Virus-Induced Gene Silencing Identifies an Important Role of the TaRSR1 Transcription Factor in Starch Synthesis in Bread Wheat
Abstract
:1. Introduction
2. Results
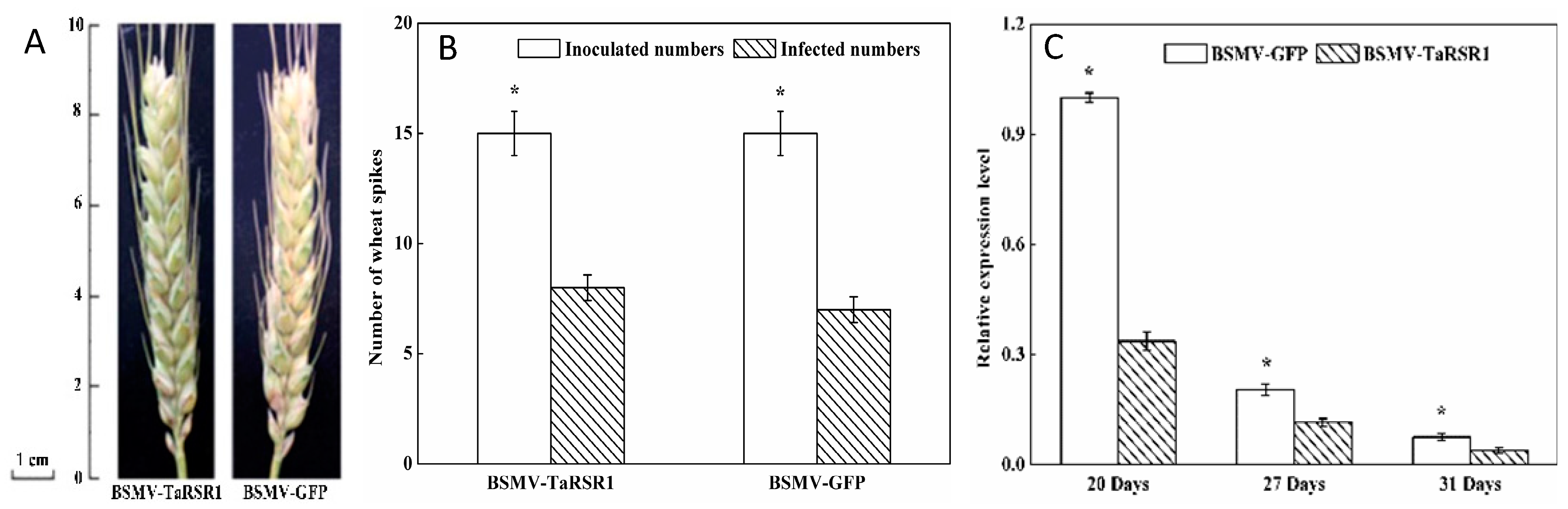
2.1. Phenotypes of Barley Stripe Mosaic Virus-Wheat Starch Regulator 1 (BSMV-TaRSR1) Infected Wheat Plants
2.2. Transcription Levels of TaRSR1 in BSMV-TaRSR1 Infected Wheat Plants
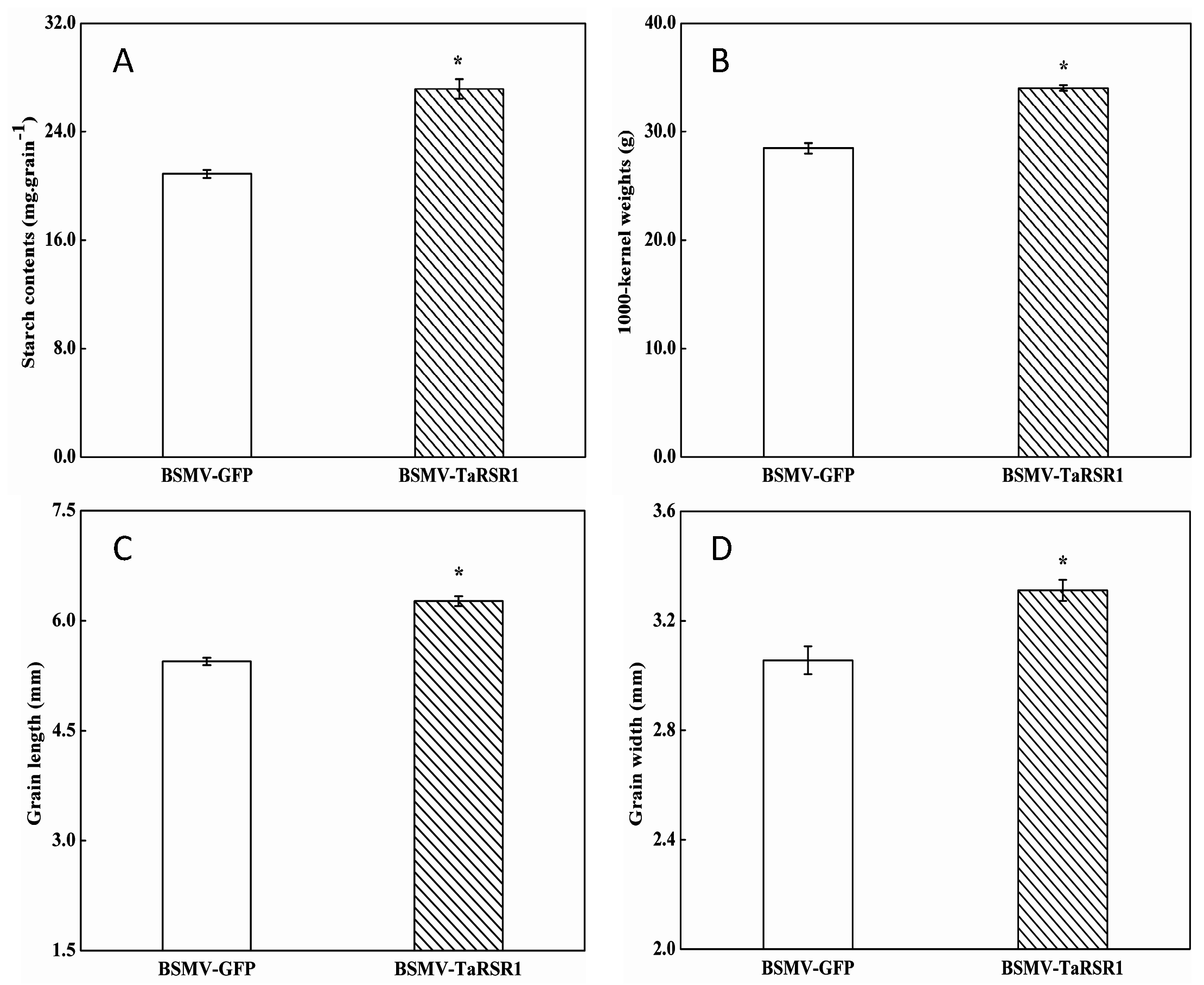
2.3. Starch Contents and One Thousand Kernel (1000-Kernel) Weights in BSMV-TaRSR1 Infected Wheat Plants
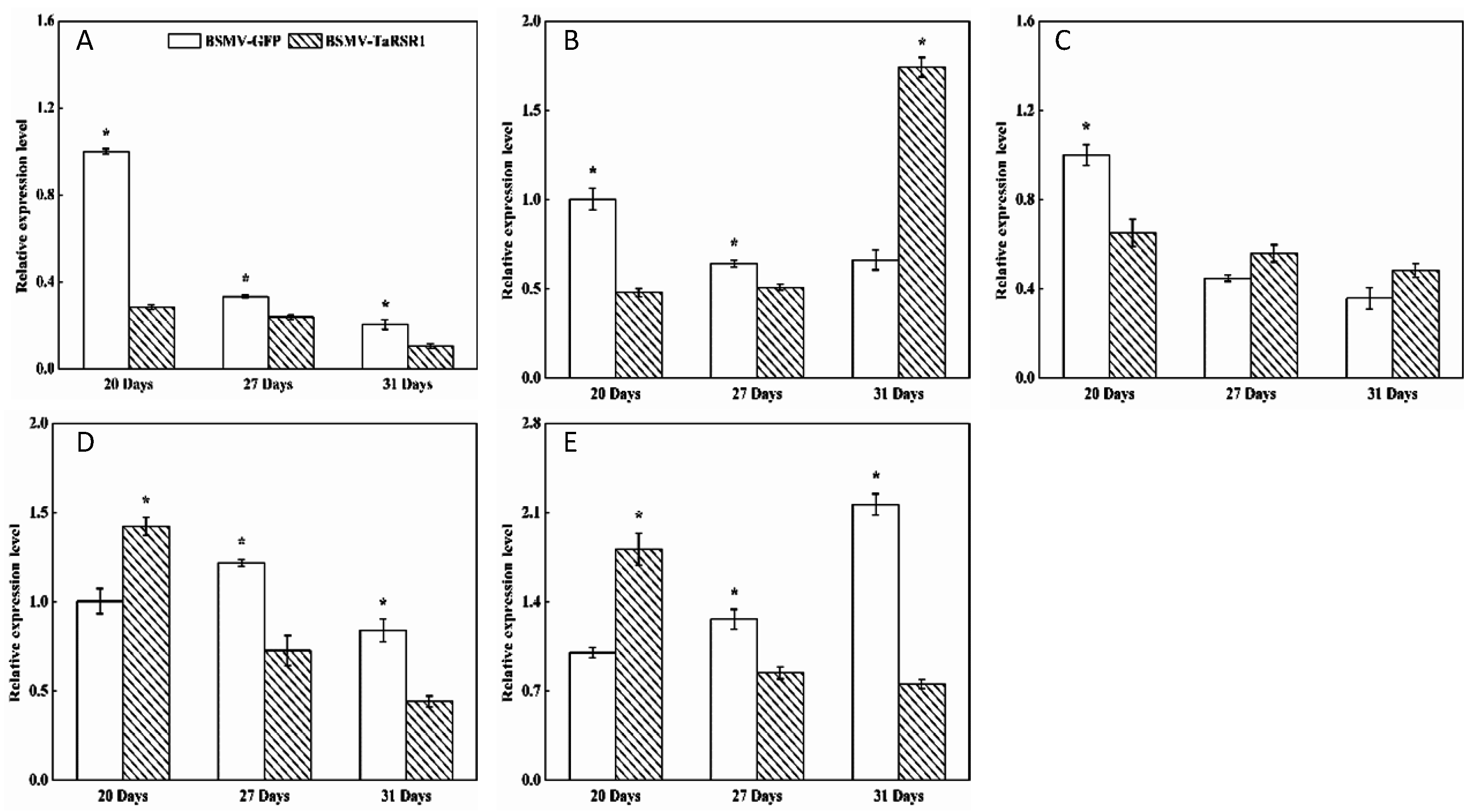
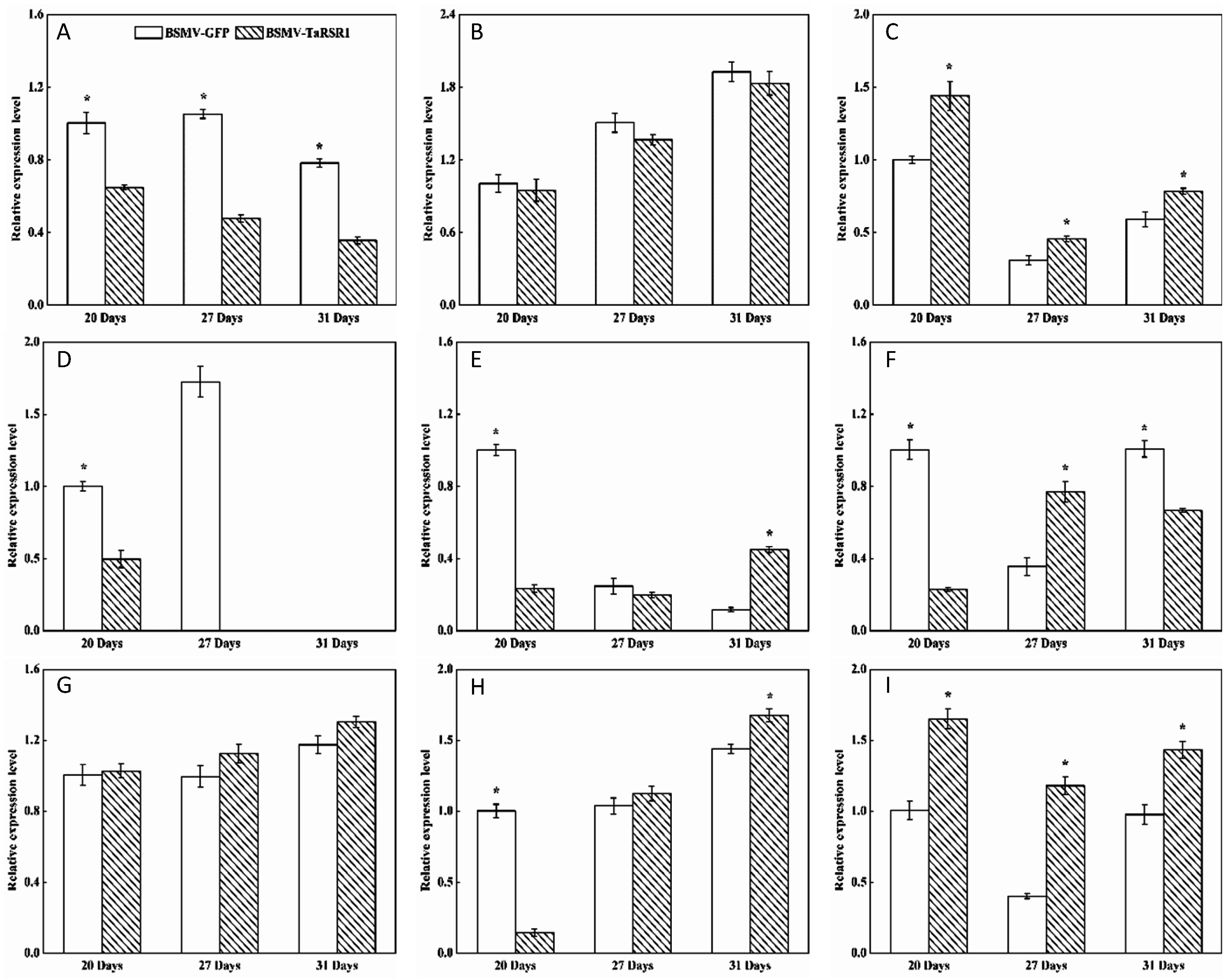
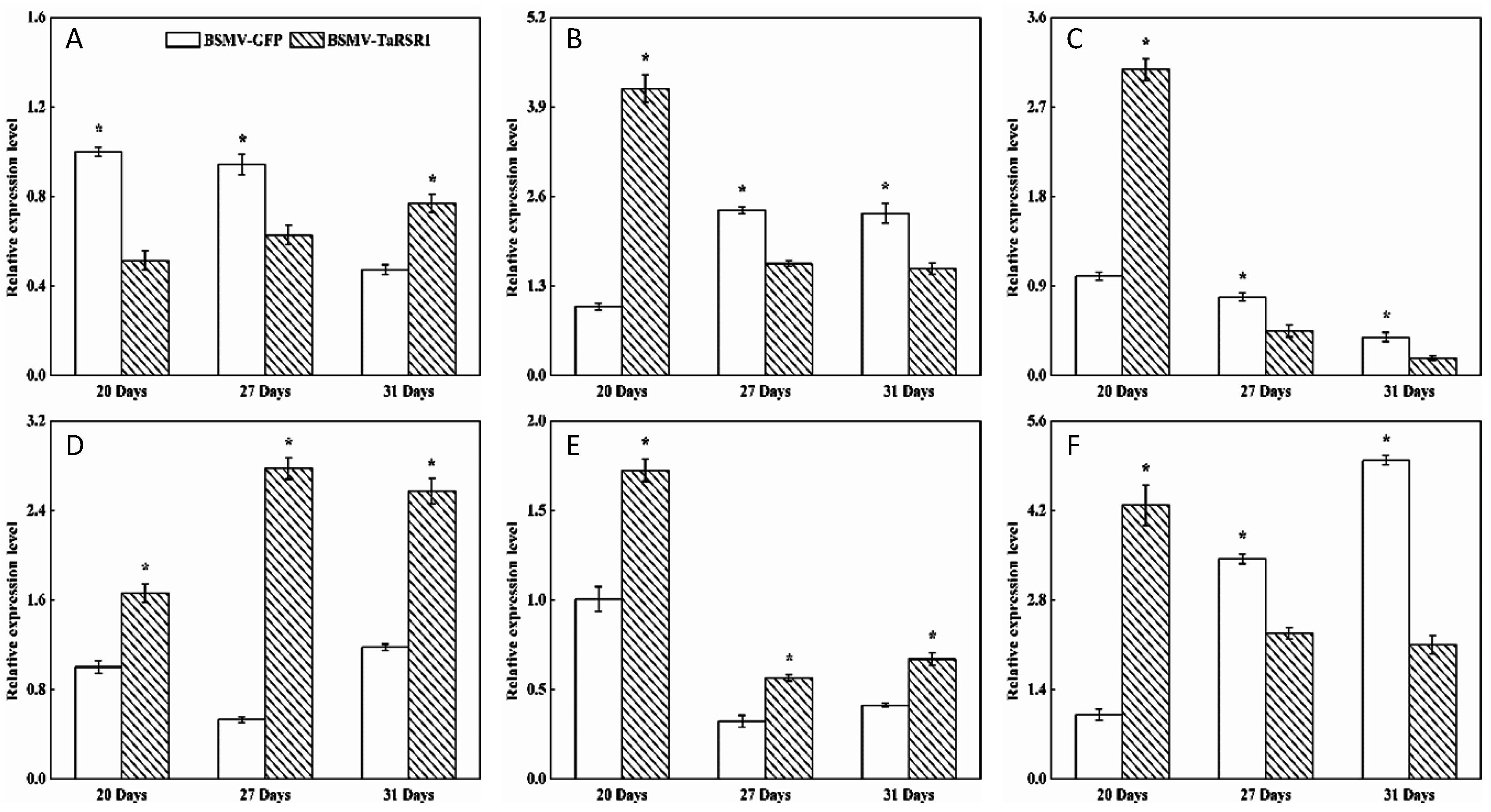
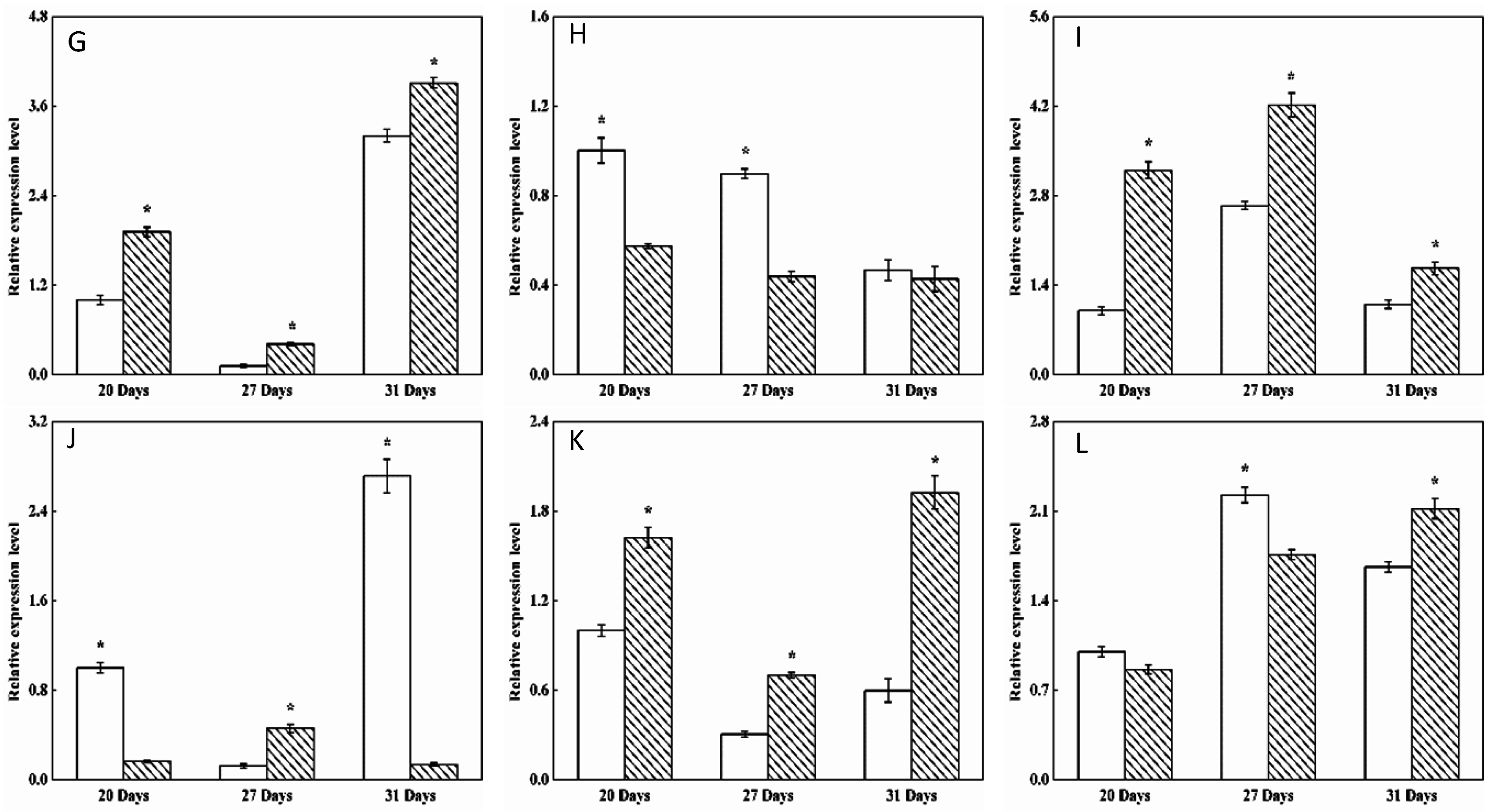
2.4. Transcription Levels of 26 Starch Synthesis-Related Enzyme Genes in BSMV-TaRSR1 Infected Wheat Plants
2.5. Putative Mechanism of TaRSR1 Regulation in Higher Plants
3. Materials and Methods
3.1. Plant Materials
3.2. Construction of BSMV-Derived Vectors
3.3. In Vitro Transcription of Viral RNAs and Plant Inoculation
3.4. Determination of Transcription Levels of TaRSR1 and 26 Starch Synthesis-Related Enzyme Genes
3.5. Assays on Wheat Grains Parameters
3.6. Statistical Analysis
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Stamova, B.S.; Laudencia-Chingcuanco, D.; Beckles, D.M. Transcriptomic analysis of starch biosynthesis in the developing grain of hexaploid wheat. Int. J. Plant Genom. 2009, 2009, 407426. [Google Scholar] [CrossRef] [PubMed]
- Jeon, J.S.; Ryoo, N.; Hahn, T.R.; Walia, H.; Nakamura, Y. Starch biosynthesis in cereal endosperm. Plant Physiol. Biochem. 2010, 48, 383–392. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Chen, Y.; Tong, C.; Huang, Y.; Xu, F.; Li, K.; Corke, H.; Sun, M.; Bao, J. Association mapping of starch physicochemical properties with starch synthesis-related gene markers in nonwaxy rice (Oryza Sativa L.). Mol. Breed. 2014, 34, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Satoh, H.; Shibahara, K.; Tokunaga, T.; Nishi, A.; Tasaki, M.; Hwang, S.K.; Okita, T.W.; Kaneko, N.; Fujita, N.; Yoshida, M.; et al. Mutation of the plastidial a-glucan phosphorylase gene in rice affects the synthesis and structure of starch in the endosperm. Plant Cell 2008, 20, 1833–1849. [Google Scholar] [CrossRef] [PubMed]
- Ohdan, T.; Francisco, P.B.J.; Sawada, T.; Hirose, T.; Terao, T.; Satoh, H.; Nakamura, Y. Expression profiling of genes involved in starch synthesis in sink and source organs of rice. J. Exp. Bot. 2005, 56, 3229–3244. [Google Scholar] [CrossRef] [PubMed]
- Yan, H.B.; Pan, X.X.; Jiang, H.W.; Wu, G.J. Comparison of the starch synthesis genes between maize and rice: Copies, chromosome location and expression divergence. Theor. Appl. Genet. 2009, 119, 815–825. [Google Scholar] [CrossRef] [PubMed]
- Kang, G.; Xu, W.; Liu, G.; Peng, X.; Guo, T. Comprehensive analysis on the transcript levels of starch synthesis genes and RSR1 transcript factor in endosperm of wheat. Genome 2013, 56, 115–122. [Google Scholar] [CrossRef] [PubMed]
- Kang, G.; Peng, X.; Wang, L.; Yang, Y.; Wang, C.; Zhu, Y.; Guo, T. Ultrastructural observation of mesophyll cells and temporal expression profiles of the genes involved in transitory starch metabolism in flag leaves of wheat after anthesis. Physiol. Plant 2015, 153, 12–19. [Google Scholar] [CrossRef] [PubMed]
- Radchuk, V.V.; Borisjuk, L.; Sreenivasulu, N.; Merx, K.; Mock, H.P.; Rolletschek, H.; Wobus, U.; Weschke, W. Spatiotemporal profiling of starch biosynthesis and degradation in the developing barley grain. Plant Physiol. 2009, 150, 190–204. [Google Scholar] [CrossRef] [PubMed]
- Tombuloglu, H.; Kekec, G.; Sakcali, M.S.; Unver, T. Transcriptome-wide identification of R2R3-MYB transcription factors in barley with their boron responsive expression analysis. Mol. Genet. Genom. 2013, 288, 141–155. [Google Scholar] [CrossRef] [PubMed]
- Quan, R.D.; Hu, S.J.; Zhang, Z.L.; Zhang, H.W.; Zhang, Z.J.; Huang, R.F. Overexpression of an ERF transcription factor TSRF1 improves rice drought tolerance. Plant Biotechnol. J. 2010, 8, 476–488. [Google Scholar] [CrossRef] [PubMed]
- Morran, S.; Eini, O.; Pyvovarenko, T.; Parent, B.; Singh, R.; Ismagul, A.; Eliby, S.; Shirley, N.; Langridge, P.; Lopato, S. Improvement of stress tolerance of wheat and barley by modulation of expression of DREB/CBF factor. Plant Biotechnol. J. 2011, 9, 230–249. [Google Scholar] [CrossRef] [PubMed]
- Rong, W.; Qi, L.; Wang, A.; Ye, X.; Du, L.; Liang, H.; Xin, Z.; Zhang, Z. The ERF transcription factor TaERF3 promotes tolerance to salt and drought stresses in wheat. Plant Biotechnol. J. 2014, 12, 468–479. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, M.; Bonnett, D.; Chapman, S.C.; Furbank, R.T.; Manes, Y.; Mather, D.E.; Parry, M.A.J. Raising yield potential of wheat. I. Overview of a consortium approach and breeding strategies. J. Exp. Bot. 2011, 62, 439–453. [Google Scholar] [CrossRef] [PubMed]
- Feldman, M.; Levy, A.A.; Fahima, T.; Korol, A. Genomic asymmetry in allopolyploid plants: Wheat as a model. J. Exp. Bot. 2012, 63, 5045–5059. [Google Scholar] [CrossRef] [PubMed]
- Pfeifer, M.; Kugler, K.G.; Sandve, S.R.; Zhan, B.; Rudi, H.; Hvidsten, T.R.; Mayer, K.F.; Olsen, O.A.; International Wheat Genome Sequencing Consortium. Genome interplay in the grain transcriptome of hexaploid bread wheat. Science 2014, 345, 1250091. [Google Scholar] [CrossRef] [PubMed]
- Flagel, L.E.; Chen, L.; Chaudhary, B.; Wendel, J.F. Coordinated and fine-scale control of homoeologous gene expression in allotetraploid cotton. J. Hered. 2009, 100, 487–490. [Google Scholar] [CrossRef] [PubMed]
- International Wheat Genome Sequencing Consortium. A chromosome-based draft sequence of the hexaploid bread wheat genome. Science 2014, 345, 1251788. [Google Scholar]
- Pingault, L.; Choulet, F.; Alberti, A.; Glover, N.; Wincker, P.; Feuillet, C.; Paux, E. Deep transcriptome sequencing provides new insights into the structural and functional organization of the wheat genome. Genome Biol. 2015, 16, 29. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Soltis, P.S.; Wall, K.; Soltis, D.E. Phylogeny and domain Evolution in the APETALA2-likegenefamily. Mol. Biol. Evol. 2006, 23, 107–120. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.Y.; Lee, J.W.; Moon, S.; Park, S.Y.; An, G. The rice heterochronic gene SUPERNUMERARY BRACT regulates the transition from spikelet meristem to floral meristem. Plant J. 2006, 49, 64–78. [Google Scholar] [CrossRef] [PubMed]
- Fu, F.F.; Xue, H.W. Coexpression analysis identifies rice starch regulator1, a rice AP2/EREBP family transcription factor, as a novel rice starch biosynthesis regulator. Plant Physiol. 2010, 154, 927–938. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Sparks, C.; Amoah, B.; Jones, H.D. Factors influencing successful Agrobacterium-mediated genetic transformation of wheat. Plant Cell Rep. 2003, 21, 659–668. [Google Scholar] [PubMed]
- Zhao, T.; Zhao, S.; Chen, H.; Zhao, Q.; Hu, Z.; Hou, B.; Xia, G. Transgenic wheat frogeny resistant to powdery mildew generated by Agrobacterium inoculumto the basal portion of wheat seedling. Plant Cell Rep. 2006, 25, 1199–1204. [Google Scholar] [CrossRef] [PubMed]
- Bhullar, N.K.; Keller, B. Virus induced gene silencing in wheat: A review. In Advances in Biotechnology; Bentham Science Publishers Ltd.: Hilversum, The Netherlands, 2010; pp. 185–203. [Google Scholar]
- Tufan, H.A.; Stefanato, F.L.; McGrann, G.R.D.; MacCormack, R.; Boyd, L.A. The barley stripe mosaic virus system used for virus-induced gene silencing in cereals differentially affects susceptibility to fungal pathogens in wheat. J. Plant Physiol. 2011, 168, 990–994. [Google Scholar] [CrossRef] [PubMed]
- Pfleeger, T.; Olszyk, D.; Lee, E.H.; Plocher, M. Comparing effects of low levels of herbicides on greenhouse and field-grown potatoes (Solanum tuberosum L.), soybeans (Glycine max L.), and peas (Pisum sativum L.). Environ. Toxicol. Chem. 2011, 30, 455–468. [Google Scholar] [CrossRef] [PubMed]
- Ma, M.; Yan, Y.; Huang, L.; Chen, M.; Zhao, H. Virus-induced gene-silencing in wheat spikes and grains and its application in functional analysis of HMW-GS-encoding genes. BMC Plant Biol. 2012, 12, 141. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Li, H.; Chen, Y.; Zheng, Q.; Li, B.; Li, Z. TaSCL14, a novel wheat (Triticum aestivum L.) GRAS gene, regulates plant growth, photosynthesis, tolerance to photooxidative stress, and senescence. J. Genet. Genom. 2015, 42, 21–32. [Google Scholar] [CrossRef] [PubMed]
- Renner, T.; Bragg, J.; Driscoll, H.E.; Jackson, A.O.; Specht, C.D. Virus-induced gene silencing in the Culinary Ginger (Zingiber officinale): An effective mechanism for down-regulating gene expression in tropical monocots. Mol. Plant 2009, 2, 1084–1094. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, P.M.; Wang, M.B.; Lough, T. Gene silencing as an adaptive defence against viruses. Nature 2001, 411, 834–842. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.S.; Rudd, J.J.; Kanyuka, K. Virus induced gene silencing (VIGS) for functional analysis of wheat genes involved in Zymoseptoria tritici susceptibility and resistance. Fungal Genet. Biol. 2015, 79, 84–88. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.S.; Hammond-Kosack, K.E.; Kanyuka, K. Barley stripe mosaic virus-mediated tools for investigating gene function in cereal plants and their pathogens: Virus-induced gene silencing, host-mediated gene silencing, and virus-mediated overexpression of heterologous proteins. Plant Physiol. 2012, 160, 582–590. [Google Scholar] [CrossRef] [PubMed]
- Scofield, S.R.; Huang, L.; Brandt, A.S.; Gill, B.S. Development of a virus-induced gene-silencing system for hexaploid wheat and its use in functional analysis of the lr21-mediated leaf rust resistance pathway. Plant Physiol. 2005, 138, 2165–2173. [Google Scholar] [CrossRef] [PubMed]
- Liang, J.; Chen, X.; Zhao, H.; Yu, S.; Long, H.; Deng, G.; Pan, Z.; Yu, M. The impacts of BSMV on vegetative growth and water status in hulless barley (Hordeum vulgarevar nudum) in VIGS study. Acta Soc. Bot. Pol. 2015, 84, 43–51. [Google Scholar] [CrossRef]
- Pearce, S.; Tabbita, F.; Cantu, D.; Buffalo, V.; Avni, R.; Vazquez-Gross, H.; Zhao, R.; Conley, C.J.; Distelfeld, A.; Dubcovksy, J. Regulation of Zn and Fe transporters by the GPC1 gene during early wheat monocarpic senescence. BMC Plant Biol. 2014, 14, 368. [Google Scholar] [CrossRef] [PubMed]
- Sato, H.; Todaka, D.; Kudo, M.; Mizoi, J.; Kidokoro, S.; Zhao, Yu.; Shinozaki, K.; Yamaguchi-Shinozaki, K. The Arabidopsis transcriptional regulator DPB3-1 enhances heat stress tolerance without growth retardation in rice. Plant Biotechnol. J. 2016, 1–12. [Google Scholar]
- Wang, J.C.; Xu, H.; Zhu, Y.; Liu, Q.Q.; Cai, X.L. OsbZIP58, a basic leucine zipper transcription factor, regulates starch biosynthesis in rice endosperm. J. Exp. Bot. 2013, 64, 3453–3466. [Google Scholar] [CrossRef] [PubMed]
- Mizoi, J.; Shinozaki, K.; Yamaguchi-Shinozaki, K. AP2/ERF family transcription factors in plant abiotic stress responses. Biochim. Biophys. Acta 2012, 1819, 86–96. [Google Scholar] [CrossRef] [PubMed]
- Nakano, T.; Suzuki, K.; Fujimura, T.; Shinshi, H. Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol. 2006, 140, 411–432. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.L.; Lv, H.; Li, L.; Liu, J.; Mu, S.H.; Li, X.P.; Gao, J. Genome-wide analysis of the AP2/ERF transcription factors family and the expression patterns of DREB genes in Moso Bamboo (Phyllostachys edulis). PLoS ONE 2015, 10, e0126657. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.L. AP2/EREBP: A special transcription factor family in plant. J. Qinghai Norm. Univ. 2006, 3, 80–83, In Chinese with English Abstract. [Google Scholar]
- Gong, W.; He, K.; Covington, M.; Dinesh-Kumar, S.P.; Snyder, M.; Harme, S.L.; Zhu, Y.X.; Deng, X.W. The development of protein microarrays and their applications in DNA-protein and protein-protein interaction analyses of Arabidopsis transcription factors. Mol. Plant 2008, 1, 27–41. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Je, J.Y.; Song, C.; Hwang, J.E.; Lim, C.O. A proximal promoter region of Arabidopsis DREB2C confers tissue-specific expression under heat stress. J. Integr. Plant Biol. 2012, 54, 640–651. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.; Xie, D.L.; Wang, Z.Y.; Hong, M.M. Interaction of rice bZIP protein REB with the 5′-upstream region of both rice sbe1 gene and waxy gene. Chin. Sci. Bull. 2002, 47, 310–314. [Google Scholar] [CrossRef]
- Chen, J.; Yi, Q.; Cao, Y.; Wei, B.; Zheng, L.J.; Xiao, Q.L.; Xie, Y.; Gu, Y.; Li, Y.P.; Huang, H.H.; et al. ZmbZIP91 regulates expression of starch synthesis-related genes by binding to ACTCAT elements in their promoters. J. Exp. Bot. 2016, 67, 1327–1338. [Google Scholar] [CrossRef] [PubMed]
- Ma, M.; Wang, Q.; Li, Z.; Cheng, H.; Li, Z.; Liu, X.; Song, W.; Appels, R.; Zhao, H. Expression of TaCYP78A3, a gene encoding cytochrome P450CYP78A3 protein in wheat (Triticum aestivum L.), affects seed size. Plant J. 2015, 83, 312–325. [Google Scholar] [CrossRef] [PubMed]
- Pacak, A.; Geisler, K.; Jørgensen, B.; Barciszewska-Pacak, M.; Nilsson, L.; Nielsen, T.H.; Johansen, E.; Grønlund, M.; Jakobsen, I.; Albrechtsen, M. Investigations of barley stripe mosaic virus as a gene silencing vector in barley roots and in Brachypodium distachyon and oat. Plant Methods 2010, 6, 26. [Google Scholar] [CrossRef] [PubMed]
- Ma, H.Z.; Liu, G.Q.; Li, C.W.; Kang, G.Z.; Guo, T.C. Identification of the TaBTF3 gene in wheat (Triticum aestivum L.) and the effect of its silencing on wheat chloroplast, mitochondria and mesophyll cell development. Biochem. Biophys. Res. Commun. 2012, 426, 608–614. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Dai, T.; Jiang, D.; Cao, W. Effects of high temperature on key enzymes involved in starch and protein formation in grains of two wheat cultivars. J. Agron. Crop. Sci. 2008, 194, 47–54. [Google Scholar] [CrossRef]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, G.; Wu, Y.; Xu, M.; Gao, T.; Wang, P.; Wang, L.; Guo, T.; Kang, G. Virus-Induced Gene Silencing Identifies an Important Role of the TaRSR1 Transcription Factor in Starch Synthesis in Bread Wheat. Int. J. Mol. Sci. 2016, 17, 1557. https://doi.org/10.3390/ijms17101557
Liu G, Wu Y, Xu M, Gao T, Wang P, Wang L, Guo T, Kang G. Virus-Induced Gene Silencing Identifies an Important Role of the TaRSR1 Transcription Factor in Starch Synthesis in Bread Wheat. International Journal of Molecular Sciences. 2016; 17(10):1557. https://doi.org/10.3390/ijms17101557
Chicago/Turabian StyleLiu, Guoyu, Yufang Wu, Mengjun Xu, Tian Gao, Pengfei Wang, Lina Wang, Tiancai Guo, and Guozhang Kang. 2016. "Virus-Induced Gene Silencing Identifies an Important Role of the TaRSR1 Transcription Factor in Starch Synthesis in Bread Wheat" International Journal of Molecular Sciences 17, no. 10: 1557. https://doi.org/10.3390/ijms17101557
APA StyleLiu, G., Wu, Y., Xu, M., Gao, T., Wang, P., Wang, L., Guo, T., & Kang, G. (2016). Virus-Induced Gene Silencing Identifies an Important Role of the TaRSR1 Transcription Factor in Starch Synthesis in Bread Wheat. International Journal of Molecular Sciences, 17(10), 1557. https://doi.org/10.3390/ijms17101557




