| NFE | Non-Finnish European |
| NM | RefSeq mRNA accession prefix used in HGVS nomenclature |
| PAH | Phenylalanine hydroxylase (gene) |
| PKU | Phenylketonuria |
| PLP | Pathogenic or likely pathogenic |
| RO | Romania/Romanian |
| SD | Standard deviation |
| SMA | Spinal muscular atrophy |
| Sy | Syndrome |
| TG | Thymine–guanine repeat tract (CFTR intron 8 polymorphism) |
| UK | United Kingdom |
| US | United States |
| WES | Whole-exome sequencing |
References
- Fridman, H.; Yntema, H.G.; Mägi, R.; Andreson, R.; Metspalu, A.; Mezzavila, M.; Tyler-Smith, C.; Xue, Y.; Carmi, S.; Levy-Lahad, E.; et al. The landscape of autosomal-recessive pathogenic variants in European populations reveals phenotype-specific effects. Am. J. Hum. Genet. 2021, 108, 608–619. [Google Scholar] [CrossRef]
- Muller, H.J. Our load of mutations. Am. J. Hum. Genet. 1950, 2, 111–176. [Google Scholar]
- Morton, N.E.; Crow, J.F.; Muller, H.J. An estimate of the mutational damage in man from data on consanguineous marriages. Proc. Natl. Acad. Sci. USA 1956, 42, 855–863. [Google Scholar] [CrossRef]
- Kondrashov, A.S. Contamination of the genome by very slightly deleterious mutations: Why have we not died 100 times over? J. Theor. Biol. 1995, 175, 583–594. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z.; Waggoner, D.; Stephens, M.; Ober, C.; Przeworski, M. An estimate of the average number of recessive lethal mutations carried by humans. Genetics 2015, 199, 1243–1254. [Google Scholar] [CrossRef] [PubMed]
- Lazarin, G.A.; Haque, I.S.; Nazareth, S.; Iori, K.; Patterson, A.S.; Jacobson, J.L.; Marshall, J.R.; Seltzer, W.K.; Patrizio, P.; Evans, E.A.; et al. An empirical estimate of carrier frequencies for 400+ causal Mendelian variants: Results from an ethnically diverse clinical sample of 23,453 individuals. Genet. Med. 2013, 15, 178–186. [Google Scholar] [CrossRef]
- Deignan, J.L.; Chung, W.K.; Kearney, H.M.; Monaghan, K.G.; Rehder, C.W.; Chao, E.C.; ACMG Laboratory Quality Assurance Committee. Points to consider in the reevaluation and reanalysis of genomic test results: A statement of the American College of Medical Genetics and Genomics (ACMG). Genet. Med. 2019, 21, 1267–1270. [Google Scholar] [CrossRef]
- Antonarakis, S.E.; Chakravarti, A.; Cohen, J.C.; Hardy, J. Mendelian disorders and multifactorial traits: The big divide or one for all? Nat. Rev. Genet. 2010, 11, 380–384. [Google Scholar] [CrossRef]
- ACOG Committee on Genetics. ACOG Committee Opinion No. 442: Preconception and prenatal carrier screening for genetic diseases in individuals of Eastern European Jewish descent. Obstet. Gynecol. 2009, 114, 950–953. [Google Scholar] [CrossRef]
- Grody, W.W.; Thompson, B.H.; Gregg, A.R.; Bean, L.H.; Monaghan, K.G.; Schneider, A.; Lebo, R.V. ACMG position statement on prenatal/preconception expanded carrier screening. Genet. Med. 2013, 15, 482–483. [Google Scholar] [CrossRef]
- Zytsar, M.V.; Barashkov, N.A.; Bady-Khoo, M.S.; Shubina-Olejnik, O.A.; Danilenko, N.G.; Bondar, A.A.; Morozov, I.V.; Solovyev, A.V.; Danilchenko, V.Y.; Maximov, V.N.; et al. Updated carrier rates for c.35delG (GJB2) associated with hearing loss in Russia and common c.35delG haplotypes in Siberia. BMC Med. Genet. 2018, 19, 138. [Google Scholar] [CrossRef] [PubMed]
- Waters, D.; Adeloye, D.; Woolham, D.; Wastnedge, E.; Patel, S.; Rudan, I. Global birth prevalence and mortality from inborn errors of metabolism: A systematic analysis of the evidence. J. Glob. Health 2018, 8, 021102. [Google Scholar] [CrossRef]
- Dima, V. Actualities in neonatal endocrine and metabolic screening. Acta Endocrinol. 2021, 17, 416–421. [Google Scholar] [CrossRef]
- Dima, V. Newborn Screening in Romania–Present and Future. Rom. J. Prev. Med. 2022, 1, 20–26. [Google Scholar] [CrossRef]
- Qiao, L.; Ge, J.; Li, C.; Liu, Y.; Hu, C.; Hu, S.; Li, W.; Li, T. Pathogenic gene variation spectrum and carrier screening for Wilson’s disease in Qingdao area. Mol. Genet. Genom. Med. 2021, 00, e1741. [Google Scholar] [CrossRef]
- Coppin, H.; Bensaid, M.; Fruchon, S.; Borot, N.; Blanché, H.; Roth, M.P. Longevity and carrying the C282Y mutation for haemochromatosis on the HFE gene: Case control study of 492 French centenarians. BMJ 2003, 327, 132–133. [Google Scholar] [CrossRef]
- Garewal, G.; Das, R.; Ahluwalia, J.; Marwaha, R.K. Prevalence of the H63D mutation of the HFE in north India: Its presence does not cause iron overload in beta thalassemia trait. Eur. J. Haematol. 2005, 74, 333–336. [Google Scholar] [CrossRef] [PubMed]
- De Boeck, K.; Amaral, M.D. Classification of CFTR mutation classes—Authors’ reply. Lancet Respir. Med. 2016, 4, e39. [Google Scholar] [CrossRef] [PubMed]
- Cutting, G. Cystic fibrosis genetics: From molecular understanding to clinical application. Nat. Rev. Genet. 2015, 16, 45–56. [Google Scholar] [CrossRef]
- Claustres, M.; Thèze, C.; des Georges, M.; Baux, D.; Girodon, E.; Bienvenu, T.; Audrezet, M.P.; Dugueperoux, I.; Férec, C.; Lalau, G.; et al. CFTR-France, a national relational patient database for sharing genetic and phenotypic data associated with rare CFTR variants. Hum. Mutat. 2017, 38, 1297–1315. [Google Scholar] [CrossRef]
- Bareil, C.; Bergougnoux, A. CFTR gene variants, epidemiology and molecular pathology. Arch. Pediatr. 2020, 27 (Suppl. 1), eS8–eS12. [Google Scholar] [CrossRef]
- Swango, K.L.; Demirkol, M.; Hüner, G.; Pronicka, E.; Sykut-Cegielska, J.; Schulze, A.; Mayatepek, E.; Wolf, B. Partial biotinidase deficiency is usually due to the D444H mutation in the biotinidase gene. Hum. Genet. 1998, 102, 571–575. [Google Scholar] [CrossRef]
- Karaca, M.; Özgül, R.K.; Ünal, Ö.; Yücel-Yılmaz, D.; Kılıç, M.; Hişmi, B.; Tokatlı, A.; Coşkun, T.; Dursun, A.; Sivri, H.S. Detection of biotinidase gene mutations in Turkish patients ascertained by newborn and family screening. Eur. J. Pediatr. 2015, 174, 1077–1084. [Google Scholar] [CrossRef]
- Çıkı, K.; Alavanda, C.; Ceylan, E.İ.; Tanyalçın, T.; Kılavuz, S. Comprehensive analysis of genotypic and phenotypic characteristics of biotinidase deficiency patients in the eastern region of Türkiye. Turk. J. Pediatr. 2024, 66, 608–617. [Google Scholar] [CrossRef]
- Almenabawy, N.; Bahl, S.; Ostlund, A.L.; Ghai-Jain, S.; Sosova, I.; Chan, A.; Mercimek-Andrews, S. Clinical and biochemical phenotypes, genotypes, and long-term outcomes of individuals with galactosemia type I from a single metabolic genetics center in Alberta. Mol. Genet. Metab. Rep. 2024, 38, 101055. [Google Scholar] [CrossRef]
- Baş, F.; Kayserili, H.; Darendeliler, F.; Uyguner, O.; Günöz, H.; Apak, M.Y.; Atalar, F.; Bundak, R.; Wilson, R.C.; New, M.I.; et al. CYP21A2 gene mutations in congenital adrenal hyperplasia: Genotype-phenotype correlation in Turkish children. J. Clin. Res. Pediatr. Endocrinol. 2009, 1, 116–128. [Google Scholar] [CrossRef]
- Robichaud, P.P.; Allain, E.P.; Belbraouet, S.; Bhérer, C.; Mamelona, J.; Harquail, J.; Crapoulet, S.; Crapoulet, N.; Bélanger, M.; Ben Amor, M. Pathogenic variants carrier screening in New Brunswick: Acadians reveal high carrier frequency for multiple genetic disorders. BMC Med. Genom. 2022, 15, 98. [Google Scholar] [CrossRef] [PubMed]
- Hendrickson, B.C.; Donohoe, C.; Akmaev, V.R.; Sugarman, E.A.; Labrousse, P.; Boguslavskiy, L.; Flynn, K.; Rohlfs, E.M.; Walker, A.; Allitto, B.; et al. Differences in SMN1 allele frequencies among ethnic groups within North America. J. Med. Genet. 2009, 46, 641–644. [Google Scholar] [CrossRef] [PubMed]
- Sandahl, T.D.; Laursen, T.L.; Munk, D.E.; Vilstrup, H.; Weiss, K.H.; Ott, P. The prevalence of Wilson’s disease: An update. Hepatology 2020, 71, 722–732. [Google Scholar] [CrossRef] [PubMed]
- Collet, C.; Laplanche, J.L.; Page, J.; Morel, H.; Woimant, F.; Poujois, A. High genetic carrier frequency of Wilson’s disease in France: Discrepancies with clinical prevalence. BMC Med. Genet. 2018, 19, 143. [Google Scholar] [CrossRef]
- Guazzarotti, L.; Tadini, G.; Mancini, G.E.; Sani, I.; Pisanelli, S.; Galderisi, F.; D’Auria, E.; Secondi, R.; Bottero, A.; Zuccotti, G.V. WNT10A gene is the second molecular candidate in a cohort of young Italian subjects with ectodermal derivative impairment (EDI). Clin. Genet. 2018, 93, 693–698. [Google Scholar] [CrossRef] [PubMed]
- Quinn, S.; Walsh, N.; Streata, I.; Ververi, A.; Kulshrestha, S.; Puri, R.D.; Riza, A.L.; Walsh, A.; Gorman, K.; Crushell, E.; et al. Catalogue of inherited autosomal recessive disorders found amongst the Roma population of Europe. Eur. J. Med. Genet. 2025, 73, 104989. [Google Scholar] [CrossRef] [PubMed]
- Karczewski, K.J.; Francioli, L.C.; Tiao, G.; Cummings, B.B.; Alföldi, J.; Wang, Q.; Collins, R.L.; Laricchia, K.M.; Ganna, A.; Birnbaum, D.P.; et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 2020, 581, 434–443, Correction in Nature 2021, 590, E53. [Google Scholar] [CrossRef]
- American College of Obstetricians and Gynecologists; American College of Medical Genetics. Preconception and Prenatal Carrier Screening for Cystic Fibrosis. Clinical and Laboratory Guidelines; American College of Obstetricians and Gynecologists: Washington, DC, USA, 2001. [Google Scholar]
- Gug, M.; Rațiu, A.; Andreescu, N.; Farcaș, S.; Laitin, S.; Gug, C. Approach and Management of Pregnancies with Risk Identified by Non-Invasive Prenatal Testing. J. Pers. Med. 2024, 14, 366. [Google Scholar] [CrossRef] [PubMed]
- Gug, C.; Mozos, I.; Ratiu, A.; Tudor, A.; Gorduza, E.V.; Caba, L.; Gug, M.; Cojocariu, C.; Furau, C.; Furau, G.; et al. Genetic Counseling and Management: The First Study to Report NIPT Findings in a Romanian Population. Medicina 2022, 58, 79. [Google Scholar] [CrossRef]
- Gurdasani, D.; Barroso, I.; Zeggini, E.; Sandhu, M.S. Genomics of disease risk in globally diverse populations. Nat. Rev. Genet. 2019, 20, 520–535, Erratum in Nat. Rev. Genet. 2019, 20, 562. [Google Scholar] [CrossRef]
- The Mediterranean Founder Mutation Database. Available online: http://mfmd.pasteur.ma/index.php?genestart=W (accessed on 9 October 2025).
- Available online: https://gnomad.broadinstitute.org/ (accessed on 6 November 2025).
- Lincoln, S.E.; Truty, R.; Lin, C.F.; Zook, J.M.; Paul, J.; Ramey, V.H.; Salit, M.; Rehm, H.L.; Nussbaum, R.L.; Lebo, M.S. A Rigorous Interlaboratory Examination of the Need to Confirm Next-Generation Sequencing-Detected Variants with an Orthogonal Method in Clinical Genetic Testing. J. Mol. Diagn. 2019, 21, 318–329. [Google Scholar] [CrossRef]
| Disorder | Nomenclature | Inheritance | Gene | West RO 1 Carrier Frequency | Non-Finnish European Carrier Frequency |
|---|---|---|---|---|---|
| Hereditary hemochromatosis type 1 | NM 2_000410.3 | AR 3 | HFE | 1:5 | 1:6 |
| CFTR-related conditions | NM_000492.3 | AR | CFTR | 1:9 | 1:9 |
| Biotinidase deficiency | NM_000060.3 | AR | BTD | 1:16 | 1: 25 |
| GJB2-related conditions | NM_004004.5 | AR | GJB2 | 1:17 | 1: 42 |
| Galactosemia (GALT-related) | NM_000155.3 | AR | GALT | 1:19 | 1:19 |
| Congenital adrenal hyperplasia due to 21-hydroxylase deficiency | NM_000500.7 | AR | CYP21A2 | 1:19 | 1:17 |
| Alpha-1 antitrypsin deficiency | NM_000295.4 | AR | SERPINA1 | 1:26 | 1: 18 |
| Phenylalanine hydroxylase deficiency | NM_000277.1 | AR | PAH | 1:27 | 1: 50 |
| Spinal muscular atrophy | NM_000344.3 | AR | SMN1 | 1:30 | 1: 45 |
| Wilson disease (AR) | NM_000053.3 | AR | ATP7B | 1:36 | 1:50 |
| USH2A-related conditions | NM_206933.2 | AR | USH2A | 1:43 | 1: 70 |
| WNT10A-related conditions | NM_025216.2 | AR | WNT10A | 1:46 | 1:33 |
| Gene | Variants 1 | Number of Individuals | Variant Frequency in the Cohort (%) |
|---|---|---|---|
| HFE | c.187C>G (p.His63Asp) (H63D) | 100 | 12.47 |
| BTD | c.1330G>C (p.Asp444His) | 36 | 4.48 |
| CFTR | c.1210-34TG[11]T[5] (Intronic) | 27 | 3.37 |
| GALT | c.-119_-116del (intronic) | 25 | 3.12 |
| GJB2 | c.35del (p.Gly12Valfs*2) | 21 | 2.62 |
| HFE | c.845G>A (p.Cys282Tyr) § 2 = C282Y | 20 | 2.49 |
| CFTR | c.1521_1523del (p.Phe508del) (F508del) | 19 | 2.37 |
| SMN1 | Exon 7 + 8 deletion | 14 | 1.75 |
| WNT1OA | c.682T>A (p.Phe228Ile) | 13 | 1.62 |
| CYP21A2 | c.1360C>T (p.Pro454Ser) | 10 | 1.24 |
| HBA1 | Deletion (Entire coding sequence) | 9 | 1.12 |
| Gene | Variant 1 | Legacy Name | Location | Number of Individuals |
|---|---|---|---|---|
| CFTR | c.1210-34TG[11]T[5] | - | Intronic | 27 |
| c.1521_1523del (p.Phe508del) | F508del | Exonic | 19 | |
| c.1408A>G (p.Met470Val) | M470V | Exonic | 5 | |
| c.1210-34TG[12]T[5] | - | Intronic | 3 | |
| c.1807G>A (p.Val603Ile) | V603I | Exonic | 2 | |
| c.3472 C>G (Arg1158*) | R1158X | Exonic | 2 | |
| c.3909C>G (p.Asn1303Lys) | N1303K | Exonic | 2 | |
| c.377G>A (p.Gly126Asp) | G126D | Exonic | 1 | |
| c.1210-7_1210-6del | - | Intronic | 1 | |
| c.1210-11delinsGTG | - | Intronic | 1 | |
| c.1624G>T (p.Gly542*) | G542X | Exonic | 1 | |
| c.2813T>G (p.Val938Gly) | V938G | Exonic | 1 | |
| c.3846G>A (p.Trp1282*) | W1282X | Exonic | 1 | |
| PAH | c.1222C>T (p.Arg408Trp) | R408W | Exonic | 5 |
| c.898G>T (p.Ala300Ser) | A300S | Exonic | 4 | |
| c.143T>C (p.Leu48Ser) | L48S | Exonic | 2 | |
| c.673C>A (p.Pro225Thr) | P225T | Exonic | 2 | |
| c.529G>C (p.Val177Leu) | V177L | Exonic | 1 | |
| c.533A>G (p.Glu178Gly) | E178G | Exonic | 1 | |
| c.545A>G (p.Glu182Gly) | E182G | Exonic | 1 | |
| c.734T>C (p.Val245Ala) | V245A | Exonic | 1 | |
| c.844G>T (p.Val282Leu) | V282L | Exonic | 1 | |
| c.1066-11G>A | - | Intronic | 1 | |
| c.1208C>T (p.Ala403Val) | A403V | Exonic | 1 | |
| c.1315 + 1G>A | - | Intronic | 1 | |
| USH2A | c.11864G>A (p.Trp3955*) | W3955* | Exonic | 2 |
| c.12332C>T (p.Ser4111Phe) | S4111F | Exonic | 2 | |
| c.2296T>C (p.Cys766Arg) | C766R | Exonic | 1 | |
| c.2802T>G (p.Cys934Trp) | C934W | Exonic | 1 | |
| c.6937G>T (p.Gly2313Cys) | G2313C | Exonic | 1 | |
| c.7524del (p.Arg2509Glyfs*19) | R2509G | Exonic | 1 | |
| c.8618T>G (p.Leu2873*) | L2873* | Exonic | 1 | |
| c.8682-9A>G | - | Intronic | 1 | |
| c.10073G>A (p.Cys3358Tyr) | C3358Y | Exonic | 1 | |
| c.12268C>A (p.Pro4090Thr) | P4090T | Exonic | 1 | |
| c.12569T>C (p.Val4190Ala) | V4190A | Exonic | 1 | |
| c.14803C>T (p.Arg4935*) | R4935* | Exonic | 1 | |
| ATP7B | c.2817G>T (p.Trp939Cys) | W939C | Exonic | 4 |
| c.3207C>A (p.His1069Gln) | H1069Q | Exonic | 4 | |
| c.19_20del (p.Gln7Aspfs*14) | Q7D | Exonic | 2 | |
| c.347T>C (p.Ile116Thr) | I116T | Exonic | 2 | |
| c.1877G>C (p.Gly626Ala) | G626A | Exonic | 1 | |
| c.2305A>G (p.Met769Val) | M769V | Exonic | 1 | |
| c.2532delA (p.Val845Serfs*28) | V845S | Exonic | 1 | |
| c.2605G>A (p.Gly869Arg) | G869R | Exonic | 1 | |
| c.2906G>A (p.Arg969Gln) | R969Q | Exonic | 1 | |
| CYP21A2 | c.1360C>T (p.Pro454Ser) | P454S | Exonic | 10 |
| c.844G>T (p.Val282Leu) | V282L | Exonic | 6 | |
| c.955C>T (p.Gln319*) | Q319* | Exonic | 5 | |
| c.293-13C>G | - | Intronic | 4 | |
| c.332_339del (p.Gly111Valfs*21) | G111V | Exonic | 2 | |
| c.188A>T (p.His63Leu) | H63L | Exonic | 1 | |
| c.1069C>T (p.Arg357Trp) | R357W | Exonic | 1 |
| Gene Frequency in Western Romania | Gene | Physiological System/Type | Disease | Observations Related to Morbidity and Mortailty |
|---|---|---|---|---|
| High frequency | HFE | Metabolic | Hemochromatosis | Variable severity; organ damage if untreated. |
| CFTR | Metabolic Pulmonary | Cystic fibrosis | Severe disease; early mortality without treatment. | |
| BTD | Metabolic | Biotinidase deficiency | Treatable; untreated may cause neurological symptoms. | |
| GJB2 | Sensory (Hearing/Skin) | Vohwinkel syndrome/ Keratitis-ichthyosis-deafness | Non-fatal; significant sensory impact. | |
| CYP21A2 | Endocrine | Congenital adrenal hyperplasia | Potential neonatal mortality without therapy. | |
| GALT | Metabolic | Classical galactosemia | Lethal in neonatal form if untreated. | |
| SERPINA1 | Liver Lung | Alpha-1 antitrypsin deficiency | Early emphysema or liver failure; variable course. | |
| PAH | Metabolic | Phenylketonuria | Non-fatal with treatment; untreated causes severe disability. | |
| SMN1 | Neuromuscular | Spinal muscular atrophy | Infantile forms fatal; treatable with gene therapy. | |
| ATB7B | Metabolic | Wilson’s disease | Fatal disease if not diagnosed and treated (copper accumulation in liver/brain). | |
| USH2A | Sensory (Vision/Hearing) | Usher syndrome type II | Non-fatal; dual sensory impairment. | |
| WNT10A | Craniodental Skin | Ectodermal dysplasia | Non-lethal; impacts quality of life. | |
| Moderate frequency | ACADM | Metabolic | Medium-chain acyl-CoA dehydrogenase deficiency | Infant mortality risk if undiagnosed. |
| ALDOB | Metabolic | Hereditary fructose intolerance | Severe hypoglycemia in infancy if untreated; treatable by dietary restriction. | |
| DHCR7 | Metabolic Developmental | Smith–Lemli–Opitz syndrome | Lethal in severe forms; survivable with cholesterol supplementation. | |
| GAA | Metabolic Neuromuscular | Pompe disease | Infantile form lethal; treatable. | |
| HBA1 | Hematologic | Alpha-thalassemia | Hydrops fetalis is lethal; trait forms are mild. | |
| EVC | Skeletal Growth | Ellis–van Creveld syndrome | Neonatal lethal forms. | |
| SLC26A2 | Skeletal Growth | Diastrophic dysplasia | Severe skeletal dysplasia; perinatal lethal variants exist. | |
| TPP1 | Neurodegenerative | Neuronal ceroid lipofuscinosis type 2 | Early-onset neurodegeneration; fatal in childhood. | |
| COL7A1 | Skin Connective tissue | Dystrophic epidermolysis bullosa | Severe forms fatal in childhood. | |
| CYP11B2 | Endocrine | Aldosterone synthase deficiency | Can cause neonatal salt-wasting; treatable. | |
| GBA1 | Metabolic Lysosomal | Gaucher disease | Infantile form lethal; chronic forms manageable. | |
| NEB | Neuromuscular | Nemaline myopathy | Severe neonatal forms fatal; variable severity. | |
| NR2E3 | Sensory (Vision) | Neural ceroid lipofuscinosis | Non-lethal; causes visual impairment. | |
| Low frequency | ACAD9 | Metabolic Mitochondrial | ACAD9 deficiency | Variable severity; can cause cardiomyopathy and early death if untreated. |
| BBS1 | Multisystem Developmental | Bardet–Biedl syndrome | Non-lethal; multisystem disorder affecting vision, obesity, and kidneys. | |
| CAPN3 | Neuromuscular | Limb-girdle muscular dystrophy type 2A | Progressive; may shorten lifespan. | |
| GALC | Neurodegenerative | Krabbe disease | Infantile form is fatal. | |
| SLC12A3 | Renal Electrolyte | Gitelman syndrome | Non-lethal; chronic electrolyte imbalance manageable with therapy. | |
| SLC22A5 | Metabolic | Primary carnitine deficiency | Potentially fatal cardiac involvement if untreated; treatable with supplementation. | |
| ARSA | Neurodegenerative | Metachromatic leukodystrophy | Lethal infantile form. | |
| CPT2 | Metabolic Neuromuscular | CPT II deficiency | Neonatal form lethal; adult form benign. | |
| CRB1 | Sensory (Vision) | Retinal dystrophy Leber congenital amaurosis | Non-fatal; severe vision loss early in life. | |
| EYS | Sensory (Vision) | Retinitis pigmentosa | Non-fatal; progressive blindness. | |
| G6PD | Metabolic Hematologic | G6PD deficiency | Hemolytic anemia; rarely fatal if managed. | |
| GBE1 | Metabolic | Glycogen storage disease type IV (Andersen disease) | Hepatic and neuromuscular forms; infantile form is often fatal. | |
| HEXA | Neurodegenerative | Tay–Sachs disease | Fatal in childhood. | |
| LAMA2 | Neuromuscular | LAMA2 muscular dystrophy | Congenital forms severe and lethal. | |
| LDLR | Metabolic Cardiovascular | Familial hypercholesterolemia | Premature cardiovascular disease; treatable with statins. | |
| LIPA | Metabolic | Lysosomal acid lipase deficiency | Wolman disease lethal in infancy. | |
| MEFV | Inflammatory Autoinflammatory | Familial Mediterranean fever | Non-fatal with treatment; risk of amyloidosis if untreated. | |
| NPC1 | Neurodegenerative | Niemann–Pick type C | Infantile forms fatal; variable course. | |
| VPS13B | Developmental Neurological | Cohen syndrome | Non-fatal; developmental delay and visual impairment. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gug, M.; Andreescu, N.; Caba, L.; Popoiu, T.-A.; Mozos, I.; Gug, C. The Landscape of Genetic Variation and Disease Risk in Romania: A Single-Center Study of Autosomal Recessive Carrier Frequencies and Molecular Variants. Int. J. Mol. Sci. 2025, 26, 10912. https://doi.org/10.3390/ijms262210912
Gug M, Andreescu N, Caba L, Popoiu T-A, Mozos I, Gug C. The Landscape of Genetic Variation and Disease Risk in Romania: A Single-Center Study of Autosomal Recessive Carrier Frequencies and Molecular Variants. International Journal of Molecular Sciences. 2025; 26(22):10912. https://doi.org/10.3390/ijms262210912
Chicago/Turabian StyleGug, Miruna, Nicoleta Andreescu, Lavinia Caba, Tudor-Alexandru Popoiu, Ioana Mozos, and Cristina Gug. 2025. "The Landscape of Genetic Variation and Disease Risk in Romania: A Single-Center Study of Autosomal Recessive Carrier Frequencies and Molecular Variants" International Journal of Molecular Sciences 26, no. 22: 10912. https://doi.org/10.3390/ijms262210912
APA StyleGug, M., Andreescu, N., Caba, L., Popoiu, T.-A., Mozos, I., & Gug, C. (2025). The Landscape of Genetic Variation and Disease Risk in Romania: A Single-Center Study of Autosomal Recessive Carrier Frequencies and Molecular Variants. International Journal of Molecular Sciences, 26(22), 10912. https://doi.org/10.3390/ijms262210912








